Darunavir-Resistant HIV-1 Protease Constructs Uphold a Conformational Selection Hypothesis for Drug Resistance
Abstract
1. Introduction
2. Materials and Methods
2.1. Cloning and Mutagenesis
2.2. Protein Expression, Purification, and Spin-Labeling
2.3. Sample Preparation, DEER Data Collection, and Analysis
3. Results
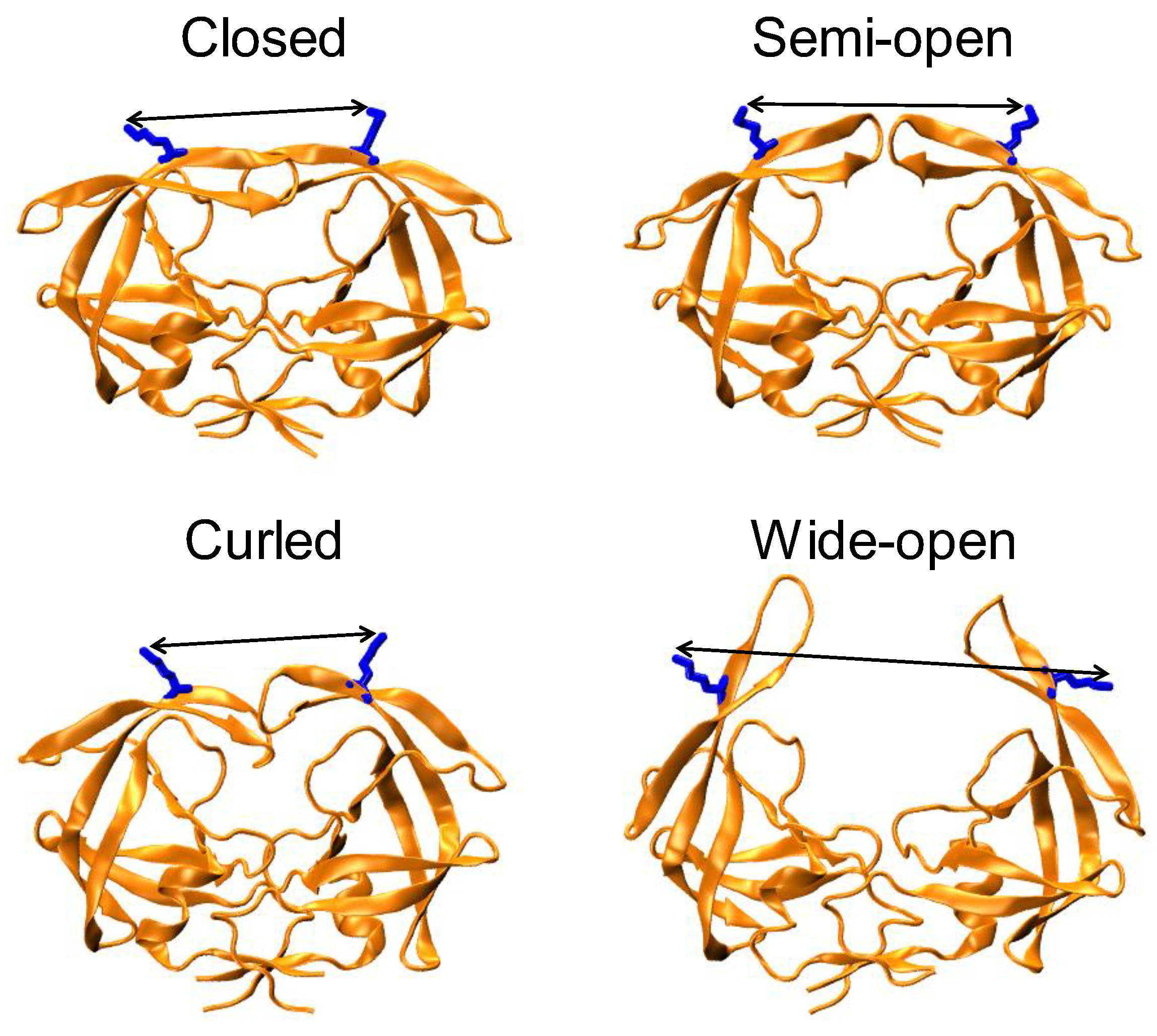
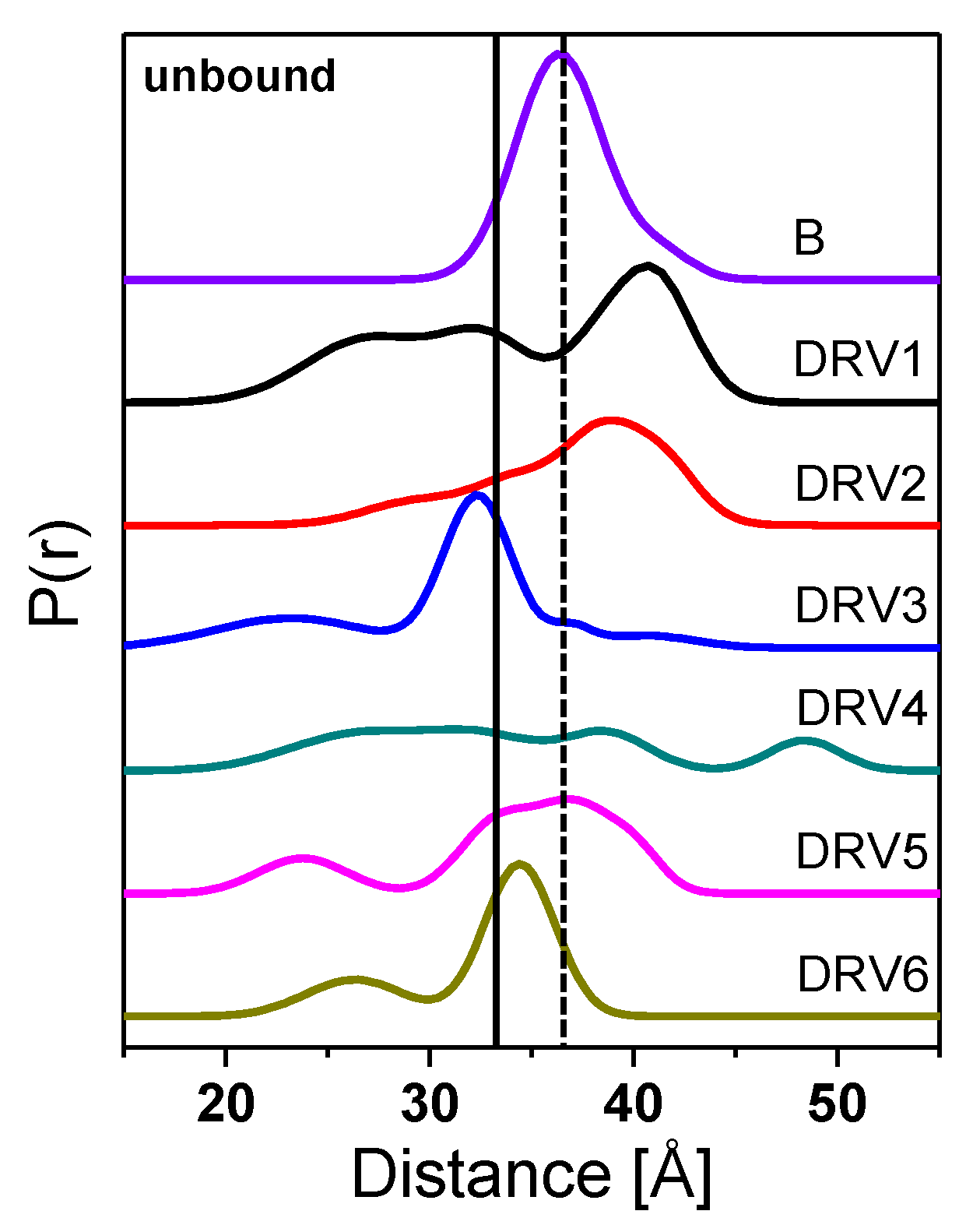
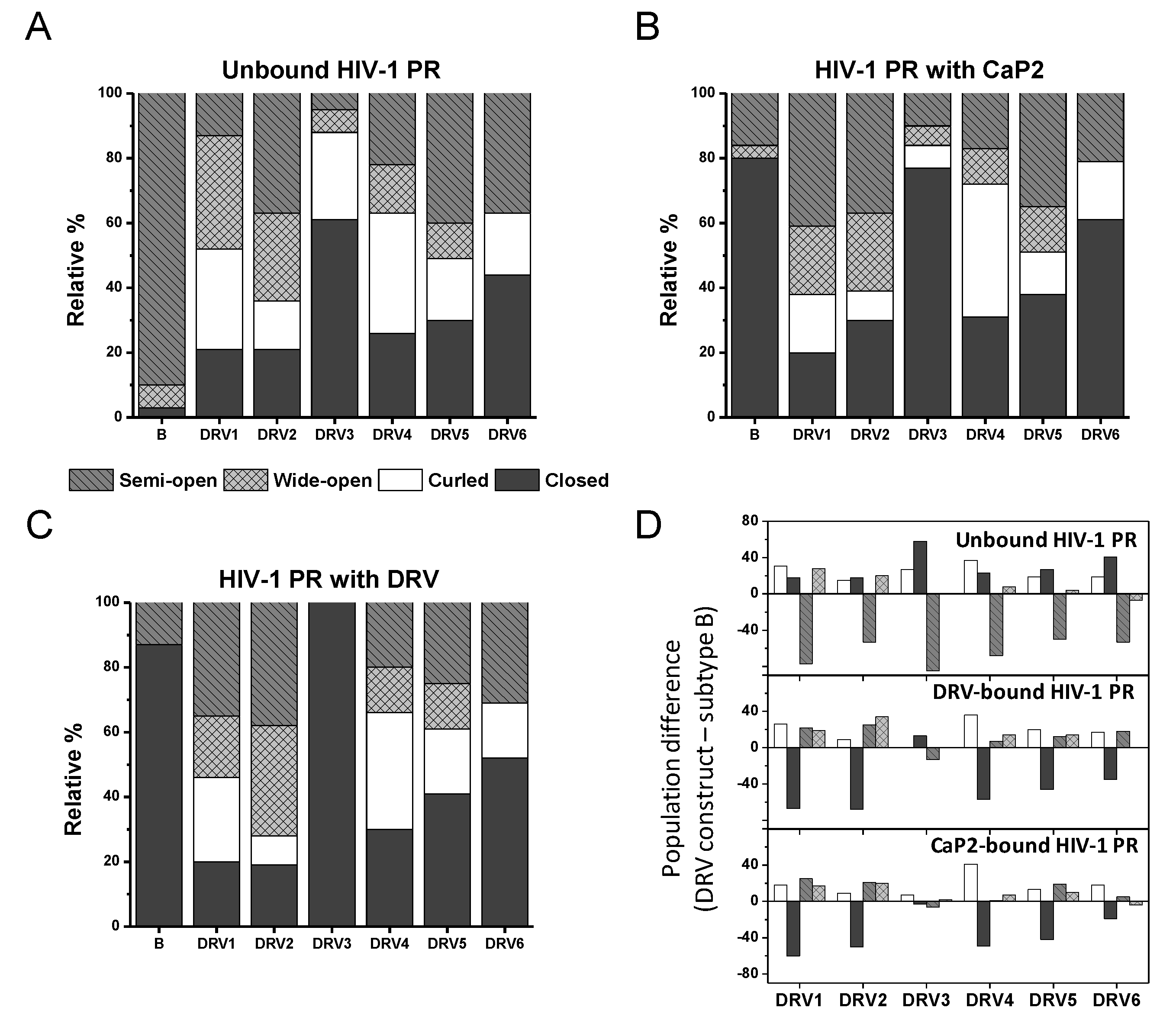
3.1. DRV-Resistant Constructs Sample High Fractional Occupancy of Open-Like and Closed State Compared to PI-Naïve Subtype B
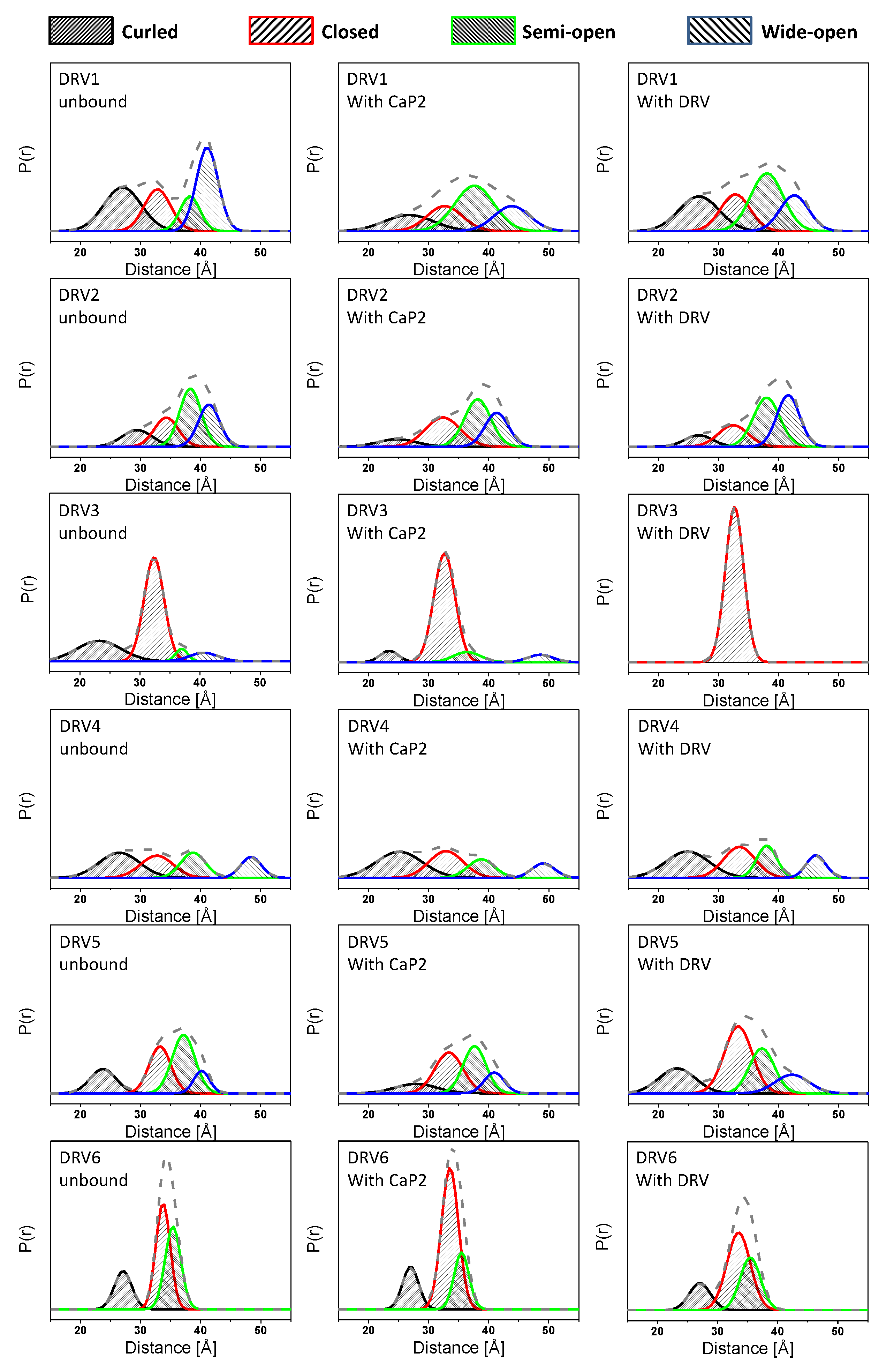
3.2. The Conformational Landscape of Most DRV-Resistant Constructs Is Not Altered by the Addition of DRV or Substrate Mimic
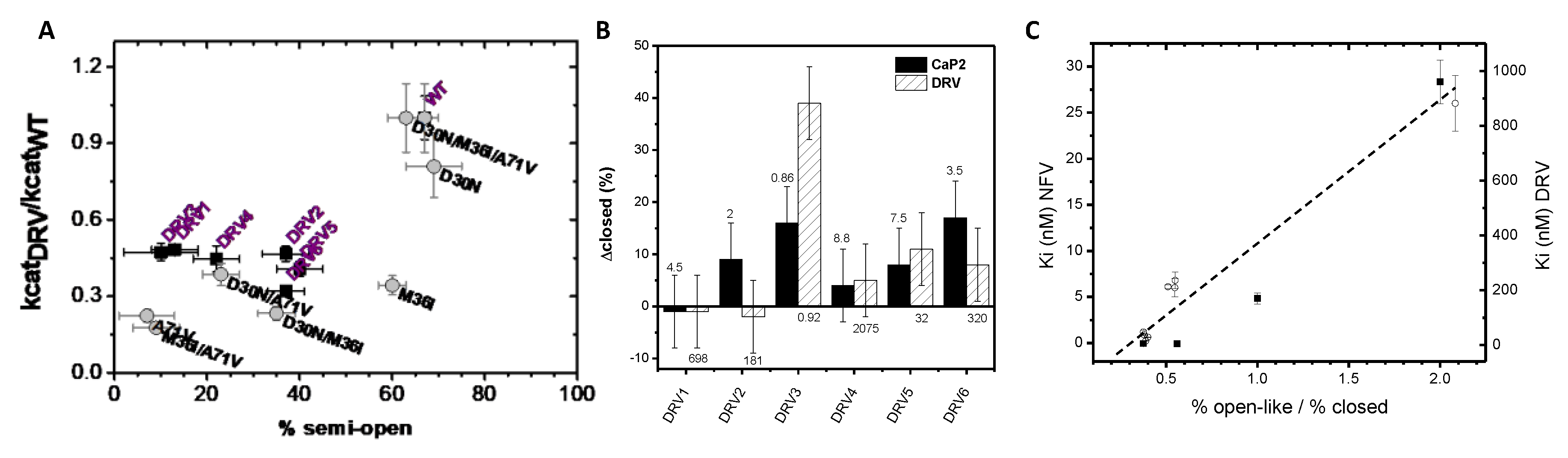
3.3. Conformational Landscape Hypothesis for Catalytic Turnover Is Upheld
4. Discussion
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Kramer, R.A.; Schaber, M.D.; Skalka, A.M.; Ganguly, K.; Wong-Staal, F.; Reddy, E.P. HTLV-III gag protein is processed in yeast cells by the virus pol-protease. Science 1986, 231, 1580–1584. [Google Scholar] [CrossRef] [PubMed]
- Weber, I.T.; Miller, M.; Jaskolski, M.; Leis, J.; Skalka, A.M.; Wlodawer, A. Molecular Modeling of the Hiv-1 Protease and Its Substrate Binding-Site. Science 1989, 243, 928–931. [Google Scholar] [CrossRef] [PubMed]
- Wlodawer, A.; Erickson, J.W. Structure-Based Inhibitors of Hiv-1 Protease. Annu. Rev. Biochem. 1993, 62, 543–585. [Google Scholar] [CrossRef] [PubMed]
- Yin, P.D.; Das, D.; Mitsuya, H. Overcoming HIV drug resistance through rational drug design based on molecular, biochemical, and structural profiles of HIV resistance. Cell Mol. Life Sci. 2006, 63, 1706–1724. [Google Scholar] [CrossRef]
- Lefebvre, E.; Schiffer, C.A. Resilience to resistance of HIV-1 protease inhibitors: Profile of darunavir. Aids Rev. 2008, 10, 131–142. [Google Scholar] [PubMed]
- Clutter, D.S.; Jordan, M.R.; Bertagnolio, S.; Shafer, R.W. HIV-1 drug resistance and resistance testing. Infect. Genet. Evol. 2016, 46, 292–307. [Google Scholar] [CrossRef]
- Wensing, A.M.; Calvez, V.; Gunthard, H.F.; Johnson, V.A.; Paredes, R.; Pillay, D.; Shafer, R.W.; Richman, D.D. 2017 Update of the Drug Resistance Mutations in HIV-1. Top. Antivir. Med. 2016, 24, 132–133. [Google Scholar]
- Feder, A.F.; Rhee, S.Y.; Holmes, S.P.; Shafer, R.W.; Petrov, D.A.; Pennings, P.S. More effective drugs lead to harder selective sweeps in the evolution of drug resistance in HIV-1. Elife 2016, 5. [Google Scholar] [CrossRef]
- Jeschke, G.; Chechik, V.; Ionita, P.; Godt, A.; Zimmermann, H.; Banham, J.; Timmel, C.R.; Hilger, D.; Jung, H. DeerAnalysis2006—A Comprehensive Software Package for Analyzing Pulsed ELDOR Data. Appl. Mag. Reson. 2006, 30, 473–498. [Google Scholar] [CrossRef]
- Jeschke, G.; Polyhach, Y. Distance measurements on spin-labelled biomacromolecules by pulsed electron paramagnetic resonance. Phys. Chem. Chem. Phys. 2007, 9, 1895–1910. [Google Scholar] [CrossRef]
- Chiang, Y.W.; Borbat, P.P.; Freed, J.H. The determination of pair distance distributions by pulsed ESR using Tikhonov regularization. J. Magn. Reson. 2005, 172, 279–295. [Google Scholar] [CrossRef]
- Carter, J.D.; Mathias, J.D.; Gomez, E.F.; Ran, Y.; Xu, F.; Galiano, L.; Tran, N.Q.; D’Amore, P.W.; Wright, C.S.; Chakravorty, D.K.; et al. Characterizing Solution Surface Loop Conformational Flexibility of the GM2 Activator Protein. J. Phys. Chem. B 2014, 118118, 10607–10617. [Google Scholar] [CrossRef]
- de Vera, I.M.; Blackburn, M.E.; Fanucci, G.E. Correlating conformational shift induction with altered inhibitor potency in a multidrug resistant HIV-1 protease variant. Biochemistry 2012, 5151, 7813–7815. [Google Scholar] [CrossRef][Green Version]
- De Vera, I.M.; Smith, A.N.; Dancel, M.C.; Huang, X.; Dunn, B.M.; Fanucci, G.E. Elucidating a relationship between conformational sampling and drug resistance in HIV-1 protease. Biochemistry 2013, 52, 3278–3288. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Galiano, L.; Ding, F.; Veloro, A.M.; Blackburn, M.E.; Simmerling, C.; Fanucci, G.E. Drug pressure selected mutations in HIV-1 protease alter flap conformations. J. Am. Chem. Soc. 2009, 131, 430–431. [Google Scholar] [CrossRef]
- Huang, X.; Britto, M.D.; Kear-Scott, J.L.; Boone, C.D.; Rocca, J.R.; Simmerling, C.; McKenna, R.; Bieri, M.; Gooley, P.R.; Dunn, B.M.; et al. The role of select subtype polymorphisms on HIV-1 protease conformational sampling and dynamics. J. Biol. Chem. 2014, 289, 17203–17214. [Google Scholar] [CrossRef]
- Huang, X.; de Vera, I.M.; Veloro, A.M.; Blackburn, M.E.; Kear, J.L.; Carter, J.D.; Rocca, J.R.; Simmerling, C.; Dunn, B.M.; Fanucci, G.E. Inhibitor-induced conformational shifts and ligand-exchange dynamics for HIV-1 protease measured by pulsed EPR and NMR spectroscopy. J. Phys. Chem. B 2012, 116, 14235–14244. [Google Scholar] [CrossRef]
- Kear, J.L.; Blackburn, M.E.; Veloro, A.M.; Dunn, B.M.; Fanucci, G.E. Subtype polymorphisms among HIV-1 protease variants confer altered flap conformations and flexibility. J. Am. Chem. Soc. 2009, 131, 14650–14651. [Google Scholar] [CrossRef]
- Liu, Z.; Casey, T.M.; Blackburn, M.E.; Huang, X.; Pham, L.; de Vera, I.M.; Carter, J.D.; Kear-Scott, J.L.; Veloro, A.M.; Galiano, L.; et al. Pulsed EPR characterization of HIV-1 protease conformational sampling and inhibitor-induced population shifts. Phys. Chem. Chem. Phys. 2016, 18, 5819–5831. [Google Scholar] [CrossRef]
- Blackburn, M.E.; Veloro, A.M.; Fanucci, G.E. Monitoring inhibitor-induced conformational population shifts in HIV-1 protease by pulsed EPR spectroscopy. Biochemistry 2009, 48, 8765–8767. [Google Scholar] [CrossRef]
- Wong-Sam, A.; Wang, Y.F.; Zhang, Y.; Ghosh, A.K.; Harrison, R.W.; Weber, I.T. Drug Resistance Mutation L76V Alters Nonpolar Interactions at the Flap-Core Interface of HIV-1 Protease. ACS Omega 2018, 3, 12132–12140. [Google Scholar] [CrossRef]
- Louis, J.M.; Zhang, Y.; Sayer, J.M.; Wang, Y.F.; Harrison, R.W.; Weber, I.T. The L76V drug resistance mutation decreases the dimer stability and rate of autoprocessing of HIV-1 protease by reducing internal hydrophobic contacts. Biochemistry 2011, 50, 4786–4795. [Google Scholar] [CrossRef]
- Goldfarb, N.E.; Ohanessian, M.; Biswas, S.; McGee, T.D., Jr.; Mahon, B.P.; Ostrov, D.A.; Garcia, J.; Tang, Y.; McKenna, R.; Roitberg, A.; et al. Defective hydrophobic sliding mechanism and active site expansion in HIV-1 protease drug resistant variant Gly48Thr/Leu89Met: Mechanisms for the loss of saquinavir binding potency. Biochemistry 2015, 54, 422–433. [Google Scholar] [CrossRef]
- Ragland, D.A.; Nalivaika, E.A.; Nalam, M.N.; Prachanronarong, K.L.; Cao, H.; Bandaranayake, R.M.; Cai, Y.; Kurt-Yilmaz, N.; Schiffer, C.A. Drug resistance conferred by mutations outside the active site through alterations in the dynamic and structural ensemble of HIV-1 protease. J. Am. Chem. Soc. 2015, 136, 11956–11963. [Google Scholar] [CrossRef]
- Cai, Y.; Myint, W.; Paulsen, J.L.; Schiffer, C.A.; Ishima, R.; Kurt Yilmaz, N. Drug Resistance Mutations Alter Dynamics of Inhibitor-Bound HIV-1 Protease. J. Chem. Theory Comput. 2014, 10, 3438–3448. [Google Scholar] [CrossRef] [PubMed]
- Foulkes-Murzycki, J.E.; Scott, W.R.; Schiffer, C.A. Hydrophobic sliding: A possible mechanism for drug resistance in human immunodeficiency virus type 1 protease. Structure 2007, 15, 225–233. [Google Scholar] [CrossRef]
- Agniswamy, J.; Louis, J.M.; Roche, J.; Harrison, R.W.; Weber, I.T. Structural Studies of a Rationally Selected Multi-Drug Resistant HIV-1 Protease Reveal Synergistic Effect of Distal Mutations on Flap Dynamics. PLoS ONE 2016, 11, e0168616. [Google Scholar] [CrossRef]
- Henes, M.; Lockbaum, G.J.; Kosovrasti, K.; Leidner, F.; Nachum, G.S.; Nalivaika, E.A.; Lee, S.K.; Spielvogel, E.; Zhou, S.; Swanstrom, R.; et al. Picomolar to Micromolar: Elucidating the Role of Distal Mutations in HIV-1 Protease in Conferring Drug Resistance. ACS Chem. Biol. 2019, 14, 2441–2452. [Google Scholar] [CrossRef]
- Laco, G.S. HIV-1 protease substrate-groove: Role in substrate recognition and inhibitor resistance. Biochimie 2015, 118, 90–103. [Google Scholar] [CrossRef]
- Clemente, J.C.; Hemrajani, R.; Blum, L.E.; Goodenow, M.M.; Dunn, B.M. Secondary mutations M36I and A71V in the human immunodeficiency virus type 1 protease can provide an advantage for the emergence of the primary mutation D30N. Biochemistry 2003, 42, 15029–15035. [Google Scholar] [CrossRef]
- Saskova, K.G.; Kozisek, M.; Rezacova, P.; Brynda, J.; Yashina, T.; Kagan, R.M.; Konvalinka, J. Molecular characterization of clinical isolates of human immunodeficiency virus resistant to the protease inhibitor darunavir. J. Virol. 2009, 83, 8810–8818. [Google Scholar] [CrossRef]
- Altman, M.D.; Ali, A.; Reddy, G.S.; Nalam, M.N.; Anjum, S.G.; Cao, H.; Chellappan, S.; Kairys, V.; Fernandes, M.X.; Gilson, M.K.; et al. HIV-1 protease inhibitors from inverse design in the substrate envelope exhibit subnanomolar binding to drug-resistant variants. J. Am. Chem. Soc. 2008, 130, 6099–6113. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Chang, Y.C.; Louis, J.M.; Wang, Y.F.; Harrison, R.W.; Weber, I.T. Structures of darunavir-resistant HIV-1 protease mutant reveal atypical binding of darunavir to wide open flaps. ACS Chem. Biol. 2014, 9, 1351–1358. [Google Scholar] [CrossRef]
- Weber, I.T.; Kneller, D.W.; Wong-Sam, A. Highly resistant HIV-1 proteases and strategies for their inhibition. Future Med. Chem. 2015, 7, 1023–1038. [Google Scholar] [CrossRef] [PubMed]
- Kurt Yilmaz, N.; Swanstrom, R.; Schiffer, C.A. Improving Viral Protease Inhibitors to Counter Drug Resistance. Trends Microbiol. 2016, 24, 547–557. [Google Scholar] [CrossRef]
- Kneller, D.W.; Agniswamy, J.; Ghosh, A.K.; Weber, I.T. Potent antiviral HIV-1 protease inhibitor combats highly drug resistant mutant PR20. Biochem. Biophys. Res. Commun. 2019, 519, 61–66. [Google Scholar] [CrossRef]
- Agniswamy, J.; Kneller, D.W.; Brothers, R.; Wang, Y.F.; Harrison, R.W.; Weber, I.T. Highly Drug-Resistant HIV-1 Protease Mutant PRS17 Shows Enhanced Binding to Substrate Analogues. ACS Omega 2019, 4, 8707–8719. [Google Scholar] [CrossRef]
- Pawar, S.D.; Freas, C.; Weber, I.T.; Harrison, R.W. Analysis of drug resistance in HIV protease. BMC Bioinformatics 2018, 19 (Suppl. 11), 362. [Google Scholar] [CrossRef]
- Humpolickova, J.; Weber, J.; Starkova, J.; Masinova, E.; Gunterova, J.; Flaisigova, I.; Konvalinka, J.; Majerova, T. Inhibition of the precursor and mature forms of HIV-1 protease as a tool for drug evaluation. Sci. Rep. 2018, 8, 10438. [Google Scholar] [CrossRef]
- Lockbaum, G.J.; Leidner, F.; Rusere, L.N.; Henes, M.; Kosovrasti, K.; Nachum, G.S.; Nalivaika, E.A.; Ali, A.; Yilmaz, N.K.; Schiffer, C.A. Structural Adaptation of Darunavir Analogues against Primary Mutations in HIV-1 Protease. ACS Infect. Dis. 2019, 5, 316–325. [Google Scholar] [CrossRef]
- Matthew, A.N.; Leidner, F.; Newton, A.; Petropoulos, C.J.; Huang, W.; Ali, A.; KurtYilmaz, N.; Schiffer, C.A. Molecular Mechanism of Resistance in a Clinically Significant Double-Mutant Variant of HCV NS3/4A Protease. Structure 2018, 26, 1360–1372.e5. [Google Scholar] [CrossRef]
- Khan, S.N.; Persons, J.D.; Paulsen, J.L.; Guerrero, M.; Schiffer, C.A.; Kurt-Yilmaz, N.; Ishima, R. Probing Structural Changes among Analogous Inhibitor-Bound Forms of HIV-1 Protease and a Drug-Resistant Mutant in Solution by Nuclear Magnetic Resonance. Biochemistry 2018, 57, 1652–1662. [Google Scholar] [CrossRef]
- Agniswamy, J.; Shen, C.H.; Wang, Y.F.; Ghosh, A.K.; Rao, K.V.; Xu, C.X.; Sayer, J.M.; Louis, J.M.; Weber, I.T. Extreme multidrug resistant HIV-1 protease with 20 mutations is resistant to novel protease inhibitors with P1′-pyrrolidinone or P2-tris-tetrahydrofuran. J. Med. Chem. 2013, 56, 4017–4027. [Google Scholar] [CrossRef]
- Agniswamy, J.; Shen, C.H.; Aniana, A.; Sayer, J.M.; Louis, J.M.; Weber, I.T. HIV-1 protease with 20 mutations exhibits extreme resistance to clinical inhibitors through coordinated structural rearrangements. Biochemistry 2012, 51, 2819–2828. [Google Scholar] [CrossRef][Green Version]
- Kozisek, M.; Lepsik, M.; Grantz Saskova, K.; Brynda, J.; Konvalinka, J.; Rezacova, P. Thermodynamic and structural analysis of HIV protease resistance to darunavir—analysis of heavily mutated patient-derived HIV-1 proteases. FEBS J. 2014, 281, 1834–1847. [Google Scholar] [CrossRef]
- Mildner, A.M.; Rothrock, D.J.; Leone, J.W.; Bannow, C.A.; Lull, J.M.; Reardon, I.M.; Sarcich, J.L.; Howe, W.J.; Tomich, C.S.; Smith, C.W. The HIV-1 protease as enzyme and substrate: Mutagenesis of autolysis sites and generation of a stable mutant with retained kinetic properties. Biochemistry 1994, 33, 9405–9413. [Google Scholar] [CrossRef]
- Galiano, L.; Bonora, M.; Fanucci, G.E. Interflap distances in HIV-1 protease determined by pulsed EPR measurements. J. Am. Chem. Soc. 2007, 129, 11004–11005. [Google Scholar] [CrossRef]
- Shao, W.; Everitt, L.; Manchester, M.; Loeb, D.D.; Hutchison, C.A., 3rd; Swanstrom, R. Sequence requirements of the HIV-1 protease flap region determined by saturation mutagenesis and kinetic analysis of flap mutants. Proc. Natl. Acad. Sci. USA 1997, 94, 2243–2248. [Google Scholar] [CrossRef]
- Galiano, L.; Blackburn, M.E.; Veloro, A.M.; Bonora, M.; Fanucci, G.E. Solute effects on spin labels at an aqueous-exposed site in the flap region of HIV-1 protease. J. Phys. Chem. B 2009, 113, 1673–1680. [Google Scholar] [CrossRef]
- Carter, J.D.; Gonzales, E.G.; Huang, X.; Smith, A.N.; de Vera, I.M.; D’Amore, P.W.; Rocca, J.R.; Goodenow, M.M.; Dunn, B.M.; Fanucci, G.E. Effects of PRE and POST therapy drug-pressure selected mutations on HIV-1 protease conformational sampling. FEBS Lett. 2014, 588, 3123–3128. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Huang, X.; Hu, L.; Pham, L.; Poole, K.M.; Tang, Y.; Mahon, B.P.; Tang, W.; Li, K.; Goldfarb, N.E.; et al. Effects of Hinge-region Natural Polymorphisms on Human Immunodeficiency Virus-Type 1 Protease Structure, Dynamics, and Drug Pressure Evolution. J. Biol. Chem. 2016, 291, 22741–22756. [Google Scholar] [CrossRef] [PubMed]
- Ding, F.; Layten, M.; Simmerling, C. Solution structure of HIV-1 protease flaps probed by comparison of molecular dynamics simulation ensembles and EPR experiments. J. Am. Chem. Soc. 2008, 130, 7184–7185. [Google Scholar] [CrossRef][Green Version]
- Louis, J.M.; Clore, G.M.; Gronenborn, A.M. Autoprocessing of HIV-1 protease is tightly coupled to protein folding. Nat. Struct. Biol. 1999, 6, 868–875. [Google Scholar] [CrossRef] [PubMed]
- Kear, J.L.; Galiano, L.; Veloro, A.M.; Harris, J.; Busenlehner, L.S.; Fanucci, G.E. Monitoring the autoproteolysis of HIV-1 Protease by Site-Directed Spin-Labeling and Electron Paramagnetic Resonance Spectroscopy. J. Biophys. Chem. 2011, 2, 137–146. [Google Scholar] [CrossRef]
- Casey, T.M.; Fanucci, G.E. Spin labeling and Double Electron-Electron Resonance (DEER) to Deconstruct Conformational Ensembles of HIV Protease. In Methods in Enzymology; Academic Press: Cambridge, MA, USA, 2015; Volume 564, pp. 153–187. [Google Scholar] [CrossRef]
- De Vera, I.M.; Blackburn, M.E.; Galiano, L.; Fanucci, E. Pulsed EPR distance measurements in soluble proteins by site-directed spin labeling (SDSL). Curr. Prot. Protein Sci. 2013, 74, 17. [Google Scholar] [CrossRef] [PubMed]
- Tran, T.T.; Liu, Z.; Fanucci, G.E. Conformational landscape of non-B variants of HIV-1 protease: A pulsed EPR study. Biochem. Biophys. Res. Commun. 2020, 532, 219–224. [Google Scholar] [CrossRef]
- Sayer, J.M.; Liu, F.; Ishima, R.; Weber, I.T.; Louis, J.M. Effect of the active site D25N mutation on the structure, stability, and ligand binding of the mature HIV-1 protease. J. Biol. Chem. 2008, 283, 13459–13470. [Google Scholar] [CrossRef]
| Unbound | CaP2 | DRV | ||||
|---|---|---|---|---|---|---|
| HIV-1 Constructs | Most Probable Distance (Å) (Error ± 0.2) | Average Distance (Å) (Error ± 0.2) | Most Probable Distance (Å) (Error ± 0.2) | Average Distance (Å) (Error ± 0.2) | Most Probable Distance (Å) (Error ± 0.2) | Average Distance (Å) (Error ± 0.2) |
| DRV1 | 40.7 | 34.6 | 36.8 | 35.9 | 38.7 | 34.9 |
| DRV2 | 39.1 | 36.9 | 39.1 | 36.0 | 40.5 | 37.1 |
| DRV3 | 32.2 | 31.6 | 32.6 | 33.3 | 32.6 | 32.7 |
| DRV4 | 30.6 | 34.2 | 32.3 | 32.5 | 37.4 | 33.0 |
| DRV5 | 36.9 | 33.8 | 37.3 | 35.2 | 34.5 | 33.5 |
| DRV6 | 34.4 | 32.4 | 34.0 | 32.3 | 34.5 | 33.0 |
| PI-naïve B 1 | 36.2 | 36.2 | 33.1 | 33.9 | 33.2 | 33.6 |
| Constructs | States | Relative Populations (±5%) | |||
|---|---|---|---|---|---|
| Curled/Tucked | Closed | Semi-Open | Wide-Open | ||
| DRV1 | unbound | 31 | 21 | 13 | 35 |
| CaP2 | 18 | 20 | 41 | 21 | |
| DRV | 26 | 20 | 35 | 19 | |
| DRV2 | unbound | 15 | 21 | 37 | 27 |
| CaP2 | 9 | 30 | 37 | 24 | |
| DRV | 9 | 19 | 38 | 34 | |
| DRV3 | unbound | 27 | 61 | 5 | 7 |
| CaP2 | 7 | 77 | 10 | 6 | |
| DRV | 0 | 100 | 0 | 0 | |
| DRV4 | unbound | 37 | 26 | 22 | 15 |
| CaP2 | 41 | 31 | 17 | 11 | |
| DRV | 36 | 30 | 20 | 14 | |
| DRV5 | unbound | 19 | 30 | 40 | 11 |
| CaP2 | 13 | 38 | 35 | 14 | |
| DRV | 20 | 41 | 25 | 14 | |
| DRV6 | unbound | 19 | 44 | 37 | 0 |
| CaP2 | 18 | 61 | 21 | 0 | |
| DRV | 17 | 52 | 31 | 0 | |
| B 1 | Unbound | 0 | 3 | 90 | 7 |
| CaP2 | 0 | 80 | 16 | 4 | |
| DRV | 0 | 87 | 13 | 0 | |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liu, Z.; Tran, T.T.; Pham, L.; Hu, L.; Bentz, K.; Savin, D.A.; Fanucci, G.E. Darunavir-Resistant HIV-1 Protease Constructs Uphold a Conformational Selection Hypothesis for Drug Resistance. Viruses 2020, 12, 1275. https://doi.org/10.3390/v12111275
Liu Z, Tran TT, Pham L, Hu L, Bentz K, Savin DA, Fanucci GE. Darunavir-Resistant HIV-1 Protease Constructs Uphold a Conformational Selection Hypothesis for Drug Resistance. Viruses. 2020; 12(11):1275. https://doi.org/10.3390/v12111275
Chicago/Turabian StyleLiu, Zhanglong, Trang T. Tran, Linh Pham, Lingna Hu, Kyle Bentz, Daniel A. Savin, and Gail E. Fanucci. 2020. "Darunavir-Resistant HIV-1 Protease Constructs Uphold a Conformational Selection Hypothesis for Drug Resistance" Viruses 12, no. 11: 1275. https://doi.org/10.3390/v12111275
APA StyleLiu, Z., Tran, T. T., Pham, L., Hu, L., Bentz, K., Savin, D. A., & Fanucci, G. E. (2020). Darunavir-Resistant HIV-1 Protease Constructs Uphold a Conformational Selection Hypothesis for Drug Resistance. Viruses, 12(11), 1275. https://doi.org/10.3390/v12111275





