In Silico Identification of Novel Aromatic Compounds as Potential HIV-1 Entry Inhibitors Mimicking Cellular Receptor CD4
Abstract
1. Introduction
2. Materials and Methods
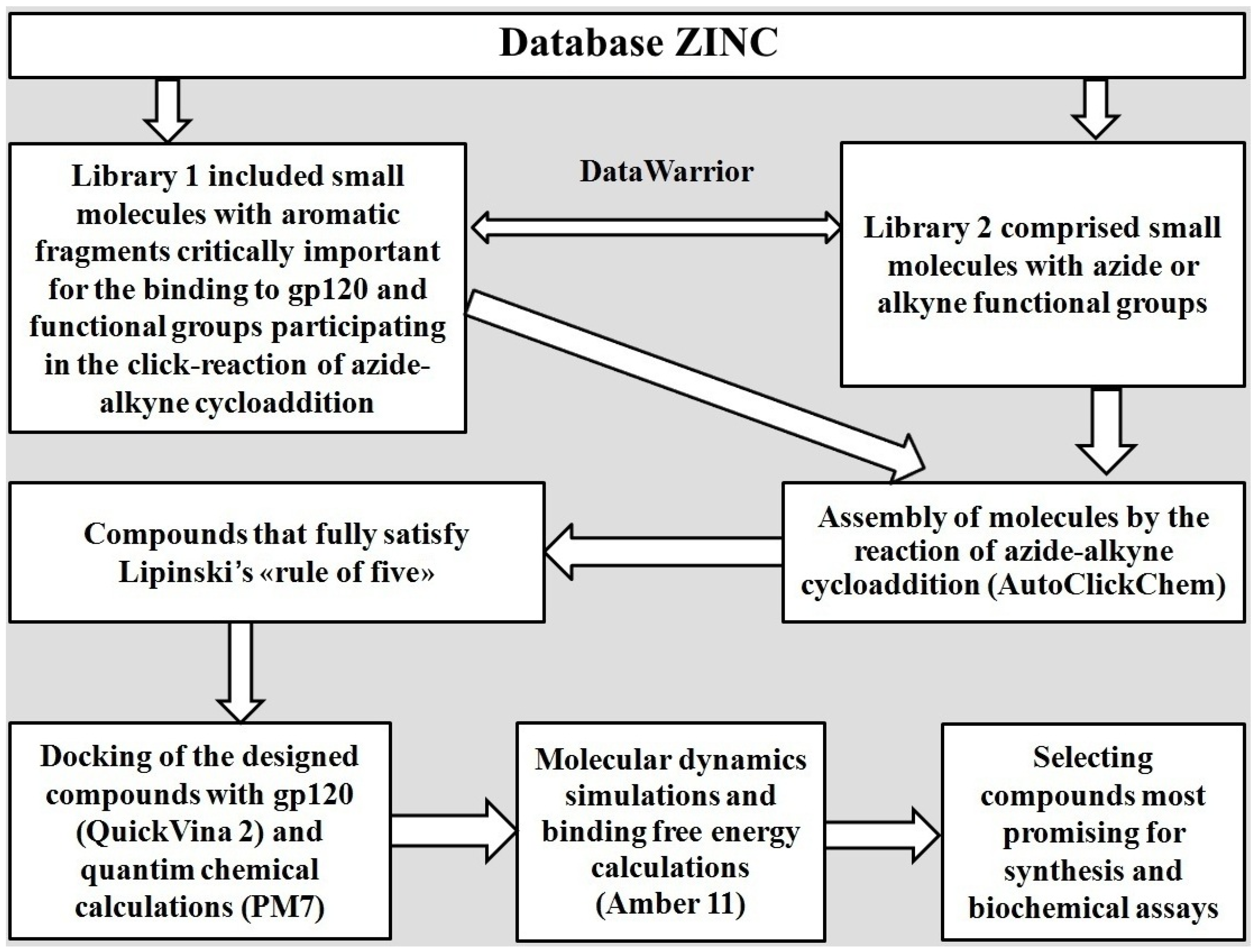
2.1. In Silico Design of Small-Molecule CD4-Mimetic Candidates
2.2. Molecular Docking
2.3. Quantum Chemical Studies
2.4. Analysis of the PM7-Based Ligand/gp120 Complexes
2.5. Molecular Dynamics Simulations
2.6. Binding Free Energy Calculations
3. Results and Discussion
4. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Sharp, P.M.; Hahn, B.H. Origins of HIV and the AIDS pandemic. Cold Spring Harb. Perspect. Med. 2011, 1, a006841. [Google Scholar] [CrossRef] [PubMed]
- UNAIDS|AIDSinfo. Available online: http://aidsinfo.unaids.org/ (accessed on 27 June 2019).
- U.S. Food and Drug Administration. Available online: https://www.fda.gov/ (accessed on 27 June 2019).
- de Clercq, E. New approaches toward anti-HIV chemotherapy. J. Med. Chem. 2005, 48, 1297–1313. [Google Scholar] [CrossRef] [PubMed]
- Este, J.A.; Telenti, A. HIV entry inhibitors. Lancet 2007, 370, 81–88. [Google Scholar] [CrossRef]
- Rusconi, S.; Scozzafava, A.; Mastrolorenzo, A.; Supuran, C.T. An update in the development of HIV entry inhibitors. Curr. Top. Med. Chem. 2007, 7, 1273–1289. [Google Scholar] [CrossRef] [PubMed]
- Ryser, H.J.-P.; Fluckiger, R. Progress in targeting HIV-1 entry. Drug Discov. Today 2005, 10, 1085–1094. [Google Scholar] [CrossRef]
- Adamson, C.S.; Freed, E.O. Novel approaches to inhibiting HIV-1 replication. Antiviral Res. 2010, 85, 119–141. [Google Scholar] [CrossRef] [PubMed]
- Santos, A.F.; Soares, M.A. HIV genetic diversity and drug resistance. Viruses 2010, 2, 503–531. [Google Scholar] [CrossRef] [PubMed]
- Abram, M.E.; Ferris, A.L.; Das, K.; Quinoñes, O.; Shao, W.; Tuske, S.; Alvord, G.; Arnold, E.; Hughes, S.H. Mutations in HIV-1 reverse transcriptase affect the errors made in a single cycle of viral replication. J. Virol. 2014, 88, 7589–7601. [Google Scholar] [CrossRef]
- Arts, E.J.; Hazuda, D.J. HIV-1 antiretroviral drug therapy. Cold Spring Harb. Perspect. Med. 2012, 2, a007161. [Google Scholar] [CrossRef]
- Kumari, G.; Singh, R.K. Highly active antiretroviral therapy for treatment of HIV/AIDS patients: Current status and future prospects and the Indian scenario. HIV Aids Rev. 2012, 11, 5–14. [Google Scholar] [CrossRef]
- Li, W.; Lu, L.; Li, W.; Jiang, S. Small-molecule HIV-1 entry inhibitors targeting gp120 and gp41: A patent review (2010–2015). Expert Opin. Ther. Pat. 2017, 27, 707–719. [Google Scholar] [CrossRef]
- Su, S.; Wang, Q.; Xu, W.; Yu, F.; Hua, C.; Zhu, Y.; Jiang, S.; Lu, L. A novel HIV-1 gp41 tripartite model for rational design of HIV-1 fusion inhibitors with improved antiviral activity. AIDS 2017, 31, 885–894. [Google Scholar] [CrossRef]
- Wilen, C.B.; Tilton, J.C.; Doms, R.W. HIV: Cell binding and entry. Cold Spring Harb. Perspect. Med. 2012, 2, a006866. [Google Scholar] [CrossRef]
- Kwong, P.D.; Wyatt, R.; Robinson, J.; Sweet, R.W.; Sodroski, J.; Hendrickson, W.A. Structure of an HIV gp120 envelope glycoprotein in complex with the CD4 receptor and a neutralizing human antibody. Nature 1998, 393, 648–659. [Google Scholar] [CrossRef]
- Matthews, T.; Salgo, M.; Greenberg, M.; Chung, J.; DeMasi, R.; Bolognesi, D. Enfuvirtide: The first therapy to inhibit the entry of HIV-1 into host CD4 lymphocytes. Nat. Rev. Drug Discov. 2004, 3, 215–225. [Google Scholar] [CrossRef]
- MacArthur, R.D.; Novak, R.M. Reviews of anti-infective agents: Maraviroc: The first of a new class of antiretroviral agents. Clin. Infect. Dis. 2008, 47, 236–241. [Google Scholar] [CrossRef]
- Zhao, Q.; Ma, L.; Jiang, S.; Lu, H.; Liu, S.; He, Y.; Strick, N.; Neamati, N.; Debnath, A.K. Identification of N-phenyl-N’-(2,2,6,6-tetramethyl-piperidin-4-yl)-oxalamides as a new class of HIV-1 entry inhibitors that prevent gp120 binding to CD4. Virology 2005, 339, 213–225. [Google Scholar] [CrossRef]
- Schön, A.; Madani, N.; Klein, J.C.; Hubicki, A.; Ng, D.; Yang, X.; Smith, A.B., III; Sodroski, J.; Freire, E. Thermodynamics of binding of a low-molecular-weight CD4 mimetic to HIV-1 gp120. Biochemistry 2006, 45, 10973–10980. [Google Scholar] [CrossRef]
- Lalonde, J.M.; Kwon, Y.D.; Jones, D.M.; Sun, A.W.; Courter, J.R.; Soeta, T.; Kobayashi, T.; Princiotto, A.M.; Wu, X.; Schön, A.; et al. Structure-based design, synthesis, and characterization of dual hotspot small-molecule HIV-1 entry inhibitors. J. Med. Chem. 2012, 55, 4382–4396. [Google Scholar] [CrossRef]
- Lalonde, J.M.; Le-Khac, M.; Jones, D.M.; Courter, J.R.; Park, J.; Schön, A.; Princiotto, A.M.; Wu, X.; Mascola, J.R.; Freire, E.; et al. Structure-based design and synthesis of an HIV-1 entry inhibitor exploiting X-ray and thermodynamic characterization. ACS Med. Chem. Lett. 2013, 4, 338–343. [Google Scholar] [CrossRef]
- Madani, N.; Schön, A.; Princiotto, A.M.; Lalonde, J.M.; Courter, J.R.; Soeta, T.; Ng, D.; Wang, L.; Brower, E.T.; Xiang, S.H.; et al. Small-molecule CD4 mimics interact with a highly conserved pocket on HIV-1 gp120. Structure 2008, 16, 1689–1701. [Google Scholar] [CrossRef]
- Si, Z.; Madani, N.; Cox, J.M.; Chruma, J.J.; Klein, J.C.; Schön, A.; Phan, N.; Wang, L.; Biorn, A.C.; Cocklin, S.; et al. Small molecule inhibitors of HIV-1 entry block receptor-induced conformational changes in the viral envelope glycoproteins. Proc. Natl. Acad. Sci. USA 2004, 101, 5036–5041. [Google Scholar] [CrossRef]
- Curreli, F.; Kwon, Y.D.; Zhang, H.; Yang, Y.; Scacalossi, D.; Kwong, P.D.; Debnath, A.K. Binding mode characterization of NBD series CD4-mimetic HIV-1 entry inhibitors by X-ray structure and resistance study. Antimicrob. Agents Chemother. 2014, 58, 5478–5491. [Google Scholar] [CrossRef]
- Curreli, F.; Choudhury, S.; Pyatkin, I.; Zagorodnikov, V.P.; Bulay, A.K.; Altieri, A.; Kwon, Y.D.; Kwong, P.D.; Debnath, A.K. Design, synthesis and antiviral activity of entry inhibitors that target the CD4-binding site of HIV-1. J. Med. Chem. 2012, 55, 4764–4775. [Google Scholar] [CrossRef]
- Narumi, T.; Ochiai, C.; Yoshimura, K.; Harada, S.; Tanaka, T.; Nomura, W.; Arai, H.; Ozaki, T.; Ohashi, N.; Matsushita, S.; et al. CD4 mimics targeting the HIV entry mechanism and their hybrid molecules with a CXCR4 antagonist. Bioorg. Med. Chem. Lett. 2010, 20, 5853–5858. [Google Scholar] [CrossRef]
- Narumi, T.; Arai, H.; Yoshimura, K.; Harada, S.; Nomura, W.; Matsushita, S.; Tamamura, H. Small molecular CD4 mimics as HIV entry inhibitors. Bioorg. Med. Chem. 2011, 19, 6735–6742. [Google Scholar] [CrossRef]
- Narumi, T.; Arai, H.; Yoshimura, K.; Harada, S.; Hirota, Y.; Ohashi, N.; Hashimoto, C.; Nomura, W.; Matsushita, S.; Tamamura, H. CD4 mimics as HIV entry inhibitors: Lead optimization studies of the aromatic substituents. Bioorg. Med. Chem. 2013, 21, 2518–2526. [Google Scholar] [CrossRef]
- Yamada, Y.; Ochiai, C.; Yoshimura, K.; Tanaka, T.; Ohashi, N.; Narumi, T.; Nomura, W.; Harada, S.; Matsushita, S.; Tamamura, H. CD4 mimics targeting the mechanism of HIV entry. Bioorg. Med. Chem. Lett. 2010, 20, 354–358. [Google Scholar] [CrossRef]
- Hashimoto, C.; Narumi, T.; Otsuki, H.; Hirota, Y.; Arai, H.; Yoshimura, K.; Harada, S.; Ohashi, N.; Nomura, W.; Miura, T.; et al. A CD4 mimic as an HIV entry inhibitor: Pharmacokinetics. Bioorg. Med. Chem. 2013, 21, 7884–7889. [Google Scholar] [CrossRef]
- Yoshimura, K.; Harada, S.; Shibata, J.; Hatada, M.; Yamada, Y.; Ochiai, C.; Tamamura, H.; Matsushita, S. Enhanced exposure of human immunodeficiency virus type 1 primary isolate neutralization epitopes through binding of CD4 mimetic compounds. J. Virol. 2010, 84, 7558–7568. [Google Scholar] [CrossRef]
- Courter, J.R.; Madani, N.; Sodroski, J.; Schön, A.; Freire, E.; Kwong, P.D.; Hendrickson, W.A.; Chaiken, I.M.; LaLonde, J.M.; Smith, A.B., III. Structure-based design, synthesis and validation of CD4-mimetic small molecule inhibitors of HIV-1 entry: Conversion of a viral entry agonist to an antagonist. Acc. Chem. Res. 2014, 47, 1228–1237. [Google Scholar] [CrossRef]
- Liu, Y.; Schön, A.; Freire, E. Optimization of CD4/gp120 inhibitors by thermodynamic-guided alanine-scanning mutagenesis. Chem. Biol. Drug Des. 2013, 81, 72–78. [Google Scholar] [CrossRef]
- Curreli, F.; Kwon, Y.D.; Zhanga, H.; Scacalossia, D.; Belov, D.S.; Tikhonov, A.A.; Andreev, I.A.; Altieric, A.; Kurkin, A.V.; Kwong, P.D.; et al. Structure-based design of a small molecule CD4-antagonist with broad spectrum anti-HIV-1 activity. J. Med. Chem. 2015, 58, 6909–6927. [Google Scholar] [CrossRef]
- Morellato-Castillo, L.; Acharya, P.; Combes, O.; Michiels, J.; Descours, A.; Ramos, O.H.P.; Yang, Y.; Guido Vanham, G.; Ariën, K.K.; Kwong, P.D.; et al. Interfacial cavity filling to optimize CD4–mimetic miniprotein interactions with HIV-1 surface glycoprotein. J. Med. Chem. 2013, 56, 5033–5047. [Google Scholar] [CrossRef][Green Version]
- Acharya, P.; Lusvarghi, S.; Bewley, C.A.; Kwong, P.D. HIV-1 gp120 as a therapeutic target: Navigating a moving labyrinth. Expert Opin. Ther. Targets 2015, 19, 1–19. [Google Scholar] [CrossRef]
- Curreli, F.; Kwon, Y.D.; Belov, D.S.; Ramesh, R.R.; Kurkin, A.V.; Altieri, A.; Kwong, P.D.; Debnath, A.K. Synthesis, antiviral potency, in vitro ADMET, and X-ray structure of potent CD4 mimics as entry inhibitors that target the Phe43 cavity of HIV-1 gp120. J. Med. Chem. 2017, 60, 3124–3153. [Google Scholar] [CrossRef]
- Curreli, F.; Belov, D.S.; Kwon, Y.D.; Ramesh, R.; Furimsky, A.M.; O’Loughlin, K.; Byrge, P.C.; Iyer, L.V.; Mirsalis, J.C.; Kurkin, A.V.; et al. Structure-based lead optimization to improve antiviral potency and ADMET properties of phenyl-1H-pyrrole-carboxamide entry inhibitors targeted to HIV-1 gp120. Eur. J. Med. Chem. 2018, 154, 367–391. [Google Scholar] [CrossRef]
- Kwon, Y.D.; Finzi, A.; Wu, X.; Dogo-Isonagie, C.; Lee, L.K.; Moore, L.R.; Schmidt, D.; Stuckey, J.; Yang, Y.; Zhou, T.; et al. Unliganded HIV-1 gp120 core structures assume the CD4-bound conformation with regulation by quaternary interactions and variable loops. Proc. Natl. Acad. Sci. USA 2012, 109, 5663–5668. [Google Scholar] [CrossRef]
- Kwon, Y.D.; Lalonde, J.M.; Yang, Y.; Elban, M.A.; Sugawara, A.; Courter, J.R.; Jones, D.M.; Smith, A.B., III; Debnath, A.K.; Kwong, P.D. Crystal structures of HIV-1 gp120 envelope glycoprotein in complex with NBD analogues that target the CD4-binding site. PLoS ONE 2014, 9, e85940. [Google Scholar] [CrossRef]
- Wadood, A.; Ahmed, N.; Shah, L.; Ahmad, A.; Hassan, H.; Shams, S. In-silico drug design: An approach which revolutionarised the drug discovery process. OA Drug Des. Deliv. 2013, 1, 3–7. [Google Scholar]
- Sliwoski, G.; Kothiwale, S.; Meiler, J.; Lowe, E.W., Jr. Computational Methods in Drug Discovery. Pharmacol. Rev. 2014, 66, 334–395. [Google Scholar] [CrossRef]
- Mallipeddi, P.L.; Kumar, G.; White, S.W.; Webb, T.R. Recent advances in computer-aided drug design as applied to anti-influenza drug discovery. Curr. Top. Med. Chem. 2014, 14, 1875–1889. [Google Scholar] [CrossRef]
- Leelananda, S.P.; Lindert, S. Computational methods in drug discovery. Beilstein J. Org. Chem. 2016, 12, 2694–2718. [Google Scholar] [CrossRef]
- Katsila, T.; Spyroulias, G.A.; Patrinos, G.P.; Matsoukas, M.-T. Computational approaches in target identification and drug discovery. Comput. Struct. Biotechnol. J. 2016, 14, 177–184. [Google Scholar] [CrossRef]
- Knight-Schrijver, V.R.; Chelliah, V.; Cucurull-Sanchez, L.; Le Novère, N. The promises of quantitative systems pharmacology modelling for drug development. Comput. Struct. Biotechnol. J. 2016, 14, 363–370. [Google Scholar] [CrossRef]
- Shoichet, B.K. Virtual screening of chemical libraries. Nature 2004, 432, 862–865. [Google Scholar] [CrossRef]
- Banegas-Luna, A.-J.; Cerón-Carrasco, J.P.; Pérez-Sánchez, H. A review of ligand-based virtual screening web tools and screening algorithms in large molecular databases in the age of big data. Future Med. Chem. 2018, 10, 2641–2658. [Google Scholar] [CrossRef]
- Huang, H.; Zhang, G.; Zhou, Y.; Lin, C.; Chen, S.; Lin, Y.; Mai, S.; Huang, Z. Reverse screening methods to search for the protein targets of chemopreventive compounds. Front. Chem. 2018, 6, 138. [Google Scholar] [CrossRef]
- Kolb, H.C.; Finn, M.G.; Sharpless, K.B. Click chemistry: Diverse chemical function from a few good reactions. Angew. Chem. Int. Ed. 2001, 40, 2004–2021. [Google Scholar] [CrossRef]
- Moses, J.E.; Moorhouse, A.D. The growing applications of click chemistry. Chem. Soc. Rev. 2007, 36, 1249–1262. [Google Scholar] [CrossRef]
- Hein, C.D.; Liu, X.-M.; Wang, D. Click chemistry, a powerful tool for pharmaceutical sciences. Pharm. Res. 2008, 10, 2216–2230. [Google Scholar] [CrossRef]
- Thirumurugan, P.; Matosiuk, D.; Jozwiak, K. Click chemistry for drug development and diverse chemical–biology applications. Chem. Rev. 2013, 113, 4905–4979. [Google Scholar] [CrossRef]
- Durrant, J.D.; McCammon, J.A. AutoClickChem: Click chemistry in silico. PLoS Comput. Biol. 2012, 8, e1002397. [Google Scholar] [CrossRef]
- Irwin, J.J.; Sterling, T.; Mysinger, M.M.; Bolstad, E.S.; Coleman, R.G. ZINC: A free tool to discover chemistry for biology. J. Chem. Inf. Model. 2012, 52, 1757–1768. [Google Scholar] [CrossRef]
- Sander, T.; Freyss, J.; von Korff, M.; Rufener, C. DataWarrior: An open-source program for chemistry aware data visualization and analysis. J. Chem. Inf. Model. 2015, 55, 460–473. [Google Scholar] [CrossRef]
- Lipinski, C.A.; Lombardo, F.; Dominy, B.W.; Feeney, P.J. Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Adv. Drug Deliv. Rev. 2001, 46, 3–26. [Google Scholar] [CrossRef]
- Handoko, S.D.; Ouyang, X.; Su, C.T.T.; Kwoh, C.K.; Ong, Y.S. QuickVina: Accelerating AutoDock Vina using gradient-based heuristics for global optimization. IEEE/ACM Trans. Comput. Biol. Bioinform. 2012, 9, 1266–1272. [Google Scholar] [CrossRef]
- Open Babel. Available online: http://openbabel.org/wiki/Main_Page (accessed on 27 June 2019).
- Rappe, A.K.; Casewit, C.J.; Colwell, K.S.; Goddard, W.A., III; Skiff, W.M. UFF, a full periodic table force field for molecular mechanics and molecular dynamics simulations. J. Am. Chem. Soc. 1992, 114, 10024–10035. [Google Scholar] [CrossRef]
- Stewart, J.J.P. Optimization of parameters for semiempirical methods VI: More modifications to the NDDO approximations and re-optimization of parameters. J. Mol. Model. 2013, 19, 1–32. [Google Scholar] [CrossRef]
- OpenMOPAC. Available online: http://OpenMOPAC.net (accessed on 27 June 2019).
- Klamt, A.; Schüürmann, G. COSMO: A new approach to dielectric screening in solvents with explicit expressions for the screening energy and its gradient. J. Chem. Soc. Perkin Trans. 2 1993, 2, 799–805. [Google Scholar] [CrossRef]
- Klamt, A. COSMO-RS: From Quantum Chemistry to Fluid Phase Thermodynamics and Drug Design, 1st ed.; Elsevier: Boston, MA, USA, 2005; p. 246. [Google Scholar]
- Klamt, A.; Moya, C.; Palomar, J. A comprehensive comparison of the IEFPCM and SS(V)PE continuum solvation methods with the COSMO approach. J. Chem. Theory Comput. 2015, 11, 4220–4225. [Google Scholar] [CrossRef]
- Høyvik, I.-M.; Jansik, B.; Jørgensen, P. Trust region minimization of orbital localization functions. J. Chem. Theory Comput. 2012, 8, 3137–3146. [Google Scholar] [CrossRef]
- Lehtola, S.; Jónsson, H. Unitary optimization of localized molecular orbitals. J. Chem. Theory Comput. 2013, 9, 5365–5372. [Google Scholar] [CrossRef]
- Durrant, J.D.; McCammon, J.A. Binana: A novel algorithm for ligand-binding characterization. J. Mol. Graph. Model. 2011, 29, 888–893. [Google Scholar] [CrossRef]
- Pettersen, E.F.; Goddard, T.D.; Huang, C.C.; Couch, G.S.; Greenblatt, D.M.; Meng, E.C.; Ferrin, T.E. UCSF Chimera—A visualization system for exploratory research and analysis. J. Comput. Chem. 2004, 13, 16051–16612. [Google Scholar] [CrossRef]
- McDonald, I.K.; Thornton, J.M. Satisfying hydrogen bonding potential in proteins. J. Mol. Biol. 1994, 238, 777–793. [Google Scholar] [CrossRef]
- Durrant, J.D.; McCammon, J.A. NNScore 2.0: A neural-network receptor-ligand scoring function. J. Chem. Inf. Model. 2011, 51, 2897–2903. [Google Scholar] [CrossRef]
- Case, D.A.; Darden, T.A.; Cheatham, T.E., III; Simmerling, C.L.; Wang, J.; Duke, R.E. AMBER 11; University of California: San Francisco, CA, USA, 2010. [Google Scholar]
- Wang, J.; Wolf, R.M.; Caldwell, J.W.; Kollman, P.A.; Case, D.A. Development and testing of a general Amber force field. J. Comput. Chem. 2004, 25, 1157–1174. [Google Scholar] [CrossRef]
- Jorgensen, W.L.; Chandrasekhar, J.; Madura, J.D.; Impey, R.W.; Klein, M.L. Comparison of simple potential functions for simulating liquid water. J. Chem. Phys. 1983, 79, 926–935. [Google Scholar] [CrossRef]
- Ryckaert, J.P.; Ciccotti, G.; Berendsen, H.J.C. Numerical integration of the Cartesian equations of motion of a system with constraints: Molecular dynamics of n-alkanes. J. Comput. Phys. 1977, 23, 327–341. [Google Scholar] [CrossRef]
- Essmann, U.; Perera, L.; Berkowitz, M.L.; Darden, T.; Lee, H.; Pedersen, L.G. A smooth particle mesh Ewald method. J. Chem. Phys. 1995, 103, 8577–8593. [Google Scholar] [CrossRef]
- Sun, H.; Li, Y.; Tian, S.; Xu, L.; Hou, T. Assessing the performance of MM/PBSA and MM/GBSA methods. 4. Accuracies of MM/PBSA and MM/GBSA methodologies evaluated by various simulation protocols using PDBbind data set. Phys. Chem. Chem. Phys. 2014, 16, 16719–16729. [Google Scholar] [CrossRef]
- Xu, L.; Sun, H.; Li, Y.; Wang, J.; Hou, T. Assessing the performance of MM/PBSA and MM/GBSA methods. 3. The impact of force fields and ligand charge models. J. Phys. Chem. B 2013, 117, 8408–8421. [Google Scholar] [CrossRef]
- Sun, H.; Li, Y.; Shen, M.; Tian, S.; Xu, L.; Pan, P.; Guan, Y.; Hou, T. Assessing the performance of MM/PBSA and MM/GBSA methods. 5. Improved docking performance using high solute dielectric constant MM/GBSA and MM/PBSA rescoring. Phys. Chem. Chem. Phys. 2014, 16, 22035–22045. [Google Scholar] [CrossRef]
- Lindorff-Larsen, K.; Piana, S.; Palmo, K.; Maragakis, P.; Klepeis, J.L.; Dror, R.O.; Shaw, D.E. Improved side-chain torsion potentials for the Amber ff99SB protein force field. Proteins 2010, 78, 1950–1958. [Google Scholar] [CrossRef]
- Drug Likeness Tool (DruLiTo). Available online: http://www.niper.gov.in/pi_dev_tools/DruLiToWeb/ (accessed on 27 June 2019).
- Moebius, U.; Clayton, L.K.; Abraham, S.; Harrison, S.C.; Reinherz, E.L. The human immunodeficiency virus-gp120 binding-site on CD4—Delineation by quantitative equilibrium and kinetic binding studies of mutants in conjunction with a high-resolution CD4 atomic-structure. J. Exp. Med. 1992, 176, 507–517. [Google Scholar] [CrossRef]
- Olshevsky, U.; Helseth, E.; Furman, C.; Li, J.; Haseltine, W.; Sodroski, J. Identification of individual human-immunodeficiency-virus type-1 gp120 amino-acids important for CD4 receptor-binding. J. Virol. 1990, 64, 5701–5707. [Google Scholar]
- Sulimov, A.V.; Kutov, D.C.; Katkova, E.V.; Sulimov, V.B. Combined docking with classical force field and quantum chemical semiempirical method PM7. Adv. Bioinform. 2017, 5, 1–6. [Google Scholar] [CrossRef]
- Sharma, G.; First, E.A. Thermodynamic Analysis Reveals a Temperature-dependent Change in the Catalytic Mechanism of Bacillus stearothermophilus Tyrosyl-tRNA Synthetase. J. Biol. Chem. 2009, 284, 4179–4190. [Google Scholar] [CrossRef]
- Myszka, D.G.; Sweet, R.W.; Hensley, P.; Brigham-Burke, M.; Kwong, P.D.; Hendrickson, W.A.; Wyatt, R.; Sodroski, J.; Doyle, M.L. Energetics of the HIV gp120-CD4 binding reaction. Proc. Natl. Acad. Sci. USA 2000, 97, 9026–9031. [Google Scholar] [CrossRef]
- Mao, Y.; Wang, L.; Gu, C.; Herschhorn, A.; Xiang, S.-H.; Haim, H.; Yang, X.; Sodroski, J. Subunit organization of the membrane-bound HIV-1 envelope glycoprotein trimer. Nat. Struct. Mol. Biol. 2012, 19, 893–899. [Google Scholar] [CrossRef]
- Mao, Y.; Wang, L.; Gu, C.; Herschhorn, A.; Désormeaux, A.; Finzi, A.; Xiang, S.-H.; Sodroski, J.G. Molecular architecture of the uncleaved HIV-1 envelope glycoprotein trimer. Proc. Natl. Acad. Sci. USA 2013, 110, 12438–12443. [Google Scholar] [CrossRef]
- Tran, E.E.H.; Borgnia, M.J.; Kuybeda, O.; Schauder, D.M.; Bartesaghi, A.; Frank, G.A.; Sapiro, G.; Milne, J.L.S.; Subramaniam, S. Structural mechanism of trimeric HIV-1 envelope glycoprotein activation. PLoS Pathog. 2012, 8, e1002797. [Google Scholar] [CrossRef]
- Moraca, F.; Acharya, K.; Melillo, B.; Smith, A.B., III; Chaiken, I.; Abrams, C.F. Computational evaluation of HIV-1 gp120 conformations of soluble trimeric gp140 structures as targets for de novo docking of first- and second-generation small-molecule CD4 mimics. J. Chem. Inf. Model. 2016, 56, 2069–2079. [Google Scholar] [CrossRef]
- LaLonde, J.M.; Elban, M.A.; Courter, J.R.; Sugawara, A.; Soeta, T.; Madani, N.; Princiotto, A.M.; Kwon, Y.D.; Kwong, P.D.; Schön, A.; et al. Design, synthesis and biological evaluation of small molecule inhibitors of CD4-gp120 binding based on virtual screening. Bioorg. Med. Chem. 2011, 19, 91–101. [Google Scholar] [CrossRef]
- Julien, J.P.; Cupo, A.; Sok, D.; Stanfield, R.L.; Lyumkis, D.; Deller, M.C.; Klasse, P.J.; Burton, D.R.; Sanders, R.W.; Moore, J.P.; et al. Crystal structure of a soluble cleaved HIV-1 envelope trimer. Science 2013, 342, 1477–1483. [Google Scholar] [CrossRef]
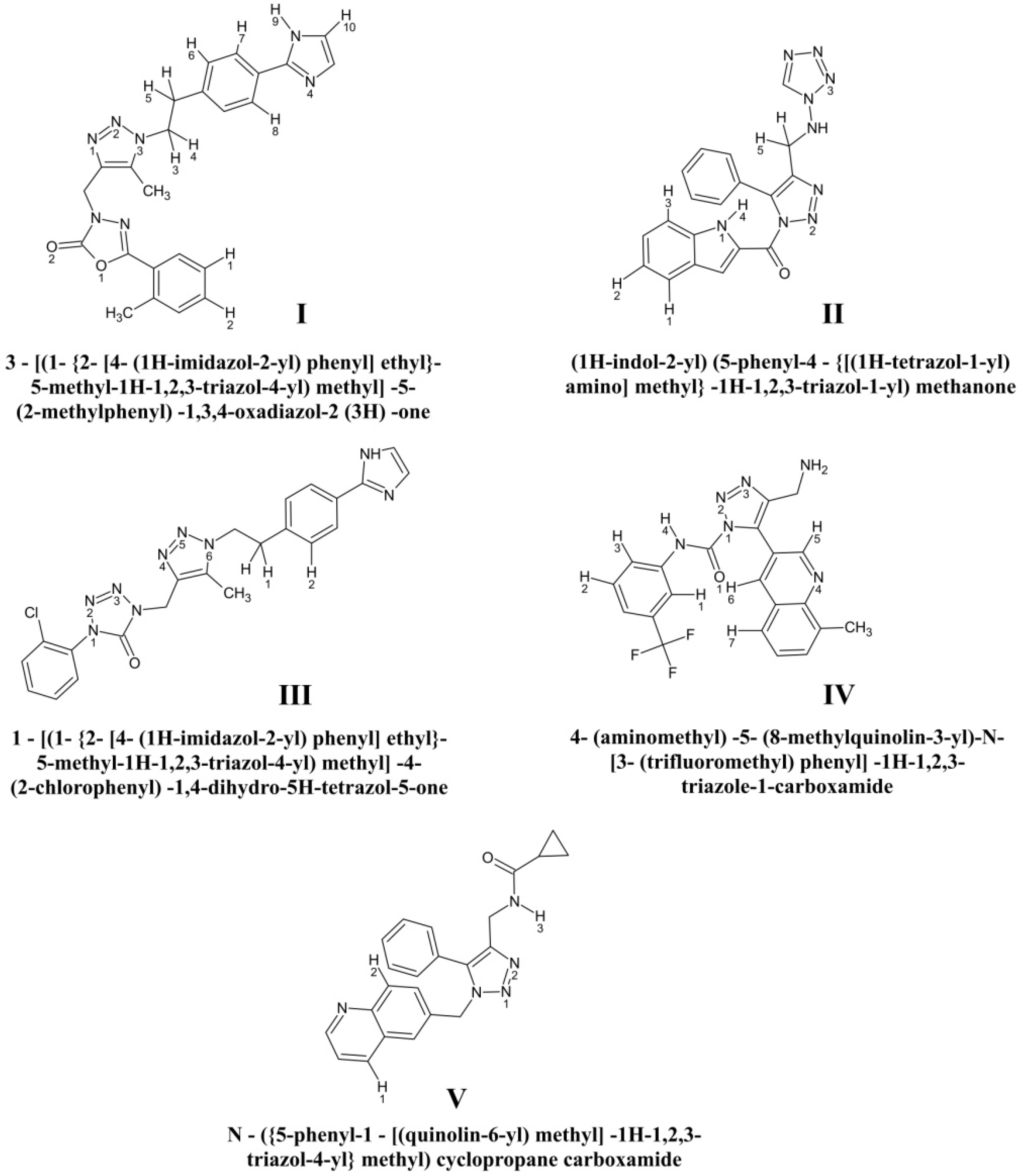
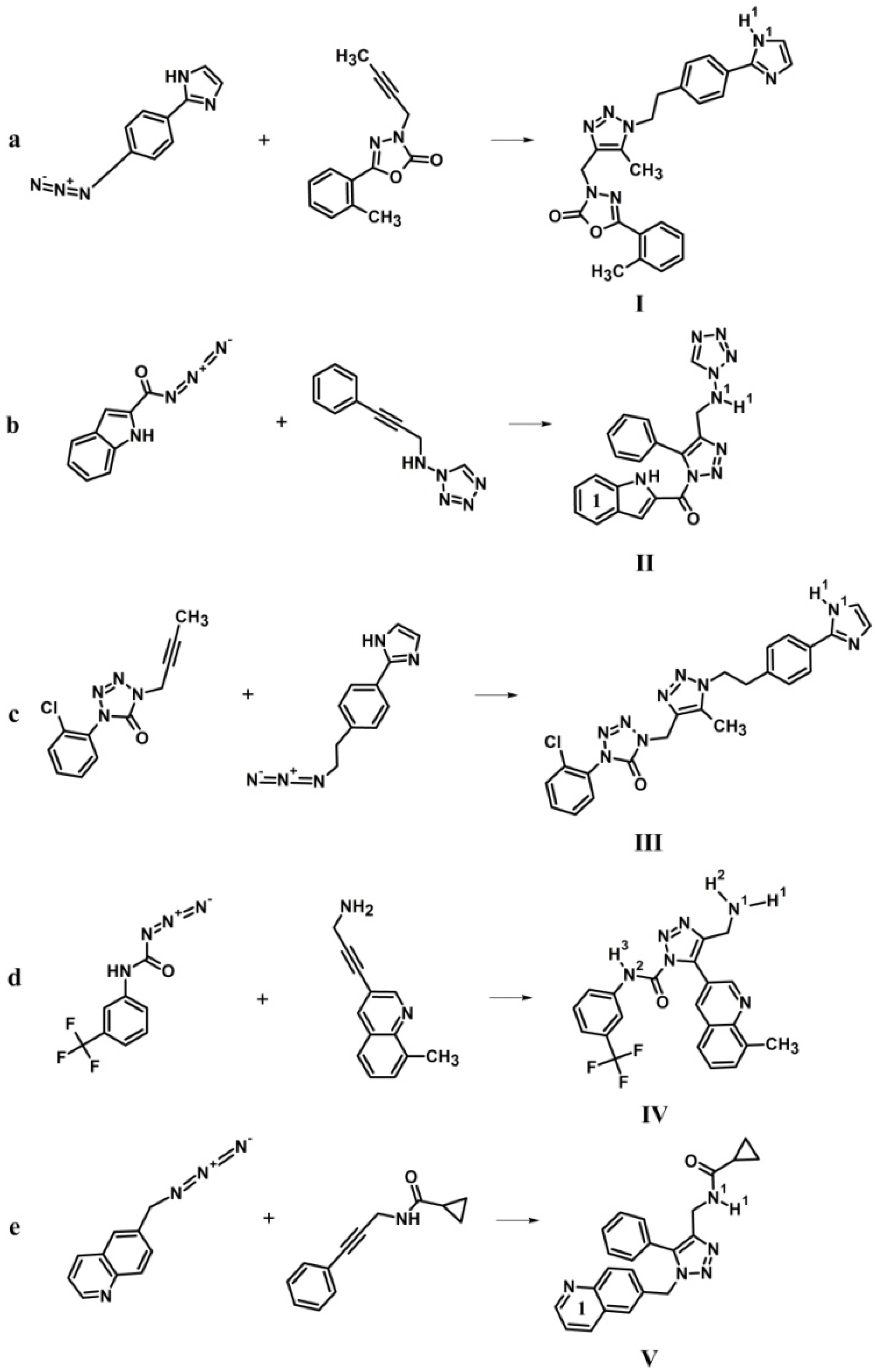
| Compound | Systematic Name | Chemical Formula | Molecular Mass (Da) | Log P 2 | Number of H-Bond Donors | Number of H-Bond Acceptors |
|---|---|---|---|---|---|---|
| I | 3-[(1-{2-[4-(1H-imidazol-2-yl) phenyl]ethyl}-5-methyl-1H-1,2,3-triazol-4-yl) methyl]-5-(2-methylphenyl)-1,3,4-oxadiazol-2 (3H)-one | C24H23N7O2 | 419.02 | 2.634 | 1 | 9 |
| II | (1H-indol-2-yl) (5-phenyl-4-{[(1H-tetrazol-1-yl)amino]methyl}-1H-1,2,3-triazol-1-yl) methanone | C19H15N9O | 372.04 | 1.131 | 1 | 10 |
| III | 1-[(1-{2-[4-(1H-imidazol-2-yl) phenyl] ethyl}-5-methyl-1H-1,2,3-triazol-4-yl) methyl]-4-(2-chlorophenyl)-1,4-dihydro-5H-tetrazol-5-one | C22H20ClN9O | 419.03 | 0.125 | 1 | 8 |
| IV | 4-(aminomethyl)-5-(8-methylquinolin-3-yl)-N-[3-(trifluoromethyl)phenyl]-1H-1,2,3-triazole-1-carboxamide | C21H17F3N6 | 412.03 | 0.905 | 2 | 7 |
| V | N-({5-phenyl-1-[(quinolin-6-yl) methyl]-1H-1,2,3-triazol-4-yl}methyl)cyclopropane carboxamide | C23H21N5O | 363.02 | 1.120 | 1 | 6 |
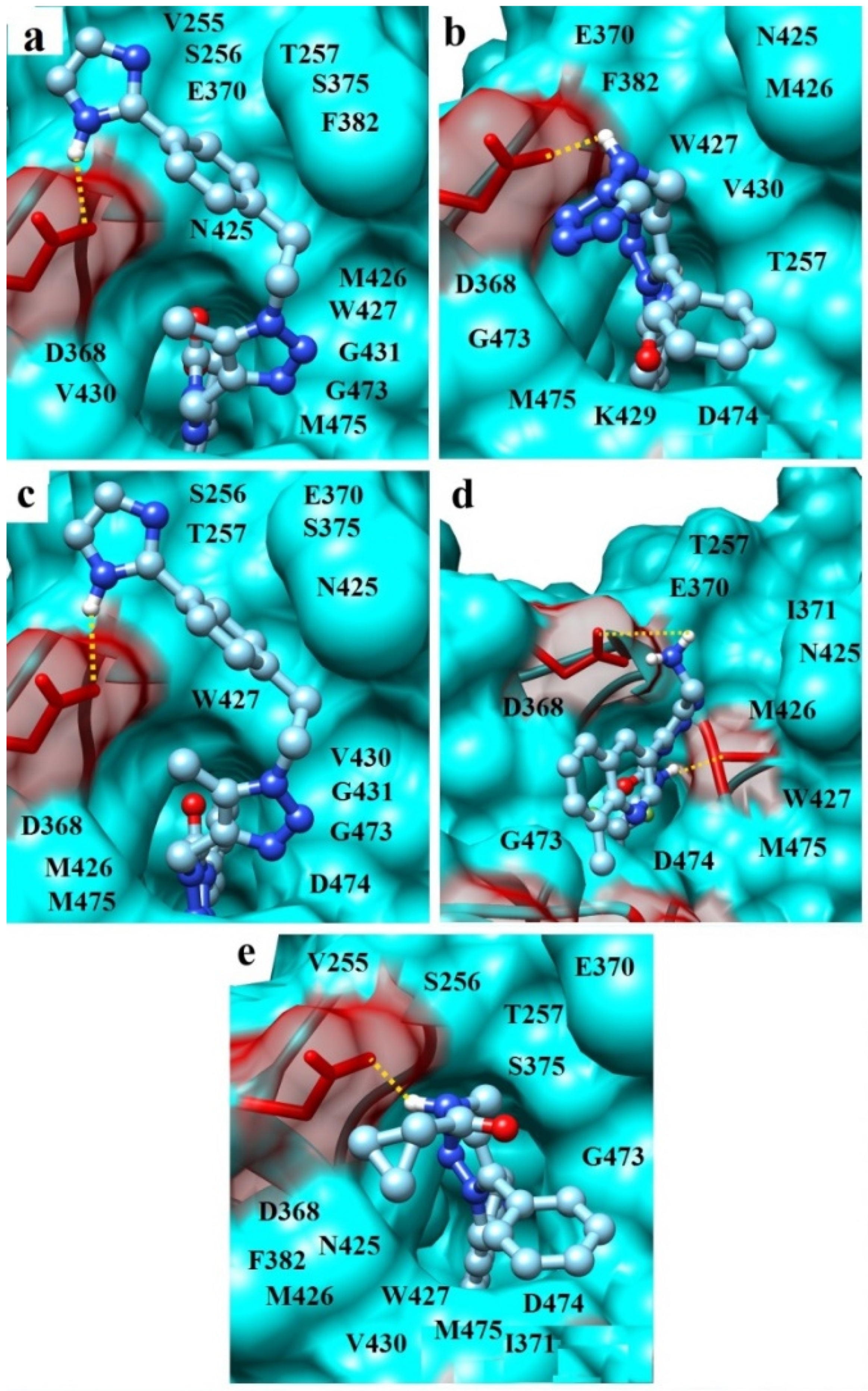
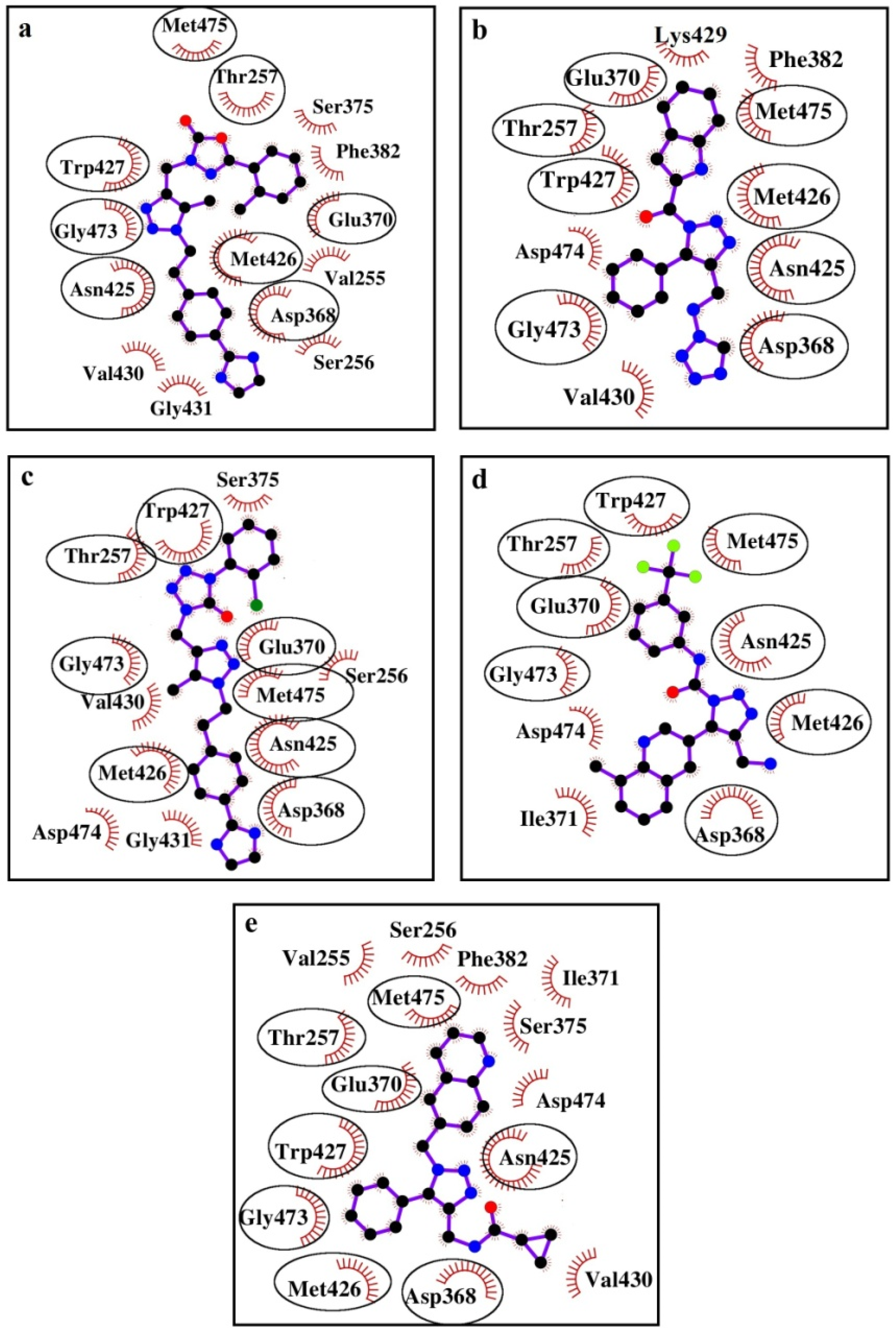
| Compound | Hydrogen Bond 1 | Van der Waals Contacts 2 | Salt Bridges and π-π-Interactions 3 |
|---|---|---|---|
| I | N1H1 ...OD2[D368] | V255(5), S256(2), T257(3), D368(5), E370(8), S375(5), F382(2), N425(11), M426(5), W427(12), V430(7), G431(3), G473(4), M475(8) | - |
| II | N1H1 ...OD1[D368] | D368(4), E370(7), F382(2), N425(8), M426(5), W427(7), V430(5), G473(7), M475(5), D474(7), K429(5), T257(4) | 1...W427 (T-shaped-interaction) |
| III | N1H1 ...OD2[D368] | S256(2), T257(3), D368(5), E370(8), S375(2), N425(7), M426(8), W427(13), V430(5), G431(3), G473(4), M475(2), D474(8) | - |
| IV | N1H1...OD2[D368] N2H3...O[M426] | T257(2), D368(6), E370(8), I371(7), N425(6), M426(4), W427(9), M475(6), G473(8), D474(5) | N1H1H2...D368 (salt bridge) |
| V | N1H1 ...OD2[D368] | V255(3), S256(2), T257(5), D368(3), E370(8), S375(3), F382(2), N425(8), M426(4), W427(9), V430(6), G473(12), M475(7), D474(5), I371(3) | 1...W427 (T-shaped-interaction) |
| Compound | I | II | III | IV | V | NBD-11021 |
|---|---|---|---|---|---|---|
| ∆G, kcal/mol 1 | −10.6 | −9.8 | −9.9 | −9.5 | −9.8 | −7.8 |
| Kd (µM) 2 | 0.0075 | 0.2767 | 0.4675 | 0.5086 | 0.0593 | 2.1 |
| ∆G, kcal/mol 3 | −11.1 | −9.0 | −8.7 | −8.6 | −9.9 | −8.0 |
| Compound | <ΔH> kcal/mol | (ΔH)STD kcal/mol | <TΔS> kcal/mol | (TΔS)STD kcal/mol | <ΔG> kcal/mol | ΔGSTD kcal/mol |
|---|---|---|---|---|---|---|
| I | −49.40 | 5.06 | −20.89 | 7.01 | −28.51 | 7.65 |
| II | −32.64 | 3.95 | −18.61 | 6.28 | −14.03 | 7.42 |
| III | −34.30 | 3.03 | −23.69 | 9.72 | −10.61 | 7.18 |
| IV | −29.00 | 4.51 | −19.50 | 8.12 | −9.50 | 6.29 |
| V | −27.93 | 4.02 | −20.35 | 9.06 | −7.58 | 5.91 |
| NBD-11021 | −30.41 | 3.60 | −22.42 | 9.89 | −7.99 | 7.52 |
| Residue of gp120 | CD4-Mimetic Candidate | ||||
|---|---|---|---|---|---|
| I | II | III | IV | V | |
| Residue Contribution to the Binding Enthalpy (kcal/mol) 2, 3 | |||||
| Gly-128 | −0.99 | - | - | - | - |
| Ala-129 | −0.78 | - | - | - | |
| Gly-194 | −0.59 | - | - | - | - |
| Val-255 | −1.23 | −1.32 | −1.35 | −1.08 | −1.04 |
| Ser-256 | −0.68 | −1.03 | - | - | −0.80 |
| Thr-257 | −1.14 | −1.65 | - | −1.39 | −1.29 |
| Asp-368 | −0.92 | −1.10 | −0.15 | −0.85 | −1.42 |
| Glu-370 | −2.00 | −2.63 | −1.70 | −3.10 | - |
| Ile-371 | - | −0.82 | - | - | −0.91 |
| Ser-375 | −1.76 | −2.36 | - | −0.82 | −0.98 |
| Phe-376 | −0.66 | −0.59 | - | - | - |
| Phe-382 | −0.78 | −0.69 | −0.68 | −0.65 | - |
| Tyr-384 | −0.75 | −0.64 | - | - | -0.68 |
| Ile-424 | −0.76 | −0.74 | −1.09 | −0.88 | −0.59 |
| Asn-425 | −6.49 | −3.48 | −2.68 | −2.96 | −2.76 |
| Met-426 | −1.70 | −1.63 | −2.17 | −3.00 | −1.57 |
| Trp-427 | −3.89 | −2.73 | −4.49 | −5.1 | −3.18 |
| Gln-428 | −0.68 | - | −0.99 | −0.88 | - |
| Lys-429 | −1.85 | - | −1.96 | −1.78 | - |
| Val-430 | −3.63 | - | −1.50 | −2.10 | −0.72 |
| Gly-431 | −1.10 | - | - | - | - |
| Gly-472 | - | -1.01 | - | - | -0.54 |
| Gly-473 | −1.54 | −1.54 | −1.29 | −1.24 | −2.83 |
| Asp-474 | −3.07 | −1.21 | −2.14 | −1.56 | −2.71 |
| Met-475 | −3.60 | −1.93 | −3.39 | −1.65 | −2.55 |
| Arg-476 | - | - | −0.54 | - | - |
| Ligand | Hydrogen Bond 1 |
|---|---|
| I | O1...HG2[Glu370; 71,3%], O2...HG2[Glu370; 20,9%], O1...HB3[Asn425; 15,6%], N4...HD21[Asn425; 99,8%], O2...HD22[Asn425; 39,7% ], N1...HB2[Trp427; 51,9%], O1...HB3[Trp427; 36,6%], N4...HA[Val430; 83,5%], N4...H[Gly431; 85,0%], N1...HA[Asp474; 35,0%], N2...HA[Asp474; 77,3%], N3...HA[Trp427; 71,1%], N1...H[Met475; 78,3%]; N2...H[Met475; 94,7%], N1...HB2[Met475; 52,0%], N2...HB2[Met475; 16,6%], N1...HG2[Met475; 32,9%], H1...O[Val255; 49,4%], H1...O[Ser375; 37,4%], H1... OD2[Asp368; 15,3%], H2...N[Phe376; 19,1%], H3...OD2[Asp474; 20,0%], H4...O[Gly473; 26,3%], H5...O[Trp427; 71,5%], H6...O[Trp427; 71,3%], H7...ND2[Asn425; 49,3%], H7...O[Met426; 71,5%], H8...O[Gly128; 39,0%], H9...O[Gly128; 42,4%], H10...O[Gly194; 22,3%] |
| II | N2...HG2[Glu370; 26,0%], N1...HG2[Glu370; 34,7%], N3...HA[Asp474; 26,2%], H1...O[Val255; 27,5%], H1... OD1[Asp368; 17,1%], H2...O[Phe376; 42,7%], H3...OH[Tyr384; 26,3%], H4...O[Asn425; 47,2%], H5...OD1[Asp368; 1,3%] |
| III | N2...HB3[Asn425; 44,2%], N3...HB3[Asn425; 25,5%], N5...HA[Trp427; 42,7%], N6...HA[Trp427; 50,2%], N4...HB2[Trp427; 39,5%], N1...HB3[Trp427; 31,4], N2...HB3[Trp427; 39,4%], N4...HA[Asp474; 82,8%], N5...HA[Asp474; 27,2%], N4...H[Met475; 84,6%], N5...H[Met475; 32,6%], N4...HB2[Met475; 30,7%], H1... OD2[Asp368; 18,7%], H1...O[Trp427; 68,3%], H2...O[Trp427; 45,3%] |
| IV | N1...HB3[Asn425; 26,8%], N2...HB3[Asn425; 53,5%], N3...HA[Val430; 44,2%], N3...H[Gly431; 40,9%], O1...HA2[Gly473; 45,4%], N4...HA3[Gly473; 36,2%], H4...O[Met426; 95,8%], H5...O[Gly473; 52,7%], H1...O[Asn425; 62,4%], H1... OD2[Asp368; 12,5%], H2...SD[Met475; 26,1%], H3...O[Gly473; 50,1%], H7...OD2[Asp368; 16.3%], H7...OD1[Asp368; 12.1%], H6...OD2[Asp368; 7.8%], H6…OD1[Asp368; 5.4%] |
| V | N2...HB2[Trp427; 29,5%], N1...HB2[Trp427; 27,5%], N2...HA[Asp474; 47,2%], N2...H[Met475; 48,8%], N1...HB2[Met475; 26,9%], H1...O[Ser256; 30,5%], H1... OD2[Asp368; 15,3%], H2...O[Asn425; 73,6%], H3...O[Gly473; 44,8%] |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Andrianov, A.M.; Nikolaev, G.I.; Kornoushenko, Y.V.; Xu, W.; Jiang, S.; Tuzikov, A.V. In Silico Identification of Novel Aromatic Compounds as Potential HIV-1 Entry Inhibitors Mimicking Cellular Receptor CD4. Viruses 2019, 11, 746. https://doi.org/10.3390/v11080746
Andrianov AM, Nikolaev GI, Kornoushenko YV, Xu W, Jiang S, Tuzikov AV. In Silico Identification of Novel Aromatic Compounds as Potential HIV-1 Entry Inhibitors Mimicking Cellular Receptor CD4. Viruses. 2019; 11(8):746. https://doi.org/10.3390/v11080746
Chicago/Turabian StyleAndrianov, Alexander M., Grigory I. Nikolaev, Yuri V. Kornoushenko, Wei Xu, Shibo Jiang, and Alexander V. Tuzikov. 2019. "In Silico Identification of Novel Aromatic Compounds as Potential HIV-1 Entry Inhibitors Mimicking Cellular Receptor CD4" Viruses 11, no. 8: 746. https://doi.org/10.3390/v11080746
APA StyleAndrianov, A. M., Nikolaev, G. I., Kornoushenko, Y. V., Xu, W., Jiang, S., & Tuzikov, A. V. (2019). In Silico Identification of Novel Aromatic Compounds as Potential HIV-1 Entry Inhibitors Mimicking Cellular Receptor CD4. Viruses, 11(8), 746. https://doi.org/10.3390/v11080746






