Characterization of EIAV env Quasispecies during Long-Term Passage In Vitro: Gradual Loss of Pathogenicity
Abstract
1. Introduction
2. Materials and Methods
2.1. Study Subjects
2.2. Viral RNA Extraction and cDNA Synthesis
2.3. Bulk PCR
2.4. Probe Design and Real-Time PCR
2.5. SGA (Single Genome Amplification)
2.6. Data Processing and Analysis
3. Results
3.1. General Characteristics of EIAV env Variability during the Course of EIAV Attenuation Using Bulk PCR
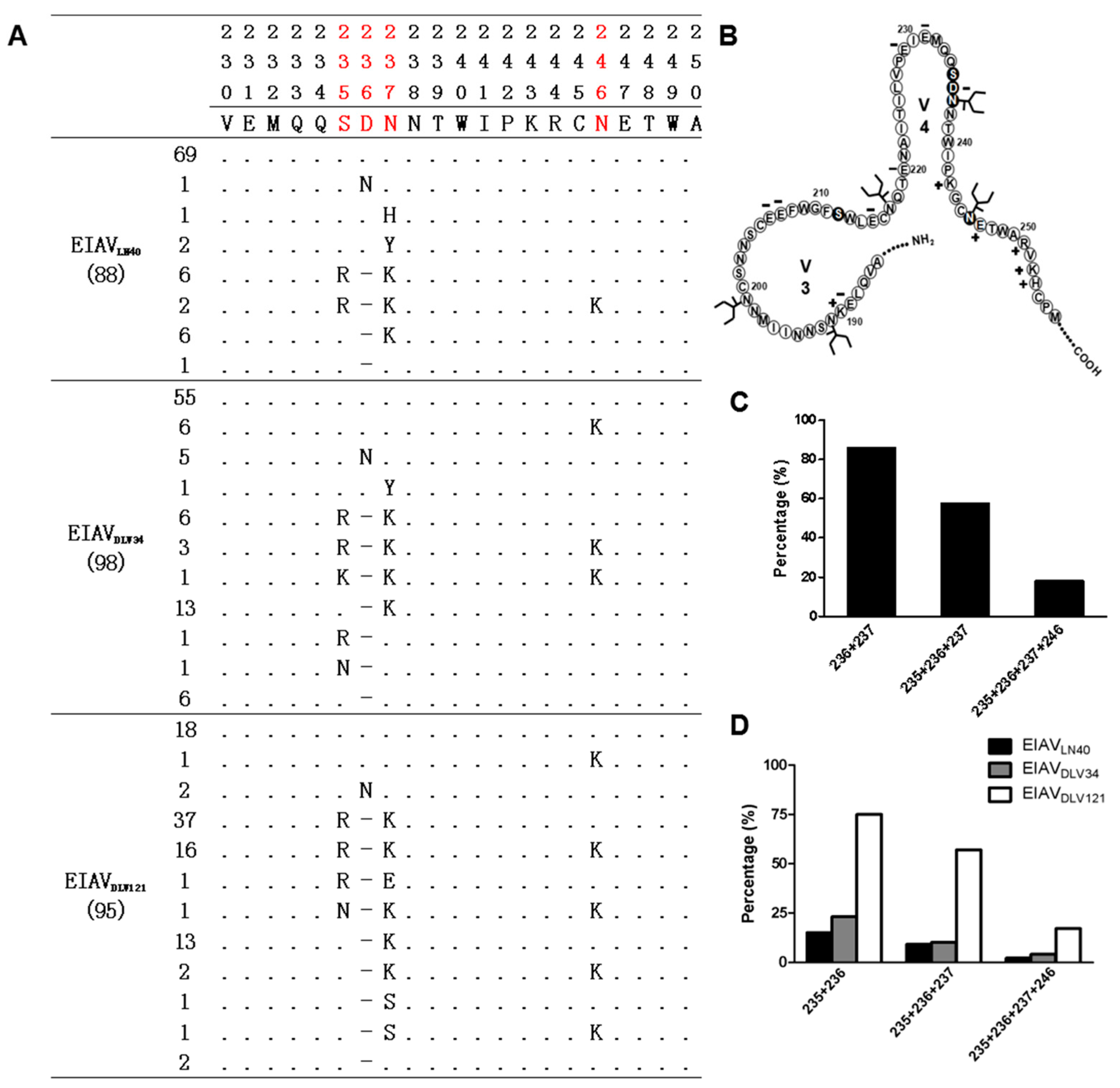
3.2. Precise Characteristics of the EIAV env Mutation Distribution during the Attenuation Process
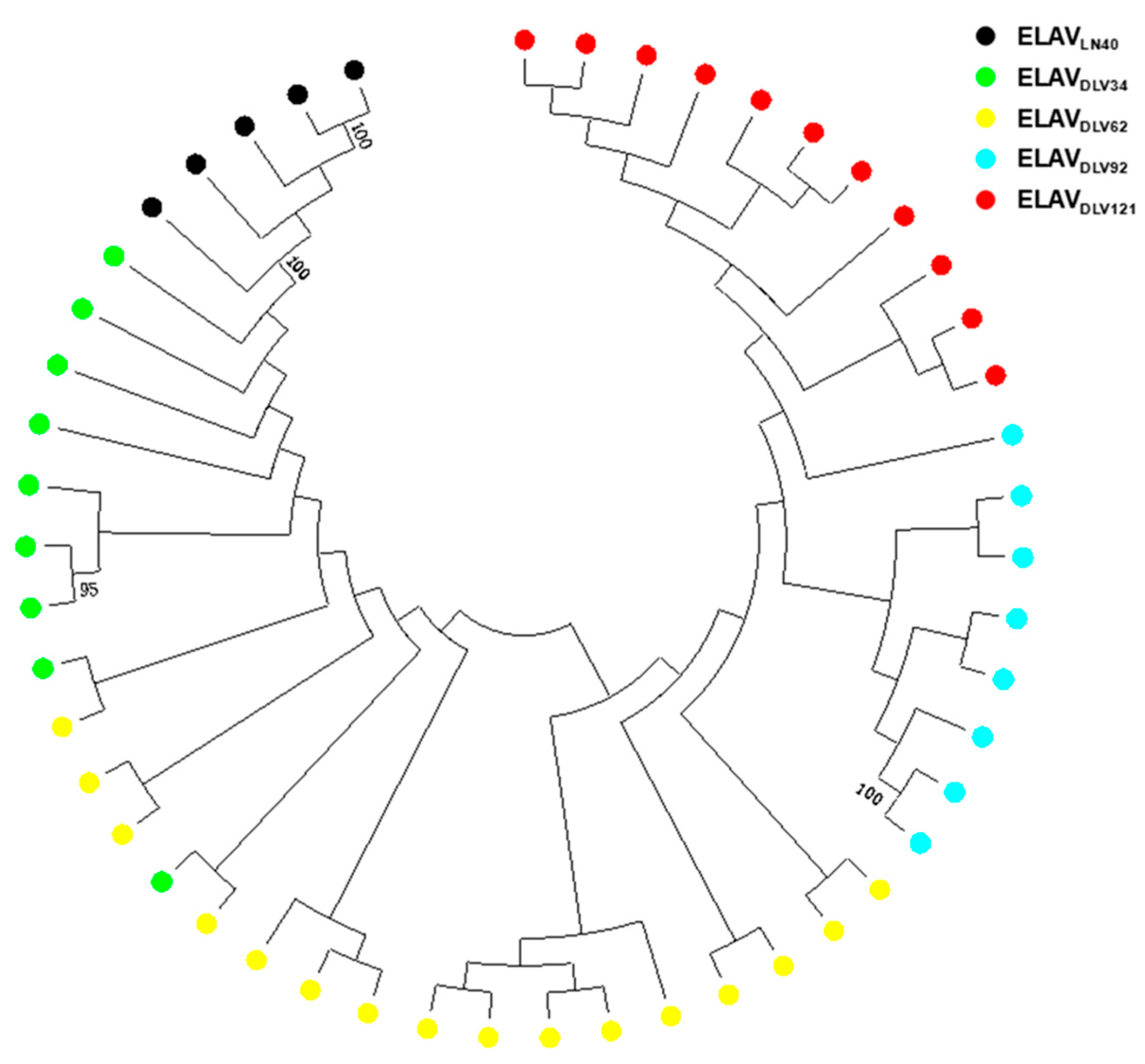
3.3. The Relationship between EIAV Attenuation In Vitro and env Gene Quasispecies
4. Discussion
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Yuan, Q.; Liu, C.; Liang, Z.; Chen, X.; Diao, D.; Kong, X. The comparison of genetic variation in the envelope protein between various immunodeficiency viruses and equine infectious anemia virus. Virol. Sin. 2012, 27, 241–247. [Google Scholar] [CrossRef]
- Cook, R.F.; Leroux, C.; Issel, C.J. Equine infectious anemia and equine infectious anemia virus in 2013: A review. Vet. Microbiol. 2013, 167, 181–204. [Google Scholar] [CrossRef]
- Ma, J.; Shi, N.; Jiang, C.G.; Lin, Y.Z.; Wang, X.F.; Wang, S.; Lv, X.L.; Zhao, L.P.; Shao, Y.M.; Kong, X.G.; et al. A proviral derivative from a reference attenuated eiav vaccine strain failed to elicit protective immunity. Virology 2011, 410, 96–106. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Wang, S.; Lin, Y.; Jiang, C.; Ma, J.; Zhao, L.; Lv, X.; Wang, F.; Shen, R.; Kong, X.; et al. Genomic comparison between attenuated chinese equine infectious anemia virus vaccine strains and their parental virulent strains. Arch. Virol. 2011, 156, 353–357. [Google Scholar] [CrossRef]
- Wang, X.F.; Lin, Y.Z.; Li, Q.; Liu, Q.; Zhao, W.W.; Du, C.; Chen, J.; Wang, X.; Zhou, J.H. Genetic evolution during the development of an attenuated EIAV vaccine. Retrovirology 2016, 13, 9. [Google Scholar] [CrossRef]
- Bingham, R.J.; Dykeman, E.C.; Twarock, R. RNA virus evolution via a quasispecies-based model reveals a drug target with a high barrier to resistance. Viruses 2017, 9, 347. [Google Scholar] [CrossRef] [PubMed]
- Colizzi, E.S.; Hogeweg, P. Evolution of functional diversification within quasispecies. Genome Biol. Evol. 2014, 6, 1990–2007. [Google Scholar] [CrossRef]
- Park, J.M.; Niestemski, L.R.; Deem, M.W. Quasispecies theory for evolution of modularity. Phys. Rev. E Stat. Nonlin. Soft Matter Phys. 2015, 91, 012714. [Google Scholar] [CrossRef]
- Domingo, E.; Sheldon, J.; Perales, C. Viral quasispecies evolution. Microbiol. Mol. Biol. Rev. 2012, 76, 159–216. [Google Scholar] [CrossRef]
- Andino, R.; Domingo, E. Viral quasispecies. Virology 2015, 479–480, 46–51. [Google Scholar] [CrossRef] [PubMed]
- Domingo, E.; Martin, V.; Perales, C.; Grande-Perez, A.; Garcia-Arriaza, J.; Arias, A. Viruses as quasispecies: Biological implications. Curr. Top. Microbiol. 2006, 299, 51–82. [Google Scholar]
- Craigo, J.K.; Ezzelarab, C.; Cook, S.J.; Chong, L.; Horohov, D.; Issel, C.J.; Montelaro, R.C. Envelope determinants of equine lentiviral vaccine protection. PLoS ONE 2013, 8, e66093. [Google Scholar] [CrossRef]
- Du, J.S.; Wang, X.F.; Ma, J.; Wang, J.X.; Qin, Y.Y.; Zhu, C.H.; Liu, F.; Shao, Y.M.; Zhou, J.H.; Qiao, W.T.; et al. Structural and biochemical insights into the v/i505t mutation found in the eiav gp45 vaccine strain. Retrovirology 2014, 11. [Google Scholar] [CrossRef][Green Version]
- Duan, L.; Du, J.; Wang, X.; Zhou, J.; Wang, X.; Liu, X. Structural and functional characterization of EIAV gp45 fusion peptide proximal region and asparagine-rich layer. Virology 2016, 491, 64–72. [Google Scholar] [CrossRef]
- Wang, X.F.; Liu, Q.; Wang, Y.H.; Wang, S.; Chen, J.; Lin, Y.Z.; Ma, J.; Zhou, J.H.; Wang, X. Characterization of equine infectious anemia virus long terminal repeat quasispecies in vitro and in vivo. J. Virol. 2018, 92. [Google Scholar] [CrossRef]
- Lin, Y.Z.; Shen, R.X.; Zhu, Z.Y.; Deng, X.L.; Cao, X.Z.; Wang, X.F.; Ma, J.; Jiang, C.G.; Zhao, L.P.; Lv, X.L.; et al. An attenuated eiav vaccine strain induces significantly different immune responses from its pathogenic parental strain although with similar in vivo replication pattern. Antiviral Res. 2011, 92, 292–304. [Google Scholar] [CrossRef]
- Jiang, C.G.; Gao, X.; Ma, J.; Lin, Y.Z.; Wang, X.F.; Zhao, L.P.; Hua, Y.P.; Liu, D.; Zhou, J.H. C-terminal truncation of the transmembrane protein of an attenuated lentiviral vaccine alters its in vitro but not in vivo replication and weakens its potential pathogenicity. Virus Res. 2011, 158, 235–245. [Google Scholar] [CrossRef]
- Hayat, M.J.; Higgins, M. Understanding poisson regression. J. Nurs. Educ. 2014, 53, 207–215. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Tamura, K. Mega7: Molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef]
- Craigo, J.K.; Montelaro, R.C. Eiav envelope diversity: Shaping viral persistence and encumbering vaccine efficacy. Curr. HIV Res. 2010, 8, 81–86. [Google Scholar] [CrossRef]
- Etemad, B.; Ghulam-Smith, M.; Gonzalez, O.; White, L.F.; Sagar, M. Single genome amplification and standard bulk pcr yield hiv-1 envelope products with similar genotypic and phenotypic characteristics. J. Virol. Methods 2015, 214, 46–53. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Wang, X.F.; Wang, S.; Liu, Q.; Lin, Y.Z.; Du, C.; Tang, Y.D.; Na, L.; Wang, X.; Zhou, J.H. A unique evolution of the s2 gene of equine infectious anemia virus in hosts correlated with particular infection statuses. Viruses 2014, 6, 4265–4279. [Google Scholar] [CrossRef]
- Ntziora, F.; Paraskevis, D.; Haida, C.; Magiorkinis, E.; Manesis, E.; Papatheodoridis, G.; Manolakopoulos, S.; Beloukas, A.; Chryssoy, S.; Magiorkinis, G.; et al. Quantitative detection of the m204v hepatitis B virus minor variants by amplification refractory mutation system real-time PCR combined with molecular beacon technology. J. Clin. Microbiol. 2009, 47, 2544–2550. [Google Scholar] [CrossRef]
- Chen, Q.; Belmonte, I.; Buti, M.; Nieto, L.; Garcia-Cehic, D.; Gregori, J.; Perales, C.; Ordeig, L.; Llorens, M.; Soria, M.E.; et al. New real-time-pcr method to identify single point mutations in hepatitis C virus. World J. Gastroenterol. 2016, 22, 9604–9612. [Google Scholar] [CrossRef] [PubMed]
- Salazar-Gonzalez, J.F.; Bailes, E.; Pham, K.T.; Salazar, M.G.; Guffey, M.B.; Keele, B.F.; Derdeyn, C.A.; Farmer, P.; Hunter, E.; Allen, S.; et al. Deciphering human immunodeficiency virus type 1 transmission and early envelope diversification by single-genome amplification and sequencing. J. Virol. 2008, 82, 3952–3970. [Google Scholar] [CrossRef]
- Geng, Q.M.; Li, H.P.; Bao, Z.Y.; Liu, Y.J.; Zhuang, D.M.; Li, L.; Liu, S.Y.; Li, J.Y. Indinavir resistance evolution in one human immunodeficiency virus type 1 infected patient revealed by single-genome amplification. Virol. Sin. 2010, 25, 316–328. [Google Scholar] [CrossRef]
- Guinoiseau, T.; Moreau, A.; Hohnadel, G.; Ngo-Giang-Huong, N.; Brulard, C.; Vourc’h, P.; Goudeau, A.; Gaudy-Graffin, C. Deep sequencing is an appropriate tool for the selection of unique hepatitis C virus (HCV) variants after single genomic amplification. PLoS ONE 2017, 12, e0174852. [Google Scholar] [CrossRef]
- Presloid, J.B.; Novella, I.S. Rna viruses and rnai: Quasispecies implications for viral escape. Viruses 2015, 7, 3226–3240. [Google Scholar] [CrossRef] [PubMed]
- Liu, F.; Wu, X.; Li, L.; Zou, Y.; Liu, S.; Wang, Z. Evolutionary characteristics of morbilliviruses during serial passages in vitro: Gradual attenuation of virus virulence. Comp. Immunol. Microbiol. Infect. Dis. 2016, 47, 7–18. [Google Scholar] [CrossRef] [PubMed]
- Vignuzzi, M.; Stone, J.K.; Arnold, J.J.; Cameron, C.E.; Andino, R. Quasispecies diversity determines pathogenesis through cooperative interactions in a viral population. Nature 2006, 439, 344–348. [Google Scholar] [CrossRef]
- Martella, V.; Blixenkrone-Moller, M.; Elia, G.; Lucente, M.S.; Cirone, F.; Decaro, N.; Nielsen, L.; Banyai, K.; Carmichael, L.E.; Buonavoglia, C. Lights and shades on an historical vaccine canine distemper virus, the rockborn strain. Vaccine 2011, 29, 1222–1227. [Google Scholar] [CrossRef]
- Han, X.; Zhang, P.; Yu, W.; Xiang, W.; Li, X. Amino acid mutations in the env gp90 protein that modify n-linked glycosylation of the chinese eiav vaccine strain enhance resistance to neutralizing antibodies. Virus Genes 2016, 52, 814–822. [Google Scholar] [CrossRef] [PubMed]
- Glebova, O.; Knyazev, S.; Melnyk, A.; Artyomenko, A.; Khudyakov, Y.; Zelikovsky, A.; Skums, P. Inference of genetic relatedness between viral quasispecies from sequencing data. BMC Genomics 2017, 18, 918. [Google Scholar] [CrossRef]
| 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | Mutation Rate (236D/-) | Morbidity (Horses) | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 3 | 3 | 3 | 3 | 3 | 3 | 3 | 3 | 3 | 3 | 4 | 4 | 4 | 4 | 4 | 4 | 4 | 4 | 4 | 4 | 5 | 5 | 5 | |||
| 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 0 | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 0 | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 0 | 1 | 2 | |||
| EIAVLN40 | N29 | A29 | I29 | T29 | I29 | L29 | V29 | P29 | E29 | V29 | E29 | M28 | Q29 | Q28 | S29 | D29 | N28 | N29 | T29 | W29 | I29 | P29 | K29 | R29 | C27 | N29 | E29 | T29 | W29 | A29 | R29 | V29 | 0.00% | 100% |
| I1 | R1 | S1 | Y2 | |||||||||||||||||||||||||||||||
| EIAVDLV34 | S9 | A16 | I15 | T15 | I16 | L16 | V16 | P16 | E16 | V10 | E16 | M16 | Q16 | Q14 | S16 | D14 | N16 | N16 | T16 | W16 | I16 | P16 | K16 | G8 | C16 | N14 | E13 | T16 | W16 | A16 | R12 | V16 | 6.25% | 100% |
| N7 | V1 | A1 | I6 | K1 | N1 | R8 | K2 | K3 | K4 | |||||||||||||||||||||||||
| R1 | -1 | |||||||||||||||||||||||||||||||||
| EIAVDLV62 | N9 | A16 | I16 | T16 | I16 | L16 | V16 | P16 | E16 | V9 | E16 | M16 | Q16 | Q14 | S16 | D9 | N16 | N16 | T16 | W16 | I16 | P16 | K16 | R11 | C16 | N13 | E13 | T16 | W16 | A16 | R12 | V16 | 18.5% | 100% |
| S7 | I7 | R2 | N4 | G5 | K3 | K2 | K4 | |||||||||||||||||||||||||||
| -3 | G1 | |||||||||||||||||||||||||||||||||
| EIAVDLV92 | N11 | A19 | I19 | T16 | I19 | L19 | V19 | P19 | E19 | V18 | E19 | M19 | Q19 | Q17 | S19 | D3 | N13 | N18 | T16 | W19 | I18 | P19 | K19 | R17 | C19 | N14 | E18 | T18 | W19 | A19 | R12 | V19 | 42.1% | 9.1% |
| S8 | A3 | I1 | R1 | N8 | K5 | D1 | A3 | V1 | G2 | K5 | K1 | A1 | K7 | |||||||||||||||||||||
| K1 | -8 | R1 | ||||||||||||||||||||||||||||||||
| EIAVDLV121 | N11 | A17 | I18 | T17 | I18 | L18 | V18 | P18 | E17 | V17 | E17 | M18 | Q18 | Q18 | R15 | -18 | K17 | N18 | T18 | W18 | I17 | P18 | K18 | R17 | C18 | K9 | E10 | T18 | W18 | A18 | K13 | V17 | 100% | 0 |
| S7 | T1 | A1 | K1 | I1 | K1 | S3 | S1 | V1 | G1 | N8 | K8 | R5 | I1 | |||||||||||||||||||||
| E1 |
| 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | Mutation Rate (236D/-) | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 3 | 3 | 3 | 3 | 3 | 3 | 3 | 3 | 3 | 3 | 4 | 4 | 4 | 4 | 4 | 4 | 4 | 4 | 4 | 4 | 5 | 5 | 5 | ||
| 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 0 | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 0 | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 0 | 1 | 2 | ||
| EIAVLN40 | N86 | A88 | I88 | T88 | I88 | L88 | V88 | P88 | E85 | V87 | E88 | M88 | Q87 | Q88 | S80 | D72 | N71 | N88 | T88 | W88 | I88 | P88 | K87 | R80 | C88 | N86 | E87 | T87 | W88 | A88 | R81 | V88 | 17.0% |
| S2 | V1 | I1 | H1 | R8 | -15 | K14 | N1 | G8 | K2 | K1 | I1 | K7 | |||||||||||||||||||||
| K1 | N1 | H1 | |||||||||||||||||||||||||||||||
| Q1 | |||||||||||||||||||||||||||||||||
| EIAVDLV34 | N51 | A97 | I98 | T98 | I98 | L97 | V98 | P97 | E94 | V60 | E96 | M98 | Q98 | Q82 | S86 | D63 | N75 | N98 | T98 | W95 | I95 | P97 | K96 | R57 | C97 | N85 | E79 | T96 | W94 | A94 | R75 | V94 | 30.6% |
| S47 | T1 | V1 | L1 | K3 | I38 | K2 | K8 | R10 | -30 | K22 | S1 | S2 | L1 | Q1 | G41 | W1 | K10 | K17 | A1 | C3 | C2 | K20 | I4 | ||||||||||
| D1 | R8 | K1 | N5 | Y1 | C2 | C1 | P1 | S3 | G2 | I1 | R1 | S1 | T3 | ||||||||||||||||||||
| N1 | T1 | ||||||||||||||||||||||||||||||||
| EIAVDLV121 | N76 | A95 | I95 | T92 | I95 | L95 | V94 | P95 | E92 | I10 | E93 | M94 | Q94 | Q91 | R54 | -74 | K69 | N95 | T95 | W95 | I95 | P95 | K95 | R68 | C95 | N74 | E83 | T94 | W95 | A93 | K55 | V95 | 77.9% |
| S18 | A3 | I1 | K1 | V84 | K1 | V1 | H1 | R3 | S40 | D19 | N23 | G25 | K21 | K11 | A1 | T2 | R40 | ||||||||||||||||
| K1 | D1 | L1 | V1 | K1 | N1 | N2 | S2 | K2 | G1 | ||||||||||||||||||||||||
| Q1 | E1 | ||||||||||||||||||||||||||||||||
| pLGFD3-8 | N100 | A100 | I100 | T100 | I100 | L100 | V100 | P100 | E99 | V97 | E99 | M100 | Q98 | Q100 | S87 | -100 | K100 | N100 | T97 | W98 | I99 | P100 | K100 | G88 | C99 | K99 | K99 | T100 | W99 | A100 | R97 | V100 | 100% |
| Q1 | I3 | K1 | H1 | R13 | I3 | C2 | V1 | R12 | G1 | N1 | E1 | C1 | K2 | ||||||||||||||||||||
| R1 | M1 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liu, C.; Wang, X.-F.; Wang, Y.; Chen, J.; Zhong, Z.; Lin, Y.; Wang, X. Characterization of EIAV env Quasispecies during Long-Term Passage In Vitro: Gradual Loss of Pathogenicity. Viruses 2019, 11, 380. https://doi.org/10.3390/v11040380
Liu C, Wang X-F, Wang Y, Chen J, Zhong Z, Lin Y, Wang X. Characterization of EIAV env Quasispecies during Long-Term Passage In Vitro: Gradual Loss of Pathogenicity. Viruses. 2019; 11(4):380. https://doi.org/10.3390/v11040380
Chicago/Turabian StyleLiu, Cong, Xue-Feng Wang, Yan Wang, Jie Chen, Zhaohua Zhong, Yuezhi Lin, and Xiaojun Wang. 2019. "Characterization of EIAV env Quasispecies during Long-Term Passage In Vitro: Gradual Loss of Pathogenicity" Viruses 11, no. 4: 380. https://doi.org/10.3390/v11040380
APA StyleLiu, C., Wang, X.-F., Wang, Y., Chen, J., Zhong, Z., Lin, Y., & Wang, X. (2019). Characterization of EIAV env Quasispecies during Long-Term Passage In Vitro: Gradual Loss of Pathogenicity. Viruses, 11(4), 380. https://doi.org/10.3390/v11040380






