Avian Influenza Virus Subtype H9N2 Affects Intestinal Microbiota, Barrier Structure Injury, and Inflammatory Intestinal Disease in the Chicken Ileum
Abstract
1. Introduction
2. Materials and Methods
2.1. Ethics Statement
2.2. Virus Subtype and Experimental Animals
2.3. Animal Experiments
2.4. Extraction of Metagenomic DNA
2.5. Illumina Sequencing
2.6. Histological Examination of Intestinal Segments and Villus Conditions
2.7. Extraction of Total RNA
2.8. Quantitative Real-Time PCR
2.9. Determination of Butyrate
2.10. Statistics
3. Results
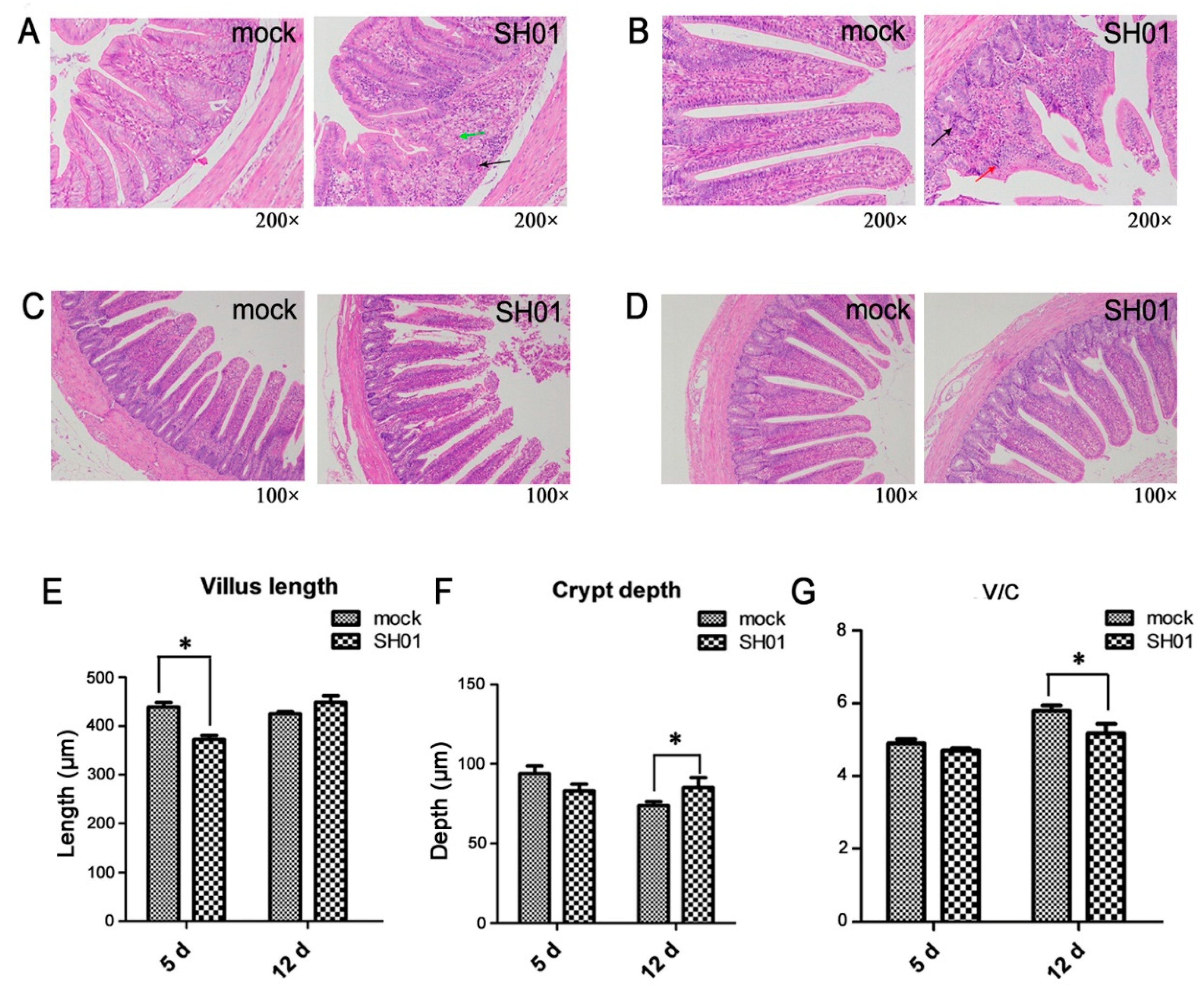
3.1. H9N2 AIV Infection Causes Intestinal Structure Injury
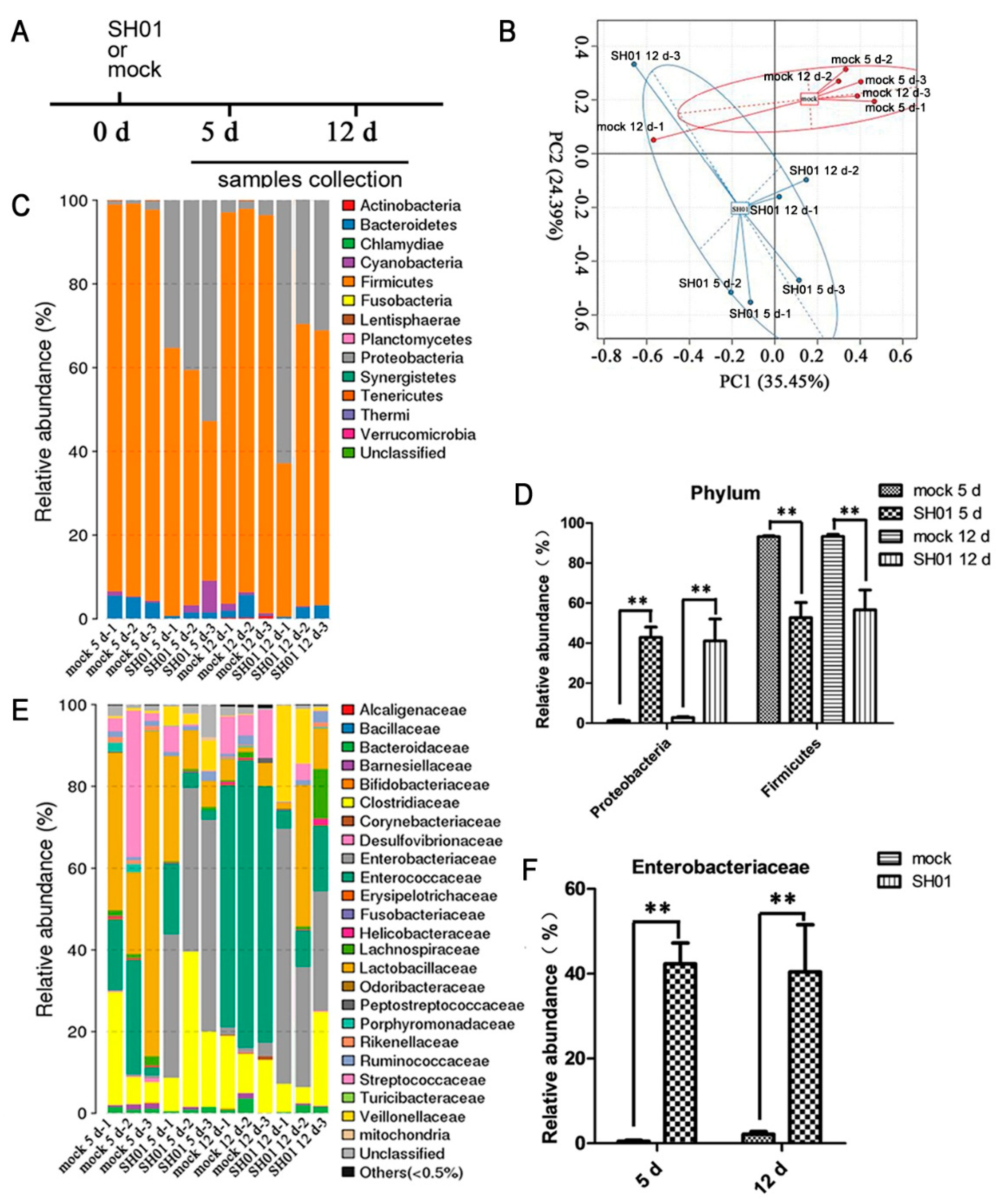
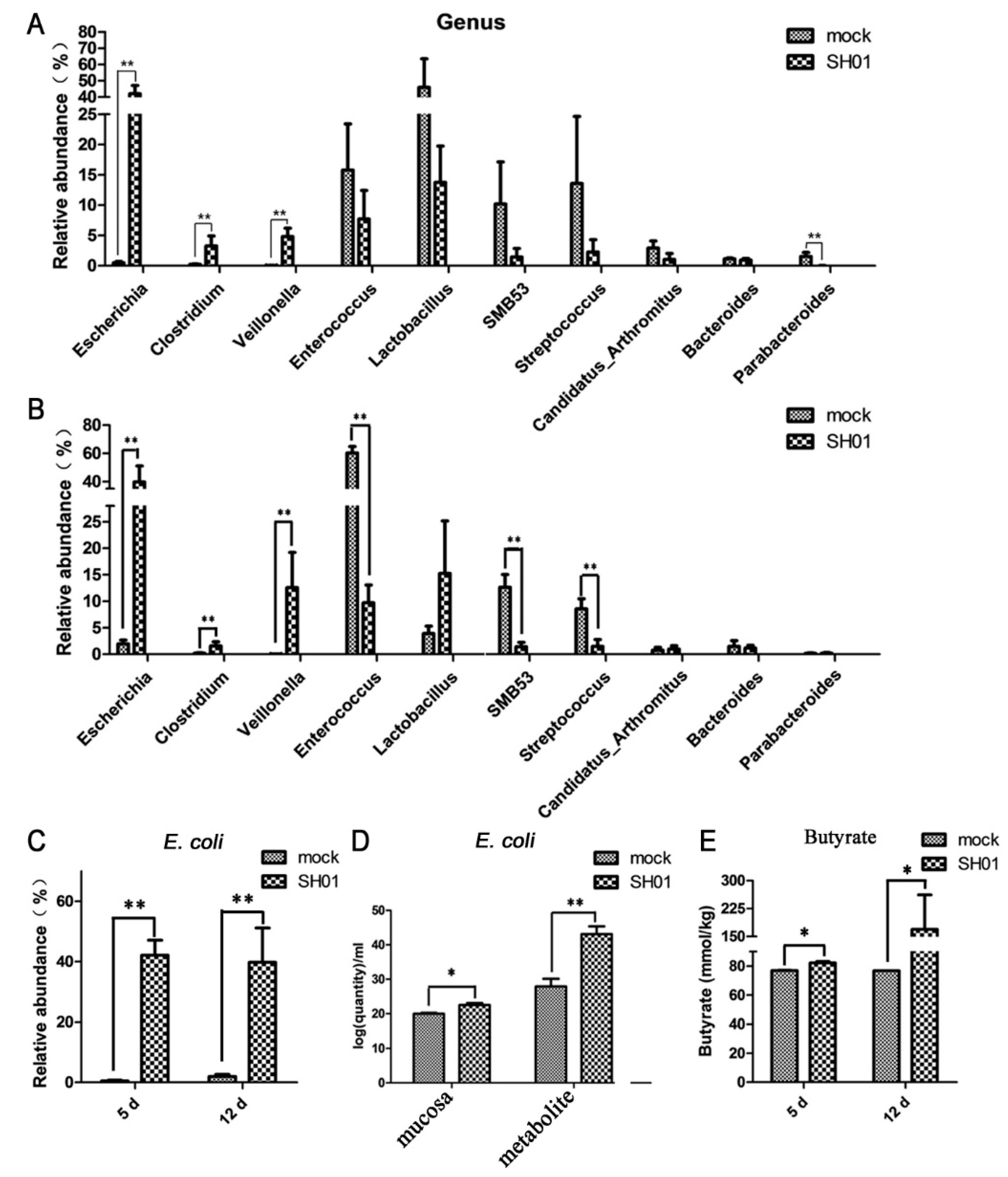
3.2. Intestinal Bacterial Microbiota Composition Differs between H9N2-Infected and Control Groups
3.3. Abundance of E. coli Increased Sharply in the Ileum after H9N2 AIV Infection
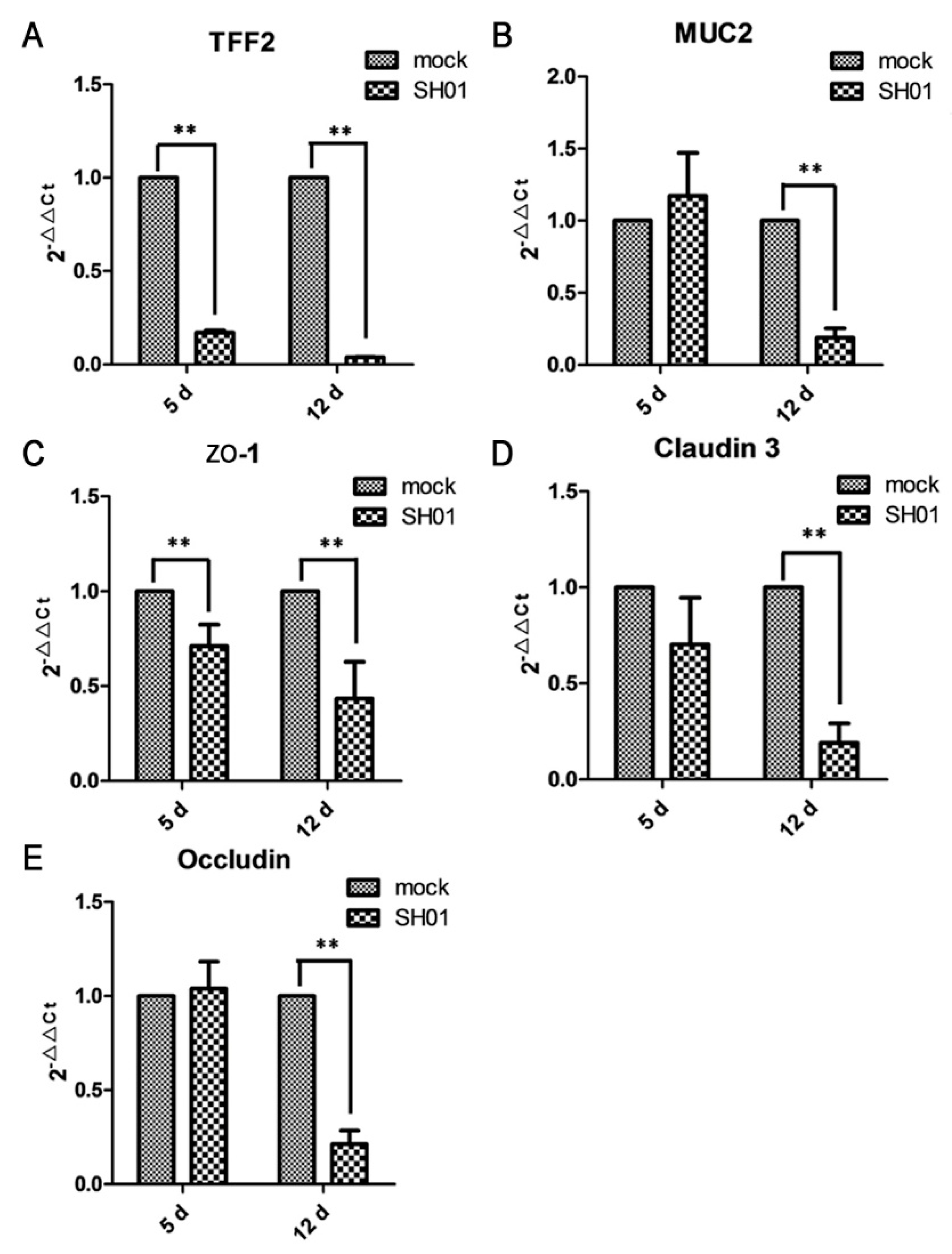
3.4. H9N2 AIV Infection Damages Ileal Mucous Layer Construction and Tight Junctions
3.5. H9N2 AIV Infection Promotes mRNA Expression of Proinflammatory Cytokines IFN-γ, IL-22, IFN-α, and IL-17A
4. Discussion
Author Contributions
Acknowledgments
Conflicts of Interest
References
- Peiris, M.; Yuen, K.Y.; Leung, C.W.; Chan, K.H.; Ip, P.L.S.; Lai, R.W.M.; Orr, W.K.; Shortridge, K.F. Human infection with influenza H9N2. Lancet (North American Edition) 1999, 354, 916–917. [Google Scholar] [CrossRef]
- Biswas, P.K.; Christensen, J.P.; Ahmed, S.S.U.; Barua, H.; Das, A.; Rahman, M.H.; Giasuddin, M.; Hannan, A.S.M.A.; Habib, M.A.; Ahad, A.; et al. Avian influenza outbreaks in chickens, Bangladesh. Emerg. Infect. Dis. 2008, 14, 1909–1912. [Google Scholar] [CrossRef] [PubMed]
- Barbour, E.K.; Mastori, F.A.; Nour, A.M.A.; Shaib, H.A.; Jaber, L.S.; Yaghi, R.H.; Sabra, A.; Sleiman, F.T.; Sawaya, R.K.; Niedzwieck, A.; et al. Standardisation of a new model of H9N2/Escherichia coli challenge in broilers in the Lebanon. Vet. Ital. 2009, 45, 317–322. [Google Scholar] [PubMed]
- Nagarajan, S.; Rajukumar, K.; Tosh, C.; Ramaswamy, V.; Purohit, K.; Saxena, G.; Behera, P.; Pattnaik, B.; Pradhan, H.K.; Dubey, S.C. Isolation and pathotyping of H9N2 avian influenza viruses in Indian poultry. Vet. Microbiol. 2009, 133, 154–163. [Google Scholar] [CrossRef] [PubMed]
- Monto, A.S.; Gravenstein, S.; Elliott, M.; Colopy, M.; Schweinle, J. Clinical signs and symptoms predicting influenza infection. Arch. Intern. Med. 2000, 160, 3243–3247. [Google Scholar] [CrossRef] [PubMed]
- Baden, L.R.; Drazen, J.M.; Kritek, P.A.; Curfman, G.D.; Morrissey, S.; Campion, E.W. H1N1 influenza a disease-information for health professionals. N. Engl. J. Med. 2009, 360, 2666–2667. [Google Scholar] [CrossRef] [PubMed]
- Dilantika, C.; Sedyaningsih, E.R.; Kasper, M.R.; Agtini, M.; Listiyaningsih, E.; Uyeki, T.M.; Burgess, T.H.; Blair, P.J.; Putnam, S.D. Influenza virus infection among pediatric patients reporting diarrhea and influenza-like illness. BMC Infect. Dis. 2010, 10, 3. [Google Scholar] [CrossRef] [PubMed]
- Yitbarek, A.; Weese, J.S.; Alkie, T.N.; Parkinson, J.; Sharif, S. Influenza A virus subtype H9N2 infection disrupts the composition of intestinal microbiota of chickens. FEMS Microbiol. Ecol. 2018, 94, 1. [Google Scholar] [CrossRef] [PubMed]
- Deriu, E.; Boxx, G.M.; He, X.S.; Pan, C.; Benavidez, S.D.; Cen, L.J.; Rozengurt, N.; Shi, W.Y.; Cheng, G.H. Influenza virus affects intestinal microbiota and secondary Salmonella infection in the gut through Type I interferons. PLOS Pathog. 2016, 12, e1005572. [Google Scholar] [CrossRef] [PubMed]
- Hooper, L.V.; Gordon, J.I. Commensal host-bacterial relationships in the gut. Science 2001, 292, 1115–1118. [Google Scholar] [CrossRef] [PubMed]
- Backhed, F.; Ley, R.E.; Sonnenburg, J.L.; Peterson, D.A.; Gordon, J.I. Host-bacterial mutualism in the human intestine. Science 2005, 307, 1915–1920. [Google Scholar] [CrossRef] [PubMed]
- Chervonsky, A. Innate receptors and microbes in induction of autoimmunity. Curr. Opin. Immunol. 2009, 21, 641–647. [Google Scholar] [CrossRef] [PubMed]
- Van der Waaij, D.; Berghuis-de Vries, J.M.; Lekkerkerk, L. Colonization resistance of the digestive tract in conventional and antibiotic-treated mice. J. Hyg. 1971, 69, 405–411. [Google Scholar] [CrossRef] [PubMed]
- Garrett, W.S.; Lord, G.M.; Punit, S.; Lugo-Villarino, G.; Mazmanian, S.K.; Ito, S.; Glickman, J.N.; Glimcher, L.H. Communicable ulcerative colitis induced by T-bet deficiency in the innate immune system. Cell 2007, 131, 33–45. [Google Scholar] [CrossRef] [PubMed]
- Winter, S.E.; Winter, M.G.; Xavier, M.N.; Thiennimitr, P.; Poon, V.; Keestra, A.M.; Laughlin, R.C.; Gomez, G.; Wu, J.; Lawhon, S.D.; et al. Host-derived nitrate boosts growth of E. coli in the inflamed gut. Science 2013, 339, 708–711. [Google Scholar] [CrossRef] [PubMed]
- Turnbaugh, P.J.; Ley, R.E.; Mahowald, M.A.; Magrini, V.; Mardis, E.R.; Gordon, J.I. An obesity-associated gut microbiome with increased capacity for energy harvest. Nature 2006, 444, 1027–1031. [Google Scholar] [CrossRef] [PubMed]
- Santhakumar, D.; Rubbenstroth, D.; Martinez-Sobrido, L.; Munir, M. Avian interferons and their antiviral effectors. Front. Immunol. 2017, 8, 49. [Google Scholar] [CrossRef] [PubMed]
- Shahangian, A.; Chow, E.K.; Tian, X.L.; Kang, J.R.; Ghaffari, A.; Liu, S.Y.; Belperio, J.A.; Cheng, G.H.; Deng, J.C. Type I IFNs mediate development of postinfluenza bacterial pneumonia in mice. J. Clin. Investig. 2009, 119, 1910–1920. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Li, F.Q.; Wei, H.M.; Lian, Z.X.; Sun, R.; Tian, Z.G. Respiratory influenza virus infection induces intestinal immune injury via microbiota-mediated Th17 cell-dependent inflammation. J. Exp. Med. 2014, 211, 2397–2410. [Google Scholar] [CrossRef] [PubMed]
- Human Microbiome Project Consortiu. Structure, function and diversity of the healthy human microbiome. Nature 2012, 486, 207–214. [Google Scholar]
- Ruby, E.G. Symbiotic conversations are revealed under genetic interrogation. Nat. Rev. Microbiol. 2008, 6, 752–762. [Google Scholar] [CrossRef] [PubMed]
- Su, X.N.; Xie, Q.M.; Liao, C.T.; Yan, Z.Q.; Chen, W.G.; Bi, Y.Z.; Chen, F. Sequence and phylogenetic analysis of hemagglutinin genes of H9N2 influenza viruses isolated from chicken in China from 2013 to 2015. J. Integr. Agric. 2016, 15, 2604–2612. [Google Scholar] [CrossRef]
- Liu, L.; Li, Y.; Li, S.; Hu, N.; He, Y.; Pong, R.; Lin, D.; Lu, L.; Law, M. Comparison of next-generation sequencing systems. J. Biomed. Biotechnol. 2012, 2012, 11. [Google Scholar] [CrossRef] [PubMed]
- He, C.; Yang, Z.; Cheng, D.D.; Xie, C.; Zhu, Y.; Ge, Z.M.; Luo, Z.J.; Lu, N.H. Helicobacter pylori infection aggravates diet-induced insulin resistance in association with gut microbiota of mice. Ebiomedicine 2016, 12, 247–254. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.Y.; Li, H.M.; Xie, Q.M.; Shang, H.Q.; Ji, J.; Bai, S.W.; Cao, Y.C.; Ma, Y.L.; Bi, Y.Z. Transcriptional profiling of host gene expression in chicken liver tissues infected with oncogenic Marek’s disease virus. J. Gen. Virol. 2011, 92, 2724–2733. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.Y.; Xie, Q.M.; Xue, Y.; Ji, J.; Chang, S.; Ma, J.Y.; Bi, Y.Z. Characterization of cytotoxicity-related gene expression in response to virulent Marek’s disease virus infection in the bursa of Fabricius. Res. Vet. Sci. 2013, 94, 496–503. [Google Scholar] [CrossRef] [PubMed]
- Sirdaarta, J.; Maen, A.; Rayan, P.; Matthews, B.; Cock, I.E. High performance liquid chromatography-mass spectrometry analysis of high antioxidant Australian fruits with antiproliferative activity against cancer cells. Pharmacogn. Mag. 2016, 122, S181–S194. [Google Scholar]
- Seksik, P.; Rigottier-Gois, L.; Gramet, G.; Sutren, M.; Pochart, P.; Marteau, P.; Jian, R.; Dore, J. Alterations of the dominant faecal bacterial groups in patients with Crohn’s disease of the colon. Gut 2003, 52, 237–242. [Google Scholar] [CrossRef] [PubMed]
- Gophna, U.; Sommerfeld, K.; Gophna, S.; Doolittle, W.F.; van Zanten, S.J.O.V. Differences between tissue-associated intestinal microfloras of patients with Crohn’s disease and ulcerative colitis. J. Clin. Microbiol. 2006, 44, 4136–4141. [Google Scholar] [CrossRef] [PubMed]
- Garrett, W.S.; Gallini, C.A.; Yatsunenko, T.; Michaud, M.; DuBois, A.; Delaney, M.L.; Punit, S.; Karlsson, M.; Bry, L.; Glickman, J.N.; et al. Enterobacteriaceae act in concert with the gut microbiota to induce spontaneous and maternally transmitted colitis. Cell Host Microbe 2010, 8, 292–300. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.; Faris, L.; Cox, C.M.; Sumners, L.H.; Jenkins, M.C.; Fetterer, R.H.; Miska, K.B.; Dalloul, R.A. Molecular characterization and immunological roles of avian IL-22 and its soluble receptor IL-22 binding protein. Cytokine 2012, 60, 815–827. [Google Scholar] [CrossRef] [PubMed]
- Del Cacho, E.; Gallego, M.; Lillehoj, H.S.; Quilez, J.; Lillehoj, E.P.; Ramo, A.; Sanchez-Acedo, C. IL-17A regulates Eimeria tenella schizont maturation and migration in avian coccidiosis. Vet. Res. 2014, 45, 25. [Google Scholar] [CrossRef] [PubMed]
- Choi, J.H.; Kim, G.B.; Cha, C.J. Spatial heterogeneity and stability of bacterial community in the gastrointestinal tracts of broiler chickens. Poult. Sci. 2014, 93, 1942–1950. [Google Scholar] [CrossRef] [PubMed]
- Baumgart, M.; Dogan, B.; Rishniw, M.; Weitzman, G.; Bosworth, B.; Yantiss, R.; Orsi, R.H.; Wiedmann, M.; McDonough, P.; Kim, S.G.; et al. Culture independent analysis of ileal mucosa reveals a selective increase in invasive Escherichia coli of novel phylogeny relative to depletion of Clostridiales in Crohn’s disease involving the ileum. ISME J. 2007, 1, 403–418. [Google Scholar] [CrossRef] [PubMed]
- Pei, J.; Sekellick, M.J.; Marcus, P.I.; Choi, I.; Collisson, E.W. Chicken interferon type I inhibits infectious bronchitis virus replication and associated respiratory illness. J. Interferon Cytokine Res. 2001, 21, 1071–1077. [Google Scholar] [CrossRef] [PubMed]
- Levy, A.M.; Heller, E.D.; Leitner, G.; Davidson, I. Effect of native chicken interferon on MDV replication. Acta Virol. 1999, 43, 121–127. [Google Scholar] [PubMed]
- Xia, C.; Liu, J.; Wu, Z.G.; Lin, C.Y.; Wang, M. The interferon-alpha genes from three chicken lines and its effects on H9N2 influenza viruses. Anim. Biotechnol. 2004, 15, 77–88. [Google Scholar] [CrossRef] [PubMed]
- Zhao, G.H.; Cheng, W.Y.; Wang, W.; Jia, Y.Q.; Fang, Y.Q.; Du, S.Z.; Yu, S.K. The expression dynamics of IL-17 and Th17 response relative cytokines in the trachea and spleen of chickens after infection with Cryptosporidium baileyi. Parasite Vector 2014, 7, 212. [Google Scholar] [CrossRef] [PubMed]
- Mallick, A.I.; Parvizi, P.; Read, L.R.; Nagy, E.; Behboudi, S.; Sharif, S. Enhancement of immunogenicity of a virosome-based avian influenza vaccine in chickens by incorporating CpG-ODN. Vaccine 2011, 29, 1657–1665. [Google Scholar] [CrossRef] [PubMed]
- Baumler, A.J.; Sperandio, V. Interactions between the microbiota and pathogenic bacteria in the gut. Nature 2016, 535, 85–93. [Google Scholar] [CrossRef] [PubMed]
- Raffatellu, M.; George, M.D.; Akiyama, Y.; Hornsby, M.J.; Nuccio, S.P.; Paixao, T.A.; Butler, B.P.; Chu, H.T.; Santos, R.L.; Berger, T.; et al. Lipocalin-2 resistance confers an advantage to salmonella enterica serotype Typhimurium for growth and survival in the inflamed intestine. Cell Host Microbe 2009, 5, 476–486. [Google Scholar] [CrossRef] [PubMed]
- Hansson, G.C. Role of mucus layers in gut infection and inflammation. Curr. Opin. Microbiol. 2012, 15, 57–62. [Google Scholar] [CrossRef] [PubMed]
- Taupin, D.R.; Kinoshita, K.; Podolsky, D.K. Intestinal trefoil factor confers colonic epithelial resistance to apoptosis. Proc. Natl. Acad. Sci. USA 2000, 97, 799–804. [Google Scholar] [CrossRef] [PubMed]
- Sands, B.E.; Podolsky, D.K. The trefoil peptide family. Annu. Rev. Physiol. 1996, 58, 253–273. [Google Scholar] [CrossRef] [PubMed]
- Gadde, U.D.; Oh, S.; Lee, Y.; Davis, E.; Zimmerman, N.; Rehberger, T.; Lillehoj, H.S. Dietary bacillus subtilis-based direct-fed microbials alleviate LPS-induced intestinal immunological stress and improve intestinal barrier gene expression in commercial broiler chickens. Res. Vet. Sci. 2017, 114, 236–243. [Google Scholar] [CrossRef] [PubMed]
- Kitessa, S.M.; Nattrass, G.S.; Forder, R.E.A.; McGrice, H.A.; Wu, S.B.; Hughes, R.J. Mucin gene mRNA levels in broilers challenged with Eimeria and/or Clostridium perfringens. Avian Dis. 2014, 58, 408–414. [Google Scholar] [CrossRef] [PubMed]
- Giraud, A.S.; Pereira, P.M.; Thim, L.; Parker, L.M.; Judd, L.M. TFF-2 inhibits iNOS/NO in monocytes, and nitrated protein in healing colon after colitis. Peptides 2004, 25, 803–809. [Google Scholar] [CrossRef] [PubMed]
- Furuse, M.; Hirase, T.; Itoh, M.; Nagafuchi, A.; Yonemura, S.; Tsukita, S.; Tsukita, S. Occludin: A novel integral membrane protein localizing at tight junctions. J. Cell Biol. 1993, 123, 1777–1788. [Google Scholar] [CrossRef] [PubMed]
- Furuse, M.; Fujita, K.; Hiriagi, T.; Fujimoto, K.; Tsukita, S. Claudin-1 and -2: Novel integral membrane proteins localizing at tight junctions with no sequence similarity to occludin. J. Cell Biol. 1998, 141, 1539–1549. [Google Scholar] [CrossRef] [PubMed]
- Gumbiner, B.; Lowenkopf, T.; Apatira, D. Identification of a 160-kDa polypeptide that binds to the tight junction protein ZO-1. Proc. Natl. Acad. Sci. USA 1991, 88, 3460–3464. [Google Scholar] [CrossRef] [PubMed]
- Willott, E.; Balda, M.S.; Fanning, A.S.; Jameson, B.; Van Itallie, C.; Anderson, J.M. The tight junction protein ZO-1 is homologous to the drosophila discs-large tumor suppressor protein of septate junctions. Proc. Natl. Acad. Sci. USA 1993, 90, 7834–7838. [Google Scholar] [CrossRef] [PubMed]
- Haskkins, J.; Gu, L.; Wittchen, E.S.; Hibbard, J.; Stevenson, B.R. ZO-3, a novel member of the MAGUK protein family found at the tight junction, interacts with ZO-1 and occludin. J. Cell Biol. 1998, 141, 199–208. [Google Scholar] [CrossRef]
- Suzuki, T. Regulation of intestinal epithelial permeability by tight junctions. Cell. Mol. Life Sci. 2013, 70, 631–659. [Google Scholar] [CrossRef] [PubMed]
- Turner, J.R. Intestinal mucosal barrier function in health and disease. Nat. Rev. Immunol. 2009, 9, 799–809. [Google Scholar] [CrossRef] [PubMed]
- Hennings, S.J.; Hird, F.J. Concentrations and metabolism of volatile fatty acids in the fermentative organs of two species of kangaroo and the guinea-pig. Br. J. Nutr. 1970, 24, 145–155. [Google Scholar] [CrossRef] [PubMed]
- Firmansyah, A.; Penn, D.; Lebenthal, E. Isolated colonocyte metabolism of glucose, glutamine, n-butyrate, and beta-hydroxybutyrate in malnutrition. Gastroenterology 1989, 97, 622–629. [Google Scholar] [CrossRef]
- Song, B.C.; Li, H.X.; Wu, Y.Y.; Zhen, W.R.; Wang, Z.; Xia, Z.F.; Guo, Y.M. Effect of microencapsulated sodium butyrate dietary supplementation on growth performance and intestinal barrier function of broiler chickens infected with necrotic enteritis. Anim. Feed Sci. Technol. 2017, 232, 6–15. [Google Scholar] [CrossRef]
- Zumbrun, S.D.; Melton-Celsa, A.R.; Smith, M.A.; Gilbreath, J.J.; Merrell, D.S.; O’Brien, A.D. Dietary choice affects Shiga toxin-producing Escherichia coli (STEC) O157:H7 colonization and disease. Proc. Natl. Acad. Sci. USA 2013, 110, E2126–E2133. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.D.; Bayir, H.O.; Cosby, D.E.; Cox, N.A.; Williams, S.M.; Fowler, J. Evaluation of encapsulated sodium butyrate on growth performance, energy digestibility, gut development, and Salmonella colonization in broilers. Poult. Sci. 2017, 96, 3638–3644. [Google Scholar] [CrossRef] [PubMed]

| Name | Sense Strand/Sense Primer (5′–3′) | Antisense Strand/Antisense Primer (5′–3′) |
|---|---|---|
| TFF2 | CCCTGCTGATCCTCGTAT | GCTGTTATTTCCCAGTTGA |
| MUC2 | AATGCTGAGTTCTTGCCTAA | GTTGCAGTTCATATCCTGGT |
| ZO-1 | GCCTGAATCAAACCCAGCAA | TATGCGGCGGTAAGGATGAT |
| Claudin-3 | GAAGGGCTGTGGATGAACTG | GAGACGATGGTGATCTTGGC |
| Occludin | GATGGACAGCATCAACGACC | CATGCGCTTGATGTGGAAGA |
| IFN-γ | ATCATACTGAGCCAGATTGTTTCG | TCTTTCACCTTCTTCACGCCAT |
| IL-22 | CAGGAATCGCACCTACACCT | TCATGTAGCAGCGGTTGTTC |
| IFN-α | CCAGCACCTCGAGCAAT | GGCGCTGTAATCGTTGTCT |
| IL-17A | CCATTCCAGGTGCGTGAACT | TTTCTTCTCCAGGCGGTACG |
| GAPDH | AGGCTGAGAACGGGAAACTTG | CACCTGCATCTGCCCATTTG |
| E. coli | GTTAATACCTTTGCTCATTGA | ACCAGGGTATCTTAATCCTGTT |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Li, H.; Liu, X.; Chen, F.; Zuo, K.; Wu, C.; Yan, Y.; Chen, W.; Lin, W.; Xie, Q. Avian Influenza Virus Subtype H9N2 Affects Intestinal Microbiota, Barrier Structure Injury, and Inflammatory Intestinal Disease in the Chicken Ileum. Viruses 2018, 10, 270. https://doi.org/10.3390/v10050270
Li H, Liu X, Chen F, Zuo K, Wu C, Yan Y, Chen W, Lin W, Xie Q. Avian Influenza Virus Subtype H9N2 Affects Intestinal Microbiota, Barrier Structure Injury, and Inflammatory Intestinal Disease in the Chicken Ileum. Viruses. 2018; 10(5):270. https://doi.org/10.3390/v10050270
Chicago/Turabian StyleLi, Hongxin, Xiaolin Liu, Feiyang Chen, Kejing Zuo, Che Wu, Yiming Yan, Weiguo Chen, Wencheng Lin, and Qingmei Xie. 2018. "Avian Influenza Virus Subtype H9N2 Affects Intestinal Microbiota, Barrier Structure Injury, and Inflammatory Intestinal Disease in the Chicken Ileum" Viruses 10, no. 5: 270. https://doi.org/10.3390/v10050270
APA StyleLi, H., Liu, X., Chen, F., Zuo, K., Wu, C., Yan, Y., Chen, W., Lin, W., & Xie, Q. (2018). Avian Influenza Virus Subtype H9N2 Affects Intestinal Microbiota, Barrier Structure Injury, and Inflammatory Intestinal Disease in the Chicken Ileum. Viruses, 10(5), 270. https://doi.org/10.3390/v10050270





