High Levels of Dual-Class Drug Resistance in HIV-Infected Children Failing First-Line Antiretroviral Therapy in Southern Ethiopia
Abstract
1. Introduction
2. Materials and Methods
2.1. Study Participants
2.2. Ethics Statement
2.3. Specimen Collection, Handling and Storage
2.4. Nucleic Acid Extraction and Viral Genotyping
2.5. Drug Resistance Genotyping
2.6. Statistical Analyses
3. Results
3.1. Patient Characteristics
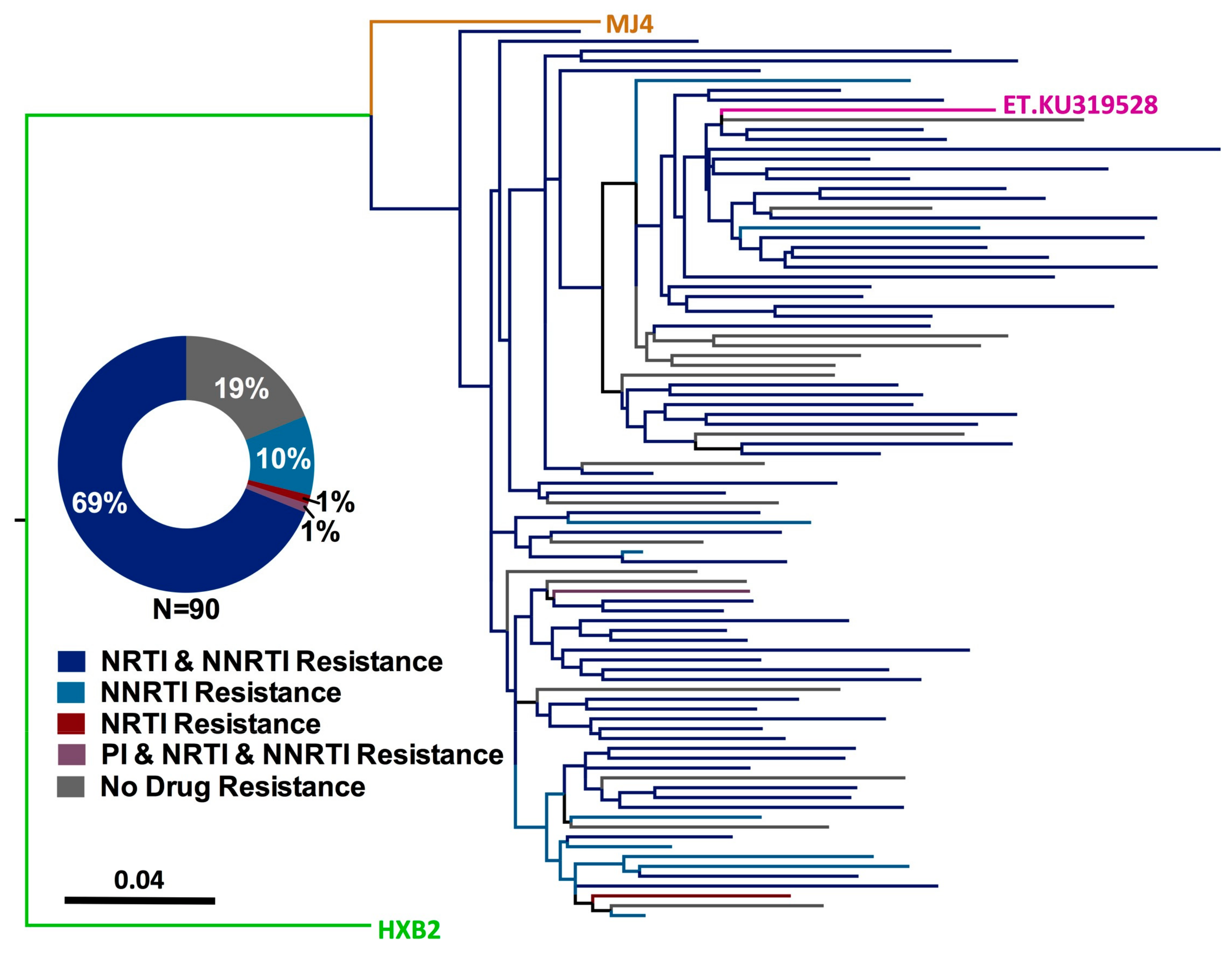
3.2. HIV-1 Genotyping from Dried Blood Spots
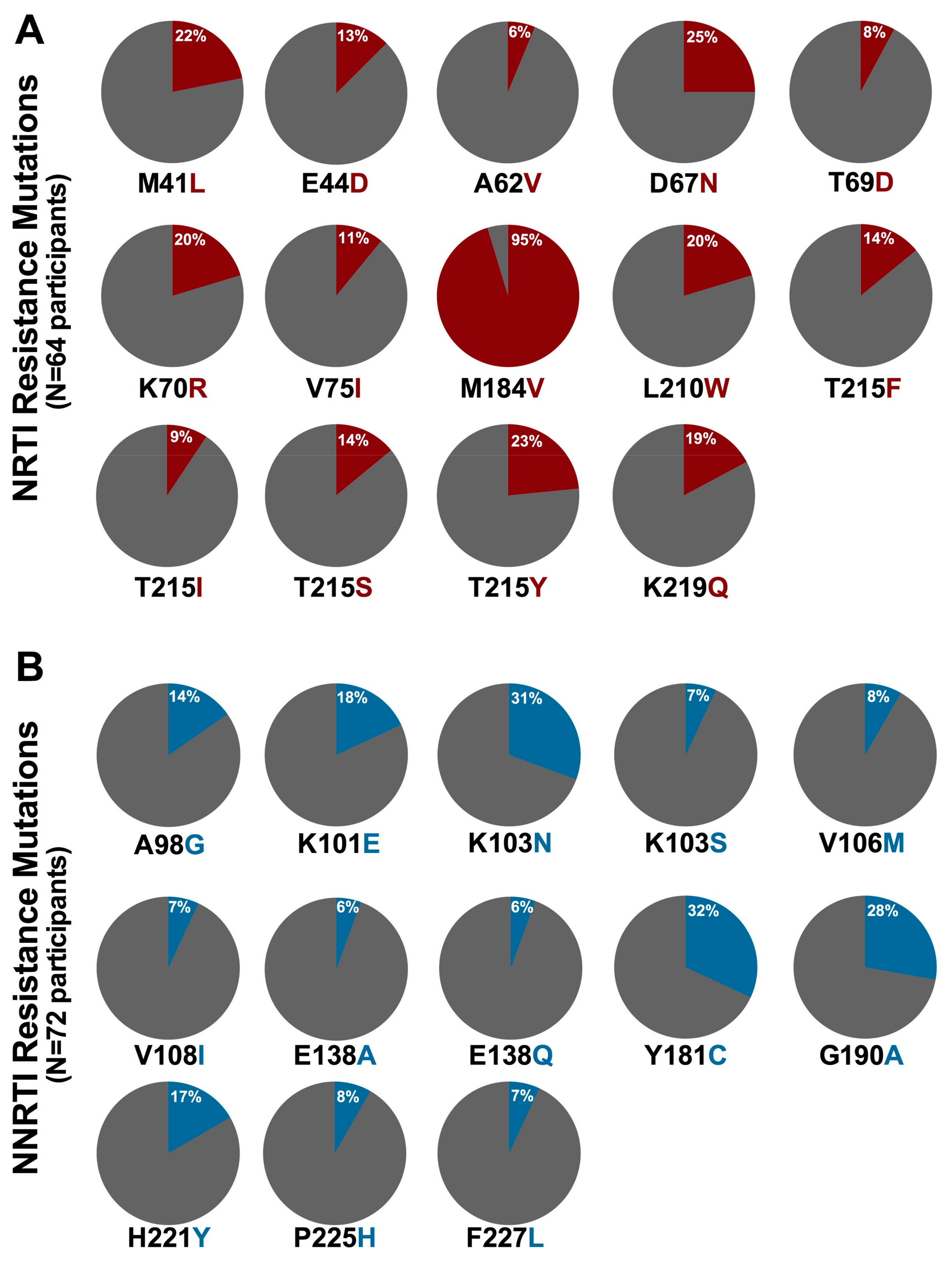
3.3. Prevalence of HIV-1 Drug Resistance among Ethiopian Children Exhibiting Virologic Treatment Failure of First-Line cART
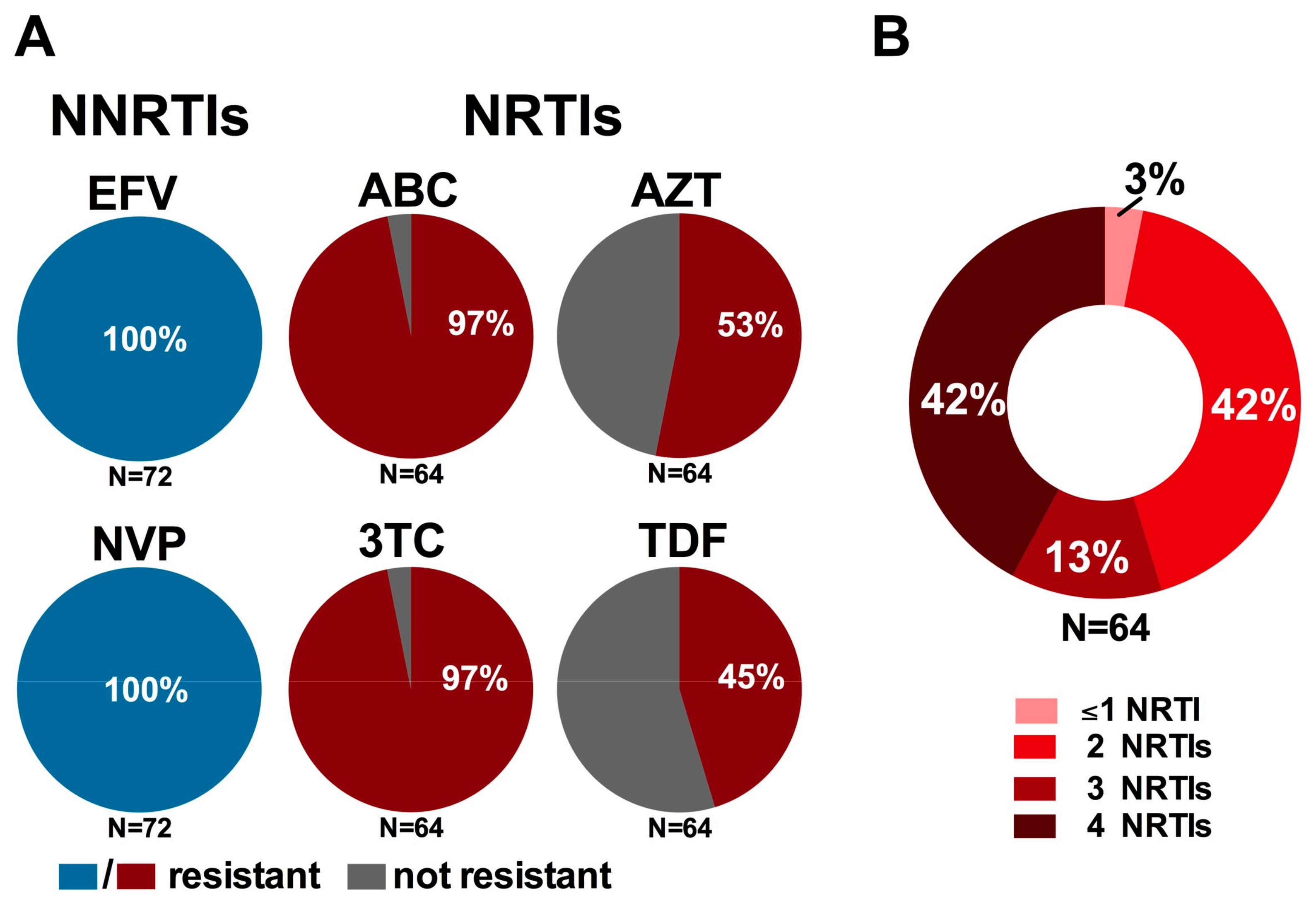
3.4. Impact on WHO-Recommended First- and Second-Line Regimens
3.5. Factors Associated with HIV-1 Drug Resistance among Ethiopian Children Experiencing Virologic Failure of First-Line cART
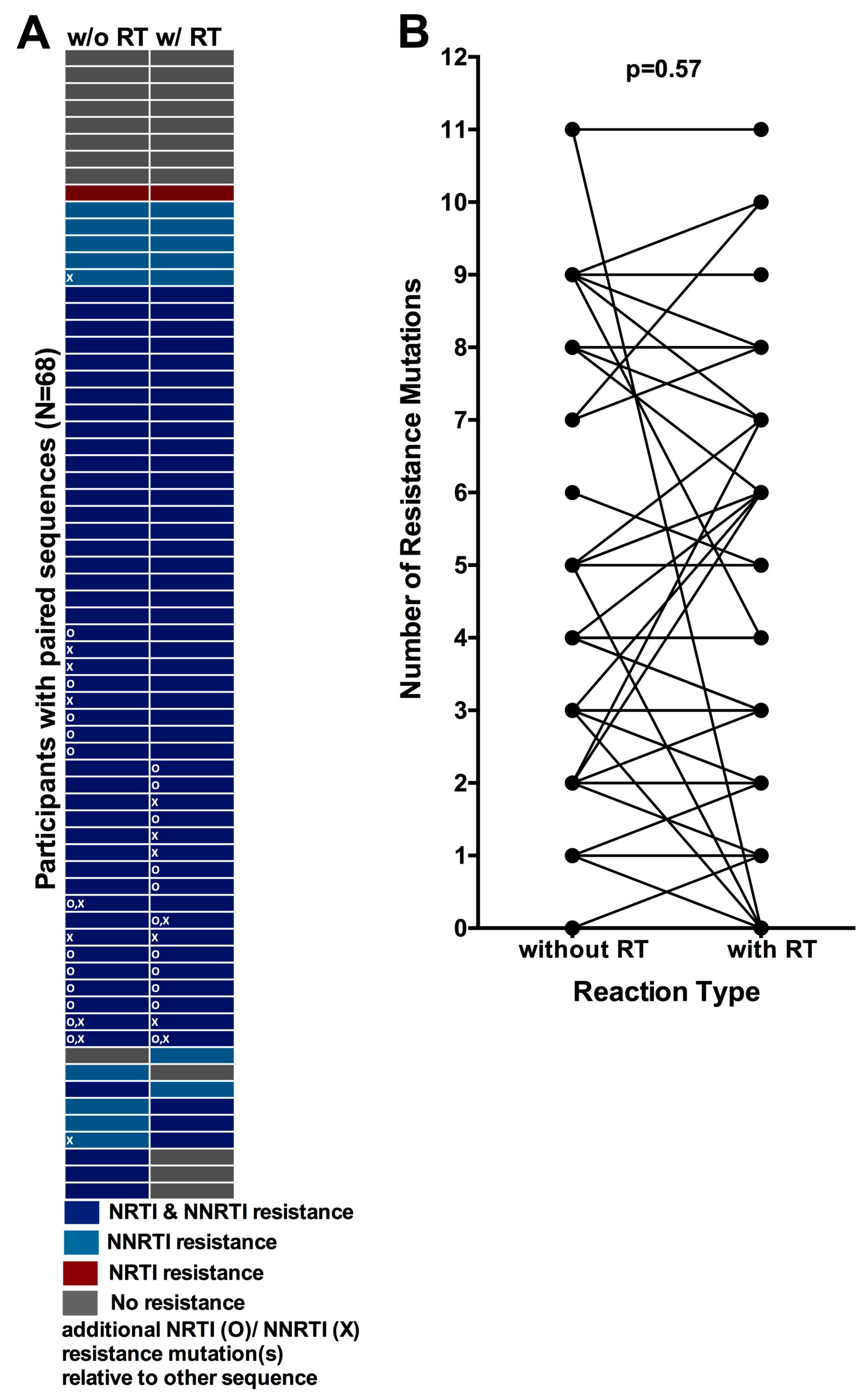
3.6. Comparison of HIV-1 Drug Resistance Genotypes Obtained with and without an Initial Reverse Transcription Step
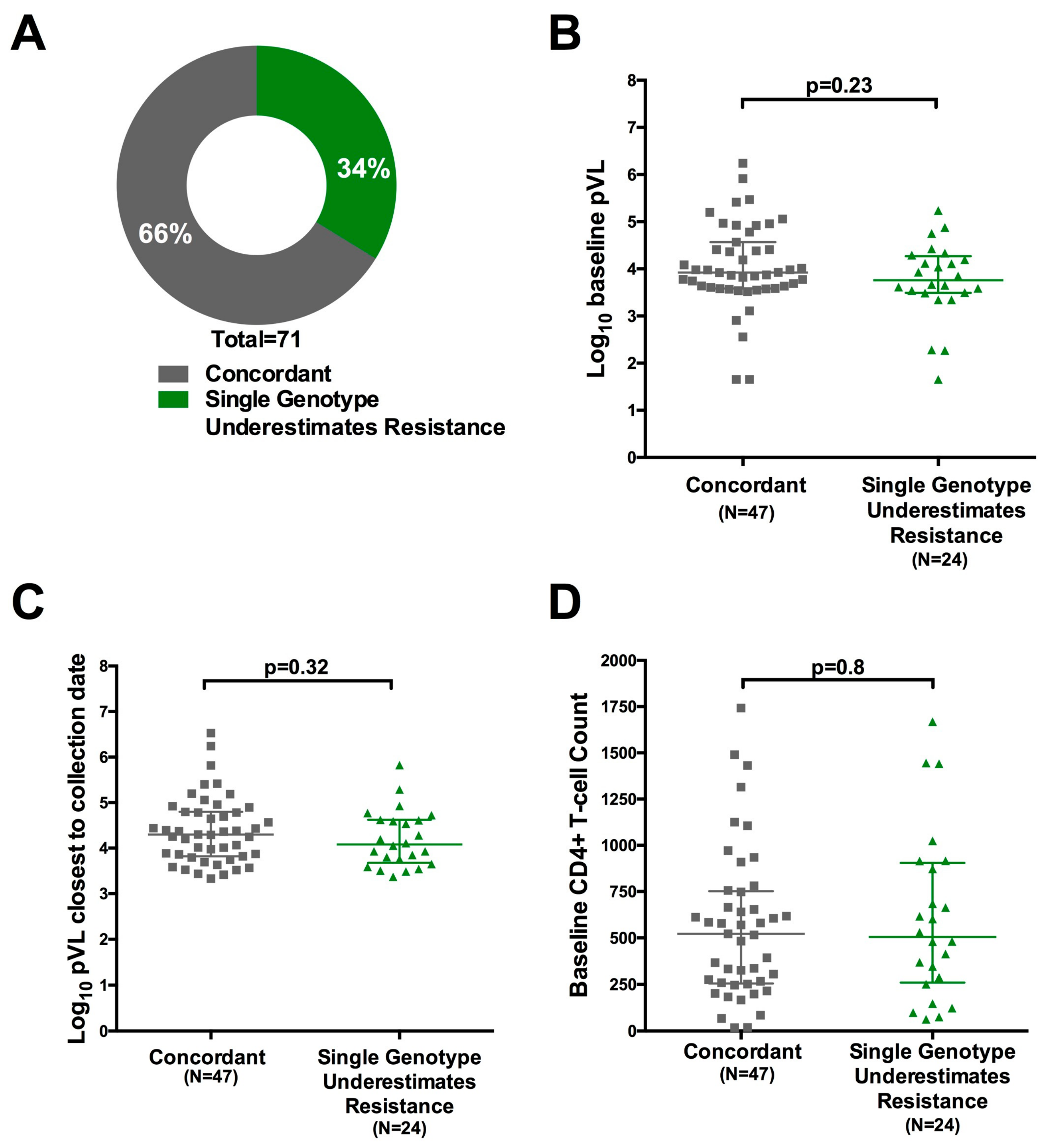
3.7. Quantification of the Degree of Underestimation of Resistance from Single versus Replicate Genotypes from Dried Blood Spots
4. Discussion
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- The Joint United Nations Programme on HIV/AIDS (UNAIDS). Fact Sheet—Global HIV Statistics; UNAIDS Joint United Nations Programme on HIV/AIDS: Geneva, Switzerland, 2017. [Google Scholar]
- The Joint United Nations Programme on HIV/AIDS (UNAIDS). Global AIDS Update; UNAIDS Joint United Nations Programme on HIV/AIDS: Geneva, Switzerland, 2016. [Google Scholar]
- Federal Ministry of Health Ethiopian Public Health Institute. HIV Related Estimates and Projections for Ethiopia-2017; Public Health Institute, Ethiopian Federal Ministry of Health: Addis Ababa, Ethiopia, 2017.
- Resino, S.; Bellon, J.M.; Resino, R.; Navarro, M.L.; Tomas Ramos, J.; de Jose, M.I.; Mellado, M.J.; Munoz-Fernandez, M.A. Extensive implementation of highly active antiretroviral therapy shows great effect on survival and surrogate markers in vertically HIV-infected children. Clin. Infect. Dis. 2004, 38, 1605–1612. [Google Scholar] [CrossRef] [PubMed]
- Berk, D.R.; Falkovitz-Halpern, M.S.; Hill, D.W.; Albin, C.; Arrieta, A.; Bork, J.M.; Cohan, D.; Nilson, B.; Petru, A.; Ruiz, J.; et al. Temporal trends in early clinical manifestations of perinatal HIV infection in a population-based cohort. JAMA 2005, 293, 2221–2231. [Google Scholar] [CrossRef] [PubMed]
- George, E.; Noel, F.; Bois, G.; Cassagnol, R.; Estavien, L.; Rouzier Pde, M.; Verdier, R.I.; Johnson, W.D.; Pape, J.W.; Fitzgerald, D.W.; et al. Antiretroviral therapy for HIV-1-infected children in Haiti. J. Infect. Dis. 2007, 195, 1411–1418. [Google Scholar] [CrossRef] [PubMed]
- O’Brien, D.P.; Sauvageot, D.; Zachariah, R.; Humblet, P. In resource-limited settings good early outcomes can be achieved in children using adult fixed-dose combination antiretroviral therapy. AIDS 2006, 20, 1955–1960. [Google Scholar] [CrossRef] [PubMed]
- Sauvageot, D.; Schaefer, M.; Olson, D.; Pujades-Rodriguez, M.; O’Brien, D.P. Antiretroviral therapy outcomes in resource-limited settings for HIV-infected children <5 years of age. Pediatrics 2010, 125, e1039–e1047. [Google Scholar] [CrossRef] [PubMed]
- Goetghebuer, T.; Haelterman, E.; Le Chenadec, J.; Dollfus, C.; Gibb, D.; Judd, A.; Green, H.; Galli, L.; Ramos, J.T.; Giaquinto, C.; et al. Effect of early antiretroviral therapy on the risk of AIDS/death in HIV-infected infants. AIDS 2009, 23, 597–604. [Google Scholar] [CrossRef] [PubMed]
- Musoke, P.M.; Mudiope, P.; Barlow-Mosha, L.N.; Ajuna, P.; Bagenda, D.; Mubiru, M.M.; Tylleskar, T.; Fowler, M.G. Growth, immune and viral responses in HIV infected African children receiving highly active antiretroviral therapy: A prospective cohort study. BMC Pediatr. 2010, 10, 56. [Google Scholar] [CrossRef] [PubMed]
- Verweel, G.; van Rossum, A.M.; Hartwig, N.G.; Wolfs, T.F.; Scherpbier, H.J.; de Groot, R. Treatment with highly active antiretroviral therapy in human immunodeficiency virus type 1-infected children is associated with a sustained effect on growth. Pediatrics 2002, 109, E25. [Google Scholar] [CrossRef] [PubMed]
- Connor, E.M.; Sperling, R.S.; Gelber, R.; Kiselev, P.; Scott, G.; O’Sullivan, M.J.; VanDyke, R.; Bey, M.; Shearer, W.; Jacobson, R.L.; et al. Reduction of maternal-infant transmission of human immunodeficiency virus type 1 with zidovudine treatment. Pediatric AIDS Clinical Trials Group Protocol 076 Study Group. N. Engl. J. Med. 1994, 331, 1173–1180. [Google Scholar] [CrossRef] [PubMed]
- Cooper, E.R.; Charurat, M.; Mofenson, L.; Hanson, I.C.; Pitt, J.; Diaz, C.; Hayani, K.; Handelsman, E.; Smeriglio, V.; Hoff, R.; et al. Combination antiretroviral strategies for the treatment of pregnant HIV-1-infected women and prevention of perinatal HIV-1 transmission. J. Acquir. Immune Defic. Syndr. 2002, 29, 484–494. [Google Scholar] [CrossRef]
- Lallemant, M.; Jourdain, G.; Le Coeur, S.; Mary, J.Y.; Ngo-Giang-Huong, N.; Koetsawang, S.; Kanshana, S.; McIntosh, K.; Thaineua, V. Single-dose perinatal nevirapine plus standard zidovudine to prevent mother-to-child transmission of HIV-1 in Thailand. N. Engl. J. Med. 2004, 351, 217–228. [Google Scholar] [CrossRef] [PubMed]
- Nduati, E.W.; Hassan, A.S.; Knight, M.G.; Muema, D.M.; Jahangir, M.N.; Mwaringa, S.L.; Etyang, T.J.; Rowland-Jones, S.; Urban, B.C.; Berkley, J.A. Outcomes of prevention of mother to child transmission of the human immunodeficiency virus-1 in rural Kenya—A cohort study. BMC Public Health 2015, 15, 1008. [Google Scholar] [CrossRef] [PubMed]
- Eley, B. Metabolic complications of antiretroviral therapy in HIV-infected children. Expert Opin. Drug Metab. Toxicol. 2008, 4, 37–49. [Google Scholar] [CrossRef] [PubMed]
- Eley, B.; Nuttall, J. Antiretroviral therapy for children: Challenges and opportunities. Ann. Trop. Paediatr. 2007, 27, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Belzer, M.E.; Fuchs, D.N.; Luftman, G.S.; Tucker, D.J. Antiretroviral adherence issues among HIV-positive adolescents and young adults. J. Adolesc. Health 1999, 25, 316–319. [Google Scholar] [CrossRef]
- European Paediatric Lipodystrophy Group. Antiretroviral therapy, fat redistribution and hyperlipidaemia in HIV-infected children in Europe. AIDS 2004, 18, 1443–1451. [Google Scholar]
- Schlatter, A.F.; Deathe, A.R.; Vreeman, R.C. The Need for Pediatric Formulations to Treat Children with HIV. AIDS Res. Treat. 2016, 2016, 1654938. [Google Scholar] [CrossRef] [PubMed]
- Workneh, N.; Girma, T.; Woldie, M. Immunologic and clinical outcomes of children on HAART: A retrospective cohort analysis at JImma University Specialized Hospital. Ethiop. J. Health Sci. 2011, 19, 75–82. [Google Scholar] [CrossRef]
- De Baets, A.J.; Ramet, J.; Msellati, P.; Lepage, P. The unique features of pediatric HIV-1 in sub-Saharan Africa. Curr. HIV Res. 2008, 6, 351–362. [Google Scholar] [CrossRef] [PubMed]
- Scott, G.B.; Hutto, C.; Makuch, R.W.; Mastrucci, M.T.; O’Connor, T.; Mitchell, C.D.; Trapido, E.J.; Parks, W.P. Survival in children with perinatally acquired human immunodeficiency virus type 1 infection. N. Engl. J. Med. 1989, 321, 1791–1796. [Google Scholar] [CrossRef] [PubMed]
- Yassin, S.; Gebretekle, G.B. Magnitude and predictors of antiretroviral treatment failure among HIV-infected children in Fiche and Kuyu hospitals, Oromia region, Ethiopia: A retrospective cohort study. Pharmacol. Res. Perspect. 2017, 5, e00296. [Google Scholar] [CrossRef] [PubMed]
- Biadgilign, S.; Deribew, A.; Amberbir, A.; Deribe, K. Adherence to highly active antiretroviral therapy and its correlates among HIV infected pediatric patients in Ethiopia. BMC Pediatr. 2008, 8, 53. [Google Scholar] [CrossRef] [PubMed]
- Kumarasamy, N.; Krishnan, S. Beyond first-line HIV treatment regimens: The current state of antiretroviral regimens, viral load monitoring, and resistance testing in resource-limited settings. Curr. Opin. HIV AIDS 2013, 8, 586–590. [Google Scholar] [CrossRef] [PubMed]
- World Health Organization (WHO). Consolidated Guidelines on the Use of Antiretroviral Drugs for Treating and Preventing HIV Infection, Recommendations for a Public Health Approach; World Health Organization: Geneva, Switzerland, 2013. [Google Scholar]
- Assefa, Y.; Gilks, C.F.; Lynen, L.; Williams, O.; Hill, P.S.; Tolera, T.; Malvia, A.; Van Damme, W. Performance of the Antiretroviral Treatment Program in Ethiopia, 2005–2015: Strengths and weaknesses toward ending AIDS. Int. J. Infect. Dis. 2017, 60, 70–76. [Google Scholar] [CrossRef] [PubMed]
- Babajide, K.; Ayemoba, O.; Terfa, K.; Ake, J.; Crowell, T.A.; Adamu, Y.; Mohammed, T.; Okoye, I.; Odeyemi, S.; Crawford, K.; et al. Virological Suppression and Patterns of Resistance Amongst Patients on Antiretroviral Therapy at 4 Nigerian Military Hospitals. Curr. HIV Res. 2017, 15, 146–151. [Google Scholar] [CrossRef] [PubMed]
- Brooks, K.; Diero, L.; DeLong, A.; Balamane, M.; Reitsma, M.; Kemboi, E.; Orido, M.; Emonyi, W.; Coetzer, M.; Hogan, J.; et al. Treatment failure and drug resistance in HIV-positive patients on tenofovir-based first-line antiretroviral therapy in western Kenya. J. Int. AIDS Soc. 2016, 19, 20798. [Google Scholar] [CrossRef] [PubMed]
- Hassan, A.S.; Nabwera, H.M.; Mwaringa, S.M.; Obonyo, C.A.; Sanders, E.J.; Rinke de Wit, T.F.; Cane, P.A.; Berkley, J.A. HIV-1 virologic failure and acquired drug resistance among first-line antiretroviral experienced adults at a rural HIV clinic in coastal Kenya: A cross-sectional study. AIDS Res. Ther. 2014, 11, 9. [Google Scholar] [CrossRef] [PubMed]
- Mulu, A.; Maier, M.; Liebert, U.G. Upward trends of acquired drug resistances in Ethiopian HIV-1C isolates: A decade longitudinal study. PLoS ONE 2017, 12, e0186619. [Google Scholar] [CrossRef] [PubMed]
- Karade, S.K.; Ghate, M.V.; Chaturbhuj, D.N.; Kadam, D.B.; Shankar, S.; Gaikwad, N.; Gurav, S.; Joshi, R.; Sane, S.S.; Kulkarni, S.S.; et al. Cross-sectional study of virological failure and multinucleoside reverse transcriptase inhibitor resistance at 12 months of antiretroviral therapy in Western India. Medicine 2016, 95, e4886. [Google Scholar] [CrossRef] [PubMed]
- Jiamsakul, A.; Sungkanuparph, S.; Law, M.; Kantor, R.; Praparattanapan, J.; Li, P.C.; Phanuphak, P.; Merati, T.; Ratanasuwan, W.; Lee, C.K.; et al. HIV multi-drug resistance at first-line antiretroviral failure and subsequent virological response in Asia. J. Int. AIDS Soc. 2014, 17, 19053. [Google Scholar] [CrossRef] [PubMed]
- Mendoza, Y.; Castillo Mewa, J.; Martinez, A.A.; Zaldivar, Y.; Sosa, N.; Arteaga, G.; Armien, B.; Bautista, C.T.; Garcia-Morales, C.; Tapia-Trejo, D.; et al. HIV-1 Antiretroviral Drug Resistance Mutations in Treatment Naive and Experienced Panamanian Subjects: Impact on National Use of EFV-Based Schemes. PLoS ONE 2016, 11, e0154317. [Google Scholar] [CrossRef] [PubMed]
- Gupta, R.K.; Hill, A.; Sawyer, A.W.; Cozzi-Lepri, A.; von Wyl, V.; Yerly, S.; Lima, V.D.; Gunthard, H.F.; Gilks, C.; Pillay, D. Virological monitoring and resistance to first-line highly active antiretroviral therapy in adults infected with HIV-1 treated under WHO guidelines: A systematic review and meta-analysis. Lancet Infect. Dis. 2009, 9, 409–417. [Google Scholar] [CrossRef]
- Germanaud, D.; Derache, A.; Traore, M.; Madec, Y.; Toure, S.; Dicko, F.; Coulibaly, H.; Traore, M.; Sylla, M.; Calvez, V.; et al. Level of viral load and antiretroviral resistance after 6 months of non-nucleoside reverse transcriptase inhibitor first-line treatment in HIV-1-infected children in Mali. J. Antimicrob. Chemother. 2010, 65, 118–124. [Google Scholar] [CrossRef] [PubMed]
- Amani-Bosse, C.; Dahourou, D.L.; Malateste, K.; Amorissani-Folquet, M.; Coulibaly, M.; Dattez, S.; Emieme, A.; Barry, M.; Rouzioux, C.; N’Gbeche, S.; et al. Virological response and resistances over 12 months among HIV-infected children less than two years receiving first-line lopinavir/ritonavir-based antiretroviral therapy in Cote d’Ivoire and Burkina Faso: The MONOD ANRS 12206 cohort. J. Int. AIDS Soc. 2017, 20, 21362. [Google Scholar] [CrossRef] [PubMed]
- Fokam, J.; Salpini, R.; Santoro, M.M.; Cento, V.; Perno, C.F.; Colizzi, V.; Ndumbe, P.M.; Fokunang Ntungen, C.; Ndiang Tetang, S.M.; Nanfack, A.J.; et al. Drug resistance among drug-naive and first-line antiretroviral treatment-failing children in Cameroon. Pediatr. Infect. Dis. J. 2011, 30, 1062–1068. [Google Scholar] [CrossRef] [PubMed]
- Boender, T.S.; Kityo, C.M.; Boerma, R.S.; Hamers, R.L.; Ondoa, P.; Wellington, M.; Siwale, M.; Nankya, I.; Kaudha, E.; Akanmu, A.S.; et al. Accumulation of HIV-1 drug resistance after continued virological failure on first-line ART in adults and children in sub-Saharan Africa. J. Antimicrob. Chemother. 2016, 71, 2918–2927. [Google Scholar] [CrossRef] [PubMed]
- Mulu, A.; Liebert, U.G.; Maier, M. Virological efficacy and immunological recovery among Ethiopian HIV-1 infected adults and children. BMC Infect. Dis. 2014, 14, 28. [Google Scholar] [CrossRef] [PubMed]
- Tadesse, B.T.; Foster, B.A.; Jerene, D.; Ruff, A. Cohort profile: Improving treatment of HIV-infected Ethiopian children through better detection of treatment failure in southern Ethiopia. BMJ Open 2017, 7, e013528. [Google Scholar] [CrossRef] [PubMed]
- Buckton, A.J.; Prabhu, D.P.; Cane, P.A.; Pillay, D. No evidence for cross-contamination of dried blood spots excised using an office hole-punch for HIV-1 drug resistance genotyping. J. Antimicrob. Chemother. 2009, 63, 615–616. [Google Scholar] [CrossRef] [PubMed]
- Woods, C.K.; Brumme, C.J.; Liu, T.F.; Chui, C.K.; Chu, A.L.; Wynhoven, B.; Hall, T.A.; Trevino, C.; Shafer, R.W.; Harrigan, P.R. Automating HIV drug resistance genotyping with RECall, a freely accessible sequence analysis tool. J. Clin. Microbiol. 2012, 50, 1936–1942. [Google Scholar] [CrossRef] [PubMed]
- Gaschen, B.; Kuiken, C.; Korber, B.; Foley, B. Retrieval and on-the-fly alignment of sequence fragments from the HIV database. Bioinformatics 2001, 17, 415–418. [Google Scholar] [CrossRef] [PubMed]
- Larsson, A. AliView: A fast and lightweight alignment viewer and editor for large datasets. Bioinformatics 2014, 30, 3276–3278. [Google Scholar] [CrossRef] [PubMed]
- Guindon, S.; Dufayard, J.F.; Lefort, V.; Anisimova, M.; Hordijk, W.; Gascuel, O. New algorithms and methods to estimate maximum-likelihood phylogenies: Assessing the performance of PhyML 3.0. Syst. Biol. 2010, 59, 307–321. [Google Scholar] [CrossRef] [PubMed]
- Shafer, R.W.; Rhee, S.Y.; Pillay, D.; Miller, V.; Sandstrom, P.; Schapiro, J.M.; Kuritzkes, D.R.; Bennett, D. HIV-1 protease and reverse transcriptase mutations for drug resistance surveillance. AIDS 2007, 21, 215–223. [Google Scholar] [CrossRef] [PubMed]
- Fourment, M.; Gibbs, M.J. PATRISTIC: A program for calculating patristic distances and graphically comparing the components of genetic change. BMC Evol. Biol. 2006, 6, 1. [Google Scholar] [CrossRef] [PubMed]
- Siepel, A.C.; Halpern, A.L.; Macken, C.; Korber, B.T. A computer program designed to screen rapidly for HIV type 1 intersubtype recombinant sequences. AIDS Res. Hum. Retrovir. 1995, 11, 1413–1416. [Google Scholar] [CrossRef] [PubMed]
- Liu, T.F.; Shafer, R.W. Web resources for HIV type 1 genotypic-resistance test interpretation. Clin. Infect. Dis. 2006, 42, 1608–1618. [Google Scholar] [CrossRef] [PubMed]
- Bennett, D.E.; Camacho, R.J.; Otelea, D.; Kuritzkes, D.R.; Fleury, H.; Kiuchi, M.; Heneine, W.; Kantor, R.; Jordan, M.R.; Schapiro, J.M.; et al. Drug resistance mutations for surveillance of transmitted HIV-1 drug-resistance: 2009 update. PLoS ONE 2009, 4, e4724. [Google Scholar] [CrossRef] [PubMed]
- Harrigan, P.R.; Wynhoven, B.; Brumme, Z.L.; Brumme, C.J.; Sattha, B.; Major, J.C.; de la Rosa, R.; Montaner, J.S. HIV-1 drug resistance: Degree of underestimation by a cross-sectional versus a longitudinal testing approach. J. Infect. Dis. 2005, 191, 1325–1330. [Google Scholar] [CrossRef] [PubMed]
- Steegen, K.; Luchters, S.; Demecheleer, E.; Dauwe, K.; Mandaliya, K.; Jaoko, W.; Plum, J.; Temmerman, M.; Verhofstede, C. Feasibility of detecting human immunodeficiency virus type 1 drug resistance in DNA extracted from whole blood or dried blood spots. J. Clin. Microbiol. 2007, 45, 3342–3351. [Google Scholar] [CrossRef] [PubMed]
- Le, T.; Chiarella, J.; Simen, B.B.; Hanczaruk, B.; Egholm, M.; Landry, M.L.; Dieckhaus, K.; Rosen, M.I.; Kozal, M.J. Low-abundance HIV drug-resistant viral variants in treatment-experienced persons correlate with historical antiretroviral use. PLoS ONE 2009, 4, e6079. [Google Scholar] [CrossRef] [PubMed]
- Garcia, F.; Alvarez, M.; Fox, Z.; Garcia-Diaz, A.; Guillot, V.; Johnson, M.; Chueca, N.; Phillips, A.; Hernandez-Quero, J.; Geretti, A.M. Predicting antiretroviral drug resistance from the latest or the cumulative genotype. Antivir. Ther. 2011, 16, 373–382. [Google Scholar] [CrossRef] [PubMed]
- Reuter, S.; Oette, M.; Sichtig, N.; Kaiser, R.; Balduin, M.; Jensen, B.; Haussinger, D. Changes in the HIV-1 mutational profile before first-line HAART in the RESINA cohort. J. Med. Virol. 2011, 83, 187–195. [Google Scholar] [CrossRef] [PubMed]
- Badri, S.M.; Adeyemi, O.M.; Max, B.E.; Hota, B.N.; Barker, D.E. Utility of repeat genotypic resistance testing and clinical response in patients with three class resistance and virologic treatment failure. AIDS Patient Care STDs 2007, 21, 544–550. [Google Scholar] [CrossRef] [PubMed]
- Contreras, G.A.; Delbianco, G.; Bell, C.S.; Kleinosky, M.T.; Murphy, J.R.; Heresi, G.P. Single genotypes underestimate the prevalence of antiretroviral resistance in patients with perinatally acquired HIV. J. Infect. 2012, 64, 125–126. [Google Scholar] [CrossRef] [PubMed]
- Punyacam, P.; Iemwimangsa, N.; Chantratita, W.; Sukasem, C.; Sungkanuparph, S. HIV drug resistance interpreted by cumulative versus last genotypes in HIV-infected patients with multiple treatment failures. Curr. HIV Res. 2012, 10, 271–274. [Google Scholar] [CrossRef] [PubMed]
- World Health Organization (WHO). WHO Global Database on Child Growth and Malnutrition; World Health Organization: Geneva, Switzerland, 1997. [Google Scholar]
- World Health Organization (WHO). WHO Case Definitions of HIV for Surveillance and Revised Clinical Staging and Immunological Classification of HIV-Related Disease in Adults and Children; World Health Organization: Geneva, Switzerland, 2007. [Google Scholar]
- Ministry of Health Federal Democratic Republic of Ethiopia. National Guidelines for Comprehensive HIV Prevention, Care and Treatment; Ethiopian Ministry of Health: Addis Ababa, Ethiopia, 2014.
- Rottinghaus, E.K.; Ugbena, R.; Diallo, K.; Bassey, O.; Azeez, A.; Devos, J.; Zhang, G.; Aberle-Grasse, J.; Nkengasong, J.; Yang, C. Dried blood spot specimens are a suitable alternative sample type for HIV-1 viral load measurement and drug resistance genotyping in patients receiving first-line antiretroviral therapy. Clin. Infect. Dis. 2012, 54, 1187–1195. [Google Scholar] [CrossRef] [PubMed]
- Youngpairoj, A.S.; Masciotra, S.; Garrido, C.; Zahonero, N.; de Mendoza, C.; Garcia-Lerma, J.G. HIV-1 drug resistance genotyping from dried blood spots stored for 1 year at 4 degrees C. J. Antimicrob. Chemother. 2008, 61, 1217–1220. [Google Scholar] [CrossRef] [PubMed]
- Ziemniak, C.; George-Agwu, A.; Moss, W.J.; Ray, S.C.; Persaud, D. A sensitive genotyping assay for detection of drug resistance mutations in reverse transcriptase of HIV-1 subtypes B and C in samples stored as dried blood spots or frozen RNA extracts. J. Virol. Methods 2006, 136, 238–247. [Google Scholar] [CrossRef] [PubMed]
- McNulty, A.; Jennings, C.; Bennett, D.; Fitzgibbon, J.; Bremer, J.W.; Ussery, M.; Kalish, M.L.; Heneine, W.; Garcia-Lerma, J.G. Evaluation of dried blood spots for human immunodeficiency virus type 1 drug resistance testing. J. Clin. Microbiol. 2007, 45, 517–521. [Google Scholar] [CrossRef] [PubMed]
- Yang, C.; McNulty, A.; Diallo, K.; Zhang, J.; Titanji, B.; Kassim, S.; Wadonda-Kabondo, N.; Aberle-Grasse, J.; Kibuka, T.; Ndumbe, P.M.; et al. Development and application of a broadly sensitive dried-blood-spot-based genotyping assay for global surveillance of HIV-1 drug resistance. J. Clin. Microbiol. 2010, 48, 3158–3164. [Google Scholar] [CrossRef] [PubMed]
- Masciotra, S.; Garrido, C.; Youngpairoj, A.S.; McNulty, A.; Zahonero, N.; Corral, A.; Heneine, W.; de Mendoza, C.; Garcia-Lerma, J.G. High concordance between HIV-1 drug resistance genotypes generated from plasma and dried blood spots in antiretroviral-experienced patients. AIDS 2007, 21, 2503–2511. [Google Scholar] [CrossRef] [PubMed]
- Hallack, R.; Doherty, L.E.; Wethers, J.A.; Parker, M.M. Evaluation of dried blood spot specimens for HIV-1 drug-resistance testing using the Trugene HIV-1 genotyping assay. J. Clin. Virol. 2008, 41, 283–287. [Google Scholar] [CrossRef] [PubMed]
- Lira, R.; Valdez-Salazar, H.; Vazquez-Rosales, G.; Rojas-Montes, O.; Ruiz-Tachiquin, M.; Torres-Ibarra, R.; Cano-Dominguez, C.; Maldonado-Rodriguez, A.; Gomez, A.; Munoz, O.; et al. Genotypic testing for HIV-1 drug resistance using dried blood samples. Arch. Virol. 2010, 155, 1117–1125. [Google Scholar] [CrossRef] [PubMed]
- Ndung’u, T.; Renjifo, B.; Novitsky, V.A.; McLane, M.F.; Gaolekwe, S.; Essex, M. Molecular cloning and biological characterization of full-length HIV-1 subtype C from Botswana. Virology 2000, 278, 390–399. [Google Scholar] [CrossRef] [PubMed]
- Hemelaar, J.; Gouws, E.; Ghys, P.D.; Osmanov, S. Global trends in molecular epidemiology of HIV-1 during 2000–2007. AIDS 2011, 25, 679–689. [Google Scholar] [CrossRef] [PubMed]
- Kassu, A.; Fujino, M.; Matsuda, M.; Nishizawa, M.; Ota, F.; Sugiura, W. Molecular epidemiology of HIV type 1 in treatment-naive patients in north Ethiopia. AIDS Res. Hum. Retrovir. 2007, 23, 564–568. [Google Scholar] [CrossRef] [PubMed]
- Tully, D.C.; Wood, C. Chronology and evolution of the HIV-1 subtype C epidemic in Ethiopia. AIDS 2010, 24, 1577–1582. [Google Scholar] [CrossRef] [PubMed]
- Kalu, A.W.; Telele, N.F.; Gebreselasie, S.; Fekade, D.; Abdurahman, S.; Marrone, G.; Sonnerborg, A. Monophylogenetic HIV-1C epidemic in Ethiopia is dominated by CCR5-tropic viruses-an analysis of a prospective country-wide cohort. BMC Infect. Dis. 2017, 17, 37. [Google Scholar] [CrossRef] [PubMed]
- Delatorre, E.O.; Bello, G. Phylodynamics of HIV-1 subtype C epidemic in east Africa. PLoS ONE 2012, 7, e41904. [Google Scholar] [CrossRef] [PubMed]
- Mulu, A.; Lange, T.; Liebert, U.G.; Maier, M. Clade homogeneity and Pol gene polymorphisms in chronically HIV-1 infected antiretroviral treatment naive patients after the roll out of ART in Ethiopia. BMC Infect. Dis. 2014, 14, 158. [Google Scholar] [CrossRef] [PubMed]
- Rhee, S.Y.; Jordan, M.R.; Raizes, E.; Chua, A.; Parkin, N.; Kantor, R.; Van Zyl, G.U.; Mukui, I.; Hosseinipour, M.C.; Frenkel, L.M.; et al. HIV-1 Drug Resistance Mutations: Potential Applications for Point-of-Care Genotypic Resistance Testing. PLoS ONE 2015, 10, e0145772. [Google Scholar] [CrossRef] [PubMed]
- Wensing, A.M.; Calvez, V.; Gunthard, H.F.; Johnson, V.A.; Paredes, R.; Pillay, D.; Shafer, R.W.; Richman, D.D. 2017 Update of the Drug Resistance Mutations in HIV-1. Top. Antivir. Med. 2017, 24, 132–133. [Google Scholar] [PubMed]
- Schuurman, R.; Nijhuis, M.; van Leeuwen, R.; Schipper, P.; de Jong, D.; Collis, P.; Danner, S.A.; Mulder, J.; Loveday, C.; Christopherson, C.; et al. Rapid changes in human immunodeficiency virus type 1 RNA load and appearance of drug-resistant virus populations in persons treated with lamivudine (3TC). J. Infect. Dis. 1995, 171, 1411–1419. [Google Scholar] [CrossRef] [PubMed]
- Wainberg, M.A.; Salomon, H.; Gu, Z.; Montaner, J.S.; Cooley, T.P.; McCaffrey, R.; Ruedy, J.; Hirst, H.M.; Cammack, N.; Cameron, J.; et al. Development of HIV-1 resistance to (−)2′-deoxy-3′-thiacytidine in patients with AIDS or advanced AIDS-related complex. AIDS 1995, 9, 351–357. [Google Scholar] [CrossRef] [PubMed]
- Wainberg, M.A.; Lewis, L.; Salomon, H.; Gu, Z.; Keller, A.; Cammack, N.; Goldsmith, J.; Church, J.; Spira, B.; Wheeler, S.; et al. Resistance to (−)-2′,3′-dideoxy-3′-thiacytidine (3TC) in HIV-1 isolated from paediatric patients. Antivir. Ther. 1996, 1, 98–104. [Google Scholar] [PubMed]
- Petrella, M.; Oliveira, M.; Moisi, D.; Detorio, M.; Brenner, B.G.; Wainberg, M.A. Differential maintenance of the M184V substitution in the reverse transcriptase of human immunodeficiency virus type 1 by various nucleoside antiretroviral agents in tissue culture. Antimicrob. Agents Chemother. 2004, 48, 4189–4194. [Google Scholar] [CrossRef] [PubMed]
- Shafer, R.W.; Schapiro, J.M. HIV-1 Drug Resistance Mutations: An Updated Framework for the Second Decade of HAART. AIDS Rev. 2008, 10, 67–84. [Google Scholar] [PubMed]
- Wainberg, M.A. The impact of the M184V substitution on drug resistance and viral fitness. Expert Rev. Anti-Infect. Ther. 2004, 2, 147–151. [Google Scholar] [CrossRef] [PubMed]
- Meintjes, G.; Dunn, L.; Coetsee, M.; Hislop, M.; Leisegang, R.; Regensberg, L.; Maartens, G. Third-line antiretroviral therapy in Africa: Effectiveness in a Southern African retrospective cohort study. AIDS Res. Ther. 2015, 12, 39. [Google Scholar] [CrossRef] [PubMed]
- Tang, M.W.; Kanki, P.J.; Shafer, R.W. A review of the virological efficacy of the 4 World Health Organization-recommended tenofovir-containing regimens for initial HIV therapy. Clin. Infect. Dis. 2012, 54, 862–875. [Google Scholar] [CrossRef] [PubMed]
- Keiser, P.; Nassar, N.; White, C.; Koen, G.; Moreno, S. Comparison of nevirapine- and efavirenz-containing antiretroviral regimens in antiretroviral-naive patients: A cohort study. HIV Clin. Trials 2002, 3, 296–303. [Google Scholar] [PubMed]
- Amico, K.R.; Fisher, W.A.; Cornman, D.H.; Shuper, P.A.; Redding, C.G.; Konle-Parker, D.J.; Barta, W.; Fisher, J.D. Visual analog scale of ART adherence: Association with 3-day self report and adherence barriers. J. Acquir. Immune Defic. Syndr. 2006, 42, 455–459. [Google Scholar] [CrossRef] [PubMed]
- Paquet, A.C.; Baxter, J.; Weidler, J.; Lie, Y.; Lawrence, J.; Kim, R.; Bates, M.; Coakley, E.; Chappey, C. Differences in reversion of resistance mutations to wild-type under structured treatment interruption and related increase in replication capacity. PLoS ONE 2011, 6, e14638. [Google Scholar] [CrossRef] [PubMed]
- Boucher, S.; Recordon-Pinson, P.; Neau, D.; Ragnaud, J.M.; Titier, K.; Faure, M.; Fleury, H.; Masquelier, B. Clonal analysis of HIV-1 variants in proviral DNA during treatment interruption in patients with multiple therapy failures. J. Clin. Virol. 2005, 34, 288–294. [Google Scholar] [CrossRef] [PubMed]
- Kaye, S.; Comber, E.; Tenant-Flowers, M.; Loveday, C. The appearance of drug resistance-associated point mutations in HIV type 1 plasma RNA precedes their appearance in proviral DNA. AIDS Res. Hum. Retrovir. 1995, 11, 1221–1225. [Google Scholar] [CrossRef] [PubMed]
- Republic du Mali Ministere de la Sante et de l’Hygiene Publique. Politique et Protocoles de Prise en Charge Antiretrovirale du VIH et du SIDA; Ministry of Public Health and Hygiene, Republic of Mali: Bamako, Mali, 2013.
- Republic of Cameroon Ministry of Public Health. National Guidelines on the Prevention and Management of HIV in Cameroon; Ministry of Health, Republic of Cameroon: Yaounde, Cameroon, 2015.
- World Health Organization (WHO). WHO Manual for HIV Drug Resistance Testing using Dried Blood Spot Specimens; World Health Organization: Geneva, Switzerland, 2012. [Google Scholar]
| Variable | Summary Statistic | N | |||
|---|---|---|---|---|---|
| Age at baseline, years, Median (IQR) | 12 (9–14) | 94 | |||
| Gender, % male | 60% | 94 | |||
| pVL at baseline, log10 copies/mL, Median (IQR) | 3.9 (3.6–4.6) | 94 | |||
| pVL at failure, log10 copies/mL, Median (IQR) | 4.2 (3.8–4.7) | 94 | |||
| CD4+ T-cell count at baseline, cells/mm3, Median (IQR) | 500 (247–781) | 92 | |||
| CD4+ T-cell count at ART initiation, cells/mm3, Median (IQR) | 276 (163–613) | 89 | |||
| ART duration at baseline, months, Median (IQR) | 35 (18–70.5) | 89 | |||
| Weight for age at baseline, Z-score *, Median (IQR) | −1.5 (−2.1–(−0.6)) | 89 | |||
| Height for age at baseline, Z-score *, Median (IQR) | −1.3 (2.1–(−0.4)) | 89 | |||
| combination Antiretrovial T regimen at baseline, % patients | |||||
| NRTI | 93 | ||||
| 3TC+ | AZT | 66% | |||
| d4T | 29% | ||||
| TDF | 3% | ||||
| ABC | 2% | ||||
| NNRTI | 93 | ||||
| EFV | 22% | ||||
| NVP | 77% | ||||
| WHO Clinical Stage at baseline, % patients Stage 1 | 85% | 94 | |||
| Variable 1 | NRTI Resistance | NNRTI Resistance | Any Resistance | |||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Yes (N = 64) | No (N = 26) | p | Yes (N = 72) | No (N = 18) | p | Yes (N = 73) | No (N = 17) | p | ||
| Age at baseline, years, Median (IQR), [N = 90] | 11 (8.0–14.0) | 12.8 (11.0–14.0) | 0.07 | 12.0 (9.0–14.0) | 12.0 (9.0–13.0) | 0.73 | 12.0 (9.0–14.0) | 12.0 (10.0–13.0) | 0.51 | |
| Sex, % Male, [N = 90] | 66% | 54% | 0.34 | 60% | 72% | 0.42 | 60% | 71% | 0.58 | |
| ART duration at baseline, months Median [IQR], [N = 86] | 49.5 (23.0–71.8) | 25.5 (7.3–61.5) | 0.11 | 48.5 (23.0–72.0) | 20.5 (4.8–51.8) | 0.03 | 48.0 (22.5–72.0) | 24.0 (6.9–54.5) | 0.08 | |
| WHO Clinical Stage at baseline, % Stage 1, [N = 90] | 86% | 81% | 0.54 | 86% | 79% | 0.48 | 85% | 82% | 0.72 | |
| CD4 count at baseline, cells/mm3 Median [IQR], [N = 88] | 481 (224–752) | 483 (270–781) | 0.79 | 481 (248–674) | 584 (270–910) | 0.50 | 482 (250–700) | 485 (257–957) | 0.66 | |
| CD4 count at ART initiation cells/mm3, Median[IQR], [N = 84] | 280 (136–646) | 265 (195–482) | 0.71 | 245 (149–643) | 292 (200–731) | 0.44 | 261 (151–628) | 270 (198–581) | 0.67 | |
| Baseline Weight-for-age Z-score * Median[IQR], [N = 89] | −1.5 (−2.1–(−0.6)) | −1.7 (−2.2–(−0.9)) | 0.59 | −1.5 (−2.1–(−0.7)) | −1.8 (−2.4–(−1.2)) | 0.2 | −1.5 (−2.1–(−0.6)) | −1.8 (−2.6–(−1.1)) | 0.31 | |
| Baseline Height-for-age Z-score * Median[IQR], [N = 89] | −1.3 (−2.0–(−0.3)) | −1.6 (−2.3–(−0.6)) | 0.32 | −1.3 (−2.1–(−0.4)) | −1.5 (−2.1–(−0.6)) | 0.63 | −1.3 (−2.1–(−0.4)) | −1.4 (−2.1–(−0.6)) | 0.77 | |
| Baseline ART regimen [N = 89] | % AZT-based ϕ | 66% | 64% | 1.00 | 68% | 56% | 0.41 | 67% | 59% | 0.58 |
| % D4T-based ϕ | 33% | 20% | 0.31 | 30% | 28% | 1.00 | 29% | 29% | 1.00 | |
| % NVP-based | 81% | 71% | 0.40 | 83% | 60% | 0.08 | 83% | 60% | 0.08 | |
| % EFV-based | 19% | 29% | 0.40 | 17% | 40% | 0.08 | 17% | 40% | 0.08 | |
| Drug substitution, %Yes, [N = 84] | 57% | 30% | 0.04 | 55% | 27% | 0.08 | 54% | 29% | 0.14 | |
| Adherence to ART, % sub-optimal #, [N = 90] | 34% | 27% | 0.62 | 64% | 17% | 0.16 | 36% | 18% | 0.25 | |
| Treatment for Tuberculosis, %Yes [N = 90] | 33% | 31% | 1.00 | 31% | 39% | 0.58 | 30% | 41% | 0.40 | |
| Viral load at baseline, Log10 copies/mL, Median[IQR], [N = 90] | 3.8 (3.5–4.4) | 3.9 (3.7–4.7) | 0.47 | 3.8 (3.5–4.3) | 4.1 (3.8–5.1) | 0.16 | 3.8 (3.6–4.4) | 4.0 (3.8–5.1) | 0.29 | |
| Viral load at failure, Log10 copies/mL, Median[IQR], [N = 90] | 4.2 (3.8–4.7) | 4.3 (3.9–4.8) | 0.42 | 4.2 (3.8–4.7) | 4.4 (3.9–5.1) | 0.20 | 4.2 (3.8–4.8) | 4.8 (3.9–5.0) | 0.33 | |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tadesse, B.T.; Kinloch, N.N.; Baraki, B.; Lapointe, H.R.; Cobarrubias, K.D.; Brockman, M.A.; Brumme, C.J.; Foster, B.A.; Jerene, D.; Makonnen, E.; et al. High Levels of Dual-Class Drug Resistance in HIV-Infected Children Failing First-Line Antiretroviral Therapy in Southern Ethiopia. Viruses 2018, 10, 60. https://doi.org/10.3390/v10020060
Tadesse BT, Kinloch NN, Baraki B, Lapointe HR, Cobarrubias KD, Brockman MA, Brumme CJ, Foster BA, Jerene D, Makonnen E, et al. High Levels of Dual-Class Drug Resistance in HIV-Infected Children Failing First-Line Antiretroviral Therapy in Southern Ethiopia. Viruses. 2018; 10(2):60. https://doi.org/10.3390/v10020060
Chicago/Turabian StyleTadesse, Birkneh Tilahun, Natalie N. Kinloch, Bemuluyigza Baraki, Hope R. Lapointe, Kyle D. Cobarrubias, Mark A. Brockman, Chanson J. Brumme, Byron A. Foster, Degu Jerene, Eyasu Makonnen, and et al. 2018. "High Levels of Dual-Class Drug Resistance in HIV-Infected Children Failing First-Line Antiretroviral Therapy in Southern Ethiopia" Viruses 10, no. 2: 60. https://doi.org/10.3390/v10020060
APA StyleTadesse, B. T., Kinloch, N. N., Baraki, B., Lapointe, H. R., Cobarrubias, K. D., Brockman, M. A., Brumme, C. J., Foster, B. A., Jerene, D., Makonnen, E., Aklillu, E., & Brumme, Z. L. (2018). High Levels of Dual-Class Drug Resistance in HIV-Infected Children Failing First-Line Antiretroviral Therapy in Southern Ethiopia. Viruses, 10(2), 60. https://doi.org/10.3390/v10020060





