A Mechanism Underlying Attenuation of Recombinant Influenza A Viruses Carrying Reporter Genes
Abstract
1. Introduction
2. Materials and Methods
2.1. Cell Lines and Viruses
2.2. Single-Cycle and Multiple-Cycle Replication Assay
2.3. qPCR Analysis of Packaged vRNAs
2.4. Cell Based RdRp Assay
2.5. Analysis of vRNA Replication in Infected Cells
3. Results
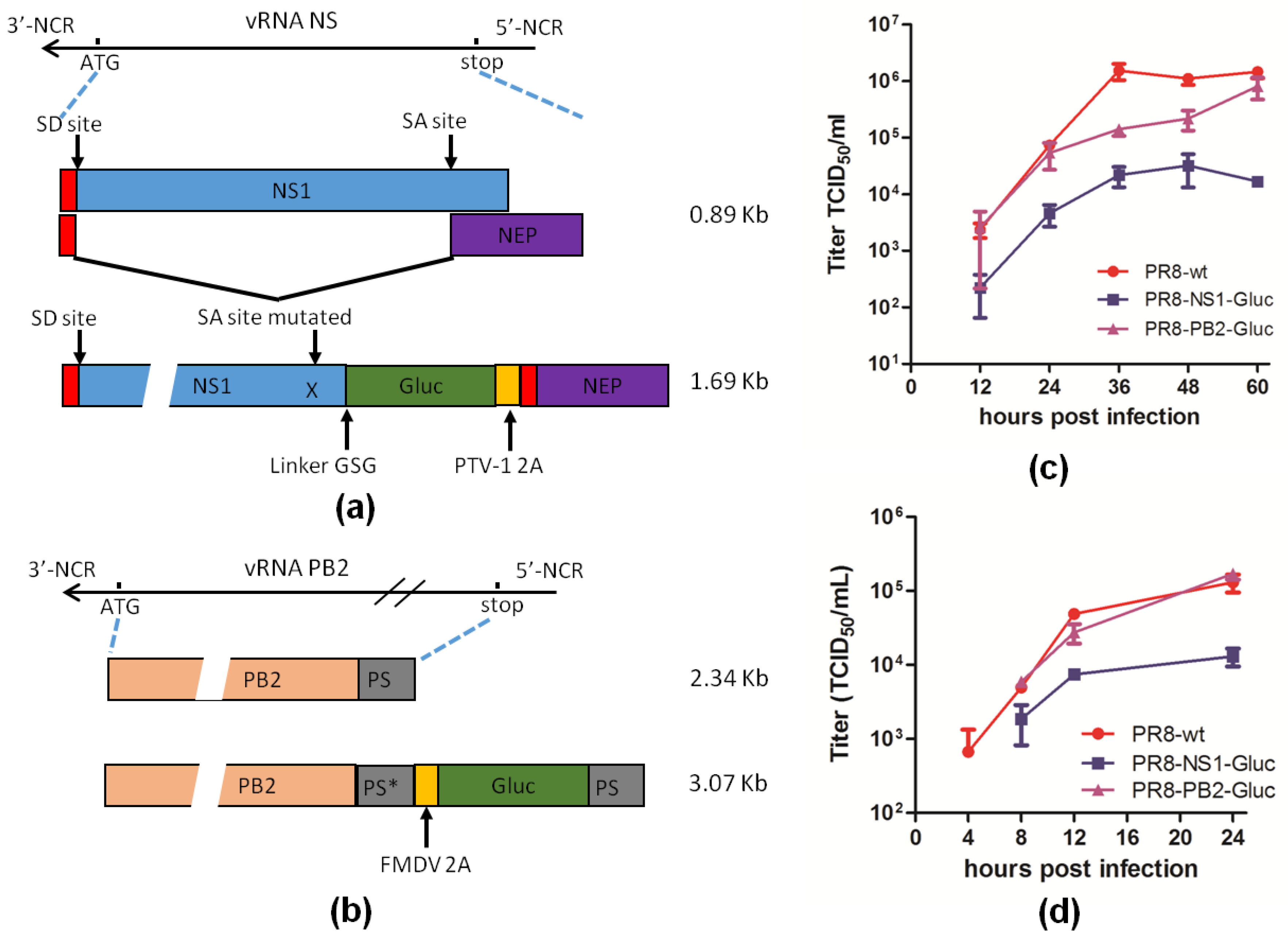
3.1. Comparison of the Replication Kinetics of Two Reporter IAVs PR8-NS1-Gluc and PR8-PB2-Gluc
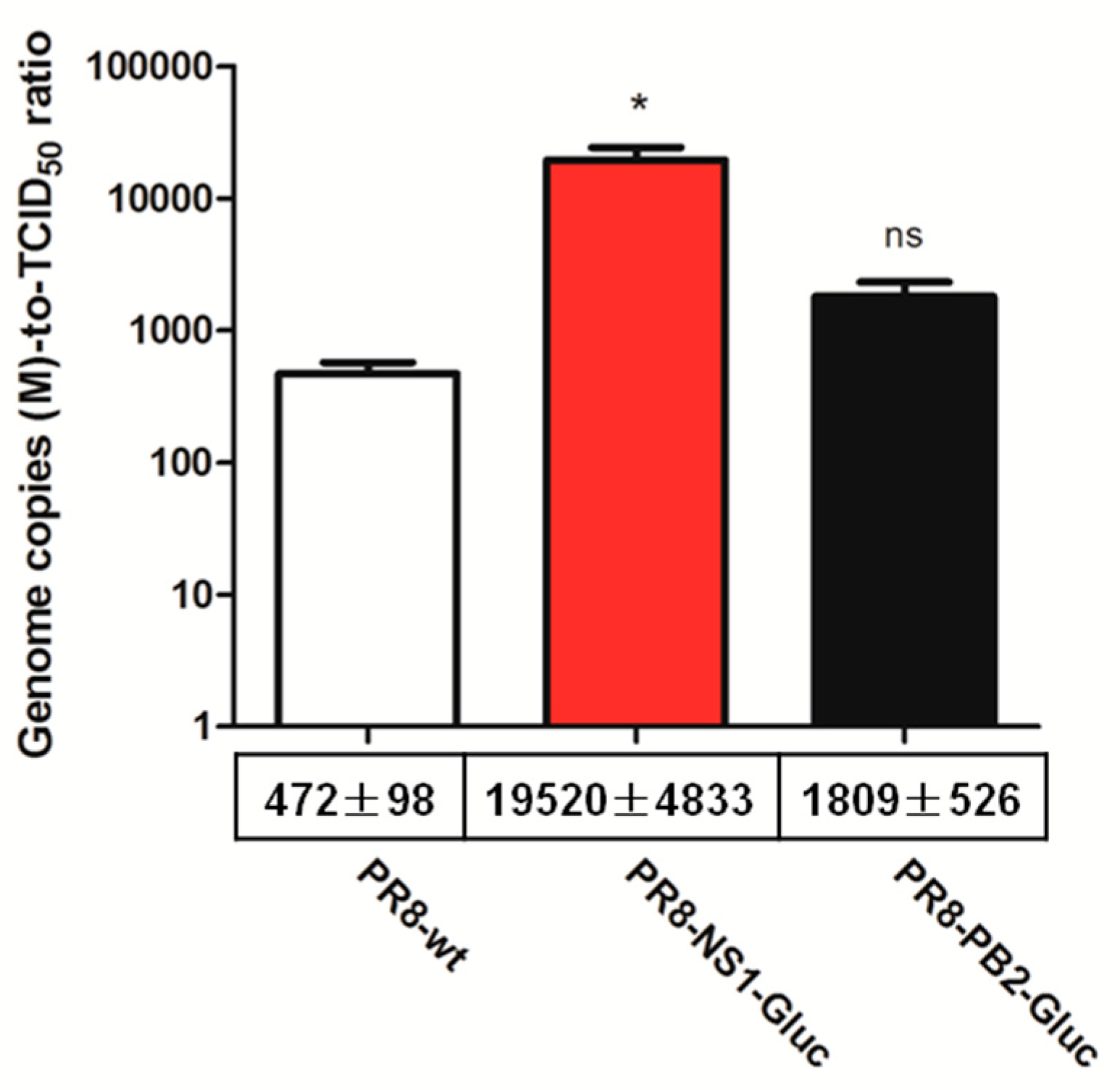
3.2. The Genome Copies-to- TCID50 Ratio of PR8-NS1-Gluc is Greatly Increased
3.3. PR8-NS1-Gluc Produces Large Amounts of NS-Null Noninfectious Particles
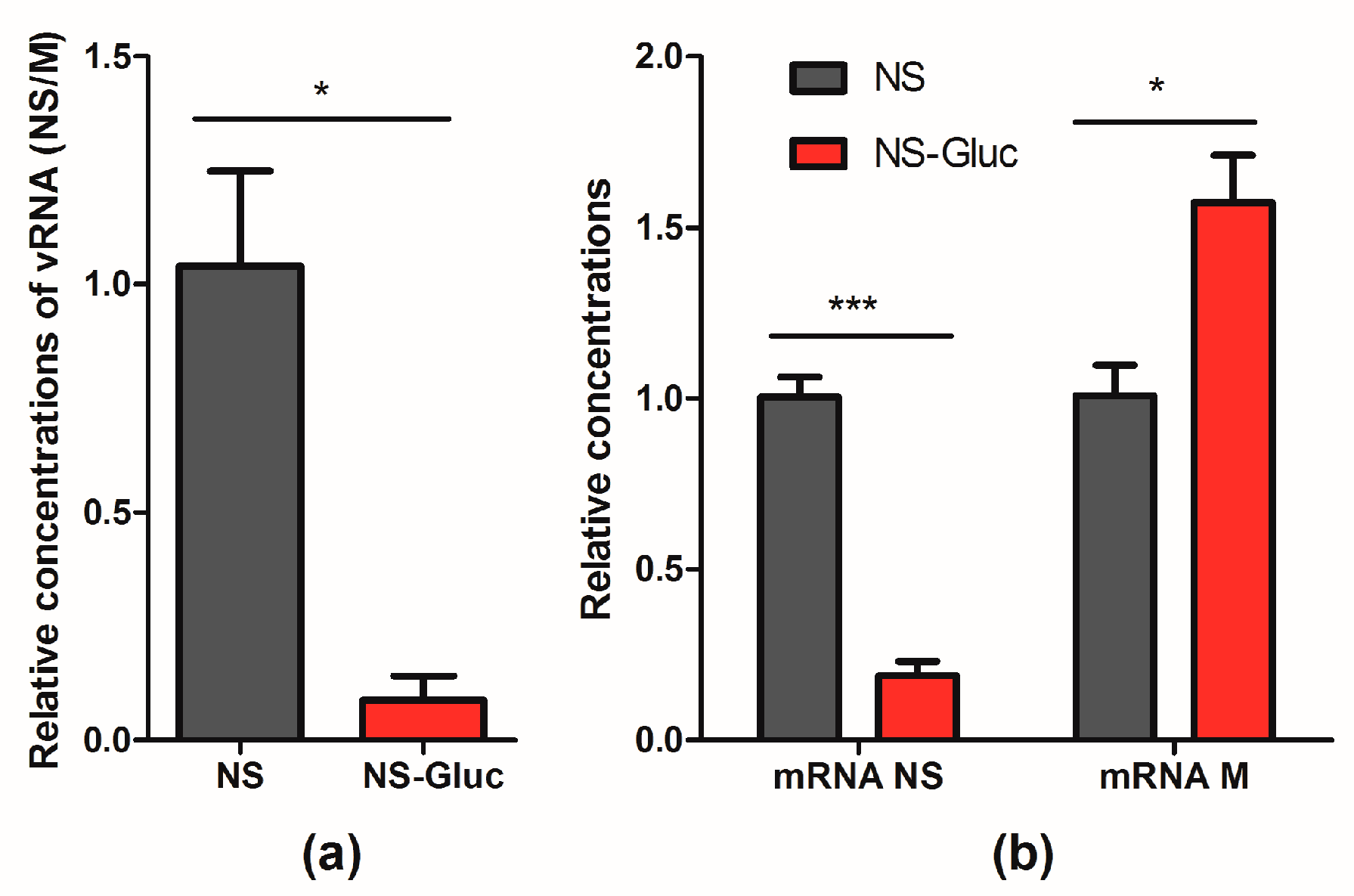
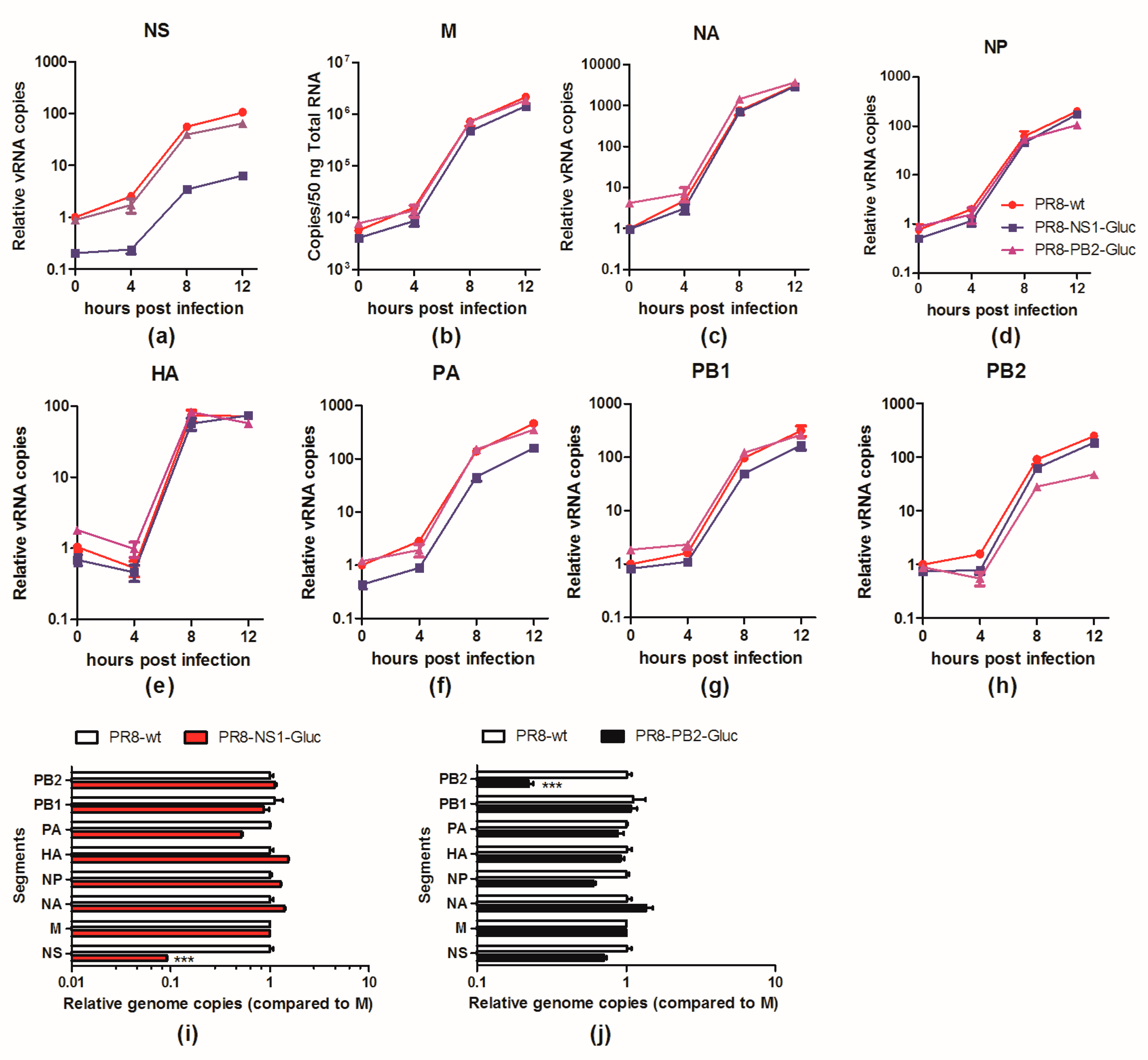
3.4. The vRNA NS-Gluc is Less Competitive in Replication
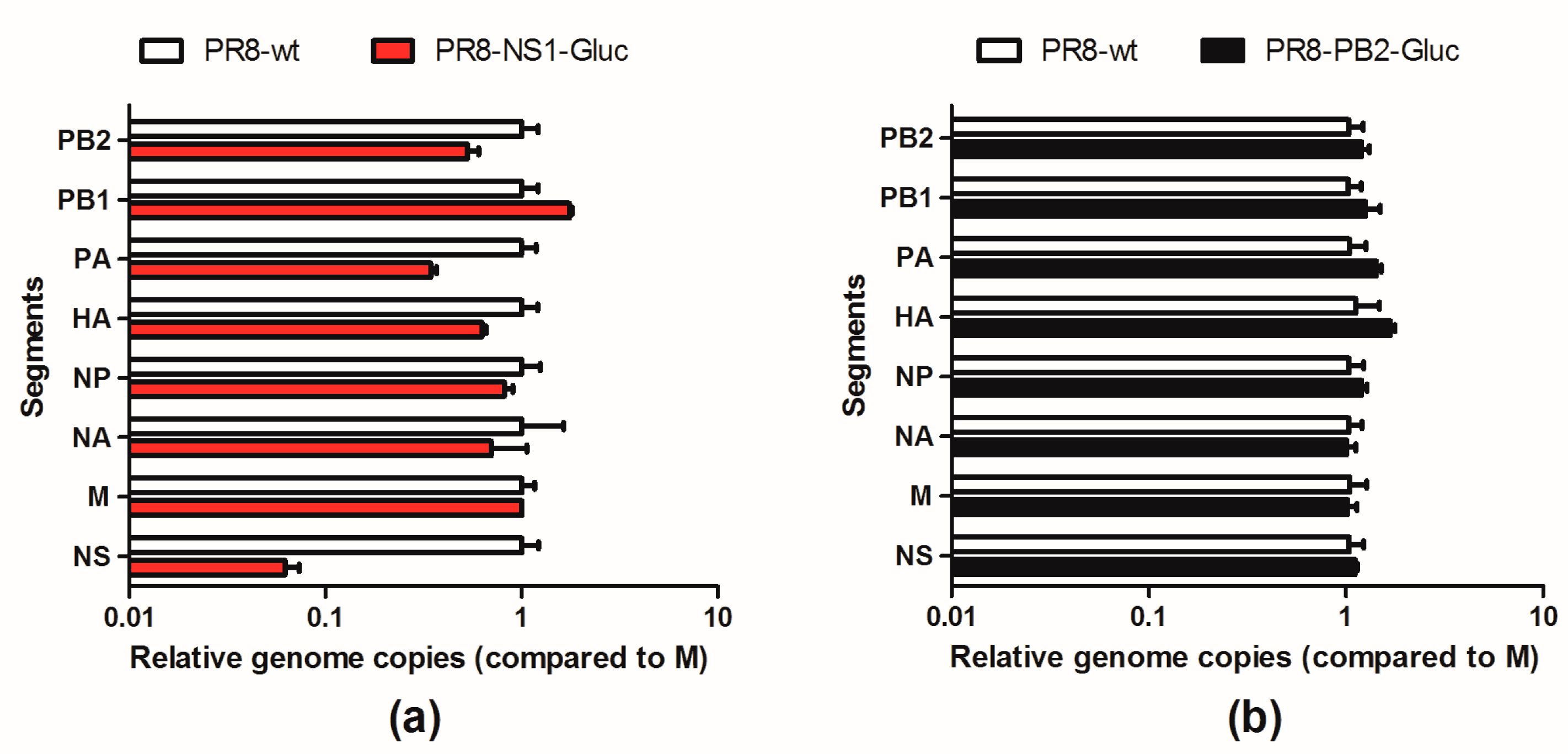
3.5. The Balance of the 8 Segments of Reporter IAVs Is Impaired
4. Discussion
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- WHO. 2018 Influenza (Seasonal) Fact Sheet. WHO: Geneva, Switzerland, 2018; Available online: http://www.who.int/mediacentre/factsheets/fs211/en/ (accessed on 15 October 2018).
- Resa-Infante, P.; Jorba, N.; Coloma, R.; Ortin, J. The influenza virus RNA synthesis machine: Advances in its structure and function. RNA Biol. 2011, 8, 207–215. [Google Scholar] [CrossRef] [PubMed]
- Eisfeld, A.J.; Neumann, G.; Kawaoka, Y. At the centre: Influenza A virus ribonucleoproteins. Nat. Rev. Microbiol. 2015, 13, 28–41. [Google Scholar] [CrossRef] [PubMed]
- Neumann, G.; Brownlee, G.G.; Fodor, E.; Kawaoka, Y. Orthomyxovirus replication, transcription, and polyadenylation. Curr. Top. Microbiol. Immunol. 2004, 283, 121–143. [Google Scholar] [PubMed]
- Desselberger, U.; Racaniello, V.R.; Zazra, J.J.; Palese, P. The 3′ and 5′-terminal sequences of influenza A, B and C virus RNA segments are highly conserved and show partial inverted complementarity. Gene 1980, 8, 315–328. [Google Scholar] [CrossRef]
- Fodor, E.; Pritlove, D.C.; Brownlee, G.G. The influenza virus panhandle is involved in the initiation of transcription. J. Virol. 1994, 68, 4092–4096. [Google Scholar] [PubMed]
- Ferhadian, D.; Contrant, M.; Printz-Schweigert, A.; Smyth, R.P.; Paillart, J.C.; Marquet, R. Structural and Functional Motifs in Influenza Virus RNAs. Front. Microbiol. 2018, 9, 559. [Google Scholar] [CrossRef] [PubMed]
- Perez, D.R.; Angel, M.; Gonzalez-Reiche, A.S.; Santos, J.; Obadan, A.; Martinez-Sobrido, L. Plasmid-Based Reverse Genetics of Influenza A Virus. Methods Mol. Biol. 2017, 1602, 251–273. [Google Scholar] [PubMed]
- Neumann, G.; Watanabe, T.; Ito, H.; Watanabe, S.; Goto, H.; Gao, P.; Hughes, M.; Perez, D.R.; Donis, R.; Hoffmann, E.; et al. Generation of influenza A viruses entirely from cloned cDNAs. Proc. Natl. Acad. Sci. USA 1999, 96, 9345–9350. [Google Scholar] [CrossRef] [PubMed]
- Isel, C.; Munier, S.; Naffakh, N. Experimental Approaches to Study Genome Packaging of Influenza A Viruses. Viruses 2016, 8, 218. [Google Scholar] [CrossRef] [PubMed]
- Marsh, G.A.; Hatami, R.; Palese, P. Specific residues of the influenza A virus hemagglutinin viral RNA are important for efficient packaging into budding virions. J. Virol. 2007, 81, 9727–9736. [Google Scholar] [CrossRef] [PubMed]
- Basler, C.F.; Aguilar, P.V. Progress in identifying virulence determinants of the 1918 H1N1 and the Southeast Asian H5N1 influenza A viruses. Antiv. Res. 2008, 79, 166–178. [Google Scholar] [CrossRef] [PubMed]
- Pica, N.; Langlois, R.A.; Krammer, F.; Margine, I.; Palese, P. NS1-truncated live attenuated virus vaccine provides robust protection to aged mice from viral challenge. J. Virol. 2012, 86, 10293–10301. [Google Scholar] [CrossRef] [PubMed]
- Mueller, S.; Coleman, J.R.; Papamichail, D.; Ward, C.B.; Nimnual, A.; Futcher, B.; Skiena, S.; Wimmer, E. Live attenuated influenza virus vaccines by computer-aided rational design. Nat. Biotechnol. 2010, 28, 723–726. [Google Scholar] [CrossRef] [PubMed]
- Yang, C.; Skiena, S.; Futcher, B.; Mueller, S.; Wimmer, E. Deliberate reduction of hemagglutinin and neuraminidase expression of influenza virus leads to an ultraprotective live vaccine in mice. Proc. Natl. Acad. Sci. USA 2013, 110, 9481–9486. [Google Scholar] [CrossRef] [PubMed]
- Pena, L.; Sutton, T.; Chockalingam, A.; Kumar, S.; Angel, M.; Shao, H.; Chen, H.; Li, W.; Perez, D.R. Influenza viruses with rearranged genomes as live-attenuated vaccines. J. Virol. 2013, 87, 5118–5127. [Google Scholar] [CrossRef] [PubMed]
- Nogales, A.; DeDiego, M.L.; Topham, D.J.; Martinez-Sobrido, L. Rearrangement of Influenza Virus Spliced Segments for the Development of Live-Attenuated Vaccines. J. Virol. 2016, 90, 6291–6302. [Google Scholar] [CrossRef] [PubMed]
- Hai, R.; Garcia-Sastre, A.; Swayne, D.E.; Palese, P. A Reassortment-Incompetent Live Attenuated Influenza Virus Vaccine for Protection against Pandemic Virus Strains. J. Virol. 2011, 85, 6832–6843. [Google Scholar] [CrossRef] [PubMed]
- Hai, R.; Krammer, F.; Tan, G.S.; Pica, N.; Eggink, D.; Maamary, J.; Margine, I.; Albrecht, R.A.; Palese, P. Influenza viruses expressing chimeric hemagglutinins: Globular head and stalk domains derived from different subtypes. J. Virol. 2012, 86, 5774–5781. [Google Scholar] [CrossRef] [PubMed]
- Breen, M.; Nogales, A.; Baker, S.F.; Martinez-Sobrido, L. Replication-Competent Influenza A Viruses Expressing Reporter Genes. Viruses 2016, 8, 179. [Google Scholar] [CrossRef] [PubMed]
- Fukuyama, S.; Katsura, H.; Zhao, D.; Ozawa, M.; Ando, T.; Shoemaker, J.E.; Ishikawa, I.; Yamada, S.; Neumann, G.; Watanabe, S.; et al. Multi-spectral fluorescent reporter influenza viruses (Color-flu) as powerful tools for in vivo studies. Nat. Commun. 2015, 6, 6600. [Google Scholar] [CrossRef] [PubMed]
- Lo, M.K.; Nichol, S.T.; Spiropoulou, C.F. Evaluation of luciferase and GFP-expressing Nipah viruses for rapid quantitative antiviral screening. Antiv. Res. 2014, 106, 53–60. [Google Scholar] [CrossRef] [PubMed]
- Czako, R.; Vogel, L.; Lamirande, E.W.; Bock, K.W.; Moore, I.N.; Ellebedy, A.H.; Ahmed, R.; Mehle, A.; Subbarao, K. In Vivo Imaging of Influenza Virus Infection in Immunized Mice. mBio 2017, 8, e00714–e00717. [Google Scholar] [CrossRef] [PubMed]
- Heaton, N.S.; Leyva-Grado, V.H.; Tan, G.S.; Eggink, D.; Hai, R.; Palese, P. In vivo bioluminescent imaging of influenza a virus infection and characterization of novel cross-protective monoclonal antibodies. J. Virol. 2013, 87, 8272–8281. [Google Scholar] [CrossRef] [PubMed]
- Manicassamy, B.; Manicassamy, S.; Belicha-Villanueva, A.; Pisanelli, G.; Pulendran, B.; Garcia-Sastre, A. Analysis of in vivo dynamics of influenza virus infection in mice using a GFP reporter virus. Proc. Natl. Acad. Sci. USA 2010, 107, 11531–11536. [Google Scholar] [CrossRef] [PubMed]
- Tran, V.; Moser, L.A.; Poole, D.S.; Mehle, A. Highly sensitive real-time in vivo imaging of an influenza reporter virus reveals dynamics of replication and spread. J. Virol. 2013, 87, 13321–13329. [Google Scholar] [CrossRef] [PubMed]
- Reuther, P.; Gopfert, K.; Dudek, A.H.; Heiner, M.; Herold, S.; Schwemmle, M. Generation of a variety of stable Influenza A reporter viruses by genetic engineering of the NS gene segment. Sci. Rep. 2015, 5, 11346. [Google Scholar] [CrossRef] [PubMed]
- Cai, H.; Liu, M.; Russell, C.J. Directed evolution of an influenza reporter virus to restore replication and virulence and enhance non-invasive bioluminescence imaging in mice. J. Virol. 2018. [Google Scholar] [CrossRef] [PubMed]
- Li, P.; Cui, Q.; Wang, L.; Zhao, X.; Zhang, Y.; Manicassamy, B.; Yang, Y.; Rong, L.; Du, R. A Simple and Robust Approach for Evaluation of Antivirals Using a Recombinant Influenza Virus Expressing Gaussia Luciferase. Viruses 2018, 10, 325. [Google Scholar] [CrossRef] [PubMed]
- Gao, Q.; Chou, Y.Y.; Doganay, S.; Vafabakhsh, R.; Ha, T.; Palese, P. The influenza A virus PB2, PA, NP, and M segments play a pivotal role during genome packaging. J. Virol. 2012, 86, 7043–7051. [Google Scholar] [CrossRef] [PubMed]
- Widjaja, I.; de Vries, E.; Rottier, P.J.; de Haan, C.A. Competition between influenza A virus genome segments. PLoS ONE 2012, 7, e47529. [Google Scholar] [CrossRef] [PubMed]
- Falzarano, D.; Groseth, A.; Hoenen, T. Development and application of reporter-expressing mononegaviruses: Current challenges and perspectives. Antiv. Res. 2014, 103, 78–87. [Google Scholar] [CrossRef] [PubMed]
- Shi, X.; van Mierlo, J.T.; French, A.; Elliott, R.M. Visualizing the replication cycle of bunyamwera orthobunyavirus expressing fluorescent protein-tagged Gc glycoprotein. J. Virol. 2010, 84, 8460–8469. [Google Scholar] [CrossRef] [PubMed]
- Hotard, A.L.; Shaikh, F.Y.; Lee, S.; Yan, D.; Teng, M.N.; Plemper, R.K.; Crowe, J.E., Jr.; Moore, M.L. A stabilized respiratory syncytial virus reverse genetics system amenable to recombination-mediated mutagenesis. Virology 2012, 434, 129–136. [Google Scholar] [CrossRef] [PubMed]
- Fujii, Y.; Goto, H.; Watanabe, T.; Yoshida, T.; Kawaoka, Y. Selective incorporation of influenza virus RNA segments into virions. Proc. Natl. Acad. Sci. USA 2003, 100, 2002–2007. [Google Scholar] [CrossRef] [PubMed]
- Noda, T.; Kawaoka, Y. Structure of influenza virus ribonucleoprotein complexes and their packaging into virions. Rev. Med. Virol. 2010, 20, 380–391. [Google Scholar] [CrossRef] [PubMed]
- Noda, T.; Sugita, Y.; Aoyama, K.; Hirase, A.; Kawakami, E.; Miyazawa, A.; Sagara, H.; Kawaoka, Y. Three-dimensional analysis of ribonucleoprotein complexes in influenza A virus. Nat. Commun. 2012, 3, 639. [Google Scholar] [CrossRef] [PubMed]
- Fernandez-Sesma, A.; Marukian, S.; Ebersole, B.J.; Kaminski, D.; Park, M.S.; Yuen, T.; Sealfon, S.C.; Garcia-Sastre, A.; Moran, T.M. Influenza virus evades innate and adaptive immunity via the NS1 protein. J. Virol. 2006, 80, 6295–6304. [Google Scholar] [CrossRef] [PubMed]
- Fodor, E. The RNA polymerase of influenza A virus: Mechanisms of viral transcription and replication. Acta Virol. 2013, 57, 113–122. [Google Scholar] [CrossRef] [PubMed]
| Segments | Forward Primer | Reverse Primer |
|---|---|---|
| PB2 | 5’-GTGCCTTCATCTGGGTCT -3’ | 5’-TTGGGACATTTGATACCG -3’ |
| PB1 | 5’-AACTGGAGCACCGCAACT-3’ | 5’-AAGGAAAGCCATCGCCTC-3’ |
| PA | 5’-GATTCCTTTCGTCAGTCC-3’ | 5’-TCTAAGTGGTCGTGGTGT-3’ |
| HA | 5’-GAACTATTACTGGACCTTGC-3’ | 5’-CCTATTGTGACTGGGTGTAT-3’ |
| NP | 5’-GTATGGACCTGCCGTAGC-3’ | 5’-CTCTTGGGAGCACCTTCG-3’ |
| NA | 5’-AAACGGAGTAAAGGGATT-3’ | 5’-CAGGATGTTGAACGAAAC-3’ |
| M | 5’-CTTCTAACCGAGGTCGAAAC-3’ | 5’-CGTCTACGCTGCAGTCCTC-3’ |
| NS | 5’-ATTTCACCATTGCCTTCT-3’ | 5’-GGTCTCCCATTCTCATTAC-3’ |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhao, X.; Wang, L.; Cui, Q.; Li, P.; Wang, Y.; Zhang, Y.; Yang, Y.; Rong, L.; Du, R. A Mechanism Underlying Attenuation of Recombinant Influenza A Viruses Carrying Reporter Genes. Viruses 2018, 10, 679. https://doi.org/10.3390/v10120679
Zhao X, Wang L, Cui Q, Li P, Wang Y, Zhang Y, Yang Y, Rong L, Du R. A Mechanism Underlying Attenuation of Recombinant Influenza A Viruses Carrying Reporter Genes. Viruses. 2018; 10(12):679. https://doi.org/10.3390/v10120679
Chicago/Turabian StyleZhao, Xiujuan, Lin Wang, Qinghua Cui, Ping Li, Yanyan Wang, Yingying Zhang, Yong Yang, Lijun Rong, and Ruikun Du. 2018. "A Mechanism Underlying Attenuation of Recombinant Influenza A Viruses Carrying Reporter Genes" Viruses 10, no. 12: 679. https://doi.org/10.3390/v10120679
APA StyleZhao, X., Wang, L., Cui, Q., Li, P., Wang, Y., Zhang, Y., Yang, Y., Rong, L., & Du, R. (2018). A Mechanism Underlying Attenuation of Recombinant Influenza A Viruses Carrying Reporter Genes. Viruses, 10(12), 679. https://doi.org/10.3390/v10120679







