Research Models and Tools for the Identification of Antivirals and Therapeutics against Zika Virus Infection
Abstract
1. Introduction
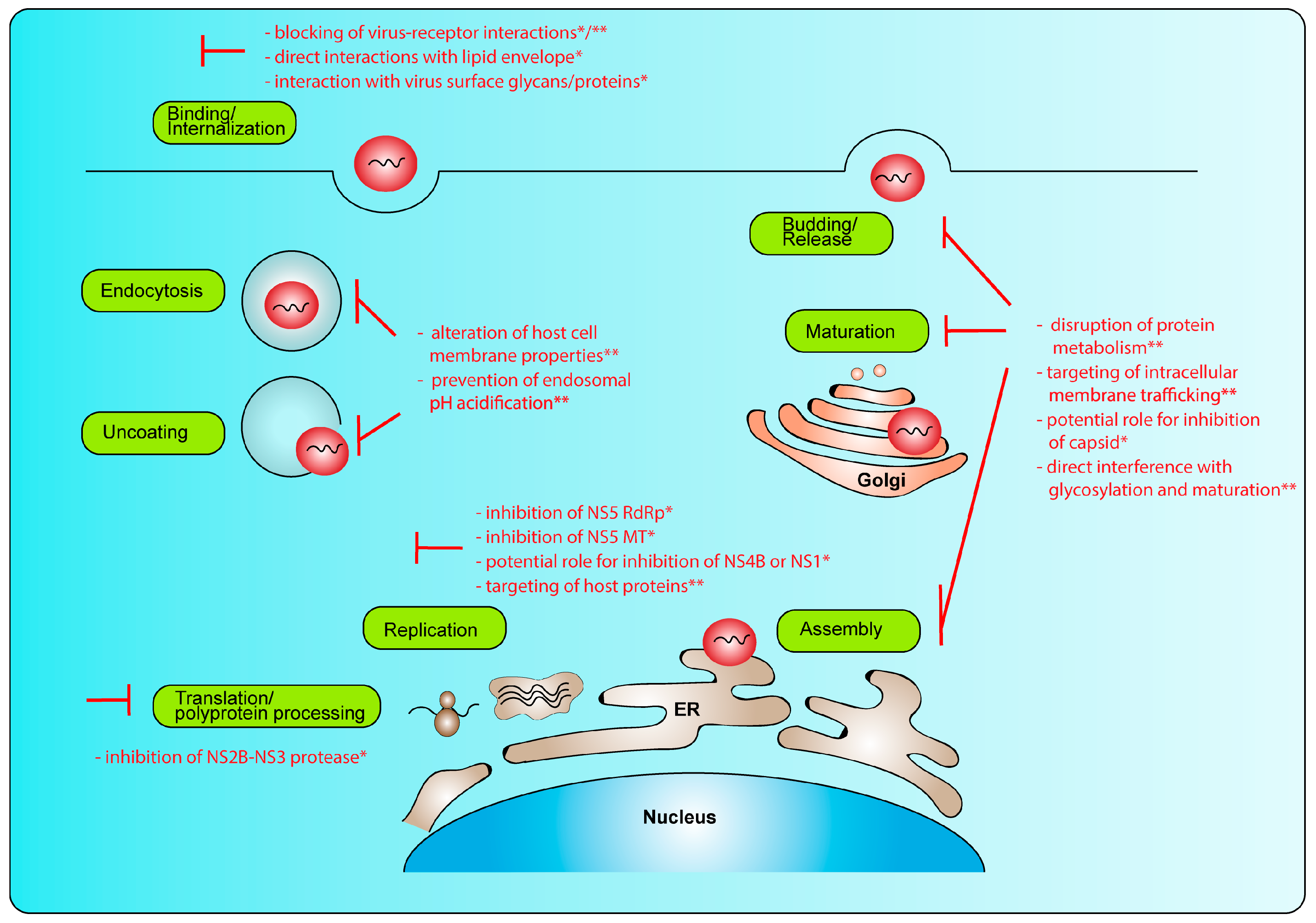
2. Replication Cycle and Potential Intervention Strategies
2.1. Genomic Organization
2.2. Replication Cycle
2.3. Intervention Strategies
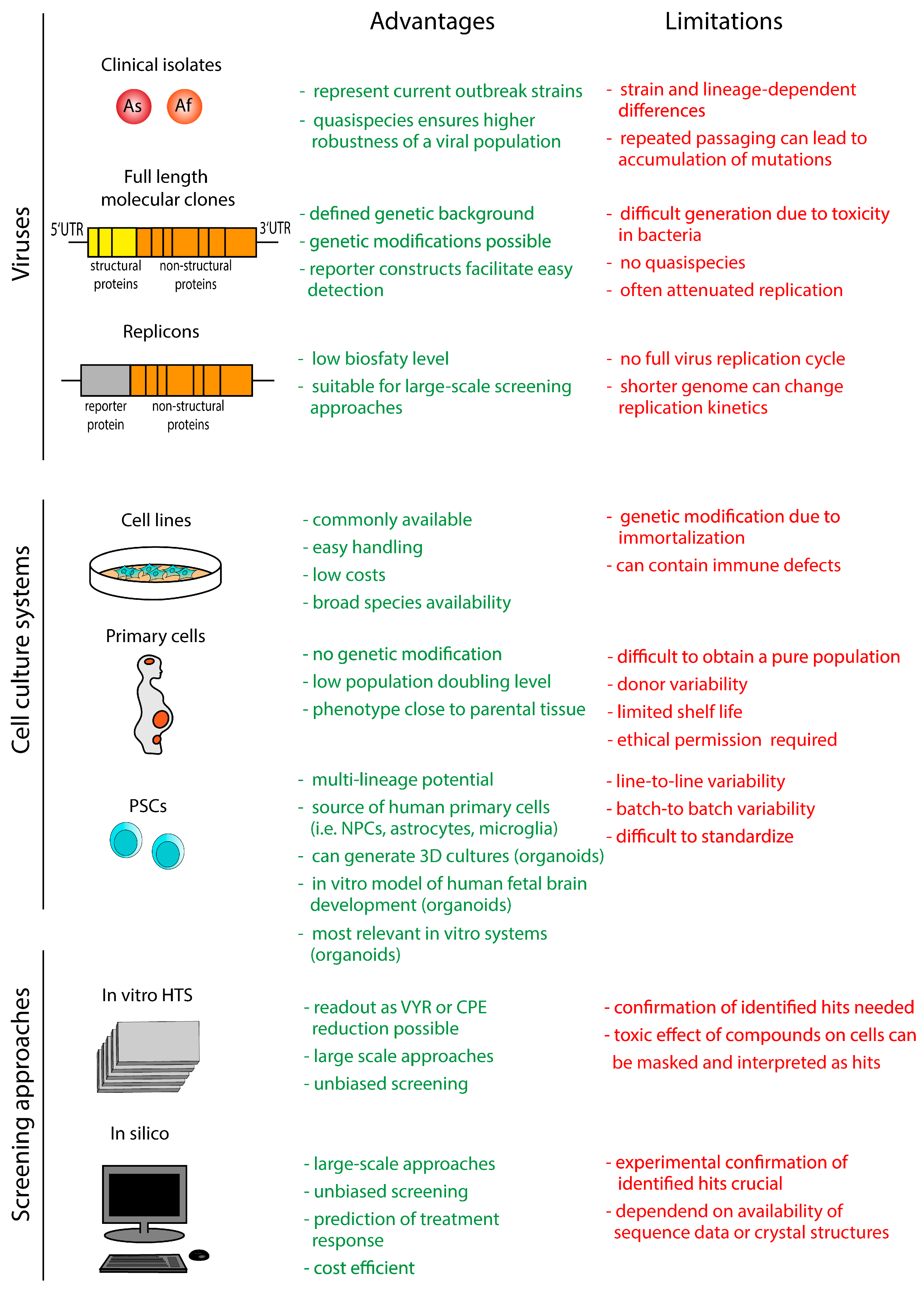
3. In Vitro Models and Screening Approaches
3.1. Viruses
3.1.1. Clinical Viral Isolates
3.1.2. Full-Length Infectious Clones
3.1.3. Replicons
3.2. Cell Culture Systems
3.2.1. Human and Animal Cell Lines
3.2.2. Human Primary Cell Cultures
3.2.3. Human Pluripotent Stem Cell-Derived Cultures
3.3. Screening Approaches
3.3.1. In Vitro Screening Approaches
3.3.2. In Silico Screening Approaches
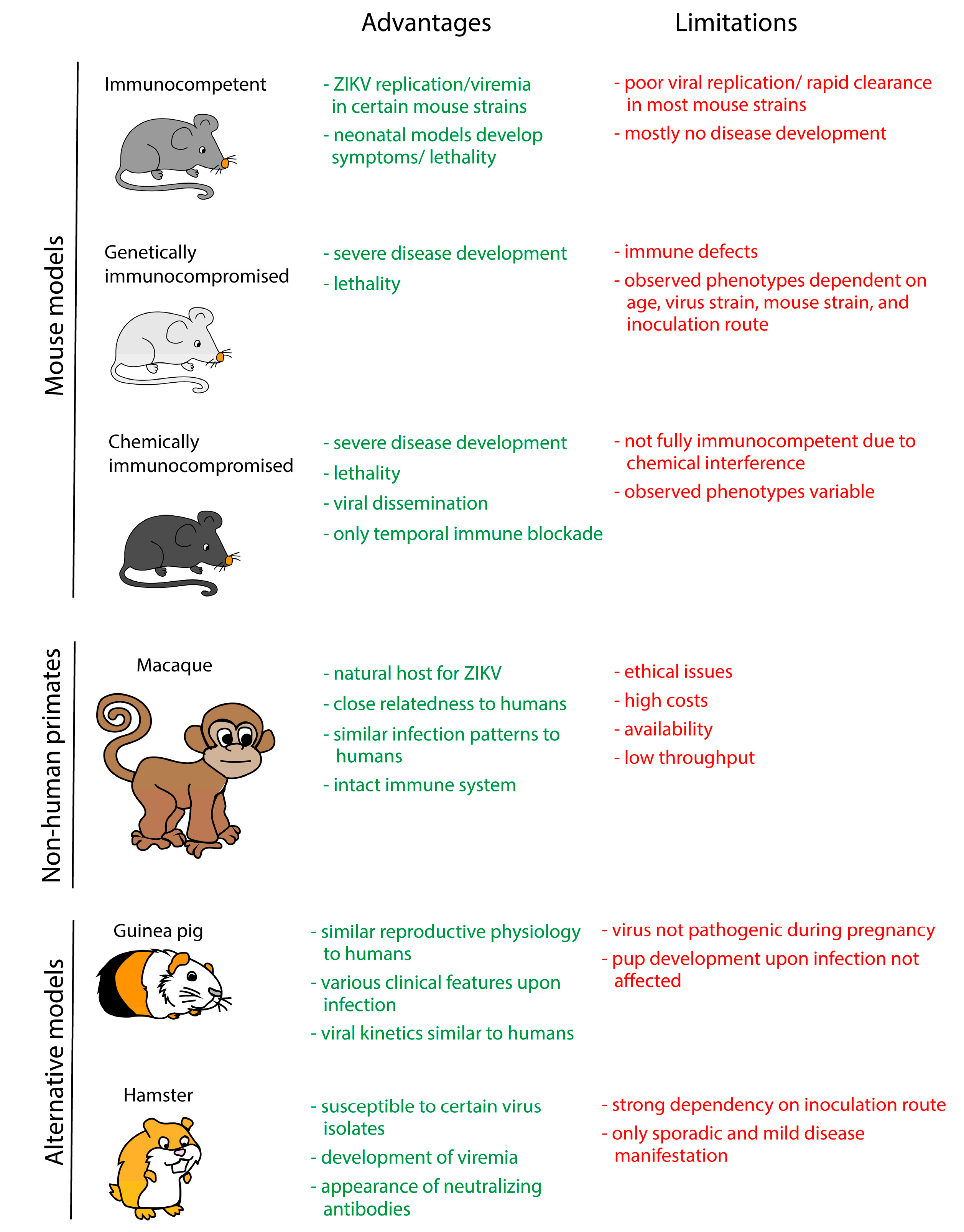
4. In Vivo Models
4.1. Mouse Models
4.2. Non-Human Primate Models
4.3. Alternative Animal Models
5. Concluding Remarks
Author Contributions
Funding
Conflicts of Interest
References
- Dick, G.W. Epidemiological notes on some viruses isolated in Uganda; Yellow fever, Rift Valley fever, Bwamba fever, West Nile, Mengo, Semliki forest, Bunyamwera, Ntaya, Uganda S and Zika viruses. Trans. R. Soc. Trop. Med. Hyg. 1953, 47, 13–48. [Google Scholar] [CrossRef]
- Dick, G.W.; Kitchen, S.F.; Haddow, A.J. Zika virus. I. Isolations and serological specificity. Trans. R. Soc. Trop. Med. Hyg. 1952, 46, 509–520. [Google Scholar] [CrossRef]
- Weinbren, M.P.; Williams, M.C. Zika virus: Further isolations in the Zika area, and some studies on the strains isolated. Trans. R. Soc. Trop. Med. Hyg. 1958, 52, 263–268. [Google Scholar] [CrossRef]
- Haddow, A.J.; Williams, M.C.; Woodall, J.P.; Simpson, D.I.; Goma, L.K. Twelve Isolations of Zika Virus from Aedes (Stegomyia) Africanus (Theobald) Taken in and above a Uganda Forest. Bull. World Health Organ. 1964, 31, 57–69. [Google Scholar] [PubMed]
- Simpson, D.I. Zika Virus Infection in Man. Trans. R. Soc. Trop. Med. Hyg. 1964, 58, 335–338. [Google Scholar] [CrossRef]
- Faye, O.; Freire, C.C.; Iamarino, A.; Faye, O.; de Oliveira, J.V.; Diallo, M.; Zanotto, P.M.; Sall, A.A. Molecular evolution of Zika virus during its emergence in the 20(th) century. PLoS Negl. Trop. Dis. 2014, 8, e2636. [Google Scholar] [CrossRef] [PubMed]
- Duffy, M.R.; Chen, T.H.; Hancock, W.T.; Powers, A.M.; Kool, J.L.; Lanciotti, R.S.; Pretrick, M.; Marfel, M.; Holzbauer, S.; Dubray, C.; et al. Zika virus outbreak on Yap Island, Federated States of Micronesia. N. Engl. J. Med. 2009, 360, 2536–2543. [Google Scholar] [CrossRef] [PubMed]
- Cao-Lormeau, V.M.; Roche, C.; Teissier, A.; Robin, E.; Berry, A.L.; Mallet, H.P.; Sall, A.A.; Musso, D. Zika virus, French polynesia, South pacific, 2013. Emerg. Infect. Dis. 2014, 20, 1085–1086. [Google Scholar] [CrossRef] [PubMed]
- Jouannic, J.M.; Friszer, S.; Leparc-Goffart, I.; Garel, C.; Eyrolle-Guignot, D. Zika virus infection in French Polynesia. Lancet 2016, 387, 1051–1052. [Google Scholar] [CrossRef]
- Campos, G.S.; Bandeira, A.C.; Sardi, S.I. Zika Virus Outbreak, Bahia, Brazil. Emerg. Infect. Dis. 2015, 21, 1885–1886. [Google Scholar] [CrossRef] [PubMed]
- Schuler-Faccini, L.; Ribeiro, E.M.; Feitosa, I.M.; Horovitz, D.D.; Cavalcanti, D.P.; Pessoa, A.; Doriqui, M.J.; Neri, J.I.; Neto, J.M.; Wanderley, H.Y.; et al. Possible Association Between Zika Virus Infection and Microcephaly—Brazil, 2015. MMWR Morb. Mortal. Wkly. Rep. 2016, 65, 59–62. [Google Scholar] [CrossRef] [PubMed]
- Martines, R.B.; Bhatnagar, J.; Keating, M.K.; Silva-Flannery, L.; Muehlenbachs, A.; Gary, J.; Goldsmith, C.; Hale, G.; Ritter, J.; Rollin, D.; et al. Notes from the Field: Evidence of Zika Virus Infection in Brain and Placental Tissues from Two Congenitally Infected Newborns and Two Fetal Losses—Brazil, 2015. MMWR Morb. Mortal. Wkly. Rep. 2016, 65, 159–160. [Google Scholar] [CrossRef] [PubMed]
- Boyer, S.; Calvez, E.; Chouin-Carneiro, T.; Diallo, D.; Failloux, A.B. An overview of mosquito vectors of Zika virus. Microbes Infect. 2018. [Google Scholar] [CrossRef] [PubMed]
- Grischott, F.; Puhan, M.; Hatz, C.; Schlagenhauf, P. Non-vector-borne transmission of Zika virus: A systematic review. Travel Med. Infect. Dis. 2016, 14, 313–330. [Google Scholar] [CrossRef] [PubMed]
- Hills, S.L.; Russell, K.; Hennessey, M.; Williams, C.; Oster, A.M.; Fischer, M.; Mead, P. Transmission of Zika Virus Through Sexual Contact with Travelers to Areas of Ongoing Transmission—Continental United States, 2016. MMWR Morb. Mortal. Wkly. Rep. 2016, 65, 215–216. [Google Scholar] [CrossRef] [PubMed]
- Musso, D.; Nhan, T.; Robin, E.; Roche, C.; Bierlaire, D.; Zisou, K.; Shan Yan, A.; Cao-Lormeau, V.M.; Broult, J. Potential for Zika virus transmission through blood transfusion demonstrated during an outbreak in French Polynesia, November 2013 to February 2014. Euro Surveill 2014, 19, 20761. [Google Scholar] [CrossRef] [PubMed]
- Gallian, P.; Cabie, A.; Richard, P.; Paturel, L.; Charrel, R.N.; Pastorino, B.; Leparc-Goffart, I.; Tiberghien, P.; de Lamballerie, X. Zika virus in asymptomatic blood donors in Martinique. Blood 2017, 129, 263–266. [Google Scholar] [CrossRef] [PubMed]
- Blohm, G.M.; Lednicky, J.A.; Marquez, M.; White, S.K.; Loeb, J.C.; Pacheco, C.A.; Nolan, D.J.; Paisie, T.; Salemi, M.; Rodriguez-Morales, A.J.; et al. Evidence for Mother-to-Child Transmission of Zika Virus Through Breast Milk. Clin. Infect. Dis. 2018, 66, 1120–1121. [Google Scholar] [CrossRef] [PubMed]
- Tan, J.J.L.; Balne, P.K.; Leo, Y.S.; Tong, L.; Ng, L.F.P.; Agrawal, R. Persistence of Zika virus in conjunctival fluid of convalescence patients. Sci. Rep. 2017, 7, 11194. [Google Scholar] [CrossRef] [PubMed]
- Petersen, L.R.; Jamieson, D.J.; Powers, A.M.; Honein, M.A. Zika Virus. N. Engl. J. Med. 2016, 374, 1552–1563. [Google Scholar] [CrossRef] [PubMed]
- Cao-Lormeau, V.M.; Blake, A.; Mons, S.; Lastere, S.; Roche, C.; Vanhomwegen, J.; Dub, T.; Baudouin, L.; Teissier, A.; Larre, P.; et al. Guillain-Barre Syndrome outbreak associated with Zika virus infection in French Polynesia: A case-control study. Lancet 2016, 387, 1531–1539. [Google Scholar] [CrossRef]
- Carteaux, G.; Maquart, M.; Bedet, A.; Contou, D.; Brugieres, P.; Fourati, S.; Cleret de Langavant, L.; de Broucker, T.; Brun-Buisson, C.; Leparc-Goffart, I.; et al. Zika Virus Associated with Meningoencephalitis. N. Engl. J. Med. 2016, 374, 1595–1596. [Google Scholar] [CrossRef] [PubMed]
- Mecharles, S.; Herrmann, C.; Poullain, P.; Tran, T.H.; Deschamps, N.; Mathon, G.; Landais, A.; Breurec, S.; Lannuzel, A. Acute myelitis due to Zika virus infection. Lancet 2016, 387, 1481. [Google Scholar] [CrossRef]
- Costello, A.; Dua, T.; Duran, P.; Gulmezoglu, M.; Oladapo, O.T.; Perea, W.; Pires, J.; Ramon-Pardo, P.; Rollins, N.; Saxena, S. Defining the syndrome associated with congenital Zika virus infection. Bull. World Health Organ. 2016, 94, 406–406A. [Google Scholar] [CrossRef] [PubMed]
- Kuno, G.; Chang, G.J. Full-length sequencing and genomic characterization of Bagaza, Kedougou, and Zika viruses. Arch. Virol. 2007, 152, 687–696. [Google Scholar] [CrossRef] [PubMed]
- Chambers, T.J.; Hahn, C.S.; Galler, R.; Rice, C.M. Flavivirus genome organization, expression, and replication. Annu. Rev. Microbiol. 1990, 44, 649–688. [Google Scholar] [CrossRef] [PubMed]
- Bressanelli, S.; Stiasny, K.; Allison, S.L.; Stura, E.A.; Duquerroy, S.; Lescar, J.; Heinz, F.X.; Rey, F.A. Structure of a flavivirus envelope glycoprotein in its low-pH-induced membrane fusion conformation. EMBO J. 2004, 23, 728–738. [Google Scholar] [CrossRef] [PubMed]
- Cortese, M.; Goellner, S.; Acosta, E.G.; Neufeldt, C.J.; Oleksiuk, O.; Lampe, M.; Haselmann, U.; Funaya, C.; Schieber, N.; Ronchi, P.; et al. Ultrastructural Characterization of Zika Virus Replication Factories. Cell Rep. 2017, 18, 2113–2123. [Google Scholar] [CrossRef] [PubMed]
- Hasan, S.S.; Sevvana, M.; Kuhn, R.J.; Rossmann, M.G. Structural biology of Zika virus and other flaviviruses. Nat. Struct. Mol. Biol. 2018, 25, 13–20. [Google Scholar] [CrossRef] [PubMed]
- Mottin, M.; Borba, J.; Braga, R.C.; Torres, P.H.M.; Martini, M.C.; Proenca-Modena, J.L.; Judice, C.C.; Costa, F.T.M.; Ekins, S.; Perryman, A.L.; et al. The A-Z of Zika drug discovery. Drug Discov. Today 2018. [Google Scholar] [CrossRef] [PubMed]
- da Silva, S.; Oliveira Silva Martins, D.; Jardim, A.C.G. A Review of the Ongoing Research on Zika Virus Treatment. Viruses 2018, 10, 255. [Google Scholar] [CrossRef] [PubMed]
- Qadir, A.; Riaz, M.; Saeed, M.; Shahzad-Ul-Hussan, S. Potential targets for therapeutic intervention and structure based vaccine design against Zika virus. Eur. J. Med. Chem. 2018, 156, 444–460. [Google Scholar] [CrossRef] [PubMed]
- Carvalho, C.A.M.; Casseb, S.M.M.; Goncalves, R.B.; Silva, E.V.P.; Gomes, A.M.O.; Vasconcelos, P.F.C. Bovine lactoferrin activity against Chikungunya and Zika viruses. J. Gen. Virol. 2017, 98, 1749–1754. [Google Scholar] [CrossRef] [PubMed]
- Rausch, K.; Hackett, B.A.; Weinbren, N.L.; Reeder, S.M.; Sadovsky, Y.; Hunter, C.A.; Schultz, D.C.; Coyne, C.B.; Cherry, S. Screening Bioactives Reveals Nanchangmycin as a Broad Spectrum Antiviral Active against Zika Virus. Cell Rep. 2017, 18, 804–815. [Google Scholar] [CrossRef] [PubMed]
- Carneiro, B.M.; Batista, M.N.; Braga, A.C.S.; Nogueira, M.L.; Rahal, P. The green tea molecule EGCG inhibits Zika virus entry. Virology 2016, 496, 215–218. [Google Scholar] [CrossRef] [PubMed]
- Tricarico, P.M.; Caracciolo, I.; Gratton, R.; D’Agaro, P.; Crovella, S. 25-hydroxycholesterol reduces inflammation, viral load and cell death in ZIKV-infected U-87 MG glial cell line. Inflammopharmacology 2018. [Google Scholar] [CrossRef] [PubMed]
- Delvecchio, R.; Higa, L.M.; Pezzuto, P.; Valadao, A.L.; Garcez, P.P.; Monteiro, F.L.; Loiola, E.C.; Dias, A.A.; Silva, F.J.; Aliota, M.T.; et al. Chloroquine, an Endocytosis Blocking Agent, Inhibits Zika Virus Infection in Different Cell Models. Viruses 2016, 8, 322. [Google Scholar] [CrossRef] [PubMed]
- Xu, M.; Lee, E.M.; Wen, Z.; Cheng, Y.; Huang, W.K.; Qian, X.; Tcw, J.; Kouznetsova, J.; Ogden, S.C.; Hammack, C.; et al. Identification of small-molecule inhibitors of Zika virus infection and induced neural cell death via a drug repurposing screen. Nat. Med. 2016, 22, 1101–1107. [Google Scholar] [CrossRef] [PubMed]
- Shiryaev, S.A.; Mesci, P.; Pinto, A.; Fernandes, I.; Sheets, N.; Shresta, S.; Farhy, C.; Huang, C.T.; Strongin, A.Y.; Muotri, A.R.; et al. Repurposing of the anti-malaria drug chloroquine for Zika Virus treatment and prophylaxis. Sci. Rep. 2017, 7, 15771. [Google Scholar] [CrossRef] [PubMed]
- Li, C.; Zhu, X.; Ji, X.; Quanquin, N.; Deng, Y.Q.; Tian, M.; Aliyari, R.; Zuo, X.; Yuan, L.; Afridi, S.K.; et al. Chloroquine, a FDA-approved Drug, Prevents Zika Virus Infection and its Associated Congenital Microcephaly in Mice. EBioMedicine 2017, 24, 189–194. [Google Scholar] [CrossRef] [PubMed]
- Kuivanen, S.; Bespalov, M.M.; Nandania, J.; Ianevski, A.; Velagapudi, V.; De Brabander, J.K.; Kainov, D.E.; Vapalahti, O. Obatoclax, saliphenylhalamide and gemcitabine inhibit Zika virus infection in vitro and differentially affect cellular signaling, transcription and metabolism. Antiviral Res. 2017, 139, 117–128. [Google Scholar] [CrossRef] [PubMed]
- Byler, K.G.; Ogungbe, I.V.; Setzer, W.N. In-silico screening for anti-Zika virus phytochemicals. J. Mol. Graph. Model. 2016, 69, 78–91. [Google Scholar] [CrossRef] [PubMed]
- Lim, S.P.; Wang, Q.Y.; Noble, C.G.; Chen, Y.L.; Dong, H.; Zou, B.; Yokokawa, F.; Nilar, S.; Smith, P.; Beer, D.; et al. Ten years of dengue drug discovery: Progress and prospects. Antiviral Res. 2013, 100, 500–519. [Google Scholar] [CrossRef] [PubMed]
- Coutard, B.; Barral, K.; Lichiere, J.; Selisko, B.; Martin, B.; Aouadi, W.; Lombardia, M.O.; Debart, F.; Vasseur, J.J.; Guillemot, J.C.; et al. Zika Virus Methyltransferase: Structure and Functions for Drug Design Perspectives. J. Virol. 2017, 91, e02202-16. [Google Scholar] [CrossRef] [PubMed]
- Stephen, P.; Baz, M.; Boivin, G.; Lin, S.X. Structural Insight into NS5 of Zika Virus Leading to the Discovery of MTase Inhibitors. J. Am. Chem. Soc. 2016, 138, 16212–16215. [Google Scholar] [CrossRef] [PubMed]
- Zhang, C.; Feng, T.; Cheng, J.; Li, Y.; Yin, X.; Zeng, W.; Jin, X.; Li, Y.; Guo, F.; Jin, T. Structure of the NS5 methyltransferase from Zika virus and implications in inhibitor design. Biochem. Biophys. Res. Commun. 2017, 492, 624–630. [Google Scholar] [CrossRef] [PubMed]
- Duan, W.; Song, H.; Wang, H.; Chai, Y.; Su, C.; Qi, J.; Shi, Y.; Gao, G.F. The crystal structure of Zika virus NS5 reveals conserved drug targets. EMBO J. 2017, 36, 919–933. [Google Scholar] [CrossRef] [PubMed]
- Hercik, K.; Brynda, J.; Nencka, R.; Boura, E. Structural basis of Zika virus methyltransferase inhibition by sinefungin. Arch. Virol. 2017, 162, 2091–2096. [Google Scholar] [CrossRef] [PubMed]
- Xie, X.; Wang, Q.Y.; Xu, H.Y.; Qing, M.; Kramer, L.; Yuan, Z.; Shi, P.Y. Inhibition of dengue virus by targeting viral NS4B protein. J. Virol. 2011, 85, 11183–11195. [Google Scholar] [CrossRef] [PubMed]
- van Cleef, K.W.; Overheul, G.J.; Thomassen, M.C.; Kaptein, S.J.; Davidson, A.D.; Jacobs, M.; Neyts, J.; van Kuppeveld, F.J.; van Rij, R.P. Identification of a new dengue virus inhibitor that targets the viral NS4B protein and restricts genomic RNA replication. Antiviral Res. 2013, 99, 165–171. [Google Scholar] [CrossRef] [PubMed]
- Hernandez-Morales, I.; Geluykens, P.; Clynhens, M.; Strijbos, R.; Goethals, O.; Megens, S.; Verheyen, N.; Last, S.; McGowan, D.; Coesemans, E.; et al. Characterization of a dengue NS4B inhibitor originating from an HCV small molecule library. Antiviral Res. 2017, 147, 149–158. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.; Liu, Y.; Guo, J.; Wang, P.; Zhang, L.; Xiao, G.; Wang, W. Screening of FDA-Approved Drugs for Inhibitors of Japanese Encephalitis Virus Infection. J. Virol. 2017, 91. [Google Scholar] [CrossRef] [PubMed]
- Watterson, D.; Modhiran, N.; Young, P.R. The many faces of the flavivirus NS1 protein offer a multitude of options for inhibitor design. Antiviral Res. 2016, 130, 7–18. [Google Scholar] [CrossRef] [PubMed]
- Bekerman, E.; Neveu, G.; Shulla, A.; Brannan, J.; Pu, S.Y.; Wang, S.; Xiao, F.; Barouch-Bentov, R.; Bakken, R.R.; Mateo, R.; et al. Anticancer kinase inhibitors impair intracellular viral trafficking and exert broad-spectrum antiviral effects. J. Clin. Investig. 2017, 127, 1338–1352. [Google Scholar] [CrossRef] [PubMed]
- Albulescu, I.C.; Kovacikova, K.; Tas, A.; Snijder, E.J.; van Hemert, M.J. Suramin inhibits Zika virus replication by interfering with virus attachment and release of infectious particles. Antiviral Res. 2017, 143, 230–236. [Google Scholar] [CrossRef] [PubMed]
- Kota, S.; Takahashi, V.; Ni, F.; Snyder, J.K.; Strosberg, A.D. Direct binding of a hepatitis C virus inhibitor to the viral capsid protein. PLoS ONE 2012, 7, e32207. [Google Scholar] [CrossRef] [PubMed]
- Scaturro, P.; Trist, I.M.; Paul, D.; Kumar, A.; Acosta, E.G.; Byrd, C.M.; Jordan, R.; Brancale, A.; Bartenschlager, R. Characterization of the mode of action of a potent dengue virus capsid inhibitor. J. Virol. 2014, 88, 11540–11555. [Google Scholar] [CrossRef] [PubMed]
- Oliveira, E.R.A.; Mohana-Borges, R.; de Alencastro, R.B.; Horta, B.A.C. The flavivirus capsid protein: Structure, function and perspectives towards drug design. Virus Res. 2017, 227, 115–123. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Valderramos, S.G.; Wu, A.; Ouyang, S.; Li, C.; Brasil, P.; Bonaldo, M.; Coates, T.; Nielsen-Saines, K.; Jiang, T.; et al. From Mosquitos to Humans: Genetic Evolution of Zika Virus. Cell Host Microbe 2016, 19, 561–565. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Zhao, H.; Oliva, S.M.; Zhu, H. Modeling the transmission and control of Zika in Brazil. Sci. Rep. 2017, 7, 7721. [Google Scholar] [CrossRef] [PubMed]
- Haddow, A.D.; Schuh, A.J.; Yasuda, C.Y.; Kasper, M.R.; Heang, V.; Huy, R.; Guzman, H.; Tesh, R.B.; Weaver, S.C. Genetic characterization of Zika virus strains: Geographic expansion of the Asian lineage. PLoS Negl. Trop. Dis. 2012, 6, e1477. [Google Scholar] [CrossRef] [PubMed]
- Beaver, J.T.; Lelutiu, N.; Habib, R.; Skountzou, I. Evolution of Two Major Zika Virus Lineages: Implications for Pathology, Immune Response, and Vaccine Development. Front. Immunol. 2018, 9, 1640. [Google Scholar] [CrossRef] [PubMed]
- Pettersson, J.H.; Eldholm, V.; Seligman, S.J.; Lundkvist, A.; Falconar, A.K.; Gaunt, M.W.; Musso, D.; Nougairede, A.; Charrel, R.; Gould, E.A.; et al. How Did Zika Virus Emerge in the Pacific Islands and Latin America? MBio 2016, 7, e01239-16. [Google Scholar] [CrossRef] [PubMed]
- Anfasa, F.; Siegers, J.Y.; van der Kroeg, M.; Mumtaz, N.; Stalin Raj, V.; de Vrij, F.M.S.; Widagdo, W.; Gabriel, G.; Salinas, S.; Simonin, Y.; et al. Phenotypic Differences between Asian and African Lineage Zika Viruses in Human Neural Progenitor Cells. mSphere 2017, 2, e00292-17. [Google Scholar] [CrossRef] [PubMed]
- Simonin, Y.; van Riel, D.; Van de Perre, P.; Rockx, B.; Salinas, S. Differential virulence between Asian and African lineages of Zika virus. PLoS Negl. Trop. Dis. 2017, 11, e0005821. [Google Scholar] [CrossRef] [PubMed]
- Hamel, R.; Ferraris, P.; Wichit, S.; Diop, F.; Talignani, L.; Pompon, J.; Garcia, D.; Liegeois, F.; Sall, A.A.; Yssel, H.; et al. African and Asian Zika virus strains differentially induce early antiviral responses in primary human astrocytes. Infect. Genet. Evol. 2017, 49, 134–137. [Google Scholar] [CrossRef] [PubMed]
- Zhang, F.; Wang, H.J.; Wang, Q.; Liu, Z.Y.; Yuan, L.; Huang, X.Y.; Li, G.; Ye, Q.; Yang, H.; Shi, L.; et al. American Strain of Zika Virus Causes More Severe Microcephaly Than an Old Asian Strain in Neonatal Mice. EBioMedicine 2017, 25, 95–105. [Google Scholar] [CrossRef] [PubMed]
- Smith, D.R.; Sprague, T.R.; Hollidge, B.S.; Valdez, S.M.; Padilla, S.L.; Bellanca, S.A.; Golden, J.W.; Coyne, S.R.; Kulesh, D.A.; Miller, L.J.; et al. African and Asian Zika Virus Isolates Display Phenotypic Differences Both In Vitro and In Vivo. Am. J. Trop. Med. Hyg. 2018, 98, 432–444. [Google Scholar] [CrossRef] [PubMed]
- Simonin, Y.; Loustalot, F.; Desmetz, C.; Foulongne, V.; Constant, O.; Fournier-Wirth, C.; Leon, F.; Moles, J.P.; Goubaud, A.; Lemaitre, J.M.; et al. Zika Virus Strains Potentially Display Different Infectious Profiles in Human Neural Cells. EBioMedicine 2016, 12, 161–169. [Google Scholar] [CrossRef] [PubMed]
- Tripathi, S.; Balasubramaniam, V.R.; Brown, J.A.; Mena, I.; Grant, A.; Bardina, S.V.; Maringer, K.; Schwarz, M.C.; Maestre, A.M.; Sourisseau, M.; et al. A novel Zika virus mouse model reveals strain specific differences in virus pathogenesis and host inflammatory immune responses. PLoS Pathog. 2017, 13, e1006258. [Google Scholar] [CrossRef] [PubMed]
- Dowall, S.D.; Graham, V.A.; Rayner, E.; Hunter, L.; Atkinson, B.; Pearson, G.; Dennis, M.; Hewson, R. Lineage-dependent differences in the disease progression of Zika virus infection in type-I interferon receptor knockout (A129) mice. PLoS Negl. Trop. Dis. 2017, 11, e0005704. [Google Scholar] [CrossRef] [PubMed]
- Shao, Q.; Herrlinger, S.; Zhu, Y.N.; Yang, M.; Goodfellow, F.; Stice, S.L.; Qi, X.P.; Brindley, M.A.; Chen, J.F. The African Zika virus MR-766 is more virulent and causes more severe brain damage than current Asian lineage and dengue virus. Development 2017, 144, 4114–4124. [Google Scholar] [CrossRef] [PubMed]
- Vielle, N.J.; Zumkehr, B.; Garcia-Nicolas, O.; Blank, F.; Stojanov, M.; Musso, D.; Baud, D.; Summerfield, A.; Alves, M.P. Silent infection of human dendritic cells by African and Asian strains of Zika virus. Sci. Rep. 2018, 8, 5440. [Google Scholar] [CrossRef] [PubMed]
- Meda, N.; Salinas, S.; Kagone, T.; Simonin, Y.; Van de Perre, P. Zika virus epidemic: Africa should not be neglected. Lancet 2016, 388, 337–338. [Google Scholar] [CrossRef]
- Wetsman, N. The missing pieces: Lack of Zika data from Africa complicates search for answers. Nat. Med. 2017, 23, 904–906. [Google Scholar] [CrossRef] [PubMed]
- Aubry, F.; Nougairede, A.; Gould, E.A.; de Lamballerie, X. Flavivirus reverse genetic systems, construction techniques and applications: A historical perspective. Antiviral Res. 2015, 114, 67–85. [Google Scholar] [CrossRef] [PubMed]
- Ruggli, N.; Rice, C.M. Functional cDNA clones of the Flaviviridae: Strategies and applications. Adv. Virus Res. 1999, 53, 183–207. [Google Scholar] [PubMed]
- Tsetsarkin, K.A.; Kenney, H.; Chen, R.; Liu, G.; Manukyan, H.; Whitehead, S.S.; Laassri, M.; Chumakov, K.; Pletnev, A.G. A Full-Length Infectious cDNA Clone of Zika Virus from the 2015 Epidemic in Brazil as a Genetic Platform for Studies of Virus-Host Interactions and Vaccine Development. MBio 2016, 7, e01114-16. [Google Scholar] [CrossRef] [PubMed]
- Schwarz, M.C.; Sourisseau, M.; Espino, M.M.; Gray, E.S.; Chambers, M.T.; Tortorella, D.; Evans, M.J. Rescue of the 1947 Zika Virus Prototype Strain with a Cytomegalovirus Promoter-Driven cDNA Clone. mSphere 2016, 1, e00246-16. [Google Scholar] [CrossRef] [PubMed]
- Shan, C.; Xie, X.; Muruato, A.E.; Rossi, S.L.; Roundy, C.M.; Azar, S.R.; Yang, Y.; Tesh, R.B.; Bourne, N.; Barrett, A.D.; et al. An Infectious cDNA Clone of Zika Virus to Study Viral Virulence, Mosquito Transmission, and Antiviral Inhibitors. Cell Host Microbe 2016, 19, 891–900. [Google Scholar] [CrossRef] [PubMed]
- Yun, S.I.; Song, B.H.; Frank, J.C.; Julander, J.G.; Olsen, A.L.; Polejaeva, I.A.; Davies, C.J.; White, K.L.; Lee, Y.M. Functional Genomics and Immunologic Tools: The Impact of Viral and Host Genetic Variations on the Outcome of Zika Virus Infection. Viruses 2018, 10, 422. [Google Scholar] [CrossRef] [PubMed]
- Widman, D.G.; Young, E.; Yount, B.L.; Plante, K.S.; Gallichotte, E.N.; Carbaugh, D.L.; Peck, K.M.; Plante, J.; Swanstrom, J.; Heise, M.T.; et al. A Reverse Genetics Platform That Spans the Zika Virus Family Tree. MBio 2017, 8, e02014-16. [Google Scholar] [CrossRef] [PubMed]
- Mutso, M.; Saul, S.; Rausalu, K.; Susova, O.; Zusinaite, E.; Mahalingam, S.; Merits, A. Reverse genetic system, genetically stable reporter viruses and packaged subgenomic replicon based on a Brazilian Zika virus isolate. J. Gen. Virol. 2017, 98, 2712–2724. [Google Scholar] [CrossRef] [PubMed]
- Aubry, F.; Nougairede, A.; de Fabritus, L.; Querat, G.; Gould, E.A.; de Lamballerie, X. Single-stranded positive-sense RNA viruses generated in days using infectious subgenomic amplicons. J. Gen. Virol. 2014, 95, 2462–2467. [Google Scholar] [CrossRef] [PubMed]
- Gadea, G.; Bos, S.; Krejbich-Trotot, P.; Clain, E.; Viranaicken, W.; El-Kalamouni, C.; Mavingui, P.; Despres, P. A robust method for the rapid generation of recombinant Zika virus expressing the GFP reporter gene. Virology 2016, 497, 157–162. [Google Scholar] [CrossRef] [PubMed]
- Atieh, T.; Baronti, C.; de Lamballerie, X.; Nougairede, A. Simple reverse genetics systems for Asian and African Zika viruses. Sci. Rep. 2016, 6, 39384. [Google Scholar] [CrossRef] [PubMed]
- Bos, S.; Viranaicken, W.; Turpin, J.; El-Kalamouni, C.; Roche, M.; Krejbich-Trotot, P.; Despres, P.; Gadea, G. The structural proteins of epidemic and historical strains of Zika virus differ in their ability to initiate viral infection in human host cells. Virology 2018, 516, 265–273. [Google Scholar] [CrossRef] [PubMed]
- Quan, J.; Tian, J. Circular polymerase extension cloning. Methods Mol. Biol. 2014, 1116, 103–117. [Google Scholar] [CrossRef] [PubMed]
- Setoh, Y.X.; Prow, N.A.; Peng, N.; Hugo, L.E.; Devine, G.; Hazlewood, J.E.; Suhrbier, A.; Khromykh, A.A. De Novo Generation and Characterization of New Zika Virus Isolate Using Sequence Data from a Microcephaly Case. mSphere 2017, 2, e00190-17. [Google Scholar] [CrossRef] [PubMed]
- Weger-Lucarelli, J.; Duggal, N.K.; Bullard-Feibelman, K.; Veselinovic, M.; Romo, H.; Nguyen, C.; Ruckert, C.; Brault, A.C.; Bowen, R.A.; Stenglein, M.; et al. Development and Characterization of Recombinant Virus Generated from a New World Zika Virus Infectious Clone. J. Virol. 2017, 91, e01765-16. [Google Scholar] [CrossRef] [PubMed]
- Munster, M.; Plaszczyca, A.; Cortese, M.; Neufeldt, C.J.; Goellner, S.; Long, G.; Bartenschlager, R. A Reverse Genetics System for Zika Virus Based on a Simple Molecular Cloning Strategy. Viruses 2018, 10, 368. [Google Scholar] [CrossRef] [PubMed]
- de Visser, J.A.; Hermisson, J.; Wagner, G.P.; Ancel Meyers, L.; Bagheri-Chaichian, H.; Blanchard, J.L.; Chao, L.; Cheverud, J.M.; Elena, S.F.; Fontana, W.; et al. Perspective: Evolution and detection of genetic robustness. Evolution 2003, 57, 1959–1972. [Google Scholar] [CrossRef] [PubMed]
- Elena, S.F.; Carrasco, P.; Daros, J.A.; Sanjuan, R. Mechanisms of genetic robustness in RNA viruses. EMBO Rep. 2006, 7, 168–173. [Google Scholar] [CrossRef] [PubMed]
- Kummerer, B.M. Establishment and Application of Flavivirus Replicons. Adv. Exp. Med. Biol. 2018, 1062, 165–173. [Google Scholar] [CrossRef] [PubMed]
- Xie, X.; Zou, J.; Shan, C.; Yang, Y.; Kum, D.B.; Dallmeier, K.; Neyts, J.; Shi, P.Y. Zika Virus Replicons for Drug Discovery. EBioMedicine 2016, 12, 156–160. [Google Scholar] [CrossRef] [PubMed]
- Li, J.Q.; Deng, C.L.; Gu, D.; Li, X.; Shi, L.; He, J.; Zhang, Q.Y.; Zhang, B.; Ye, H.Q. Development of a replicon cell line-based high throughput antiviral assay for screening inhibitors of Zika virus. Antiviral Res. 2018, 150, 148–154. [Google Scholar] [CrossRef] [PubMed]
- Lee, I.; Bos, S.; Li, G.; Wang, S.; Gadea, G.; Despres, P.; Zhao, R.Y. Probing Molecular Insights into Zika Virus−Host Interactions. Viruses 2018, 10, 233. [Google Scholar] [CrossRef] [PubMed]
- Wells, M.F.; Salick, M.R.; Wiskow, O.; Ho, D.J.; Worringer, K.A.; Ihry, R.J.; Kommineni, S.; Bilican, B.; Klim, J.R.; Hill, E.J.; et al. Genetic Ablation of AXL Does Not Protect Human Neural Progenitor Cells and Cerebral Organoids from Zika Virus Infection. Cell Stem Cell 2016, 19, 703–708. [Google Scholar] [CrossRef] [PubMed]
- Hamel, R.; Dejarnac, O.; Wichit, S.; Ekchariyawat, P.; Neyret, A.; Luplertlop, N.; Perera-Lecoin, M.; Surasombatpattana, P.; Talignani, L.; Thomas, F.; et al. Biology of Zika Virus Infection in Human Skin Cells. J. Virol. 2015, 89, 8880–8896. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.A.; Seong, R.K.; Son, S.W.; Shin, O.S. Insights into ZIKV-Mediated Innate Immune Responses in Human Dermal Fibroblasts and Epidermal Keratinocytes. J. Investig. Dermatol. 2018. [Google Scholar] [CrossRef] [PubMed]
- Sun, X.; Hua, S.; Chen, H.R.; Ouyang, Z.; Einkauf, K.; Tse, S.; Ard, K.; Ciaranello, A.; Yawetz, S.; Sax, P.; et al. Transcriptional Changes during Naturally Acquired Zika Virus Infection Render Dendritic Cells Highly Conducive to Viral Replication. Cell Rep. 2017, 21, 3471–3482. [Google Scholar] [CrossRef] [PubMed]
- Bowen, J.R.; Zimmerman, M.G.; Suthar, M.S. Taking the defensive: Immune control of Zika virus infection. Virus Res. 2018, 254, 21–26. [Google Scholar] [CrossRef] [PubMed]
- Bowen, J.R.; Quicke, K.M.; Maddur, M.S.; O’Neal, J.T.; McDonald, C.E.; Fedorova, N.B.; Puri, V.; Shabman, R.S.; Pulendran, B.; Suthar, M.S. Zika Virus Antagonizes Type I Interferon Responses during Infection of Human Dendritic Cells. PLoS Pathog. 2017, 13, e1006164. [Google Scholar] [CrossRef] [PubMed]
- Lum, F.M.; Lee, D.; Chua, T.K.; Tan, J.J.L.; Lee, C.Y.P.; Liu, X.; Fang, Y.; Lee, B.; Yee, W.X.; Rickett, N.Y.; et al. Zika Virus Infection Preferentially Counterbalances Human Peripheral Monocyte and/or NK Cell Activity. mSphere 2018, 3, e00120-18. [Google Scholar] [CrossRef] [PubMed]
- Carlin, A.F.; Vizcarra, E.A.; Branche, E.; Viramontes, K.M.; Suarez-Amaran, L.; Ley, K.; Heinz, S.; Benner, C.; Shresta, S.; Glass, C.K. Deconvolution of pro- and antiviral genomic responses in Zika virus-infected and bystander macrophages. Proc. Natl. Acad. Sci. USA 2018, 115, E9172–E9181. [Google Scholar] [CrossRef] [PubMed]
- Michlmayr, D.; Andrade, P.; Gonzalez, K.; Balmaseda, A.; Harris, E. CD14+CD16+ monocytes are the main target of Zika virus infection in peripheral blood mononuclear cells in a paediatric study in Nicaragua. Nat. Microbiol. 2017, 2, 1462–1470. [Google Scholar] [CrossRef] [PubMed]
- Foo, S.S.; Chen, W.; Chan, Y.; Bowman, J.W.; Chang, L.C.; Choi, Y.; Yoo, J.S.; Ge, J.; Cheng, G.; Bonnin, A.; et al. Asian Zika virus strains target CD14+ blood monocytes and induce M2-skewed immunosuppression during pregnancy. Nat. Microbiol. 2017, 2, 1558–1570. [Google Scholar] [CrossRef] [PubMed]
- Tang, H.; Hammack, C.; Ogden, S.C.; Wen, Z.; Qian, X.; Li, Y.; Yao, B.; Shin, J.; Zhang, F.; Lee, E.M.; et al. Zika Virus Infects Human Cortical Neural Progenitors and Attenuates Their Growth. Cell Stem Cell 2016, 18, 587–590. [Google Scholar] [CrossRef] [PubMed]
- Qian, X.; Nguyen, H.N.; Song, M.M.; Hadiono, C.; Ogden, S.C.; Hammack, C.; Yao, B.; Hamersky, G.R.; Jacob, F.; Zhong, C.; et al. Brain-Region-Specific Organoids Using Mini-bioreactors for Modeling ZIKV Exposure. Cell 2016, 165, 1238–1254. [Google Scholar] [CrossRef] [PubMed]
- Meertens, L.; Labeau, A.; Dejarnac, O.; Cipriani, S.; Sinigaglia, L.; Bonnet-Madin, L.; Le Charpentier, T.; Hafirassou, M.L.; Zamborlini, A.; Cao-Lormeau, V.M.; et al. Axl Mediates ZIKA Virus Entry in Human Glial Cells and Modulates Innate Immune Responses. Cell Rep. 2017, 18, 324–333. [Google Scholar] [CrossRef] [PubMed]
- Stefanik, M.; Formanova, P.; Bily, T.; Vancova, M.; Eyer, L.; Palus, M.; Salat, J.; Braconi, C.T.; Zanotto, P.M.A.; Gould, E.A.; et al. Characterisation of Zika virus infection in primary human astrocytes. BMC Neurosci. 2018, 19, 5. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Yang, Y.F.; Yang, Y.; Zou, P.; Chen, J.; He, Y.; Shui, S.L.; Cui, Y.R.; Bai, R.; Liang, Y.J.; et al. AXL promotes Zika virus infection in astrocytes by antagonizing type I interferon signalling. Nat. Microbiol. 2018, 3, 302–309. [Google Scholar] [CrossRef] [PubMed]
- Mladinich, M.C.; Schwedes, J.; Mackow, E.R. Zika Virus Persistently Infects and Is Basolaterally Released from Primary Human Brain Microvascular Endothelial Cells. MBio 2017, 8, e00952-17. [Google Scholar] [CrossRef] [PubMed]
- El Costa, H.; Gouilly, J.; Mansuy, J.M.; Chen, Q.; Levy, C.; Cartron, G.; Veas, F.; Al-Daccak, R.; Izopet, J.; Jabrane-Ferrat, N. ZIKA virus reveals broad tissue and cell tropism during the first trimester of pregnancy. Sci. Rep. 2016, 6, 35296. [Google Scholar] [CrossRef] [PubMed]
- Jurado, K.A.; Simoni, M.K.; Tang, Z.; Uraki, R.; Hwang, J.; Householder, S.; Wu, M.; Lindenbach, B.D.; Abrahams, V.M.; Guller, S.; et al. Zika virus productively infects primary human placenta-specific macrophages. JCI Insight 2016, 1. [Google Scholar] [CrossRef] [PubMed]
- Tabata, T.; Petitt, M.; Puerta-Guardo, H.; Michlmayr, D.; Wang, C.; Fang-Hoover, J.; Harris, E.; Pereira, L. Zika Virus Targets Different Primary Human Placental Cells, Suggesting Two Routes for Vertical Transmission. Cell Host Microbe 2016, 20, 155–166. [Google Scholar] [CrossRef] [PubMed]
- Quicke, K.M.; Bowen, J.R.; Johnson, E.L.; McDonald, C.E.; Ma, H.; O’Neal, J.T.; Rajakumar, A.; Wrammert, J.; Rimawi, B.H.; Pulendran, B.; et al. Zika Virus Infects Human Placental Macrophages. Cell Host Microbe 2016, 20, 83–90. [Google Scholar] [CrossRef] [PubMed]
- Bayer, A.; Lennemann, N.J.; Ouyang, Y.; Bramley, J.C.; Morosky, S.; Marques, E.T., Jr.; Cherry, S.; Sadovsky, Y.; Coyne, C.B. Type III Interferons Produced by Human Placental Trophoblasts Confer Protection against Zika Virus Infection. Cell Host Microbe 2016, 19, 705–712. [Google Scholar] [CrossRef] [PubMed]
- Sheridan, M.A.; Yunusov, D.; Balaraman, V.; Alexenko, A.P.; Yabe, S.; Verjovski-Almeida, S.; Schust, D.J.; Franz, A.W.; Sadovsky, Y.; Ezashi, T.; et al. Vulnerability of primitive human placental trophoblast to Zika virus. Proc. Natl. Acad. Sci. USA 2017, 114, e1587–e1596. [Google Scholar] [CrossRef] [PubMed]
- Aagaard, K.M.; Lahon, A.; Suter, M.A.; Arya, R.P.; Seferovic, M.D.; Vogt, M.B.; Hu, M.; Stossi, F.; Mancini, M.A.; Harris, R.A.; et al. Primary Human Placental Trophoblasts are Permissive for Zika Virus (ZIKV) Replication. Sci. Rep. 2017, 7, 41389. [Google Scholar] [CrossRef] [PubMed]
- Fink, S.L.; Vojtech, L.; Wagoner, J.; Slivinski, N.S.J.; Jackson, K.J.; Wang, R.; Khadka, S.; Luthra, P.; Basler, C.F.; Polyak, S.J. The Antiviral Drug Arbidol Inhibits Zika Virus. Sci. Rep. 2018, 8, 8989. [Google Scholar] [CrossRef] [PubMed]
- Siemann, D.N.; Strange, D.P.; Maharaj, P.N.; Shi, P.Y.; Verma, S. Zika Virus Infects Human Sertoli Cells and Modulates the Integrity of the In Vitro Blood-Testis Barrier Model. J. Virol. 2017, 91, e00623-17. [Google Scholar] [CrossRef] [PubMed]
- Strange, D.P.; Green, R.; Siemann, D.N.; Gale, M., Jr.; Verma, S. Immunoprofiles of human Sertoli cells infected with Zika virus reveals unique insights into host-pathogen crosstalk. Sci. Rep. 2018, 8, 8702. [Google Scholar] [CrossRef] [PubMed]
- Kumar, A.; Jovel, J.; Lopez-Orozco, J.; Limonta, D.; Airo, A.M.; Hou, S.; Stryapunina, I.; Fibke, C.; Moore, R.B.; Hobman, T.C. Human Sertoli cells support high levels of Zika virus replication and persistence. Sci. Rep. 2018, 8, 5477. [Google Scholar] [CrossRef] [PubMed]
- Joguet, G.; Mansuy, J.M.; Matusali, G.; Hamdi, S.; Walschaerts, M.; Pavili, L.; Guyomard, S.; Prisant, N.; Lamarre, P.; Dejucq-Rainsford, N.; et al. Effect of acute Zika virus infection on sperm and virus clearance in body fluids: A prospective observational study. Lancet Infect. Dis. 2017, 17, 1200–1208. [Google Scholar] [CrossRef]
- Matusali, G.; Houzet, L.; Satie, A.P.; Mahe, D.; Aubry, F.; Couderc, T.; Frouard, J.; Bourgeau, S.; Bensalah, K.; Lavoue, S.; et al. Zika virus infects human testicular tissue and germ cells. J. Clin. Investig. 2018. [Google Scholar] [CrossRef] [PubMed]
- Robinson, C.L.; Chong, A.C.N.; Ashbrook, A.W.; Jeng, G.; Jin, J.; Chen, H.; Tang, E.I.; Martin, L.A.; Kim, R.S.; Kenyon, R.M.; et al. Male germ cells support long-term propagation of Zika virus. Nat. Commun. 2018, 9, 2090. [Google Scholar] [CrossRef] [PubMed]
- Roach, T.; Alcendor, D.J. Zika virus infection of cellular components of the blood-retinal barriers: Implications for viral associated congenital ocular disease. J. Neuroinflamm. 2017, 14, 43. [Google Scholar] [CrossRef] [PubMed]
- Mosca, J.D.; Pitha, P.M. Transcriptional and posttranscriptional regulation of exogenous human beta interferon gene in simian cells defective in interferon synthesis. Mol. Cell. Biol. 1986, 6, 2279–2283. [Google Scholar] [CrossRef] [PubMed]
- Chan, J.F.; Yip, C.C.; Tsang, J.O.; Tee, K.M.; Cai, J.P.; Chik, K.K.; Zhu, Z.; Chan, C.C.; Choi, G.K.; Sridhar, S.; et al. Differential cell line susceptibility to the emerging Zika virus: Implications for disease pathogenesis, non-vector-borne human transmission and animal reservoirs. Emerg. Microbes Infect. 2016, 5, e93. [Google Scholar] [CrossRef] [PubMed]
- Offerdahl, D.K.; Dorward, D.W.; Hansen, B.T.; Bloom, M.E. Cytoarchitecture of Zika virus infection in human neuroblastoma and Aedes albopictus cell lines. Virology 2017, 501, 54–62. [Google Scholar] [CrossRef] [PubMed]
- Himmelsbach, K.; Hildt, E. Identification of various cell culture models for the study of Zika virus. World J. Virol. 2018, 7, 10–20. [Google Scholar] [CrossRef] [PubMed]
- Barr, K.L.; Anderson, B.D.; Prakoso, D.; Long, M.T. Working with Zika and Usutu Viruses In Vitro. PLoS Negl. Trop. Dis. 2016, 10, e0004931. [Google Scholar] [CrossRef] [PubMed]
- Barrows, N.J.; Campos, R.K.; Powell, S.T.; Prasanth, K.R.; Schott-Lerner, G.; Soto-Acosta, R.; Galarza-Munoz, G.; McGrath, E.L.; Urrabaz-Garza, R.; Gao, J.; et al. A Screen of FDA-Approved Drugs for Inhibitors of Zika Virus Infection. Cell Host Microbe 2016, 20, 259–270. [Google Scholar] [CrossRef] [PubMed]
- Takahashi, K.; Yamanaka, S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 2006, 126, 663–676. [Google Scholar] [CrossRef] [PubMed]
- Papapetrou, E.P. Induced pluripotent stem cells, past and future. Science 2016, 353, 991–992. [Google Scholar] [CrossRef] [PubMed]
- Lanko, K.; Eggermont, K.; Patel, A.; Kaptein, S.; Delang, L.; Verfaillie, C.M.; Neyts, J. Replication of the Zika virus in different iPSC-derived neuronal cells and implications to assess efficacy of antivirals. Antiviral Res. 2017, 145, 82–86. [Google Scholar] [CrossRef] [PubMed]
- Zhou, T.; Tan, L.; Cederquist, G.Y.; Fan, Y.; Hartley, B.J.; Mukherjee, S.; Tomishima, M.; Brennand, K.J.; Zhang, Q.; Schwartz, R.E.; et al. High-Content Screening in hPSC-Neural Progenitors Identifies Drug Candidates that Inhibit Zika Virus Infection in Fetal-like Organoids and Adult Brain. Cell Stem Cell 2017, 21, 274–283.e5. [Google Scholar] [CrossRef] [PubMed]
- Krencik, R.; Weick, J.P.; Liu, Y.; Zhang, Z.J.; Zhang, S.C. Specification of transplantable astroglial subtypes from human pluripotent stem cells. Nat. Biotechnol. 2011, 29, 528–534. [Google Scholar] [CrossRef] [PubMed]
- Shaltouki, A.; Peng, J.; Liu, Q.; Rao, M.S.; Zeng, X. Efficient generation of astrocytes from human pluripotent stem cells in defined conditions. Stem Cells 2013, 31, 941–952. [Google Scholar] [CrossRef] [PubMed]
- Muffat, J.; Li, Y.; Yuan, B.; Mitalipova, M.; Omer, A.; Corcoran, S.; Bakiasi, G.; Tsai, L.H.; Aubourg, P.; Ransohoff, R.M.; et al. Efficient derivation of microglia-like cells from human pluripotent stem cells. Nat. Med. 2016, 22, 1358–1367. [Google Scholar] [CrossRef] [PubMed]
- Lancaster, M.A.; Renner, M.; Martin, C.A.; Wenzel, D.; Bicknell, L.S.; Hurles, M.E.; Homfray, T.; Penninger, J.M.; Jackson, A.P.; Knoblich, J.A. Cerebral organoids model human brain development and microcephaly. Nature 2013, 501, 373–379. [Google Scholar] [CrossRef] [PubMed]
- Garcez, P.P.; Loiola, E.C.; Madeiro da Costa, R.; Higa, L.M.; Trindade, P.; Delvecchio, R.; Nascimento, J.M.; Brindeiro, R.; Tanuri, A.; Rehen, S.K. Zika virus impairs growth in human neurospheres and brain organoids. Science 2016, 352, 816–818. [Google Scholar] [CrossRef] [PubMed]
- Cugola, F.R.; Fernandes, I.R.; Russo, F.B.; Freitas, B.C.; Dias, J.L.; Guimaraes, K.P.; Benazzato, C.; Almeida, N.; Pignatari, G.C.; Romero, S.; et al. The Brazilian Zika virus strain causes birth defects in experimental models. Nature 2016, 534, 267–271. [Google Scholar] [CrossRef] [PubMed]
- Watanabe, M.; Buth, J.E.; Vishlaghi, N.; de la Torre-Ubieta, L.; Taxidis, J.; Khakh, B.S.; Coppola, G.; Pearson, C.A.; Yamauchi, K.; Gong, D.; et al. Self-Organized Cerebral Organoids with Human-Specific Features Predict Effective Drugs to Combat Zika Virus Infection. Cell Rep. 2017, 21, 517–532. [Google Scholar] [CrossRef] [PubMed]
- Pan, T.; Peng, Z.; Tan, L.; Zou, F.; Zhou, N.; Liu, B.; Liang, L.; Chen, C.; Liu, J.; Wu, L.; et al. Non-Steroidal Anti-Inflammatory Drugs (NSAIDs) Potently Inhibit the Replication of Zika Viruses by Inducing the Degradation of AXL. J. Virol. 2018, 92, e01018-18. [Google Scholar] [CrossRef] [PubMed]
- Pascoalino, B.S.; Courtemanche, G.; Cordeiro, M.T.; Gil, L.H.; Freitas-Junior, L. Zika antiviral chemotherapy: Identification of drugs and promising starting points for drug discovery from an FDA-approved library. F1000Research 2016, 5, 2523. [Google Scholar] [CrossRef] [PubMed]
- Goebel, S.; Snyder, B.; Sellati, T.; Saeed, M.; Ptak, R.; Murray, M.; Bostwick, R.; Rayner, J.; Koide, F.; Kalkeri, R. A sensitive virus yield assay for evaluation of Antivirals against Zika Virus. J. Virol. Methods 2016, 238, 13–20. [Google Scholar] [CrossRef] [PubMed]
- Eyer, L.; Nencka, R.; Huvarova, I.; Palus, M.; Joao Alves, M.; Gould, E.A.; De Clercq, E.; Ruzek, D. Nucleoside Inhibitors of Zika Virus. J. Infect. Dis. 2016, 214, 707–711. [Google Scholar] [CrossRef] [PubMed]
- Muller, J.A.; Harms, M.; Schubert, A.; Mayer, B.; Jansen, S.; Herbeuval, J.P.; Michel, D.; Mertens, T.; Vapalahti, O.; Schmidt-Chanasit, J.; et al. Development of a high-throughput colorimetric Zika virus infection assay. Med. Microbiol. Immunol. 2017, 206, 175–185. [Google Scholar] [CrossRef] [PubMed]
- Adcock, R.S.; Chu, Y.K.; Golden, J.E.; Chung, D.H. Evaluation of anti-Zika virus activities of broad-spectrum antivirals and NIH clinical collection compounds using a cell-based, high-throughput screen assay. Antiviral Res. 2017, 138, 47–56. [Google Scholar] [CrossRef] [PubMed]
- Bernatchez, J.A.; Yang, Z.; Coste, M.; Li, J.; Beck, S.; Liu, Y.; Clark, A.E.; Zhu, Z.; Luna, L.A.; Sohl, C.D.; et al. Development and Validation of a Phenotypic High-Content Imaging Assay for Assessing the Antiviral Activity of Small-Molecule Inhibitors Targeting Zika Virus. Antimicrob. Agents Chemother. 2018, 62. [Google Scholar] [CrossRef] [PubMed]
- Micewicz, E.D.; Khachatoorian, R.; French, S.W.; Ruchala, P. Identification of novel small-molecule inhibitors of Zika virus infection. Bioorg. Med. Chem. Lett. 2017. [Google Scholar] [CrossRef] [PubMed]
- Roy, A.; Lim, L.; Srivastava, S.; Lu, Y.; Song, J. Solution conformations of Zika NS2B-NS3pro and its inhibition by natural products from edible plants. PLoS ONE 2017, 12, e0180632. [Google Scholar] [CrossRef] [PubMed]
- Xu, H.T.; Hassounah, S.A.; Colby-Germinario, S.P.; Oliveira, M.; Fogarty, C.; Quan, Y.; Han, Y.; Golubkov, O.; Ibanescu, I.; Brenner, B.; et al. Purification of Zika virus RNA-dependent RNA polymerase and its use to identify small-molecule Zika inhibitors. J. Antimicrob. Chemother. 2017, 72, 727–734. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.; Brecher, M.; Deng, Y.Q.; Zhang, J.; Sakamuru, S.; Liu, B.; Huang, R.; Koetzner, C.A.; Allen, C.A.; Jones, S.A.; et al. Existing drugs as broad-spectrum and potent inhibitors for Zika virus by targeting NS2B-NS3 interaction. Cell Res. 2017, 27, 1046–1064. [Google Scholar] [CrossRef] [PubMed]
- Murgueitio, M.S.; Bermudez, M.; Mortier, J.; Wolber, G. In silico virtual screening approaches for anti-viral drug discovery. Drug Discov. Today Technol. 2012, 9, e219–e225. [Google Scholar] [CrossRef] [PubMed]
- Franca, T.C. Homology modeling: An important tool for the drug discovery. J. Biomol. Struct. Dyn. 2015, 33, 1780–1793. [Google Scholar] [CrossRef] [PubMed]
- Yuan, S.; Chan, J.F.; den-Haan, H.; Chik, K.K.; Zhang, A.J.; Chan, C.C.; Poon, V.K.; Yip, C.C.; Mak, W.W.; Zhu, Z.; et al. Structure-based discovery of clinically approved drugs as Zika virus NS2B-NS3 protease inhibitors that potently inhibit Zika virus infection in vitro and in vivo. Antiviral Res. 2017, 145, 33–43. [Google Scholar] [CrossRef] [PubMed]
- Singh, A.; Jana, N.K. Discovery of potential Zika virus RNA polymerase inhibitors by docking-based virtual screening. Comput. Biol. Chem. 2017, 71, 144–151. [Google Scholar] [CrossRef] [PubMed]
- Airapetian, K.V.; Nikitin, S.S.; Pavlov, E.V. [Electroneuromyography in patients with severe diphtheric polyneuropathy under conditions of artificial ventilation of the lungs]. Anesteziol. Reanimatol. 1996, 21–23. [Google Scholar]
- Feher, M. Consensus scoring for protein-ligand interactions. Drug Discov. Today 2006, 11, 421–428. [Google Scholar] [CrossRef] [PubMed]
- Sirohi, D.; Chen, Z.; Sun, L.; Klose, T.; Pierson, T.C.; Rossmann, M.G.; Kuhn, R.J. The 3.8 A resolution cryo-EM structure of Zika virus. Science 2016, 352, 467–470. [Google Scholar] [CrossRef] [PubMed]
- Lei, J.; Hansen, G.; Nitsche, C.; Klein, C.D.; Zhang, L.; Hilgenfeld, R. Crystal structure of Zika virus NS2B-NS3 protease in complex with a boronate inhibitor. Science 2016, 353, 503–505. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Yang, K.; Wu, C.; Chen, C.; Hu, C.; Buzovetsky, O.; Wang, Z.; Ji, X.; Xiong, Y.; Yang, H. Mechanisms of activation and inhibition of Zika virus NS2B-NS3 protease. Cell Res. 2016, 26, 1260–1263. [Google Scholar] [CrossRef] [PubMed]
- Onawole, A.T.; Sulaiman, K.O.; Adegoke, R.O.; Kolapo, T.U. Identification of potential inhibitors against the Zika virus using consensus scoring. J. Mol. Graph. Model. 2017, 73, 54–61. [Google Scholar] [CrossRef] [PubMed]
- Jayanthi, C.J.; Tosatti, E.; Fasolino, A. Erratum: Self-consistent phonons, thermal properties, and vibrational instability of the copper crystal. Phys. Rev. B Condens. Matter 1985, 31, 7465. [Google Scholar] [CrossRef] [PubMed]
- McInnes, C. Virtual screening strategies in drug discovery. Curr. Opin. Chem. Biol. 2007, 11, 494–502. [Google Scholar] [CrossRef] [PubMed]
- Rohini, K.; Agarwal, P.; Preethi, B.; Shanthi, V.; Ramanathan, K. Exploring the Lead Compounds for Zika Virus NS2B-NS3 Protein: An e-Pharmacophore-Based Approach. Appl. Biochem. Biotechnol. 2018. [Google Scholar] [CrossRef] [PubMed]
- Zitzmann, C.; Kaderali, L. Mathematical Analysis of Viral Replication Dynamics and Antiviral Treatment Strategies: From Basic Models to Age-Based Multi-Scale Modeling. Front. Microbiol. 2018, 9, 1546. [Google Scholar] [CrossRef] [PubMed]
- Best, K.; Guedj, J.; Madelain, V.; de Lamballerie, X.; Lim, S.Y.; Osuna, C.E.; Whitney, J.B.; Perelson, A.S. Zika plasma viral dynamics in nonhuman primates provides insights into early infection and antiviral strategies. Proc. Natl. Acad. Sci. USA 2017, 114, 8847–8852. [Google Scholar] [CrossRef] [PubMed]
- Dick, G.W. Zika virus. II. Pathogenicity and physical properties. Trans. R. Soc. Trop. Med. Hyg. 1952, 46, 521–534. [Google Scholar] [CrossRef]
- Bell, T.M.; Field, E.J.; Narang, H.K. Zika virus infection of the central nervous system of mice. Arch. Gesamte Virusforsch. 1971, 35, 183–193. [Google Scholar] [CrossRef] [PubMed]
- Way, J.H.; Bowen, E.T.; Platt, G.S. Comparative studies of some African arboviruses in cell culture and in mice. J. Gen. Virol. 1976, 30, 123–130. [Google Scholar] [CrossRef] [PubMed]
- Rossi, S.L.; Tesh, R.B.; Azar, S.R.; Muruato, A.E.; Hanley, K.A.; Auguste, A.J.; Langsjoen, R.M.; Paessler, S.; Vasilakis, N.; Weaver, S.C. Characterization of a Novel Murine Model to Study Zika Virus. Am. J. Trop. Med. Hyg. 2016, 94, 1362–1369. [Google Scholar] [CrossRef] [PubMed]
- Lazear, H.M.; Govero, J.; Smith, A.M.; Platt, D.J.; Fernandez, E.; Miner, J.J.; Diamond, M.S. A Mouse Model of Zika Virus Pathogenesis. Cell Host Microbe 2016, 19, 720–730. [Google Scholar] [CrossRef] [PubMed]
- Grant, A.; Ponia, S.S.; Tripathi, S.; Balasubramaniam, V.; Miorin, L.; Sourisseau, M.; Schwarz, M.C.; Sanchez-Seco, M.P.; Evans, M.J.; Best, S.M.; et al. Zika Virus Targets Human STAT2 to Inhibit Type I Interferon Signaling. Cell Host Microbe 2016, 19, 882–890. [Google Scholar] [CrossRef] [PubMed]
- Morrison, T.E.; Diamond, M.S. Animal Models of Zika Virus Infection, Pathogenesis, and Immunity. J. Virol. 2017, 91. [Google Scholar] [CrossRef] [PubMed]
- Larocca, R.A.; Abbink, P.; Peron, J.P.; Zanotto, P.M.; Iampietro, M.J.; Badamchi-Zadeh, A.; Boyd, M.; Ng’ang’a, D.; Kirilova, M.; Nityanandam, R.; et al. Vaccine protection against Zika virus from Brazil. Nature 2016, 536, 474–478. [Google Scholar] [CrossRef] [PubMed]
- Manangeeswaran, M.; Ireland, D.D.; Verthelyi, D. Zika (PRVABC59) Infection Is Associated with T cell Infiltration and Neurodegeneration in CNS of Immunocompetent Neonatal C57Bl/6 Mice. PLoS Pathog. 2016, 12, e1006004. [Google Scholar] [CrossRef] [PubMed]
- Yu, J.; Liu, X.; Ke, C.; Wu, Q.; Lu, W.; Qin, Z.; He, X.; Liu, Y.; Deng, J.; Xu, S.; et al. Effective Suckling C57BL/6, Kunming, and BALB/c Mouse Models with Remarkable Neurological Manifestation for Zika Virus Infection. Viruses 2017, 9, 165. [Google Scholar] [CrossRef] [PubMed]
- Yuan, L.; Huang, X.Y.; Liu, Z.Y.; Zhang, F.; Zhu, X.L.; Yu, J.Y.; Ji, X.; Xu, Y.P.; Li, G.; Li, C.; et al. A single mutation in the prM protein of Zika virus contributes to fetal microcephaly. Science 2017, 358, 933–936. [Google Scholar] [CrossRef] [PubMed]
- Li, S.; Armstrong, N.; Zhao, H.; Hou, W.; Liu, J.; Chen, C.; Wan, J.; Wang, W.; Zhong, C.; Liu, C.; et al. Zika Virus Fatally Infects Wild Type Neonatal Mice and Replicates in Central Nervous System. Viruses 2018, 10, 49. [Google Scholar] [CrossRef] [PubMed]
- Hutchings, P.R.; Varey, A.M.; Cooke, A. Immunological defects in SJL mice. Immunology 1986, 59, 445–450. [Google Scholar] [PubMed]
- Izumi, K.; Mine, K.; Inoue, Y.; Teshima, M.; Ogawa, S.; Kai, Y.; Kurafuji, T.; Hirakawa, K.; Miyakawa, D.; Ikeda, H.; et al. Reduced Tyk2 gene expression in beta-cells due to natural mutation determines susceptibility to virus-induced diabetes. Nat. Commun. 2015, 6, 6748. [Google Scholar] [CrossRef] [PubMed]
- Haller, O.; Kochs, G.; Weber, F. The interferon response circuit: Induction and suppression by pathogenic viruses. Virology 2006, 344, 119–130. [Google Scholar] [CrossRef] [PubMed]
- Dowall, S.D.; Graham, V.A.; Rayner, E.; Atkinson, B.; Hall, G.; Watson, R.J.; Bosworth, A.; Bonney, L.C.; Kitchen, S.; Hewson, R. A Susceptible Mouse Model for Zika Virus Infection. PLoS Negl. Trop. Dis. 2016, 10, e0004658. [Google Scholar] [CrossRef] [PubMed]
- Yockey, L.J.; Varela, L.; Rakib, T.; Khoury-Hanold, W.; Fink, S.L.; Stutz, B.; Szigeti-Buck, K.; Van den Pol, A.; Lindenbach, B.D.; Horvath, T.L.; et al. Vaginal Exposure to Zika Virus during Pregnancy Leads to Fetal Brain Infection. Cell 2016, 166, 1247–1256.e4. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Saucedo-Cuevas, L.; Regla-Nava, J.A.; Chai, G.; Sheets, N.; Tang, W.; Terskikh, A.V.; Shresta, S.; Gleeson, J.G. Zika Virus Infects Neural Progenitors in the Adult Mouse Brain and Alters Proliferation. Cell Stem Cell 2016, 19, 593–598. [Google Scholar] [CrossRef] [PubMed]
- Kawiecki, A.B.; Mayton, E.H.; Dutuze, M.F.; Goupil, B.A.; Langohr, I.M.; Del Piero, F.; Christofferson, R.C. Tissue tropisms, infection kinetics, histologic lesions, and antibody response of the MR766 strain of Zika virus in a murine model. Virol. J. 2017, 14, 82. [Google Scholar] [CrossRef] [PubMed]
- Aliota, M.T.; Caine, E.A.; Walker, E.C.; Larkin, K.E.; Camacho, E.; Osorio, J.E. Characterization of Lethal Zika Virus Infection in AG129 Mice. PLoS Negl. Trop. Dis. 2016, 10, e0004682. [Google Scholar] [CrossRef] [PubMed]
- Miner, J.J.; Cao, B.; Govero, J.; Smith, A.M.; Fernandez, E.; Cabrera, O.H.; Garber, C.; Noll, M.; Klein, R.S.; Noguchi, K.K.; et al. Zika Virus Infection during Pregnancy in Mice Causes Placental Damage and Fetal Demise. Cell 2016, 165, 1081–1091. [Google Scholar] [CrossRef] [PubMed]
- Kamiyama, N.; Soma, R.; Hidano, S.; Watanabe, K.; Umekita, H.; Fukuda, C.; Noguchi, K.; Gendo, Y.; Ozaki, T.; Sonoda, A.; et al. Ribavirin inhibits Zika virus (ZIKV) replication in vitro and suppresses viremia in ZIKV-infected STAT1-deficient mice. Antiviral Res. 2017, 146, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Zhao, H.; Fernandez, E.; Dowd, K.A.; Speer, S.D.; Platt, D.J.; Gorman, M.J.; Govero, J.; Nelson, C.A.; Pierson, T.C.; Diamond, M.S.; et al. Structural Basis of Zika Virus-Specific Antibody Protection. Cell 2016, 166, 1016–1027. [Google Scholar] [CrossRef] [PubMed]
- Smith, D.R.; Hollidge, B.; Daye, S.; Zeng, X.; Blancett, C.; Kuszpit, K.; Bocan, T.; Koehler, J.W.; Coyne, S.; Minogue, T.; et al. Neuropathogenesis of Zika Virus in a Highly Susceptible Immunocompetent Mouse Model after Antibody Blockade of Type I Interferon. PLoS Negl. Trop. Dis. 2017, 11, e0005296. [Google Scholar] [CrossRef] [PubMed]
- Gorman, M.J.; Caine, E.A.; Zaitsev, K.; Begley, M.C.; Weger-Lucarelli, J.; Uccellini, M.B.; Tripathi, S.; Morrison, J.; Yount, B.L.; Dinnon, K.H., 3rd; et al. An Immunocompetent Mouse Model of Zika Virus Infection. Cell Host Microbe 2018, 23, 672–685.e6. [Google Scholar] [CrossRef] [PubMed]
- Chan, J.F.; Zhang, A.J.; Chan, C.C.; Yip, C.C.; Mak, W.W.; Zhu, H.; Poon, V.K.; Tee, K.M.; Zhu, Z.; Cai, J.P.; et al. Zika Virus Infection in Dexamethasone-immunosuppressed Mice Demonstrating Disseminated Infection with Multi-organ Involvement Including Orchitis Effectively Treated by Recombinant Type I Interferons. EBioMedicine 2016, 14, 112–122. [Google Scholar] [CrossRef] [PubMed]
- Zmurko, J.; Marques, R.E.; Schols, D.; Verbeken, E.; Kaptein, S.J.; Neyts, J. The Viral Polymerase Inhibitor 7-Deaza-2’-C-Methyladenosine Is a Potent Inhibitor of In Vitro Zika Virus Replication and Delays Disease Progression in a Robust Mouse Infection Model. PLoS Negl. Trop. Dis. 2016, 10, e0004695. [Google Scholar] [CrossRef] [PubMed]
- Deng, Y.Q.; Zhang, N.N.; Li, C.F.; Tian, M.; Hao, J.N.; Xie, X.P.; Shi, P.Y.; Qin, C.F. Adenosine Analog NITD008 Is a Potent Inhibitor of Zika Virus. Open Forum Infect. Dis. 2016, 3, ofw175. [Google Scholar] [CrossRef] [PubMed]
- Bullard-Feibelman, K.M.; Govero, J.; Zhu, Z.; Salazar, V.; Veselinovic, M.; Diamond, M.S.; Geiss, B.J. The FDA-approved drug sofosbuvir inhibits Zika virus infection. Antiviral Res. 2017, 137, 134–140. [Google Scholar] [CrossRef] [PubMed]
- Julander, J.G.; Siddharthan, V.; Evans, J.; Taylor, R.; Tolbert, K.; Apuli, C.; Stewart, J.; Collins, P.; Gebre, M.; Neilson, S.; et al. Efficacy of the broad-spectrum antiviral compound BCX4430 against Zika virus in cell culture and in a mouse model. Antiviral Res. 2017, 137, 14–22. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.; Liu, Y.; Wang, S.; Sun, J.; Wang, P.; Xin, Q.; Zhang, L.; Xiao, G.; Wang, W. Antiviral activity of peptide inhibitors derived from the protein E stem against Japanese encephalitis and Zika viruses. Antiviral Res. 2017, 141, 140–149. [Google Scholar] [CrossRef] [PubMed]
- Costa, V.V.; Del Sarto, J.L.; Rocha, R.F.; Silva, F.R.; Doria, J.G.; Olmo, I.G.; Marques, R.E.; Queiroz-Junior, C.M.; Foureaux, G.; Araujo, J.M.S.; et al. N-Methyl-d-Aspartate (NMDA) Receptor Blockade Prevents Neuronal Death Induced by Zika Virus Infection. MBio 2017, 8, e00350-17. [Google Scholar] [CrossRef] [PubMed]
- Li, C.; Deng, Y.Q.; Wang, S.; Ma, F.; Aliyari, R.; Huang, X.Y.; Zhang, N.N.; Watanabe, M.; Dong, H.L.; Liu, P.; et al. 25-Hydroxycholesterol Protects Host against Zika Virus Infection and Its Associated Microcephaly in a Mouse Model. Immunity 2017, 46, 446–456. [Google Scholar] [CrossRef] [PubMed]
- Ferreira, A.C.; Zaverucha-do-Valle, C.; Reis, P.A.; Barbosa-Lima, G.; Vieira, Y.R.; Mattos, M.; Silva, P.P.; Sacramento, C.; de Castro Faria Neto, H.C.; Campanati, L.; et al. Sofosbuvir protects Zika virus-infected mice from mortality, preventing short- and long-term sequelae. Sci. Rep. 2017, 7, 9409. [Google Scholar] [CrossRef] [PubMed]
- Shiryaev, S.A.; Farhy, C.; Pinto, A.; Huang, C.T.; Simonetti, N.; Elong Ngono, A.; Dewing, A.; Shresta, S.; Pinkerton, A.B.; Cieplak, P.; et al. Characterization of the Zika virus two-component NS2B-NS3 protease and structure-assisted identification of allosteric small-molecule antagonists. Antiviral Res. 2017, 143, 218–229. [Google Scholar] [CrossRef] [PubMed]
- Pattnaik, A.; Palermo, N.; Sahoo, B.R.; Yuan, Z.; Hu, D.; Annamalai, A.S.; Vu, H.L.X.; Correas, I.; Prathipati, P.K.; Destache, C.J.; et al. Discovery of a non-nucleoside RNA polymerase inhibitor for blocking Zika virus replication through in silico screening. Antiviral Res. 2018, 151, 78–86. [Google Scholar] [CrossRef] [PubMed]
- Govero, J.; Esakky, P.; Scheaffer, S.M.; Fernandez, E.; Drury, A.; Platt, D.J.; Gorman, M.J.; Richner, J.M.; Caine, E.A.; Salazar, V.; et al. Zika virus infection damages the testes in mice. Nature 2016, 540, 438–442. [Google Scholar] [CrossRef] [PubMed]
- Bueno, M.G.; Martinez, N.; Abdalla, L.; Duarte Dos Santos, C.N.; Chame, M. Animals in the Zika Virus Life Cycle: What to Expect from Megadiverse Latin American Countries. PLoS Negl. Trop. Dis. 2016, 10, e0005073. [Google Scholar] [CrossRef] [PubMed]
- McCrae, A.W.; Kirya, B.G. Yellow fever and Zika virus epizootics and enzootics in Uganda. Trans. R. Soc. Trop. Med. Hyg. 1982, 76, 552–562. [Google Scholar] [CrossRef]
- Osuna, C.E.; Lim, S.Y.; Deleage, C.; Griffin, B.D.; Stein, D.; Schroeder, L.T.; Omange, R.W.; Best, K.; Luo, M.; Hraber, P.T.; et al. Zika viral dynamics and shedding in rhesus and cynomolgus macaques. Nat. Med. 2016, 22, 1448–1455. [Google Scholar] [CrossRef] [PubMed]
- Li, X.F.; Dong, H.L.; Huang, X.Y.; Qiu, Y.F.; Wang, H.J.; Deng, Y.Q.; Zhang, N.N.; Ye, Q.; Zhao, H.; Liu, Z.Y.; et al. Characterization of a 2016 Clinical Isolate of Zika Virus in Non-human Primates. EBioMedicine 2016, 12, 170–177. [Google Scholar] [CrossRef] [PubMed]
- Dudley, D.M.; Aliota, M.T.; Mohr, E.L.; Weiler, A.M.; Lehrer-Brey, G.; Weisgrau, K.L.; Mohns, M.S.; Breitbach, M.E.; Rasheed, M.N.; Newman, C.M.; et al. A rhesus macaque model of Asian-lineage Zika virus infection. Nat. Commun. 2016, 7, 12204. [Google Scholar] [CrossRef] [PubMed]
- Aliota, M.T.; Dudley, D.M.; Newman, C.M.; Mohr, E.L.; Gellerup, D.D.; Breitbach, M.E.; Buechler, C.R.; Rasheed, M.N.; Mohns, M.S.; Weiler, A.M.; et al. Heterologous Protection against Asian Zika Virus Challenge in Rhesus Macaques. PLoS Negl. Trop. Dis. 2016, 10, e0005168. [Google Scholar] [CrossRef] [PubMed]
- Koide, F.; Goebel, S.; Snyder, B.; Walters, K.B.; Gast, A.; Hagelin, K.; Kalkeri, R.; Rayner, J. Development of a Zika Virus Infection Model in Cynomolgus Macaques. Front. Microbiol. 2016, 7, 2028. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, S.M.; Antony, K.M.; Dudley, D.M.; Kohn, S.; Simmons, H.A.; Wolfe, B.; Salamat, M.S.; Teixeira, L.B.C.; Wiepz, G.J.; Thoong, T.H.; et al. Highly efficient maternal-fetal Zika virus transmission in pregnant rhesus macaques. PLoS Pathog. 2017, 13, e1006378. [Google Scholar] [CrossRef] [PubMed]
- Rayner, J.O.; Kalkeri, R.; Goebel, S.; Cai, Z.; Green, B.; Lin, S.; Snyder, B.; Hagelin, K.; Walters, K.B.; Koide, F. Comparative Pathogenesis of Asian and African-Lineage Zika Virus in Indian Rhesus Macaque’s and Development of a Non-Human Primate Model Suitable for the Evaluation of New Drugs and Vaccines. Viruses 2018, 10, 229. [Google Scholar] [CrossRef] [PubMed]
- Haddow, A.D.; Nalca, A.; Rossi, F.D.; Miller, L.J.; Wiley, M.R.; Perez-Sautu, U.; Washington, S.C.; Norris, S.L.; Wollen-Roberts, S.E.; Shamblin, J.D.; et al. High Infection Rates for Adult Macaques after Intravaginal or Intrarectal Inoculation with Zika Virus. Emerg. Infect. Dis. 2017, 23, 1274–1281. [Google Scholar] [CrossRef] [PubMed]
- Grigsby, P.L. Animal Models to Study Placental Development and Function throughout Normal and Dysfunctional Human Pregnancy. Semin. Reprod. Med. 2016, 34, 11–16. [Google Scholar] [CrossRef] [PubMed]
- Mysorekar, I.U.; Diamond, M.S. Modeling Zika Virus Infection in Pregnancy. N. Engl. J. Med. 2016, 375, 481–484. [Google Scholar] [CrossRef] [PubMed]
- Adams Waldorf, K.M.; Stencel-Baerenwald, J.E.; Kapur, R.P.; Studholme, C.; Boldenow, E.; Vornhagen, J.; Baldessari, A.; Dighe, M.K.; Thiel, J.; Merillat, S.; et al. Fetal brain lesions after subcutaneous inoculation of Zika virus in a pregnant nonhuman primate. Nat. Med. 2016, 22, 1256–1259. [Google Scholar] [CrossRef] [PubMed]
- Abbink, P.; Larocca, R.A.; De La Barrera, R.A.; Bricault, C.A.; Moseley, E.T.; Boyd, M.; Kirilova, M.; Li, Z.; Ng’ang’a, D.; Nanayakkara, O.; et al. Protective efficacy of multiple vaccine platforms against Zika virus challenge in rhesus monkeys. Science 2016, 353, 1129–1132. [Google Scholar] [CrossRef] [PubMed]
- Dowd, K.A.; Ko, S.Y.; Morabito, K.M.; Yang, E.S.; Pelc, R.S.; DeMaso, C.R.; Castilho, L.R.; Abbink, P.; Boyd, M.; Nityanandam, R.; et al. Rapid development of a DNA vaccine for Zika virus. Science 2016, 354, 237–240. [Google Scholar] [CrossRef] [PubMed]
- Muthumani, K.; Griffin, B.D.; Agarwal, S.; Kudchodkar, S.B.; Reuschel, E.L.; Choi, H.; Kraynyak, K.A.; Duperret, E.K.; Keaton, A.A.; Chung, C.; et al. In vivo protection against ZIKV infection and pathogenesis through passive antibody transfer and active immunisation with a prMEnv DNA vaccine. NPJ Vaccines 2016, 1, 16021. [Google Scholar] [CrossRef] [PubMed]
- Pardi, N.; Hogan, M.J.; Pelc, R.S.; Muramatsu, H.; Andersen, H.; DeMaso, C.R.; Dowd, K.A.; Sutherland, L.L.; Scearce, R.M.; Parks, R.; et al. Zika virus protection by a single low-dose nucleoside-modified mRNA vaccination. Nature 2017, 543, 248–251. [Google Scholar] [CrossRef] [PubMed]
- Kumar, M.; Krause, K.K.; Azouz, F.; Nakano, E.; Nerurkar, V.R. A guinea pig model of Zika virus infection. Virol. J. 2017, 14, 75. [Google Scholar] [CrossRef] [PubMed]
- Deng, Y.Q.; Zhang, N.N.; Li, X.F.; Wang, Y.Q.; Tian, M.; Qiu, Y.F.; Fan, J.W.; Hao, J.N.; Huang, X.Y.; Dong, H.L.; et al. Intranasal infection and contact transmission of Zika virus in guinea pigs. Nat. Commun. 2017, 8, 1648. [Google Scholar] [CrossRef] [PubMed]
- Bierle, C.J.; Fernandez-Alarcon, C.; Hernandez-Alvarado, N.; Zabeli, J.C.; Janus, B.C.; Putri, D.S.; Schleiss, M.R. Assessing Zika virus replication and the development of Zika-specific antibodies after a mid-gestation viral challenge in guinea pigs. PLoS ONE 2017, 12, e0187720. [Google Scholar] [CrossRef] [PubMed]
- Padilla-Carlin, D.J.; McMurray, D.N.; Hickey, A.J. The guinea pig as a model of infectious diseases. Comp. Med. 2008, 58, 324–340. [Google Scholar] [PubMed]
- Griffith, B.P.; McCormick, S.R.; Fong, C.K.; Lavallee, J.T.; Lucia, H.L.; Goff, E. The placenta as a site of cytomegalovirus infection in guinea pigs. J. Virol. 1985, 55, 402–409. [Google Scholar] [PubMed]
- Mess, A. The Guinea pig placenta: Model of placental growth dynamics. Placenta 2007, 28, 812–815. [Google Scholar] [CrossRef] [PubMed]
- Miller, L.J.; Nasar, F.; Schellhase, C.W.; Norris, S.L.; Kimmel, A.E.; Valdez, S.M.; Wollen-Roberts, S.E.; Shamblin, J.D.; Sprague, T.R.; Lugo-Roman, L.A.; et al. Zika Virus Infection in Syrian Golden Hamsters and Strain 13 Guinea Pigs. Am. J. Trop. Med. Hyg. 2018, 98, 864–867. [Google Scholar] [CrossRef] [PubMed]
| ZIKV Isolate | Lineage | Isolation | Cloning Strategy | Plasmid Name | Reporter | Tested | Reference |
|---|---|---|---|---|---|---|---|
| Paraiba_01/2015 | Asian | Brazil, 2015 | Synthetic intron insertion | ZIKV-1 | - | Cell culture | [78] |
| ZIKV-ICD | |||||||
| ZIKV-NS3m | |||||||
| MR766 | African | Uganda, 1947 | Synthetic intron insertion | - | Cell culture | [79] | |
| FSS13025 | Asian | Cambodia, 2010 | Subgenomic fragments | pFLZIKV | RLuc | Cell culture mosquito-mouse model | [80] |
| MR766 | African | Uganda, 1947 | Subgenomic fragments | - | Cell culture mouse model | [82] | |
| H/PF/2013 | Asian | French Polynesia, 2013 | |||||
| SPH2015 | Asian | Brazil, 2015 | |||||
| BeH819015 | Asian | Brazil, 2015 | |||||
| BeH819015 | Asian | Brazil, 2015 | Subgenomic fragments | icDNA BeH819015 | nLuc | Cell culture | [83] |
| GFP | |||||||
| mCherry | |||||||
| MR766NIID | African | Uganda 1947 | Subgenomic fragments | ZIKV-MR766NIID-MC | GFP | Cell culture | [85] |
| ZIKVGFP | |||||||
| PF | Asian | French Polynesia, 2013 | Subgenomic fragments | PF | - | Cell culture | [86] |
| DAK | African | Dakar, 1984 | DAK | ||||
| (MART) | Asian | Martinique, 2015 | PF/DAK | ||||
| DAK/PF | |||||||
| PF/MART | |||||||
| MR766M | African | Uganda, 1947 | Subgenomic fragments | MR766MC | - | Cell culture | [87] |
| BeH819015 | Asian | Brazil, 2015 | BR15MC | ||||
| CHIM | |||||||
| ZIKVNatal | Asian | Natal, 2015 | Subgenomic fragments | Cell culture | [89] | ||
| PRVABC59 | Asian | Puerto Rico, 2015 | Subgenomic fragments | Cell culture mosquito-mouse model | [90] | ||
| MR-766 | African | Uganda, 1947 | Subgenomic fragments | pBac/MR-766, | - | Cell culture | [81] |
| P6-740 | Asian | Malaysia, 1966 | pBac/P6-740 | ||||
| PRVABC-59 | Asian | Puerto Rico, 2015 | pBac/PRVABC-59 | ||||
| MR766 | African | Uganda 1947 | Mutational inactivation CEPs | synZIKV-MR766 | RLuc | Cell culture | [91] |
| H/PF/2013 | Asian | French Polynesia, 2013 | synZIKV-H/PF/2013 | FP635 |
| Primary Cell Type | Tissue/Source | Reference |
|---|---|---|
| Dermal fibroblasts | Skin | [99,100] |
| Epidermal keratinocytes | Skin | [99] |
| Blood dendritic cells | Peripheral blood | [101] |
| Monocyte-derived dendritic cells | Peripheral blood | [73,99,102,103] |
| Monocyte-derived macrophages | Peripheral blood | [104,105] |
| Monocytes | Peripheral blood | [104,106,107] |
| NPCs | Brain/PSCs | [108,109] |
| Astrocytes | Brain | [66,110,111,112] |
| Microglia | Brain | [110] |
| Endothelial cells | Brain | [113] |
| Hofbauer cells | Placenta | [114,115,116,117] |
| Trophoblasts | Placenta | [114,116,118,119,120] |
| Fibroblasts | Placenta | [116] |
| Endothelial cells | Placenta | [116] |
| Fibroblasts | Uterus | [34,114] |
| Mesenchymal stem cells | Umbilical cord | [114] |
| Epithelial cells | Vagina and cervix | [121] |
| Sertoli cells | Testis | [122,123,124] |
| Spermatozoa | Testis | [125] |
| Germ cells | Testis | [126,127] |
| Retinal endothelial cells | Eye | [128] |
| Retinal pericytes | Eye | [128] |
| Retinal pigmented cells | Eye | [128] |
| Model | Strain | Deficiency | ZIKV Inoculation | Pathogenesis | Reference |
|---|---|---|---|---|---|
| Immunocompetent | Balb/c | ZIKV2015 (Brazil, 2015) | ZIKV replication | [179] | |
| PRVABC59 (Puerto Rico, 2015) | Viremia | ||||
| Lethality: No | |||||
| SJL | Immunological defects | ZIKV2015 (Brazil, 2015) | ZIKV replication | [179] | |
| Viremia | |||||
| Lethality: No | |||||
| ZIKVBR (Brazil, 2015) | Whole-body growth delay or intra-uterine growth restriction (IUGR) in pups | [144] | |||
| Neonatal | C57BL/6 | PRVABC59 (Puerto Rico, 2015) | Neurological symptoms | [180] | |
| C57BL/6 | MR766 (Uganda, 1947) | Neurological symptoms | [183] | ||
| Kunming | PRVABC59 (Puerto Rico, 2015) | Lethality: Yes (age and dose-dependent) | |||
| ICR | SZ-WIV01 (China, 2016) | ||||
| Balb/c | Z16006 (China, 2016) | Neurological symptoms | [181] | ||
| C57BL/6 | Lethality: Yes/No (mouse strain-specific differences) | ||||
| Kunming | |||||
| Balb/c | MRS_OPY_Martinique_PaRi_2015 | Neurological symptoms | [182] | ||
| (Martinique, 2015) | Lethality: Yes (viral strain-specific differences) | ||||
| GZ01 (Venezuela, 2016) | |||||
| SZ01 (Samoa, 2016) | |||||
| FSS13025 (Cambodia, 2010) | |||||
| C57Bl/6 | Dakar 41519 (Senegal, 1984) | Lethality: Yes (partly) | [176] | ||
| Partially immunocompetent | hSTAT2 KI Mice | hSTAT2 under control mStat2 promotor | Mouse adapted ZIKV-Dak-41525 (=ZIKV-Dak-MA) (Senegal, 1984) | Placental transmission | [196] |
| (C57BL/6) | Lethality: Yes (partially) | ||||
| Genetically Immunocompromised | A192 (129sV) | Ifnar1−/− | MP1751 (Uganda, 1962) | Sever disease | [187] |
| Lethality: Yes | |||||
| FSS13025 (Cambodia, 2010) | Signs of illness | [175] | |||
| Severe disease (age dependent) | |||||
| Lethality: Yes (age dependent) | |||||
| IFNAR−/− | Ifnar−/− | MR 766 (Uganda, 1947) | Severe disease | [176] | |
| (C57BL/6) | Dakar 41519, 41667, 4167 (Senegal, 1984) | Neurological symptoms | |||
| H/PF/2013 (FP, 2013) | Lethality: Yes (age dependent) | ||||
| PRVABC59 (Puerto Rico, 2015) | Severe disease | [180] | |||
| Lethality: Yes | |||||
| FSS13025 (Cambodia, 2010) | Severe disease | [188] | |||
| Lethality: Yes | |||||
| MR 766 (Uganda, 1947 | Severe disease | [70] | |||
| DAKAR 41519 (Senegal, 1984) | Neurological symptoms | ||||
| P6-740 (Malaysia, 1966) | Lethality: Yes | ||||
| FSS13025 (Cambodia, 2010) | (virus strain-specific differences in morbidity and lethality) | ||||
| PRVABC59 (Puerto Rico, 2015) | |||||
| Irf3−/−, Irf5−/−, Irf7−/− TKO | Irf3−/−, Irf5−/−, Irf7−/− | MR 766 (Uganda, 1947), | Severe Disease | [176] | |
| (C57BL/6) | H/PF/2013 (French Polynesia, 2013) | Neurological symptoms | |||
| Lethality: Yes | |||||
| FSS13025 (Cambodia, 2010) | Signs of disease | [189] | |||
| Neurological symptoms | |||||
| Irf3−/−, Irf7−/− DKO | Irf3−/−, Irf7−/− | MR766 (Uganda, 1947) | Viremia | [190] | |
| (C57BL/6) | Lethality: Infrequent | ||||
| AG129 (129/Sv) | Ifnar−/−, Ifngr1−/− | FSS13025 (Cambodia, 2010) | Severe disease | [175] | |
| Neurological symptoms | |||||
| Lethality: Yes | |||||
| H/PF/2013 (French Polynesia, 2013) | Severe disease | [191] | |||
| Lethality: Yes | |||||
| Stat2−/− (C57BL/6) | Stat2−/− | MR 766 (Uganda, 1947) | Severe disease | [70] | |
| DAKAR 41519 (Senegal, 1984) | Neurological symptoms | ||||
| P6-740 (Malaysia, 1966) | Lethality: Yes | ||||
| FSS13025 (Cambodia, 2010) | (virus strain-specific differences in morbidity and lethality) | ||||
| PRVABC59 (Puerto Rico, 2015) | |||||
| Stat1−/− | Stat1−/− | MR 766 (Uganda, 1947) | Viremia | [193] | |
| Disease development | |||||
| Lethality: Yes | |||||
| Chemically immunocompromised | C57Bl/6 | IFNAR1-blocking monoclonal antibody (MAb-5A3 | H/PF/2013 (French Polynesia, 2013) | Viremia/ Increased replication | [176] |
| Lethality: No | |||||
| DAK AR D 41525 (Senegal, 1984) | Viremia | [195] | |||
| Severe disease | |||||
| Lethality: Yes (differences depending on inoculation route) | |||||
| Mouse adapted ZIKV-Dak-41525 (=ZIKV-Dak-MA) (Senegal, 1984) | Viremia | [194] | |||
| Severe Disease | |||||
| Lethality: Yes | |||||
| ZIKV-DAK-41525 (Senegal, 1984) | Lethality: Yes | [196] | |||
| ZIKV-DAK-MA (mouse adapted) | |||||
| C57BL/6 | H/PF/2013 (French Polynesia, 2013) | Viremia in testis and epididymis | [208] | ||
| Mouse adapted ZIKV-Dak-41525 (=ZIKV-Dak-MA) (Senegal, 1984) | |||||
| Balb/c | Dexamethasone | PRVABC59 (Puerto Rico, 2015) | Viral dissemination | [197] | |
| Severe disease (after withdrawal) | |||||
| Lethality: Yes (after withdrawal) |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Alves, M.P.; Vielle, N.J.; Thiel, V.; Pfaender, S. Research Models and Tools for the Identification of Antivirals and Therapeutics against Zika Virus Infection. Viruses 2018, 10, 593. https://doi.org/10.3390/v10110593
Alves MP, Vielle NJ, Thiel V, Pfaender S. Research Models and Tools for the Identification of Antivirals and Therapeutics against Zika Virus Infection. Viruses. 2018; 10(11):593. https://doi.org/10.3390/v10110593
Chicago/Turabian StyleAlves, Marco P., Nathalie J. Vielle, Volker Thiel, and Stephanie Pfaender. 2018. "Research Models and Tools for the Identification of Antivirals and Therapeutics against Zika Virus Infection" Viruses 10, no. 11: 593. https://doi.org/10.3390/v10110593
APA StyleAlves, M. P., Vielle, N. J., Thiel, V., & Pfaender, S. (2018). Research Models and Tools for the Identification of Antivirals and Therapeutics against Zika Virus Infection. Viruses, 10(11), 593. https://doi.org/10.3390/v10110593





