Estimation of Forest Above-Ground Biomass by Geographically Weighted Regression and Machine Learning with Sentinel Imagery
Abstract
1. Introduction
2. Materials and Methods
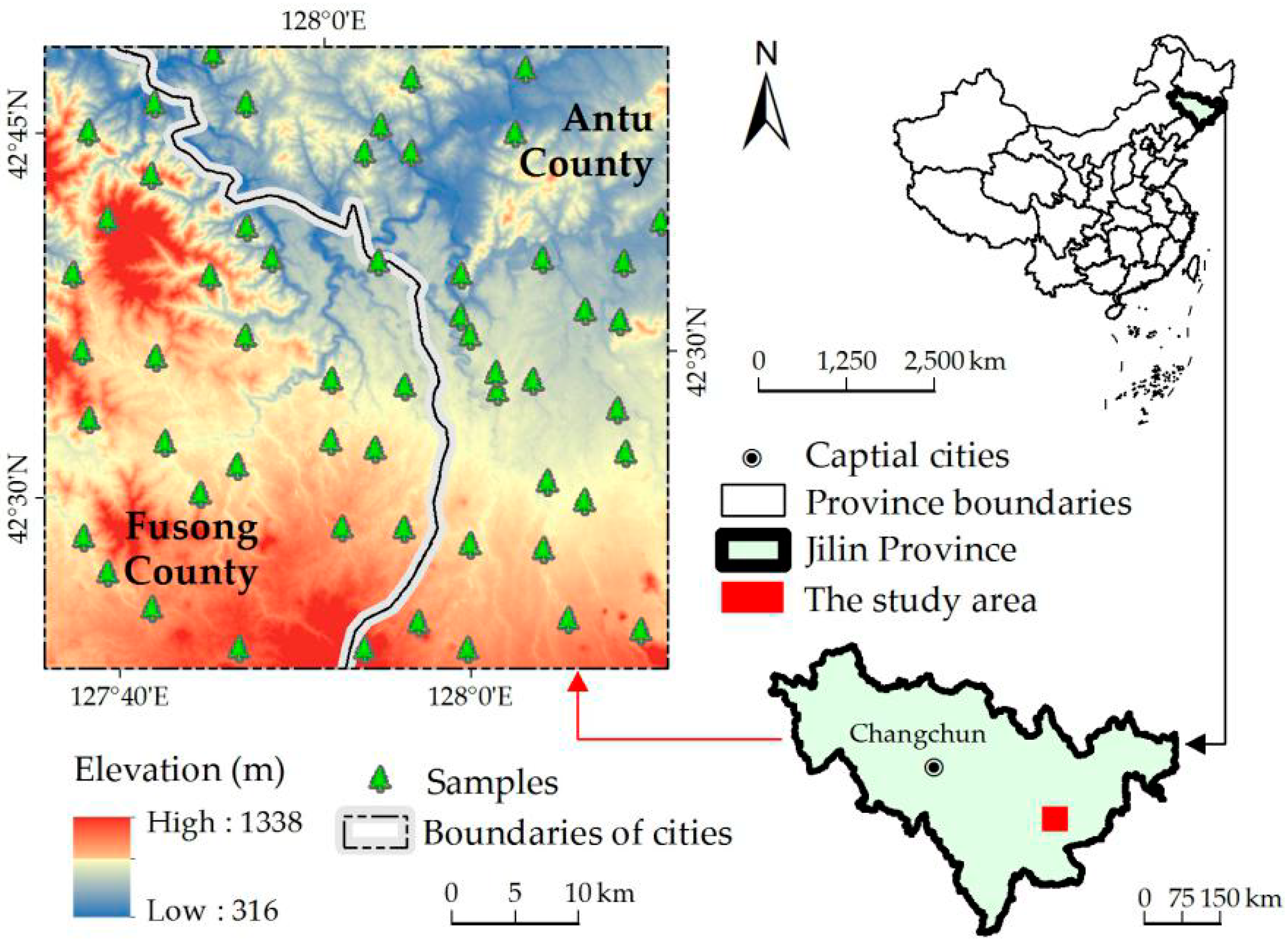
2.1. Study Site
2.2. Field Observations
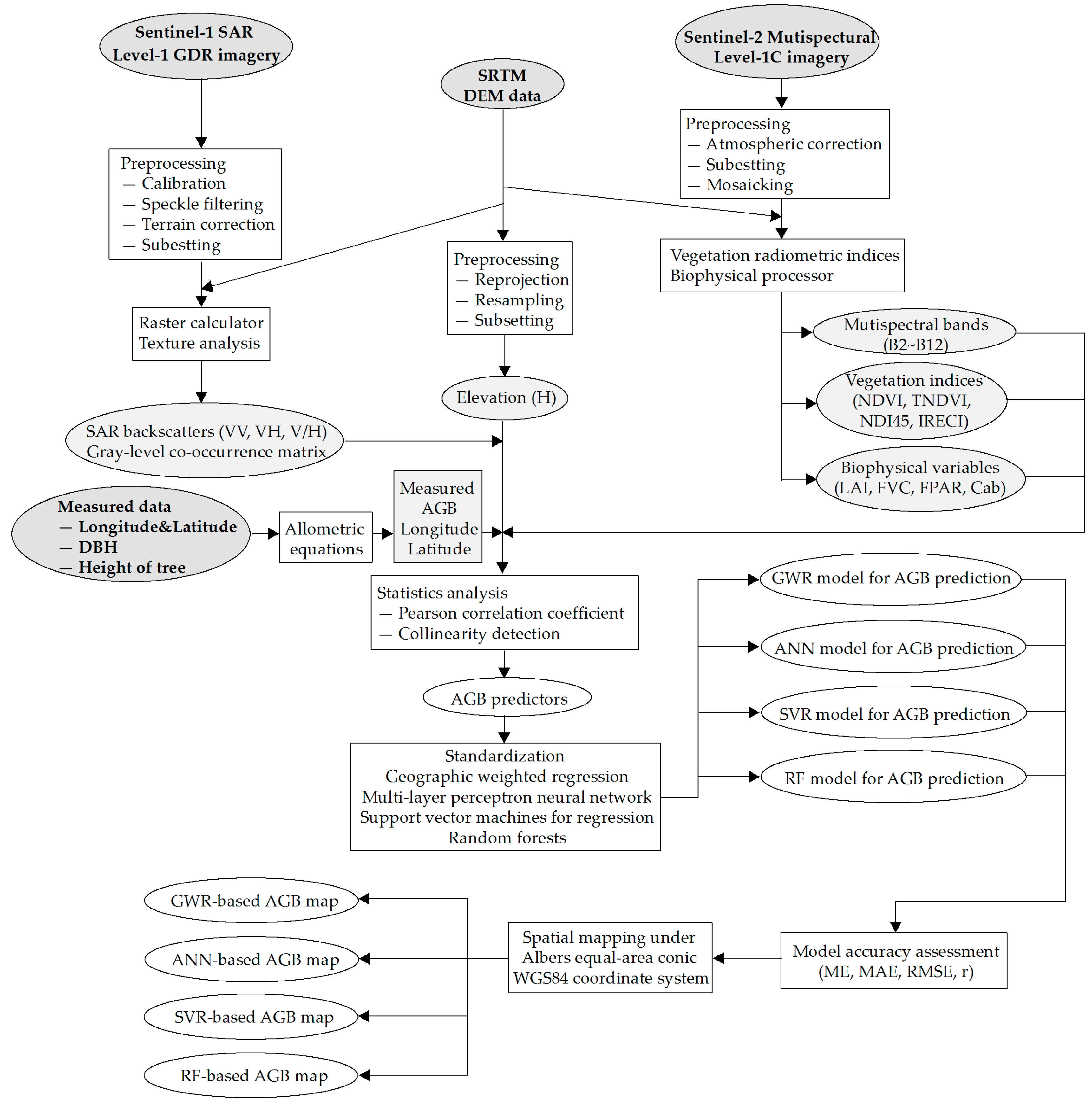
2.3. Satellite Data Pre-Processing and Predictors Derived
3. Modeling the Relationship between Field AGB and Satellite Data
3.1. Geographically Weighted Regression
3.2. Machine Learning Methods
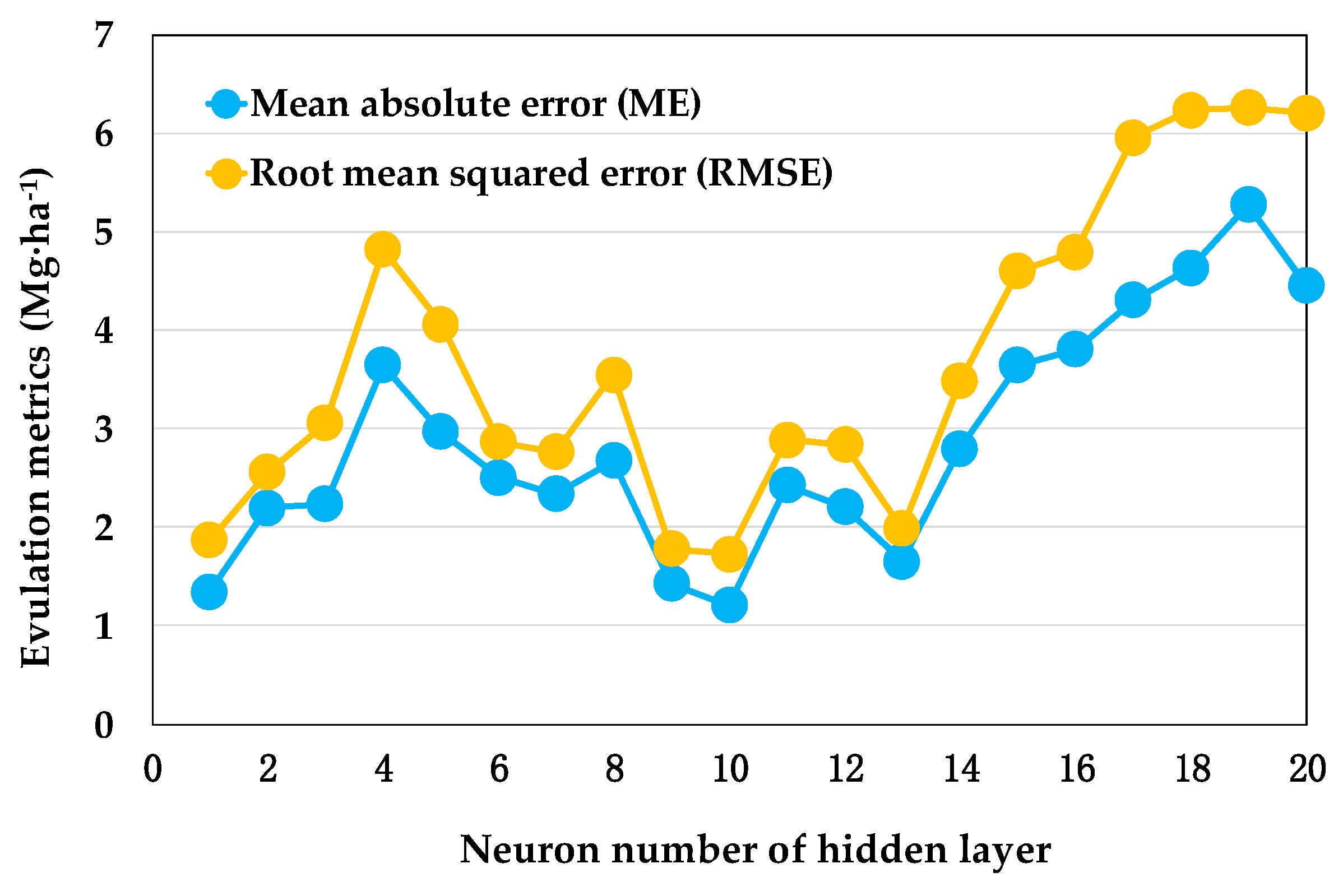
3.2.1. Multi-Layer Perception Neural Network
3.2.2. Support Vector Machines for Regression
3.2.3. Random Forests
3.3. Evaluation of ABG Models
4. Results
4.1. Statistics Analysis
4.2. Models of GWR and ML
4.3. Models Evaluation and Mapping of AGB
4.3.1. Models Assessment by Evaluation Indices
4.3.2. Mapping of Four AGB Models
5. Discussion
5.1. Sentinel-Derived Predictors
5.2. The Comparison of Models
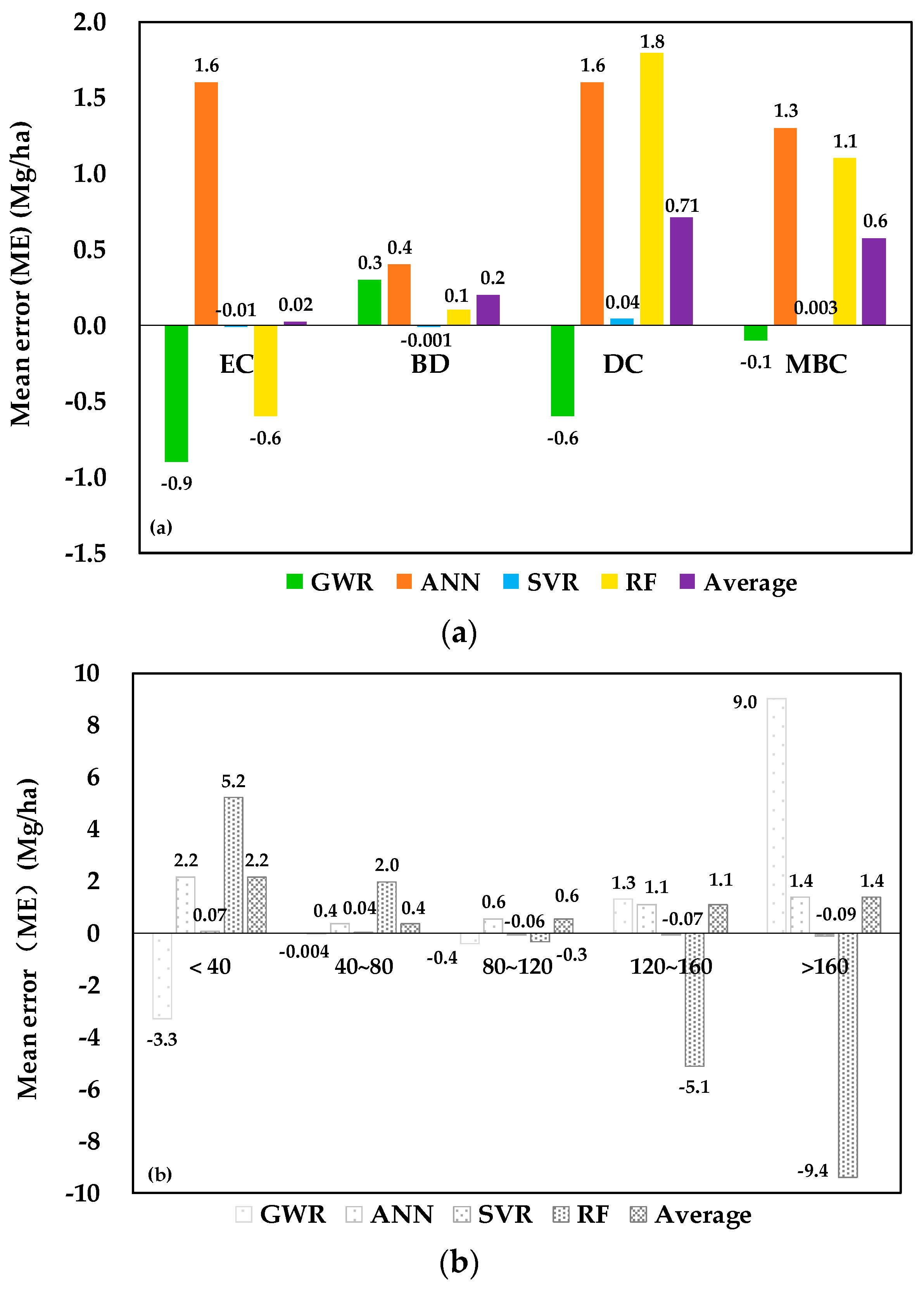
5.3. Model Evaluation by Forest Types and Measured AGB
6. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Olson, J.S.; Watts, J.; Allison, L.J. Carbon in Live Vegetation of Major World Ecosystems; Oak Ridge National Laboratory: Oak Ridge, TN, USA, 1983.
- Liu, K.; Wang, J.D.; Zeng, W.S.; Song, J.L. Comparison and evaluation of three methods for estimating forest above ground biomass using TM and GLAS data. Remote Sens. 2017, 9, 341. [Google Scholar] [CrossRef]
- Cairns, M.A.; Brown, S.; Helmer, E.H.; Baumgardner, G.A. Root biomass allocation in the world’s upland forests. Oecologia 1997, 111, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Brown, S.; Schroeder, P.; Birdsey, R. Aboveground biomass distribution of US eastern hardwood forests and the use of large trees as an indicator of forest development. For. Ecol. Manag. 1997, 96, 37–47. [Google Scholar] [CrossRef]
- Deb, D.; Singh, J.P.; Deb, S.; Datta, D.; Ghosh, A.; Chaurasia, R.S. An alternative approach for estimating above ground biomass using Resourcesat-2 satellite data and artificial neural network in Bundelkhand region of India. Environ. Monit. Assess. 2017, 189, 576. [Google Scholar] [CrossRef] [PubMed]
- Kaasalainen, S.; Holopainen, M.; Karjalainen, M.; Vastaranta, M.; Kankare, V.; Karila, K.; Osmanoglu, B. Combining Lidar and Synthetic Aperture Radar data to estimate forest biomass: Status and prospects. Forests 2015, 6, 252–270. [Google Scholar] [CrossRef]
- Pan, Y.; Birdsey, R.A.; Fang, J.; Houghton, R.; Kauppi, P.E.; Kurz, W.A.; Phillips, O.L.; Shvidenko, A.; Lewis, S.L.; Canadell, J.G.; et al. A large and persistent carbon sink in the world’s forests. Science 2011, 333, 988–993. [Google Scholar] [CrossRef] [PubMed]
- Saatchi, S.S.; Harris, N.L.; Brown, S.; Lefsky, M.; Mitchard, E.T.A.; Salas, W.; Zutta, B.R.; Buermann, W.; Lewis, S.L.; Hagen, S.; et al. Benchmark map of forest carbon stocks in tropical regions across three continents. Proc. Natl. Acad. Sci. USA 2011, 108, 9899–9904. [Google Scholar] [CrossRef] [PubMed]
- Deo, R.K.; Russell, M.B.; Domke, G.M.; Andersen, H.E.; Cohen, W.B.; Woodall, C.W. Evaluating site-specific and generic spatial models of aboveground forest biomass based on Landsat time-series and LiDAR strip samples in the Eastern USA. Remote Sens. 2017, 9, 598. [Google Scholar] [CrossRef]
- Ene, L.T.; Naesset, E.; Gobakken, T.; Gregoire, T.G.; Stahl, G.; Holm, S. A simulation approach for accuracy assessment of two-phase post-stratified estimation in large-area LiDAR biomass surveys. Remote Sens. Environ. 2013, 133, 210–224. [Google Scholar] [CrossRef]
- Wu, C.F.; Shen, H.H.; Shen, A.H.; Deng, J.S.; Gan, M.Y.; Zhu, J.X.; Xu, H.W.; Wang, K. Comparison of machine-learning methods for above-ground biomass estimation based on Landsat imagery. J. Appl. Remote Sens. 2016, 10, 035010. [Google Scholar] [CrossRef]
- McRoberts, R.E.; Næsset, E.; Gobakken, T. Inference for lidar-assisted estimation of forest growing stock volume. Remote Sens. Environ. 2013, 128, 268–275. [Google Scholar] [CrossRef]
- Zhao, P.P.; Lu, D.S.; Wang, G.X.; Liu, L.J.; Li, D.Q.; Zhu, J.R.; Yu, S.Q. Forest aboveground biomass estimation in Zhejiang Province using the integration of Landsat TM and ALOS PALSAR data. Int. J. Appl. Earth Obs. 2016, 53, 1–15. [Google Scholar] [CrossRef]
- Kumar, L.; Sinha, P.; Taylor, S.; Alqurashi, A.F. Review of the use of remote sensing for biomass estimation to support renewable energy generation. J. Appl. Remote Sens. 2015, 9, 097696. [Google Scholar] [CrossRef]
- Lin, Y.; West, G. Reflecting conifer phenology using mobile terrestrial LiDAR: A case study of Pinus sylvestris growing under the Mediterranean climate in Perth, Australia. Ecol. Indic. 2016, 70, 1–9. [Google Scholar] [CrossRef]
- Blackard, J.A.; Finco, M.V.; Helmer, E.H.; Holden, G.R.; Hoppus, M.L.; Jacobs, D.M.; Lister, A.J.; Moisen, G.G.; Nelson, M.D.; Riemann, R.; et al. Mapping us forest biomass using nationwide forest inventory data and moderate resolution information. Remote Sens. Environ. 2008, 112, 1658–1677. [Google Scholar] [CrossRef]
- Cao, L.D.; Pan, J.J.; Li, R.J.; Li, J.L.; Li, Z.F. Integrating airborne LiDAR and optical data to estimate forest aboveground biomass in arid and semi-arid regions of China. Remote Sens. 2018, 10, 532. [Google Scholar] [CrossRef]
- Thenkabail, P.S.; Stucky, N.; Griscom, B.W.; Ashton, M.S.; Diels, J.; van der Meer, B.; Enclona, E. Biomass estimations and carbon stock calculations in the oil palm plantations of African derived savannas using IKONOS data. Int. J. Remote Sens. 2004, 25, 5447–5472. [Google Scholar] [CrossRef]
- Sun, G.; Ni, W.; Zhang, Z.; Xiong, C. Forest Aboveground Biomass Mapping using Spaceborne Stereo Imagery Acquired by Chinese ZY-3. In Proceedings of the AGU Fall Meeting, San Francisco, CA, USA, 14–18 December 2015; Volume 12, p. 2089. [Google Scholar]
- Kumar, L.; Mutanga, O. Remote sensing of above-ground biomass. Remote Sens. 2017, 9, 935. [Google Scholar] [CrossRef]
- Avitabile, V.; Baccini, A.; Friedl, M.A.; Schmullius, C. Capabilities and limitations of Landsat and land cover data for aboveground woody biomass estimation of Uganda. Remote Sens. Environ. 2012, 117, 366–380. [Google Scholar] [CrossRef]
- Santi, E.; Paloscia, S.; Pettinato, S.; Fontanelli, G.; Mura, M.; Zolli, C.; Maselli, F.; Chiesi, M.; Bottai, L.; Chirici, G. The potential of multifrequency SAR images for estimating forest biomass in Mediterranean areas. Remote Sens. Environ. 2017, 200, 63–73. [Google Scholar] [CrossRef]
- Laurin, G.V.; Chen, Q.; Lindsell, J.A.; Coomes, D.A.; Del Frate, F.; Guerriero, L.; Pirotti, F.; Valentini, R. Above ground biomass estimation in an African tropical forest with LiDAR and hyperspectral data. ISPRS J. Photogramm. Remote Sens. 2014, 89, 49–58. [Google Scholar] [CrossRef]
- Chi, H.; Sun, G.Q.; Huang, J.L.; Guo, Z.F.; Ni, W.J.; Fu, A.M. National forest aboveground biomass mapping from ICESat/GLAS data and MODIS imagery in China. Remote Sens. 2015, 7, 5534–5564. [Google Scholar] [CrossRef]
- Guo, Q.H.; Liu, J.; Tao, S.L.; Xue, B.L.; Li, L.; Xu, G.C.; Li, W.K.; Wu, F.F.; Li, Y.M.; Chen, L.H.; et al. Perspectives and prospects of LiDAR in forest ecosystem monitoring and modeling. Chin. Sci. Bull. 2014, 59, 459–478. [Google Scholar] [CrossRef]
- Wilkes, P.; Disney, M.; Vicari, M.B.; Calders, K.; Burt, A. Estimating urban above ground biomass with multi-scale LiDAR. Carbon Balance Manag. 2018, 13, 10. [Google Scholar] [CrossRef] [PubMed]
- Malenovsky, Z.; Rott, H.; Cihlar, J.; Schaepman, M.E.; Garcia-Santos, G.; Fernandes, R.; Berger, M. Sentinels for science: Potential of Sentinel-1, -2, and -3 missions for scientific observations of ocean, cryosphere, and land. Remote Sens. Environ. 2012, 120, 91–101. [Google Scholar] [CrossRef]
- Torres, R.; Snoeij, P.; Geudtner, D.; Bibby, D.; Davidson, M.; Attema, E.; Potin, P.; Rommen, B.; Floury, N.; Brown, M.; et al. GMES Sentinel-1 mission. Remote Sens. Environ. 2012, 120, 9–24. [Google Scholar] [CrossRef]
- Chang, J.S.; Shoshany, M. Mediterranean Shrublands Biomass Estimation using Sentinel-1 and Sentinel-2. In Proceedings of the 36th IEEE International Geoscience and Remote Sensing Symposium (IGARSS), Beijing, China, 10–15 July 2016; pp. 5300–5303. [Google Scholar]
- Laurin, G.V.; Puletti, N.; Hawthorne, W.; Liesenberg, V.; Corona, P.; Papale, D.; Chen, Q.; Valentini, R. Discrimination of tropical forest types, dominant species, and mapping of functional guilds by hyperspectral and simulated multispectral Sentinel-2 data. Remote Sens. Environ. 2016, 176, 163–176. [Google Scholar] [CrossRef]
- Liu, Y.A.; Gong, W.S.; Hu, X.Y.; Gong, J.Y. Forest type identification with random forest using Sentinel-1A, Sentinel-2A, multi-temporal Landsat-8 and DEM data. Remote Sens. 2018, 10, 946. [Google Scholar] [CrossRef]
- Tesfamichael, S.G.; Newete, S.W.; Adam, E.; Dubula, B. Field spectroradiometer and simulated multispectral bands for discriminating invasive species from morphologically similar cohabitant plants. GISci. Remote Sens. 2018, 55, 417–436. [Google Scholar] [CrossRef]
- Battude, M.; Al Bitar, A.; Morin, D.; Cros, J.; Huc, M.; Sicre, C.M.; Le Dantec, V.; Demarez, V. Estimating maize biomass and yield over large areas using high spatial and temporal resolution Sentinel-2 like remote sensing data. Remote Sens. 2016, 184, 668–681. [Google Scholar] [CrossRef]
- Su, W.; Hou, N.; Li, Q.; Zhang, M.Z.; Zhao, X.F.; Jiang, K.P. Retrieving leaf area index of corn canopy based on Sentinel-2 remote sensing image. Trans. Chin. Soc. Agric. Mach. 2018, 49, 151–156. [Google Scholar]
- Sanches, I.D.; Feitosa, R.Q.; Diaz, P.M.A.; Soares, M.D.; Luiz, A.J.B.; Schultz, B.; Maurano, L.E.P. Campo verde database: Seeking to improve agricultural remote sensing of tropical areas. IEEE Geosci. Remote Sens. Lett. 2018, 15, 369–373. [Google Scholar] [CrossRef]
- Sibanda, M.; Mutanga, O.; Rouget, M. Examining the potential of Sentinel-2 MSI spectral resolution in quantifying above ground biomass across different fertilizer treatments. ISPRS J. Photogramm. 2015, 110, 55–65. [Google Scholar] [CrossRef]
- Sakowska, K.; Juszczak, R.; Gianelle, D. Remote sensing of grassland biophysical parameters in the context of the Sentinel-2 satellite mission. J. Sens. 2016, 2016, 4612809. [Google Scholar] [CrossRef]
- Ali, I.; Cawkwell, F.; Dwyer, E.; Green, S. Modeling managed grassland biomass estimation by using multitemporal remote sensing data-A machine learning approach. IEEE J. Sel. Top. Appl. Earth Obs. Remote Sens. 2017, 10, 3254–3264. [Google Scholar] [CrossRef]
- Hawrylo, P.; Wezyk, P. Predicting growing stock volume of scots pine stands using Sentinel-2 satellite imagery and airborne image-derived point clouds. Forests 2018, 9, 274. [Google Scholar] [CrossRef]
- Mura, M.; Bottalico, F.; Giannetti, F.; Bertani, R.; Giannini, R.; Mancini, M.; Orlandini, S.; Travaglini, D.; Chirici, G. Exploiting the capabilities of the Sentinel-2 multi spectral instrument for predicting growing stock volume in forest ecosystems. Int. J. Appl. Earth Obs. 2018, 66, 126–134. [Google Scholar] [CrossRef]
- Plank, S. Rapid damage assessment by means of multi-temporal SAR-A comprehensive review and outlook to Sentinel-1. Remote Sens. 2014, 6, 4870–4906. [Google Scholar] [CrossRef]
- Chemura, A.; Mutanga, O.; Dube, T. Separability of coffee leaf rust infection levels with machine learning methods at Sentinel-2 MSI spectral resolutions. Precis. Agric. 2017, 18, 859–881. [Google Scholar] [CrossRef]
- Mallinis, G.; Mitsopoulos, I.; Chrysafi, I. Evaluating and comparing Sentinel 2A and Landsat-8 Operational Land Imager (OLI) spectral indices for estimating fire severity in a Mediterranean pine ecosystem of Greece. GISci. Remote Sens. 2018, 55, 1–18. [Google Scholar] [CrossRef]
- Baccini, A.; Goetz, S.J.; Walker, W.S.; Laporte, N.T.; Sun, M.; Sulla-Menashe, D.; Hackler, J.; Beck, P.S.A.; Dubayah, R.; Friedl, M.A.; et al. Estimated carbon dioxide emissions from tropical deforestation improved by carbon-density maps. Nat. Clim. Chang. 2012, 2, 182–185. [Google Scholar] [CrossRef]
- Lu, D. The potential and challenge of remote sensing–based biomass estimation. Int. J. Remote Sens. 2006, 27, 1297–1328. [Google Scholar] [CrossRef]
- Fassnacht, F.E.; Hartig, F.; Latifi, H.; Berger, C.; Hernandez, J.; Corvalan, P.; Koch, B. Importance of sample size, data type and prediction method for remote sensing-based estimations of aboveground forest biomass. Remote Sens. Environ. 2014, 154, 102–114. [Google Scholar] [CrossRef]
- Fotheringham, A.S.; Brunsdon, C.; Charlton, M.E. Geographically Weighted Regression: The Analysis of Spatially Varying Relationships; Wiley: Chichester, UK, 2002. [Google Scholar]
- Propastin, P. Modifying geographically weighted regression for estimating aboveground biomass in tropical rainforests by multispectral remote sensing data. Int. J. Appl. Earth Obs. 2012, 18, 82–90. [Google Scholar] [CrossRef]
- Gao, Y.K.; Lu, D.S.; Li, G.Y.; Wang, G.X.; Chen, Q.; Liu, L.J.; Li, D.Q. Comparative analysis of modeling algorithms for forest aboveground biomass estimation in a subtropical region. Remote Sens. 2018, 10, 627. [Google Scholar] [CrossRef]
- Gao, Y.; Bian, J.M.; Song, C. Study on the dynamic relation between spring discharge and precipitation in Fusong County, Changbai Mountain, Jilin Province of China. Water Sci. Technol. Water Supply 2016, 16, 428–437. [Google Scholar] [CrossRef]
- Yang, F.; Yao, Z.F.; Sun, J.L.; Zhu, Y.Q.; Wang, Z.M. The landscape pattern changes analysis of Changbai Mountain forest based on RS and GIS—A case study in Fusong and Antu Counties. Syst. Sci. Compr. Stud. Agric. 2010, 26, 431–437. (In Chinese) [Google Scholar]
- Chen, L.; Ren, C.Y.; Zhang, B.; Wang, Z.M.; Liu, M.Y. Quantifying urban land sprawl and its driving forces in Northeast China from 1990 to 2015. Sustainability 2018, 10, 188. [Google Scholar] [CrossRef]
- Li, X.N.; Guo, Q.X.; Wang, X.C.; Zheng, H.F. Allometry of understory tree species in a natural secondary forest in Northeast China. Sci. Silvae Sin. 2010, 46, 22–32. (In Chinese) [Google Scholar]
- Tang, X.G. Estimation of Forest Aboveground Biomass by Integrating ICESat/GLAS Waveform and TM Data. Ph.D. Thesis, University of Chinese Academy of Sciences, Beijing, China, 2013. (In Chinese). [Google Scholar]
- Castillo, J.A.A.; Apan, A.A.; Maraseni, T.N.; Salmo, S.G. Estimation and mapping of above-ground biomass of mangrove forests and their replacement land uses in the Philippines using Sentinel imagery. ISPRS J. Photogramm. Remote Sens. 2017, 134, 70–85. [Google Scholar] [CrossRef]
- Veci, L. Sentinel-1 Toolbox: SAR Basics Tutorial; ARRAY Systems Computing, Inc. and European Space Agency: Paris, France, 2015. [Google Scholar]
- Liu, C. Analysis of Sentinel-1 SAR Data for Mapping Standing Water in the Twente Region. Master’s Thesis, University of Twente, Enschede, The Netherlands, 2016. [Google Scholar]
- Pan, L.; Sun, Y.J. Estimation of Cunninghamia lanceolata forest biomass based on Sentinel-1 image texture information. J. Northeast For. Univ. 2018, 46, 58–62. (In Chinese) [Google Scholar]
- Jacquemoud, S.; Verhoef, W.; Baret, F.; Bacour, C.; Zarco-Tejada, P.J.; Asner, G.P.; François, C.; Ustin, S.L. PROSPECT + SAIL models: A review of use for vegetation characterization. Remote Sens. Environ. 2009, 113, S56–S66. [Google Scholar] [CrossRef]
- Haralick, R.M.; Shanmugam, K.; Denstien, I. Textural features for image classification. IEEE Trans. Syst. Man Cybern. 1973, 3, 610–621. [Google Scholar] [CrossRef]
- Rouse, J.W.; Haas, R.H.; Schell, J.A.; Deering, D.W. Monitoring Vegetation Systems in the Great Plains with ERTS. In Proceedings of the Third Earth Resources Technology Satellite-1 Symposium, Washington, DC, USA, 10–14 December 1973; Volume 1, pp. 309–317. [Google Scholar]
- Delegido, J.; Verrelst, J.; Alonso, L.; Moreno, J. Evaluation of Sentinel-2 red-edge bands for empirical estimation of green LAI and chlorophyll content. Sensors 2011, 11, 7063–7081. [Google Scholar] [CrossRef] [PubMed]
- Clevers, J.G.P.W.; De Jong, S.M.; Epema, G.F.; Van der Meer, F.D.; Bakker, W.H.; Skidmore, A.K.; Scholte, K.H. Derivation of the red edge index using the MERIS standard band setting. Int. J. Remote Sens. 2002, 23, 3169–3184. [Google Scholar] [CrossRef]
- Deering, D.W.; Rouse, J.W.; Haas, R.H.; Schell, J.A. Measuring Forage Production of Grazing Units from Landsat MSS data. In Proceedings of the Tenth International Symposium on Remote Sensing of Environment, Ann Arbor, MI, USA, 6 October 1975; Volume 2, pp. 1169–1178. [Google Scholar]
- O’brien, R.M. A caution regarding rules of thumb for variance inflation factors. Qual. Quant. 2007, 41, 673–690. [Google Scholar] [CrossRef]
- Were, K.; Bui, D.T.; Dick, Ø.B.; Singh, B.R. A comparative assessment of support vector regression, artificial neural networks, and random forests for predicting and mapping soil organic carbon stocks across an Afromontane landscape. Ecol. Indic. 2015, 52, 394–403. [Google Scholar] [CrossRef]
- Cheadle, C.; Vawter, M.P.; Freed, W.J.; Becker, K.G. Analysis of microarray data using Z score transformation. J. Mol. Diagn. 2003, 5, 73–81. [Google Scholar] [CrossRef]
- Brunsdon, C.; Fotheringham, S.; Charlton, M. Geographically Weighted Regression–Modelling Spatial Non-stationarity. In Workshop on Local Indicators of Spatial Association; University of Leicester: Leicester, UK, 1998; Volume 47, pp. 431–443. [Google Scholar]
- Kumar, S.; Lal, R.; Liu, D. A geographically weighted regression kriging approach for mapping soil organic carbon stock. Geoderma 2012, 189, 627–634. [Google Scholar] [CrossRef]
- Shin, J.; Temesgen, H.; Strunk, J.L.; Hilker, T. Comparing modeling methods for predicting forest attributes using LiDAR metrics and ground measurements. Can. J. Remote Sens. 2016, 42, 739–765. [Google Scholar] [CrossRef]
- Zhang, C.S.; Tang, Y.; Xu, X.L.; Kiely, G. Towards spatial geochemical modelling: Use of geographically weighted regression for mapping soil organic carbon contents in Ireland. Appl. Geochem. 2011, 26, 1239–1248. [Google Scholar] [CrossRef]
- Peter, M. Efficient statistical classification of satellite measurements. Int. J. Remote Sens. 2011, 32, 6109–6132. [Google Scholar]
- Ahmed, M.A.A.; Abd-Elrahman, A.; Escobedo, F.J.; Cropper, W.P.; Martin, T.A.; Timilsina, N. Spatially-explicit modeling of multi-scale drivers of aboveground forest biomass and water yield in watersheds of the Southeastern United States. J. Environ. Manag. 2017, 199, 158–171. [Google Scholar] [CrossRef] [PubMed]
- Nakaya, T.; Charlton, M.; Lewis, P.; Brunsdon, C.; Yao, J.; Fotheringham, S. GWR4 User Manual, Windows Application for Geographically Weighted Regression Modelling; Ritsumeikan University: Kyoto, Japan, 2014. [Google Scholar]
- Haykin, S.S. Neural Networks: A Comprehensive Foundation; Tsinghua University Press: Beijing, China, 2001. [Google Scholar]
- Zhu, Y.H.; Liu, K.; Liu, L.; Wang, S.G.; Liu, H.X. Retrieval of mangrove aboveground biomass at the individual species level with WorldView-2 images. Remote Sens. 2015, 7, 12192–12214. [Google Scholar] [CrossRef]
- Hornik, K. Multilayer feed forward network are universal approximators. Neural Netw. 1989, 2, 359–366. [Google Scholar] [CrossRef]
- Santi, E.; Paloscia, S.; Pettinato, S.; Chirici, G.; Mura, M.; Maselli, F. Application of Neural Networks for the retrieval of forest woody volume from SAR multifrequency data at L and C bands. Eur. J. Remote Sens. 2015, 48, 673–687. [Google Scholar] [CrossRef]
- Lee, S.; Evangelista, D.G. Earthquake-induced landslide susceptibility mapping using an artificial neural network. Nat. Hazards Earth Syst. Sci. 2006, 6, 687–695. [Google Scholar] [CrossRef]
- Ottoy, S.; De Vos, B.; Sindayihebura, A.; Hermy, M.; Van Orshoven, J. Assessing soil organic carbon stocks under current and potential forest cover using digital soil mapping and spatial generalisation. Ecol. Indic. 2017, 77, 139–150. [Google Scholar] [CrossRef]
- Vapnik, V.N. The Nature of Statistical Learning Theory. Statistics for Engineering and Information Science; Springer: New York, NY, USA, 2000. [Google Scholar]
- Li, M.; Im, J.; Quackenbush, L.J.; Liu, T. Forest biomass and carbon stock quantification using airborne LiDAR data: A case study over Huntington wildlife forest in the Adirondack Park. IEEE J. Sel. Top. Appl. Earth Obs. Remote Sens. 2014, 7, 3143–3156. [Google Scholar] [CrossRef]
- Meng, S.L.; Pang, Y.; Zhang, Z.J.; Jia, W.; Li, Z.Y. Mapping aboveground biomass using texture indices from aerial photos in a temperate forest of Northeastern China. Remote Sens. 2015, 7, 12192–12214. [Google Scholar] [CrossRef]
- Sharifi, A.; Amini, J.; Tateishi, R. Estimation of forest biomass using multivariate relevance vector regression. Photogramm. Eng. Remote Sens. 2016, 82, 41–49. [Google Scholar] [CrossRef]
- Platt, J. Fast Training of Support Vector Machines Using Sequential Minimal Optimization. In Advances in Kernel Methods–Support Vector Learning; MIT Press: Cambridge, MA, USA, 1999. [Google Scholar]
- Zeng, Z.; Hsieh, W.W.; Shabbar, A.; Burrows, W.R. Seasonal prediction of winter extreme precipitation over Canada by support vector regression. Hydrol. Earth Syst. Sci. 2011, 15, 65–74. [Google Scholar] [CrossRef]
- Singh, M.; Evans, D.; Friess, D.A.; Tan, B.S.; Nin, C.S. Mapping above-ground biomass in a tropical forest in Cambodia using canopy textures derived from Google Earth. Remote Sens. 2015, 7, 5057–5076. [Google Scholar] [CrossRef]
- Rabe, A.; van der Linden, S.; Hostert, P. Simplifying Support Vector Machines for Regression Analysis of Hyperspectral Imagery. In Proceedings of the 1st Workshop on Hyperspectral Image and Signal Processing–Evolution in Remote Sensing, Grenoble, France, 26–28 August 2009. [Google Scholar]
- Breiman, L. Random Forests. Mach. Learn. 2001, 45, 5–32. [Google Scholar] [CrossRef]
- Breiman, L. Bagging predictors. Mach. Learn. 1996, 24, 123–140. [Google Scholar] [CrossRef]
- Amit, Y.; Geman, D. Shape quantization and recognition with randomized trees. Neural Comput. 1997, 9, 1545–1588. [Google Scholar] [CrossRef]
- Dhanda, P.; Nandy, S.; Kushwaha, S.P.S.; Ghosh, S.; Murthy, Y.V.N.K.; Dadhwal, V.K. Optimizing spaceborne LiDAR and very high resolution optical sensor parameters for biomass estimation at ICESat/GLAS footprint level using regression algorithms. Prog. Phys. Geogr. 2017, 41, 247–267. [Google Scholar] [CrossRef]
- Genuer, R.; Poggi, J.-M.; Tuleau-Malot, C.; Villa-Vialaneix, N. Random forests for big data. Big Data Res. 2017, 9, 28–46. [Google Scholar] [CrossRef]
- Jovic, A.; Bogunovic, N. Electrocardiogram analysis using a combination of statistical, geometric, and nonlinear heart rate variability features. Artif. Intell. Med. 2011, 51, 175–186. [Google Scholar] [CrossRef] [PubMed]
- Mutanga, O.; Adam, E.; Cho, M.A. High density biomass estimation for wetland vegetation using WorldView-2 imagery and random forest regression algorithm. Int. J. Appl. Earth. Obs. 2012, 18, 399–406. [Google Scholar] [CrossRef]
- Isaaks, E.H.; Srivastava, R.M. An Introduction to Applied Geostatistics; Oxford University Press: Oxford, UK, 1989. [Google Scholar]
- Wang, D.L.; Xin, X.P.; Shao, Q.Q.; Brolly, M.; Zhu, Z.L.; Chen, J. Modeling aboveground biomass in Hulunber grassland ecosystem by using unmanned aerial vehicle discrete Lidar. Sensors 2017, 17, 180. [Google Scholar] [CrossRef] [PubMed]
- Bourgoin, C.; Blanc, L.; Bailly, J.S.; Cornu, G.; Berenguer, E.; Oszwald, J.; Tritsch, I.; Laurent, F.; Hasan, A.F.; Sist, P.; et al. The potential of multisource remote sensing for mapping the biomass of a degraded Amazonian forest. Forests 2018, 9, 303. [Google Scholar] [CrossRef]
- Ghosh, S.M.; Behera, M.D. Aboveground biomass estimation using multi-sensor data synergy and machine learning algorithms in a dense tropical forest. Appl. Geogr. 2018, 96, 29–40. [Google Scholar] [CrossRef]
- Berninger, A.; Lohberger, S.; Stangel, M.; Siegert, F. SAR-based estimation of above-ground biomass and its changes in tropical forests of Kalimantan using L- and C-band. Remote Sens. 2018, 10, 831. [Google Scholar] [CrossRef]
- Vafaei, S.; Soosani, J.; Adeli, K.; Fadaei, H.; Naghavi, H.; Pham, T.D.; Bui, D.T. Improving accuracy estimation of forest aboveground biomass based on incorporation of ALOS-2 PALSAR-2 and Sentinel-2A imagery and machine learning: A case study of the Hyrcanian forest area (Iran). Remote Sens. 2018, 10, 172. [Google Scholar] [CrossRef]
- Sinha, S.; Jeganathan, C.; Sharma, L.K. A review of radar remote sensing for biomass estimation. Int. J. Environ. Sci. Technol. 2015, 12, 1779–1792. [Google Scholar] [CrossRef]



| Mission | Product | Observation Date | Cell Size (m) | Uniform Resource Identifier (URI) |
|---|---|---|---|---|
| Sentinel-1B | Level-1 GRD-HR | 22 September 2017 | 10 | S1B_IW_GRDH_1SDV_20170922T213003_20170922T213028_007510_00D425_4962.SAFE |
| Sentinel-2A | Multispectral image Level-1C | 3 May 2017 | 10 | S2A_MSIL1C_20170503T021611_N0205_R003_T52TDN_20170503T022350.SAFE |
| Sentinel-2A | Multispectral image Level-1C | 25 July 2017 | 10 | S2A_MSIL1C_20170725T022551_N0205_R046_T52TCM_20170725T023524.SAFE |
| Sentinel-2A | Multispectral image Level-1C | 23 September 2017 | 10 | S2A_MSIL1C_20170923T022551_N0205_R046_T52TCN_20170923T023519.SAFE |
| Sentinel-2A | Multispectral image Level-1C | 23 September 2017 | 10 | S2A_MSIL1C_20170923T022551_N0205_R046_T52TDM_20170923T023519.SAFE |
| Tree Species | Allometric Equations |
|---|---|
| Betula platyphylla Suk. | AGB = T + B + L = 0.04939 × (D2 × H)0.9011 + 0.01417 × (D2 × H)0.7686 + 0.0109 × (D2 × H)0.6472 |
| Acer mono Maxim. | AGB = T + B + L = 0.3274 × (D2 × H)0.7218 + 0.01349 × (D2 × H)0.7198 + 0.02347 × (D2 × H)0.6929 |
| Tilia amurensis Rupr. | AGB = T + B + L = 0.01275 × (D2 × H)1.0094 + 0.00182 × (D2 × H)0.9746 + 0.00024 × (D2 × H)0.9907 |
| Mongolian oak (Quercus spp.) | AGB = T + B + L = 0.03147 × (D2 × H)0.7329 + 0.002127 × D2.9504 + 0.00321 × D2.47349 |
| Ulmus japonica Sarg. | AGB = T + B + L = 0.031457 × (D2 × H)1.032 + 0.007429 × D2.6745 + 0.002754 × D2.4965 |
| Fraxinus mandschurica Rupr | AGB = T + B + L = 1.416 × D1.71 + 1.154 × D1.549 + 0.7655 × D0.886 |
| Populus cathayana Rehd. | AGB = T + B + L = 0.3642 × D2.0043 + 0.0317 × D2.6398 + 0.0149 × D2.2541 |
| Juglans mandshurica Maxim. | AGB = 0.099 × (D2H)0.841 |
| Prunus padus L. | AGB = 0.09 × D2.696 |
| Pinus koraiensis Sieb. et Zucc. | AGB = T + B + L = 0.0144 × (D2 × H)1.0004 + 0.0332 × (D2 × H)0.6941 + 0.0866 × (D2 × H)0.4696 |
| Larix gmelinii var. japonica | AGB = T + B + L = 0.025 × (D2 × H)0.96 + 0.0021 × (D2 × H)0.9638 + 0.00126 × (D2 × H)0.9675 |
| Source Image | Relevant Predictors | Description | |
|---|---|---|---|
| Sentinel-1 | Polarization | VV | vertical transmit-vertical channel |
| VH | vertical transmit-Horizontal channel | ||
| V/H 1 | VV/VH | ||
| Texture 2 | VH_CON, VV_CON | Contrast | |
| VH_DIS, VV_DIS | Dissimilarity | ||
| VH_HOM, VV_HOM | Homogeneity | ||
| VH_ASM, VV_ASM | Angular Second Moment | ||
| VH_ENE, VV_ENE | Energy | ||
| VH_MAX, VV_MAX | Maximum Probability | ||
| VH_ENT, VV_ENT | Entropy | ||
| VH_MEA, VV_MEA | GLCM Mean | ||
| VH_VAR, VV_VAR | GLCM Variance | ||
| VH_COR, VV_COR | GLCM Correlation | ||
| Sentinel-2 | Multispectral bands | B2 | Blue, 490 nm |
| B3 | Green, 560 nm | ||
| B4 | Red, 665 nm | ||
| B5 | Red edge, 705 nm | ||
| B6 | Red edge, 749 nm | ||
| B7 | Red edge, 783 nm | ||
| B8 | Near Infrared, 842 nm | ||
| B8a | Near Infrared, 865 nm | ||
| B11 | Short Wave IR, 1610 nm | ||
| B12 | Short Wave IR, 2190 nm | ||
| Vegetation indices | NDVI 3 | (Band 8 − Band 4)/(Band 8 + Band 4) | |
| NDI45 4 | (Band 5 − Band 4)/(Band 5 + Band 4) | ||
| IRECI 5 | (Band 7 − Band 4)/(Band 5/Band 6) | ||
| TNDVI 6 | [(Band 8 − Band 4)/(Band 8 + Band 4) + 0.5]1/2 | ||
| Vegetation biophysical variables | LAI | Leaf Area Index | |
| FVC | Fraction of Vegetation Cover | ||
| FPAR | Fraction of Absorbed Photosynthetically Active Radiation | ||
| Cab | Chlorophyll content in the leaf | ||
| SRTM DEM | Elevation | H | Elevation, 30 m resolution |
| Evaluation Index | GWR | ANN | SVR | RF |
|---|---|---|---|---|
| ME | 0.04 | 0.84 | 4 × 10−3 | 0.55 |
| MAE | 4.01 | 1.21 | 0.07 | 3.48 |
| RMSE | 5.26 | 1.73 | 0.08 | 4.43 |
| r | 0.995 | 1 | 1 | 0.999 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chen, L.; Ren, C.; Zhang, B.; Wang, Z.; Xi, Y. Estimation of Forest Above-Ground Biomass by Geographically Weighted Regression and Machine Learning with Sentinel Imagery. Forests 2018, 9, 582. https://doi.org/10.3390/f9100582
Chen L, Ren C, Zhang B, Wang Z, Xi Y. Estimation of Forest Above-Ground Biomass by Geographically Weighted Regression and Machine Learning with Sentinel Imagery. Forests. 2018; 9(10):582. https://doi.org/10.3390/f9100582
Chicago/Turabian StyleChen, Lin, Chunying Ren, Bai Zhang, Zongming Wang, and Yanbiao Xi. 2018. "Estimation of Forest Above-Ground Biomass by Geographically Weighted Regression and Machine Learning with Sentinel Imagery" Forests 9, no. 10: 582. https://doi.org/10.3390/f9100582
APA StyleChen, L., Ren, C., Zhang, B., Wang, Z., & Xi, Y. (2018). Estimation of Forest Above-Ground Biomass by Geographically Weighted Regression and Machine Learning with Sentinel Imagery. Forests, 9(10), 582. https://doi.org/10.3390/f9100582








