Analysis of Microbial Diversity and Greenhouse Gas Production of Decaying Pine Logs
Abstract
:1. Introduction
2. Materials and Methods
2.1. Study Area
2.2. Experimental Design and Deadwood Sampling
2.3. GHG Potential Production from Deadwood
2.4. DNA Extraction and PCR-DGGE Analysis of Microbial Communities
2.5. Analysis of Physical and Chemical Properties
2.6. Enzyme Assays
2.7. Statistical Analysis
3. Results
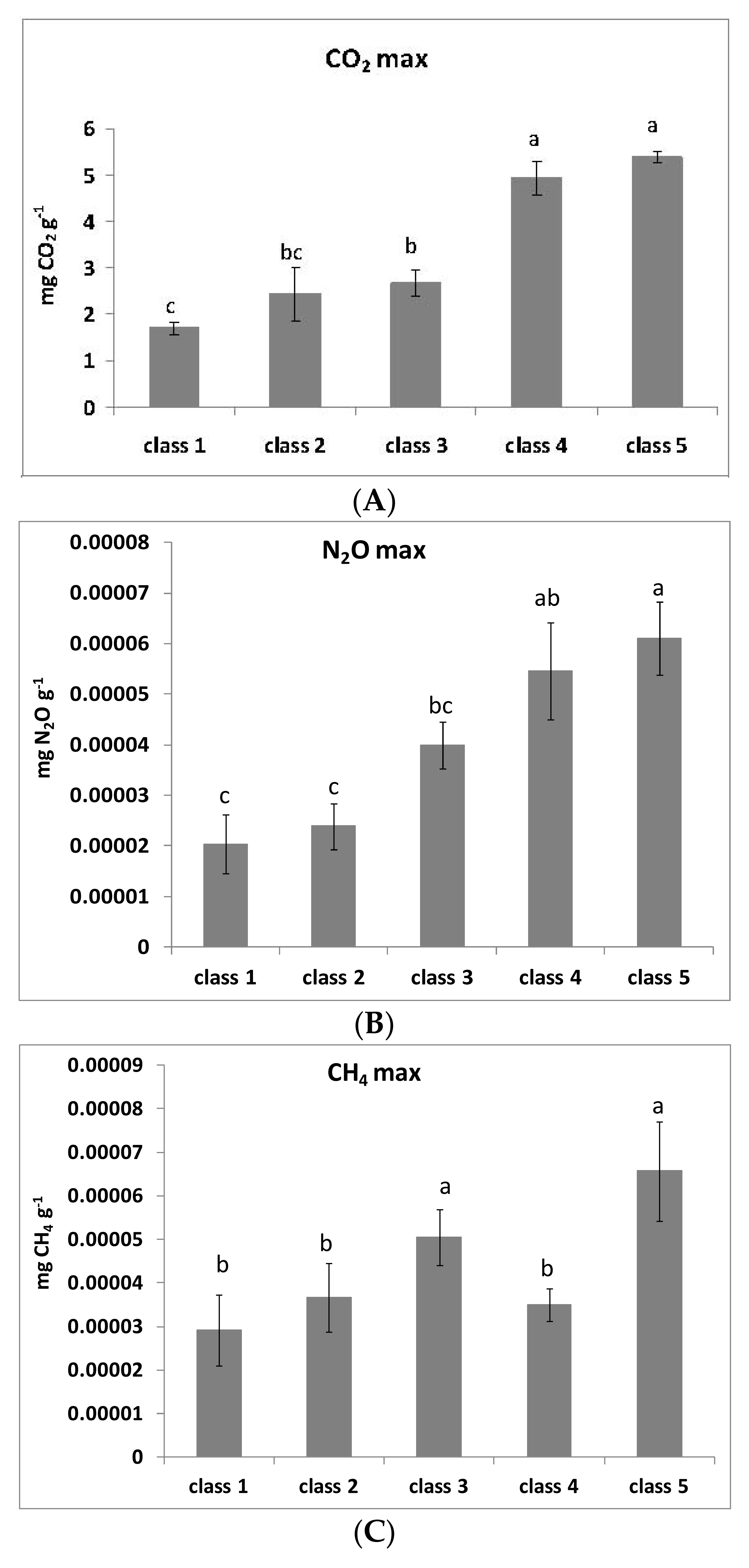
3.1. GHG Potential Production from Deadwood
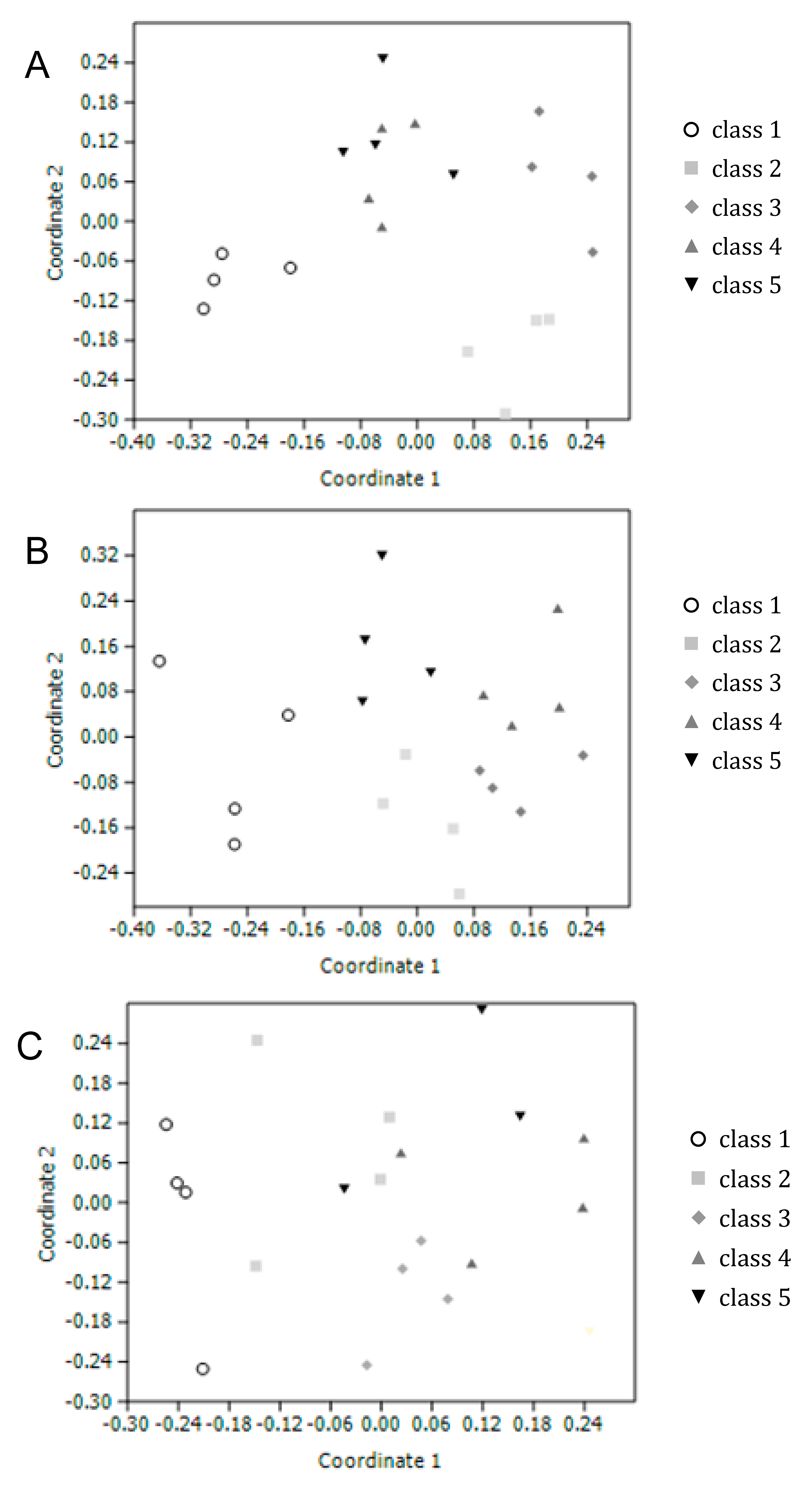
3.2. PCR-DGGE Analysis
3.3. Physical and Chemical Analysis
3.4. Potential Enzymatic Activities
3.5. Correlation of Chemical, Microbiological and Biochemical Data
4. Discussion
5. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Holvoet, B.; Muys, B. Sustainable forest management worldwide: A comparative assessment of standards. Int. For. Rev. 2004, 6, 99–122. [Google Scholar] [CrossRef]
- Wang, S.; Wilson, B. Pluralism in the economics of sustainable forest management. For. Policy Econ. 2007, 9, 743–750. [Google Scholar] [CrossRef]
- Paletto, A.; De Meo, I.; Di Salvatore, U.; Ferretti, F. Perceptions of sustainable forest management practices: An application from the forestry sector in southern Italy. Int. For. Rev. 2014, 16, 55–66. [Google Scholar] [CrossRef]
- Luckert, M.K.; Williamson, T. Should sustained yield be part of sustainable forest management? Can. J. For. Res. 2005, 35, 356–364. [Google Scholar] [CrossRef]
- Putz, F.E. Approaches to Sustainable Forest Management; Working Paper 4; Center for International Forestry Research (CIFOR): Bangor, ME, USA, 1994; pp. 1–6. [Google Scholar]
- Paletto, A.; Ferretti, F.; De Meo, I.; Cantiani, P.; Focacci, M. Ecological and environmental role of deadwood in managed and unmanaged forests. In Sustainable Forest Management—Current Research; Garcia, J.M., Diez, J.J., Eds.; InTech Open Access Publisher: Rijeka, Croatia, 2012; pp. 219–238. [Google Scholar]
- Food and Agriculture Organization (FAO). Global Forest Resources Assessment Update 2005: Terms and Definitions; Forest Resources Assessment Programme, Working Papers 83/E; FAO: Rome, Italy, 2004. [Google Scholar]
- Ligot, G.; Lejeune, P.; Rondeux, J.; Hébert, J. Assessing and harmonizing lying deadwood volume with regional forest inventory data in Wallonia (Southern region of Belgium). Open For. Sci. J. 2012, 5, 15–22. [Google Scholar] [CrossRef]
- Paletto, A.; Tosi, V. Deadwood density variation with decay class in seven tree species of the Italian Alps. Scand. J. For. Res. 2010, 25, 164–173. [Google Scholar] [CrossRef]
- Holub, S.M.; Spears, J.D.H.; Lajtha, K. A reanalysis of nutrient dynamics in coniferous coarse woody debris. Can. J. For. Res. 2001, 31, 1894–1902. [Google Scholar] [CrossRef]
- Laiho, R.; Prescott, C.E. The contribution of coarse woody debris to carbon, nitrogen, and phosphorus cycles in three Rocky Mountain coniferous forests. Can. J. For. Res. 1999, 29, 1592–1603. [Google Scholar] [CrossRef]
- Persiani, A.M.; Lombardi, F.; Lunghini, D.; Granito, V.M.; Tognetti, R.; Maggi, O.; Pioli, S.; Marchetti, M. Stand structure and deadwood amount influences saproxylic fungal biodiversity in Mediterranean mountain unmanaged forests. iForest 2015, 9, 115–124. [Google Scholar] [CrossRef]
- Harmon, M.E.; Franklin, J.F.; Swanson, F.J.; Sollins, P.; Gregory, S.V.; Lattin, J.D.; Anderson, N.H.; Cline, S.P.; Aumen, N.G.; Sedell, J.R.; et al. Ecology of coarse woody debris in temperate ecosystem. Adv. Ecol. Res. 1986, 15, 133–302. [Google Scholar]
- Renvall, P. Community structure and dynamics of wood-rotting basidiomycetes on decomposing conifer trunks in northern Finland. Karstenia 1995, 35, 1–51. [Google Scholar]
- Hoppe, B. Microbial Diversity and Community Structure in Deadwood of Fagus sylvatica L. and Picea abies (L.) H. Karst. Ph.D. Dissertation, Albert-Ludwigs-Universität Freiburg, Freiburg, Germany, 2015. [Google Scholar]
- Boddy, L. Fungal community, ecology and wood decomposition processes in angiosperms: From standing tree to complete decay of coarse woody debris. Ecol. Bull. 2001, 49, 43–56. [Google Scholar]
- Kahl, T.; Baber, K.; Otto, P.; Wirth, C.; Bauhus, J. Drivers of CO2 emission rates from dead wood logs of 13 tree species in the initial decomposition phase. Forests 2015, 6, 2484–2504. [Google Scholar] [CrossRef]
- Hoppe, B.; Purahong, W.; Wubet, T.; Kahl, T.; Bauhus, J.; Arnstadt, T.; Hofrichter, M.; Buscot, F.; Krüger, D. Linking molecular deadwood-inhabiting fungal diversity and community dynamics to ecosystem functions and processes in Central European forests. Fungal Divers. 2016, 77, 367–379. [Google Scholar] [CrossRef]
- Clausen, C.A. Bacterial associations with decaying wood: A review. Int. Biodeterior. Biodegrad. 1996, 37, 101–107. [Google Scholar] [CrossRef]
- Purahong, W.; Arnstadt, T.; Kahl, T.; Bauhus, J.; Kellner, H.; Hofrichter, M.; Krüger, D.; Buscot, F.; Hoppe, B. Are correlations between deadwood fungal community structure, wood physico-chemical properties and lignin-modifying enzymes stable across different geographical regions? Fungal Ecol. 2016, 22, 98–105. [Google Scholar] [CrossRef]
- Kahl, T.; Arnstadt, T.; Baber, K.; Bässler, C.; Bauhus, J.; Borken, W.; Buscot, F.; Floren, A.; Heibl, C.; Hessenmöller, D.; et al. Wood decay rates of 13 temperate tree species in relation to wood properties, enzyme activities and organismic diversities. For. Ecol. Manag. 2017, 391, 86–95. [Google Scholar] [CrossRef]
- Rock, J.; Badeck, F.-W.; Harmon, M.E. Estimating decomposition rate constants for European tree species from literature sources. Eur. J. For. Res. 2008, 127, 301–313. [Google Scholar] [CrossRef]
- Hale, C.M.; Pastor, J. Nitrogen content, decay rates, and decompositional dynamics of hollow versus solid hardwood logs in hardwood forests of Minnesota, USA. Can. J. For. Res. 1998, 28, 1276–1285. [Google Scholar] [CrossRef]
- Alban, D.H.; Pastor, J. Decomposition of aspen, spruce, and pine boles on two sites in Minnesota. Can. J. For. Res. 1993, 23, 1744–1749. [Google Scholar] [CrossRef]
- Nordén, B.; Ryberg, M.; Götmark, F.; Olausson, B. Relative importance of coarse and fine woody debris for the diversity of wood-inhabiting fungi in temperate broadleaf forests. Biol. Conserv. 1994, 117, 1–10. [Google Scholar] [CrossRef]
- Nordén, B.; Paltto, H. Wood-decay fungi in hazel wood: Species richness correlated to stand age and dead wood features. Biol. Conserv. 2001, 101, 1–8. [Google Scholar] [CrossRef]
- Christensen, M.; Heilmann-Clausen, J.; Walleyn, R.; Adamcik, S. Wood-inhabiting fungi as indicators of nature value in European beech forests. In Monitoring and Indicator of Forest Biodiversity in Europe. From Ideas to Operationality; Marchetti, M., Ed.; European Forest Institute (EFI): Joensuu, Finland, 2004; pp. 229–237. [Google Scholar]
- Bader, P.; Jansson, S.; Jonsson, B.G. Wood-inhabiting fungi and substratum decline in selectively logged boreal spruce forests. Biol. Conserv. 1995, 72, 355–362. [Google Scholar] [CrossRef]
- Gómez-Brandón, M.; Ascher-Jenull, J.; Bardelli, T.; Fornasier, F.; Fravolini, G.; Arfaioli, P.; Ceccherini, M.T.; Pietramellara, G.; Lamorski, K.; Sławiński, C.; et al. Physico-chemical and microbiological evidence of exposure effects on Picea abies—Coarse woody debris at different stages of decay. For. Ecol. Manag. 2017, 391, 376–389. [Google Scholar] [CrossRef]
- Kan, J.; Wang, K.; Chen, F. Temporal variation and detection limit of an estaurine bacterioplankton community analyzed by denaturing gradient gel electrophoresis (DGGE). Aquat. Microb. Ecol. 2006, 42, 7–18. [Google Scholar] [CrossRef]
- Arrigoni, P.V.; Bechi, N.; Ricceri, C.; Foggi, B. Documenti per la carta della vegetazione del Monte Morello (Provincia di Firenze). Parlatorea 1997, 2, 73–100. [Google Scholar]
- Cenni, E.; Bussotti, F.; Galeotti, L. The decline of a Pinus nigra Arn. reforestation stand on a limestone substrate: The role of nutritional factors examined by means of foliar diagnosis. Ann. Sci. For. 1998, 55, 567–576. [Google Scholar] [CrossRef]
- Næsset, E. Relationship between relative wood density of Picea abies logs and simple classification systems of decayed coarse woody debris. Can. J. For. Res. 1999, 14, 454–461. [Google Scholar] [CrossRef]
- Fridman, J.; Walheim, M. Amount, structure, and dynamics of dead wood on managed forestland in Sweden. For. Ecol. Manag. 2000, 131, 23–36. [Google Scholar] [CrossRef]
- Paletto, A.; Tosi, V. Forest canopy cover and canopy closure: Comparison of assessment techniques. Eur. J. For. Res. 2009, 128, 265–272. [Google Scholar] [CrossRef]
- De Meo, I.; Agnelli, A.E.; Graziani, A.; Kitikidou, K.; Lagomarsino, A.; Milios, E.; Radoglou, K.; Paletto, A. Deadwood volume assessment in Calabrian pine (Pinus brutia Ten.) peri-urban forests: Comparison between two sampling methods. J. Sust. For. 2017. (under review). [Google Scholar]
- Vainio, E.J.; Hantula, J. Direct analysis of wood-inhabiting fungi using denaturing gradient gel electrophoresis of amplified ribosomal DNA. Mycol. Res. 2000, 8, 927–936. [Google Scholar] [CrossRef]
- Nübel, U.; Engelen, B.; Felske, A.; Snairdr, J.; Wieshuber, A.; Amann, R.I.; Ludwig, W.; Backhaus, H. Sequence heterogeneities of genes encoding 16S rRNAs in Paenibacillus polymyxa detected by temperature gradient gel electrophoresis. J. Bacteriol. 1996, 178, 5636–5643. [Google Scholar] [CrossRef] [PubMed]
- Heuer, H.; Krsek, M.; Baker, P.; Smalla, K.; Wellington, E.M.H. Analysis of actinomycetes communities by specific amplification of genes encoding 16S rRNA and gel-electrophoretic separation in denaturing gradients. Appl. Environ. Microbiol. 1997, 63, 3233–3241. [Google Scholar] [PubMed]
- Fromin, N.; Hamelin, J.; Tarnawski, S.; Roesti, D.; Jourdain-Miserez, K.; Forestier, N.; Teyssier-Cuvelle, S.; Gillet, F.; Aragno, M.; Rossi, P. Statistical analysis of denaturing gel electrophoresis (DGE) fingerprinting patterns. Environ. Microbiol. 2002, 4, 634–643. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Yang, D.; Zhang, Y.; Shen, J.; van der Gast, C.; Hahn, M.W.; Wu, Q. Do patterns of bacterial diversity along salinity gradients differ from those observed for macroorganisms? PLoS ONE 2011, 6, e27597. [Google Scholar] [CrossRef] [PubMed]
- Pastorelli, R.; Landi, S.; Trabelsi, D.; Piccolo, R.; Mengoni, A.; Bazzicalupo, M.; Pagliai, M. Effects of soil management on structure and activity of denitrifying bacterial communities. App. Soil Ecol. 2011, 49, 46–58. [Google Scholar] [CrossRef]
- Marx, M.C.; Wood, M.; Jarvis, S.C. A microplate fluorimetric assay for the study of enzyme diversity in soils. Soil Biol. Biochem. 2001, 33, 1633–1640. [Google Scholar] [CrossRef]
- Hammer, Ø.; Harper, D.A.T.; Ryan, P.D. PAST: Palaeontological statistics software package for education and data analysis. Palaeontol. Electron. 2009, 4, 9. [Google Scholar]
- Kraigher, H.; Jurc, D.; Kalan, P.; Kutnar, L.; Levanic, T.; Rupel, M.; Smolej, I. Beech coarse woody debris characteristics in two virgin forest reserves in southern Slovenia. Zb. Gozdarstva Lesar. 2002, 39, 91–134. [Google Scholar]
- Tomescu, R.; Tarziu, D.R.; Turcu, D.O. The importance of dead wood in the forest. ProEnvironment 2011, 4, 10–19. [Google Scholar]
- Green, P.; Peterken, G.F. Variation in the amount of deadwood in the woodlands of the Lower Wye Valley, UK in relation to the intensity of management. For. Ecol Manag. 1997, 98, 229–238. [Google Scholar] [CrossRef]
- Lombardi, F.; Lasserre, B.; Tognetti, B.; Marchetti, M. Deadwood in relation to stand management and forest type in central Apennines (Molise, Italy). Ecosystems 2008, 11, 882–894. [Google Scholar] [CrossRef]
- Nilsson, T.; Björdal, C. Culturing wood-degrading erosion bacteria. Int. Biodeterior. Biodegrad. 2008, 61, 3–10. [Google Scholar] [CrossRef]
- Allen, R.B.; Buchanan, P.K.; Clinton, P.W.; Cone, A.J. Composition and diversity of fungi on decaying logs in a New Zealand temperate beech (Nothofagus) forest. Can. J. For. Res. 2000, 30, 1025–1033. [Google Scholar] [CrossRef]
- Jönsson, M.; Edman, M.; Jonsson, B.G. Colonization and extinction patterns of wood-decaying fungi in a boreal old-growth Picea abies forest. J. Ecol. 2008, 96, 1065–1075. [Google Scholar] [CrossRef]
- Rajala, T.; Peltoniemi, M.; Hantula, J.; Mäkipää, R.; Pennenen, T. RNA reveals a succession of active fungi during the decay of Norway spruce logs. Fungal Ecol. 2011, 4, 437–448. [Google Scholar] [CrossRef]
- Muyzer, G.; de Waal, E.C.; Uitterlinden, A. profiling of complex microbial populations using denaturing gradient gel electrophoresis analysis of polymerase chain reaction-amplified genes coding for 16S rRNA. App. Environ. Microbiol. 1993, 59, 695–700. [Google Scholar]
- Hoppe, B.; Kahl, T.; Karasch, P.; Wubet, T.; Bauhus, J.; Buscot, F.; Krüger, D. Network analysis reveals ecological links between N-fixing bacteria and wood-decaying fungi. PLoS ONE 2014, 9, e88141. [Google Scholar] [CrossRef] [PubMed]
- Cornelissen, J.H.C.; Sass-Klaassen, U.; Poorter, L.; van Geffen, K.; van Logtestijn, R.S.P.; van Hal, J.; Goudzwaard, L.; Sterck, F.J.; Klaassen, R.K.W.M.; Freschet, G.T.; et al. Controls on coarse wood decay in temperate tree species: Birth of the LOGLIFE Experiment. AMBIO 2012, 41, 231–245. [Google Scholar] [CrossRef] [PubMed]
- Valášková, V.; de Boer, W.; Klein Gunnewiek, P.J.A.; Pospíšek, M.; Baldrian, P. Phylogenetic composition and properties of bacteria coexisting with the fungus Hypholoma fasciculare in decaying wood. ISME J. 2009, 3, 1218–1221. [Google Scholar] [CrossRef] [PubMed]
- De Boer, W.; Van der Wal, A. Interactions between saprotrophic basidiomycetes and bacteria. In Ecology of Saptrotrophic Basidiomycetes; Boddy, L., Frankland, J.C., van West, P., Eds.; Academic Press: Amsterdam, The Netherlands, 2008; pp. 143–153. [Google Scholar]
- Lonsdale, D.; Pautasso, M.; Holdenrieder, O. Wood-decaying fungi in the forest: Conservation needs and management options. Eur. J. For. Res. 2008, 127, 1–22. [Google Scholar] [CrossRef]
- Dickie, I.A.; Fukami, T.; Wilkie, J.P.; Allen, R.B.; Buchanan, P.K. Do assembly history effects attenuate from species to ecosystem properties? A field test with wood-inhabiting fungi. Ecol. Lett. 2012, 15, 133–141. [Google Scholar] [CrossRef] [PubMed]
- Fukami, T.; Dickie, I.A.; Wilkie, P.; Paulus, B.C.; Park, D.; Roberts, A.; Buchanan, P.K.; Allen, R.B. Assembly history dictates ecosystem functioning: Evidence from wood decomposer communities. Ecol. Lett. 2010, 13, 675–684. [Google Scholar] [CrossRef] [PubMed]
- Tedersoo, L.; Kõljalg, U.; Hallenberg, N.; Larsson, K.H. Fine scale distribution of ectomycorrhizal fungi and roots across substrate layers including coarse woody debris in a mixed forest. New Phytol. 2003, 159, 153–165. [Google Scholar] [CrossRef]
- White, N.A. The importance of wood-decay fungi in forest ecosystems. In Fungal Biotechnology in Agricultural, Food, and Environmental Applications; Arora, D.K., Ed.; Marcel Dekker: New York, NY, USA, 2003; pp. 375–392. [Google Scholar]
- Folman, L.B.; Klein Gunnewiek, P.J.A.; Boddy, L.; de Boer, W. Impact of white-rot fungi on numbers and community composition of bacteria colonizing beech wood from forest soil. FEMS Micorbiol. Ecol. 2008, 63, 181–191. [Google Scholar] [CrossRef] [PubMed]
- Hoppe, B.; Krüger, D.; Kahl, T.; Arnstadt, T.; Buscot, F.; Bauhus, J.; Wubet, T. A pyrosequencing insight into sprawling bacterial diversity and community dynamics in decaying deadwood logs of Fagus sylvatica and Picea abies. Sci. Rep. 2015, 5, 9456. [Google Scholar] [CrossRef] [PubMed]
- Olajuyigbe, S.O.; Tobin, B.; Gardiner, P.; Nieuwenhuis, M. Stocks and decay dynamics of above- and belowground coarse woody debris in managed Sitka spruce forests in Ireland. For. Ecol. Manag. 2011, 262, 1109–1118. [Google Scholar] [CrossRef]
- Lombardi, F.; Cherubini, P.; Tognetti, R.; Cocozza, C.; Lasserre, B.; Marchetti, M. Investigating biochemical processes to assess deadwood decay of beech and silver fir in Mediterranean mountain forests. Ann. For. Sci. 2013, 70, 101–111. [Google Scholar] [CrossRef]
- Strukelj, M.; Brais, S.; Quideau, S.A.; Angers, V.A.; Kebli, H.; Drapeau, P.; Oh, S.-W. Chemical transformations in downed logs and snags of mixed boreal species during decomposition. Can. J. For. Res. 2013, 43, 785–798. [Google Scholar] [CrossRef]
- Eriksson, K.E.L.; Blanchette, R.; Ander, P. Morphological aspects of wood degradation by fungi and bacteria. In Microbial and Anzymatic Degradation of Wood and Wood Components; Springer: Berlin, Germany, 1990; pp. 1–407. [Google Scholar]
- Noormets, A.; Epron, D.; Domec, J.C.; McNulty, S.G.; Fox, T.; Sun, G.; King, J.S. Effects of forest management on productivity and carbon sequestration: A review and hypothesis. For. Ecol. Manag. 2015, 355, 124–140. [Google Scholar] [CrossRef]
- Wang, C.; Bond-Lamberty, B.; Gower, S.T. Environmental controls on carbon dioxide flux from black spruce coarse woody debris. Oecologia 2002, 132, 374–381. [Google Scholar] [CrossRef] [PubMed]
- Bond-Lamberty, B.; Wang, C.; Gower, S. Annual carbon flux woody debris for a boreal black spruce fire chronosequence. J. Geophys. Res. 2003, 108, 8220. [Google Scholar] [CrossRef]
- Litton, C.M.; Raich, J.W.; Ryan, M.G. Carbon allocation in forest ecosystems. Glob. Chang. Biol. 2007, 13, 2089–2109. [Google Scholar] [CrossRef]
- Jomura, M.; Kominami, Y.; Dannoura, M.; Kanazawa, Y. Spatial variation in respiration from coarse woody debris in a temperate secondary broad-leaved forest in Japan. For. Ecol. Mang. 2008, 255, 149–155. [Google Scholar] [CrossRef]
- Forrester, J.A.; Mladenoff, D.J.; Gower, S.T.; Stoffel, J.L. Interactions of temperature and moisture with respiration from coarse woody debris in experimental forest canopy gaps. For. Ecol. Manag. 2012, 265, 124–132. [Google Scholar] [CrossRef]
- Covey, K.R.; Bueno de Mesquita, C.P.; Oberle, B.; Maynard, D.S.; Bettigole, C.; Crowther, T.W.; Duguid, M.C.; Steven, B.; Zanne, A.E.; Lapin, M.; et al. Greenhouse trace gases in deadwood. Biogeochemistry 2016, 130, 215–226. [Google Scholar] [CrossRef]
- Mukhin, V.A.; Voronin, P.Y. Methanogenic activity of woody debris. Russ. J. Ecol. 2009, 40, 149–153. [Google Scholar] [CrossRef]
- Palviainen, M.; Finér, L.; Laiho, R.; Shorohova, E.; Kapitsa, E.; Vanha-Majamaa, I. Carbon and nitrogen release from decomposing Scots pine, Norway spruce and Silver birch stumps. For. Ecol. Manag. 2010, 259, 390–398. [Google Scholar] [CrossRef]
- Brunner, A.; Kimmins, J.P. Nitrogen fixation in coarse woody debris of Thuja plicata and Tsuga heterophylla forests on northern Vancouver Island. Can. J. For. Res. 2003, 33, 1670–1682. [Google Scholar] [CrossRef]
| Decay Class | Description |
|---|---|
| 1 | Recently dead—bark intact, small branches, wood texture intact, visible small rotten areas under bark |
| 2 | Weakly decayed—bark intact but not attached, twigs partly present, wood texture intact, rotten areas <3 cm |
| 3 | Medium decayed—trace of bark, no twigs, wood texture partly broken, rotten area >3 cm |
| 4 | Very decayed—no bark, no twigs, wood soft, texture with blocky pieces, large rotten areas |
| 5 | Almost decomposed—no bark, no twigs, wood soft, powdery texture, very large rotten areas with musk and lichens |
| DecayClass | Fungi (18S-DGGE) | Bacteria (16S-DGGE) | Actinobacteria (16S-DGGE) | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Richness | Shannon | Richness | Shannon | Richness | Shannon | |||||||
| 1 | 20.2 (0.5) | a | 2.99 (0.02) | a | 25.0 (1.1) | b | 3.19 (0.04) | b | 16.7 (1.9) | a | 2.69 (0.05) | a |
| 2 | 16.7 (0.2) | ab | 2.80 (0.02) | ab | 29.2 (1.8) | ab | 3.33 (0.06) | ab | 13.7 (1.8) | ab | 2.58 (0.12) | a |
| 3 | 18.2 (0.8) | ab | 2.88 (0.05) | ab | 31.5 (0.9) | a | 3.42 (0.03) | a | 14.0 (2.0) | ab | 2.62 (0.14) | a |
| 4 | 17.5 (0.9) | ab | 2.83 (0.06) | ab | 25.0 (1.8) | b | 3.18 (0.07) | b | 12.5 (1.9) | ab | 2.48 (0.14) | ab |
| 5 | 15.7 (2.9) | b | 2.67 (0.20) | b | 28.2 (2.8) | ab | 3.29 (0.11) | ab | 9.0 (0.7) | b | 2.18 (0.08) | b |
| Microbial Group | ANOSIM | PERMANOVA | ||
|---|---|---|---|---|
| R | p< | F | p< | |
| Fungi, 18S-DGGE | 0.827 | 0.001 | 4.67 | 0.001 |
| Bacteria, 16S-DGGE | 0.699 | 0.001 | 3.55 | 0.001 |
| Actinobacteria, 16S-DGGE | 0.381 | 0.001 | 1.91 | 0.001 |
| Decay Class | Moisture (%) | Total N (%) | Total C (%) | C/N Ratio | ||||
|---|---|---|---|---|---|---|---|---|
| 1 | 56.4 (1.9) | a | 0.134 (0.011) | c | 47.85 (0.50) | ab | 366.1 (33) | a |
| 2 | 67.4 (10.6) | a | 0.200 (0.011) | c | 47.52 (0.38) | b | 239.9 (13) | b |
| 3 | 60.2 (4.5) | a | 0.214 (0.012) | c | 46.78 (0.31) | b | 220.7 (11) | b |
| 4 | 62.5 (5.4) | a | 0.690 (0.078) | b | 49.28 (0.75) | a | 74.3 (9) | c |
| 5 | 72.4 (3.2) | a | 0.890 (0.102) | a | 48.14 (0.43) | ab | 56.5 (7) | c |
| Class | β-Cellobiohydrolase | Acid-Phosphatase | β-Glucosidase | NAG | Xylanase | Butyrate Esterase | Acetate Esterase | Arylsulphatase | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | 0 (0) | b | 22.1 (22.1) | c | 0.1 (0.1) | b | 0 (0) | b | 0 (0) | b | 0.7 (0.7) | b | 0 (0) | b | 0 (0) | a |
| 2 | 4.9 (4.9) | b | 796.5 (272.6) | bc | 59.2 (35.6) | b | 29.3 (15.5) | ab | 18.2 (14.5) | b | 245.9 (137.8) | b | 193.5 (106.6) | b | 1.1 (1.4) | a |
| 3 | 22.3 (13.2) | b | 2326.7 (500.4) | ab | 117.6 (17.7) | b | 57.1 (14.0) | ab | 18.5 (2.6) | b | 282.4 (70.2) | b | 431.5 (111.1) | b | 5.3 (2.1) | a |
| 4 | 37.0 (20.1) | b | 1236.5 (592.2) | abc | 346.9 (180.4) | b | 185.4 (122.8) | ab | 38.7 (27.7) | b | 556.5 (371.9) | ab | 820.5 (542.5) | b | 19.6 (17.6) | a |
| 5 | 353.1 (112.7) | a | 2867.1 (916.2) | a | 1763.8 (383.8) | a | 197.2 (63.0) | a | 95.2 (21.1) | a | 1060 2 (227.3) | a | 2944.4 (962.3) | a | 20.5 (10.3) | a |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pastorelli, R.; Agnelli, A.E.; De Meo, I.; Graziani, A.; Paletto, A.; Lagomarsino, A. Analysis of Microbial Diversity and Greenhouse Gas Production of Decaying Pine Logs. Forests 2017, 8, 224. https://doi.org/10.3390/f8070224
Pastorelli R, Agnelli AE, De Meo I, Graziani A, Paletto A, Lagomarsino A. Analysis of Microbial Diversity and Greenhouse Gas Production of Decaying Pine Logs. Forests. 2017; 8(7):224. https://doi.org/10.3390/f8070224
Chicago/Turabian StylePastorelli, Roberta, Alessandro E. Agnelli, Isabella De Meo, Anna Graziani, Alessandro Paletto, and Alessandra Lagomarsino. 2017. "Analysis of Microbial Diversity and Greenhouse Gas Production of Decaying Pine Logs" Forests 8, no. 7: 224. https://doi.org/10.3390/f8070224
APA StylePastorelli, R., Agnelli, A. E., De Meo, I., Graziani, A., Paletto, A., & Lagomarsino, A. (2017). Analysis of Microbial Diversity and Greenhouse Gas Production of Decaying Pine Logs. Forests, 8(7), 224. https://doi.org/10.3390/f8070224








