Forest Inventory with Terrestrial LiDAR: A Comparison of Static and Hand-Held Mobile Laser Scanning
Abstract
:1. Introduction
2. Materials and Methods
2.1. Instrumentation
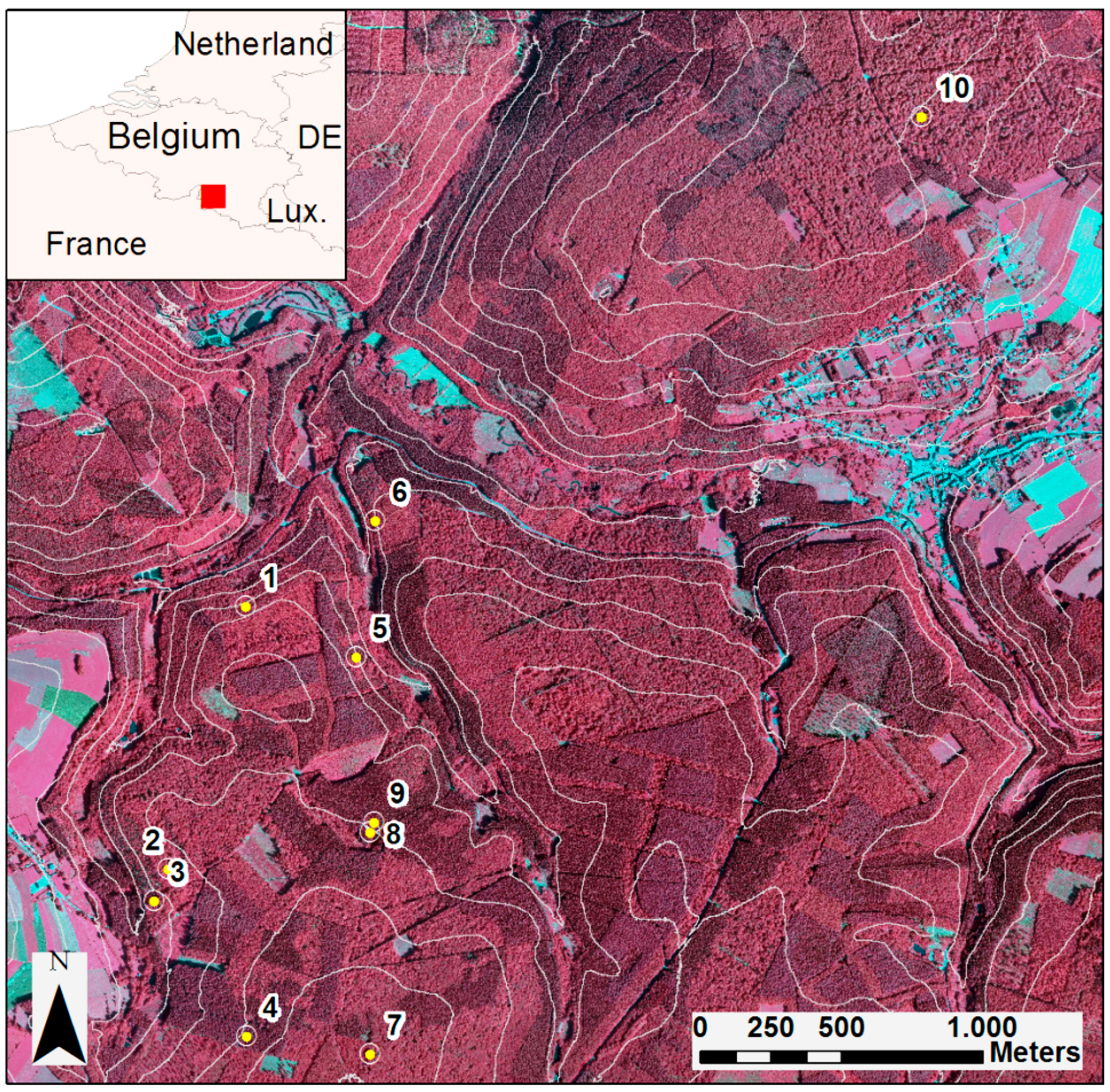
2.2. Study Area
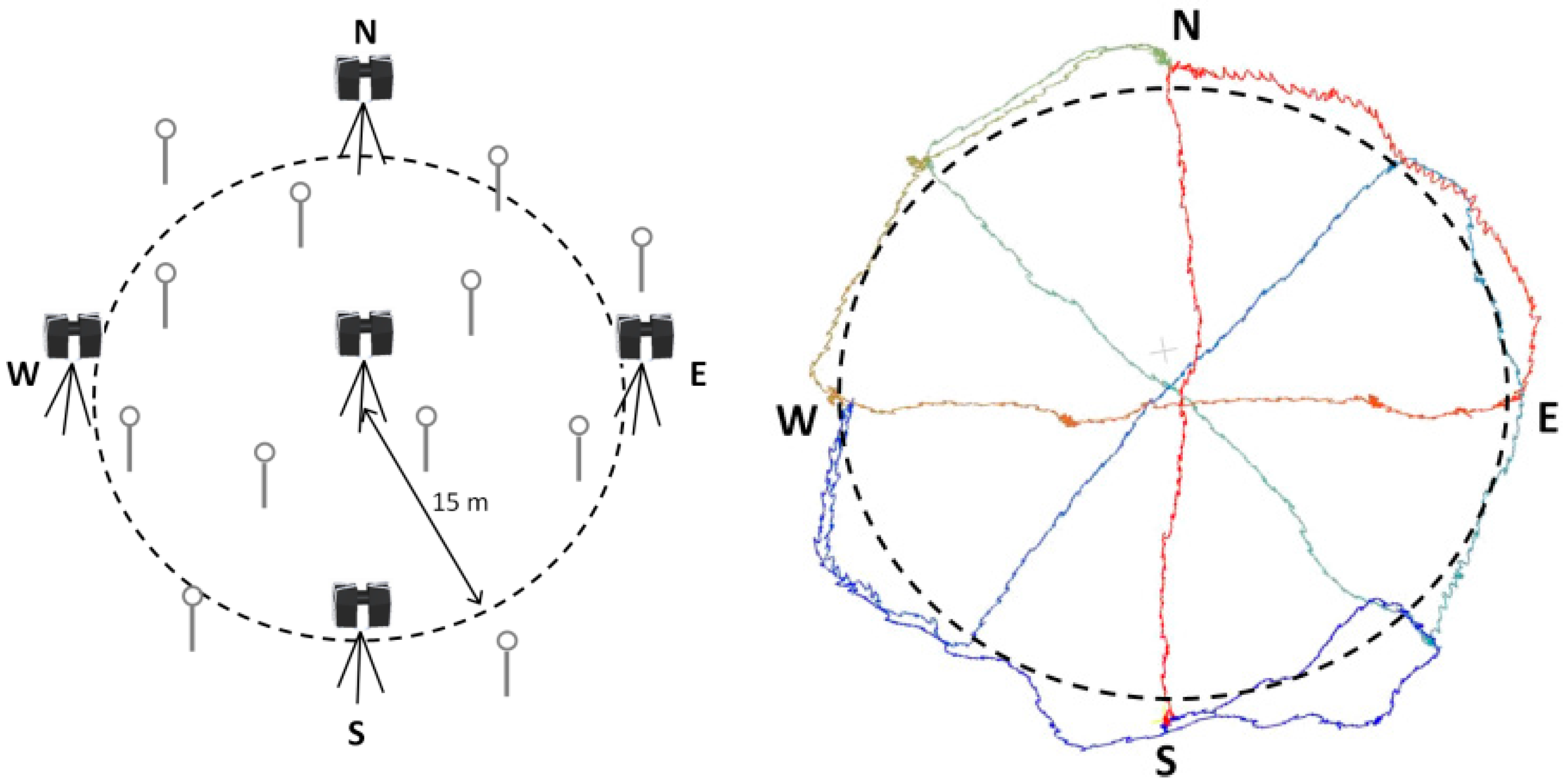
2.3. Data Collection
2.4. LiDAR Data Pre-Processing
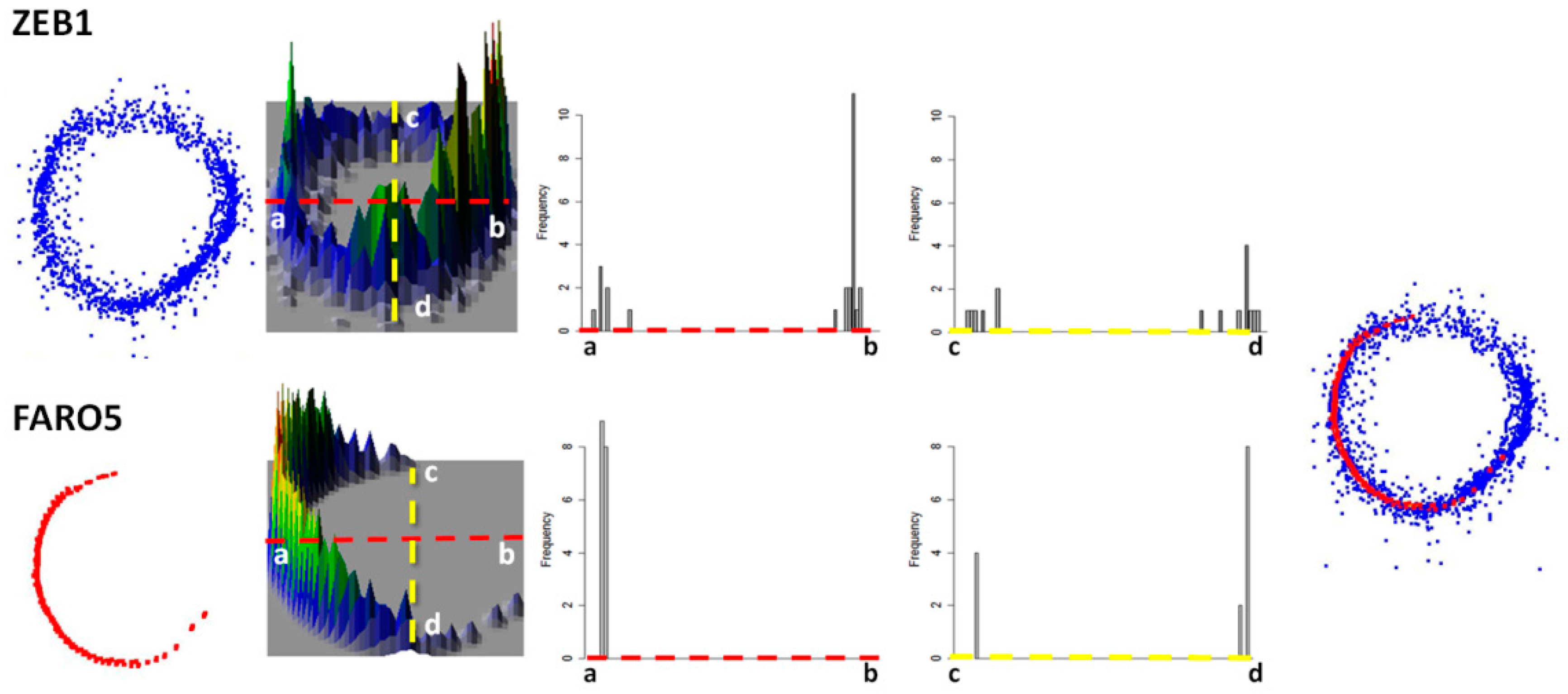
2.5. Scanning Completeness of the Trees
2.6. Extraction of Forest Parameters from the Point Clouds
2.7. Analysis
3. Results
3.1. Data Collection
3.2. TLS Pre-Processing
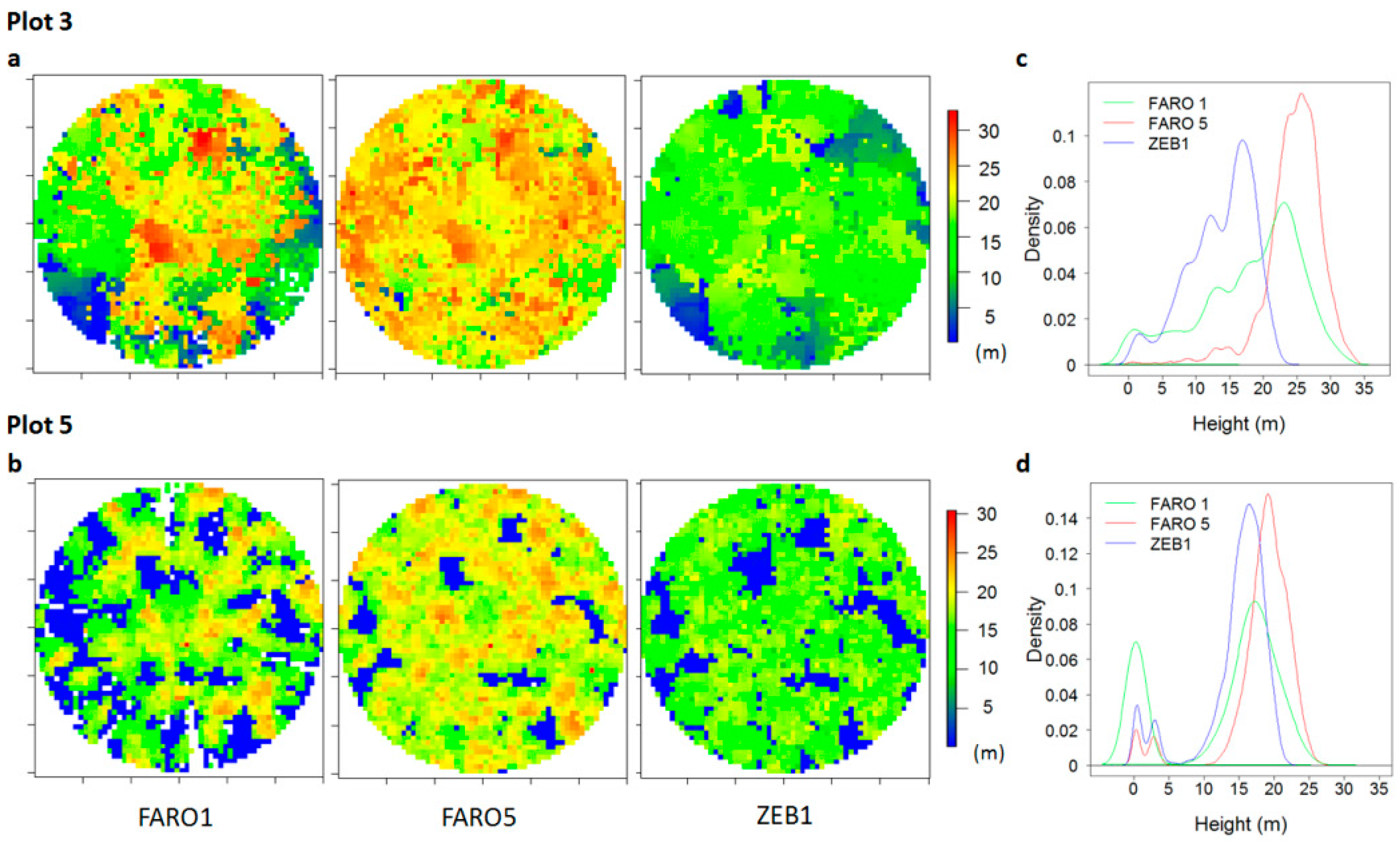
3.3. Visual Comparison
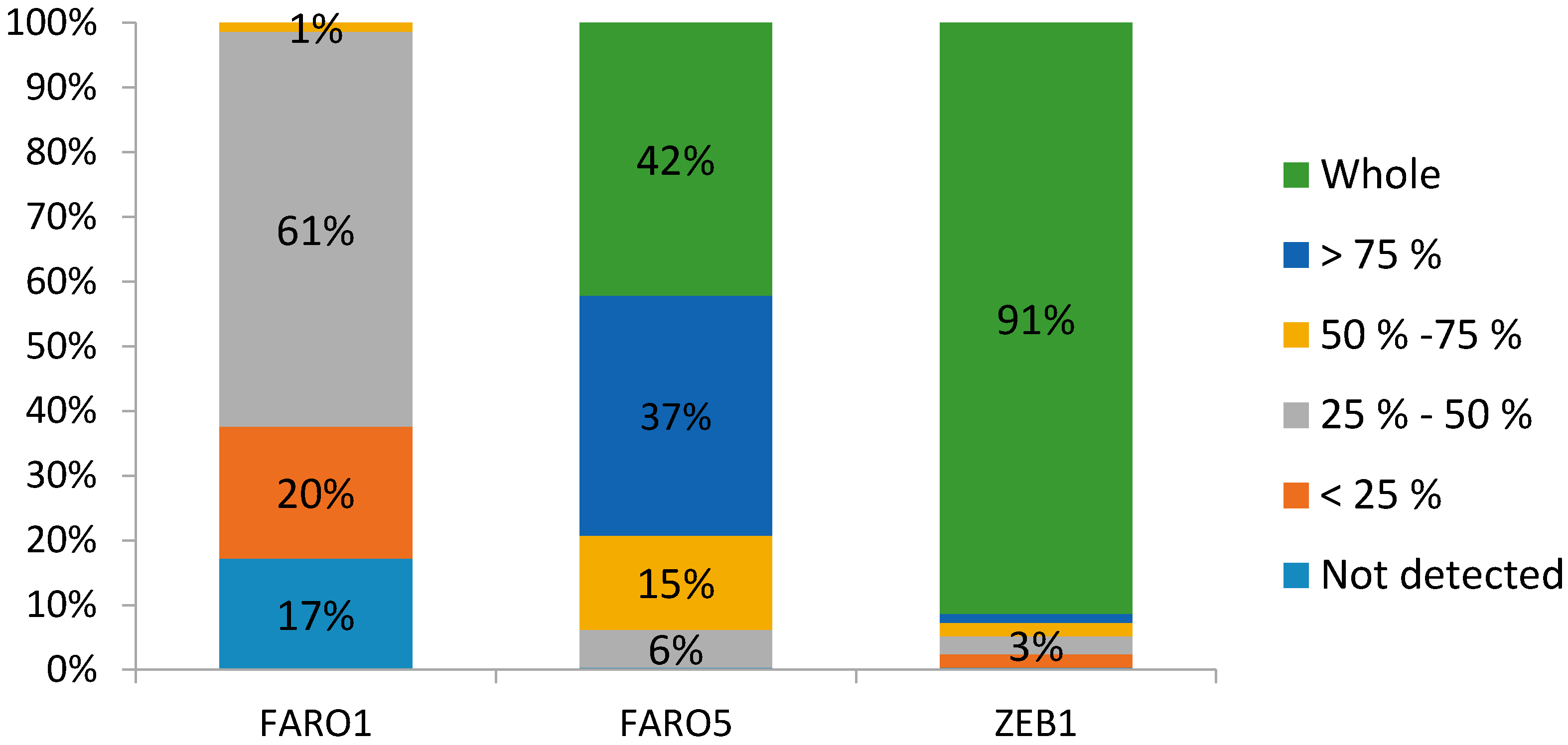
3.4. The Success of Scanning Trees
3.5. Extraction of Forest Parameters
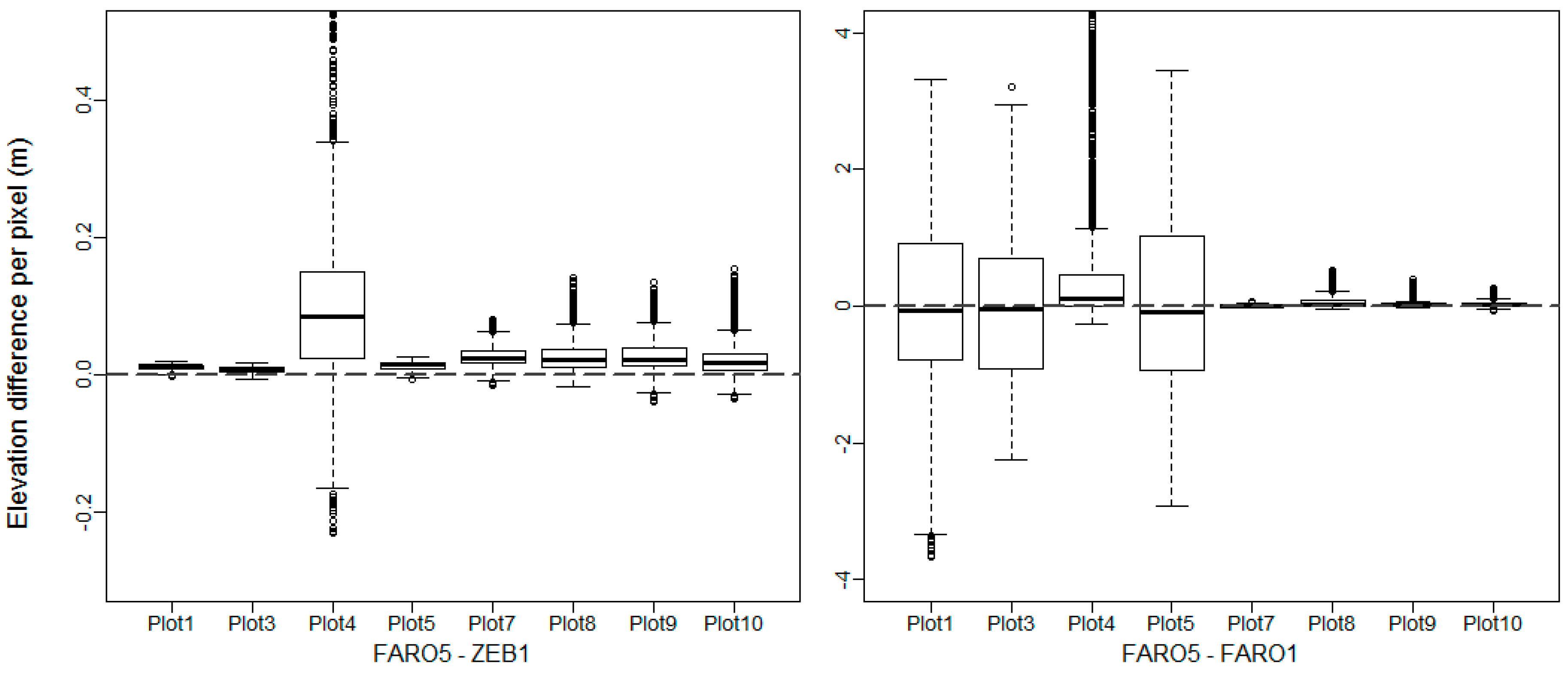
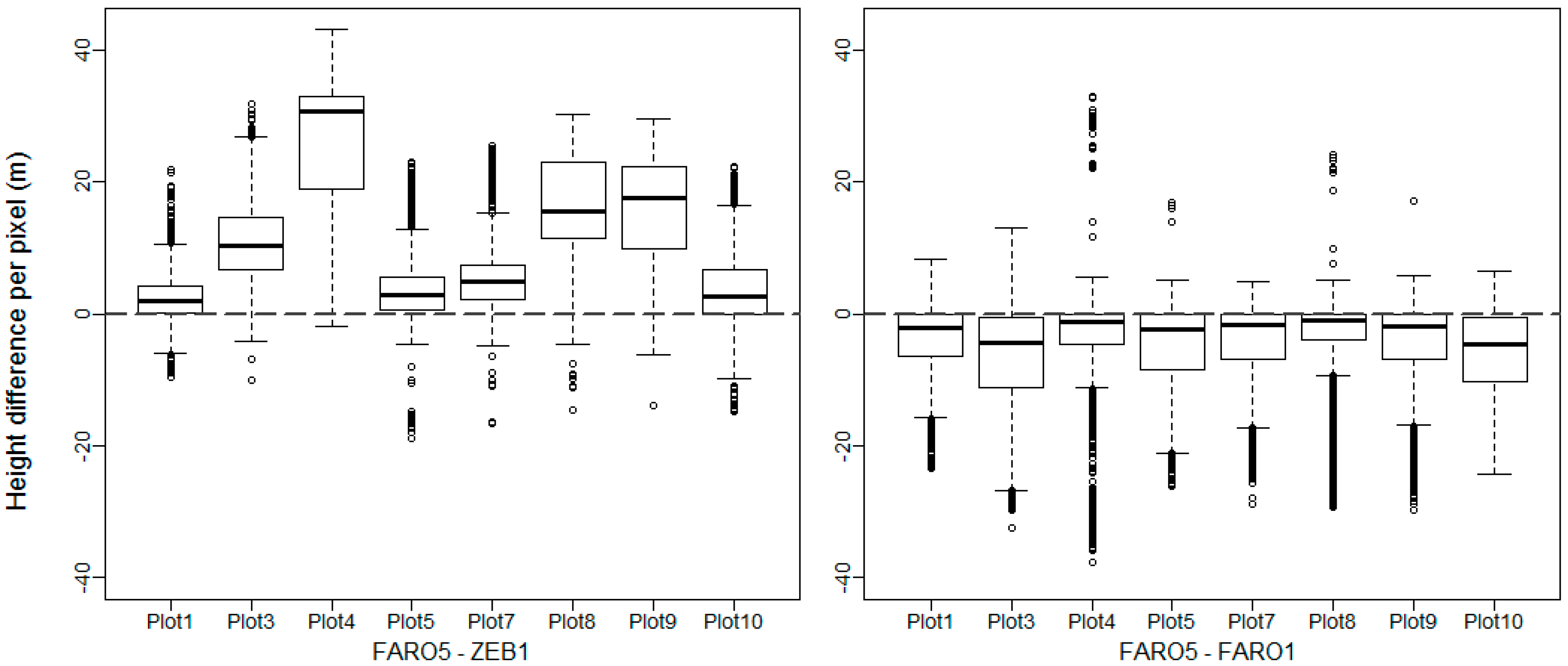
3.5.1. DTM and CHM Comparison
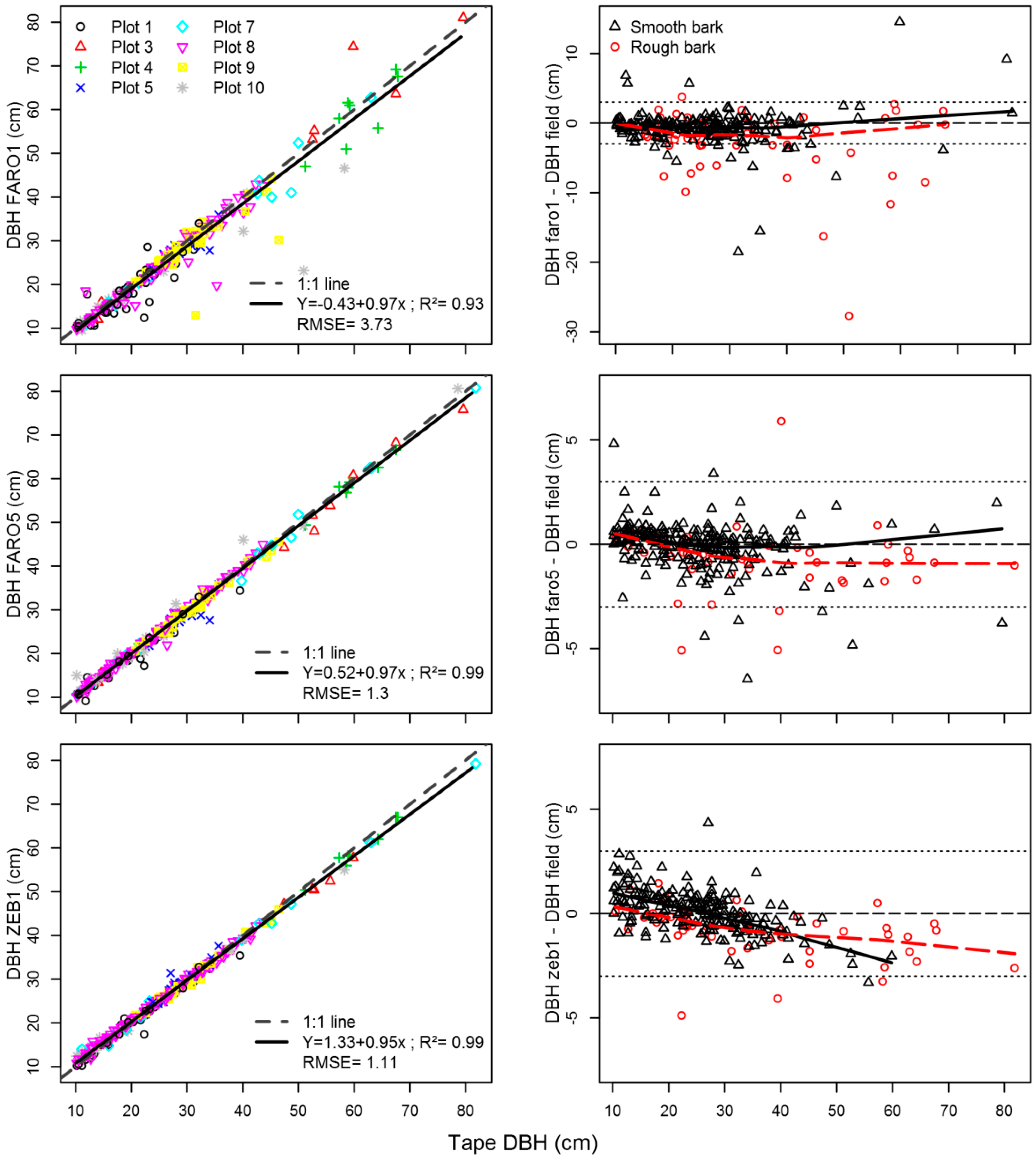
3.5.2. Stem Mapping and DBH Estimation
4. Discussion
5. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Newnham, G.J.; Armston, J.D.; Calders, K.; Disney, M.I.; Lovell, J.L.; Schaaf, C.B.; Strahler, A.H.; Danson, F.M. Terrestrial laser scanning for plot-scale forest measurement. Curr. For. Rep. 2015, 1, 239–251. [Google Scholar] [CrossRef]
- Liang, X. Feasibility of Terrestrial Laser Scanning for Plotwise Forest Inventories. Available online: http://lib.tkk.fi/Diss/2013/isbn9789517112994/isbn9789517112994.pdf (accessed on 15 June 2016).
- Lovell, J.L.; Jupp, D.L.B.; Newnham, G.J.; Culvenor, D.S. Measuring tree stem diameters using intensity profiles from ground-based scanning lidar from a fixed viewpoint. ISPRS J. Photogramm. Remote Sens. 2011, 66, 46–55. [Google Scholar] [CrossRef]
- Van Leeuwen, M.; Nieuwenhuis, M. Retrieval of forest structural parameters using LiDAR remote sensing. Eur. J. For. Res. 2010, 129, 749–770. [Google Scholar] [CrossRef]
- Erikson, M.; Karin, V. Finding tree-stems in laser range images of young mixed stands to perform selective cleaning. In Proceedings of the ScandLaser Scientific Workshop on Airborne Laser Scanning of Forest, Umea, Sweden, 3–4 September 2003.
- Simonse, M.; Aschoff, T.; Spiecker, H.; Thies, M. Automatic determination of forest inventory parameters using terrestrial laserscanning. In Proceedings of the ScandLaser Scientific Workshop on Airborne Laser Scanning of Forest, Umea, Sweden, 3–4 September 2003; Volume 2003, pp. 252–258.
- Watt, P.J.; Donoghue, D.N.M.; Dunford, R.W. Forest Parameter Extraction Using Terrestrial Laser Scanning. Available online: http://www.natscan.uni-freiburg.de/suite/pdf/030916_1642_1.pdf (accessed on 15 June 2016).
- Aschoff, T.; Thies, M.; Spiecker, H. Describing forest stands using terrestrial laser-scanning. Int. Arch. Photogramm. Remote Sens. Spat. Inf. Sci. 2004, 35, 237–241. [Google Scholar]
- Haala, N.; Reulke, R.; Thies, M.; Aschoff, T. Combination of Terrestrial Laser Scanning with High Resolution Panoramic Images for Investigations in Forest Applications and Tree Species Recognition. Available online: http://www.isprs.org/proceedings/XXXIV/5-W16/papers/PanoWS_Dresden2004_Haala.pdf (accessed on 15 June 2016).
- Hopkinson, C.; Chasmer, L.; Young-Pow, C.; Treitz, P. Assessing forest metrics with a ground-based scanning lidar. Can. J. For. Res. 2004, 34, 573–583. [Google Scholar] [CrossRef]
- Thies, M.; Spiecker, H. Evaluation and future prospects of terrestrial laser scanning for standardized forest inventories. Forest 2004, 2, 1. [Google Scholar]
- Côté, J.-F.; Widlowski, J.-L.; Fournier, R.A.; Verstraete, M.M. The structural and radiative consistency of three-dimensional tree reconstructions from terrestrial lidar. Remote Sens. Environ. 2009, 113, 1067–1081. [Google Scholar] [CrossRef]
- Fleck, S.; Mölder, I.; Jacob, M.; Gebauer, T.; Jungkunst, H.F.; Leuschner, C. Comparison of conventional eight-point crown projections with LIDAR-based virtual crown projections in a temperate old-growth forest. Ann. For. Sci. 2011, 68, 1173–1185. [Google Scholar] [CrossRef]
- Bienert, A.; Scheller, S.; Keane, E.; Mohan, F.; Nugent, C. Tree Detection and Diameter Estimations by Analysis of Forest Terrestrial Laserscanner Point Clouds. Available online: http://www.isprs.org/proceedings/XXXVI/3-W52/final_papers/Bienert_2007.pdf (accessed on 15 June 2016).
- Calders, K.; Newnham, G.; Burt, A.; Murphy, S.; Raumonen, P.; Herold, M.; Culvenor, D.; Avitabile, V.; Disney, M.; Armston, J.; et al. Nondestructive estimates of above-ground biomass using terrestrial laser scanning. Methods Ecol. Evol. 2015, 6, 198–208. [Google Scholar] [CrossRef]
- Côté, J.-F.; Fournier, R.A.; Egli, R. An architectural model of trees to estimate forest structural attributes using terrestrial LiDAR. Environ. Model. Softw. 2011, 26, 761–777. [Google Scholar] [CrossRef]
- Dassot, M.; Colin, A.; Santenoise, P.; Fournier, M.; Constant, T. Terrestrial laser scanning for measuring the solid wood volume, including branches, of adult standing trees in the forest environment. Comput. Electron. Agric. 2012, 89, 86–93. [Google Scholar] [CrossRef]
- Lefsky, M.; McHale, M.R. Volume estimates of trees with complex architecture from terrestrial laser scanning. J. Appl. Remote Sens. 2008, 2, 023521. [Google Scholar]
- Othmani, A.; Piboule, A.; Krebs, M.; Stolz, C.; Voon, L.L.Y. Towards Automated and Operational Forest Inventories with T-Lidar. Available online: https://hal.archives-ouvertes.fr/hal-00646403/document (accessed on 15 June 2016).
- Raumonen, P.; Casella, E.; Calders, K.; Murphy, S.; AAkerblom, M.; Kaasalainen, M. Massive-scale tree modelling from TLS data. ISPRS Ann. Photogramm. Remote Sens. Spat. Inf. Sci. 2015, 2, 189–196. [Google Scholar] [CrossRef]
- Huang, H.; Li, Z.; Gong, P.; Cheng, X.; Clinton, N.; Cao, C.; Ni, W.; Wang, L. Automated methods for measuring DBH and tree heights with a commercial scanning lidar. Photogramm. Eng. Remote Sens. 2011, 77, 219–227. [Google Scholar] [CrossRef]
- García, M.; Danson, F.M.; Riano, D.; Chuvieco, E.; Ramirez, F.A.; Bandugula, V. Terrestrial laser scanning to estimate plot-level forest canopy fuel properties. Int. J. Appl. Earth Obs. Geoinform. 2011, 13, 636–645. [Google Scholar] [CrossRef]
- Strahler, A.H.; Jupp, D.L.; Woodcock, C.E.; Schaaf, C.B.; Yao, T.; Zhao, F.; Yang, X.; Lovell, J.; Culvenor, D.; Newnham, G.; et al. Retrieval of forest structural parameters using a ground-based lidar instrument (Echidna®). Can. J. Remote Sens. 2008, 34, S426–S440. [Google Scholar] [CrossRef]
- Danson, F.M.; Hetherington, D.; Morsdorf, F.; Koetz, B.; Allgöwer, B. Forest canopy gap fraction from terrestrial laser scanning. IEEE Geosci. Remote Sens. Lett. 2007, 4, 157–160. [Google Scholar] [CrossRef]
- Lovell, J.L.; Haverd, V.; Jupp, D.L.B.; Newnham, G.J. The Canopy Semi-analytic P gap and Radiative Transfer (CanSPART) model: Validation using ground based LiDAR. Agric. For. Meteorol. 2012, 158, 1–12. [Google Scholar] [CrossRef]
- Hosoi, F.; Omasa, K. Factors contributing to accuracy in the estimation of the woody canopy leaf area density profile using 3D portable lidar imaging. J. Exp. Bot. 2007, 58, 3463–3473. [Google Scholar] [CrossRef] [PubMed]
- Jupp, D.L.; Culvenor, D.S.; Lovell, J.L.; Newnham, G.J. Evaluation and validation of canopy laser radar (LIDAR) systems for native and plantation forest inventory. Final Report to the Forest and Wood Products Research & Development Corporation( FWPRDC: PN 02.2902) CSIRO 2005, 20, 192–197. [Google Scholar]
- Liang, X.; Kukko, A.; Kaartinen, H.; Hyyppä, J.; Yu, X.; Jaakkola, A.; Wang, Y. Possibilities of a Personal Laser Scanning System for Forest Mapping and Ecosystem Services. Sensors 2014, 14, 1228–1248. [Google Scholar] [CrossRef] [PubMed]
- Kaartinen, H.; Hyyppä, J.; Kukko, A.; Jaakkola, A.; Hyyppä, H. Benchmarking the performance of mobile laser scanning systems using a permanent test field. Sensors 2012, 12, 12814–12835. [Google Scholar] [CrossRef]
- Jaakkola, A.; Hyyppä, J.; Kukko, A.; Yu, X.; Kaartinen, H.; Lehtomäki, M.; Lin, Y. A low-cost multi-sensoral mobile mapping system and its feasibility for tree measurements. ISPRS J. Photogramm. Remote Sens. 2010, 65, 514–522. [Google Scholar] [CrossRef]
- Rutzinger, M.; Pratihast, A.K.; Oude Elberink, S.; Vosselman, G. Detection and modelling of 3D trees from mobile laser scanning data. Int. Arch. Photogramm. Remote Sens. Spat. Inf. Sci. 2010, 38, 520–525. [Google Scholar]
- Holopainen, M.; Kankare, V.; Vastaranta, M.; Liang, X.; Lin, Y.; Vaaja, M.; Yu, X.; Hyyppä, J.; Hyyppä, H.; Kaartinen, H.; et al. Tree mapping using airborne, terrestrial and mobile laser scanning—A case study in a heterogeneous urban forest. Urban For. Urban Green. 2013, 12, 546–553. [Google Scholar] [CrossRef]
- Kukko, A.; Kaartinen, H.; Hyyppä, J.; Chen, Y. Multiplatform mobile laser scanning: Usability and performance. Sensors 2012, 12, 11712–11733. [Google Scholar] [CrossRef]
- Bosse, M.; Zlot, R.; Flick, P. Zebedee: Design of a spring-mounted 3-d range sensor with application to mobile mapping. IEEE Trans. Robot. 2012, 28, 1104–1119. [Google Scholar] [CrossRef]
- Ryding, J.; Williams, E.; Smith, M.J.; Eichhorn, M.P. Assessing Handheld Mobile Laser Scanners for Forest Surveys. Remote Sens. 2015, 7, 1095–1111. [Google Scholar] [CrossRef]
- James, M.R.; Quinton, J.N. Ultra-rapid topographic surveying for complex environments: The hand-held mobile laser scanner (HMLS). Earth Surf. Process. Landf. 2014, 39, 138–142. [Google Scholar] [CrossRef]
- Trochta, J.; Král, K.; Janík, D.; Adam, D. Arrangement of terrestrial laser scanner positions for area-wide stem mapping of natural forests. Can. J. For. Res. 2013, 43, 355–363. [Google Scholar] [CrossRef]
- FARO Scene—version 5.4. FARO Verwaltungs GmbH. 2015. Available online: http://www.faro.com/ (accessed on 15 June 2016).
- CloudCompare—version 2.6. Available online: http://www.cloudcompare.org/ (accessed on 15 June 2016).
- RIEGL LMS, RiSCAN PRO Version 2.0. Software Description and User’s Instructions. 2016. Available online: http://www.riegl.com/ (accessed on 15 June 2016).
- Dagnélie, P. Statistique Théorique et Appliquée, Tome 2: Inférences à une et à Deux Dimensions; Bruxelles-Université, Ed.; De Boeck & Larcier: Bruxelles, Belgium, 2006. [Google Scholar]
- Brolly, G.; Kiraly, G. Algorithms for stem mapping by means of terrestrial laser scanning. Acta Silv. Lignaria Hung. 2009, 5, 119–130. [Google Scholar]
- Wezyk, P.; Koziol, K.; Glista, M.; Pierzchalski, M. Terrestrial Laser Scanning versus Traditional Forest Inventory. First Results from the Polish Forests. Available online: http://www.isprs.org/proceedings/XXXVI/3-W52/final_papers/Wezyk_2007.pdf (accessed on 15 June 2016).
- Pfeifer, N.; Winterhalder, D. Modelling of tree cross sections from terrestrial laser scanning data with free-form curves. Int. Arch. Photogramm. Remote Sens. Spat. Inf. Sci. 2004, 36, 76–81. [Google Scholar]
- Murphy, G. Determining stand value and log product yields using terrestrial lidar and optimal bucking: A case study. J. For. 2008, 106, 317–324. [Google Scholar]
- Pueschel, P.; Newnham, G.; Rock, G.; Udelhoven, T.; Werner, W.; Hill, J. The influence of scan mode and circle fitting on tree stem detection, stem diameter and volume extraction from terrestrial laser scans. ISPRS J. Photogramm. Remote Sens. 2013, 77, 44–56. [Google Scholar] [CrossRef]
- Schilling, A.; Maas, H.-G.; Lingnau, C. Tree detection by row recovery on Eucalyptus spp. Plantations from TLS data. In Proceedings of the EARSeL 34th Symposium, Warsaw, Poland, 16–20 June 2014.
| Plot | Forest Type | Slope (%) | Stand Structure | Main Species | Under-story | NHA (N) (trees ha−1) | GHA (m2·ha−1) | Dmin–Dmax (cm) |
|---|---|---|---|---|---|---|---|---|
| 1 | B | 32.5 | Coppice | Carpinus betulus and Betula spp. | 1 | 835 (59) | 46.2 | 10–135 |
| 2 | B | 11 | Even-aged | Fagus sylvatica | 0 | 113 (8) | 29.8 | 44–68 |
| 3 | B | 11.2 | Uneven-aged | Fagus sylvatica | 1 | 127 (9) | 28.6 | 14–80 |
| 4 | C | 16.9 | Even-aged | Pseudotsuga menziesii | 1 | 113 (8) | 32.9 | 51–68 |
| 5 | C | 28.7 | Even-aged | Picea abies | 0 | 410 (29) | 26.3 | 23–36 |
| 6 | M | 24.9 | Even-aged | Quercus spp. and Pinus sylvestris | 2 | 439 (31) | 20.0 | 11–40 |
| 7 | B | 5.1 | Uneven-aged | Fagus sylvatica | 1 | 283 (20) | 33.5 | 10–82 |
| 8 | C | 6.2 | Even-aged | Picea abies | 0 | 1344 (95) | 75.5 | 10–44 |
| 9 | C | 6.0 | Even-aged | Picea abies | 0 | 594 (42) | 45.8 | 21–46 |
| 10 | B | 10.5 | Uneven-aged | Fagus sylvatica | 1 | 424 (30) | 30.0 | 10–79 |
| FARO1 | FARO5 | ZEB1 | Field Measurements | ||
|---|---|---|---|---|---|
| Fieldwork | |||||
| Setting up | 6 min | 40 min | 11 min | 20–45 min | |
| Scan(s) | 4–6 min | 35 min | 13 min | ||
| Total | 10 min | 1 h 15 min | 24 min | 32 min | |
| Processing data * | |||||
| Registering | 5 min | 37 min | 20 min | ||
| Computree | 4 min | 47 min | 1 h 26 min | ||
| Total | 9 min | 1 h 24 min | 1 h 46 min | 10 min | |
| Plot | FARO1 | FARO5 | ZEB1 | ||||||
|---|---|---|---|---|---|---|---|---|---|
| Max (m) | Mean (m) | CV (%) | Max (m) | Mean (m) | CV (%) | Max (m) | Mean (m) | CV (%) | |
| 1 | 22.6 | 14.1 | 34 | 29.5 | 17.4 | 17 | 22.1 | 14.9 | 17 |
| 3 | 32.8 | 19.0 | 40 | 33.5 | 24.5 | 17 | 24.0 | 14.0 | 34 |
| 4 | 40.4 | 29.9 | 38 | 45.9 | 33.0 | 39 | 24.6 | 15.8 | 29 |
| 5 | 30.6 | 17.5 | 46 | 30.4 | 18.9 | 26 | 21.9 | 15.3 | 32 |
| 7 | 26.1 | 17.5 | 43 | 29.0 | 20.1 | 21 | 21.3 | 15.2 | 29 |
| 8 | 33.3 | 23.5 | 31 | 33.3 | 25.3 | 15 | 21.0 | 12.6 | 50 |
| 9 | 32.0 | 21.0 | 43 | 30.7 | 22.9 | 23 | 22.3 | 12.7 | 52 |
| 10 | 25.7 | 10.4 | 46 | 28.6 | 15.9 | 38 | 21.7 | 12.2 | 32 |
| Setup | Bias | RMSE (cm) | RMSE (%) |
|---|---|---|---|
| FARO1 | −1.17 | 3.73 | 13.4 |
| FARO5 | −0.17 | 1.3 | 4.7 |
| ZEB1 | −0.08 | 1.11 | 4.1 |
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bauwens, S.; Bartholomeus, H.; Calders, K.; Lejeune, P. Forest Inventory with Terrestrial LiDAR: A Comparison of Static and Hand-Held Mobile Laser Scanning. Forests 2016, 7, 127. https://doi.org/10.3390/f7060127
Bauwens S, Bartholomeus H, Calders K, Lejeune P. Forest Inventory with Terrestrial LiDAR: A Comparison of Static and Hand-Held Mobile Laser Scanning. Forests. 2016; 7(6):127. https://doi.org/10.3390/f7060127
Chicago/Turabian StyleBauwens, Sébastien, Harm Bartholomeus, Kim Calders, and Philippe Lejeune. 2016. "Forest Inventory with Terrestrial LiDAR: A Comparison of Static and Hand-Held Mobile Laser Scanning" Forests 7, no. 6: 127. https://doi.org/10.3390/f7060127
APA StyleBauwens, S., Bartholomeus, H., Calders, K., & Lejeune, P. (2016). Forest Inventory with Terrestrial LiDAR: A Comparison of Static and Hand-Held Mobile Laser Scanning. Forests, 7(6), 127. https://doi.org/10.3390/f7060127







