Above-Ground Biomass and Biomass Components Estimation Using LiDAR Data in a Coniferous Forest
Abstract
:1. Introduction
2. Materials and Methods
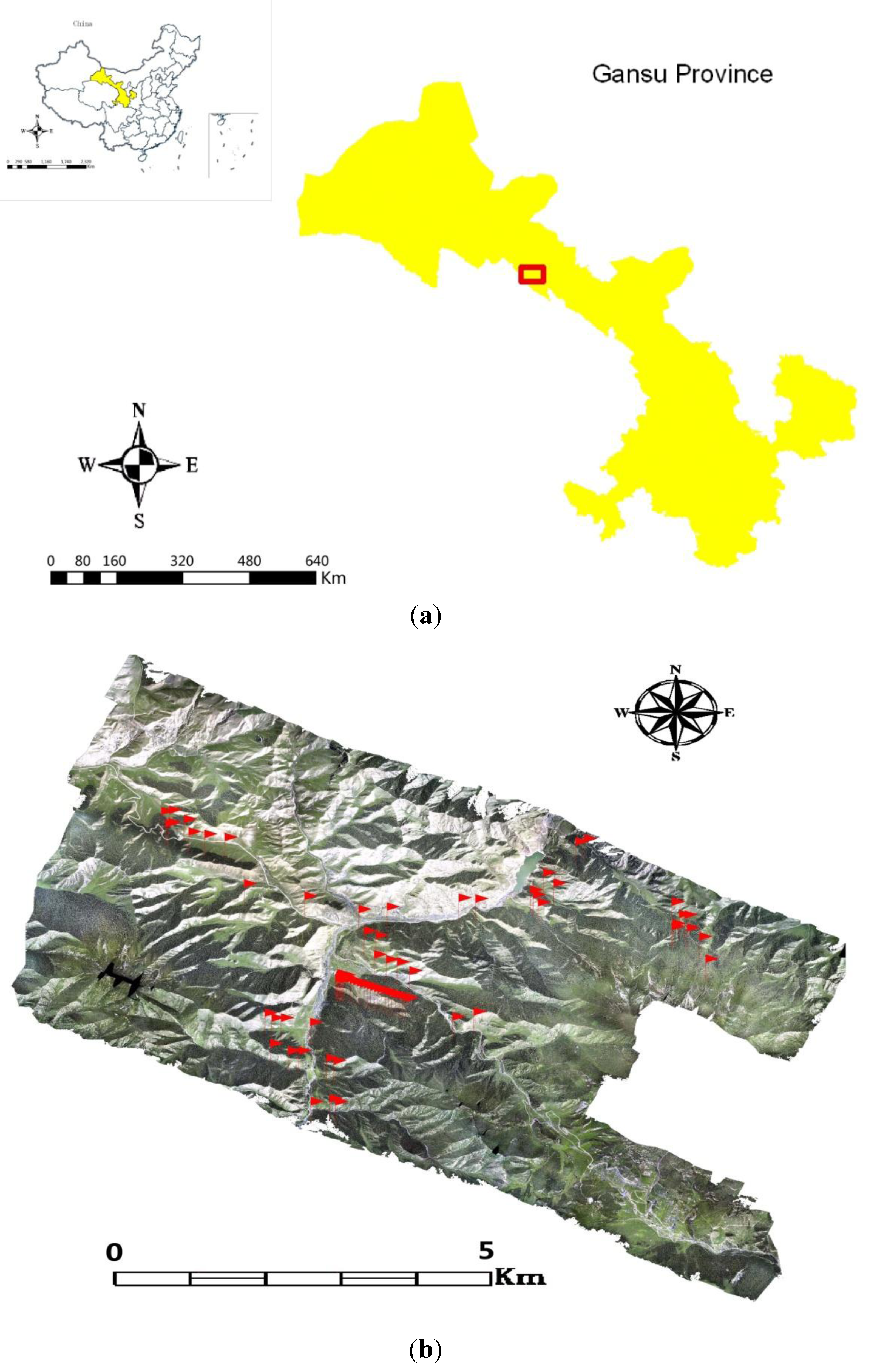
2.1. Study Area
2.2. LiDAR Data Acquisition
2.3. Plot Measurements
2.4. Field-Based Biomass Estimates
2.5. Calculation of LiDAR Metrics
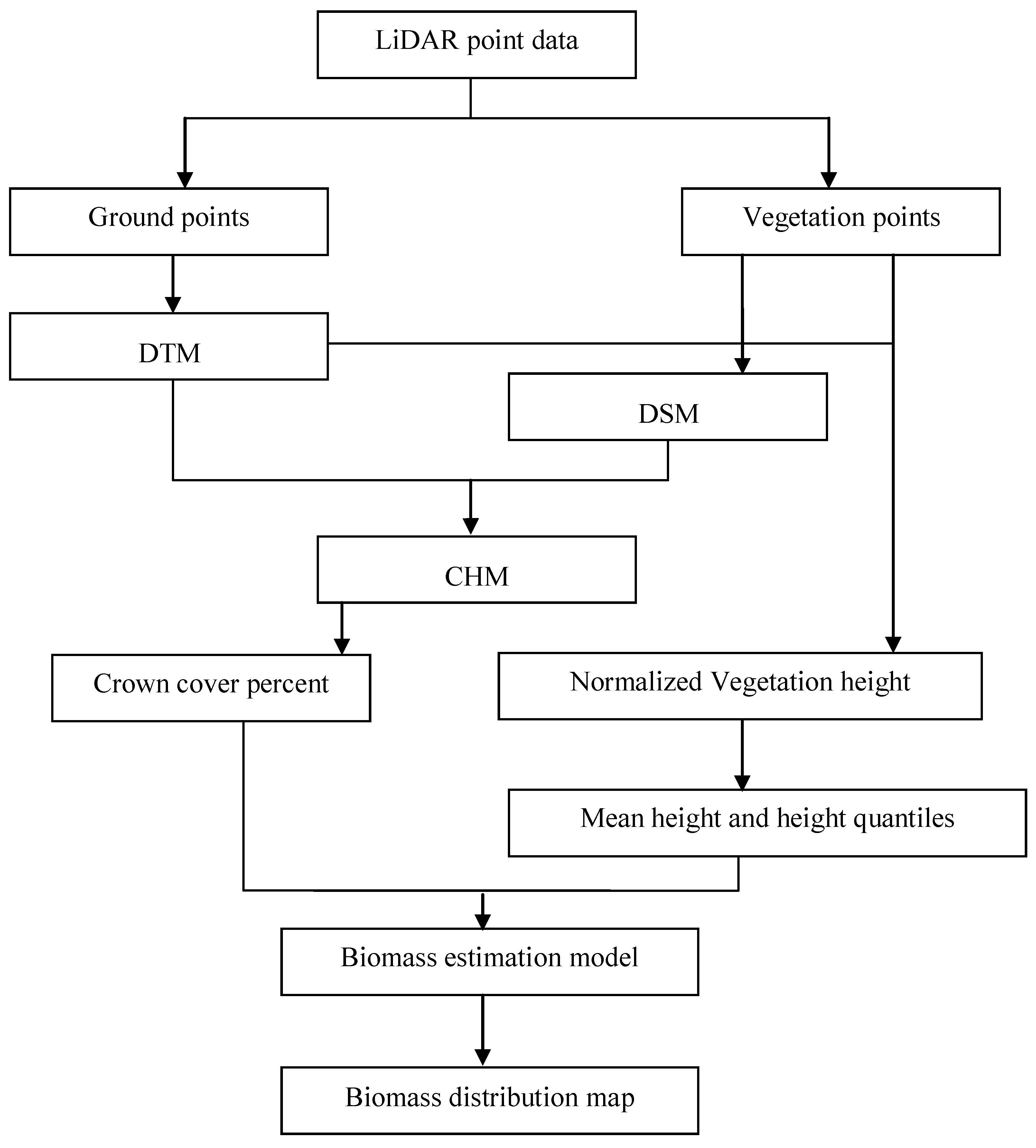
2.6. Statistical Analysis

3. Results
3.1. Biophysical Parameters Description
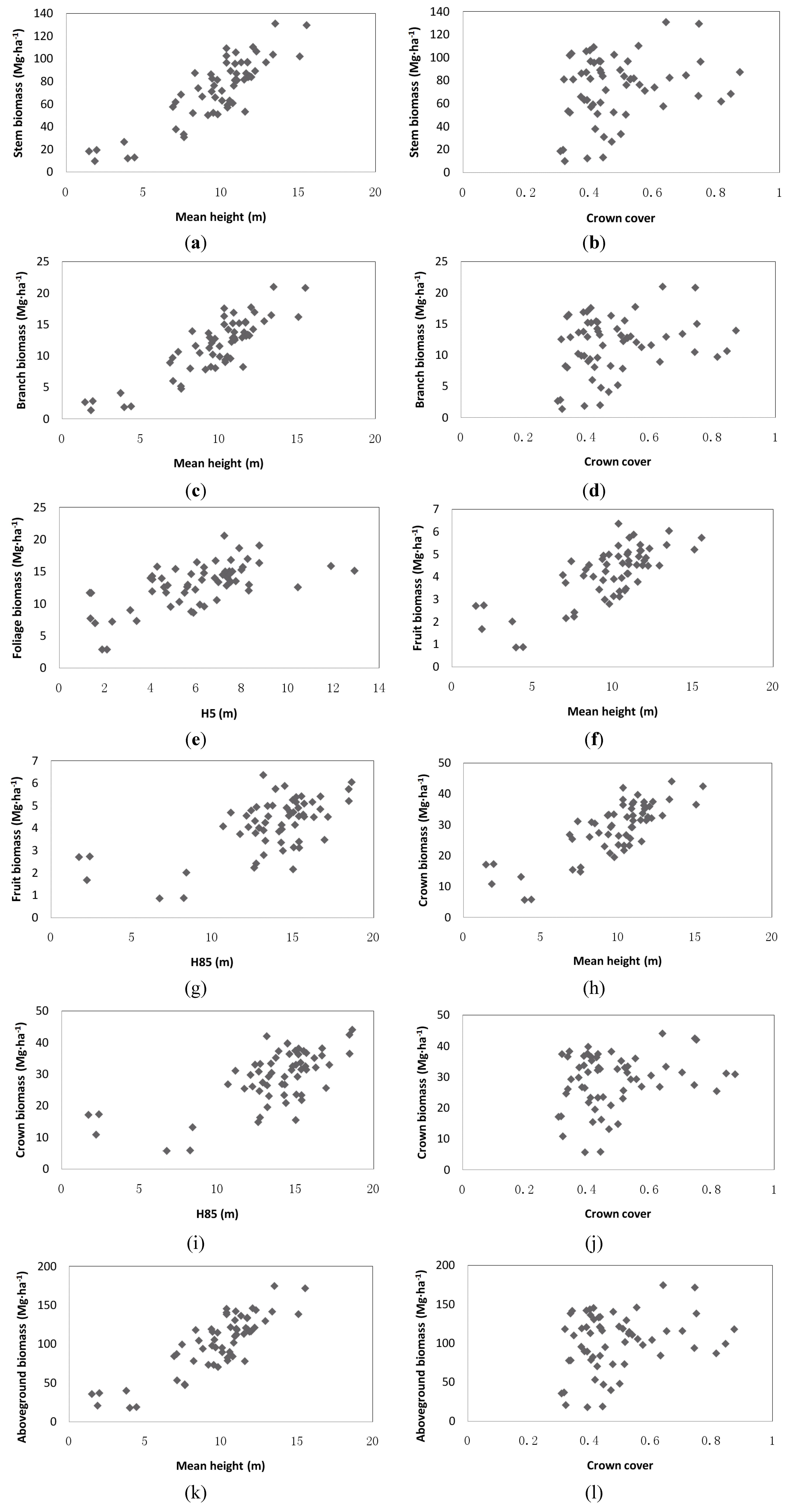
3.2. Regression Models between Biomass and LiDAR Metrics
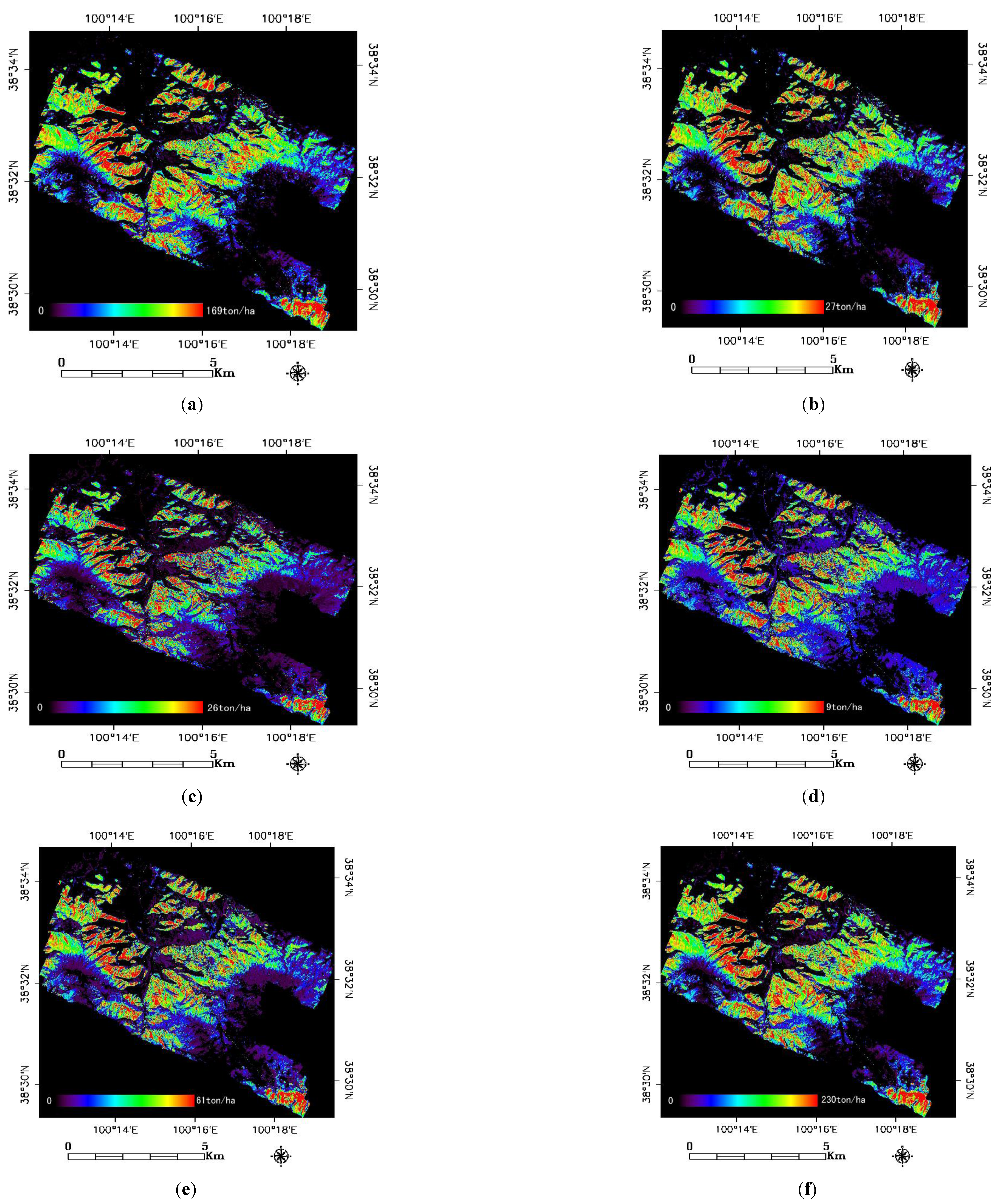
3.3. Spatial Distribution Figure of Forest Biomass
| Statistic | Minimum | Maximum | Mean | Standard deviation |
|---|---|---|---|---|
| Stem biomass | 9.80 | 130.82 | 75.19 | 26.02 |
| Branch biomass | 1.41 | 21.00 | 11.92 | 4.21 |
| Foliage biomass | 2.92 | 22.87 | 13.19 | 3.47 |
| Fruit biomass | 0.87 | 6.85 | 4.23 | 1.17 |
| Crown biomass | 5.68 | 44.42 | 29.34 | 8.23 |
| Above-ground biomass | 17.89 | 174.88 | 104.53 | 33.86 |
| Variables | Std | R2 | Adjusted R2 | Model | |
|---|---|---|---|---|---|
| Stem biomass | Mean, CC | 13.823 | 0.756 | 0.748 | −13.595 + 8.446Mean + 20.378CC |
| Branch biomass | Mean, CC | 2.229 | 0.757 | 0.749 | −2.447 + 1.367Mean + 3.300CC |
| Foliage biomass | H5 | 2.692 | 0.366 | 0.356 | 7.767 + 0.861H5 |
| Fruit biomass | Mean, H85 | 9.507 | 0.591 | 0.578 | 1.726 + 0.541Mean − 0.210H85 |
| Crown biomass | Mean, H85, CC | 5.025 | 0.664 | 0.648 | 8.017 + 4.038Mean − 1.502H85 + 7.287CC |
| Above-ground biomass | Mean, CC | 18.640 | 0.736 | 0.727 | −9.013 + 10.812Mean + 25.105CC |
| Biomass | Stem | Branch | Foliage | Fruit | Crown | Above-ground |
|---|---|---|---|---|---|---|
| RMSE(Mg·ha−1) | 9.876 | 1.520 | 3.691 | 1.022 | 5.963 | 15.237 |
| Relative RMSE(%) | 12.783 | 12.423 | 26.953 | 23.273 | 19.665 | 14.163 |
| Accuracy(%) | 87.45 | 87.60 | 80.00 | 80.15 | 82.59 | 87.08 |

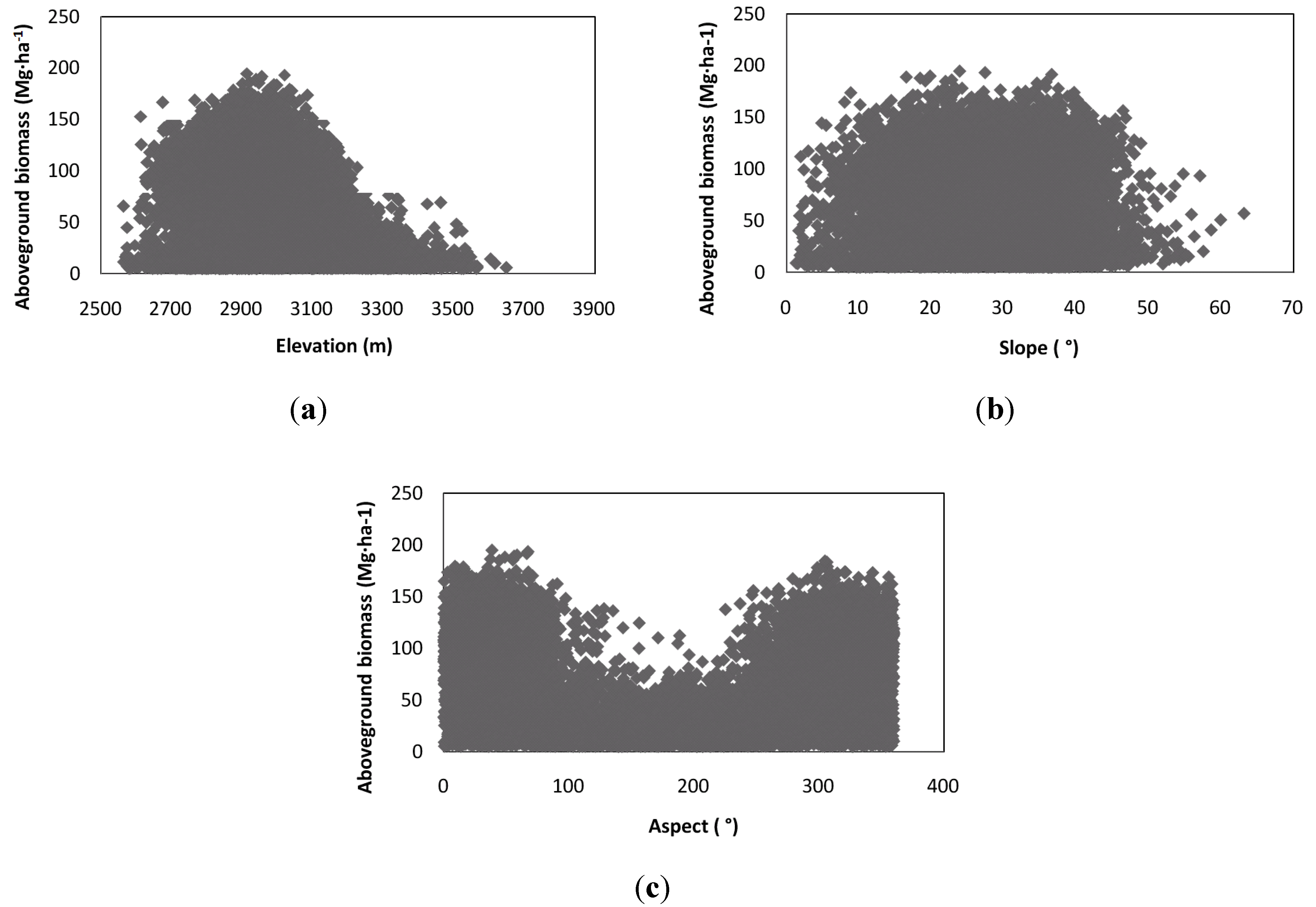
4. Discussion and Conclusions
Acknowledgments
Conflicts of Interest
References
- Zhao, F.; Guo, Q.; Kelly, M. Allometric equation choice impacts Lidar-based forest biomass estimates: A case study from the Sierra National Forest, CA. Agric. For. Meteorol. 2012, 165, 64–72. [Google Scholar] [CrossRef]
- Salas, C.; Ene, L.; Gregoire, T.G.; Naesset, E.; Gobakken, T. Modelling tree diameter from airborne laser scanning derived variables: A comparison of spatial statistical models. Remote Sens. Environ. 2010, 114, 1277–1285. [Google Scholar] [CrossRef]
- Ahmed, R.; Siqueira, P.; Hensley, S. A study of forest biomass estimates from Lidar in the northern temperate forests of New England. Remote Sens. Environ. 2013, 130, 121–135. [Google Scholar] [CrossRef]
- Allouis, T.; Durrieu, S.; Vega, C.; Couteron, P. Stem volume and above-ground biomass estimation of individual pine trees from LiDAR data: Contribution of full-waveform signals. IEEE J. Sel. Top. Appl. Earth Obs. Remote Sens. 2013, 6, 924–934. [Google Scholar] [CrossRef]
- Ene, L.T.; Naesset, E.; Gobakken, T.; Gregoire, T.G.; Stahl, G.; Holm, S. A simulation approach for accuracy assessment of two-phase post-stratified estimation in large-area LiDAR biomass surveys. Remote Sens. Environ. 2013, 133, 210–224. [Google Scholar] [CrossRef]
- Popescu, S.C.; Zhao, K.; Neuenschwander, A.; Lin, C. Satellite Lidar vs. small footprint airborne lidar: Comparing the accuracy of aboveground biomass estimates and forest structure metrics at footprint level. Remote Sens. Environ. 2011, 115, 2786–2797. [Google Scholar] [CrossRef]
- Kankare, V.; Vastaranta, M.; Holopainen, M.; Raety, M.; Yu, X.; Hyyppa, J.; Hyyppa, H.; Alho, P.; Viitala, R. Retrieval of forest aboveground biomass and stem volume with airborne scanning LiDAR. Remote Sens. 2013, 5, 2257–2274. [Google Scholar] [CrossRef] [Green Version]
- Wulder, M.A.; White, J.C.; Nelson, R.F.; Naesset, E.; Orka, H.O.; Coops, N.C.; Hilker, T.; Bater, C.W.; Gobakken, T. LiDAR sampling for large-area forest characterization: A review. Remote Sens. Environ. 2012, 121, 196–209. [Google Scholar] [CrossRef]
- Kim, Y.; Chang, A.; Kim, Y.; Song, J.; Kim, C. Estimation of forest biomass from airborne LiDAR data as measures against Global Warming-Individual Tree Unit and Forest Stand Unit. Disaster Adv. 2012, 5, 295–299. [Google Scholar]
- D’Oliveira, M.V.N.; Reutebuch, S.E.; McGaughey, R.J.; Andersen, H.-E. Estimating forest biomass and identifying low-intensity logging areas using airborne scanning Lidar in Antimary State Forest, Acre State, Western Brazilian Amazon. Remote Sens. Environ. 2012, 124, 479–491. [Google Scholar] [CrossRef]
- Gonzalez, P.; Asner, G.P.; Battles, J.J.; Lefsky, M.A.; Waring, K.M.; Palace, M. Forest carbon densities and uncertainties from LiDAR, QuickBird, and field measurements in California. Remote Sens. Environ. 2010, 114, 1561–1575. [Google Scholar] [CrossRef]
- Sexton, J.O.; Bax, T.; Siqueira, P.; Swenson, J.J.; Hensley, S. A comparison of Lidar, radar, and field measurements of canopy height in pine and hardwood forests of southeastern North America. For. Ecol. Manag. 2009, 257, 1136–1147. [Google Scholar] [CrossRef]
- Nelson, R.F.; Hyde, P.; Johnson, P.; Emessiene, B.; Imhoff, M.L.; Campbell, R.; Edwards, W. Investigating RaDAR-LiDAR synergy in a North Carolina pine forest. Remote Sens. Environ. 2007, 110, 98–108. [Google Scholar] [CrossRef]
- Hajnsek, I.; Kugler, F.; Lee, S.-K.; Papathanassiou, K.P. Tropical-forest-parameter estimation by means of Pol-InSAR: The INDREX-II campaign. IEEE Trans. Geosci. Remote Sens. 2009, 47, 481–493. [Google Scholar] [CrossRef]
- Asner, G.P.; Mascaro, J.; Muller-Landau, H.C.; Vieilledent, G.; Vaudry, R.; Rasamoelina, M.; Hall, J.S.; van Breugel, M. A universal airborne LiDAR approach for tropical forest carbon mapping. Oecologia 2012, 168, 1147–1160. [Google Scholar] [CrossRef]
- He, Q.-S.; Cao, C.-X.; Chen, E.-X.; Sun, G.-Q.; Ling, F.-L.; Pang, Y.; Zhang, H.; Ni, W.-J.; Xu, M.; Li, Z.-Y.; et al. Forest stand biomass estimation using ALOS PALSAR data based on LiDAR-derived prior knowledge in the Qilian Mountain, western China. Int. J. Remote Sens. 2012, 33, 710–729. [Google Scholar] [CrossRef]
- Santos, J.R.; Freitas, C.C.; Araujo, L.S.; Dutra, L.V.; Mura, J.C.; Gama, F.F.; Soler, L.S.; Sant’Anna, S.J.S. Airborne P-band SAR applied to the aboveground biomass studies in the Brazilian tropical rainforest. Remote Sens. Environ. 2003, 87, 482–493. [Google Scholar] [CrossRef]
- Lambert, M.C.; Ung, C.H.; Raulier, F. Canadian national tree aboveground biomass equations. Can. J. For. Res. 2005, 35, 1996–2018. [Google Scholar] [CrossRef]
- Saatchi, S.; Halligan, K.; Despain, D.G.; Crabtree, R.L. Estimation of forest fuel load from radar remote sensing. IEEE Trans. Geosci. Remote Sens. 2007, 45, 1726–1740. [Google Scholar] [CrossRef]
- Tsui, O.W.; Coops, N.C.; Wulder, M.A.; Marshall, P.L.; McCardle, A. Using multi-frequency radar and discrete-return LiDAR measurements to estimate above-ground biomass and biomass components in a coastal temperate forest. ISPRS J. Photogramm. Remote Sens. 2012, 69, 121–133. [Google Scholar] [CrossRef]
- Tian, X.; Su, Z.; Chen, E.; Li, Z.; van der Tol, C.; Guo, J.; He, Q. Estimation of forest above-ground biomass using multi-parameter remote sensing data over a cold and arid area. Int. J.Appl. Earth Obs. Geoinf. 2012, 14, 160–168. [Google Scholar] [CrossRef]
- He, Q.; Xu, H.; Zhang, Y. Estimation of Forest Biophysical Parameters Using Small-Footprint LiDAR with low Density in a Coniferous Forest. In Proceedings of International Symposium on Lidar and Radar Mapping 2011: Technologies and Applications, Nanjing, China, 26–29 May 2011; Volume 8286.
- Zhou, Y.; Zhu, Q.; Chen, J.M.; Wang, Y.Q.; Liu, J.; Sun, R.; Tang, S. Observation and simulation of net primary productivity in Qilian Mountain, western China. J. Environ. Manag. 2007, 85, 574–584. [Google Scholar] [CrossRef]
- Wang, J.Y.; Ju, K.J.; Fu, H.E.; Chang, X.X.; He, H.Y. Study on biomass of water conservation forest on North Slope of Qilian Mountains. J. Fujian Coll. For. 1998, 18, 319–323. [Google Scholar]
- Chan, N.; Takeda, S.; Suzuki, R.; Yamamoto, S. Establishment of allometric models and estimation of biomass recovery of swidden cultivation fallows in mixed deciduous forests of the Bago Mountains, Myanmar. For. Ecol. Manag. 2013, 304, 427–436. [Google Scholar] [CrossRef]
- Kenzo, T.; Furutani, R.; Hattori, D.; Kendawang, J.J.; Tanaka, S.; Sakurai, K.; Ninomiya, I. Allometric equations for accurate estimation of above-ground biomass in logged-over tropical rainforests in Sarawak, Malaysia. J. For. Res. 2009, 14, 365–372. [Google Scholar] [CrossRef]
- Kenzo, T.; Ichie, T.; Hattori, D.; Itioka, T.; Handa, C.; Ohkubo, T.; Kendawang, J.J.; Nakamura, M.; Sakaguchi, M.; Takahashi, N.; et al. Development of allometric relationships for accurate estimation of above- and below-ground biomass in tropical secondary forests in Sarawak, Malaysia. J. Trop. Ecol. 2009, 25, 371–386. [Google Scholar] [CrossRef]
- Axelsson, P.E. Processing of laser scanner data—Algorithms and applications. ISPRS J. Photogramm. Remote Sens. 1999, 54, 138–147. [Google Scholar] [CrossRef]
- Kraus, K.; Pfeifer, N. Determination of terrain models in wooded areas with airborne laser scanner data. ISPRS J. Photogramm. Remote Sens. 1998, 53, 193–203. [Google Scholar] [CrossRef]
- Means, J.E.; Acker, S.A.; Fitt, B.J.; Renslow, M.; Emerson, L.; Hendrix, C.J. Predicting forest stand characteristics with airborne scanning Lidar. Photogramm. Eng. Remote Sens. 2000, 66, 1367–1371. [Google Scholar]
- Treitz, P.; Lim, K.; Woods, M.; Pitt, D.; Nesbitt, D.; Etheridge, D. LiDAR sampling density for forest resource inventories in Ontario, Canada. Remote Sens. 2012, 4, 830–848. [Google Scholar] [CrossRef]
- Naesset, E. Practical large-scale forest stand inventory using a small-footprint airborne scanning laser. Scand. J. For. Res. 2004, 19, 164–179. [Google Scholar] [CrossRef]
- Cao, C.; Bao, Y.; Xu, M.; Chen, W.; Zhang, H.; He, Q.; Li, Z.; Guo, H.; Li, J.; Li, X.; et al. Retrieval of forest canopy attributes based on a geometric-optical model using airborne LiDAR and optical remote-sensing data. Int. J. Remote Sens. 2012, 33, 692–709. [Google Scholar] [CrossRef]
- Li, Y.; Andersen, H.-E.; McGaughey, R. A Comparison of statistical methods for estimating forest biomass from light detection and ranging data. West. J. Appl. For. 2008, 23, 223–231. [Google Scholar]
- Vincent, G.; Sabatier, D.; Blanc, L.; Chave, J.; Weissenbacher, E.; Pelissier, R.; Fonty, E.; Molino, J.F.; Couteron, P. Accuracy of small footprint airborne LiDAR in its predictions of tropical moist forest stand structure. Remote Sens. Environ. 2012, 125, 23–33. [Google Scholar] [CrossRef]
- O’Brien, R.M. A caution regarding rules of thumb for variance inflation factors. Qual. Quant. 2007, 41, 673–690. [Google Scholar] [CrossRef]
- Popescu, S.C.; Wynne, R.H.; Nelson, R.F. Measuring individual tree crown diameter with Lidar and assessing its influence on estimating forest volume and biomass. Can. J. Remote Sens. 2003, 29, 564–577. [Google Scholar] [CrossRef]
- Lim, K.; Treitz, P.; Baldwin, K.; Morrison, I.; Green, J. Lidar remote sensing of biophysical properties of tolerant northern hardwood forests. Can. J. Remote Sens. 2003, 29, 658–678. [Google Scholar] [CrossRef]
© 2013 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
He, Q.; Chen, E.; An, R.; Li, Y. Above-Ground Biomass and Biomass Components Estimation Using LiDAR Data in a Coniferous Forest. Forests 2013, 4, 984-1002. https://doi.org/10.3390/f4040984
He Q, Chen E, An R, Li Y. Above-Ground Biomass and Biomass Components Estimation Using LiDAR Data in a Coniferous Forest. Forests. 2013; 4(4):984-1002. https://doi.org/10.3390/f4040984
Chicago/Turabian StyleHe, Qisheng, Erxue Chen, Ru An, and Yong Li. 2013. "Above-Ground Biomass and Biomass Components Estimation Using LiDAR Data in a Coniferous Forest" Forests 4, no. 4: 984-1002. https://doi.org/10.3390/f4040984
APA StyleHe, Q., Chen, E., An, R., & Li, Y. (2013). Above-Ground Biomass and Biomass Components Estimation Using LiDAR Data in a Coniferous Forest. Forests, 4(4), 984-1002. https://doi.org/10.3390/f4040984




