2.4.1. Optical Image Features
- (1)
The Surface Reflectance
The raw bands of optical remote sensing imagery directly capture the spectral characteristics of terrestrial objects, containing key information reflecting land cover and vegetation conditions. Different spectral bands correspond to distinct reflectance properties of land features, which form the basis for vegetation discrimination and parameter retrieval. In the visible spectrum, the blue band exhibits strong water penetration, effectively capturing water turbidity information; the green band is sensitive to chlorophyll content in vegetation, showing higher reflectance; the red band is closely associated with photosynthesis, where vegetation absorbs significant light energy, resulting in relatively lower reflectance. The near-infrared band offers unique advantages for vegetation detection, as healthy vegetation displays extremely high reflectance due to multiple scattering and reflection within the canopy structure.
To fully leverage the spectral potential of the GF-6 satellite, this study extracted reflectance values for all eight bands from preprocessed imagery, including four standard multispectral bands: the blue band (0.45–0.52 μm), the green band (0.52–0.59 μm), the red band (0.63–0.69 μm), and the near-infrared band (0.77–0.89 μm); and four GF-6-specific bands: the red edge 1 band (0.69–0.73 μm), the red edge 2 band (0.73–0.77 μm), the violet band (0.40–0.45 μm), and the yellow band (0.59–0.63 μm). These new bands (particularly the two red edge bands) exhibit high sensitivity to vegetation physiological parameters (e.g., chlorophyll content, nitrogen status) and phenological changes, significantly enhancing vegetation identification and parameter retrieval capabilities. For each sample plot, the final reflectance was calculated as the average value of all pixels falling within the 10 m × 10 m plot grid to mitigate within-pixel heterogeneity and improve result stability. All band information extraction was performed using the ENVI software platform. The resulting spectral data were serve as core input variables for subsequent modeling and analysis.
- (2)
Vegetation Index
Using a single band or multiple single-band data often fails to fully capture the complex spectral characteristics and variation patterns of vegetation in remote sensing imagery. To more effectively utilize remote sensing satellite data for characterizing vegetation status, this study constructed and extracted nine vegetation indices based on the spectral characteristics of typical vegetation. These indices, calculated through linear or nonlinear combinations of spectral bands (e.g., addition, subtraction, multiplication, division), enhance vegetation signals and minimize confounding factors such as soil background and atmospheric effects. These indices include NDVI, RVI, DVI, SAVI, EVI, NDREI, REDNDVI, NDVIR1, and NDVIR2 [
35,
36], with their calculation formulas shown in
Table 2.
The specific indices employed are as follows:
Normalized Difference Vegetation Index (NDVI): A standard index for vegetation greenness and coverage, closely related to vegetation cover fraction. NDVI values range from −1 to 1, with higher values indicating denser vegetation.
Ratio Vegetation Index (RVI): Sensitive to dense vegetation, this index enhances vegetation information by leveraging the strong reflectance in the near-infrared band and strong absorption in the red band.
Difference Vegetation Index (DVI): Calculated as the difference between two spectral bands, this index is sensitive to soil variations and effectively distinguishes water bodies and vegetation.
Soil-Adjusted Vegetation Index (SAVI): Designed to minimize soil brightness effects in high-density vegetation areas, SAVI reduces saturation issues common with NDVI in dense canopies.
Enhanced Vegetation Index (EVI): Optimized for high-biomass regions by incorporating the blue band and adjustment coefficients, EVI more accurately reflects vegetation status under complex conditions such as atmospheric interference or heterogeneous soil backgrounds.
Normalized Difference Red Edge Index (NDREI): This index replaces the near-infrared and red bands in NDVI with the red edge band’s peak and valley, making it sensitive to chlorophyll content.
Red Edge Normalized Difference Vegetation Index (REDNDVI): Combining the near-infrared and the first red-edge band (R1), this index is associated with vegetation chlorophyll content.
NDVIR1 and NDVIR2: These indices replace the near-infrared band in NDVI with red edge 1 (R1) and red edge 2 (R2), respectively. They are sensitive to subtle changes in the canopy layer and senescence, making them suitable for forest monitoring and precision agriculture applications.
By amplifying the reflectance differences between vegetation and other land cover types such as soil and water bodies in the red, near-infrared, and red edge bands, these vegetation indices can more sensitively reflect vegetation cover, growth status, and physiological parameters (e.g., chlorophyll content, biomass). This provides important remote sensing feature inputs for forest structure parameter extraction and biomass estimation.
- (3)
Texture Features
Texture features describe the spatial distribution patterns of grayscale or color in image pixels, serving as crucial visual indicators for characterizing the surface structure of land objects [
37]. In remote sensing image analysis, texture features provide essential spatial contextual information for land object interpretation, classification, and identification. Particularly in areas with complex forest stand structures, incorporating texture features enhances the accuracy and reliability of biomass estimation [
38]. Numerous studies have demonstrated that integrating spectral and texture features significantly improves the estimation accuracy of forest parameter inversion models [
39,
40].
Among various texture analysis methods, the Gray-Level Co-occurrence Matrix (GLCM) is widely used due to its robustness and adaptability. Originally proposed by Haralick in 1973, GLCM captures the correlation between grayscale values of pixel pairs at a specified distance and direction, reflecting comprehensive information on image orientation, variation amplitude, and spacing. Common directions include 0°, 45°, 90°, and 135°.
In this study, to reduce data dimensionality while preserving critical information, Principal Component Analysis (PCA) was first performed on the eight GF-6 spectral bands. The first principal component, which accounted for 96.5% of the total variance, was selected as the input image for texture analysis. Based on the Gray-level Co-occurrence Matrix (GLCM), eight commonly used texture features were extracted from the first principal component across four window sizes (3 × 3, 5 × 5, 7 × 7, 9 × 9), including: Mean, Entropy, Homogeneity, Variance, Dissimilarity, Angular Second Moment, Contrast, and Correlation.
The choice of window size is critical for capturing texture information at different spatial scales. Smaller windows (e.g., 3 × 3) capture fine-scale structural variations, while larger windows capture broader spatial patterns. Through iterative testing, the 3 × 3 window was found to most effectively capture the fine-scale textural characteristics of the spruce forest. Consequently, this study employed a 3 × 3 moving window with a step size of 1 and an angle of 0° for texture feature extraction.
The eight extracted texture features are defined in
Table 3 as follows:
Mean (ME): The average gray level within the window, reflecting the overall brightness level of the target area.
Entropy (ENT): A measure of randomness indicating texture complexity; higher entropy values suggest more complex canopy structures.
Homogeneity (HOM): Measures the closeness of the distribution of elements in the GLCM to the diagonal; higher values indicate more uniform texture.
Variance (VAR): A measure of gray-level dispersion; greater variance indicates richer, “rougher” texture.
Dissimilarity (DIS): Measures the absolute difference between gray levels; higher values indicate greater texture heterogeneity.
Angular Second Moment (ASM): A measure of textural uniformity (energy); higher values indicate more regular, uniform texture.
Contrast (CON): A measure of local gray-level variation; high contrast indicates significant gray-level differences.
Correlation (COR): A measure of linear dependency between gray levels, indicating the linear relationship between pixel gray values.
2.4.2. LiDAR Image Features
ALT08 product includes geographic coordinates, slope, elevation, and canopy height information for photon points. Parameters potentially influencing spruce forest biomass were screened as feature variables, as shown in
Table 4. A total of 38 feature variables were selected, including forest canopy height and elevation at different percentiles.
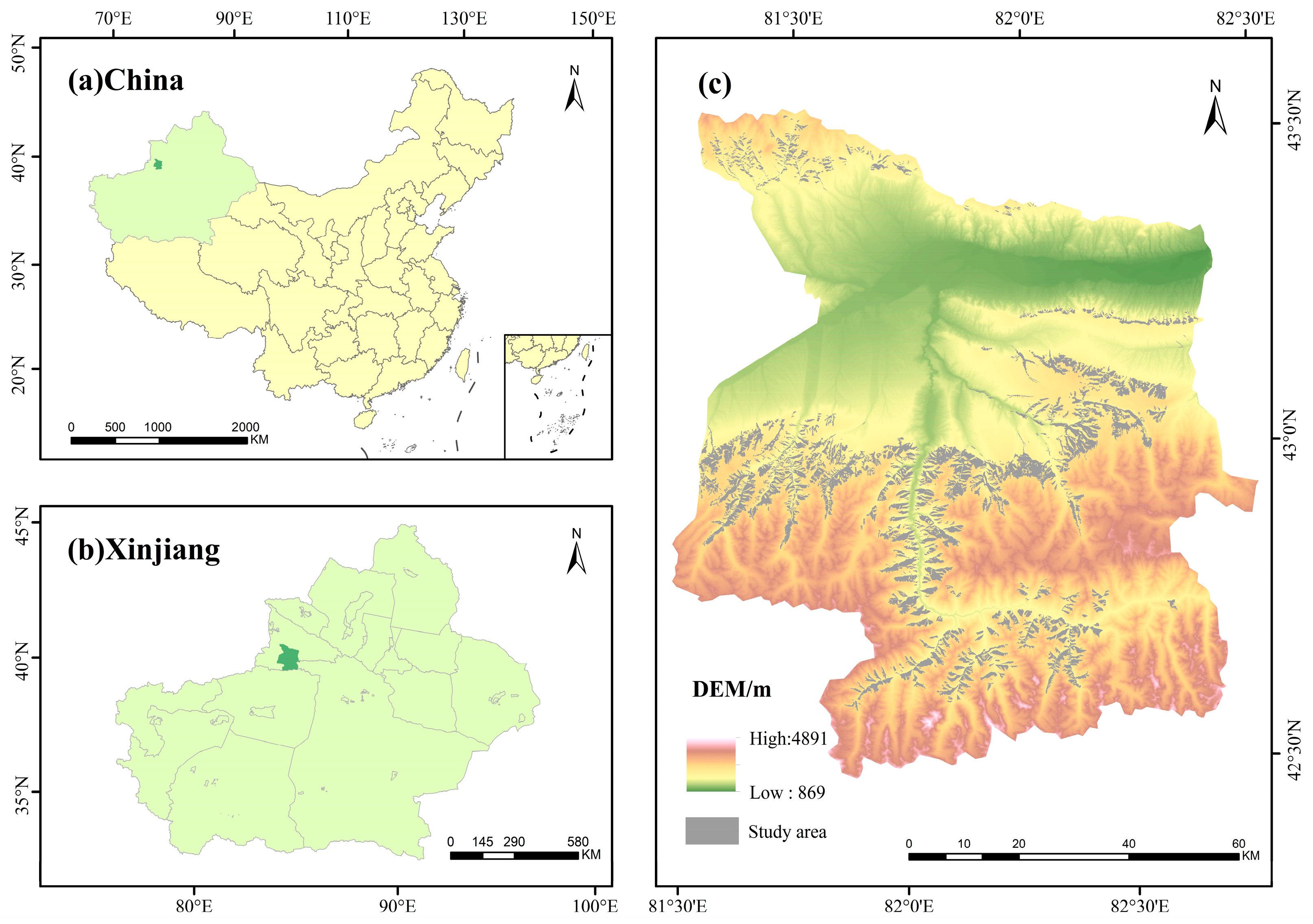
The Advanced Topographic Laser Altimeter System (ATLAS) aboard the ICESat-2 satellite emits six laser beams arranged in three parallel groups along the orbital direction. Each group consists of one high-power laser and one low-power laser. The strong beams (gt3l, gt3r) have a pulse energy of approximately 110 μJ, while the weak beams (gt1l, gt1r, gt2l, gt2r) have an energy of about 28 μJ [
41]. This energy difference directly affects the photon flux density, thereby determining the spatial sampling characteristics of the photon point cloud. In vegetation-covered areas, the strong beam typically penetrates the canopy more effectively to capture understory terrain signals, while photons from the weak beam are largely intercepted by the upper canopy layer. To mitigate the impact of laser intensity differences on spruce forest biomass estimation, this study categorizes photons into strong and weak beam groups based on beam intensity. The sc_orient field in the ATL08 product records the satellite’s attitude information during forward, backward, and transition flight modes. When sc_orien t = 1 (ICESat-2 is considered to be flying forward), strong beams are located ahead of the orbit, and weak beams are located behind. When sc_orient = 0 (ICESat-2 is considered to be flying backward), strong beams are located behind, and weak beams are located ahead.
The ICESat-2 satellite operates in a continuous day-night observation mode, where variations in solar background noise levels impact metrics such as signal-to-noise ratio (SNR). During daytime observations, significant scattering of solar radiation by the atmosphere substantially increases the number of background photons received by the detector. The background noise level of the ATLAS system typically exceeds that during nighttime operations by 1–2 orders of magnitude [
42]. This elevated background noise not only reduces the signal-to-noise ratio (SNR) but also increases the false alarm rate of photon classification algorithms (e.g., DRAGANN), particularly over low-albedo surfaces. Daytime atmospheric turbulence, convective activity, and aerosol concentrations are generally higher than at night, causing additional beam broadening and path perturbations that degrade the spatial distribution accuracy of the photon point cloud. The more stable nighttime atmosphere facilitates sub-meter elevation measurement accuracy [
43]. Daytime surface reflectance exhibits significant angular dependence, particularly for high-albedo surfaces like snow, ice, and deserts. The coupled effect of solar elevation angle and observation geometry substantially alters optical return intensity, introducing additional elevation bias [
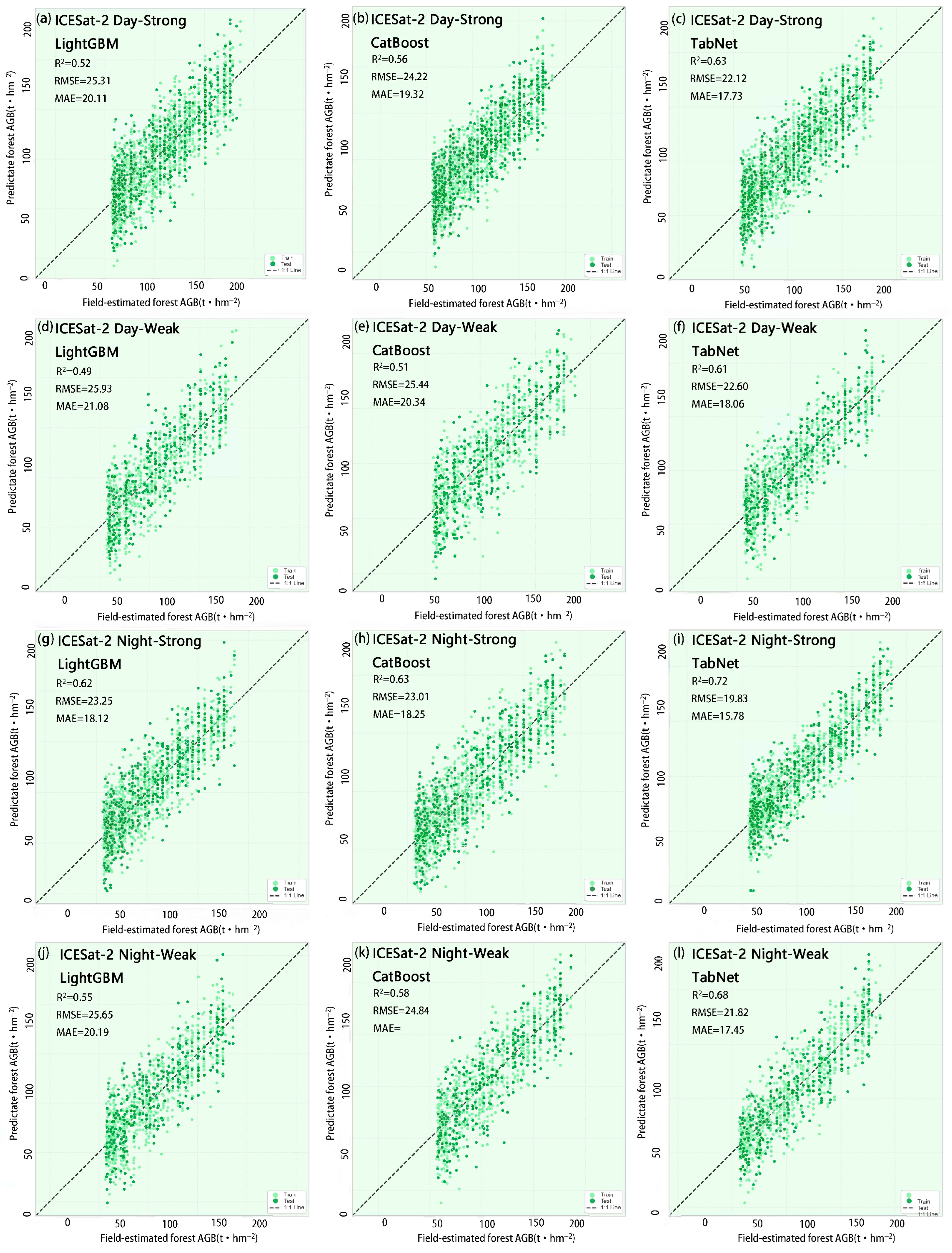
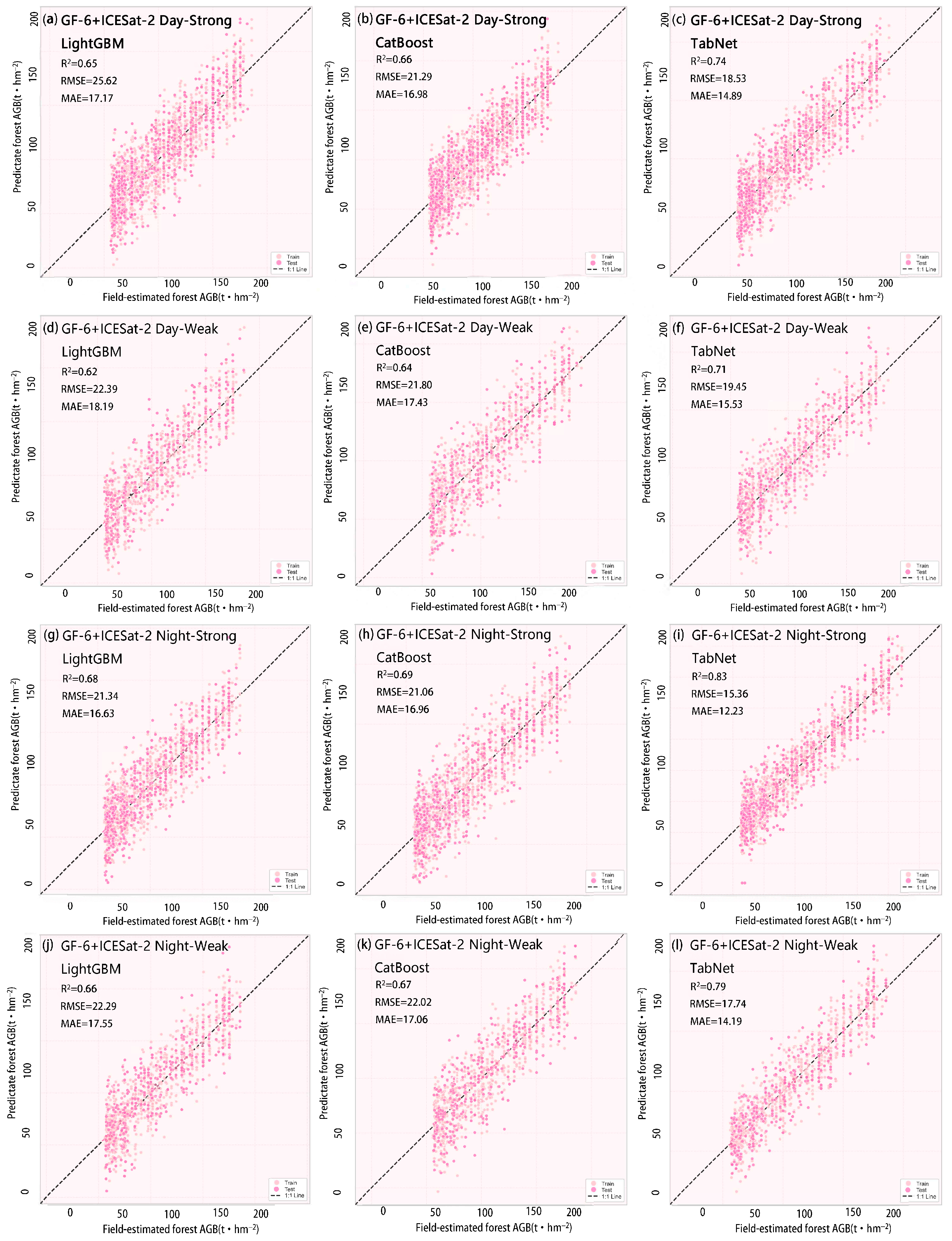
44]. Nighttime observations eliminate solar radiation interference and stabilize surface reflectance characteristics, facilitating consistent analysis of long-term time series. To mitigate the impact of solar radiation and atmospheric conditions on spruce forest biomass inversion, this study further categorizes data into daytime and nighttime segments based on observation timing. The night_flag field in the ATL08 product identifies data acquisition periods: 0 denotes daytime, 1 denotes nighttime. This flag is calculated based on the solar elevation angle within the geolocated segment. If the solar elevation angle exceeds the threshold, the observation is classified as daytime; otherwise, it is classified as nighttime. A total of 5796 valid photon points were extracted within the study area. Based on the two dimensions of observation time and beam intensity, these points were categorized into four observation modes: daytime strong beam, daytime weak beam, nighttime strong beam, and nighttime weak beam. This classification facilitates subsequent analysis of how different modes influence biomass estimation.
2.4.3. Determination of Characteristic Factors
Using Pearson’s correlation coefficient method in SPSS data analysis software (IBM SPSS Statistics v26, IBM Corp., Armonk, NY, USA), we analyzed the correlation between characteristic factors and spruce forest biomass, identifying statistically significant factors correlated with spruce forest biomass [
45]. This approach eliminates redundant variables and optimizes model inputs, enhancing the model’s predictive accuracy and computational efficiency. This study calculated Pearson correlations between measured spruce forest biomass values and the characteristic factors from GF-6 and ICESat-2, as shown in
Table 5 and
Table 6, respectively.
Among the 28 feature factors in the GF-6 dataset, 19 feature factors showed significant correlation at the 0.01 level (two-tailed), while 4 feature factors exhibited significant correlation at the 0.05 level (two-tailed). The feature factor with the highest correlation coefficient against spruce forest biomass in the study area was ME_3, with a correlation coefficient of 0.612. The top five feature factors by absolute correlation coefficient were ME_3, REDNDVI, Band6, NDREI, and Band5, with respective correlations of 0.612, −0.601, −0.583, −0.572, and −0.557. The feature factors with strong correlation coefficients were the mean statistics within the texture features, indicating that the mean statistics of the bands are highly sensitive to biomass.
Among the 38 feature factors in the ICESat-2 data, 25 feature factors showed significant correlations at the 0.01 level (two-tailed), while 4 feature factors showed significant correlations at the 0.05 level (two-tailed). Among these, the highest correlation coefficient was observed for h_canopy, reaching 0.831. The top five feature factors by absolute correlation coefficient were h_canopy, rh90, rh95, rh85, and rh80, with respective correlations of 0.831, 0.824, 0.817, 0.812, and 0.796—all significant at the 0.01 level. The results indicate that canopy height from ICESat-2 data exhibits strong sensitivity to spruce forest biomass.
Given that remote sensing features may exhibit complex nonlinear relationships with forest biomass beyond linear correlations, a secondary validation was conducted during model construction using model-based feature importance evaluation mechanisms. Specifically, during the training of LightGBM, CatBoost, and TabNet models, the built-in importance metrics—including gain and split count for LightGBM and CatBoost, and attention masks with feature contribution weights for TabNet—were employed to rank and refine the preliminarily selected features. The final feature set for modeling was determined by synthesizing the results from both correlation analysis and model-based importance evaluation.
Through the above correlation analysis, feature factors with the most significant correlations to measured spruce forest biomass values were identified from the two types of remote sensing data and retained for dataset construction. Following screening, GF-6 data retained 23 feature factors, while ICESat-2 data retained 29 feature factors.