Genome-Wide Identification and Expression Analysis of the NF-Y Transcription Factor Family in Prunus armeniaca
Abstract
1. Introduction
2. Materials and Methods
2.1. Identification of the NF-Y Gene Family in P. armeniaca
2.2. Phylogenetic, Gene Structure, and Conserved Motif Analyses
2.3. Chromosomal Distribution and Synteny Analysis
2.4. Identification of Cis-Acting Elements
2.5. Expression Analysis Using RNA-Seq Data and Verification via RT-qPCR
2.6. Oil Content Detection
3. Results
3.1. Identification of NF-Y Family Members in P. armeniaca
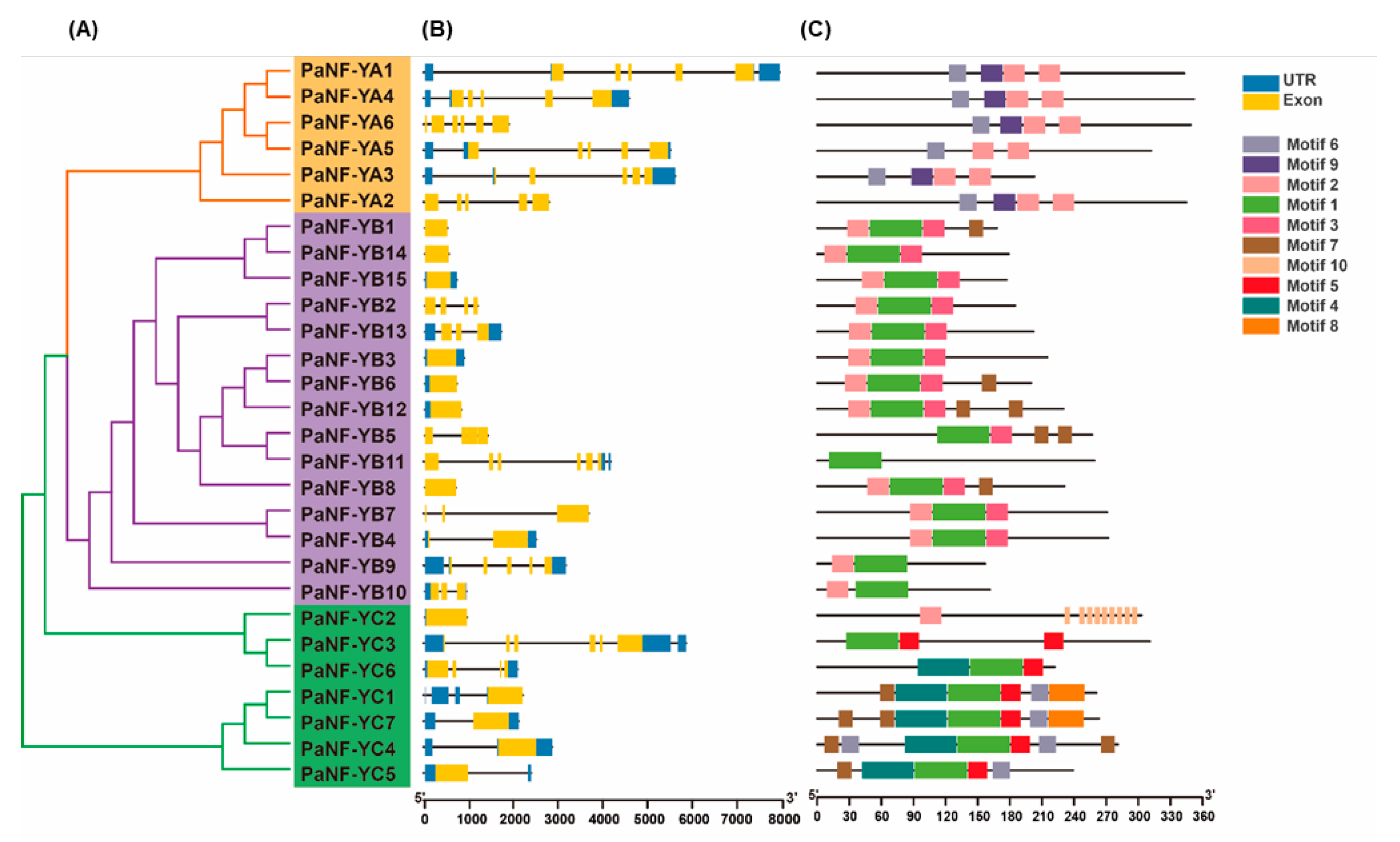
3.2. Phylogenetic Analysis of the PNF-Y Family
3.3. Gene Structure and Conserved Motif of the PNF-Y Family
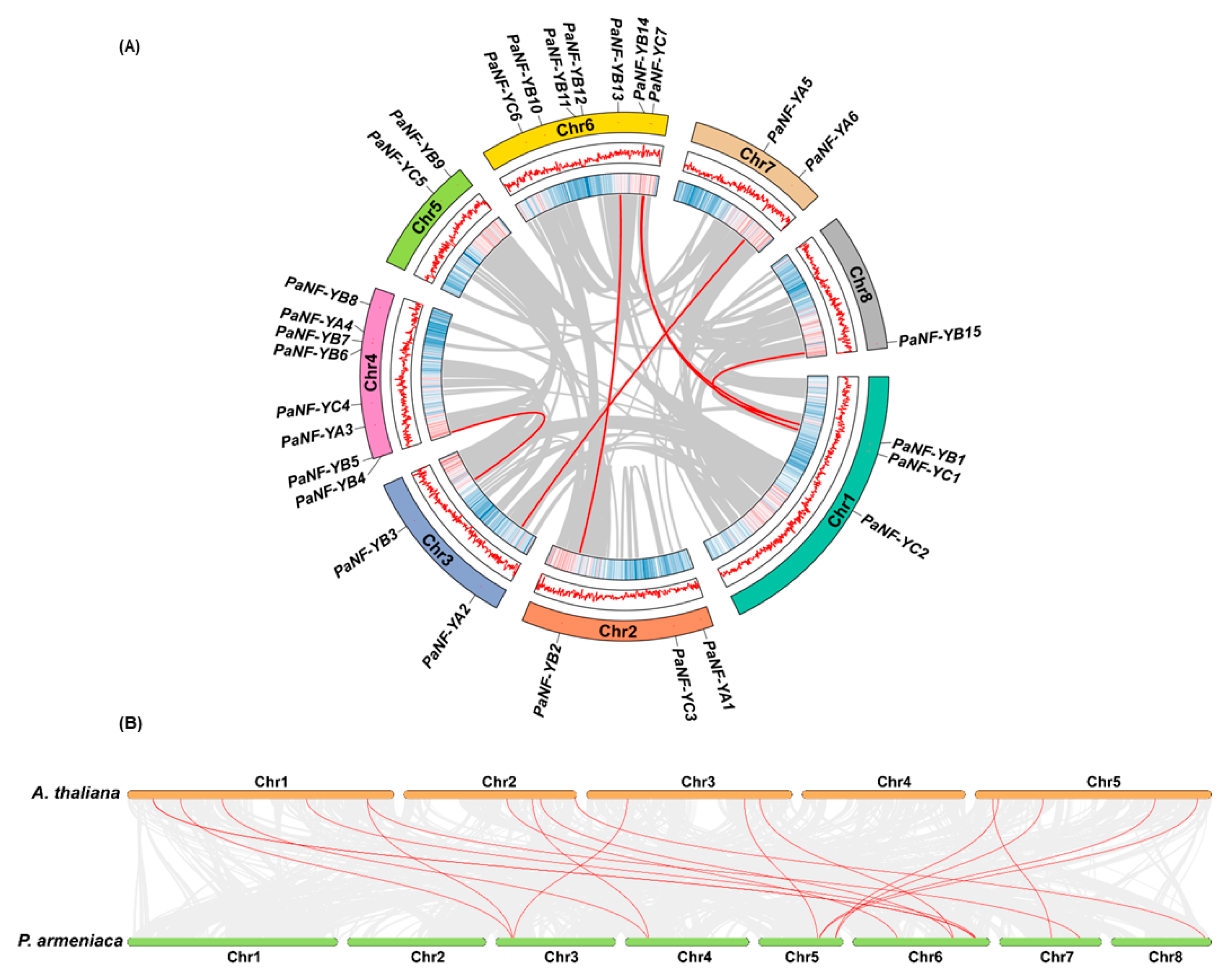
3.4. Chromosomal Distribution and Synteny Analysis of the PaNF-Y Family
3.5. Cis-Acting Elements Analysis of the PaNF-Y Family
3.6. Tissue-Specific Expression Patterns of the PaNF-Y Family
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Romier, C.; Cocchiarella, F.; Mantovani, R.; Moras, D. The NF-YB/NF-YC structure gives insight into DNA binding and transcription regulation by CCAAT factor NF-Y. J. Biol. Chem. 2003, 278, 1336–1345. [Google Scholar] [CrossRef] [PubMed]
- Mantovani, R. The molecular biology of the CCAAT-binding factor NF-Y. Gene 1999, 239, 15–27. [Google Scholar] [CrossRef] [PubMed]
- Calvenzani, V.; Testoni, B.; Gusmaroli, G.; Lorenzo, M.; Gnesutta, N.; Petroni, K.; Mantovani, R.; Tonelli, C. Interactions and CCAAT-binding of Arabidopsis thaliana NF-Y subunits. PLoS ONE 2012, 7, e42902. [Google Scholar] [CrossRef] [PubMed]
- Kahle, J.; Baake, M.; Doenecke, D.; Albig, W. Subunits of the heterotrimeric transcription factor NF-Y are imported into the nucleus by distinct pathways involving importin beta and importin 13. Mol. Cell Biol. 2005, 25, 5339–5354. [Google Scholar] [CrossRef] [PubMed]
- Oldfield, A.J.; Yang, P.; Conway, A.E.; Cinghu, S.; Freudenberg, J.M.; Yellaboina, S.; Jothi, R. Histone-fold domain protein NF-Y promotes chromatin accessibility for cell type-specific master transcription factors. Mol. Cell 2014, 55, 708–722. [Google Scholar] [CrossRef]
- Hackenberg, D.; Wu, Y.; Voigt, A.; Adams, R.; Schramm, P.; Grimm, B. Studies on differential nuclear translocation mechanism and assembly of the three subunits of the Arabidopsis thaliana transcription factor NF-Y. Mol. Plant. 2012, 5, 876–888. [Google Scholar] [CrossRef]
- Riechmann, J.L.; Heard, J.; Martin, G.; Reuber, L.; Jiang, C.; Keddie, J.; Adam, L.O.; Pineda, O.; Ratcliffe, O.; Samaha, R.; et al. Arabidopsis transcription factors: Genome-wide comparative analysis among eukaryotes. Science 2000, 290, 2105–2110. [Google Scholar] [CrossRef]
- Siefers, N.; Dang, K.K.; Kumimoto, R.W.; Bynum, W.E., IV; Tayrose, G.; Hol, B.F., III. Tissue-specific expression patterns of Arabidopsis NF-Y transcription factors suggest potential for extensive combinatorial complexity. Plant Physiol. 2009, 149, 625–641. [Google Scholar] [CrossRef]
- Stephenson, T.; McIntyre, C.; Collet, C.; Xue, G. Genome-wide identification and expression analysis of the NF-Y family of transcription factors in Triticum aestivum. Plant Mol. Biol. 2007, 65, 77–92. [Google Scholar] [CrossRef]
- Wang, J.; Li, G.; Li, C.; Zhang, C.; Cui, L.; Ai, G.; Wang, X.; Zheng, F.; Zhang, D.; Larkin, R.M. NF-Y plays essential roles in flavonoid biosynthesis by modulating histone modifications in tomato. New Phytol. 2021, 229, 3237–3252. [Google Scholar] [CrossRef]
- Wang, Y.; Xu, W.; Chen, Z.; Han, B.; Haque, M.E.; Liu, A. Gene structure, expression pattern and interaction of Nuclear Factor-Y family in castor bean (Ricinus communis). Planta 2018, 247, 559–572. [Google Scholar] [CrossRef] [PubMed]
- Quach, T.N.; Nguyen, H.; Valliyodan, B.; Joshi, T.; Xu, D.; Nguyen, H.T. Genome-wide expression analysis of soybean NF-Y genes reveals potential function in development and drought response. Mol. Genet. Genom. 2015, 290, 1095–1115. [Google Scholar] [CrossRef] [PubMed]
- Li, M.; Li, G.; Liu, W.; Dong, X.; Zhang, A. Genome-wide analysis of the NF-Y gene family in peach (Prunus persica L.). BMC Genom. 2019, 20, 612. [Google Scholar] [CrossRef]
- Li, Y.J.; Fang, Y.; Fu, Y.; Huang, J.; Wu, C.; Zheng, C.C. NFYA1 is involved in regulation of postgermination growth arrest under salt stress in Arabidopsis. PLoS ONE 2013, 8, e61289. [Google Scholar] [CrossRef] [PubMed]
- Lee, D.K.; Kim, H.I.; Jang, G.; Chung, P.J.; Jeong, J.S.; Kim, Y.S.; Bang, S.W.; Jung, H.; Choi, Y.D.; Kim, J.K. The NF-YA transcription factor OsNF-YA7 confers drought stress tolerance of rice in an abscisic acid independent manner. Plant Sci. 2015, 241, 199–210. [Google Scholar] [CrossRef] [PubMed]
- Yu, T.F.; Liu, Y.; Fu, J.D.; Ma, J.; Fang, Z.W.; Chen, J.; Zheng, L.; Lu, Z.W.; Zhou, Y.B.; Chen, M.; et al. The NF-Y-PYR module integrates the abscisic acid signal pathway to regulate plant stress tolerance. Plant Biotechnol. J. 2021, 19, 2589–2605. [Google Scholar] [CrossRef]
- Kumimoto, R.W.; Adam, L.; Hymus, G.; Repetti, P.; Reuber, T.; Marion, C.; Hempel, F.; Ratcliffe, O.J. The Nuclear Factor Y subunits NF-YB2 and NF-YB3 play additive roles in the promotion of flowering by inductive long-day photoperiods in Arabidopsis. Planta 2008, 228, 709–723. [Google Scholar] [CrossRef]
- Wenkel, S.; Turck, F.; Singer, K.; Gissot, L.; Le, G.; Samach, A.; Coupland, G. CONSTANS and the CCAAT box binding complex share a functionally important domain and interact to regulate flowering of Arabidopsis. Plant Cell 2006, 18, 2971–2984. [Google Scholar] [CrossRef]
- Nelson, D.; Repetti, P.; Adams, T.; Creelman, R.; Wu, J.; Warner, D.; Anstrom, D.; Bensen, R.; Castiglioni, P.; Donnarummo, M.; et al. Plant nuclear factor Y (NF-Y) B subunits confer drought tolerance and lead to improved corn yields on water-limited acres. Proc. Natl. Acad. Sci. USA 2007, 104, 16450–16455. [Google Scholar] [CrossRef]
- Yang, Y.; Kong, Q.; Lim, A.R.Q.; Lu, S.; Zhao, H.; Guo, L.; Yuan, L.; Ma, W. Transcriptional regulation of oil biosynthesis in seed plants: Current understanding, applications, and perspectives. Plant Commun. 2022, 3, 100328. [Google Scholar] [CrossRef]
- Mu, J.; Tan, H.; Zheng, Q.; Fu, F.; Liang, Y.; Zhang, J.; Yang, X.; Wang, T.; Chong, K.; Wang, X.; et al. LEAFY COTYLEDON1 is a key regulator of fatty acid biosynthesis in Arabidopsis. Plant Physiol. 2008, 148, 1042–1054. [Google Scholar] [CrossRef] [PubMed]
- Tan, H.; Yang, X.; Zhang, F.; Zheng, X.; Qu, C.; Mu, J.; Fu, F.; Li, J.; Guan, R.; Zhang, H.; et al. Enhanced seed oil production in canola by conditional expression of Brassica napus LEAFY COTYLEDON1 and LEC1-LIKE in developing seeds. Plant Physiol. 2011, 156, 1577–1588. [Google Scholar] [CrossRef] [PubMed]
- Lee, H.; Ahn, J.; Kwon, A.; Lee, E.; Kwak, J.; Min, Y. Chemical composition and antimicrobial activity of the essential oil of apricot seed. Phytother. Res. 2014, 28, 1867–1872. [Google Scholar] [CrossRef] [PubMed]
- Kiralan, M.; Özkan, G.; Kucukoner, E.; Ozcelik, M.M. Apricot (Prunus armeniaca L.) Oil. In Fruit Oils: Chemistry and Functionality; Ramadan, M.F., Ed.; Springer: Cham, Switzerland, 2019; Chapter 25, pp. 505–519. [Google Scholar]
- Zhou, B.; Wang, Y.; Kang, J.; Zhong, H.; Prenzler, P.D. The quality and volatile-profile changes of Longwangmo apricot (Prunus armeniaca L.) kernel oil prepared by different oil-producing processes. Eur. J. Lipid Sci. Technol. 2016, 118, 236–243. [Google Scholar] [CrossRef]
- Sharma, R.; Gupta, A.; Abrol, G.S.; Joshi, V.K. Value addition of wild apricot fruits grown in North-West Himalayan regions-a review. J. Food Sci. Technol. 2014, 51, 2917–2924. [Google Scholar] [CrossRef]
- Siddiqui, S.A.; Anwar, S.; Yunusa, B.M.; Nayik, G.A.; Khaneghah, A.M. The potential of apricot seed and oil as functional food: Composition, biological properties, health benefits & safety. Food Biosci. 2023, 51, 102336. [Google Scholar]
- Chen, C.; Chen, H.; Zhang, Y.; Thomas, H.R.; Frank, M.H.; He, Y.; Xia, R. TBtools: An integrative toolkit developed for interactive analyses of big biological data. Mol. Plant. 2020, 13, 1194–1202. [Google Scholar] [CrossRef]
- Wang, T.; Yang, Y.; Lou, S.; Wei, W.; Zhao, Z.; Ren, Y.; Lin, C.; Ma, L. Genome-wide characterization and gene expression analyses of GATA transcription factors in moso bamboo (Phyllostachys edulis). Int. J. Mol. Sci. 2019, 21, 14. [Google Scholar] [CrossRef]
- Wang, Y.; Tang, H.; Debarry, J.; Tan, X.; Li, J.; Wang, X.; Lee, T.; Jin, H.; Marler, B.; Guo, H.; et al. MCScanX: A toolkit for detection and evolutionary analysis of gene synteny and collinearity. Nucleic Acids Res. 2012, 40, e49. [Google Scholar] [CrossRef]
- Frazee, A.C.; Pertea, G.; Jaffe, A.A.; Langmead, B.; Salzberg, S.A.O.; Leek, J.T. Ballgown bridges the gap between transcriptome assembly and expression analysis. Nat. Biotechnol. 2015, 33, 243–246. [Google Scholar] [CrossRef]
- Liu, H.; Lu, Y.; Wang, J.; Hu, J.; Wuyun, T. Genome-wide screening of long non-coding RNAs involved in rubber biosynthesis in Eucommia ulmoides. J. Integr. Plant Biol. 2018, 60, 1070–1082. [Google Scholar] [CrossRef]
- Schmittgen, T.D.; Livak, K.J. Analyzing real-time PCR data by the comparative CT method. Nat. Protoc. 2008, 3, 1101–1108. [Google Scholar] [CrossRef]
- Deng, P.; Cui, B.; Zhu, H.; Phommakoun, B.; Zhang, D.; Li, Y.; Zhao, F.; Zhao, Z. Accumulation pattern of amygdalin and prunasin and its correlation with fruit and kernel agronomic characteristics during apricot (Prunus armeniaca L.) kernel development. Foods 2021, 10, 397. [Google Scholar] [CrossRef]
- Zhao, H.; Wu, D.; Kong, F.; Lin, K.; Zhang, H.; Li, G. The Arabidopsis thaliana nuclear factor Y transcription factors. Front. Plant Sci. 2017, 7, 2045. [Google Scholar] [CrossRef]
- Petroni, K.; Kumimoto, R.W.; Gnesutta, N.; Calvenzani, V.; Fornari, M.; Tonelli, C.; Holt III, B.F.; Mantovani, R. The promiscuous life of plant NUCLEAR FACTOR Y transcription factors. Plant Cell 2012, 24, 4777–4792. [Google Scholar] [CrossRef]
- Myers, Z.A.; Holt III, B.F. NUCLEAR FACTOR-Y: Still complex after all these years? Curr. Opin. Plant Biol. 2018, 45, 96–102. [Google Scholar] [CrossRef]
- Yang, W.; Lu, Z.; Xiong, Y.; Yao, J. Genome-wide identification and co-expression network analysis of the OsNF-Y gene family in rice. Crop J. 2017, 5, 21–31. [Google Scholar] [CrossRef]
- Liu, R.; Wu, M.; Liu, H.L.; Gao, Y.M.; Chen, J.; Yan, H.W.; Xiang, Y. Genome-wide identification and expression analysis of the NF-Y transcription factor family in Populus. Physiol. Plant. 2021, 171, 309–327. [Google Scholar] [CrossRef]
- Zhou, J.; Yang, L.; Chen, X.; Zhou, M.; Shi, W.; Deng, S.; Luo, Z. Genome-wide identification and characterization of the NF-YA gene family and its expression in response to different nitrogen forms in Populus × canescens. Int. J. Mol. Sci. 2022, 23, 11217. [Google Scholar] [CrossRef]
- Wei, Q.; Wen, S.; Lan, C.; Yu, Y.; Chen, G. Genome-wide identification and expression profile analysis of the NF-Y transcription factor gene family in Petunia hybrida. Plants 2020, 9, 336. [Google Scholar] [CrossRef]
- Gu, Z.; Steinmetz, L.M.; Gu, X.; Scharfe, C.; Davis, R.W.; Li, W.H. Role of duplicate genes in genetic robustness against null mutations. Nature 2003, 421, 63–66. [Google Scholar] [CrossRef]
- Cannon, S.B.; Mitra, A.; Baumgarten, A.; Young, N.D.; May, G. The roles of segmental and tandem gene duplication in the evolution of large gene families in Arabidopsis thaliana. BMC Plant Biol. 2004, 4, 10. [Google Scholar] [CrossRef]
- Paterson, A.; Wang, X.; Tang, H.; Lee, T. Synteny and genomic rearrangements. In Plant Genome Diversity; Wendel, J., Greilhuber, J., Dolezel, J., Leitch, I., Eds.; Springer: Vienna, Austria, 2012; Volume 1, pp. 195–207. [Google Scholar]
- Kim, S.; Kang, J.y.; Cho, D.I.; Park, J.H.; Kim, S.Y. ABF2, an ABRE-binding bZIP factor, is an essential component of glucose signaling and its overexpression affects multiple stress tolerance. Plant J. 2004, 40, 75–87. [Google Scholar] [CrossRef]
- Narusaka, Y.; Nakashima, K.; Shinwari, Z.K.; Sakuma, Y.; Furihata, T.; Abe, H.; Narusaka, M.; Shinozaki, K.; Yamaguchi-Shinozaki, K. Interaction between two cis-acting elements, ABRE and DRE, in ABA-dependent expression of Arabidopsis rd29A gene in response to dehydration and high-salinity stresses. Plant J. 2003, 34, 137–148. [Google Scholar] [CrossRef]
- Abe, H.; Urao, T.; Ito, T.; Seki, M.; Shinozaki, K.; Yamaguchi-Shinozaki, K. Arabidopsis AtMYC2 (bHLH) and AtMYB2 (MYB) function as transcriptional activators in abscisic acid signaling. Plant Cell 2003, 15, 63–78. [Google Scholar] [CrossRef]
- Abe, H.; Yamaguchi-Shinozaki, K.; Urao, T.; Iwasaki, T.; Hosokawa, D.; Shinozaki, K. Role of arabidopsis MYC and MYB homologs in drought-and abscisic acid-regulated gene expression. Plant Cell 1997, 9, 1859–1868. [Google Scholar]
- Scharf, K.D.; Berberich, T.; Ebersberger, I.; Nover, L. The plant heat stress transcription factor (Hsf) family: Structure, function and evolution. Biochim. Et Biophys. Acta 2012, 1819, 104–119. [Google Scholar] [CrossRef]
- Freitas, F.Z.; Virgilio, S.; Cupertino, F.B.; Kowbel, D.J.; Fioramonte, M.; Gozzo, F.C.; Glass, N.L.; Bertolini, M.C. The SEB-1 transcription factor binds to the STRE motif in Neurospora crassa and regulates a variety of cellular processes including the stress response and reserve carbohydrate metabolism. G3 (Bethesda) 2016, 6, 1327–1343. [Google Scholar] [CrossRef]
- Ebeed, H.T. Genome-wide analysis of polyamine biosynthesis genes in wheat reveals gene expression specificity and involvement of STRE and MYB-elements in regulating polyamines under drought. BMC Genom. 2022, 23, 734. [Google Scholar] [CrossRef] [PubMed]
- Li, W.X.; Oono, Y.; Zhu, J.; He, X.J.; Wu, J.M.; Iida, K.; Lu, X.Y.; Cui, X.; Jin, H.; Zhu, J.K. The Arabidopsis NFYA5 transcription factor is regulated transcriptionally and posttranscriptionally to promote drought resistance. Plant Cell 2008, 20, 2238–2251. [Google Scholar] [CrossRef]
- Leyva-González, M.A.; Ibarra-Laclette, E.; Cruz-Ramírez, A.; Herrera-Estrella, L. Functional and transcriptome analysis reveals an acclimatization strategy for abiotic stress tolerance mediated by Arabidopsis NF-YA family members. PLoS ONE 2012, 7, e48138. [Google Scholar] [CrossRef]
- Sato, H.; Suzuki, T.; Takahashi, F.; Shinozaki, K.; Yamaguchi-Shinozaki, K. NF-YB2 and NF-YB3 have functionally diverged and differentially induce drought and heat stress-specific genes. Plant Physiol. 2019, 180, 1677–1690. [Google Scholar] [CrossRef]
- Mu, J.; Tan, H.; Hong, S.; Liang, Y.; Zuo, J. Arabidopsis transcription factor genes NF-YA1, 5, 6, and 9 play redundant roles in male gametogenesis, embryogenesis, and seed development. Mol. Plant. 2013, 6, 188–201. [Google Scholar] [CrossRef] [PubMed]
- Niu, B.; Xu, J.; E, Z.; Zhang, Z.; Lu, X.; Chen, C. Ectopic expression of OsNF-YA8, an endosperm-specific nuclear factor Y transcription-factor gene, causes vegetative and reproductive development defects in rice. Crop J. 2023, 11, 1719–1730. [Google Scholar] [CrossRef]
- Zhang, M.; Zheng, H.; Jin, L.; Xing, L.; Zou, J.; Zhang, L.; Liu, C.; Chu, J.; Xu, M.; Wang, L. miR169o and ZmNF-YA13 act in concert to coordinate the expression of ZmYUC1 that determines seed size and weight in maize kernels. New Phytol. 2022, 235, 2270–2284. [Google Scholar] [CrossRef]
- Kim, S.K.; Park, H.Y.; Jang, Y.H.; Lee, K.C.; Chung, Y.S.; Lee, J.H.; Kim, J.K. OsNF-YC2 and OsNF-YC4 proteins inhibit flowering under long-day conditions in rice. Planta 2016, 243, 563–576. [Google Scholar] [CrossRef]
- Kumimoto, R.W.; Zhang, Y.; Siefers, N.; Holt, B. NF-YC3, NF-YC4 and NF-YC9 are required for CONSTANS-mediated, photoperiod-dependent flowering in Arabidopsis thaliana. Plant J. 2010, 63, 379–391. [Google Scholar] [CrossRef]
- Turan, S.; Topcu, A.; Karabulut, I.; Vural, H.; Hayaloglu, A. Fatty acid, triacylglycerol, phytosterol, and tocopherol variations in kernel oil of Malatya apricots from Turkey. J. Agric. Food Chem. 2007, 55, 10787–10794. [Google Scholar] [CrossRef]
- Yeap, W.C.; Lee, F.C.; Shabari, S.; Musa, H.; Appleton, D.R.; Kulaveerasingam, H. WRI1-1, ABI5, NF-YA3 and NF-YC2 increase oil biosynthesis in coordination with hormonal signaling during fruit development in oil palm. Plant J. 2017, 91, 97–113. [Google Scholar] [CrossRef] [PubMed]
- Lu, X.; Li, Q.T.; Xiong, Q.; Li, W.; Bi, Y.D.; Lai, Y.C.; Liu, X.L.; Man, W.Q.; Zhang, W.K.; Ma, B.; et al. The transcriptomic signature of developing soybean seeds reveals the genetic basis of seed trait adaptation during domestication. Plant J. 2016, 86, 530–544. [Google Scholar] [CrossRef]
- Junker, A.; Mönke, G.; Rutten, T.; Keilwagen, J.; Seifert, M.; Thi, T.; Renou, J.; Balzergue, S.; Viehöver, P.; Hähnel, U.; et al. Elongation-related functions of LEAFY COTYLEDON1 during the development of Arabidopsis thaliana. Plant J. 2012, 71, 427–442. [Google Scholar] [CrossRef] [PubMed]
- Shen, B.; Allen, W.; Zheng, P.; Li, C.; Glassman, K.; Ranch, J.; Nubel, D.; Tarczynski, M.C. Expression of ZmLEC1 and ZmWRI1 increases seed oil production in maize. Plant Physiol. 2010, 153, 980–987. [Google Scholar] [CrossRef] [PubMed]




Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wu, J.; He, Y.; Wang, L.; Zhao, H.; Jiang, N.; Wuyun, T.; Liu, H. Genome-Wide Identification and Expression Analysis of the NF-Y Transcription Factor Family in Prunus armeniaca. Forests 2024, 15, 1986. https://doi.org/10.3390/f15111986
Wu J, He Y, Wang L, Zhao H, Jiang N, Wuyun T, Liu H. Genome-Wide Identification and Expression Analysis of the NF-Y Transcription Factor Family in Prunus armeniaca. Forests. 2024; 15(11):1986. https://doi.org/10.3390/f15111986
Chicago/Turabian StyleWu, Jiangting, Yanguang He, Lin Wang, Han Zhao, Nan Jiang, Tana Wuyun, and Huimin Liu. 2024. "Genome-Wide Identification and Expression Analysis of the NF-Y Transcription Factor Family in Prunus armeniaca" Forests 15, no. 11: 1986. https://doi.org/10.3390/f15111986
APA StyleWu, J., He, Y., Wang, L., Zhao, H., Jiang, N., Wuyun, T., & Liu, H. (2024). Genome-Wide Identification and Expression Analysis of the NF-Y Transcription Factor Family in Prunus armeniaca. Forests, 15(11), 1986. https://doi.org/10.3390/f15111986





