1. Introduction
Forest plots where the locations of all trees are mapped contain invaluable information for revealing processes of community assembly and dynamics [
1,
2]. Neighborhood analyses are one particularly common use of such data, and involve modeling some metric of tree performance (usually growth rate) as a function of the species identities, sizes, and other aspects of neighboring trees [
3]. These neighborhood models are typically used for inferential projects, where several model structures that represent different hypotheses of a process of interest (e.g., competition) are fitted to the same dataset, and model selection is used to determine the hypothesis with most support [
4]. This approach has allowed many interesting questions to be addressed, including: How are competitive interactions influenced by environmental variables [
5]? How might tree performance respond to climate change [
6]? How are competitive interactions moderated by niche similarities and hierarchies [
7,
8]?
Current neighborhood models are structured around a set of empirically supported patterns of tree performance, but this ecological realism has some drawbacks. For example, tree growth is typically modeled as a non-linear function of tree size and the degree of crowding by neighboring trees [
4]. These non-linear relationships increase the number of parameters and the opportunity for local optima, resulting in a slow and computationally intensive model fitting process (for a linearized version, see [
9,
10]). In addition, the need to fit a separate model for each hypothesis to be tested further increases computation time and thereby limits the number of hypotheses compared; a potential problem when considering the many ways in which processes such as competition can be modeled [
2]. Another consequence of ecological realism is that the models representing different hypotheses tend not to be fully nested, meaning that information theoretic approaches such as Akaike’s information criterion (AIC) must be used for model selection. This is not ideal because AIC penalizes model complexity and can thereby lead to an overly simple final model and bias the conclusions drawn [
11].
Regularized regression is an alternative method for neighborhood analysis that avoids many of these drawbacks. The aim of regularization is the same as AIC model selection; to trade off the antithetical aims of penalizing model complexity and accurately fitting the training data. However, in regularized regression, the strength of this trade-off is determined by a regularization parameter, which is estimated through cross-validation. We focus specifically on Least Absolute Shrinkage and Selection Operator (LASSO) regularized regression [
12], which penalizes model complexity by shrinking coefficients of unhelpful covariates to zero and thereby conducts variable selection autonomously. The main benefit of regularized regression is speed; as a linear model, it can be fitted very quickly, and autonomous variable selection removes the requirement of fitting multiple models representing different hypotheses. Moreover, regularized regression is highly robust to correlated variables; therefore, it allows a single model to include many manifestations of the same process and drop all but the most influential driver [
12]. Importantly, due to its penalization of model complexity, regularization can result in an overly simple final model [
11], but it is unclear whether it leads to more bias than AIC. The main drawback of regularized regression is that it requires a linear modeling framework, which may not accurately approximate the inherently non-linear effects on tree growth [
6,
13]. Overall, regularized regression offers a distinct set of strengths and weaknesses relative to classical neighborhood models.
Another potential use of neighborhood analyses is to predict tree growth beyond the spatial or temporal limits of training data. To the best of our knowledge, neighborhood models have not been used for out-of-sample prediction, but this is likely to change as growing emphasis on dataset publication [
14,
15] and new software tools [
16] increases the availability of mapped forest plot data. Moreover, accurate predictions of tree growth are highly desirable because they could provide insight to current spatial patterns in carbon storage and the effects of land use and climate change on timber production and global carbon cycling. It is important to recognize two levels of prediction: (1)
interpolation of observations experiencing conditions contained in the training data (e.g., of trees experiencing environmental conditions similar to those of trees used for model fitting); and (2)
extrapolation to observations experiencing conditions not present in the training data (e.g., novel climates). However, neither the interpolation nor extrapolation ability of neighborhood models has been tested.
Both classical neighborhood models and regularized regression may be well-suited to prediction, but it is unclear which method will be the most accurate. The biggest pitfall in predictive modeling is applying an overly complex model that fits the sample data well, but does not generalize to other data because it includes relationships that are specific to the sample. Both AIC and regularization are designed to avoid this overfitting problem by penalizing model complexity, and therefore classical and regularized regression neighborhood models may produce accurate out-of-sample predictions. The cross-validation model selection approach used to select the regularization parameter also guards against overfitting and is growing in popularity in ecology [
17,
18]. However, cross-validation has been shown to be asymptotically equivalent to AIC [
19] and also requires models to be fitted to many subsets of the data. To ensure that resources are not wasted on applying cross-validation to classical neighborhood models, it is timely to empirically demonstrate the equivalency between AIC and cross-validation model selection.
In this study, we built a novel regularized regression model of tree growth and used it to test hypotheses regarding how tree growth is influenced by the presence and species identity of neighboring trees. This is a particularly interesting test case for a new modeling method because variation in the abundances of different neighbor species leads to an inherently unbalanced sample, which is likely to create challenges in model fitting. To evaluate the accuracy of our model’s inferences, we compared them with those of a commonly used neighborhood model ([
4] hereafter, likelihood model). In addition, we investigated the ability of both neighborhood models (i.e., regularized regression and likelihood) to interpolate the growth of trees not used in model fitting. We also evaluated the predictive performance of a cross-validated likelihood model to demonstrate the equivalency between AIC and cross-validation model selection [
19]. Overall, we found that regularized regression makes similar inferences to the likelihood model, but that the neighborhood model with the most accurate out-of-sample predictions varies between focal tree species.
2. Materials and Methods
2.1. Tree Growth Data
The data used in this study came from the mature and old-growth conifer forests of Mount Rainier National Park, WA, USA. Mount Rainier is a 4392 m high volcano and covers a large climatic gradient. Increasing elevation is associated with decreasing temperatures and increasing precipitation, although precipitation is considerably reduced on the eastern side of the volcano due to a rain shadow effect. The region experiences a temperate maritime climate with warm, dry summers and cold, wet winters.
We used data collected in 15 forest plots established in 1977 and 1978 as part of the Pacific Northwest Permanent Sample Plot Program [
20]. These plots were intentionally located to capture the diversity of climatic conditions on Mount Rainier, and therefore range in elevation from 581 to 1492 m. All plots are 1 ha (100 m × 100 m) in size and, at the time of their establishment, all trees with a diameter at breast height (1.37 m above ground level; hereafter DBH) ≥ 15 cm were tagged, identified to species, mapped on a coordinate grid, and had their DBH recorded. Between 25% and 100% of the area of each plot was also designated as a detailed plot where data were collected on all trees with a DBH ≥ 5 cm. Approximately every five years, all plots are revisited to tag new trees meeting the minimum size threshold, document tree mortality, and to re-measure the size of tagged trees, with the most recent census occurring in 2017.
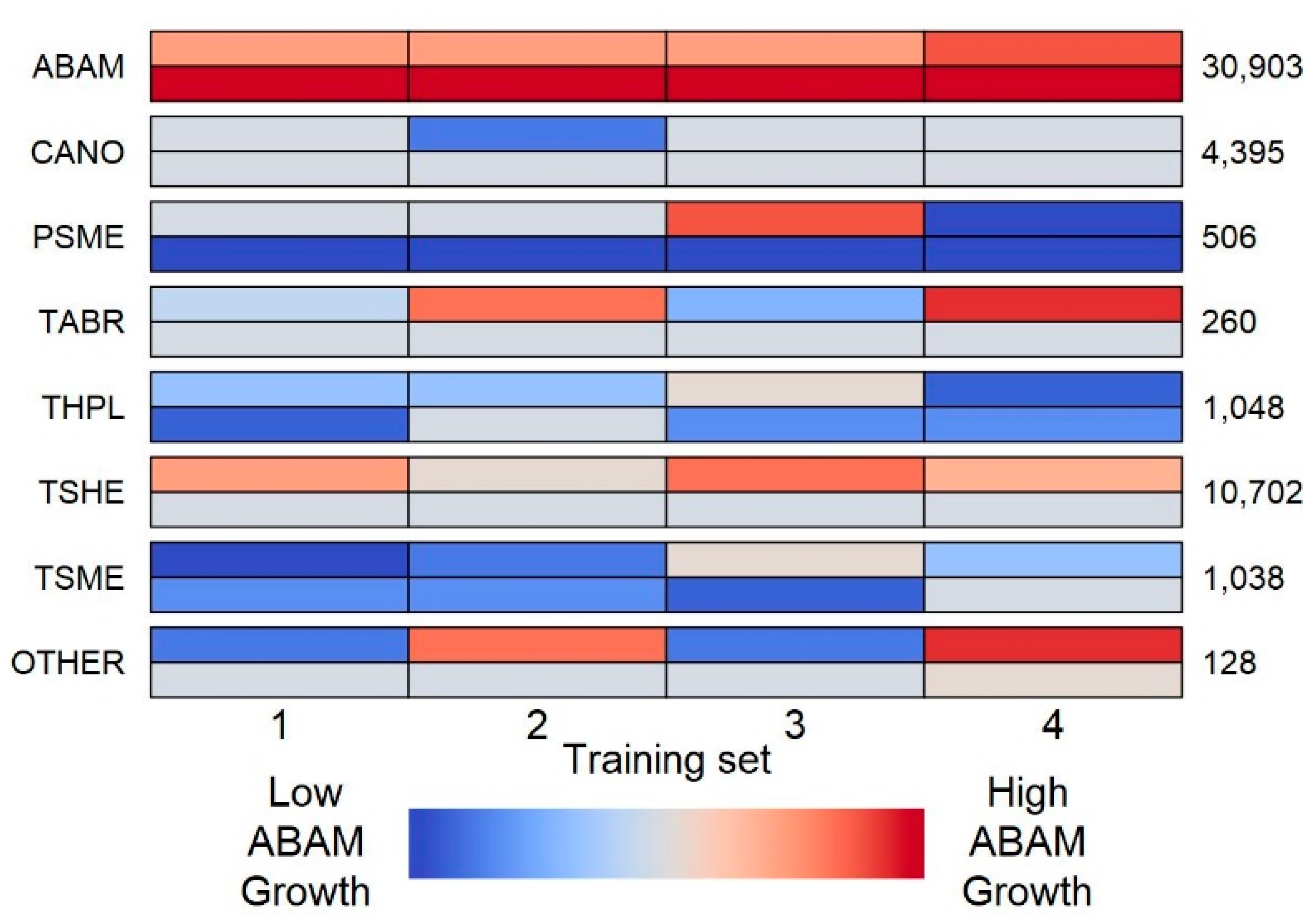
We calculated average annual growth for each tree as the difference in DBH between its earliest and most recent measurement divided by the number of years elapsed between those measurements. The slow growth rates of trees in this harsh high-elevation environment meant that measurement inaccuracies sometimes resulted in biologically impossible negative growth rates; all such trees were excluded from our analysis (1.6% of focal trees). The smaller trees recorded only in the detailed plots (5–15 cm DBH) were included in our analyses as focal trees but excluded as neighbors that could influence the growth of other focal trees to prevent systematic bias in neighbor interactions between detailed and non-detailed areas of the plots. Of the 17 tree species included in the dataset, we modeled growth for only the six species represented by at least 100 individuals:
Abies amabilis,
Callitropsis nootkatensis,
Pseudotsuga menziesii,
Thuja plicata,
Tsuga heterophylla, and
Tsuga mertensiana (hereafter: ABAM, CANO, PSME, THPL, TSHE, TSME, respectively; see
Table S1 for full species list and
Table S2 for a summary of how focal species were distributed across sample plots).
For our neighborhood models, we considered all trees growing within 15 m of a focal tree to be that focal tree’s neighbors. This neighborhood size is comparable to those used in other studies [
4,
7] and was found through our own exploratory analyses to result in the best training data fits. To avoid edge effects, all focal trees within 15 m of a forest plot boundary were excluded from our analysis. Each of the remaining focal trees was assigned to one of four test datasets at random, such that 25% of the focal trees in each plot were placed in each test set. We then defined a corresponding training dataset for each test set such that the first training set consisted of all focal trees not in the first test set (75% of all focal trees). All models were fitted to each of the four training sets to assess the robustness of conclusions; test sets were used to evaluate the predictive skill. As in previous studies, we also created separate models for each of our focal species because model parameter values are expected to differ greatly between species.
The data and code underlying all presented analyses are available on Zenodo at doi:10.5281/zenodo.5512791, reference number [
21].
2.2. Likelihood Model
The likelihood model [
4,
22] has the following generalized formula:
where
is the predicted growth,
is an estimated maximum potential growth rate in the absence of neighbors,
is a size effect,
is a climate effect, and
is a crowding effect, with subscripts indicating whether effects vary between focal trees (
) or plots (
). The size, climate and crowding effects can take any value between 0 and 1; therefore, growth predictions can take any value between 0 and
.
The size effect (
) accounts for the expectation that trees have an optimal size at which maximum growth occurs and is modeled as a lognormal distribution:
Parameter specifies the DBH at which maximum growth occurs, and parameter determines the width of the lognormal distribution. This formulation is highly flexible, allowing the relationship between focal growth and size to be monotonically increasing, monotonically decreasing, or non-monotonic.
The climate effect (
) accounts for the expectation that growth rates will differ among the plots due to their different climatic conditions. Although there are many climatic variables that differ dramatically along the elevational gradient on which our plots are situated, these variables are strongly correlated [
23], and potential evapotranspiration (PET) is informative of growth rates of our focal species in these plots [
24]. Consequently, we used PET as the sole abiotic variable in our models and calculated the average annual PET for each plot from the time of plot establishment up until the most recent tree measurements, following the protocol outlined in [
24]. The climate effect (
) is modeled as a Gaussian distribution:
where
represents the average annual PET of plot
(where the focal tree resides),
specifies the PET at which maximum growth occurs, and
determines the width of the Gaussian distribution. As with the size effect, this flexible structure allows the relationship between focal growth and PET to monotonically increase, monotonically decrease, or be non-monotonic.
To incorporate the effects of neighbors on focal tree growth, a neighborhood crowding index (NCI) was calculated for tree
as:
where
is the number of neighbor species and
is the number of trees of species
in focal tree
’s neighborhood. This formula reflects the expectation that a neighbor’s influence on focal tree growth increases with its size but decreases with its distance from the focal, and the estimated parameters
and
allow these relationships to be non-linear. The effect of neighbor size and distance is also multiplied by an estimated interaction coefficient (
), which takes a value between 0 and 1 and represents the effect of neighbors of species
on the growth of focal trees of the species being modeled.
The crowding effect (
) is calculated as a negative exponential function of
NCI:
where
is an estimated parameter that modulates the growth response of trees to varying
NCI values.
To make inferences regarding the effects of neighbors on focal growth, we fitted four variations of the likelihood model for each focal species: (1) no interactions—crowding effect () excluded; (2) equivalent interactions—no parameters included (quantitatively equivalent to a single with value 1); (3) conspecific vs. heterospecific interactions—two parameters, one for conspecific neighbors () and another for heterospecific neighbors (); and (4) species-specific interactions—estimated for each neighbor species i. In the species-specific interaction models, rare neighbor species were grouped under a single parameter. Rare neighbor species (<5% of neighbors in each focal species × training set combination) were defined as those that appeared as neighbors of the focal species fewer than 100 times, when averaged across the four training sets. We elected to use an average instead of specifying rare neighbor species separately for each training set to ensure that the fitted parameters could be compared across training sets. The four model structures for each focal species × training set combination were compared using Akaike’s information criterion corrected for a low sample size (AICc).
Parameter values were estimated using the simulated annealing algorithm implemented through the
optim function in the base library of R 4.0.2 [
25]. The optimizations were facilitated through the use of advanced computational, storage, and networking infrastructure provided by the Hyak supercomputer system at the University of Washington.
2.3. Regularized Regression Model
In our regularized regression model, focal tree growth was modeled as a linear function of: the species identity, size and proximity of neighbors; PET; and the densities of each neighbor species, and all species combined, in the neighborhood. Rare neighbor species (characterized in the same way as for the likelihood model) were assigned a species identity of “other”, and the density of “other” was also included in the model. To estimate the effects of neighbor identity, size and proximity in a linear modeling framework, each focal tree–neighbor interaction was treated as an independent observation. This resulted in a design matrix where each focal tree occupied rows, with n being the number of neighbors in its neighborhood. The growth rate, PET and densities were necessarily identical across all n rows corresponding to the same focal tree. This design matrix structure resulted in growth predictions of each focal tree based on each of its interactions and we used the arithmetic average of these predictions as the final prediction of focal growth.
To meet the assumptions of linear regression, the growth rate data were transformed to approximate a normal distribution as follows:
This transformation has the additional benefit of partially accounting for the non-linear relationship between focal size and focal growth included in the likelihood model; it allows a saturating but always monotonic relationship between growth rate and tree size.
We fitted the regularized regression models using the
cv.glmnet function of the glmnet R package [
26]. This function estimates parameter values through a stochastic cyclical coordinate descent algorithm using a set of values for the regularization parameter. It then uses 10-fold cross-validation to evaluate the models fitted with different regularization parameter values, reporting the mean square error (MSE) for each. Of the multiple output models, glmnet indicates the one with the highest regularization value (strongest regularization) that resulted in an MSE within one standard error of the model with the lowest MSE; we used this model for interpretation. In addition, the rapid fitting of the regularized regression models allowed us to fit 100 models for each focal species by training set combination to evaluate how consistent the findings of this model are in the face of the stochastic fitting process. Of these 100 models, the one with the lowest MSE was used for evaluating model fit to the training data and predictive performance. The model fitting procedure implemented by glmnet is rapid; therefore, it was conducted on a personal laptop.
2.4. Comparing Inferential Performance
To determine whether our regularized regression model could replicate the inferences of the likelihood model, we compared the conclusions each of the models would have led us to for four commonly asked questions regarding the impact of neighbors on tree growth. Separately for each focal species, we asked: (1) Is focal growth influenced by neighboring trees? (2) Is focal growth influenced by neighbor species identity? (3) Is focal growth higher in the presence of conspecific or heterospecific neighbors? and (4) Which neighbor species are associated with the highest/lowest focal growth? For each modeling approach, the conditions under which we drew particular conclusions regarding these questions are outlined in
Table 1.
2.5. Evaluating Predictive Performance
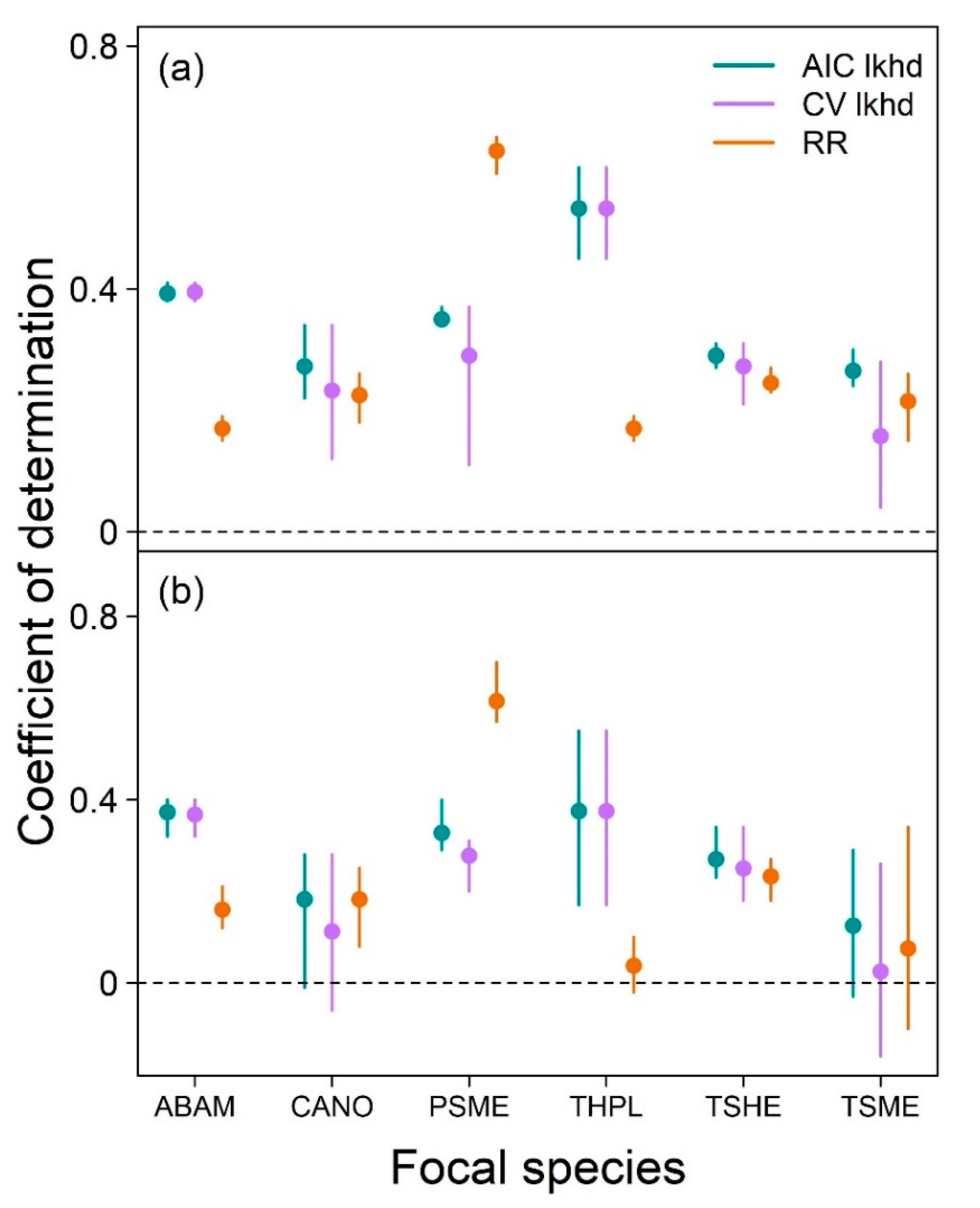
To investigate the predictive potential of neighborhood models, we evaluated the out-of-sample predictive ability for three different models for each focal species by training set combination: regularized regression, AIC likelihood and CV likelihood. Each of these predictive models was one of the models described in Methods: Inference. For the regularized regression model, we used the model with the lowest MSE (out of the 100 models run). For the AIC and CV likelihood models, we used the models with the lowest AIC and lowest cross-validated MSE, respectively, of the four model structures. To identify the lowest cross-validated MSE, we divided each training set into 10 folds (consistent with regularized regression cross-validation), fitted each of the four likelihood model structures to each possible set of 9 folds, then calculated MSE of the models’ predictions of the 10th fold. We averaged the resulting 10 MSE values to obtain the cross-validated MSE of each model structure.
We quantified the out-of-sample prediction (interpolation) ability of each model type applied to each training set by calculating the coefficient of determination (R
2) of the model when applied to its corresponding test set, which was entirely unused in the fitting of that model (see
Section 2.1 Tree growth data for how training and test sets were defined). This metric measures the proportion of variance around the mean value of the dependent variable explained by the model. The maximum possible value for a coefficient of determination is 1 (all variance explained), but negative values can exist when a model is applied to unseen test data if there is more unexplained variation around model predictions than exists around the mean growth value in the test data.
It is often advised that training and test sets be spatially or temporally separated to prevent overestimates of predictive ability that can result from spatial or temporal autocorrelation. In this study, we present results from spatially overlapping training and test sets because the predictive performance of the models was similar when the spatially separated training and test sets were used. Moreover, although we suspect that spatial and temporal autocorrelation in unmeasured variables which influence tree growth was likely in our dataset (e.g., soil conditions, pest damage), we do not know the scale of such variation—which means that it is unclear whether spatial or temporal separation would address such autocorrelation.
5. Conclusions
We have developed a regularized regression model of neighborhood-dependent tree growth that can replicate the ecological inferences of a classical likelihood model in a fraction of the time. Regularization is particularly efficient for inferential projects because it automates the process of model selection and can handle correlated explanatory variables. This feature means that our regularized regression model could also be used to select among potential explanatory variables (e.g., climate variables) and thereby streamline the development of a classical likelihood model. We encourage the investigation of regularization as a tool for modeling tree growth and other processes that have many potential covariates, such as seedling survival [
30] and mature tree mortality [
31].
We have also shown that neighborhood models, including regularized regression, can provide accurate growth predictions of trees not used in model fitting. However, we only tested the model’s predictive skill on trees that experienced similar conditions to those used in model fitting. Future research should investigate whether the findings of neighborhood models can be extrapolated to trees experiencing conditions absent from the training set (e.g., novel climates). Although the optimal neighborhood modeling approach for prediction will vary among species and systems, we believe that our regularized regression has great potential due to its rapid fitting and ability to include many explanatory variables that represent different models of complex processes such as competition. Overall, we found that regularized regression and likelihood approaches are complementary to better understand the drivers of tree growth, and suggest that regularization will be a valuable tool for advancing the field of tree growth modeling toward prediction.