Fungal Community Composition and Enzyme Activity in Different Type Bark of Pinus koraiensis
Abstract
:1. Introduction
2. Materials and Methods
2.1. Study Sites
2.2. Sample Preparation
2.3. Physico-Chemical Properties Analyses
2.4. Extracellular Enzyme Activities
2.5. DNA Extraction, Sequencing and Quality Control
2.6. Bioinformatic Analyses and Functional Annotation
2.7. Statistical Analyses
3. Results
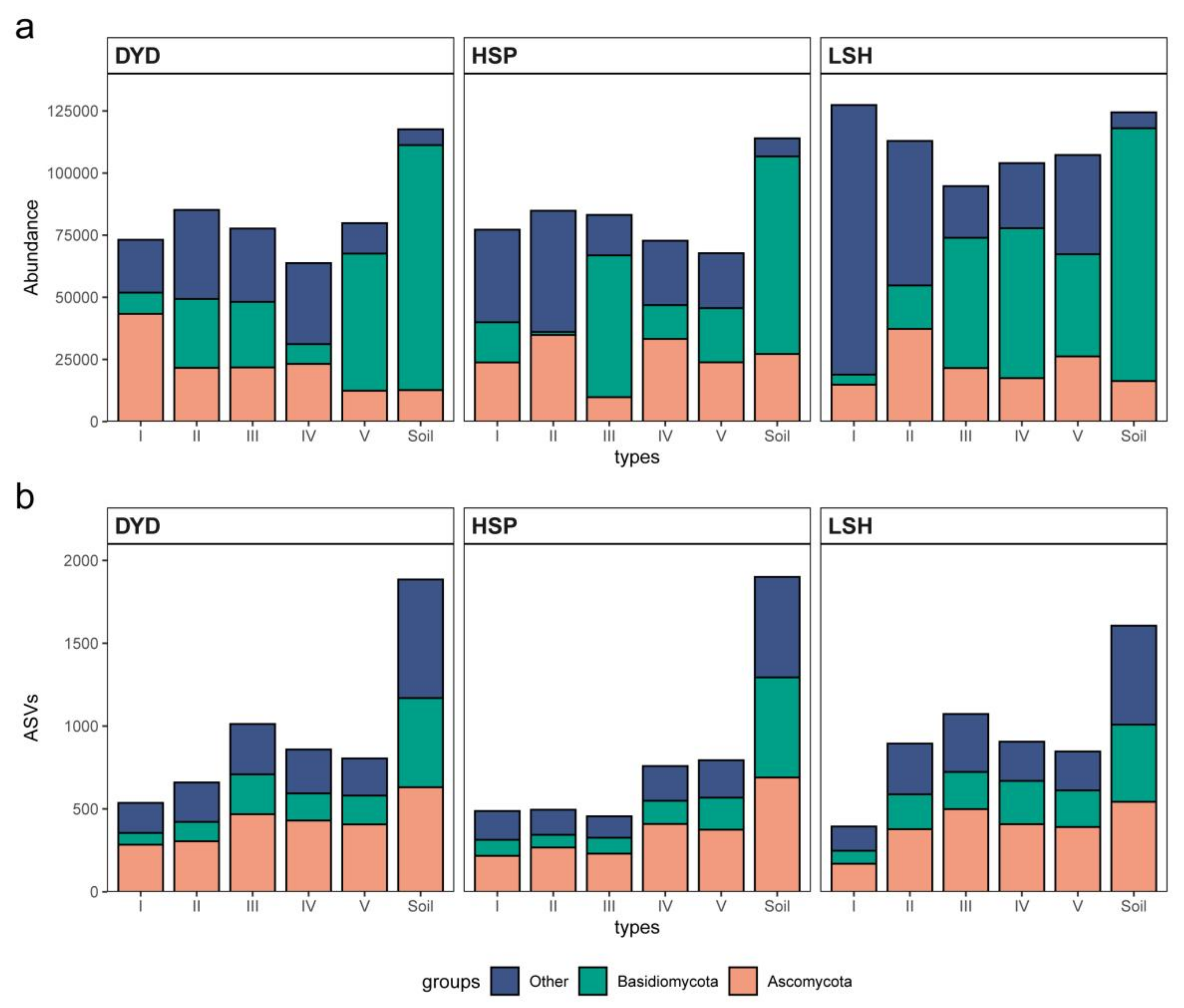
3.1. Pine Barks Fungal Species Richness
3.2. Ordination Plot of Non-Metric Multidimensional Scaling of All Fungal Communities
3.3. Physico-Chemical Properties of Pine Barks
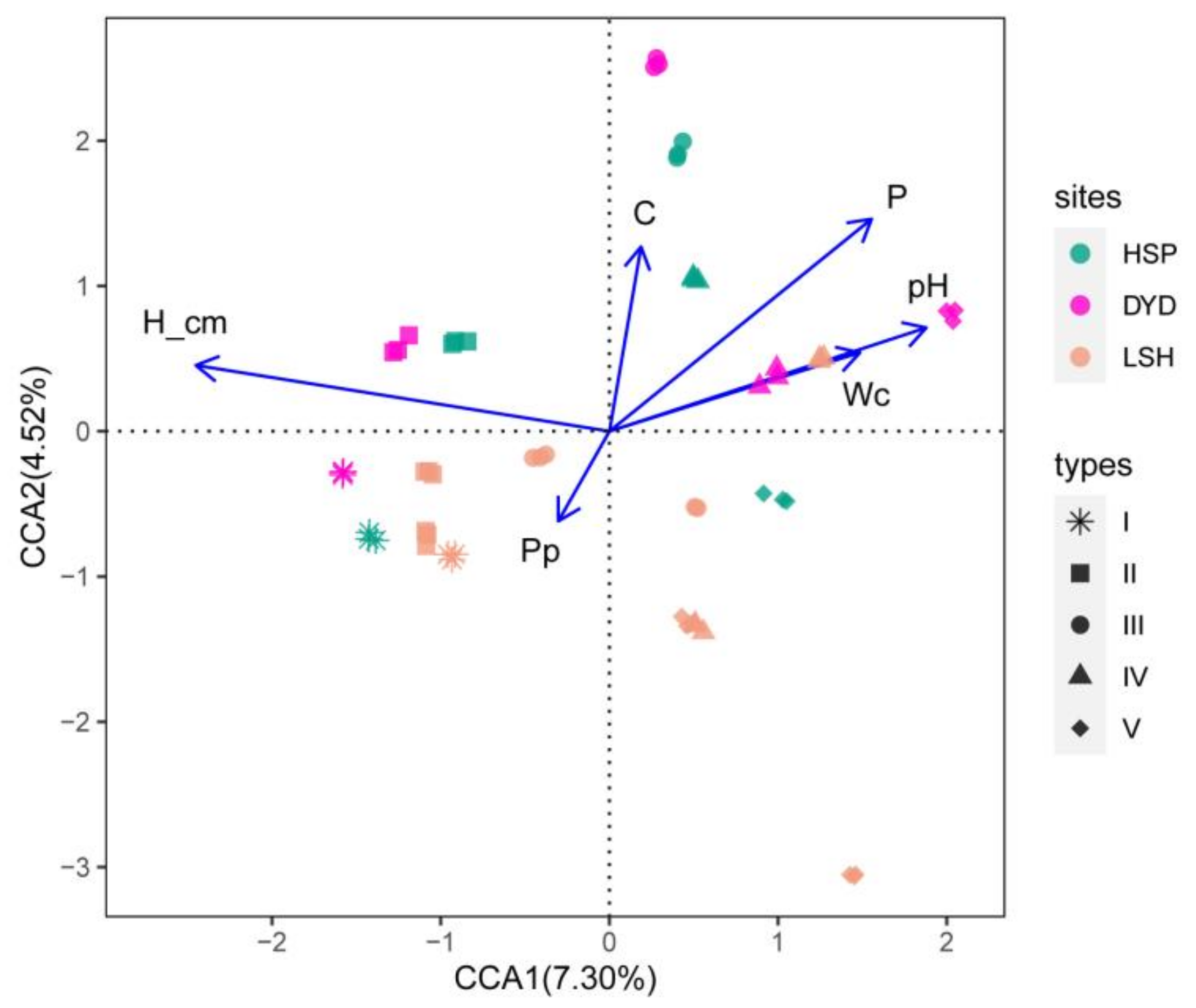
3.4. Canonical Correspondence Analysis of Korean Pine Bark Fungal Community Structure
3.5. Correlation Analysis between Dominant ASVs and Physico-Chemical Properties of Pine Bark
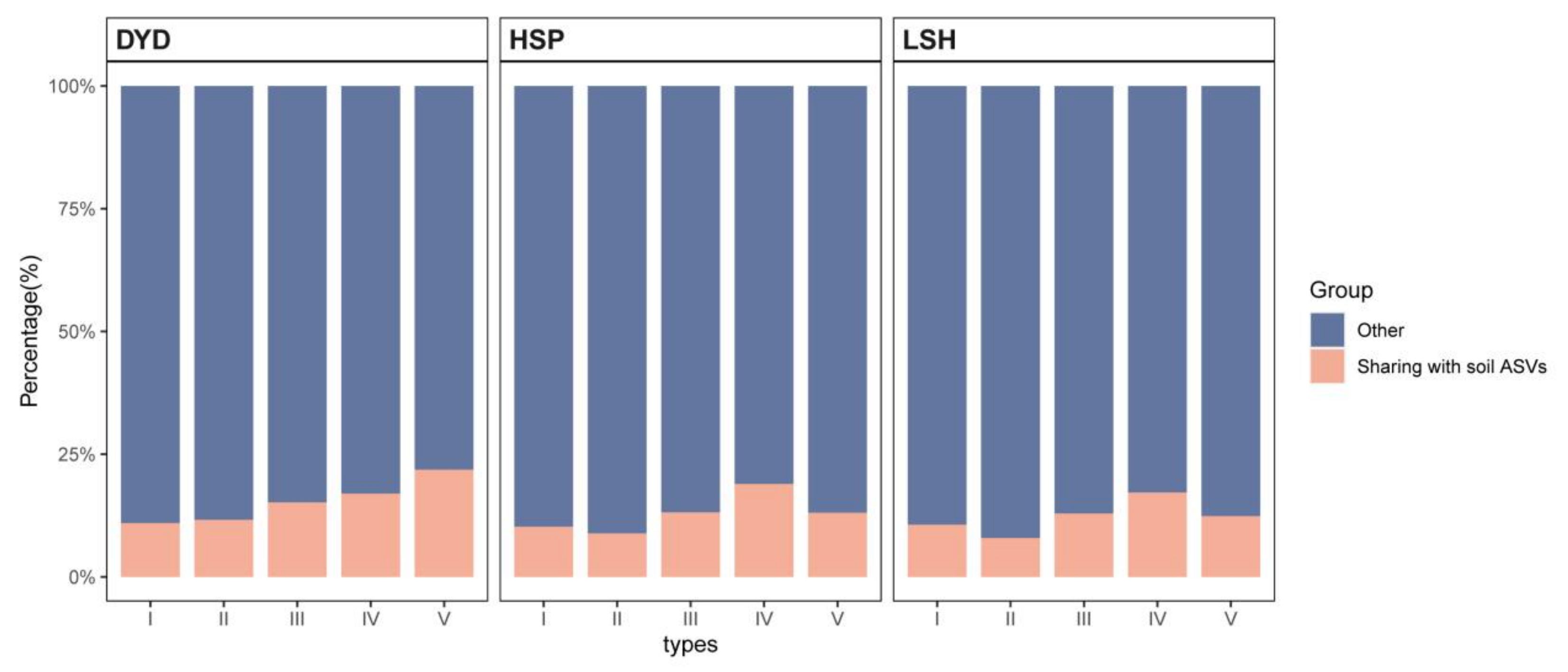
3.6. Effect of Soil Fungal Community on Pine Barks Fungal Community
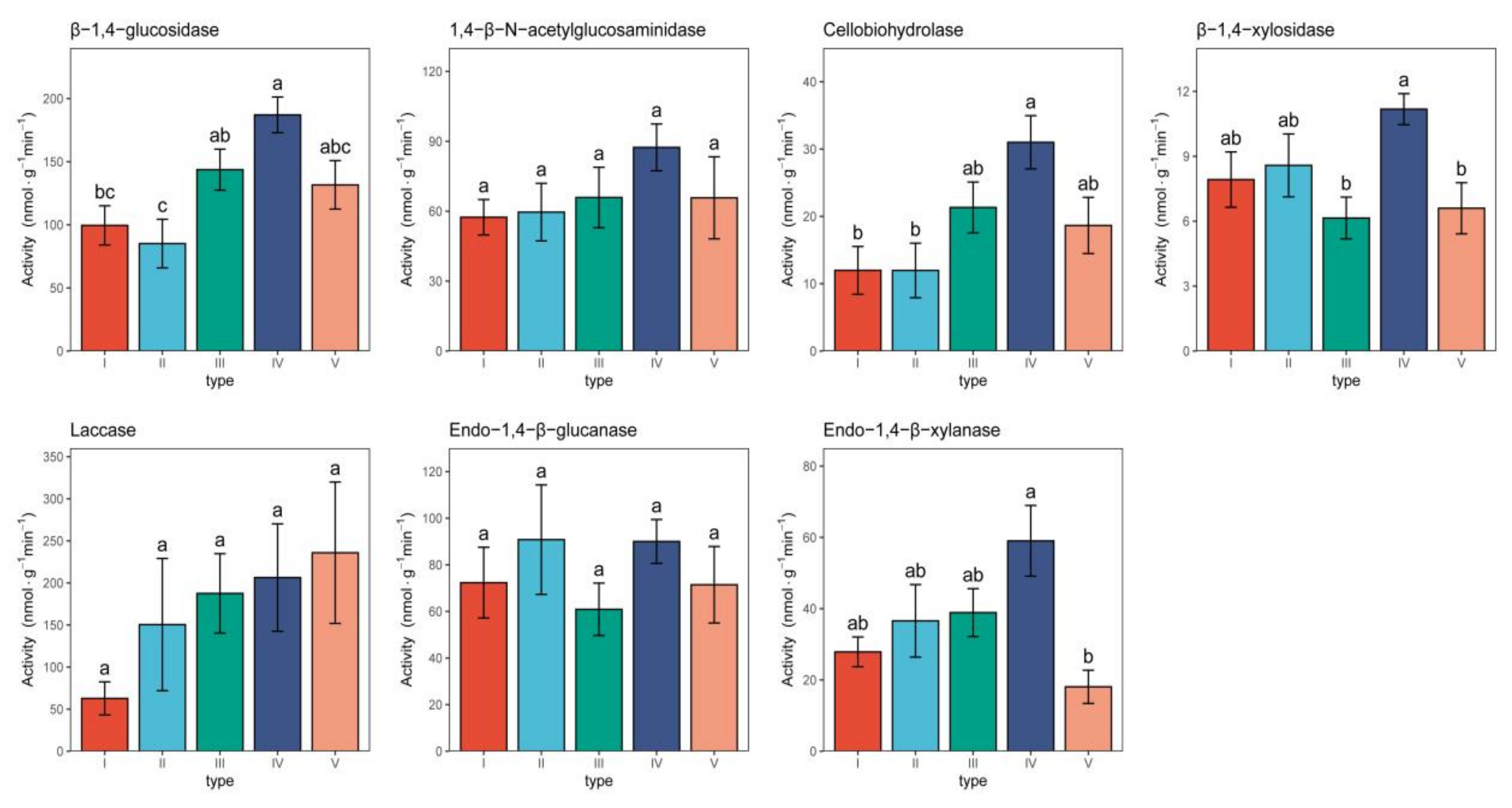
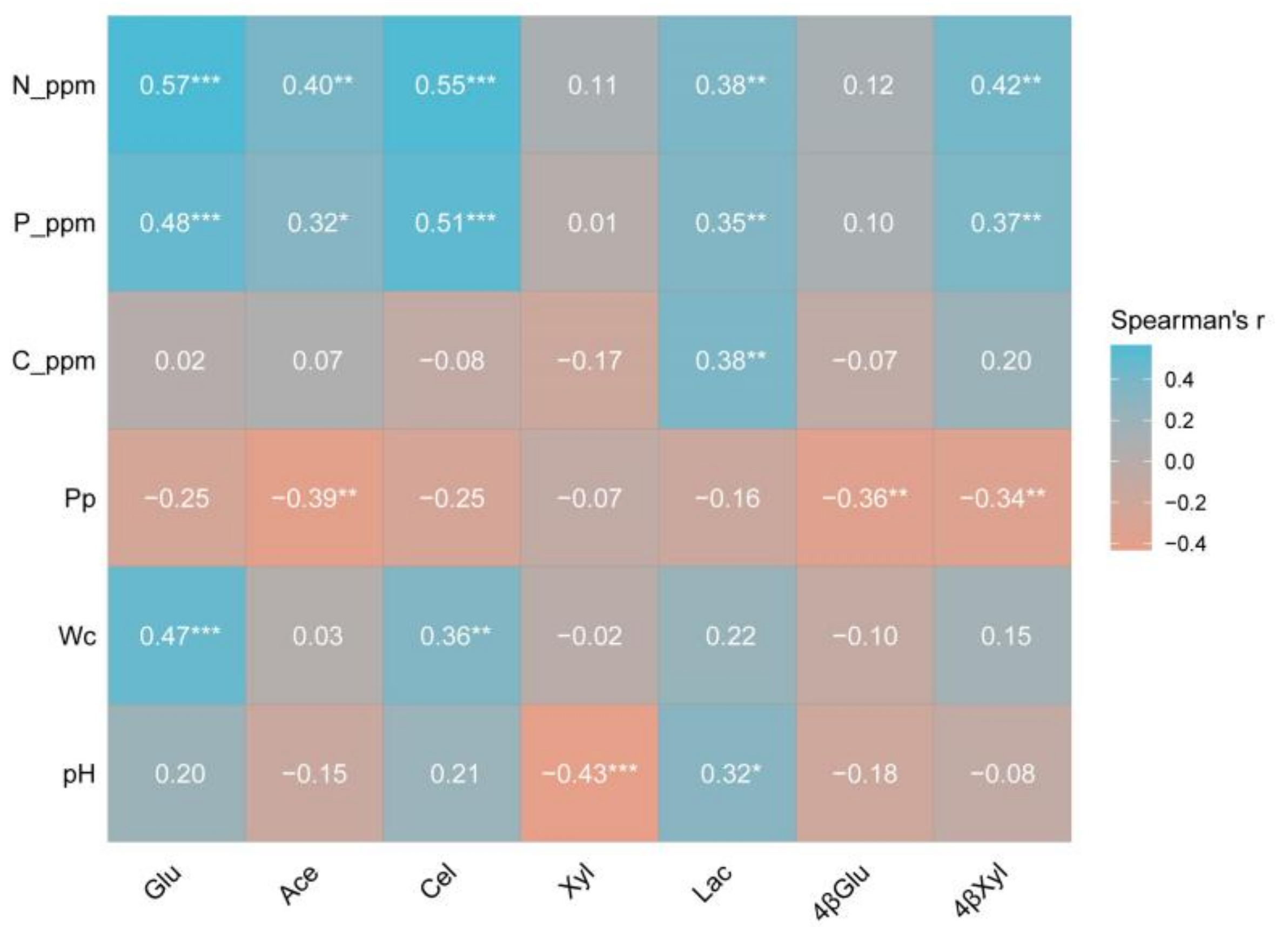
3.7. Extracellular Enzyme Activity of Different Pine Bark Types
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Siitonen, J. Microhabitats. In Biodiversity in Dead Wood, 1st ed.; Stokland, J.N., Siitonen, J., Jonsson, B.G., Eds.; Cambridge University Press: Cambridge, UK, 2012; p. 301. [Google Scholar]
- Liers, C.; Ullrich, R.; Steffen, K.T.; Hatakka, A.; Hofrichter, M. Mineralization of 14C-labelled synthetic lignin and extracellular enzyme activities of the wood-colonizing ascomycetes Xylaria hypoxylon and Xylaria polymorpha. Appl. Microbiol. Biotechnol. 2005, 69, 573–579. [Google Scholar] [CrossRef] [PubMed]
- Hofrichter, M.; Vares, T.; Kalsi, M.; Galkin, S.; Scheibner, K.; Fritsche, W.; Hatakka, A. Production of Manganese Peroxidase and Organic Acids and Mineralization of 14 C-Labelled Lignin (14 C-DHP) during Solid-State Fermentation of Wheat Straw with the White Rot Fungus Nematoloma frowardii. Appl. Environ. Microbiol. 1999, 65, 1864–1870. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Arnstadt, T.; Hoppe, B.; Kahl, T.; Kellner, H.; Krüger, D.; Bässler, C.; Bauhus, J.; Hofrichter, M. Patterns of laccase and peroxidases in coarse woody debris of Fagus sylvatica, Picea abies and Pinus sylvestris and their relation to different wood parameters. Eur. J. For. Res. 2015, 135, 109–124. [Google Scholar] [CrossRef] [Green Version]
- Paine, C.E.T.; Stahl, C.; Courtois, E.A.; Patiño, S.; Sarmiento, C.; Baraloto, C. Functional explanations for variation in bark thickness in tropical rain forest trees. Funct. Ecol. 2010, 24, 1202–1210. [Google Scholar] [CrossRef]
- Rosell, J.A. Bark thickness across the angiosperms: More than just fire. New Phytol. 2016, 211, 90–102. [Google Scholar] [CrossRef]
- Lymperopoulou, D.S.; Adams, R.I.; Lindow, S.E. Contribution of Vegetation to the Microbial Composition of Nearby Outdoor Air. Appl. Environ. Microbiol. 2016, 82, 3822–3833. [Google Scholar] [CrossRef] [Green Version]
- Shorohova, E.; Kapitsa, E.; Kazartsev, I.; Romashkin, I.; Polevoi, A.; Kushnevskaya, E. Tree species traits are the predominant control on the decomposition rate of tree log bark in a mesic old-growth boreal forest. For. Ecol. Manag. 2016, 377, 36–45. [Google Scholar] [CrossRef]
- Schmidt, O. Wood and Tree Fungi; Springer: Berlin/Heidelberg, Germany, 2006. [Google Scholar]
- Bing, L.A.; Lewis, L.C. Suppression of Ostrinia nubilalis (Huebner) (Lepidoptera: Pyralidae) by endophytic Beauveria bassiana (Balsamo) Vuillemin. Environ. Entomol. 1991, 20, 1207–1211. [Google Scholar] [CrossRef]
- Stone, J.K.; Bacon, C.W.; White, J.F. An Overview of Endophytic Microbes: Endophytism Defined. In Microbial Endophytes; Bacon, C.W., White, J.F., Jr., Eds.; Marcel Dekker Inc.: New York, NY, USA, 2000. [Google Scholar]
- Schmull, M.; Hauck, M. Element microdistribution in the bark of Abies balsamea and Picea rubens and its impact on epiphytic lichen abundance on Whiteface Mountain, New York. Flora Morphol. Distrib. Funct. Ecol. Plants 2003, 198, 293–303. [Google Scholar] [CrossRef]
- Vega, F.E.; Posada, F.; Aime, M.C.; Pava-Ripoll, M.; Infante, F.; Rehner, S.A. Entomopathogenic fungal endophytes. Biol. Control. 2008, 46, 72–82. [Google Scholar] [CrossRef] [Green Version]
- Kazartsev, I.; Shorohova, E.; Kapitsa, E.; Kushnevskaya, H. Decaying Picea abies log bark hosts diverse fungal communities. Fungal Ecol. 2018, 33, 1–12. [Google Scholar] [CrossRef]
- Watkinson, S.; Bebber, D.; Darrah, P.; Fricker, M.; Tlalka, M.; Boddy, L. The role of wood decay fungi in the carbon and nitrogen dynamics of the forest floor. In Fungi in Biogeochemical Cycles; Gadd, G.M., Ed.; Cambridge University Press (CUP): Cambridge, UK, 2006; pp. 151–181. [Google Scholar]
- Ódor, P.; Heilmann-Clausen, J.; Christensen, M.; Aude, E.; van Dort, K.W.; Piltaver, A.; Siller, I.; Veerkamp, M.T.; Walleyn, R.; Standovár, T.; et al. Diversity of dead wood inhabiting fungi and bryophytes in semi-natural beech forests in Europe. Biol. Conserv. 2006, 131, 58–71. [Google Scholar] [CrossRef]
- Mäkipää, R.; Rajala, T.; Schigel, D.; Rinne, K.T.; Pennanen, T.; Abrego, N.; Ovaskainen, O. Interactions between soil- and dead wood-inhabiting fungal communities during the decay of Norway spruce logs. ISME J. 2017, 11, 1964–1974. [Google Scholar] [CrossRef] [Green Version]
- Kubartová, A.; Ottosson, E.; Dahlberg, A.; Stenlid, J. Patterns of fungal communities among and within decaying logs, revealed by 454 sequencing. Mol. Ecol. 2012, 21, 4514–4532. [Google Scholar] [CrossRef]
- Ovaskainen, O.; Schigel, D.; Ali-Kovero, H.; Auvinen, P.; Paulin, L.; Nordén, B.; Nordén, J. Combining high-throughput sequencing with fruit body surveys reveals contrasting life-history strategies in fungi. ISME J. 2013, 7, 1696–1709. [Google Scholar] [CrossRef]
- Lindahl, B.D.; Ihrmark, K.; Boberg, J.; Trumbore, S.E.; Högberg, P.; Stenlid, J.; Finlay, R.D. Spatial separation of litter decom-position and mycorrhizal nitrogen uptake in a boreal forest. New Phytol. 2007, 173, 611–620. [Google Scholar] [CrossRef] [Green Version]
- Rajala, T.; Tuomivirta, T.; Pennanen, T.; Mäkipää, R. Habitat models of wood-inhabiting fungi along a decay gradient of Norway spruce logs. Fungal Ecol. 2015, 18, 48–55. [Google Scholar] [CrossRef] [Green Version]
- Tedersoo, L.; Kõljalg, U.; Hallenberg, N.; Larsson, K.H. Fine scale distribution of ectomycorrhizal fungi androots across sub-strate layers including coarse woody debris in a mixed forest. New Phytol. 2003, 159, 153–165. [Google Scholar] [CrossRef]
- Boddy, L.; Watkinson, S.C. Wood decomposition, higher fungi, and their role in nutrient redistribution. Can. J. Bot. 1995, 73, 1377–1383. [Google Scholar] [CrossRef]
- Philpott, T.; Prescott, C.; Chapman, W.; Grayston, S. Nitrogen translocation and accumulation by a cord-forming fungus (Hypholoma fasciculare) into simulated woody debris. For. Ecol. Manag. 2014, 315, 121–128. [Google Scholar] [CrossRef]
- Harmon, M.E.; Franklin, J.F.; Swanson, F.J.; Sollins, P.; Gregory, S.V.; Lattin, J.D.; Anderson, N.H.; Cline, S.P.; Aumen, N.G.; Sedell, J.R.; et al. Ecology of coarse woody debris in temperate ecosystems. Adv. Ecol. Res. 1986, 15, 133–302. [Google Scholar]
- Ma, J.L.; Zhuang, L.W.; Chen, D.; Li, J.W. Geographic distribution of Pinus koraiensis in the world. J. Northeast. For. Univ. 1992, 5, 40–48. [Google Scholar]
- Huang, Y.Y. The Separation, Isolation and Identification of Polyphenol Sturctures in Korean Pine (Pinus koraiensis) Bark and Evaluation of Its Antioxidant and Anticancer Activity; Northeast Forestry University: Harbin, China, 2014. [Google Scholar]
- Wei, Y.-L. Polypore species diversity, floral composition, and distribution characteristics in Changbai Mountains, Northeast China. Yingyong Shengtai Xuebao 2011, 22, 2711–2717. [Google Scholar] [PubMed]
- Valšková, V.; Šnajdr, J.; Bittner, B.; Cajthaml, T.; Merhautová, V.; Hoffichter, M.; Baldrian, P. Production of lignocellu-lose-degrading enzymes and degradation of leaf litter by saprotrophic basidiomycetes isolated from a Quercus petraea forest. Soil Biol. Biochem. 2007, 39, 2651–2660. [Google Scholar] [CrossRef]
- Bolyen, E.; Rideout, J.R.; Dillon, M.R.; Bokulich, N.A.; Abnet, C.; Alghalith, G.A.; Alexander, H.; Alm, E.; Arumugam, M.; Animozhiyan, A.; et al. QIIME 2: Reproducible, interactive, scalable, and extensible microbiome data science. Peer J. Preprints 2018, 6, e27295v2. [Google Scholar] [CrossRef]
- Callahan, B.J.; Mcmurdie, P.J.; Rosen, M.J.; Han, A.W.; Johnson, A.J.A.; Holmes, S.P. DADA2: High-resolution sample inference from Illumina amplicon data. Nat. Methods 2016, 13, 581–583. [Google Scholar] [CrossRef] [Green Version]
- Katoh, K.; Misawa, K.; Kuma, K.; Miyata, T. MAFFT: A novel method for rapid multiple sequence alignment based on fast Fourier transform (describes the FFT-NS-1, FFT-NS-2 and FFT-NS-i strategies). Nucleic Acids Res. 2002, 30, 3059–3066. [Google Scholar] [CrossRef] [Green Version]
- Price, M.N.; Dehal, P.S.; Arkin, A.P. FastTree: Computing Large Minimum Evolution Trees with Profiles instead of a Distance Matrix. Mol. Biol. Evol. 2009, 26, 1641–1650. [Google Scholar] [CrossRef]
- Bokulich, N.A.; Kaehler, B.D.; Ram, R.J.; Matthew, D.; Evan, B.; Rob, K.; Huttley, G.A.; Caporaso, J.G. Optimizing taxonomic classification of marker-gene amplicon sequences with qiime 2’s q2-feature-classifier plugin. Microbiome 2018, 6, 90. [Google Scholar] [CrossRef]
- McDonald, D.; Price, M.N.; Goodrich, J.; Nawrocki, E.P.; DeSantis, T.Z.; Probst, A.; Andersen, G.L.; Knight, R.; Hugenholtz, P. An improved Greengenes taxonomy with explicit ranks for ecological and evolutionary analyses of bacteria and archaea. ISME J. 2012, 6, 610–618. [Google Scholar] [CrossRef]
- Wang, Q.; Garrity, G.M.; Tiedje, J.M.; Cole, J.R. Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl. Environ. Microbiol. 2007, 73, 5261–5267. [Google Scholar] [CrossRef] [Green Version]
- Abarenkov, K.; Tedersoo, L.; Nilsson, R.H.; Vellak, K.; Saar, I.; Veldre, V.; Parmasto, E.; Prous, M.; Aan, A.; Ots, M.; et al. PlutoF-a web based workbench for ecological and taxonomic research, with an online implementation for fungal ITS sequences. Evol. Bioinf. 2010, 6, 189–196. [Google Scholar] [CrossRef]
- Altschul, S.F.; Madden, T.L.; Schäffer, A.A.; Zhang, J.; Zhang, Z.; Miller, W.; Lipman, D.J. Gapped BLAST and PSI-BLAST: A new generation of protein database search programs. Nucleic Acids Res. 1997, 25, 3389–3402. [Google Scholar] [CrossRef] [Green Version]
- Nguyen, N.H.; Song, Z.; Bates, S.T.; Branco, S.; Tedersoo, L.; Menke, J.; Schilling, J.S.; Kennedy, P. FUNGuild: An open anno-tation tool for parsing fungal community datasets by ecological guild. Fungal Ecol. 2016, 20, 241–248. [Google Scholar] [CrossRef]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2021; Available online: https://www.R-project.org/ (accessed on 1 November 2021).
- Oksanen, J.; Kindt, R.; Legendre, P.; O’Hara, B.; Stevens, M.H.H.; Oksanen, M.J.; Suggests, M. The Vegan Package. Community Ecol. Package 2007, 10, 631–637. [Google Scholar]
- Baber, K.; Otto, P.; Kahl, T.; Gossner, M.M.; Wirth, C.; Gminder, A.; Bässler, C. Disentangling the effects of forest-stand type and dead-wood origin of the early successional stage on the diversity of wood-inhabiting fungi. For. Ecol. Manag. 2016, 177, 161–169. [Google Scholar] [CrossRef]
- Zhou, L.W.; Hao, Z.Q.; Wang, Z.; Wang, B.; Dai, Y.C. Comparison of ecological patterns of polypores in three forest zones in China. Mycology 2011, 2, 260–275. [Google Scholar]
- Zhou, L.-W.; Dai, Y.-C. Recognizing ecological patterns of wood-decaying polypores on gymnosperm and angiosperm trees in Northeast China. Fungal Ecol. 2012, 5, 230–235. [Google Scholar] [CrossRef]
- Watkinson, S.C. Molecular Ecology. In The Fungi, 3rd ed.; Elsevier: Amsterdam, The Netherlands, 2016. [Google Scholar]
- Bader, P.; Jansson, S.; Jonsson, B. Wood-inhabiting fungi and substratum decline in selectively logged boreal spruce forests. Biol. Conserv. 1995, 72, 355–362. [Google Scholar] [CrossRef]
- Junninen, K.; Similä, M.; Kouki, J.; Kotiranta, H. Assemblages of wood-inhabiting fungi along the gradients of succession and naturalness in boreal pine-dominated forests in Fennoscandia. Ecography 2006, 29, 75–83. [Google Scholar] [CrossRef]
- Pellitier, P.T.; Zak, D.R.; Salley, S. Environmental filtering structures fungal endophyte communities in tree bark. Mol. Ecol. 2019, 28, 5188–5198. [Google Scholar] [CrossRef]
- Birch, J.D.; Lutz, J.A.; Turner, B.L.; Karst, J. Divergent, age-associated fungal communities of Pinus flexilis and Pinus longaeva. For. Ecol. Manag. 2021, 494, 119277. [Google Scholar] [CrossRef]
- Frey, S.D.; Elliott, E.T.; Paustian, K. Fungal translocation as a mechanism for soil nitrogen inputs to surface residue decompo-sition in a no-tillage agroecosystem. Soil Biol. Biochem. 2000, 32, 689–698. [Google Scholar] [CrossRef]
- Frey, S.; Six, J.; Elliott, E. Reciprocal transfer of carbon and nitrogen by decomposer fungi at the soil–litter interface. Soil Biol. Biochem. 2003, 35, 1001–1004. [Google Scholar] [CrossRef]
- Arnstadt, T.; Hoppe, B.; Kahl, T.; Kellner, H.; Krüger, D.; Bauhus, J.; Hofrichter, M. Dynamics of fungal community composi-tion, decomposition and resulting dead-wood properties in logs of Fagus sylvatica, Picea abies and Pinus sylvestris. For. Ecol. Manag. 2016, 382, 129–142. [Google Scholar] [CrossRef]


| Type | Condition | Collecting Height (cm) | Notes |
|---|---|---|---|
| I | Living tree | 150–170 | Bark was fresh and healthy. Collected clear ones from four different points and mixed. |
| II | Dead tree | 150–170 | Bark was broken partially. Collected clear ones from four different points and mixed. |
| III | Impending log | 100–120 | Bark kept perfectly. Collected clear ones from four different points and mixed. |
| IV | Impending log | 50–60 | Bark was broken partially. Some ones dropped to the soil surface and touched soil. Collected the untouched parts from four different points and mixed. |
| V | Lying log closest to ground | 0–3 | Bark was broken completely and dropped to the soil. Collected four parts close to soil and mixed. |
| ASVs | Type-Site | N | P | C | Pp | Wc | pH | H_cm | Ecological Function |
|---|---|---|---|---|---|---|---|---|---|
| Chaetosphaeriaceae | I-HSP | −0.42027 ** | −0.51154 *** | −0.10519 | −0.00214 | −0.36901 ** | −0.59416 *** | 0.58404 *** | Plant Saprotroph-Wood Saprotroph |
| Chloridium sp. | V-HSP | 0.34847 ** | 0.25285 | 0.1853 | −0.09486 | 0.39404 ** | 0.34321 ** | −0.36782 ** | Ectomycorrhizal |
| Collophora sp. | IV-DYD | 0.62952 *** | 0.58176 *** | 0.35379 ** | −0.28265 * | 0.5831 *** | 0.52576 *** | −0.4399 *** | Plant Pathogen |
| Devriesiastrelitziicola Arzanlou & Crous | I-HSP | −0.58717 *** | −0.65799 *** | −0.09294 | 0.05523 | −0.58317 *** | −0.58236 *** | 0.7771 *** | Plant Pathogen |
| Engyodontiumalbum (Limber) de Hoog | I-HSP | −0.37785 ** | −0.28339 * | −0.04966 | 0.35069 ** | −0.19738 | −0.22208 | 0.39015 ** | Animal Pathogen |
| Eucasphaeriacapensis Crous | IV-DYD | 0.34649 ** | 0.34512 ** | −0.14348 | −0.16996 | 0.33915 * | 0.18387 | −0.13009 | endophytic fungi |
| Ganoderma sp. | IV-DYD | 0.29567 * | 0.30706 * | 0.09148 | −0.03686 | 0.4461 *** | 0.21728 | −0.21755 | Wood Saprotroph |
| Gyoerffyellaentomobryoides (Boerema & Arx) Marvanová | V-HSP | 0.39435 ** | 0.53284 *** | −0.01823 | 0.1202 | 0.40469 ** | 0.65847 *** | −0.71019 *** | Undefined Saprotroph |
| Holtermanniellatakashimae Wuczk., Passoth, A.-C. Andersson, Turchetti, Prillinger, Boekhout & Libkind | IV-LSH | 0.32572 * | 0.3815 ** | −0.00476 | 0.20557 | 0.43039 *** | 0.37396 ** | −0.63874 *** | Undefined Saprotroph |
| Hymenochaetefuliginosa (Fr.) Lév. | I-DYD | −0.08828 | −0.31073 * | 0.25099 | −0.12912 | 0.0177 | −0.26346 * | 0.26839 * | Wood Saprotroph |
| Hymenochaete fuliginosa (Fr.) Lév. | II-LSH | −0.11769 | −0.15627 | 0.28638 * | −0.00318 | −0.23187 | −0.31254 * | 0.36307 ** | Wood Saprotroph |
| Kuraishiamolischiana Dlauchy, G. Péter, Tornai-Leh. & Kurtzman | II-LSH | −0.32065 * | −0.17142 | −0.48064 *** | 0.20931 | −0.33462 * | −0.3276 * | 0.07798 | Undefined Saprotroph |
| Mariannaeasamuelsii Seifert & Bissett | III-DYD | 0.41535 ** | 0.38432 ** | 0.20207 | −0.09156 | 0.57932 *** | 0.32385 * | −0.38355 ** | Undefined Saprotroph |
| Naganishiacerealis (Passoth, A.-C. Andersson, Olstorpe, Theelen, Boekhout & Schnürer) Xin Zhan Liu, F.Y. Bai, M. Groenew. & Boekhout | IV-LSH | 0.32281 * | 0.35058 ** | 0.01399 | 0.14483 | 0.29027 * | 0.1936 | −0.67137 *** | Undefined Saprotroph |
| Nakazawaea holstii (Wick.) Y. Yamada, K. Maeda & Mikata | IV-LSH | 0.33259 * | 0.37819 ** | −0.15547 | 0.07405 | 0.31148 * | 0.2894 * | −0.42209 ** | Undefined Saprotroph |
| Peterozyma toletana (Socias, C. Ramírez & Peláez) Kurtzman & Robnett | IV-DYD | 0.51026 *** | 0.58943 *** | 0.19928 | −0.07098 | 0.48773 *** | 0.52501 *** | −0.34207 ** | Undefined Saprotroph |
| Pezicula sp. | IV-DYD | 0.3436 ** | 0.33559 * | −0.02744 | −0.12116 | 0.43092 *** | 0.30086 * | −0.34954 ** | Undefined Saprotroph |
| Phialocephala fusca W.B. Kendr. | I-DYD | −0.02463 | −0.25659 | 0.37366 ** | −0.14357 | 0.01675 | −0.33018 * | 0.39015 ** | Endophyte |
| Phialophora japonica Iwatsu & Udagawa | I-DYD | −0.14267 | −0.3641 ** | 0.22505 | −0.01393 | −0.08506 | −0.32705 * | 0.31275 * | Plant Pathogen |
| Phlebiopsis gigantea (Fr.) Jülich | V-HSP | 0.28512 * | 0.33356 * | −0.04281 | −0.18003 | 0.2533 | 0.24777 | −0.21439 | Wood Saprotroph |
| Pseudoteratosphaeria sp. | I-DYD | −0.39898 ** | −0.53475 *** | −0.004 | −0.11655 | −0.48064 *** | −0.52299 *** | 0.62296 *** | Plant Pathogen |
| Sakaguchia lamellibrachiae (Nagah., Hamam., Nakase & Horikoshi) Q.M. Wang, F.Y. Bai, M. Groenew. & Boekhout | IV-LSH | 0.58176 *** | 0.65954 *** | −0.01538 | −0.00391 | 0.54981 *** | 0.52497 *** | −0.667 *** | Undefined Saprotroph |
| Serpulahimantioides (Fr.) P. Karst. | IV-LSH | 0.04183 | 0.12973 | −0.32163 * | 0.05626 | 0.06251 | −0.10271 | −0.23728 | Wood Saprotroph |
| Sistotrema sp. | III-D | 0.1768 | 0.14627 | 0.36528 ** | −0.09784 | 0.10289 | −0.03372 | 0.09446 | Wood Saprotroph |
| Teratosphaeriaencephalarti Crous & A.R. Wood | I-LSH | −0.52565 *** | −0.62286 *** | −0.16118 | −0.03546 | −0.42869 *** | −0.59112 *** | 0.68998 *** | Plant Pathogen |
| Teratosphaeriaceae | II-LSH | −0.34933 ** | −0.37335 ** | 0.07149 | 0.00367 | −0.43903 *** | −0.43732 *** | 0.44238 *** | Animal Pathogen-Plant Pathogen |
| Tremella sp. | V-DYD | 0.34527 ** | 0.36954 ** | 0.37419 ** | −0.10019 | 0.36161 ** | 0.32992 * | −0.29751 * | Fungal Parasite |
| Tremellales | V-HSP | 0.37249 ** | 0.46203 *** | −0.10606 | −0.10004 | 0.26532 * | 0.22017 | −0.48118 *** | Fungal Parasite-Undefined Saprotroph |
| Trichaptumabietinum (Pers. ex J.F. Gmel.) Ryvarden | III-DYD | 0.25117 | 0.23961 | 0.30346 * | −0.13052 | 0.20463 | 0.08076 | 0.02301 | Wood Saprotroph |
| Trichoderma sp. | III-DYD | 0.3238 * | 0.31916 * | 0.3793 ** | −0.13352 | 0.31134 * | 0.18385 | −0.21448 | Wood Saprotroph |
| Trichodermaatroviride P. Karst. | V-HSP | −0.25694 | −0.17028 | −0.16304 | 0.29729 * | −0.10514 | 0.09028 | −0.06518 | Wood Saprotroph |
| Capnodiales | I-LSH | −0.52565 *** | −0.62286 *** | −0.16118 | −0.03546 | −0.42869 *** | −0.59112 *** | 0.68998 *** | Animal Pathogen-Plant Pathogen |
| Venturia sp. | V-HSP | 0.26284 | 0.31145 * | −0.09667 | 0.15266 | 0.28038 * | 0.35911 ** | −0.37968 ** | Plant Pathogen |
| Xenopolyscytalum pinea Crous | II-HSP | −0.15217 | 0.02994 | 0.02655 | 0.37774 ** | −0.02524 | 0.20164 | −0.142 | Undefined Saprotroph |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wei, Y.-L.; Li, Q.-S.; Bai, Z.; Wu, Q.-X. Fungal Community Composition and Enzyme Activity in Different Type Bark of Pinus koraiensis. Forests 2021, 12, 1781. https://doi.org/10.3390/f12121781
Wei Y-L, Li Q-S, Bai Z, Wu Q-X. Fungal Community Composition and Enzyme Activity in Different Type Bark of Pinus koraiensis. Forests. 2021; 12(12):1781. https://doi.org/10.3390/f12121781
Chicago/Turabian StyleWei, Yu-Lian, Qiu-Shi Li, Zhen Bai, and Qing-Xue Wu. 2021. "Fungal Community Composition and Enzyme Activity in Different Type Bark of Pinus koraiensis" Forests 12, no. 12: 1781. https://doi.org/10.3390/f12121781
APA StyleWei, Y.-L., Li, Q.-S., Bai, Z., & Wu, Q.-X. (2021). Fungal Community Composition and Enzyme Activity in Different Type Bark of Pinus koraiensis. Forests, 12(12), 1781. https://doi.org/10.3390/f12121781






