Potential Link between Ectomycorrhizal Community Composition and Host Tree Phenology
Abstract
1. Introduction
2. Materials and Methods
2.1. Study Site and Phenological Observations
2.2. Mycorrhizal Sampling
2.3. Molecular Analysis
2.4. Statistical Analyses
3. Results
3.1. Ectomycorrhizal Community Composition Changes along Transition between Phenological Stages
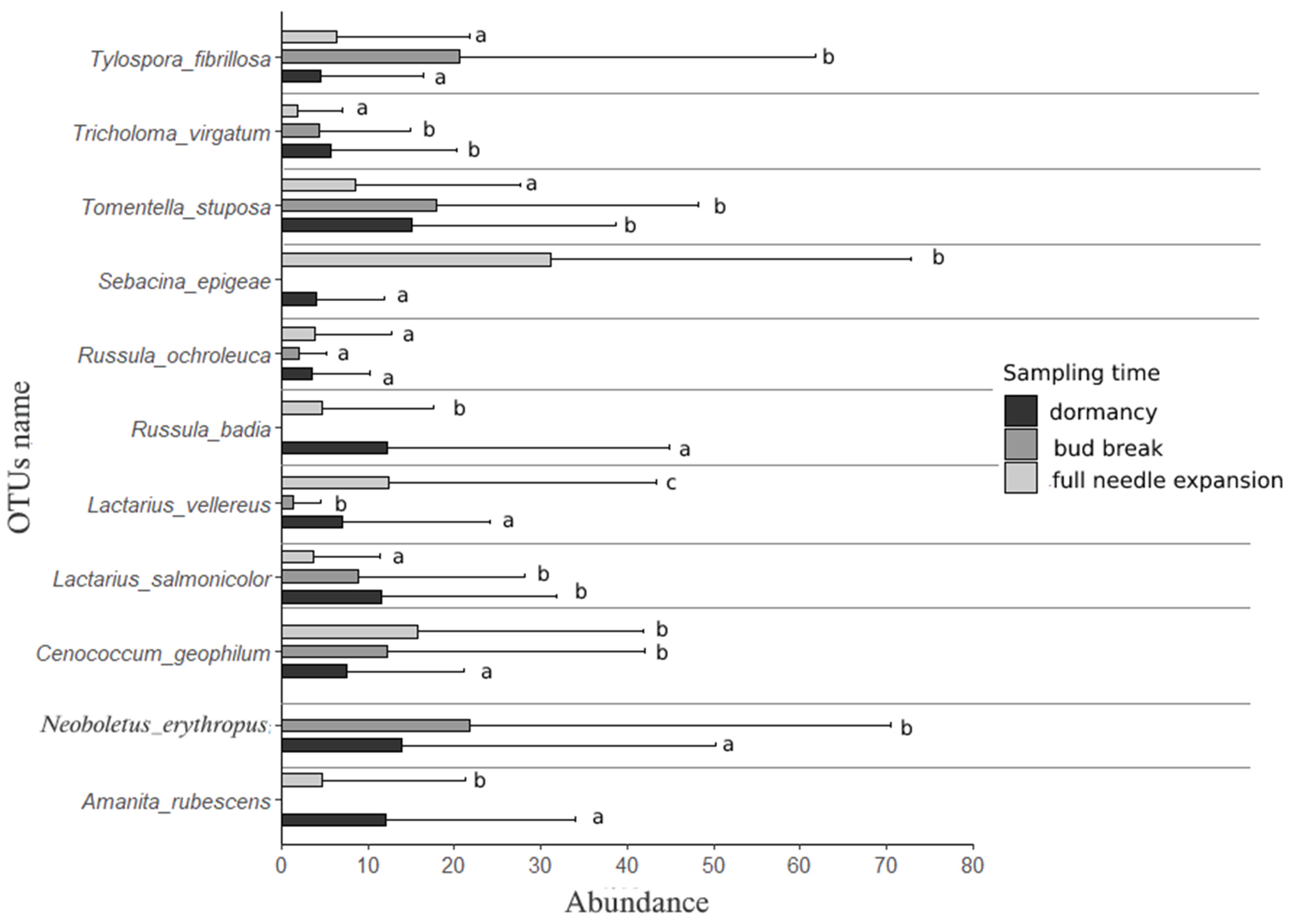
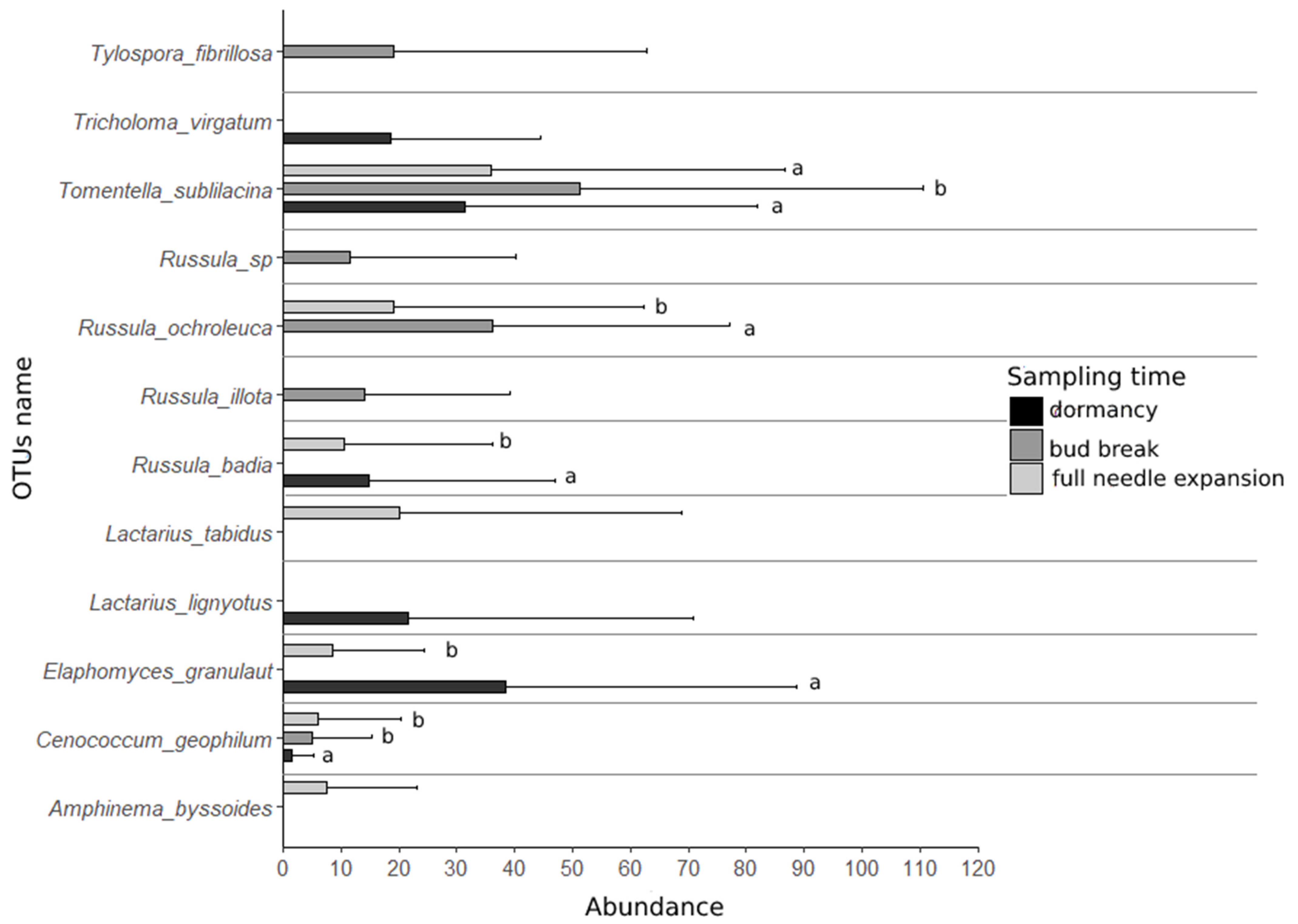
3.2. Individual Responses of Ectomycorrhizal OTUs along Transition between Phenological Stages
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Ważny, R. Ectomycorrhizal communities associated with silver fir seedlings (Abies alba Mill.) differ largely in mature silver fir stands and in Scots pine forecrops. Ann. For. Sci. 2014, 71, 801–810. [Google Scholar] [CrossRef]
- Rudawska, M.; Pietras, M.; Smutek, I.; Strzeliński, P.; Leski, T. Ectomycorrhizal fungal assemblages of Abies alba Mill. outside its native range in Poland. Mycorrhiza 2016, 26, 57–65. [Google Scholar] [CrossRef]
- Vitasse, Y.; Delzon, S.; Bresson, C.C.; Michalet, R.; Kremer, A. Altitudinal differentiation in growth and phenology among populations of temperate-zone tree species growing in a common garden. Can. J. For. Res. 2009, 39, 1259–1269. [Google Scholar] [CrossRef]
- Vitasse, Y.; Delzon, S.; Dufrêne, E.; Pontailler, J.Y.; Louvet, J.M.; Kremer, A.; Michalet, R. Leaf phenology sensitivity to temperature in European trees: Do within-species populations exhibit similar responses? Agric. For. Meteorol. 2009, 149, 735–744. [Google Scholar] [CrossRef]
- Arend, M.; Brem, A.; Kuster, T.M.; Günthardt-Goerg, M.S. Seasonal photosynthetic responses of European oaks to drought and elevated daytime temperature. Plant Biol. 2013, 15, 169–176. [Google Scholar] [CrossRef] [PubMed]
- Delpierre, N.; Vitasse, Y.; Chuine, I.; Guillemot, J.; Bazot, S.; Rutishauser, T.; Rathgeber, C.B.K. Temperate and boreal forest tree phenology: From organ-scale processes to terrestrial ecosystem models. Ann. For. Sci. 2016, 73, 5–25. [Google Scholar] [CrossRef]
- Dickie, I.A.; Montgomery, R.A.; Reich, P.B.; Schnitzer, S.A. Physiological and phenological responses of oak seedlings to oak forest soil in the absence of trees. Tree Physiol. 2007, 27, 133–140. [Google Scholar] [CrossRef][Green Version]
- Unuk, T.; Martinović, T.; Finžgar, D.; Šibanc, N.; Grebenc, T.; Kraigher, H. Root-associated fungal communities from two phenologically contrasting silver fir (Abies alba mill.) groups of trees. Front. Plant Sci. 2019, 10, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Allen, B.Y.E.B.; Allen, M.E. Water relations of xeric grasses in the field: Interactions of mycorrhizas and competition. New Phytol. 1986, 104, 559–571. [Google Scholar] [CrossRef]
- Garbaye, J. Garbaye_citation. In Mycorrhizae: Physiology and Genetics-Les Mycorrhizes: Physiologie et Genetique; Gianinazzi-Pearson, V., Gianinazzi, S., Eds.; INRA: Paris, France, 1986; pp. 493–496. [Google Scholar]
- Courty, P.E.; Bréda, N.; Garbaye, J. Relation between oak tree phenology and the secretion of organic matter degrading enzymes by Lactarius quietus ectomycorrhizas before and during bud break. Soil Biol. Biochem. 2007, 39, 1655–1663. [Google Scholar] [CrossRef]
- Waldrop, M.P.; Firestone, M.K. Seasonal dynamics of microbial community composition and function in oak canopy and open grassland soils. Microb. Ecol. 2006, 52, 470–479. [Google Scholar] [CrossRef]
- Dhuli, P.; Rohloff, J.; Strimbeck, G.R. Metabolite changes in conifer buds and needles during forced bud break in Norway spruce (Picea abies) and European silver fir (Abies alba). Front. Plant Sci. 2014, 5, 706. [Google Scholar] [CrossRef] [PubMed]
- Walker, J.F.; Miller, O.K.; Horton, J.L. Seasonal dynamics of ectomycorrhizal fungus assemblages on oak seedlings in the southeastern Appalachian Mountains. Mycorrhiza 2008, 18, 123–132. [Google Scholar] [CrossRef]
- Bajc, M.; Aravanopoulos, F.; Westergren, M.; Fussi, B.; Kavaliauskas, D.; Alizoti, P.; Kraigher, H. (Eds.) Manual for Forest Genetic Monitoring; Silva Slovenica Publishing Centre: Ljubljana, Slovenia, 2020. [Google Scholar]
- Kraigher, H. Tipi ektomikorize-taksonomija, pomen in aplikacihe-Types of ectomycorrhizae-their taxonomy, role and applocation. Acta Silvae Ligni 1996, 49, 33–66. [Google Scholar]
- Taylor, D.L.; Bruns, T.D. Community structure of ectomycorrhizal fungi in a Pinus muricata forest: Minimal overlap between the mature forest and resistant propagule communities. Mol. Ecol. 1999, 8, 1837–1850. [Google Scholar] [CrossRef]
- Baar, J.; Horton, T.R.; Kretzer, A.M.; Bruns, T.D. Mycorrhizal colonization of Pinus muricata from resistant propagules after a stand-replacing wildfire. New Phytol. 1999, 143, 409–418. [Google Scholar] [CrossRef]
- Železnik, P.; Hrenko, M.; Then, C.; Koch, N.; Grebenc, T.; Levanič, T.; Kraigher, H. CASIROZ: Root parameters and types of ectomycorrhiza of young beech plants exposed to different ozone and light regimes. Plant Biol. 2007, 9, 298–308. [Google Scholar] [CrossRef]
- Chen, W.; Koide, R.T.; Adams, T.S.; DeForest, J.L.; Cheng, L.; Eissenstat, D.M. Root morphology and mycorrhizal symbioses together shape nutrient foraging strategies of temperate trees. Proc. Natl. Acad. Sci. USA 2016, 113, 8741–8746. [Google Scholar] [CrossRef]
- Železnik, P.; Vilhar, U.; Starr, M.; de Groot, M.; Kraigher, H. Fine root dynamics in Slovenian beech forests in relation to soil temperature and water availability. Trees Struct. Funct. 2016, 30, 375–384. [Google Scholar] [CrossRef][Green Version]
- Agerer, R. Characterization of Ectomycorrhiza. Methods Microbiol. 1991, 23, 25–73. [Google Scholar] [CrossRef]
- Gardes, M.; Bruns, T.D. ITS primers with enhanced specificity for basidiomycetes-application to the identification of mycorrhizae and rusts. Mol. Ecol. 1993, 2, 113–118. [Google Scholar] [CrossRef]
- White, T.J.; Bruns, T.; Lee, S.; Taylor, J. Amplification and direct sequencing of fungal ribosomal rna genes for phylogenetics. In PCR Protocols; Elsevier: Amsterdam, The Netherlands, 1990; pp. 315–322. [Google Scholar]
- Sulzbacher, M.A.; Grebenc, T.; Cabral, T.S.; Giachini, A.J.; Goto, B.T.; Smith, M.E.; Baseia, I.G. Restingomyces, a new sequestrate genus from the Brazilian Atlantic rainforest that is phylogenetically related to early-diverging taxa in Trappeaceae (Phallales). Mycologia 2016, 108, 954–966. [Google Scholar] [CrossRef] [PubMed]
- Kearse, M.; Moir, R.; Wilson, A.; Stones-Havas, S.; Cheung, M.; Sturrock, S.; Buxton, S.; Cooper, A.; Markowitz, S.; Duran, C.; et al. Geneious Basic: An integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 2012, 28, 1647–1649. [Google Scholar] [CrossRef]
- Nilsson, R.H.; Larsson, K.H.; Taylor, A.F.S.; Bengtsson-Palme, J.; Jeppesen, T.S.; Schigel, D.; Kennedy, P.; Picard, K.; Glöckner, F.O.; Tedersoo, L.; et al. The UNITE database for molecular identification of fungi: Handling dark taxa and parallel taxonomic classifications. Nucleic Acids Res. 2019, 47, D259–D264. [Google Scholar] [CrossRef] [PubMed]
- Porras-Alfaro, A.; Liu, K.L.; Kuske, C.R.; Xiec, G. From genus to phylum: Large-subunit and internal transcribed spacer rRNA operon regions show similar classification accuracies influenced by database composition. Appl. Environ. Microbiol. 2014, 80, 829–840. [Google Scholar] [CrossRef]
- Raja, H.A.; Miller, A.N.; Pearce, C.J.; Oberlies, N.H. Fungal Identification Using Molecular Tools: A Primer for the Natural Products Research Community. J. Nat. Prod. 2017, 80, 756–770. [Google Scholar] [CrossRef]
- Rinaldi, A.C.; Comandini, O.; Kuyper, T.W. Ectomycorrhizal fungal diversity: Separating the wheat from the chaff. Fungal Divers. 2008, 33, 1–45. [Google Scholar]
- Tedersoo, L.; May, T.W.; Smith, M.E. Ectomycorrhizal lifestyle in fungi: Global diversity, distribution, and evolution of phylogenetic lineages. Mycorrhiza 2010, 20, 217–263. [Google Scholar] [CrossRef] [PubMed]
- R Core Team. R: A Language and Environment for Statistical Computing; R Core Team: Vienna, Austria, 2016. [Google Scholar]
- Oksanen, J.F.; Blanchet, G.; Friendly, M.; Kindt, R.; Legendre, P.; McGlinn, D.; Minchin, P.R.; O’Hara, R.B.; Simpson, G.L.; Solymos, P.; et al. Vegan: Community Ecology Package. 2018. Available online: https://github.com/vegandevs/vegan (accessed on 3 September 2019).
- Wang, Y.; Naumann, U.; Wright, S.T.; Warton, D.I. Mvabund—An R package for model-based analysis of multivariate abundance data. Methods Ecol. Evol. 2012, 3, 471–474. [Google Scholar] [CrossRef]
- Wickham, H. Package ‘ggplot2’: Elegant Graphics for Data Analysis; Springer: New York, NY, USA, 2016. [Google Scholar] [CrossRef]
- Hothorn, T.; Bretz, F.; Westfall, P. Simultaneous inference in general parametric models. Biom. J. 2008, 50, 346–363. [Google Scholar] [CrossRef]
- Wickham, H. Reshaping data with the reshape package. J. Stat. Softw. 2007, 21, 1–20. [Google Scholar] [CrossRef]
- Cremer, E. Population genetics of silver fir (Abies alba Mill.) in the Northern Black Forest–preconditions for the recolonization of windthrow areas and associated ectomycorrhizal communities. Phillips-Universität Marbg. 2009, 1, 103. [Google Scholar] [CrossRef]
- Schirkonyer, U.; Bauer, C.; Rothe, G.M. Ectomycorrhizal diversity at five different tree species in forests of the Taunus Mountains in Central Germany. Open J. Ecol. 2013, 3, 66–81. [Google Scholar] [CrossRef]
- Unuk Nahberger, T. Ektomikorizni Simbionti Bele Jelke (Abies alba Mill.); Univerza v Ljubljani, Biotehniška Fakulteta: Ljubljana, Slovenia, 2020. [Google Scholar]
- Hupperts, S.F.; Karst, J.; Pritsch, K.; Landhäusser, S.M. Host phenology and potential saprotrophism of ectomycorrhizal fungi in the boreal forest. Funct. Ecol. 2017, 31, 116–126. [Google Scholar] [CrossRef]
- Helmisaari, H.-S.; Ostonen, I.; Lohmus, K.; Derome, J.; Lindroos, A.-J.; Merila, P.; Nojd, P. Ectomycorrhizal root tips in relation to site and stand characteristics in Norway spruce and Scots pine stands in boreal forests. Tree Physiol. 2009, 29, 445–456. [Google Scholar] [CrossRef]
- Tierney, G.L.; Fahey, T.J.; Groffman, P.M.; Hardy, J.P.; Fitzhugh, R.D.; Driscoll, C.T. Soil freezing alters fine root dynamics in a northern hardwood forest. Biogeochemistry 2001, 56, 175–190. [Google Scholar] [CrossRef]
- Hendrick, R.L.; Pregitzer, K.S. Temporal and Depth-Related Patterns of Fine Root Dynamics in Northern Hardwood Forests. J. Ecol. 1996, 84, 167. [Google Scholar] [CrossRef]
- Fahey, T.J.; Hughes, J.W. Fine Root Dynamics in a Northern Hardwood Forest Ecosystem, Hubbard Brook Experimental Forest, NH. J. Ecol. 1994, 82, 533. [Google Scholar] [CrossRef]
- Harley, J.L. Ecology of ectotrophic mycorrhizas. In The Biology of Mycorrhiza; Polunin, N., Ed.; Leonard Hill: London, UK, 1969; pp. 150–162. [Google Scholar]
- Rygiewicz, P.T.; Johnson, M.G.; Ganio, L.M.; Tingey, D.T.; Storm, M.J. Lifetime and temporal occurrence of ectomycorrhizae on ponderosa pine (Pinus ponderosa Laws.) seedlings grown under varied atmospheric CO2 and nitrogen levels. Plant Soil 1997, 189, 275–287. [Google Scholar] [CrossRef]
- Majdi, H.; Damm, E.; Nylund, J.-E. Longevity of mycorrhizal roots depends on branching order and nutrient availability. New Phytol. 2001, 150, 195–202. [Google Scholar] [CrossRef]
- Fernandez, C.W.; McCormack, M.L.; Hill, J.M.; Pritchard, S.G.; Koide, R.T. On the persistence of Cenococcum geophilum ectomycorrhizas and its implications for forest carbon and nutrient cycles. Soil Biol. Biochem. 2013, 65, 141–143. [Google Scholar] [CrossRef]
- Kou, L.; McCormack, M.L.; Chen, W.; Guo, D.; Wang, H.; Gao, W.; Yang, H.; Li, S. Nitrogen ion form and spatio-temporal variation in root distribution mediate nitrogen effects on lifespan of ectomycorrhizal roots. Plant Soil 2017, 411, 261–273. [Google Scholar] [CrossRef]
- Cullings, K.; Ishkhanova, G.; Henson, J. Defoliation effects on enzyme activities of the ectomycorrhizal fungus Suillus granulatus in a Pinus contorta (lodgepole pine) stand in Yellowstone National Park. Oecologia 2008, 158, 77–83. [Google Scholar] [CrossRef]
- Talbot, J.M.; Allison, S.D.; Treseder, K.K. Decomposers in disguise: Mycorrhizal fungi as regulators of soil C dynamics in ecosystems under global change. Funct. Ecol. 2008, 22, 955–963. [Google Scholar] [CrossRef]
- Bödeker, I.T.M.; Nygren, C.M.R.; Taylor, A.F.S.; Olson, Å.; Lindahl, B.D. ClassII peroxidase-encoding genes are present in a phylogenetically wide range of ectomycorrhizal fungi. ISME J. 2009, 3, 1387–1395. [Google Scholar] [CrossRef] [PubMed]
- Lindahl, B.D.; Tunlid, A. Ectomycorrhizal fungi-potential organic matter decomposers, yet not saprotrophs. New Phytol. 2015, 205, 1443–1447. [Google Scholar] [CrossRef] [PubMed]
- Hibbett, D.S.; Gilbert, L.; Donoghue, M.J. Evolutionary instability of ectomycorrhizal symbioses in basidiomycetes. Nat. Cell Biol. 2000, 407, 506–508. [Google Scholar] [CrossRef]
- Chen, D.M.; Taylor, A.F.S.; Burke, R.M.; Cairney, J.W.G. Identification of genes for lignin peroxidases and manganese peroxidases in ectomycorrhizal fungi. New Phytol. 2001, 152, 151–158. [Google Scholar] [CrossRef]
- Chen, D.M.; Bastias, B.A.; Taylor, A.F.S.; Cairney, J.W.G. Identification of laccase-like genes in ectomycorrhizal basidiomycetes and transcriptional regulation by nitrogen in Piloderma byssinum. New Phytol. 2003. [Google Scholar] [CrossRef]
- Read, D.J.; Perez-Moreno, J. Mycorrhizas and nutrient cycling in ecosystems-A journey towards relevance? New Phytol. 2003, 157, 475–492. [Google Scholar] [CrossRef]
- Burke, R.M.; Cairney, J.W.G. Laccases and other polyphenol oxidases in ecto- and ericoid mycorrhizal fungi. Mycorrhiza 2002, 12, 105–116. [Google Scholar] [CrossRef]
- Chambers, S.M.; Burke, R.M.; Brooks, P.R.; Cairney, J.W.G. Molecular and biochemical evidence for manganese-dependent peroxidase activity in Tylospora fibrillosa. Mycol. Res. 1999, 103, 1098–1102. [Google Scholar] [CrossRef]
- Martin, F.; Kohler, A.; Murat, C.; Veneault-Fourrey, C.; Hibbett, D.S. Unearthing the roots of ectomycorrhizal symbioses. Nat. Rev. Microbiol. 2016, 14, 760–773. [Google Scholar] [CrossRef]
- Koide, R.T.; Shumway, D.L.; Xu, B.; Sharda, J.N. On temporal partitioning of a community of ectomycorrhizal fungi. New Phytol. 2007, 174, 420–429. [Google Scholar] [CrossRef] [PubMed]
- Aponte, C.; Marañón, T.; García, L.V. Microbial C, N and P in soils of Mediterranean oak forests: Influence of season, canopy cover and soil depth. Biogeochemistry 2010, 101, 77–92. [Google Scholar] [CrossRef]
- Richard, F.; Roy, M.; Shahin, O.; Sthultz, C.; Duchemin, M.; Joffre, R.; Selosse, M.A. Ectomycorrhizal communities in a Mediterranean forest ecosystem dominated by Quercus ilex: Seasonal dynamics and response to drought in the surface organic horizon. Ann. For. Sci. 2011, 68, 57–68. [Google Scholar] [CrossRef]
- Vořiškova, J.; Brabcová, V.; Cajthaml, T.; Baldrian, P. Seasonal dynamics of fungal communities in a temperate oak forest soil. New Phytol. 2014, 201, 269–278. [Google Scholar] [CrossRef]
- Chen, M.M.; Zhu, Y.G.; Su, Y.H.; Chen, B.D.; Fu, B.J.; Marschner, P. Effects of soil moisture and plant interactions on the soil microbial community structure. Eur. J. Soil Biol. 2007, 43, 31–38. [Google Scholar] [CrossRef]
- Schimel, J.P.; Gulledge, J.M.; Clein-Curley, J.S.; Lindstrom, J.E.; Braddock, J.F. Moisture effects on microbial activity and community structure in decomposing birch litter in the Alaskan taiga. Soil Biol. Biochem. 1999, 31, S0038–S0717. [Google Scholar] [CrossRef]
- Williams, M.A. Response of microbial communities to water stress in irrigated and drought-prone tallgrass prairie soils. Soil Biol. Biochem. 2007, 39, 2750–2757. [Google Scholar] [CrossRef]
- Erlandson, S.R.; Savage, J.A.; Cavender-Bares, J.M.; Peay, K.G. Soil moisture and chemistry influence diversity of ectomycorrhizal fungal communities associating with willow along an hydrologic gradient. FEMS Microbiol. Ecol. 2016, 92, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Hawkes, C.V.; Kivlin, S.N.; Rocca, J.D.; Huguet, V.; Thomsen, M.A.; Suttle, K.B. Fungal community responses to precipitation. Glob. Chang. Biol. 2011, 17, 1637–1645. [Google Scholar] [CrossRef]
- Castro, H.F.; Classen, A.T.; Austin, E.E.; Norby, R.J.; Schadt, C.W. Soil microbial community responses to multiple experimental climate change drivers. Appl. Environ. Microbiol. 2010, 76, 999–1007. [Google Scholar] [CrossRef]
- Bahram, M.; Põlme, S.; Kõljalg, U.; Zarre, S.; Tedersoo, L. Regional and local patterns of ectomycorrhizal fungal diversity and community structure along an altitudinal gradient in the Hyrcanian forests of northern Iran. New Phytol. 2012, 193, 465–473. [Google Scholar] [CrossRef] [PubMed]

| Phenophase|Sampling Time | Year 2016 | Year 2017 |
|---|---|---|
| Dormancy | 16 March | 20 March |
| Bud bursting | 16 May | 28 April |
| Full needle expansion | 15 June | 25 May |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Unuk Nahberger, T.; Damjanič, R.; Kraigher, H.; Grebenc, T. Potential Link between Ectomycorrhizal Community Composition and Host Tree Phenology. Forests 2021, 12, 1719. https://doi.org/10.3390/f12121719
Unuk Nahberger T, Damjanič R, Kraigher H, Grebenc T. Potential Link between Ectomycorrhizal Community Composition and Host Tree Phenology. Forests. 2021; 12(12):1719. https://doi.org/10.3390/f12121719
Chicago/Turabian StyleUnuk Nahberger, Tina, Rok Damjanič, Hojka Kraigher, and Tine Grebenc. 2021. "Potential Link between Ectomycorrhizal Community Composition and Host Tree Phenology" Forests 12, no. 12: 1719. https://doi.org/10.3390/f12121719
APA StyleUnuk Nahberger, T., Damjanič, R., Kraigher, H., & Grebenc, T. (2021). Potential Link between Ectomycorrhizal Community Composition and Host Tree Phenology. Forests, 12(12), 1719. https://doi.org/10.3390/f12121719








