PmMYB4, a Transcriptional Activator from Pinus massoniana, Regulates Secondary Cell Wall Formation and Lignin Biosynthesis
Abstract
:1. Introduction
2. Results
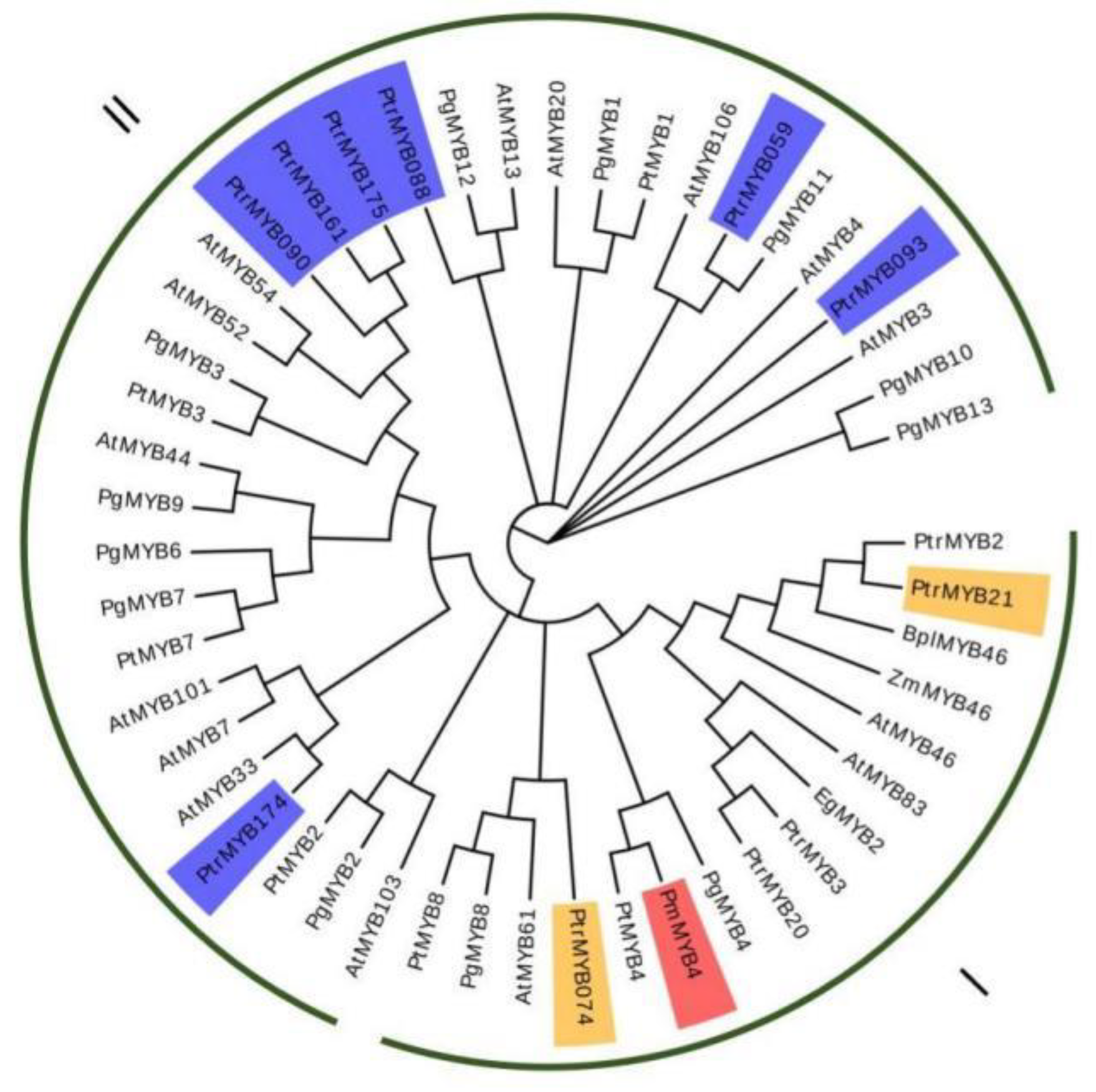
2.1. Isolation and Characterization of PmMYB4
2.2. PmMYB4 Is Primarily Expressed in Bark
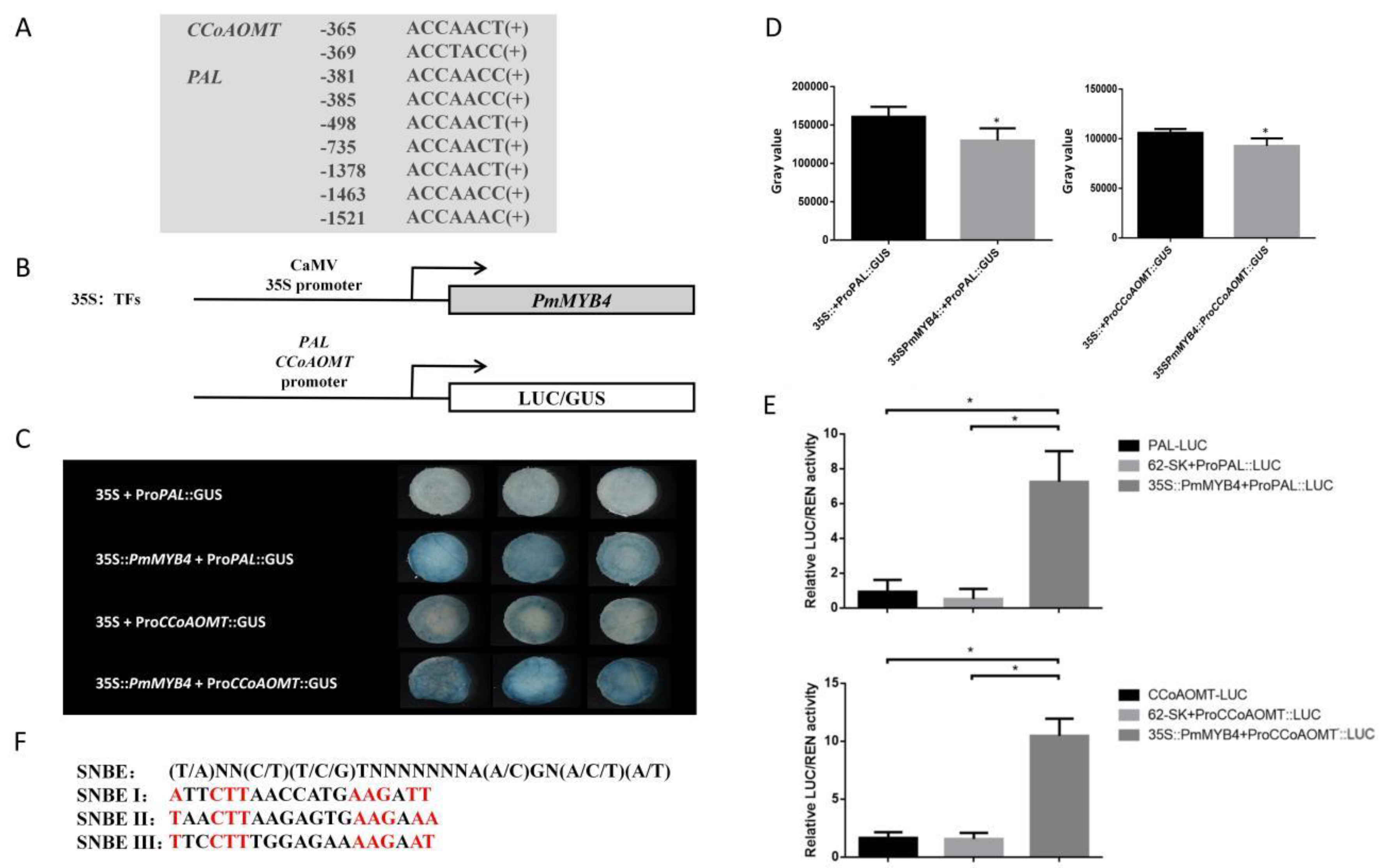
2.3. Transcriptional Activation of PmMYB4 and the Binding Ability of PmMYB4 with AC Elements
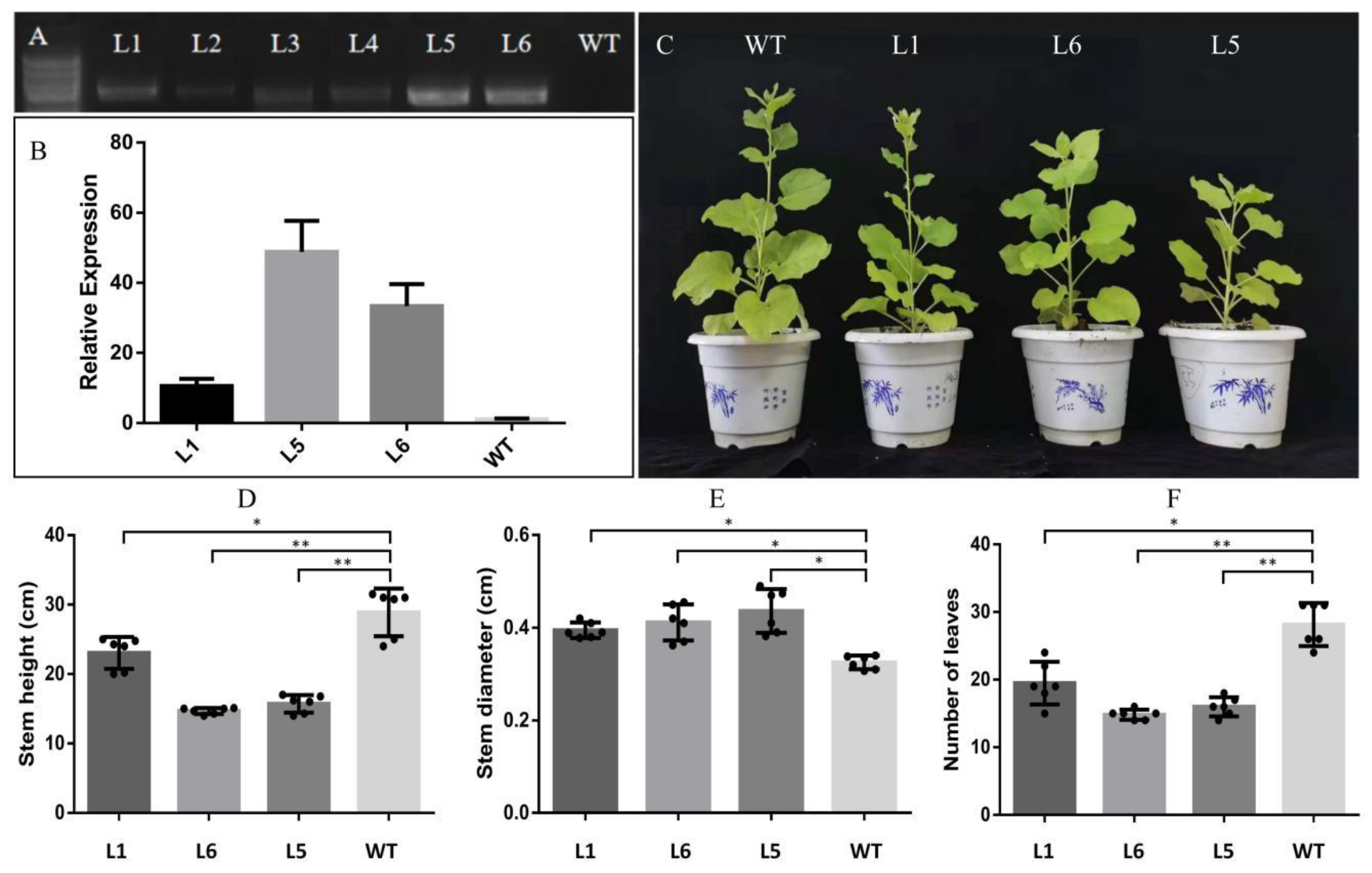
2.4. Growth and Morphological Characteristics of PmMYB4 Transgenic Tobacco
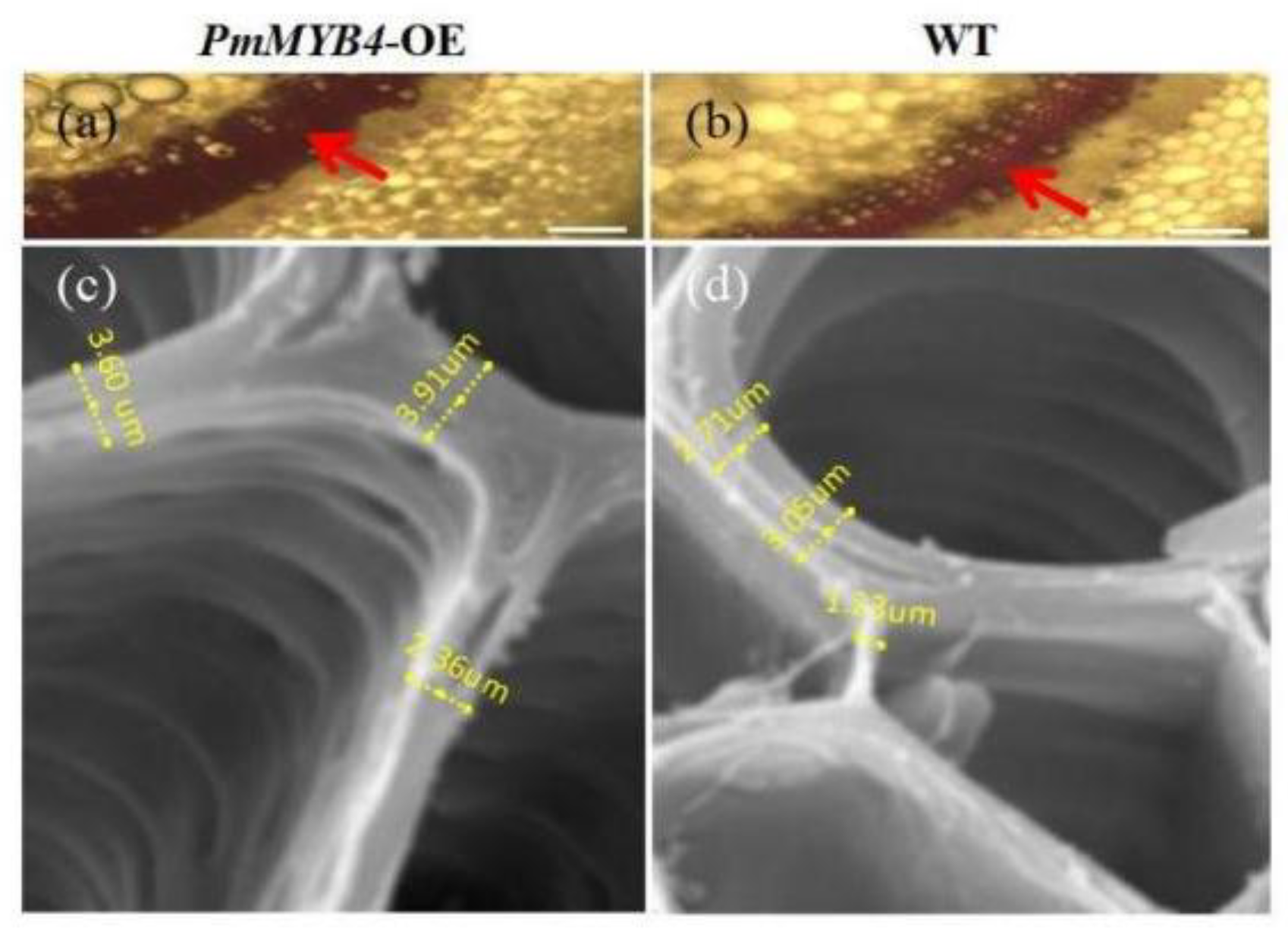
2.5. Overexpression of PmMYB4 Positively Regulates SCW in Transgenic Tobacco
2.6. Overexpression of PmMYB4 Affects the Expression of SCW Biosynthesis Genes in Transgenic Tobacco
2.7. The PmPAL and PmCCoAOMT Promoters Are Activated by PmMYB4
3. Discussion
4. Materials and Methods
4.1. Plant Materials and Stress Treatments
4.2. Cloning of the Full-Length PmMYB4 Coding Sequence (CDS) and Promoter
4.3. Sequence and Phylogenetic Analysis
4.4. Transcriptional Activation Analysis of PmMYB4
4.5. Semiquantitative RT-PCR and Quantitative Real-Time PCR
4.6. Agrobacterium-Mediated Transformation of Tobacco
4.7. Plant Height and Biomass Measurements
4.8. Microscopy and Histochemistry
4.9. Measurement of Lignin, Cellulose, and Hemicellulose Components in Transgenic Tobacco
4.10. Transient Expression and LUC/GUS Activity Assay
4.11. Statistical Analysis
4.12. GenBank Accession Numbers of Genes Used in This Study
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Plomion, C.; Leprovost, G.; Stokes, A. Wood formation in trees. Plant Physiol. 2001, 127, 1513–1523. [Google Scholar] [CrossRef]
- Blanch, H.W. Bioprocessing for biofuels. Curr. Opin. Biotechnol. 2012, 23, 390–395. [Google Scholar] [CrossRef] [PubMed]
- Cosgrove, D.J. Growth of the plant cell wall. Nat. Rev. Mol. Cell Biol. 2005, 6, 850–861. [Google Scholar] [CrossRef] [PubMed]
- Xu, C.Z. PtoMYB170 positively regulates lignin deposition during wood formation in poplar and confers drought tolerance in transgenic Arabidopsis. Tree Physiol. 2017, 37, 1713–1726. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- James, A.R.; William, R.A. Molecular evolution of the MYB family of transcription factors: Evidence for polyphyletic origin. J. Mol. Evol. 1998, 46, 74–83. [Google Scholar]
- Chen, H.; Wang, J.P.; Liu, H.Z.; Li, H.Y.; Lin, Y.C.; Shi, R.; Yang, C.M.; Gao, J.H.; Zhou, C.G.; Li, Q.Z.; et al. Hierarchical Transcription Factor and Chromatin Binding Network for Wood Formation in Populus trichocarpa. Plant Cell 2019, 31, 602–626. [Google Scholar] [CrossRef] [Green Version]
- Minoru, K.; Makiko, U.; Nobuyuki, N. Transcription switches for protoxylem and metaxylem vessel formation. Genes Dev. 2005, 19, 1855–1860. [Google Scholar]
- Zhong, R.Q.; Demura, T.; Ye, Z.H. SND1, a NAC domain transcription factor, is a key regulator of secondary wall synthesis in fibers of Arabidopsis. Plant Cell 2006, 18, 3158–3170. [Google Scholar] [CrossRef] [Green Version]
- Nobutaka, M.; Akira, I.; Hiroyuki, Y. NAC transcription factors, NST1 and NST3, are key regulators of the formation of secondary walls in woody tissues of Arabidopsis. Plant Cell 2007, 19, 270–280. [Google Scholar]
- De, R.B.; Mähönen, A.P.; Helariutta, Y. Plant vascular development: From early specification to differentiation. Nat. Rev. Mol. Cell Biol. 2016, 17, 30–40. [Google Scholar]
- Ko, J.H.; Kim, W.C.; Kim, J.Y.; Ahn, S.J.; Han, K.H. MYB46-Mediated Transcriptional Regulation of Secondary Wall Biosynthesis. Mol. Plant 2012, 5, 961–963. [Google Scholar] [CrossRef] [Green Version]
- McCarthy, R.L.; Zhong, R.; Ye, Z.H. MYB83 is a direct target of SND1 and acts redundantly with MYB46 in the regulation of secondary cell wall biosynthesis in Arabidopsis. Plant Cell Physiol. 2009, 50, 1950–1964. [Google Scholar] [CrossRef] [Green Version]
- Zhong, R.; Ye, Z.H. Regulation of cell wall biosynthesis. Curr. Opin. Plant Biol. 2007, 10, 564–572. [Google Scholar] [CrossRef]
- Zhong, R.; Ye, Z.H. MYB46 and MYB83 Bind to the SMRE Sites and Directly Activate a Suite of Transcription Factors and Secondary Wall Biosynthetic Genes. Plant Cell Physiol. 2012, 53, 368–380. [Google Scholar] [CrossRef]
- Monica, G. EgMYB2, a new transcriptional activator from Eucalyptus xylem, regulates secondary cell wall formation and lignin biosynthesis. Plant J. 2005, 43, 553–567. [Google Scholar]
- Guo, H.Y. Expression of the MYB transcription factor gene BplMYB46 affects abiotic stress tolerance and secondary cell wall deposition in Betula platyphylla. Plant Biotechnol. J. 2017, 15, 107–121. [Google Scholar] [CrossRef]
- Bomal, C.B.; Frank, C.S.; Mansfield, S.D. Involvement of Pinus taeda MYB1 and MYB8 in phenylpropanoid metabolism and secondary cell wall biogenesis: A comparative in planta analysis. J. Exp. Bot. 2008, 59, 3925–3939. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhong, R.; Mccarthy, R.L.; Haghighat, M. The poplar MYB master switches bind to the SMRE site and activate the secondary wall biosynthetic program during wood formation. PLoS ONE 2013, 8, e69219. [Google Scholar] [CrossRef]
- Zhong, R.Q. Transcriptional activation of secondary wall biosynthesis by rice and maize NAC and MYB transcription factors. Plant Cell Physiol. 2011, 52, 1856–1871. [Google Scholar] [CrossRef] [Green Version]
- Cai, S.; Guo, Y.; Li, Y. Intratree Variation in Viscoelastic Properties of Cell Walls of Masson Pine (Pinus Massoniana Lamb). J. Renew. Mater. 2022, 10, 119–133. [Google Scholar] [CrossRef]
- Ni, Z.; Han, X.; Yang, Z. Integrative analysis of wood biomass and developing xylem transcriptome provide insights into mechanisms of lignin biosynthesis in wood formation of Pinus massoniana. Int. J. Biol. Macromol. 2020, 163, 1926–1937. [Google Scholar] [CrossRef] [PubMed]
- Morse, A.M.; Peterson, D.G.; Nurul, I. Evolution of Genome Size and Complexity in Pinus. PLoS ONE 2009, 4, e4332. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yao, S.; Wu, F.; Hao, Q. Transcriptome-Wide Identification of WRKY Transcription Factors and Their Expression Profiles under Different Types of Biological and Abiotic Stress in Pinus massoniana Lamb. Genes 2020, 11, 1386. [Google Scholar] [CrossRef] [PubMed]
- Wu, F.; Sun, X.; Zou, B.; Zhu, P.; Ji, K. Transcriptional Analysis of Masson Pine (Pinus massoniana) under High CO2 Stress. Genes 2019, 10, 804. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- María, B.P.; María, T.L.; Blanca, C.B. PpNAC1, a main regulator of phenylalanine biosynthesis and utilization in maritime pine. Plant Biotechnol. J. 2017, 16, 1094–1104. [Google Scholar]
- Yoshimi, N.; Masatoshi, Y.; Hitoshi, E. NAC-MYB-based transcriptional regulation of secondary cell wall biosynthesis in land plants. Front. Plant Sci. 2015, 6, 288. [Google Scholar]
- Taylor-Teeples, M.; Lin, L.; de Lucas, M.; Turco, G.; Toal, T.W.; Gaudinier, A. An Arabidopsis gene regulatory network for secondary cell wall synthesis. Nature 2015, 517, 571–575. [Google Scholar] [CrossRef] [Green Version]
- Liu, B.G.; Wang, J.P. Tracheid-associated transcription factors in loblolly pine. Tree Physiol. 2020, 40, 700–703. [Google Scholar] [CrossRef]
- Patzlaff, A.; McInnis, S.; Courtenay, A.; Surman, C.; Newman, L.J.; Smith, C. Characterization of a pine MYB that regulates lignifification. Plant J. 2003, 36, 743–754. [Google Scholar] [CrossRef]
- Newman, L.J.; Perazza, D.E.; Juda, L.; Campbell, M.M. Involvement of the R2R3-MYB, AtMYB61, in the ectopic lignifification and darkphotomorphogenic components of the det3 mutant phenotype. Plant J. 2004, 37, 239–250. [Google Scholar] [CrossRef]
- Bedon, F.; Grima-Pettenati, J.; MacKay, J. Conifer R2R3-MYB transcription factors: Sequence analysis and gene expression in wood forming tissues of white spruce (Picea glauca). BMC Plant Biol. 2007, 7, 17. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Craven-Bartle, B.; Pascual, M.B.; Canovas, F.M.; Avila, C. A Myb transcription factor regulates genes of the phenylalanine pathway in maritime pine. Plant J. 2013, 74, 755–766. [Google Scholar] [CrossRef] [PubMed]
- Akiyoshi, N.; Yoshimi, N.; Ryosuke, S.; Yusuke, K.; Misato, O.; Taku, D. Involvement of VNS NAC-domain transcription factors in tracheid formation in Pinus taeda. Tree Physiol. 2019, 40, 704–716. [Google Scholar] [CrossRef] [PubMed]
- Schuetz, M.; Benske, A.; Smith, R.A.; Watanabe, Y.; Tobimatsu, Y.; Ralph, J.; Demura, T. Laccases direct lignification in the discrete secondary cell wall domains of protoxylem. Plant Physiol. 2014, 166, 798–807. [Google Scholar] [CrossRef] [Green Version]
- Ko, J. Ectopic expression of MYB46 identifies transcriptional regulatory genes involved in secondary wall biosynthesis in Arabidopsis. Plant J. Cell Mol. Biol. 2010, 60, 649–665. [Google Scholar] [CrossRef]
- Rogers, L.A.; Campbell, M.M. The genetic control of lignin deposition during plant growth and development. New Phytol. 2004, 1, 17–30. [Google Scholar] [CrossRef]
- Zhong, R.; Ye, Z.H. Transcriptional regulation of lignin biosynthesis. Plant Signal. Behav. 2009, 4, 1028–1034. [Google Scholar] [CrossRef] [Green Version]
- Stracke, R.; Werber, M.; Weisshaar, B. The R2R3-MYB gene family in Arabidopsis thaliana. Curr. Opin. Plant Biol. 2001, 4, 447–456. [Google Scholar] [CrossRef]
- Lin, Y.C.; Li, W.; Sun, Y.H.; Kumari, S.; Wei, H.; Li, Q.; Tunlaya-Anukit, S. SND1 Transcription factor-directed quantitative functional hierarchical genetic regulatory network in wood formation in Populus trichocarpa. Plant Cell 2013, 25, 4324–4341. [Google Scholar] [CrossRef] [Green Version]
- Wang, S.; Li, E.; Porth, I.; Chen, J.G.; Mansfifield, S.D.; Douglas, C.J. Regulation of secondary cell wall biosynthesis by poplar R2R3 MYB transcription factor PtrMYB152 in Arabidopsis. Sci. Rep. 2014, 4, 5054. [Google Scholar] [CrossRef] [Green Version]
- Maleki, S.S.; Mohammadi, K.; Movahedi, A. Increase in cell wall thickening and biomass production by overexpression of PmCesA2 in Poplar. Front. Plant Sci. 2020, 11, 110. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Altschul, S.F.; Gish, W.; Miller, W.; Myers, E.W.; Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 1990, 215, 403–410. [Google Scholar] [CrossRef]
- Gietz, R.D.; Schiestl, R.H. High-efficiency yeast transformation using the LiAc/SS carrier DNA/PEG method. Nat. Protoc. 2007, 2, 38–41. [Google Scholar] [CrossRef] [PubMed]
- Zhu, P.H.; Ma, Y.Y.; Zhu, L.Z.; Chen, Y.; Ji, K.S. Selection of suitable reference genes in Pinus massoniana Lamb. under different abiotic stresses for qPCR normalization. Forests 2019, 10, 632. [Google Scholar] [CrossRef] [Green Version]
- Holsters, M.; De Waele, D.; Depicker, A.; Messens, E.; Van Montagu, M.; Schell, J. Transfection and transformation of Agrobacterium tumefaciens. Mol. Gen. Genet. 1978, 163, 181–187. [Google Scholar] [CrossRef]
- Yu, Y.; Zhao, Y.Q.; Zhao, B.; Ren, S.; Guo, Y.D. Influencing factors and structural characterization of hyperhydricity of in vitro regeneration in Brassica oleracea var. italica. J. Plant Sci. 2011, 91, 159–165. [Google Scholar] [CrossRef]
- Li, C.; Wang, X.; Ran, L.; Tian, Q.; Fan, D.; Luo, K. PtoMYB92 is a transcriptional activator of the lignin biosynthetic pathway during secondary cell wall formation in Populus tomentosa. Plant Cell Physiol. 2015, 56, 2436–2446. [Google Scholar] [CrossRef] [Green Version]
- Iiyama, K.; Wallis, A.F. An improved acetyl bromide procedure for determining lignin in woods and wood pulps. Wood Sci. Technol. 1988, 22, 271–280. [Google Scholar] [CrossRef]
- Van Soest, P.J.; Wine, R.H. Use of detergents in the analysis of fibrous feeds IV determination of plant cell-wall constituents. J. Assoc. Anal. Chem. 1967, 50, 50–55. [Google Scholar]
- Yang, Y.; Li, R.; Qi, M. In vivo analysis of plant promoters and transcription factors by agroinfiltration of tobacco leaves. Plant J. 2000, 22, 543–551. [Google Scholar] [CrossRef]



| Samples | Lignin (%) | Cellulose (%) | Hemicellulose (%) |
|---|---|---|---|
| WT | 21.56 ± 0.16 | 42.17 ± 0.92 | 24.56 ± 1.0 |
| L1 | 22.80 ± 0.11 * | 42.38 ± 0.97 | 25.41 ± 1.7 |
| L5 | 25.09 ± 0.32 * | 44.56 ± 0.36 * | 24.16 ± 2.8 |
| L6 | 24.80 ± 0.11 * | 44.34 ± 0.97 * | 24.45 ± 1.3 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yao, S.; Chen, P.; Yu, Y.; Zhang, M.; Wang, D.; Liu, J.; Hao, Q.; Ji, K. PmMYB4, a Transcriptional Activator from Pinus massoniana, Regulates Secondary Cell Wall Formation and Lignin Biosynthesis. Forests 2021, 12, 1618. https://doi.org/10.3390/f12121618
Yao S, Chen P, Yu Y, Zhang M, Wang D, Liu J, Hao Q, Ji K. PmMYB4, a Transcriptional Activator from Pinus massoniana, Regulates Secondary Cell Wall Formation and Lignin Biosynthesis. Forests. 2021; 12(12):1618. https://doi.org/10.3390/f12121618
Chicago/Turabian StyleYao, Sheng, Peizhen Chen, Ye Yu, Mengyang Zhang, Dengbao Wang, Jiahe Liu, Qingqing Hao, and Kongshu Ji. 2021. "PmMYB4, a Transcriptional Activator from Pinus massoniana, Regulates Secondary Cell Wall Formation and Lignin Biosynthesis" Forests 12, no. 12: 1618. https://doi.org/10.3390/f12121618
APA StyleYao, S., Chen, P., Yu, Y., Zhang, M., Wang, D., Liu, J., Hao, Q., & Ji, K. (2021). PmMYB4, a Transcriptional Activator from Pinus massoniana, Regulates Secondary Cell Wall Formation and Lignin Biosynthesis. Forests, 12(12), 1618. https://doi.org/10.3390/f12121618






