Complete Chloroplast Genome of Pinus densiflora Siebold & Zucc. and Comparative Analysis with Five Pine Trees
Abstract
1. Introduction
2. Materials and Methods
2.1. Sampling, DNA Extraction and Sequencing
2.2. Assemblies of Chloroplast Genome Sequences and Annotation
2.3. Alignments and Construction of a Phylogenetic Tree
3. Results
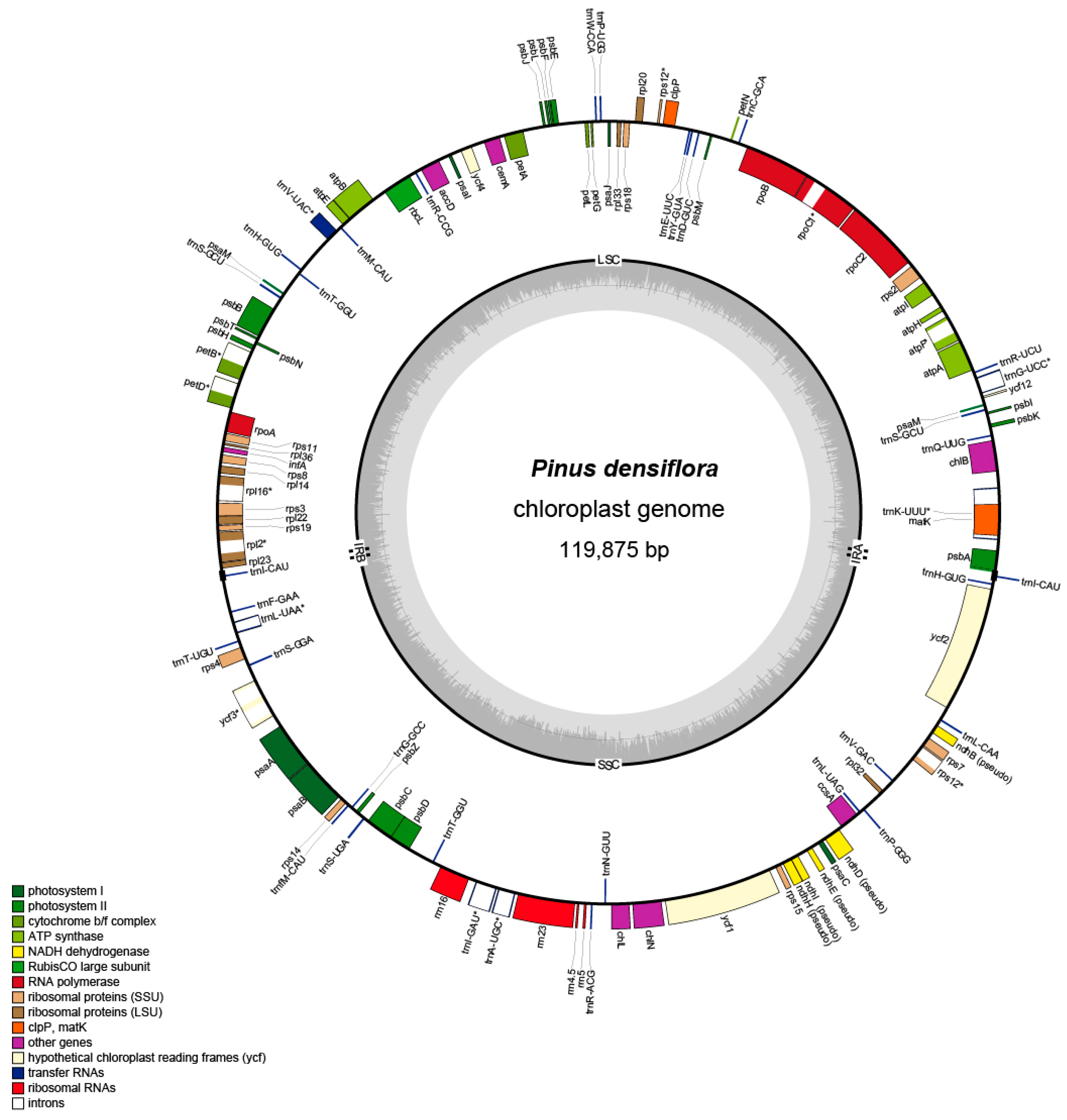
3.1. The Structure of Chloroplast Genomes of P. densiflora
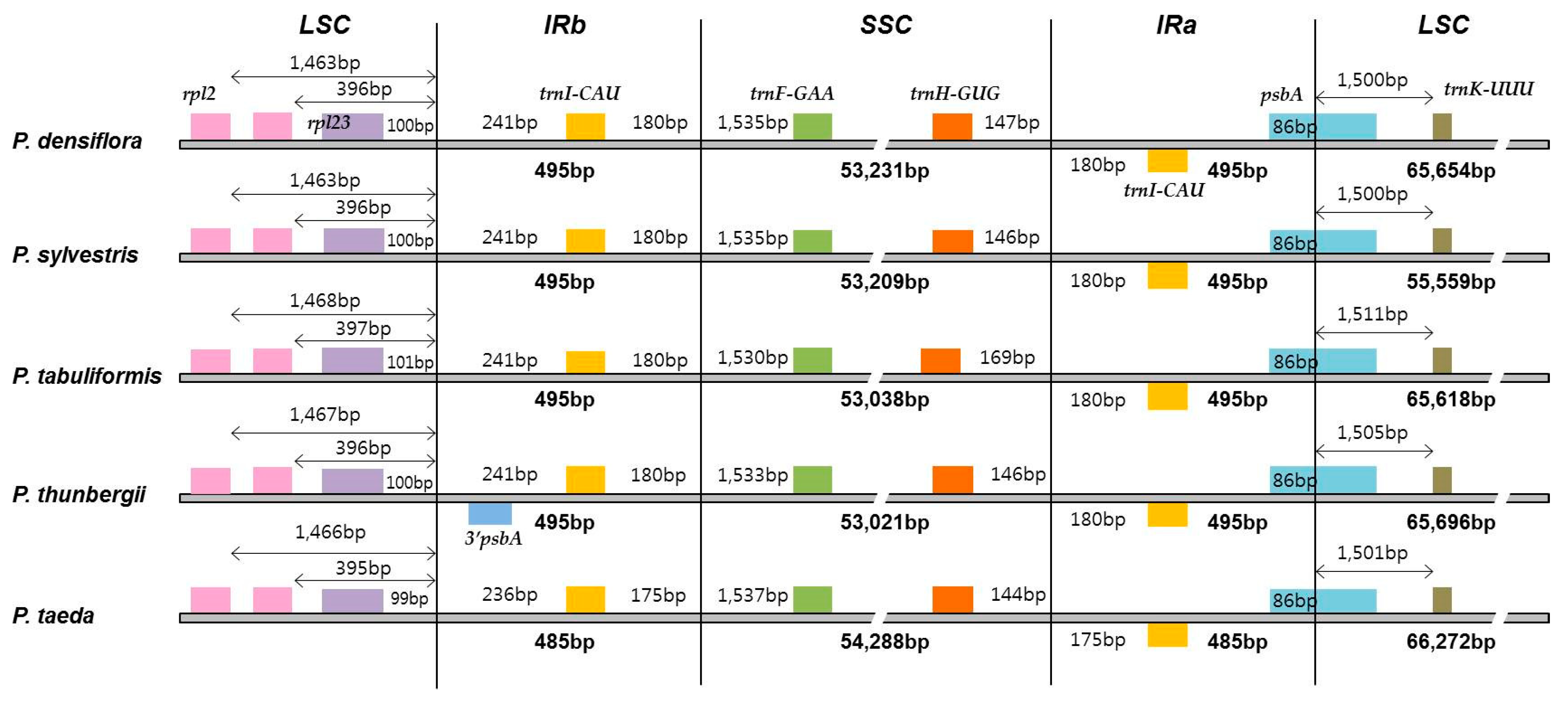
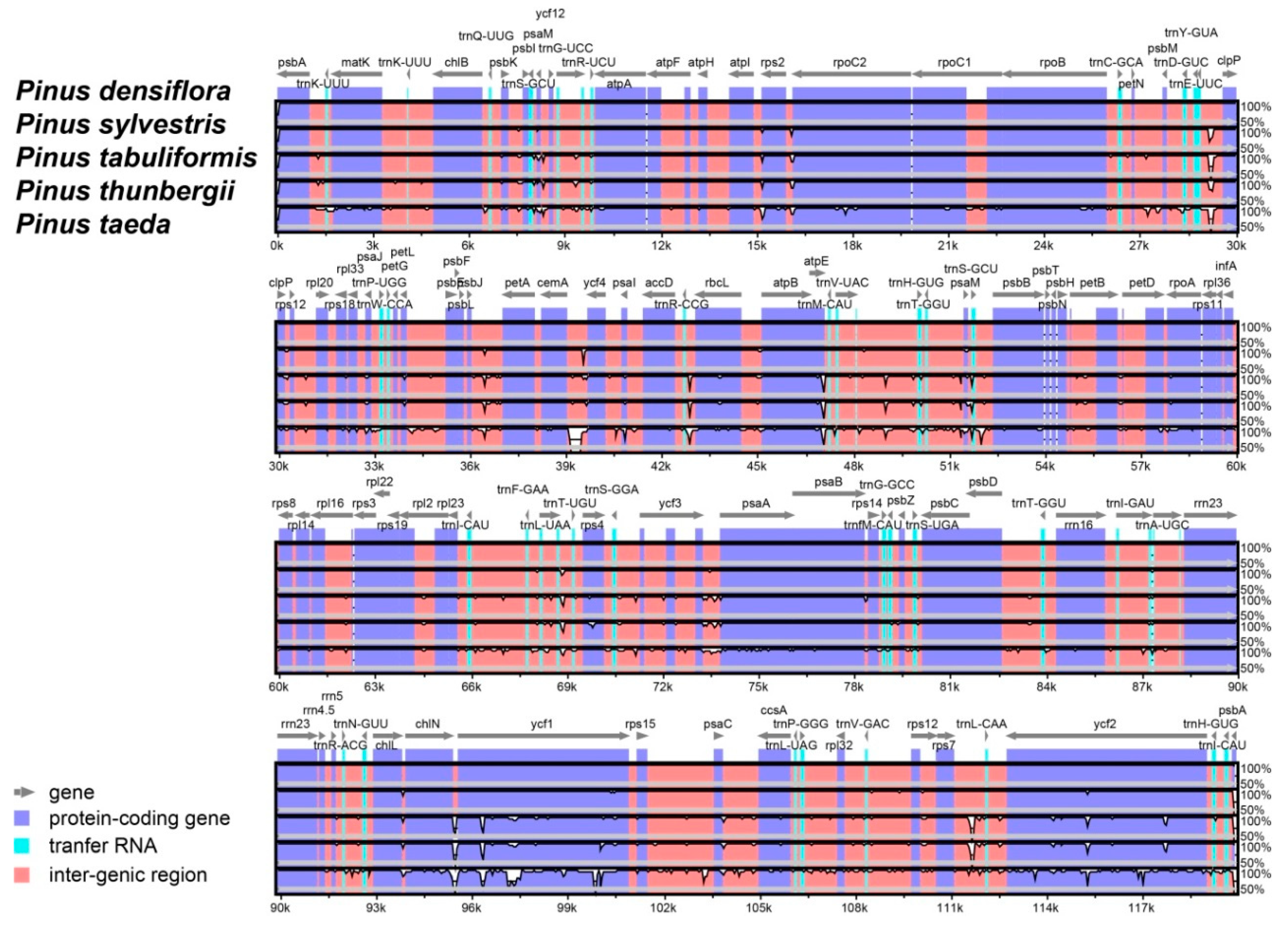
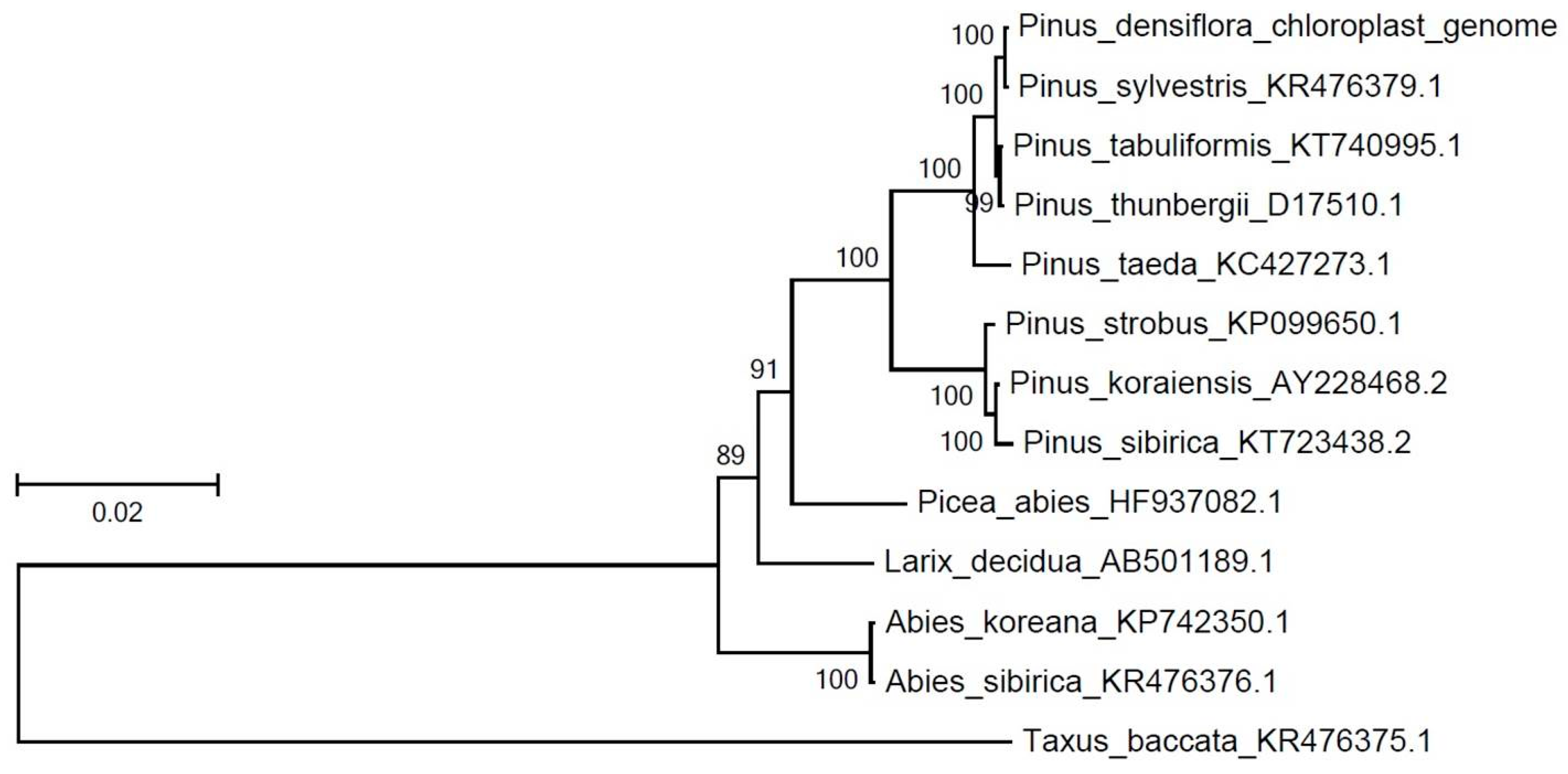
3.2. Comparative Analyses of the Chloroplast Genome with Other Pinus Species for the Identification of DNA Variation
4. Discussion and Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Ni, Z.; Ye, Y.; Bai, T.; Xu, M.; Xu, L.A. Complete chloroplast genome of pinus massoniana (Pinaceae): Gene rearrangements, loss of ndh genes, and short inverted repeats contraction, expansion. Molecules 2017, 22, 1528. [Google Scholar] [CrossRef] [PubMed]
- Szmidt, A.E.; Wang, X.R. Molecular systematic and genetic differentiation of Pinus sylvestris (L.) and P. densiflora (sieb. et Zucc.). Theor. Appl. Genet. 1993, 86, 159–165. [Google Scholar] [CrossRef] [PubMed]
- Korea Forest Service. Statistical Yearbook of Forestry; Korea Forest Service: Daejeon, Korea, 2011.
- Duan, R.Y.; Yang, L.M.; Lv, T.; Wu, G.L.; Huang, M.Y. The complete chloroplast genome sequence of Pinus dabeshanensis. Conserv. Genet. Resour. 2016, 8, 395–397. [Google Scholar] [CrossRef]
- Asaf, S.; Waqas, M.; Khan, A.L.; Khan, M.A.; Kang, S.M.; Imran, Q.M.; Shahzad, R.; Bilal, S.; Yun, B.W.; Lee, I.J. The complete chloroplast genome of wild rice (Oryza minuta) and its comparison to related species. Front. Plant Sci. 2017, 8, 304. [Google Scholar] [CrossRef] [PubMed]
- Sugiura, M. The chloroplast genome. Plant Mol. Biol. 1992, 19, 149–168. [Google Scholar] [CrossRef] [PubMed]
- Palmer, J.D.; Thompson, W.F. Chloroplast DNA rearrangements are more frequent when a large inverted repeat sequence is lost. Cell 1982, 29, 537–550. [Google Scholar] [CrossRef]
- Lin, C.P.; Wu, C.S.; Huang, Y.Y.; Chaw, S.M. The complete chloroplast genome of Ginkgo biloba reveals the mechanism of inverted repeat contraction. Genome Biol. Evol. 2012, 4, 374–381. [Google Scholar] [CrossRef] [PubMed]
- Strauss, S.H.; Palmer, J.D.; Howe, G.T.; Doerksen, A.H. Chloroplast genomes of two conifers lack a large inverted repeat and are extensively rearranged. Proc. Natl. Acad. Sci. USA 1988, 85, 3898–3902. [Google Scholar] [CrossRef]
- Perry, A.S.; Wolfe, K.H. Nucloetide substitution rates in legume chloroplast DNA depend on the presence of the inverted repeat. J. Mol. Evol. 2002, 55, 501–508. [Google Scholar] [CrossRef]
- Wu, C.S.; Wang, Y.N.; Hsu, C.Y.; Lin, C.P.; Chaw, S.M. Loss of different inverted repeat copies from the chloroplast genomes of Pinaceae and cupressophytes and influence of heterotachy on the evaluation of gymnosperm phylogeny. Genome Biol. Evol. 2011, 3, 1284–1295. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Ma, J.; Yang, B.; Li, R.; Zhu, W.; Sun, L.; Tian, J.; Zhang, L. The complete chloroplast genome sequence of Taxus chinensis var. mairei (Taxaceae): Loss of an inverted repeat region and comparative analysis with related species. Gene 2014, 540, 201–209. [Google Scholar] [CrossRef] [PubMed]
- Heather, J.M.; Chain, B. The sequence of sequencers: The history of sequencing DNA. Genomics 2016, 107, 1–8. [Google Scholar] [CrossRef] [PubMed]
- McKenna, A.; Hanna, M.; Banks, E.; Sivachenko, A.; Cibulskis, K.; Kernytsky, A.; Garimella, K.; Altshuler, D.; Gabriel, S.; Daly, M.; et al. The genome analysis toolkit: A mapreduce framework for analyzing next-generation dna sequencing data. Genome Res. 2010, 20, 1297–1303. [Google Scholar] [CrossRef] [PubMed]
- Scarcelli, N.; Mariac, C.; Couvreur, T.L.; Faye, A.; Richard, D.; Sabot, F.; Berthouly-Salazar, C.; Vigouroux, Y. Intra-individual polymorphism in chloroplasts from NGS data: Where does it come from and how to handle it? Mol. Ecol. Resour. 2016, 16, 434–445. [Google Scholar] [CrossRef] [PubMed]
- Mikheyev, A.S.; Tin, M.M. A first look at the Oxford Nanopore MinION sequencer. Mol. Ecol. Resour. 2014, 14, 1097–1102. [Google Scholar] [CrossRef] [PubMed]
- Lu, H.; Giordano, F.; Ning, Z. Oxford Nanopore MinION sequencing and genome assembly. Genom. Proteom. Bioinform. 2016, 14, 265–279. [Google Scholar] [CrossRef] [PubMed]
- Lowe, T.M.; Eddy, S.R. tRNAscan-SE: A program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Res. 1997, 25, 955–964. [Google Scholar] [CrossRef]
- Laslett, D.; Canback, B. ARAGORN, a program to detect tRNA genes and tmRNA genes in nucleotide sequences. Nucleic Acids Res. 2004, 32, 11–16. [Google Scholar] [CrossRef]
- Tillich, M.; Lehwark, P.; Pellizzer, T.; Ulbricht-Jones, E.S.; Fischer, A.; Bock, R.; Greiner, S. GeSeq - versatile and accurate annotation of organelle genomes. Nucleic Acids Res. 2017, 45, W6–W11. [Google Scholar] [CrossRef]
- Mayor, C.; Brudno, M.; Schwartz, J.R.; Poliakov, A.; Rubin, E.M.; Frazer, K.A.; Pachter, L.S.; Dubchak, I. VISTA: Visualizing global DNA sequence alignments of arbitrary length. Bioinformatics 2000, 16, 1046–1047. [Google Scholar] [CrossRef]
- Katoh, K.; Standley, D.M. MAFFT multiple sequence alignment software version 7: Improvements in performance and usability. Mol. Biol. Evol. 2013, 30, 772–780. [Google Scholar] [CrossRef]
- Goodwin, S.; Gurtowski, J.; Ethe-Sayers, S.; Deshpande, P.; Schatz, M.C.; McCombie, W.R. Oxford Nanopore sequencing, hybrid error correction, and de novo assembly of a eukaryotic genome. Genome Res. 2015, 25, 1750–1756. [Google Scholar] [CrossRef]
- Laver, T.; Harrison, J.; O’Neill, P.A.; Moore, K.; Farbos, A.; Paszkiewicz, K.; Studholme, D.J. Assessing the performance of the Oxford Nanopore Technologies MinION. Biomol. Detect Quantif. 2015, 3, 1–8. [Google Scholar] [CrossRef]
- Giordano, F.; Aigrain, L.; Quail, M.A.; Coupland, P.; Bonfield, J.K.; Davies, R.M.; Tischler, G.; Jackson, D.K.; Keane, T.M.; Li, J.; et al. De novo yeast genome assemblies from MinION, PacBio and MiSeq platforms. Sci. Rep. 2017, 7, 3935. [Google Scholar] [CrossRef]
- Ishizuka, W.; Tabata, A.; Ono, K.; Fukuda, Y.; Hara, T. Draft chloroplast genome of Larix gmelinii var. japonica: Insight into intraspecific divergence. J. For. Res. 2017, 22, 393–398. [Google Scholar] [CrossRef]
- Ranade, S.S.; Garcia-Gil, M.R.; Rossello, J.A. Non-functional plastid ndh gene fragments are present in the nuclear genome of Norway spruce (Picea abies L. Karsch): Insights from in silico analysis of nuclear and organellar genomes. Mol. Genet. Genom. 2016, 291, 935–941. [Google Scholar] [CrossRef]
- Lin, C.S.; Chen, J.J.; Chiu, C.C.; Hsiao, H.C.; Yang, C.J.; Jin, X.H.; Leebens-Mack, J.; de Pamphilis, C.W.; Huang, Y.T.; Yang, L.H.; et al. Concomitant loss of NDH complex-related genes within chloroplast and nuclear genome is some orchids. Plant J. 2017, 90, 994–1006. [Google Scholar] [CrossRef]
- Raubeson, L.A.; Jansen, R.K. A rare chloroplast-DNA structural mutation is shared by all conifers. Biochem. Syst. Ecol. 1992, 20, 17–24. [Google Scholar] [CrossRef]
- Wu, C.S.; Lin, C.P.; Hsu, C.Y.; Wang, R.J.; Chaw, S.M. Comparative chloroplast genomes of pinaceae: Insights into the mechanism of diversified genomic organizations. Genome Biol. Evol. 2011, 3, 309–319. [Google Scholar] [CrossRef]
- Wu, C.S.; Wang, Y.N.; Liu, S.M.; Chaw, S.M. Chloroplast genome (cpDNA) of Cycas taitungensis and 56 cp protein-coding genes of Gnetum parvifolium: Insights into cpDNA evolution and phylogeny of extant seed plasts. Mol. Biol. Evol. 2007, 3. [Google Scholar] [CrossRef]
- Li, J.; Gao, L.; Chen, S.; Tao, K.; Su, Y.; Wang, T. Evolution of short inverted repeat in cupressophytes, transfer of accD to nucleus in Sciadopitys verticillata and phylogenetic position of Sciadopityaceae. Sci. Rep. 2016, 6, 20934. [Google Scholar] [CrossRef]
- Moore, M.J.; Soltis, P.S.; Bell, C.D.; Burleigh, J.G.; Soltis, D.E. Phylogenetic analysis of 83 plastid genes further resolves the early diversification of eudicots. Proc. Natl. Acad. Sci. USA 2010, 107, 4623–4628. [Google Scholar] [CrossRef]
- Olsson, S.; Grivet, D.; Vian, J.C. Species-diagnostic markers in the genus Pinus: Evaluation of the chloroplast regions matK and ycf1. For. Syst. 2018, 27, 2. [Google Scholar] [CrossRef]
- Wu, F.; Li, M.; Liao, B.; Shi, X.; Xu, Y. DNA Barcoding Analysis and Phylogenetic Relation of Mangroves in Guangdong Province, China. Forests 2019, 10, 56. [Google Scholar] [CrossRef]
- Cbol Plant Working Group. A DNA barcode for land plants. Proc. Natl. Acad. Sci. USA 2009, 106, 12794–12797. [Google Scholar] [CrossRef]
- Snel, B.; Bork, P.; Huynen, M.A. Genome phylogeny based on gene content. Nat. Genet. 1999, 21, 108. [Google Scholar] [CrossRef]
- Nishiyama, T.; Wolf, P.G.; Kugita, M.; Sinclair, R.B.; Sugita, M.; Sugiura, C.; Wakasugi, T.; Yamada, K.; Yoshinaga, K.; Yamaguchi, K.; et al. Chloroplast phylogeny indicates that bryophytes are monophyletic. Mol. Biol. Evol. 2004, 21, 1813–1819. [Google Scholar] [CrossRef]
- Kosinski, G. Hybridization. In Genetics of Scots Pine; Giertych, M., Matyas, C., Eds.; Elsevier: Amsterdam, The Netherlands, 1991; pp. 183–189. [Google Scholar]
- Hizume, M.; Shibata, F.; Matsusaki, Y.; Garajova, Z. Chromosome identification and comparative karyotypic analyses of four Pinus species. Appl. Genet. 2002, 105, 491–497. [Google Scholar] [CrossRef]
- Eckert, A.J.; Benjamin, D.H. Phylogeny, historical biogeography, and patterns of diversification for Pinus (Pinaceae): Phylogenetic tests of fossil-based hypotheses. Mol. Phylogenet. Evol. 2006, 40, 166–182. [Google Scholar] [CrossRef]
- Parks, M.; Cronn, R.; Liston, A. Separating the wheat from the chaff: Mitigating the effects of noise in a plastome phylogenomic data set from Pinus L. (Pinaceae). BMC Evol. Biol. 2012, 12, 100. [Google Scholar] [CrossRef]
- Asaf, S.; Khan, A.L.; Khan, M.A.; Shahzad, R.; Lubna; Kang, S.M.; Al-Harrasi, A.; Al-Rawahi, A.; Lee, I.J. Complete chloroplast genome sequence and comparative analysis of loblolly pine (Pinus taeda L.) with related species. PLoS ONE 2018, 13, e0192966. [Google Scholar] [CrossRef]

| Sequencing Platform | Input Reads | Trimmed Reads | Raw Bases | Trimmed Bases |
|---|---|---|---|---|
| MiSeq | 49,013,296 | 40,786,223 (83.21%) | 14,563,262,097 | 10,101,761,675 (69.36%) |
| ONM | 305,965 | 306,493 (100.21%) | 2,116,530,768 | 2,089,930,503 (98.74%) |
| Genome Size (bp) | LSC Length (bp) | SSC Length (bp) | IR Length (bp) | Number of Genes | |
|---|---|---|---|---|---|
| P. densiflora (MK285358) | 119,875 | 65,654 | 53,231 | 495 | 113 |
| P. sylvestris (KR476379) | 119,758 | 65,559 | 53,209 | 495 | 112 |
| P. thunbergii (D17510) | 119,707 | 65,696 | 53,021 | 495 | 113 |
| P. tabuliformis (KT740995) | 119,646 | 65,618 | 53,038 | 495 | 114 |
| P. taeda (KC427273) | 121,530 | 66,272 | 54,288 | 485 | 110 |
| Function | Genes |
|---|---|
| RNAs, ribosomal | rrn4.5, rrn5, rrn16, rrn23, |
| RNAs, transfer | trnA-UGC *, trnC-GCA, trnD-GUC, trnE-UUC, trnF-GAA, trnG-GCC, trnG-UCC *, trnH-GUG, trnI-CAU, trnI-GAU *, trnK-UUU *,T, trnL-CAA, trnL-UAA *, trnL-UAG, trnM-CAU, trnfM-CAU, trnN-GUU, trnP-UGG, trnP-GGG, trnQ-UUG, trnR-ACG, trnR-CCG, trnR-UCU, trnS-GCU, trnS-GGA, trnS-UGA, trnT-GGU, trnT-UGU, trnV-GAC, trnV-UAC *, trnW-CCA, trnY-GUA |
| Transcription and splicing | rpoA, rpoB, rpoC1 *, rpoC2, matK |
| Translation, ribosomal proteins | |
| Small subunit | rps2, rps3, rps4, rps7, rps8, rps11, rps12 **,T, rps14, rps15, rps18, rps19 |
| Large subunit | rpl2 *, rpl14, rpl16 *, rpl20, rpl22, rpl23, rpl32, rpl33, rpl36 |
| Photosynthesis | |
| ATP synthase | atpA, atpB, atpE, atpF *, atpH, atpI |
| Photosystem Ⅰ | psaA, psaB, psaC, psaI, psaJ, psaM, ycf3 **, ycf4 |
| Photosystem Ⅱ | psbA, psbB, psbC, psbD, psbE, psbF, psbH, psbI, psbJ, psbK, psbL, psbM, psbN, psbT, psbZ |
| Calvin cycle | rbcL |
| Cytochrome complex | petA, petB *, petD *, petG, petL, petN |
| Chlorophyll biosynthesis | chlB, chlL, chlN |
| Others | clpP, accD, cemA, ccsA, infA, ycf1, ycf2, ycf12 |
| Species | Accession No. | No. of Protein Coding Genes | No. of Common Protein Coding Genes with P. densiflora |
|---|---|---|---|
| Pinus sylvestris Linn. | KR476379 | 73 | 73 |
| Pinus tabuliformis Carr. | KT740995 | 74 | 73 |
| Pinus thunbergii Parl. | D17510 | 69 | 69 |
| Pinus taeda L. | KC427273 | 71 | 71 |
| Pinus strobus Linn. | KP099650 | - | - |
| Pinus koraiensis Sieb. et Zucc. | AY228468 | 74 | 72 |
| Pinus sibirica (Loud.) Mayr | KT723438 | 77 | 73 |
| Picea abies (L.) Karst. | HF937082 | 74 | 72 |
| Larix decidua Mill | AB501189 | 72 | 71 |
| Abies koreana Wils | KP742350 | 74 | 73 |
| Abies sibirica Ledeb. | KR476376 | 74 | 73 |
| Taxus baccata L. | KR476375 | 81 | 70 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kang, H.-I.; Lee, H.O.; Lee, I.H.; Kim, I.S.; Lee, S.-W.; Yang, T.J.; Shim, D. Complete Chloroplast Genome of Pinus densiflora Siebold & Zucc. and Comparative Analysis with Five Pine Trees. Forests 2019, 10, 600. https://doi.org/10.3390/f10070600
Kang H-I, Lee HO, Lee IH, Kim IS, Lee S-W, Yang TJ, Shim D. Complete Chloroplast Genome of Pinus densiflora Siebold & Zucc. and Comparative Analysis with Five Pine Trees. Forests. 2019; 10(7):600. https://doi.org/10.3390/f10070600
Chicago/Turabian StyleKang, Hye-In, Hyun Oh Lee, Il Hwan Lee, In Sik Kim, Seok-Woo Lee, Tae Jin Yang, and Donghwan Shim. 2019. "Complete Chloroplast Genome of Pinus densiflora Siebold & Zucc. and Comparative Analysis with Five Pine Trees" Forests 10, no. 7: 600. https://doi.org/10.3390/f10070600
APA StyleKang, H.-I., Lee, H. O., Lee, I. H., Kim, I. S., Lee, S.-W., Yang, T. J., & Shim, D. (2019). Complete Chloroplast Genome of Pinus densiflora Siebold & Zucc. and Comparative Analysis with Five Pine Trees. Forests, 10(7), 600. https://doi.org/10.3390/f10070600




