1. Introduction
Unstructured clinical notes contain a substantial portion of clinically relevant patient information, yet their free-text format poses persistent challenges for automated extraction and structuring. Reliable identification of diagnoses, procedures, and medications from clinical narratives is essential for biomedical research, clinical decision support, and the development of data-driven healthcare systems [
1,
2]. However, clinical information extraction remains a challenging task in clinical natural language processing (NLP) due to high lexical variability, large and heterogeneous label spaces, frequent abbreviations, and domain-specific writing styles that often diverge from standardized terminologies [
3].
Early work in clinical named entity recognition and information extraction relied on rule-based systems and statistical models such as Conditional Random Fields, which required extensive feature engineering and exhibited limited robustness across institutions and documentation styles. The introduction of transformer-based language models, particularly Bidirectional Encoder Representations from Transformers (BERT) [
4], marked a turning point by enabling contextualized representations of clinical language. Domain-adapted variants including BioBERT [
5], ClinicalBERT, and BlueBERT [
6], trained on biomedical literature and clinical notes, consistently outperformed general-domain models on biomedical and clinical NLP tasks. Subsequent transformer-based clinical models, such as RoBERTa-MIMIC, reported strong performance on selected clinical concept extraction benchmarks, with F1-scores exceeding 0.89 in controlled settings [
7]. These advances established domain-specific pretraining as a foundational component of high-performing supervised clinical information extraction systems. In parallel, multi-label classification formulations have been shown to be effective when ground truth is provided as sets of clinical concepts rather than entity spans, particularly in settings characterized by large clinical vocabularies.
More recently, large language models (LLMs) have introduced an alternative paradigm for clinical text extraction that reduces reliance on task-specific supervision. LLMs demonstrate strong natural language understanding and generation capabilities [
8], enabling task adaptation through prompting rather than explicit fine-tuning. Few-shot prompting has been successfully applied to a range of clinical NLP tasks, allowing models to leverage limited in-context examples [
9]. However, recent empirical studies report nuanced and sometimes counterintuitive findings. Prompting performance is highly sensitive to prompt structure, with reasoning-oriented strategies such as chain-of-thought and verification-based prompting outperforming simple example-based approaches in certain clinical contexts [
10]. Moreover, evidence that zero-shot prompting can outperform few-shot configurations in biomedical tasks suggests that pretrained LLMs may already encode substantial domain knowledge, which is not always enhanced by in-context demonstrations [
11]. These observations raise important questions about when prompting alone is sufficient for structured clinical extraction and when supervised learning remains necessary.
Retrieval-augmented generation (RAG) has emerged as a complementary strategy for grounding LLM outputs in external knowledge sources, a consideration of particular importance in healthcare applications where accuracy and consistency are critical [
12]. RAG-based systems have demonstrated improvements in diagnostic reasoning, clinical decision support, and biomedical information extraction by incorporating evidence from structured knowledge bases and clinical literature [
13]. In the context of clinical entity extraction, RAG can be applied as a post-processing canonicalization mechanism to align extracted surface forms with standardized terminology. Such canonicalization can mitigate variability arising from abbreviations, brand–generic differences, and heterogeneous documentation practices, thereby improving semantic consistency and interpretability. However, when predicted entities diverge substantially from reference terminology, canonicalization alone is unlikely to recover missing extractions or significantly improve strict string-based performance.
Alongside advances in modeling, growing attention has been directed toward limitations of traditional evaluation metrics for clinical NLP. Exact string matching often underestimates extraction quality by penalizing clinically equivalent expressions that differ lexically [
14]. Fuzzy matching partially addresses this limitation but remains insufficient for capturing true clinical equivalence across diverse surface forms. Recent work demonstrates that large language model-based semantic evaluation, including LLM-as-Judge approaches, better reflects clinical correctness by accounting for synonymy, abbreviation expansion, and partial matches [
15,
16]. Comparative analyses consistently reveal large discrepancies between exact-match and semantically informed evaluation, underscoring the need for assessment frameworks that align more closely with clinical interpretation rather than surface-form similarity alone.
Despite substantial progress across supervised learning, prompting-based extraction, and retrieval-augmented approaches, there remains limited systematic understanding of their relative tradeoffs when applied to the same clinical task under controlled conditions. Existing studies often evaluate these paradigms in isolation, rely on heterogeneous datasets or metrics, or prioritize performance improvements without examining methodological implications. This study addresses these gaps by conducting a unified evaluation of three complementary strategies for structured clinical information extraction from MIMIC-IV discharge summaries: (1) prompting-based extraction using zero-shot, few-shot, and Chain-of-Verification strategies; (2) retrieval-augmented generation for entity canonicalization; and (3) supervised fine-tuning of a domain-specific transformer model. By combining exact, fuzzy, and semantic evaluation metrics, we aim to clarify algorithmic tradeoffs across approaches, quantify the discrepancy between string-based and clinically informed evaluation, and provide practical guidance for selecting extraction strategies under varying data availability and resource constraints.
2. Methodological Framework and Study Design
Figure 1 provides a high-level overview of the proposed methodological framework.
The framework is designed to enable a controlled comparison of three dominant paradigms for structured clinical entity extraction from free-text discharge summaries: prompting-based extraction using large language models, retrieval-augmented canonicalization, and supervised fine-tuning of a domain-specific transformer model. Discharge summaries from the MIMIC-IV dataset are first preprocessed and partitioned into training, validation, and test sets. The pipeline then branches into two primary extraction pathways: (1) a prompting-based pathway that applies large language models to directly generate structured outputs, and (2) a supervised learning pathway based on BioClinicalBERT.
Outputs generated by the prompting-based pathway may optionally pass through a retrieval-augmented generation (RAG)-based canonicalization stage. This stage is applied as a post-processing step and is designed to improve terminology consistency without altering extraction coverage. Outputs from all pathways are subsequently evaluated using a unified assessment protocol. By keeping data splits, entity definitions, and evaluation criteria fixed across all experiments, the framework isolates methodological differences between prompting, retrieval augmentation, and supervised learning, allowing their respective tradeoffs to be examined under comparable conditions.
Within this framework, prompting-based extraction is evaluated across multiple in-context learning strategies using both closed-source and open-source LLMs. Zero-shot, few-shot, and Chain-of-Verification prompting are applied to the same extraction task and output schema, enabling assessment of how prompt structure and model choice influence extraction behavior. This design facilitates comparison of prompting strategies without conflating performance differences with variations in task formulation or output constraints.
Retrieval-augmented canonicalization is incorporated as an entity-type-specific post-processing component. Predicted entities are embedded using biomedical sentence representations and compared against separate canonical label sets for diagnoses, procedures, and medications. Canonicalization is applied selectively based on similarity thresholds, ensuring that surface-form normalization improves interpretability and consistency while avoiding aggressive over-normalization. Treating retrieval augmentation as a modular refinement step allows its effects to be evaluated independently from the underlying extraction process.
The supervised learning pathway employs BioClinicalBERT, a transformer model pre-trained on clinical text, and formulates entity extraction as a multi-label classification problem. This formulation reflects the structure of the available ground truth, which is provided as sets of clinical concepts per admission rather than token-level annotations. Model training and inference are conducted using the same data partitions as the prompting-based experiments, enabling direct comparison between supervision-intensive and supervision-light approaches under identical conditions.
Finally, all extraction outputs are assessed using a unified evaluation framework that combines exact string matching, fuzzy lexical matching, and semantic evaluation via an LLM-based judge. This multi-level evaluation is intended to capture both surface-level accuracy and clinically meaningful correctness, and to highlight discrepancies between string-based and semantic assessment. Integrating all methods within a single experimental and evaluation framework enables systematic analysis of tradeoffs among prompting, retrieval-augmented, and supervised approaches, particularly with respect to data availability, computational cost, and evaluation sensitivity.
3. Clinical Entity Extraction Approaches
We evaluate three complementary pathways for structured clinical entity extraction: prompting-based extraction using large language models, retrieval-augmented canonicalization as a post-processing step, and supervised fine-tuning of a domain-specific transformer model.
The prompting-based pathway applies multiple in-context learning strategies using GPT-4o-mini and Qwen-2.5-7B. Three prompting paradigms commonly used in LLM-based information extraction are evaluated. In the zero-shot setting, models receive only task instructions and a fixed JSON output schema specifying three entity lists: diagnoses, procedures, and medications [
17]. Few-shot prompting is evaluated using two-shot and four-shot configurations, with annotated examples selected via stratified sampling based on entity count percentiles (25th and 75th percentiles for two-shot; 20th, 40th, 60th, and 80th percentiles for four-shot) [
8]. Chain-of-Verification (CoVE) prompting is also evaluated as a multi-stage procedure involving initial extraction, generation of verification questions, re-examination of the clinical note, error correction, and production of a final verified JSON output [
18,
19]. All prompting strategies enforce a stable output schema to ensure comparability across configurations. Prompt templates and example constructions are provided in
Appendix A.
To address surface-form variability and clinical synonymy in prompting-based outputs, retrieval-augmented generation (RAG) is applied as an optional post-processing canonicalization step. Predicted entities are embedded using S-PubMedBERT and compared against canonical label sets derived from ground-truth annotations [
12]. Separate knowledge bases are maintained for diagnoses, procedures, and medications to prevent cross-category mismatches. For each predicted entity, the top-k candidates retrieved from a FAISS IndexFlatIP index are examined. If the highest cosine similarity score exceeds a predefined threshold, the predicted surface form is replaced with the corresponding canonical label; otherwise, the original prediction is retained. This selective, entity-type-specific canonicalization improves terminology consistency while avoiding incorrect over-normalization and does not alter extraction coverage.
The supervised extraction pathway employs BioClinicalBERT, a transformer model pre-trained on clinical notes [
20]. Because ground truth annotations are provided as sets of entity labels per admission rather than token-level spans, extraction is formulated as a multi-label classification problem with three independent sigmoid output heads corresponding to diagnoses (1078 labels), procedures (344 labels), and medications (555 labels). Weighted binary cross-entropy loss is used to mitigate class imbalance [
21]. Model optimization follows a two-stage strategy consisting of initial training with frozen encoder layers, followed by progressive unfreezing [
22]. Thresholds for converting predicted probabilities to binary outputs, along with top-K output caps for each entity type, are tuned using validation data.
4. Data and Preprocessing
This study uses the publicly available MIMIC-IV dataset, which contains 331,793 discharge summaries corresponding to more than 360,000 patients treated at Beth Israel Deaconess Medical Center between 2008 and 2022 [
23]. Discharge summaries serve as the unstructured text source, while structured ground-truth information is obtained from linked clinical tables, including diagnoses (diagnoses_icd.csv), procedures (procedures_icd.csv), and medications (prescriptions.csv). Discharge summaries are linked to these tables using the hospital admission identifier (hadm_id), enabling admission-level alignment between unstructured text and structured entity references.
To ensure consistent coverage across all entity types, the dataset is restricted to admissions for which diagnoses, procedures, and medication records are all available, resulting in 194,530 eligible admissions. Additional filtering is applied to remove atypical or incomplete cases and to focus the analysis on clinically representative discharge summaries. Specifically, admissions are retained if they contain 3–20 diagnoses per admission (median: 12), 1–8 procedures (median: 2), and 10–50 medications (median: 37). These criteria exclude extreme outliers while preserving typical inpatient complexity. After filtering, the final working dataset consists of 449 discharge summaries from 300 unique patients.
While the initial cohort included 194,530 eligible admissions, the final analytic dataset consisted of 449 discharge summaries from 300 unique patients after applying filtering criteria to ensure complete entity coverage and typical inpatient complexity. These constraints were intentionally applied to enable controlled comparison across extraction paradigms under consistent entity-type availability and to prevent patient-level information leakage across splits. However, this filtering substantially reduces the sample size relative to the source corpus and may limit representativeness. Accordingly, the results should be interpreted as a controlled methodological comparison rather than a population-level performance estimate across the full MIMIC-IV dataset.
It is also important to note that ground-truth entity labels were derived from admission-level structured coding tables (diagnoses_icd, procedures_icd, prescriptions) rather than manually annotated mentions within discharge summaries. In practice, narrative documentation and billing or structured coding data may not perfectly align. Certain diagnoses or historical conditions may appear in structured tables but not be explicitly described in the discharge summary, while clinically relevant narrative mentions may not correspond to structured codes. As a result, the reference labels used in this study may contain both omissions and inclusions relative to the narrative text, introducing potential label noise.
To prevent information leakage and ensure independence across experimental splits, patient-level stratified sampling is employed such that no patient appears in more than one partition [
24]. The resulting splits include 276 discharge summaries from 180 patients for training (61.5%), 84 summaries from 60 patients for validation (18.7%), and 89 summaries from 60 patients for testing (19.8%). Ground-truth entity labels are extracted via hadm_id joins with the structured tables and stored in JSON format to ensure consistency across extraction pathways and evaluation procedures.
5. Evaluation Framework
To assess extraction performance comprehensively, we employ a three-level evaluation framework that captures surface-level accuracy as well as clinically meaningful semantic correctness. The framework combines exact string matching, fuzzy lexical matching, and semantic evaluation using a large language model-based judge. Together, these metrics enable analysis of how different extraction approaches perform under increasingly permissive notions of correctness.
5.1. Exact Match
Exact Match evaluates strict string equivalence between predicted and ground-truth entities after normalization. This metric reflects the most conservative evaluation setting and penalizes any lexical deviation, including differences in word order, abbreviation usage, or synonymous expressions. Precision, recall, and F1-score are defined as follows:
Exact Match provides a lower bound on extraction performance and is commonly used in clinical NLP benchmarks, but it may underestimate clinically valid predictions when surface forms differ. Because the reference labels are derived from structured admission-level coding tables rather than token-level manual annotations, exact string matching may penalize predictions that are clinically correct but absent from billing codes, or may reward matches that reflect coding artifacts rather than explicit narrative documentation. The inclusion of fuzzy and semantic evaluation metrics partially mitigates this limitation by assessing clinical equivalence rather than strict code-level alignment.
5.2. Fuzzy Match
Fuzzy Match relaxes strict string equivalence by accounting for lexical variation using Levenshtein similarity. The similarity between two strings is defined as follows:
Predicted and ground-truth entity pairs with similarity scores of at least 60% are considered matches. This metric partially mitigates penalties arising from minor spelling differences, abbreviations, or morphological variants, but it remains limited in capturing deeper semantic equivalence.
5.3. Semantic Evaluation Using an LLM-As-Judge
To assess semantic correctness beyond surface-form similarity, we employ a large language model-based evaluation strategy using DeepSeek-v3 (December 2025) [
15] as an automated judge. The judge compares predicted and ground-truth entities and assigns one of three correctness labels: correct (1.0), partial (0.5), or incorrect (0.0). This grading scheme allows partial credit for predictions that are clinically related but differ in specificity or phrasing.
The evaluation prompt instructs the model to consider clinical equivalence rather than lexical identity, explicitly accounting for factors such as synonymy, abbreviation expansion, reordered phrases, and brand–generic medication equivalence. For example, predictions that differ only in specificity (e.g., “heart failure” vs. “chronic heart failure”) are treated as partial rather than incorrect matches.
Using the judge-assigned scores, precision, recall, and F1-score are computed as follows:
This semantic evaluation is intended to approximate clinically informed judgment at scale and to complement string-based metrics rather than replace them. While LLM-as-Judge evaluation introduces model-dependent biases and does not substitute for expert human adjudication, prior work has shown that it provides a more faithful estimate of clinical correctness than exact or fuzzy matching alone [
15]. In this study, semantic evaluation is used to highlight discrepancies between surface-level accuracy and clinically meaningful extraction quality across methods.
This semantic evaluation relies on a single LLM judge (DeepSeek) operating under a fixed evaluation prompt. No multi-judge ensemble, cross-model agreement analysis, or calibration against a manually adjudicated gold standard was performed. As such, semantic scores may reflect model-specific evaluation tendencies. In this study, LLM-as-Judge is used as a scalable approximation of clinically informed assessment rather than a definitive substitute for expert human adjudication. To partially assess clinical validity beyond automated scoring, a clinician evaluation study was conducted on a subset of test summaries (
Section 7.6).
6. Experimental Design
All experiments are conducted using the training, validation, and test splits described in
Section 4. Prompting-based extraction, retrieval-augmented prompting, and supervised fine-tuning are evaluated on the identical test set to ensure direct comparability across approaches. The experimental workflow consists of applying prompting strategies to validation and test discharge summaries, optionally post-processing prompting outputs using retrieval-augmented canonicalization, training and validating the supervised model on the training split, and evaluating all approaches using a unified metric suite.
Prompting experiments are performed using GPT-4o-mini via the OpenAI API and Qwen-2.5-7B with local inference. To ensure reproducibility and minimize stochastic variation, all prompting runs use a temperature of 0.0, a maximum token length of 2048, batch size of 1, and a fixed random seed of 42. Qwen-2.5-7B inference is executed on a Google Colab A100 GPU with 40 GB memory. Semantic evaluation is performed using DeepSeek, and all prompting templates and example constructions are provided in
Appendix A.
Each prompting configuration, including zero-shot, two-shot, four-shot, and Chain-of-Verification prompting, is applied to all 84 validation discharge summaries and 89 test summaries for both model backbones. This results in eight prompting conditions in total (four prompting strategies across two models). Latency and API usage costs are recorded for all prompting runs to support comparison across configurations.
6.1. RAG Canonicalization Experiments
RAG was applied strictly as a post-processing step to prompting outputs. The configuration used in all RAG experiments is summarized in
Table 1.
Entity embeddings were computed using S-PubMedBERT; retrieval used FAISS IndexFlatIP with top-k = 3 and a similarity threshold of 0.7. Knowledge bases were constructed from all unique ground-truth labels in the dataset. RAG enhanced outputs were evaluated against raw prompting outputs to isolate the effect of canonicalization.
6.2. Fine-Tuning Experiments
Bio ClinicalBERT was fine-tuned on the training data using the hyperparameters shown in
Table 2.
The Model was trained for up to 50 epochs with early stopping based on validation loss (patience = 10). A 10% linear warmup schedule was used, and optimizer settings followed standard AdamW configurations. Thresholds for converting sigmoid probabilities to binary predictions were tuned on the validation set via grid search. Top K caps (20 for diagnoses, 8 for procedures, 50 for medications) were applied during inference to limit over-prediction. The best-performing checkpoint was evaluated on the test set.
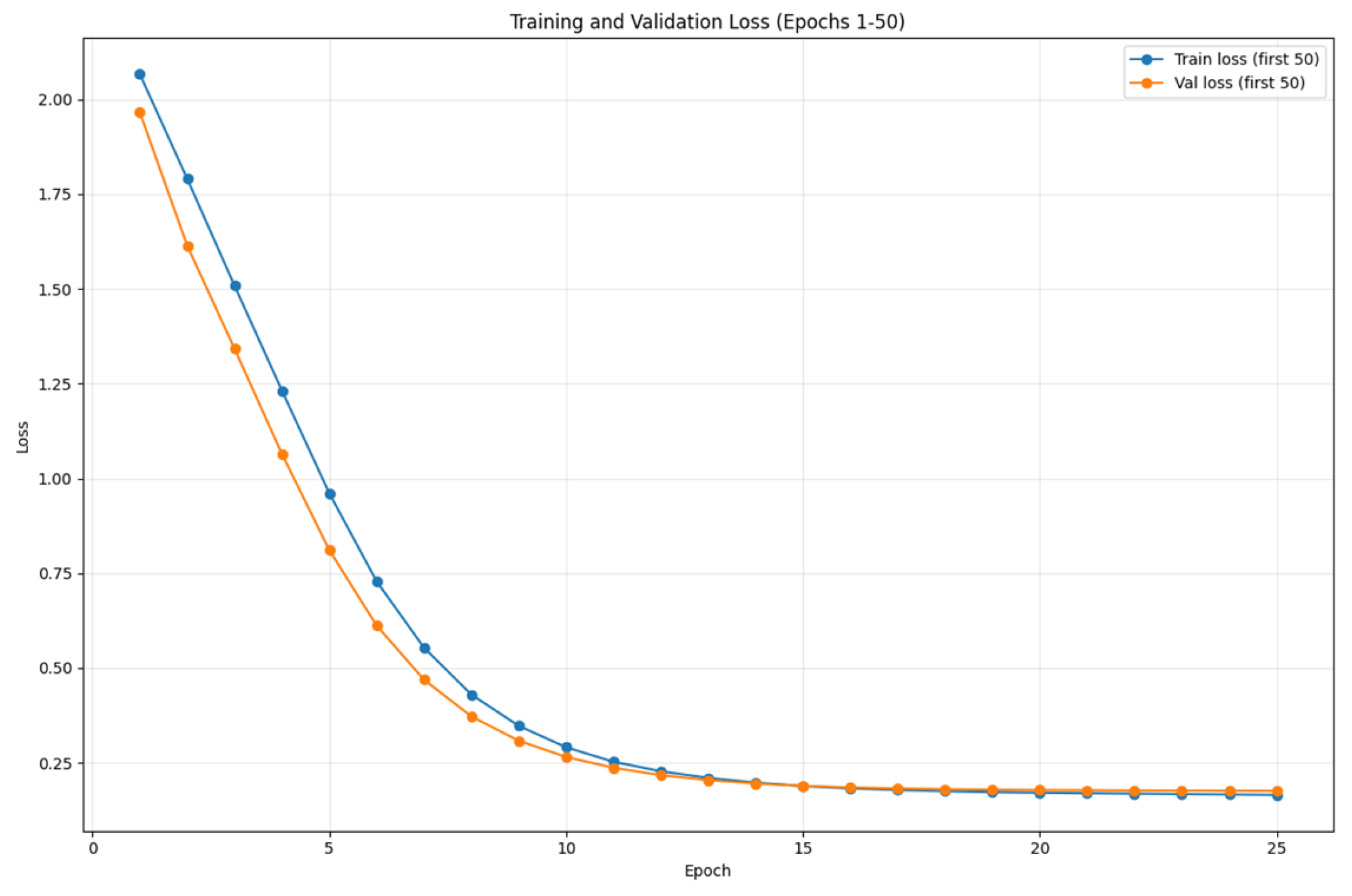
Figure 2 illustrates the training and validation loss trajectories across all epochs for Bio ClinicalBERT fine-tuning. Both losses decrease rapidly during the first 10 epochs, with training loss falling from 2.06 to 0.28 and validation loss from 1.96 to 0.26.
The curves converge closely until epoch 12, after which validation loss begins to plateau while training loss continues to decline. The minimum validation loss (0.19) occurs at epoch 15, which was selected as the optimal checkpoint under the early stopping criterion. Subsequent divergence indicates the onset of overfitting, validating the checkpoint selection procedure.
7. Experimental Results
7.1. Prompting-Based Extraction Performance
Table 3 and
Table 4 summarize the performance of GPT-4o-mini and Qwen 2.5-7B across prompting strategies and entity types. Across both models, semantic evaluation consistently yields substantially higher scores than exact or fuzzy matching, underscoring the extent to which string-based metrics underestimate clinically meaningful correctness in free-text extraction tasks.
For GPT-4o-mini (
Table 3), zero-shot prompting achieves the strongest overall semantic performance on the validation set (Judge F1 = 0.447), while few-shot prompting provides limited and inconsistent gains. Chain-of-Verification prompting does not yield systematic improvements over simpler prompting strategies. Across all prompting configurations, medication extraction consistently outperforms diagnosis and procedure extraction, achieving Judge F1 scores above 0.50. This pattern reflects lower lexical variability and clearer surface forms for medications relative to diagnoses and procedures in discharge summaries. Results for Qwen 2.5-7B (
Table 4) exhibit similar trends. CoVE prompting achieves the highest overall semantic performance (Judge F1 = 0.419), although differences across prompting strategies remain modest. As with GPT-4o-mini, medication extraction substantially outperforms other entity types. Despite running entirely locally, Qwen 2.5-7B attains approximately 94% of GPT-4o-mini’s zero-shot semantic performance, highlighting the competitiveness of open-source models under resource-constrained settings.
Across both models, the consistent gap between exact, fuzzy, and semantic scores reinforces the importance of evaluation frameworks that move beyond surface-form matching when assessing clinical extraction quality.
7.2. Effects of Retrieval-Augmented Canonicalization
Table 5 reports test-set performance for GPT-4o-mini with and without retrieval-augmented canonicalization. RAG yields localized improvements in fuzzy matching, particularly for diagnoses, where Fuzzy F1 increases from 0.123 to 0.162. Medication performance remains largely stable, while procedure extraction exhibits mixed behavior.
Overall performance differences between RAG-enhanced and non-RAG configurations are modest, indicating that retrieval-based canonicalization primarily improves output consistency rather than extraction coverage. These results suggest that RAG is most effective as a normalization mechanism for surface-form variability, rather than as a substitute for improved entity detection. It is important to emphasize that retrieval in this study is restricted to post-processing canonicalization against ground-truth-derived label sets. This configuration refines surface-form consistency but does not introduce new contextual knowledge or reasoning support during extraction. Therefore, the limited gains observed here reflect the behavior of this specific normalization-oriented implementation rather than a general limitation of retrieval-augmented generation in clinical NLP tasks.
7.3. Cost and Latency Analysis
In addition to extraction accuracy, practical deployment considerations are critical for clinical NLP systems.
Table 6 and
Table 7 summarize the monetary cost, token usage, and latency associated with each prompting configuration, enabling direct comparison of performance–efficiency trade-offs across model families and prompting strategies.
Evaluating all prompting strategies on the validation set requires a total cost of $0.366 and approximately 53 min of compute time. These results highlight the practical tradeoffs between supervision-free prompting strategies and resource consumption, particularly in large-scale or cost-sensitive deployment scenarios.
Zero-shot prompting consistently provides the strongest performance-to-cost ratio, achieving competitive semantic accuracy with the lowest token usage and inference cost. Few-shot prompting substantially increases input token counts without delivering consistent performance gains, resulting in diminishing returns under cost-sensitive settings. GPT-4o-mini exhibits lower per-request latency but incurs API cost, whereas Qwen-2.5-7B eliminates monetary cost at the expense of higher inference latency and local hardware requirements. These findings highlight trade-offs between supervision level, monetary cost, and computational overhead that must be considered in real-world deployment scenarios.
7.4. Supervised Fine-Tuning Results
Table 8 presents test-set performance for the fine-tuned BioClinicalBERT model. Compared with prompting-based approaches, supervised fine-tuning achieves substantially higher string-based performance, with an overall Exact F1 of 0.201 and Fuzzy F1 of 0.608. This improvement reflects the model’s ability to leverage labeled data to capture domain-specific terminology, even when surface forms vary. Performance gains are most pronounced for diagnoses and medications, while procedure extraction remains more challenging. The large discrepancy between Exact and Fuzzy scores further illustrates the extent of lexical variation present in discharge summaries and the limitations of strict string matching for evaluating supervised clinical NLP systems.
7.5. Comparative Analysis Across Extraction Paradigms
Table 9 directly compares the best-performing prompting configuration, its RAG-enhanced variant, and the supervised fine-tuned model. Supervised fine-tuning achieves the strongest performance across string-based metrics, improving Exact F1 from 0.125 to 0.201 and Fuzzy F1 from 0.236 to 0.608 relative to zero-shot prompting. RAG-enhanced prompting maintains similar fuzzy performance to prompting alone but does not yield consistent gains in overall extraction accuracy.
These results highlight a clear tradeoff between supervision requirements and performance: prompting-based methods provide competitive semantic correctness with minimal labeled data, while supervised fine-tuning delivers higher string-level accuracy when labeled data are available.
7.6. Clinician Evaluation of Extraction Quality
To complement automated evaluation metrics, a board-certified physician independently evaluated 46 discharge summaries sampled from the test set. For each note, the clinician was provided with the full discharge summary and the corresponding extracted diagnoses, procedures, and medications. Each extraction was rated on a 1–5 Likert scale across three dimensions: (1) extraction completeness, (2) normalization correctness, and (3) clinical plausibility.
As shown in
Table 10, across the 46 evaluated notes, mean ratings were 3.73 ± 0.54 for extraction completeness, 3.71 ± 0.51 for normalization correctness, and 3.60 ± 0.54 for clinical plausibility. Additionally, 68.9% of notes were rated ≥4 for completeness and normalization, and 57.8% were rated ≥4 for clinical plausibility. These findings provide independent expert validation of extraction quality and partially support the trends observed under semantic evaluation, while acknowledging that only a single clinician participated in the assessment.
7.7. Benchmark Contextualization
To clarify methodological positioning relative to prior EHR-LLM benchmarks,
Table 11 contrasts the present study with representative systems across dataset domain, task formulation, model families evaluated, retrieval usage, supervision strategy, evaluation methodology, and deployment analysis. This comparison emphasizes differences in experimental design rather than direct performance ranking.
Table 12 positions the reported results within the broader landscape of clinical information extraction. As expected, domain-specific transformer models fine-tuned on established benchmarks, such as RoBERTa-MIMIC, continue to achieve strong performance on curated datasets. In this context, the fine-tuned BioClinicalBERT model attains a Fuzzy F1 of 0.608 on MIMIC-IV discharge summaries, reflecting the increased lexical variability and complexity of this corpus relative to more homogeneous benchmarks.
Prompting-based LLM approaches perform competitively in zero-shot settings, with GPT-4o-mini achieving a Judge F1 of 0.447 on the proposed task. Retrieval-augmented prompting yields more limited benefits in this setting compared with prior work such as DiRAG, likely due to dataset-specific terminology patterns and differences in knowledge base construction. Together, these results emphasize that extraction performance is highly sensitive to dataset characteristics, evaluation methodology, and the availability of labeled supervision.
8. Discussion and Limitations
This study provides a systematic comparison of prompting-based extraction, retrieval-augmented canonicalization, and supervised fine-tuning for structured clinical information extraction, highlighting tradeoffs that depend on model capacity, supervision availability, entity type, and evaluation methodology.
Across both GPT-4o-mini and Qwen 2.5-7B, zero-shot prompting consistently achieves performance comparable to, and in some cases exceeding, few-shot configurations. This behavior contrasts with conventional expectations from in-context learning and suggests that pretrained LLMs may already encode sufficient medical knowledge for structured extraction tasks. In this setting, additional examples can introduce noise or distract from adherence to a fixed output schema, particularly when strict JSON formatting is required. Chain-of-Verification prompting exhibits more consistent benefits for the smaller open-source model, indicating that explicit verification and reasoning scaffolds may compensate for more limited parametric knowledge, whereas larger models appear less sensitive to prompt structure [
25].
Performance varies substantially across entity types, reflecting underlying linguistic characteristics of clinical documentation. Medication extraction is consistently strongest across all methods, likely due to more standardized naming conventions and reduced contextual ambiguity. Diagnoses exhibit greater lexical variability, leading to low exact-match scores but substantially higher fuzzy and semantic performance. Procedures remain the most challenging category, with highly variable phrasing and implicit references that complicate both extraction and normalization. These patterns align with prior clinical NLP findings and suggest that entity-type-specific modeling and evaluation strategies remain necessary.
The comparison between closed-source and open-source models indicates that locally deployable LLMs are increasingly viable for clinical extraction tasks. Qwen 2.5-7B achieves approximately 94% of GPT-4o-mini’s zero-shot semantic performance while operating entirely without API costs, albeit with higher latency. This tradeoff highlights the potential of open-source models in privacy-sensitive or resource-constrained environments, particularly when paired with structured prompting strategies. From a deployment perspective, zero-shot prompting emerges as the most efficient supervision-light strategy, while supervised fine-tuning requires labeled data and substantial upfront training cost but offers low marginal inference cost once deployed.
Evaluation methodology strongly influences observed performance. Large gaps between Exact Match and semantic scores demonstrate that string-based metrics substantially underestimate clinically meaningful correctness in free-text extraction. Fuzzy matching partially mitigates this issue but remains insufficient to capture true clinical equivalence. Semantic evaluation using an LLM-as-Judge provides a more informative assessment of extraction quality, though it introduces model-dependent biases and does not replace expert human adjudication. In this work, semantic evaluation is used to complement, rather than supersede, string-based metrics.
Retrieval-augmented canonicalization improves terminology consistency and yields localized gains, particularly for diagnoses, but does not substantially improve overall extraction accuracy [
26]. This outcome is expected given that canonicalization refines only already-extracted entities and cannot recover missed predictions. Its effectiveness depends on alignment between predicted phrases and the canonical vocabulary, and mismatches may improve semantic interpretability at the expense of strict string matching. These findings suggest that RAG is better suited for post-processing and harmonization than as a standalone accuracy-enhancing mechanism [
27]. Supervised fine-tuning of BioClinicalBERT achieves the strongest string-based performance, particularly under fuzzy evaluation, reflecting the benefits of labeled supervision for capturing domain-specific terminology. However, performance varies by entity type and exhibits recall-heavy tendencies, especially for diagnoses and procedures. These patterns likely reflect class imbalance, large label spaces, and limited annotated data, underscoring the continued importance of domain-specific pretraining, careful threshold calibration, and improved learning objectives for clinical extraction.
Several limitations should be noted. The analysis is conducted on discharge summaries from a single institution, which may limit generalizability to settings with different documentation practices. The filtered dataset, while suitable for controlled comparison, is small for supervised learning and may not capture rare conditions or procedural descriptions. Temporal variation in clinical terminology across the 2008–2022 period is not explicitly modeled. The RAG knowledge base is derived from the same dataset used for evaluation, raising potential concerns about circularity, and retrieval parameters are fixed rather than optimized per entity type. Finally, cost and resource constraints prevent full semantic evaluation of the fine-tuned model and limit latency measurements to a single hardware configuration.
Several additional limitations warrant emphasis. The analytic dataset comprised 449 discharge summaries from 300 unique patients, representing a small filtered subset of the original MIMIC-IV cohort. While this design enables controlled comparison across methods, it limits representativeness and statistical generalizability. Moreover, ground-truth labels were derived from admission-level structured coding tables rather than manually annotated discharge summaries. As a result, the reference standard does not constitute a curated gold-standard dataset at the narrative entity level and may introduce label noise due to documentation–coding mismatches. The semantic evaluation depends on a single LLM judge without ensemble verification or systematic calibration against multiple human raters. Although a clinician evaluation was conducted on a subset of notes, only one clinician participated and inter-rater reliability was not assessed. Future work should incorporate larger cohorts and multi-rater manual annotation to strengthen evaluation robustness.
Taken together, these findings emphasize that no single extraction paradigm dominates across all conditions. Prompting-based methods offer strong semantic performance with minimal supervision, supervised fine-tuning yields higher string-level accuracy when labeled data are available, and retrieval-based canonicalization improves consistency rather than coverage. The choice of extraction strategy should therefore be guided by data availability, resource constraints, and evaluation objectives, particularly in clinical settings where semantic correctness is often more relevant than surface-form matching.
9. Conclusions
This study systematically compares prompting-based extraction, retrieval-augmented canonicalization, and supervised fine-tuning for structured clinical information extraction from MIMIC-IV discharge summaries under a unified experimental and evaluation framework. The results clarify how these approaches differ in accuracy, consistency, cost, and practical applicability. Prompting-based extraction performs strongly in zero-shot settings, with both GPT-4o-mini and Qwen 2.5-7B achieving meaningful semantic accuracy without task-specific supervision. Few-shot prompting does not yield consistent gains, suggesting that pretrained language models already encode substantial clinical knowledge and benefit more from structured reasoning mechanisms, such as Chain-of-Verification, than from additional examples. Retrieval-augmented generation improves terminology consistency and interpretability but does not materially increase extraction coverage, indicating that its primary role is post-processing rather than recall enhancement. Supervised fine-tuning with BioClinicalBERT achieves the strongest string-based performance, particularly under fuzzy evaluation, though effectiveness varies by entity type due to differences in lexical variability, label space size, and data sparsity.
From a deployment perspective, open-source models show increasing practical viability. Qwen 2.5-7B approaches the semantic performance of GPT-4o-mini while operating locally and without inference cost, making it suitable for privacy-sensitive or resource-constrained clinical settings. Across all approaches, evaluation results are highly sensitive to metric choice: exact string matching substantially underestimates clinically meaningful correctness, while semantic evaluation provides a more informative, though imperfect, assessment. These findings primarily reflect controlled comparative behavior across extraction paradigms within a filtered subset of MIMIC-IV and should be validated at a larger scale in future work.