α-Amidoamids as New Replacements of Antibiotics—Research on the Chosen K12, R2–R4 E. coli Strains
Abstract
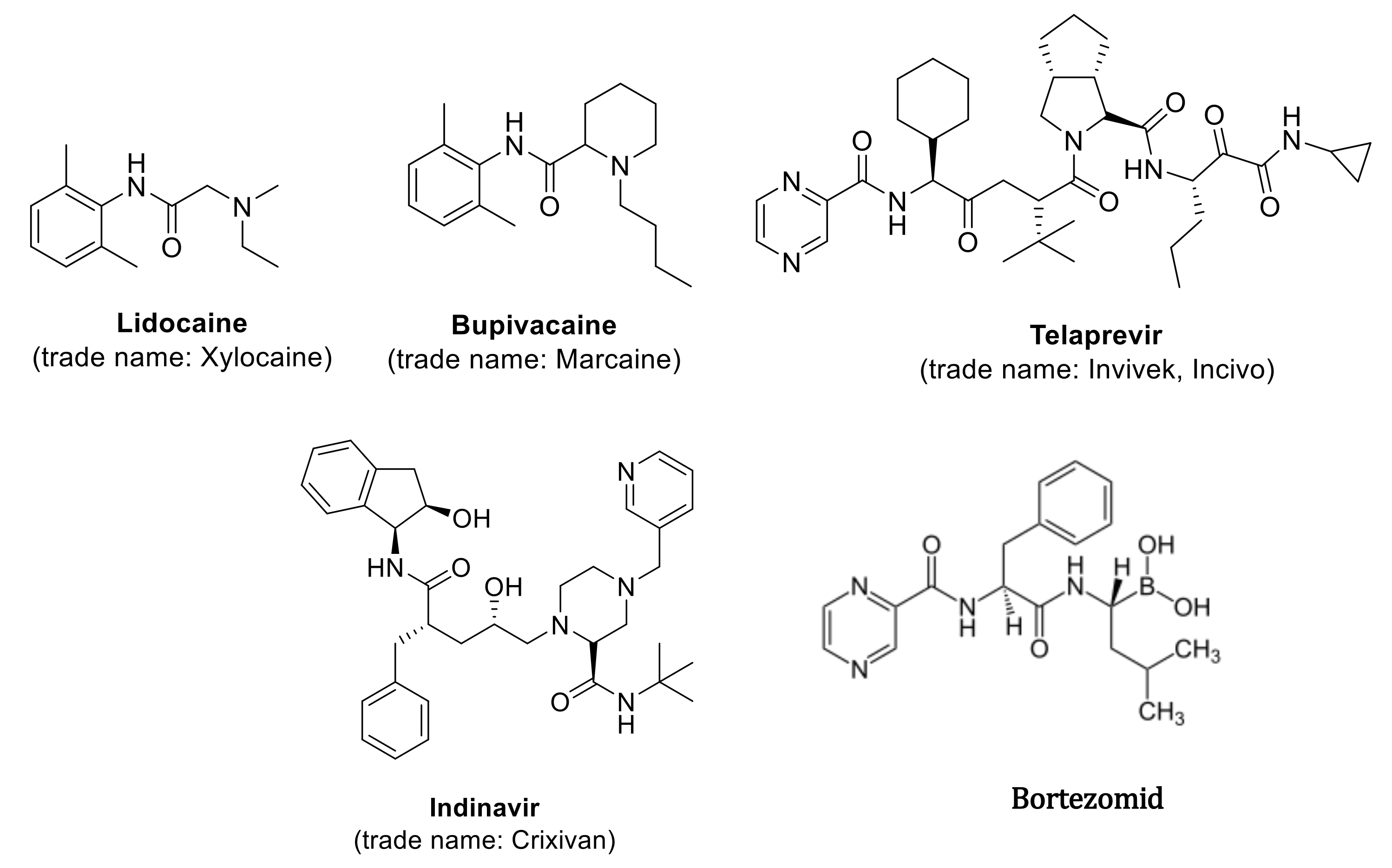
1. Introduction
2. Materials and Methods
2.1. Microorganisms and Media
2.2. Experimental Chemistry
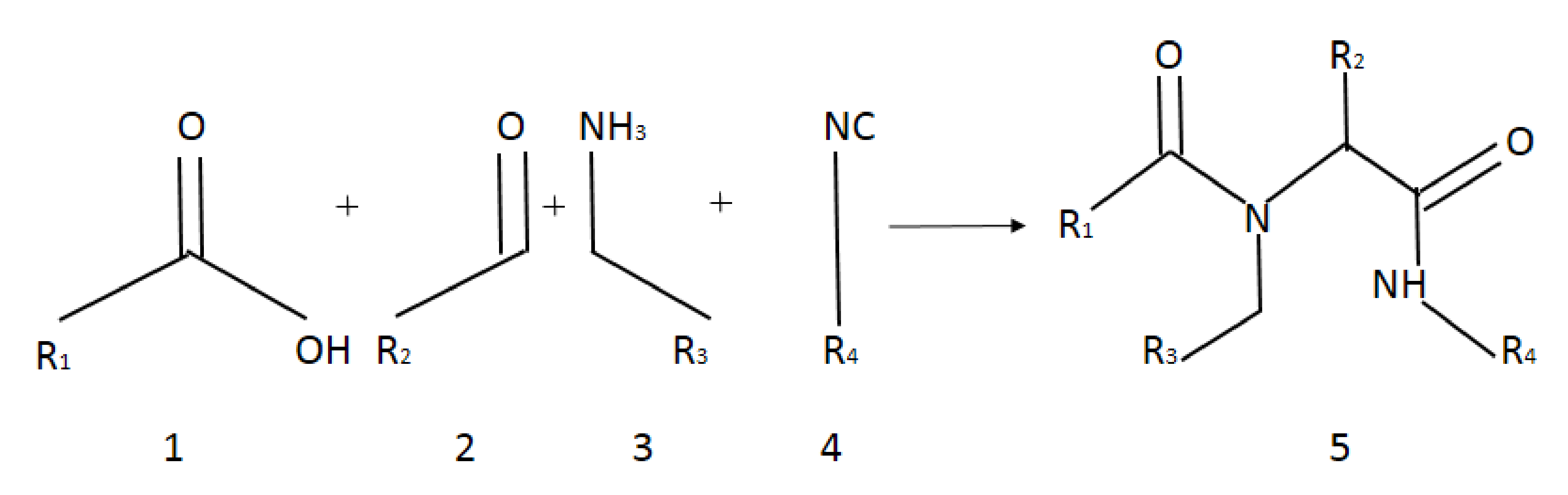
2.3. General Procedure for Synthesis of Compounds 1–20
2.3.1. Product 5a (AM 93)
2.3.2. Product 5b (AM 70)
2.3.3. Product 5c (AM 121)
2.3.4. Product 5d (AM 84)
2.3.5. Product 5e (AM 119)
2.3.6. Product 5f (AM 116)
2.3.7. Product 5g (AM 115)
2.3.8. Product 5h (AM 164)
2.3.9. Product 5i (AM 165)
2.3.10. Product 5j (AM 107)
2.3.11. Product 5k (AM 91)
2.3.12. Product 5l (AM 163)
2.3.13. Product 5m (AM 162)
2.3.14. Product 5n (AM 182)
2.3.15. Product 5o (AM 111)
2.3.16. Product 5p (AM 114)
2.3.17. Product 5r (AM 170)
2.3.18. Product 15s (AM 178)
2.3.19. Product 5t (AM 171)
2.3.20. Product 5u (AM 184)
2.4. Determination of MIC and MBC
2.4.1. Interaction of Bleomycine, Kanamycine, Streptomycine, with Bacterial Plasmid DNA Isolated from K12 and R1–R4 Strains
2.4.2. Determination of MIC and MBC after Antibiotics Treatment
2.5. Interaction of the Plasmid DNA from K12 and R4 Strains with α-Amidoamids
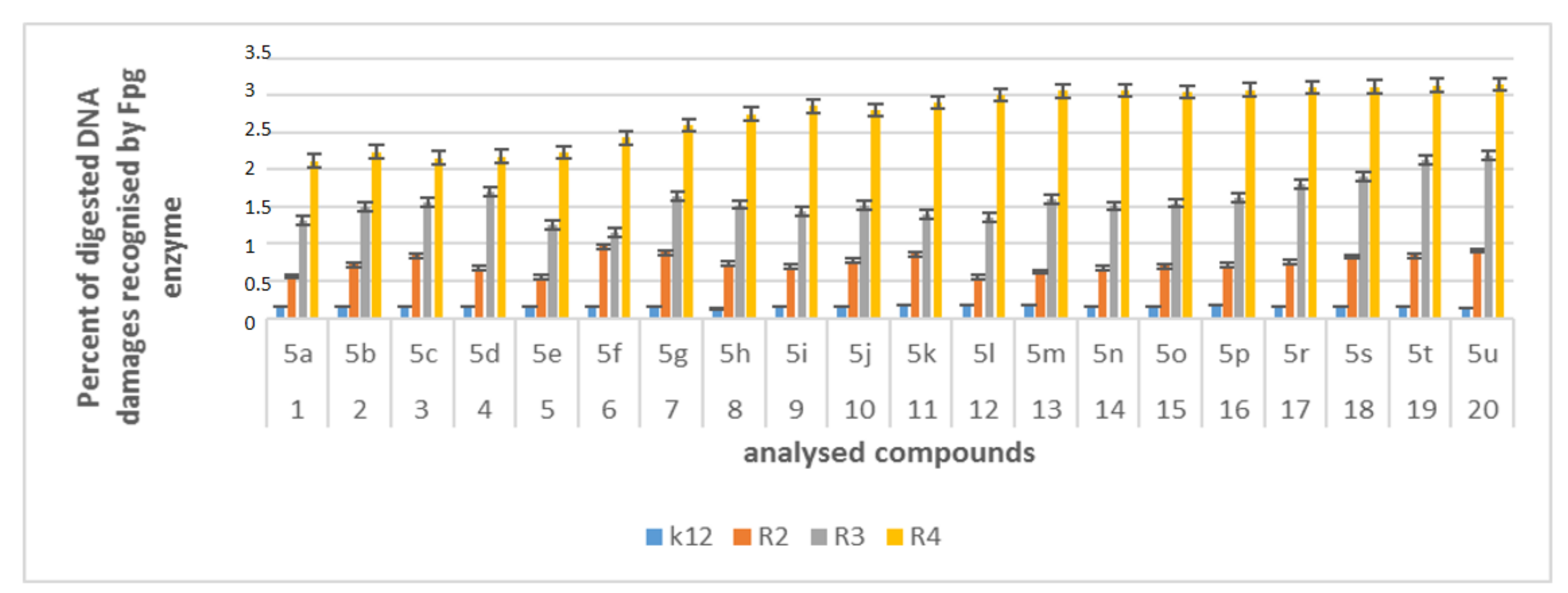
2.6. Repair and Cleavage of Oxidative DNA Damage Adducts by Fpg Protein in Bacterial Cells
2.7. Statistical Analysis
3. Results
3.1. Chemistry
3.2. Toxicity of Tested Compounds
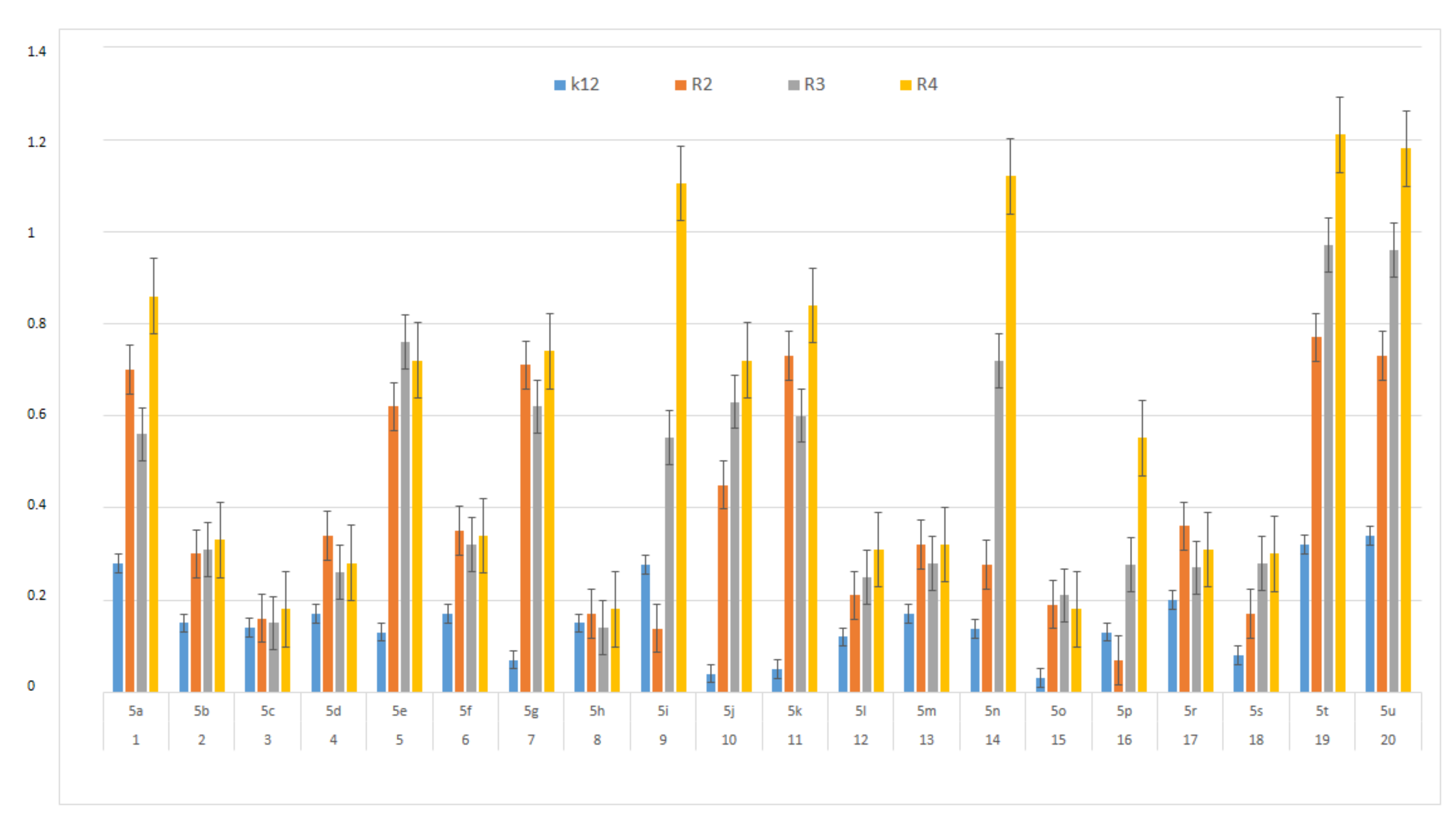
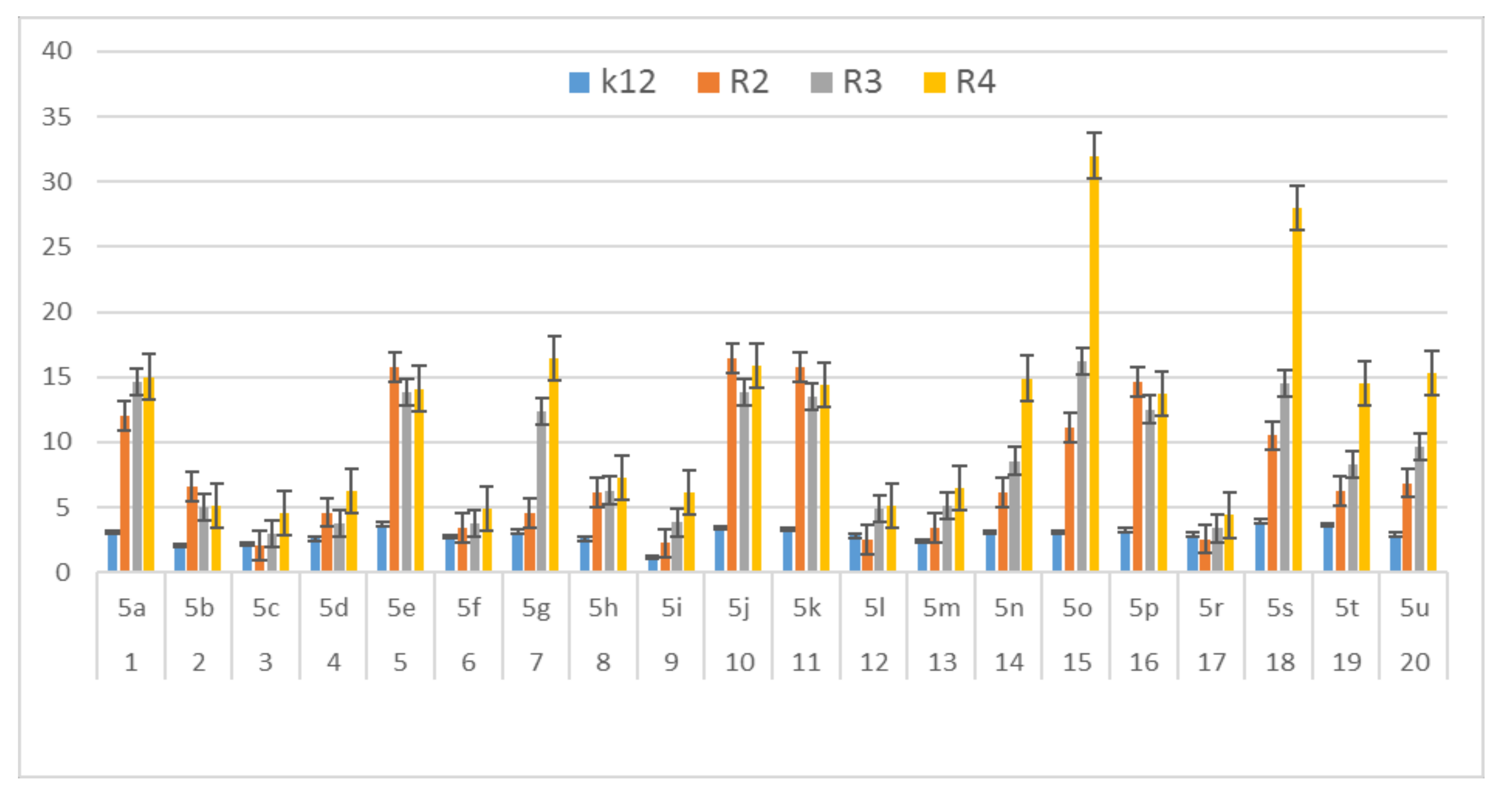
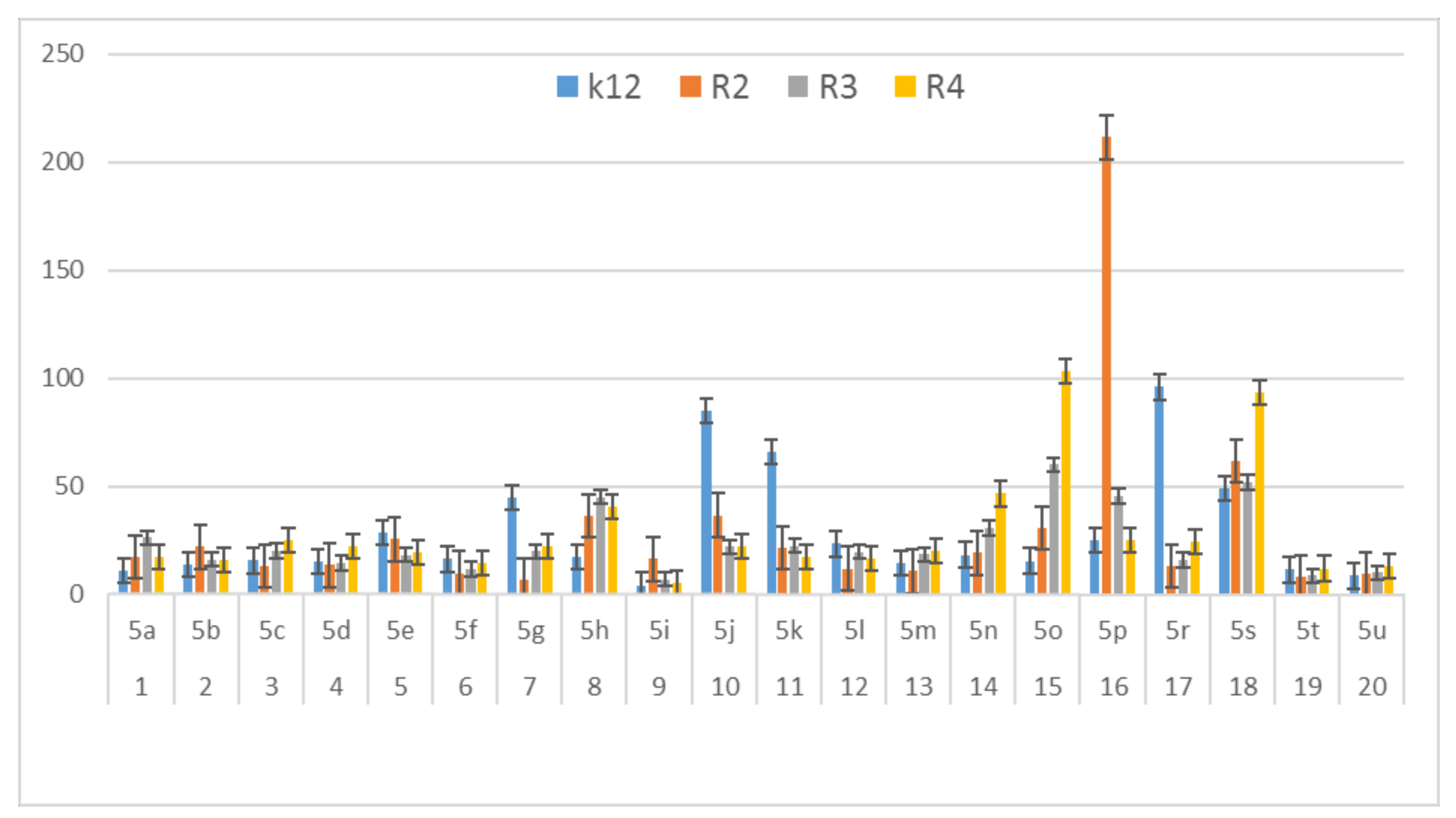
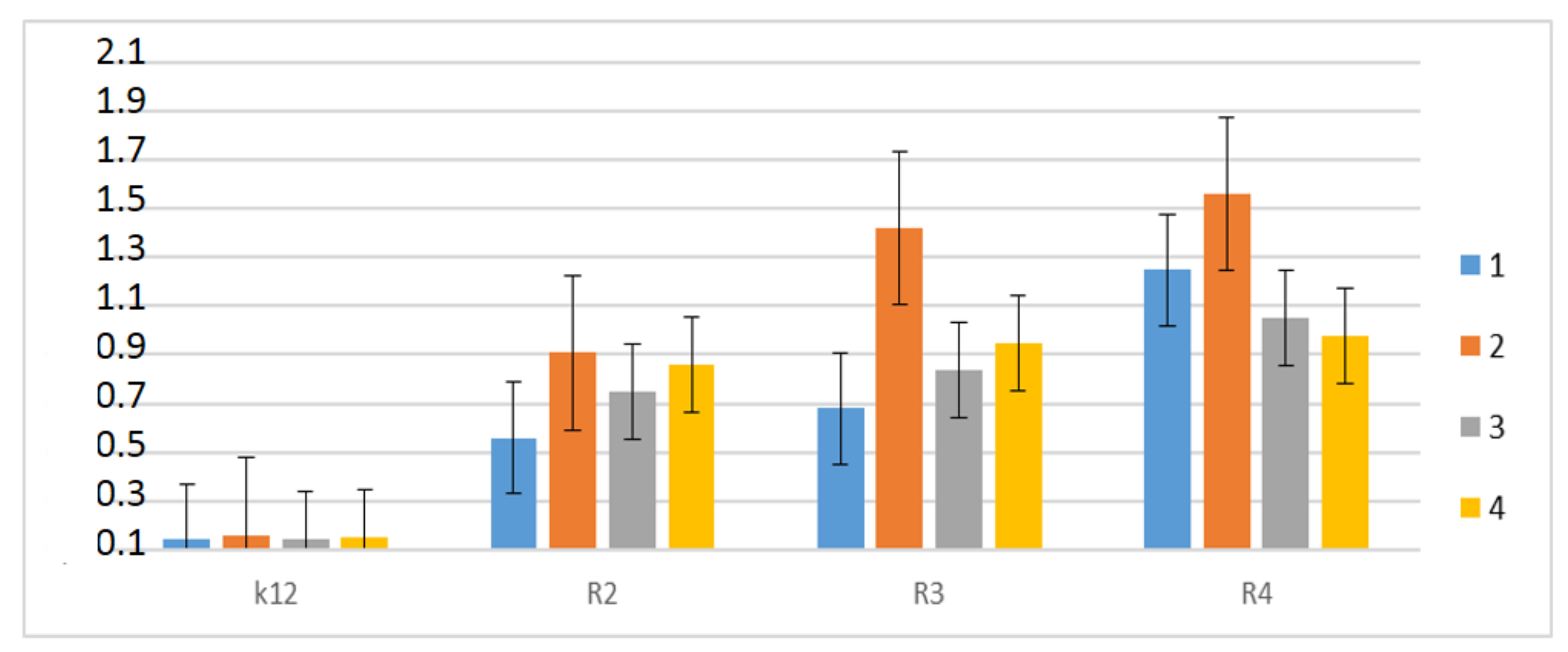
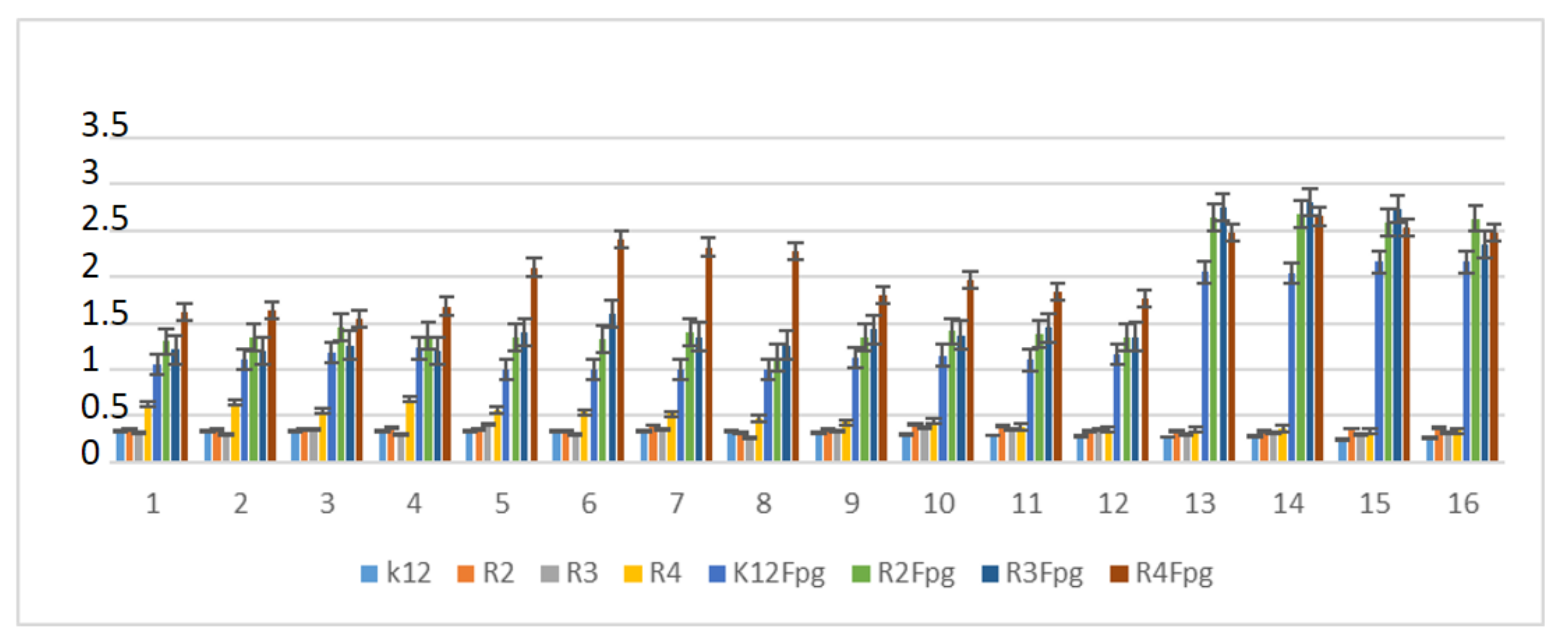
3.3. Modification of Plasmid DNA Isolated from E. coli R2–R4 Strains with Tested α-Amidoamids
4. Discussion
5. Conclusions
- -
- The R4-type strain was the most sensitive among the tested E. coli strains.
- -
- The insertion of analysed compounds into the leaflet of the outer membrane of the E. coli K-12 and rough strains showed that differences in the O-antigen and truncated oligosaccharide core may play important roles in the cellular response to alpha-amidoamids.
- -
- The toxicity of alkyl groups depends on their interaction with the membrane, which can incorporate into cell wall structures and change their hydrophobicity.
- -
- Membrane rearrangements and disruption may, in turn, result in changes of bacterial responses to other biologically active compounds such as antibiotics [79].
- -
- Plasmid DNA damage has been associated with the structure of verified peptidomimetics, suggesting that the presence of R4 tetr-butyl or 2,5-dimethoxybenzyl groups influences bacterial LPS and generates oxidative stress, which was already observed in our previous studies [50].
- -
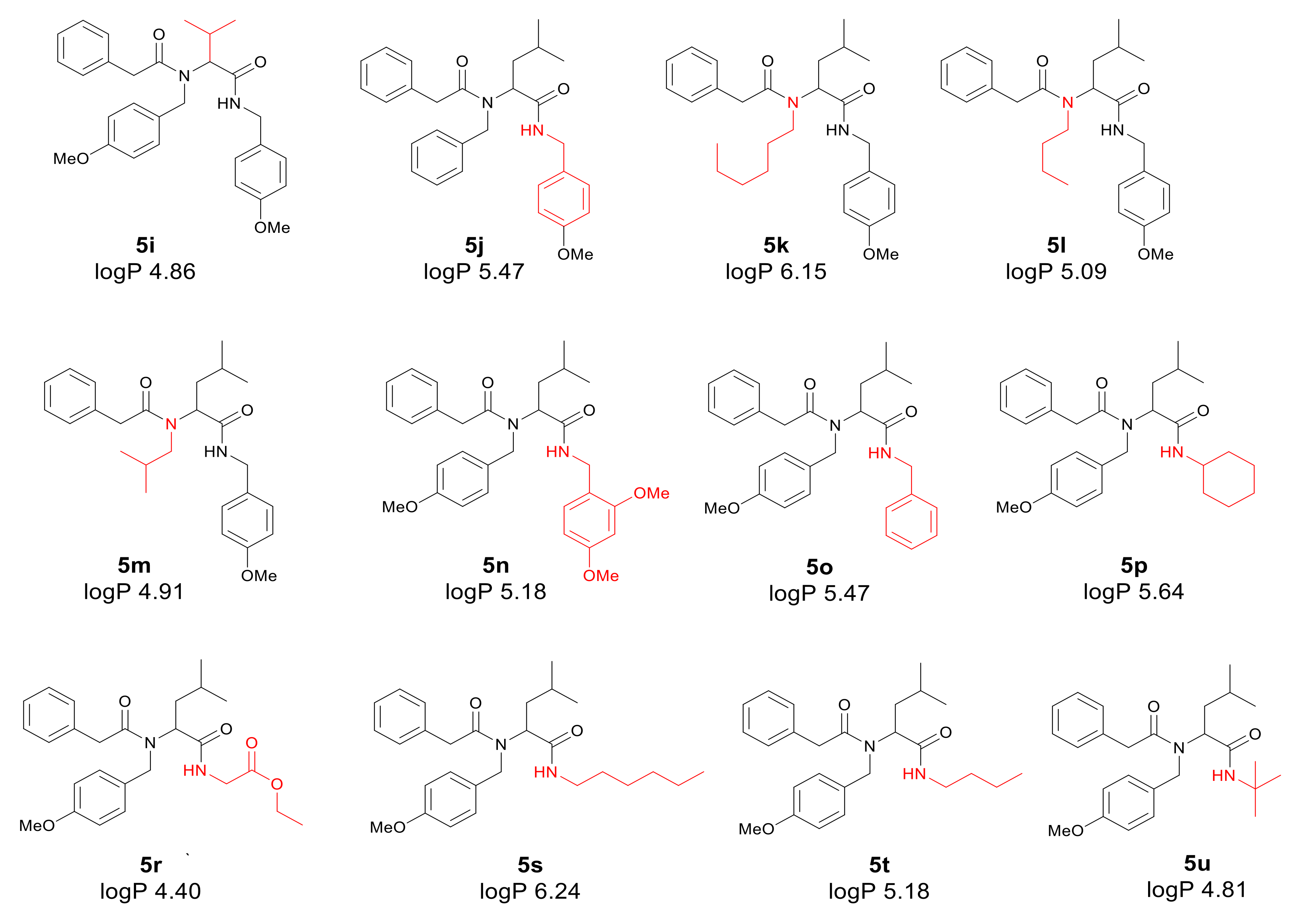
- The tested α-amidoamides 5 show a different influence on the MIC, which is strongly correlated with the steric factor of the R1, R2, R3 and R4 functional groups, and the presence of a methyl group with short alkyl chain in the structure of peptidomimetics 5 [50].
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Compliance with Ethical Standards
Abbreviations
| MIC | Minimum inhibitory concentration; |
| MBC | Minimum bactericidal concentration; |
| oc | open circle; |
| ccc | covalently-closed circle; |
| BER | base excision repair; |
| ROS | reactive oxygen species; |
| RNS | reactive nitrogen species. |
References
- Mroczkiewicz, M.; Winkler, K.; Nowis, D.; Placha, G.; Golab, J.; Ostaszewski, R. Studies of the Synthesis of All Stereoisomers of MG-132 Proteasome Inhibitors in the Tumor Targeting Approach. J. Med. Chem. 2010, 53, 1509–1518. [Google Scholar] [CrossRef] [PubMed]
- Tolomelli, A.; Squassabia, F. Peptides and Peptidomimetics in Medicine, Surgery and Biotechnology. Curr. Med. Chem. 2006, 13, 2449–2466. [Google Scholar] [CrossRef]
- Vázquez, J.; Durán, A.; Amado, I.R.; Prieto, M.; Rial, D.; Murado, M.A. Evaluation of toxic effects of several carboxylic acids on bacterial growth by toxicodynamic modelling. Microb. Cell Factories 2011, 10, 100. [Google Scholar] [CrossRef] [PubMed]
- Hatahet, Z.; Kow, Y.W.; Purmal, A.A.; Cunningham, R.P.; Wallace, S.S. New substrates for old enzymes. J. Biol. Chem. 1994, 269, 18814–18820. [Google Scholar] [PubMed]
- Ugi, I. The Multicomponent Reactions and their Libraries for Natural and Preparative Chemistry. Comb. Chem. High Throughput Screen. 1970, 4, 1–34. [Google Scholar] [CrossRef]
- Bienaymé, H.; Hulme, C.; Oddon, G.; Schmitt, P. Maximizing Synthetic Efficiency: Multi-Component Transformations Lead the Way. Chem. Eur. J. 2000, 6, 3321–3329. [Google Scholar] [CrossRef]
- El Kaïm, L.; Gizolme, M.; Grimaud, A.L.; Oble, J. Direct Access to Heterocyclic Scaffolds by New Multicomponent Ugi−Smiles Couplings. Org. Lett. 2006, 8, 4019–4021. [Google Scholar] [CrossRef]
- Tempest, P.A. Recent advances in heterocycle generation using the efficient Ugi multiple-component condensation reaction. Curr. Opin. Drug Discov. Devel. 2005, 8, 776–788. [Google Scholar] [CrossRef]
- Pirrung, M.C.; Sarma, K.D. Multicomponent Reactions Are Accelerated in Water. J. Am. Chem. Soc. 2004, 126, 444–445. [Google Scholar] [CrossRef]
- Ilyin, A.; Kysil, V.; Krasavin, M.; Kurashvili, I.; Ivachtchenko, A.V. Complexity-Enhancing Acid-Promoted Rearrangement of Tricyclic Products of Tandem Ugi 4CC/Intramolecular Diels−Alder Reaction. J. Org. Chem. 2006, 71, 9544–9547. [Google Scholar] [CrossRef]
- Dormán, G.; Frank, R. Small Molecule Microarrays—Mall and smart. QSAR Comb. Sci. 2006, 25, 1007–1008. [Google Scholar] [CrossRef]
- Özogul, Y.; Özogul, F. Chapter 1: Biogenic Amines Formation, Toxicity, Regulations in Food. In Biogenic Amines in Food: Analysis, Occurrence and Toxicity; From Book Series: Food Chemistry, Function and Analysis; Royal Society of Chemistry: London, UK, 2019; pp. 1–17. ISBN 978-1-78801-581-3. [Google Scholar] [CrossRef]
- Ugi, I.; Meyr, R.; Fetzer, U.; Steinbrückner, C. Versuche mit Isonitrilen. Angew. Chem. 1959, 71, 386. [Google Scholar]
- Ugi, I.; Steinbrückner, C. Über ein neues Kondensations-Prinzip. Angew. Chem. 1960, 72, 267–268. [Google Scholar] [CrossRef]
- Ugi, I.; Werner, B.; Dömling, A. The Chemistry of Isocyanides, their MultiComponent Reactions and their Libraries. Molecules 2003, 8, 53–66. [Google Scholar] [CrossRef]
- Zhang, J.; Jacobson, A.; Rusche, J.R.; Herlihy, W. Unique Structures Generated by Ugi 3CC Reactions Using Bifunctional Starting Materials Containing Aldehyde and Carboxylic Acid. J. Org. Chem. 1999, 64, 1074–1076. [Google Scholar] [CrossRef] [PubMed]
- Xiang, Z.; Luo, T.; Lu, K.; Cui, J.; Shi, X.; Fathi, R.; Chen, J.; Yang, Z. Novel Pd-II-mediated cascade carboxylative annulation to construct benzo[b]furan-3-carboxylic acids. Org. Lett. 2004, 6, 3155–3158. [Google Scholar] [CrossRef]
- Banfi, L.; Riva, R. The Passerini Reaction. In Organic Reactions; Overman, L.E., Ed.; Wiley: Hoboken, NJ, USA, 2005; Volume 65, ISBN 0-471-68260-8. [Google Scholar]
- Ugi, I.; Lohberger, S.; Karl, R. Comprehensive organic synthesis. In The Passerini and Ugi Reactions; Pergamon: Oxford, UK, 1991; pp. 1083–1109. ISBN 0-08-040593-2. [Google Scholar]
- Ugi, I. The α-Addition of Immonium Ions and Anions to Isonitriles Accompanied by Secondary Reactions. Angew. Chem. Int. Ed. Engl. 1962, 1, 8–21. [Google Scholar] [CrossRef]
- DömLing, A.; Ugi, I. Multicomponent reactions with isocyanides. Angew. Chem. Int. Ed. Engl. 2000, 39, 3168–3210. [Google Scholar] [CrossRef]
- Denmark, S.E.; Fan, Y. Catalytic, Enantioselective α-Additions of Isocyanides: Lewis Base Catalyzed Passerini-Type Reactions. J. Org. Chem. 2005, 70, 9667–9676. [Google Scholar] [CrossRef]
- Bossio, R.; Marcaccini, S.; Pepino, R.; Torroba, T. Synthesis studies on isocyanides and related compounds: A novel synthetic route to furan derivatives. Synthesis 1993, 8, 783–785. [Google Scholar] [CrossRef]
- Veena, K.S.; Taniya, M.S.; Ravindran, J.; Thangarasu, A.K.; Priya, S.; Lankalapalli, R.S. Semi-synthetic diversification of coronarin D, a labdane diterpene, under Ugi reaction conditions. Nat. Prod. Res. 2020, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Szymanski, W.; Ostaszewski, R. Toward stereocontrolled, chemoenzymatic synthesis of unnatural peptides. Tetrahedron 2008, 64, 3197–3203. [Google Scholar] [CrossRef]
- Szymanski, W.; Ostaszewski, R. Multicomponent diversity and enzymatic enantioselectivity as a route towards both enantiomers of α-amino acids—A model study. Tetrahedron Asymmetry 2006, 17, 2667–2671. [Google Scholar] [CrossRef]
- Rossen, K.; Pye, P.; DiMichele, L.; Volante, R.; Reider, P. An efficient asymmetric hydrogenation approach to the synthesis of the Crixivan® piperazine intermediate. Tetrahedron Lett. 1998, 39, 6823–6826. [Google Scholar] [CrossRef]
- Ghosh, A.K.; Kincaid, J.F.; Cho, W.; Walters, D.; Krishnan, K.; Hussain, K.A.; Koo, Y.; Cho, H.; Rudall, C.; Holland, L.; et al. Potent HIV protease inhibitors incorporating high-affinity P2-ligands and (R)-(hydroxyethylamino)sulfonamide isostere. Bioorganic Med. Chem. Lett. 1998, 8, 687–690. [Google Scholar] [CrossRef]
- Zhao, H.; Neamati, N.; Hong, H.; Mazumder, A.; Wang, S.; Sunder, S.; Milne, G.W.A.; Pommier, Y.; Burke, T.R. Coumarin-Based Inhibitors of HIV Integrase1. J. Med. Chem. 1997, 40, 242–249. [Google Scholar] [CrossRef] [PubMed]
- Marieb, E.N.; Hoehn, K. Human Anatomy & Physiology, 9th ed.; Benjamin Cummings: San Francisco, CA, USA, 2012; p. 599. [Google Scholar]
- Madej, A.; Paprocki, D.; Koszelewski, D.; Żądło-Dobrowolska, A.; Brzozowska-Elliott, A.; Walde, P.; Ostaszewski, R. Efficient Ugi reactions in an aqueous vesicle system. RSC Adv. 2017, 7, 33344–33354. [Google Scholar] [CrossRef]
- Dömling, A.; Wang, W.; Wang, K. Chemistry and Biology of Multicomponent Reactions. Chem. Rev. 2012, 112, 3083–3135. [Google Scholar] [CrossRef]
- Bartus, R.T.; Baker, K.L.; Heiser, A.D.; Sawyer, S.D.; Dean, R.L.; Elliott, P.J.; Straub, J.A.; Cereb, J.J. Postischemic administration of AK275, a calpain inhibitor, provides substantial protection against focal ischemic brain damage. Blood Flow Metab. 1994, 14, 537–544. [Google Scholar] [CrossRef]
- Zeldin, K.R.; Petruschke, R.A. Pharmacological and therapeutic properties of ritonavir-boosted protease inhibitor therapy in HIV-infected patients. J. Antimicrob. Chem. 2004, 53, 4–9. [Google Scholar] [CrossRef]
- Tan, C.R.C.; Abdul-Majeed, S.; Cael, B.; Barta, S.K. Clinical Pharmacokinetics and Pharmacodynamics of Bortezomib. Clin. Pharmacokinet. 2019, 58, 157–168. [Google Scholar] [CrossRef]
- Tsetlin, V. Snake venom alpha-neurotoxins and other ‘three-finger’ proteins. JBIC J. Biol. Inorg. Chem. 1999, 264, 281–286. [Google Scholar] [CrossRef] [PubMed]
- Xue, H.; Lu, X.; Zheng, P.; Liu, L.; Han, C.; Hu, J.; Liu, Z.; Ma, T.; Li, Y.; Wang, L.; et al. Highly Suppressing Wild-Type HIV-1 and Y181C Mutant HIV-1 Strains by 10-Chloromethyl-11-demethyl-12-oxo-calanolide A with Druggable Profile. J. Med. Chem. 2010, 53, 1397–1401. [Google Scholar] [CrossRef] [PubMed]
- Klossowski, S.; Muchowicz, A.; Swiech, M.; Firczuk, M.; Redzej, A.; Golab, J.; Ostaszewski, R. Studies toward novel peptidomimetic inhibitors of thioredoxin-thioredoxin reductase system. J. Med. Chem. 2012, 55, 55–67. [Google Scholar] [CrossRef] [PubMed]
- Żądło-Dobrowolska, A.; Kłossowski, S.; Koszelewski, D.; Paprocki, D.; Ostaszewski, R. Enzymatic Ugi reaction with amines and cyclic imines. Chemistry 2016, 22, 16684–16689. [Google Scholar] [CrossRef]
- Zaorska, E.; Tomasova, L.; Koszelewski, D.; Ostaszewski, R.; Ufnal, M. Hydrogen sulfide in pharmacotherapy, beyond the hydrogen sulfide-donors. Biomolecules 2020, 10, 323. [Google Scholar] [CrossRef] [PubMed]
- Von Minckwitz, G.; Huang, C.-S.; Mano, M.S.; Loibl, S.; Mamounas, E.P.; Untch, M.; Wolmark, N.; Rastogi, P.; Schneeweiss, A.; Redondo, A.; et al. Trastuzumab Emtansine for Residual Invasive HER2-Positive Breast Cancer. N. Engl. J. Med. 2019, 380, 617–628. [Google Scholar] [CrossRef]
- Von Minckwitz, G.; Procter, M.; De Azambuja, E.; Zardavas, D.; Benyunes, M.M.; Viale, G.; Suter, T.; Arahmani, A.A.; Rouchet, N.N.; Clark, E.E.; et al. Adjuvant Pertuzumab and Trastuzumab in Early HER2-Positive Breast Cancer. Steering Committee and Investigators. N. Engl. J. Med. 2017, 377, 122–131. [Google Scholar] [CrossRef]
- Szymanski, W.; Zwolińska, M.; Klossowski, S.; Mlynarczuk-Bialy, I.; Bialy, L.P.; Issat, T.; Malejczyk, J.; Ostaszewski, R. Synthesis of novel, peptidic kinase inhibitors with cytostatic/cytotoxic activity. Bioorganic Med. Chem. 2014, 22, 1773–1781. [Google Scholar] [CrossRef]
- Karmiris, E.; Vasilopoulou, M.-G.; Chalkiadaki, E. Acute Bilateral Anterior Uveitis following Cyclophosphamide/ Bortezomid/Dexamethasone (CyBorD) Protocol in a Newly Diagnosed Multiple Myeloma Patient with Concomitant Use of Zoledronic Acid. Ocul. Immunol. Inflamm. 2020, 12, 1–4. [Google Scholar] [CrossRef]
- Kowalczyk, P.; Borkowski, A.; Czerwonka, G.; Cłapa, T.; Cieśla, J.; Misiewicz, A.; Borowiec, M.; Szala, M. The microbial toxicity of quaternary ammonium ionic liquids is dependent on the type of lipopolysaccharide. J. Mol. Liq. 2018, 266, 540–547. [Google Scholar] [CrossRef]
- Appelmelk, B.J.; An, Y.; Hekker, T.A.M.; Thijs, L.G.; MacLaren, D.M.; De Graaf, J. Frequencies of lipopolysaccharide core types in Escherichia coli strains from bacteraemic patients. Microbiology 1994, 140, 1119–1124. [Google Scholar] [CrossRef] [PubMed]
- Amor, K.; Heinrichs, D.E.; Frirdich, E.; Ziebell, K.; Johnson, R.P.; Whitfield, C. Distribution of Core Oligosaccharide Types in Lipopolysaccharides from Escherichia coli. Infect. Immun. 2000, 68, 1116–1124. [Google Scholar] [CrossRef] [PubMed]
- Heinrichs, D.E.; Yethon, J.A.; Amor, P.A.; Whitfield, C. The assembly system for the outer core portion of R1 and R4 type lipopolysaccharides of Escherichia coli the r1 core specific β glucosyltransferase provides a novel attachment site for opolysaccharides. J. Biol. Chem. 1998, 273, 29497–29505. [Google Scholar] [CrossRef] [PubMed]
- Nnalue, N.A.; Khan, G.N.; Mustafa, N. Cross-reactivity between six Enterobacteriaceae complete lipopolysaccharide core chemotypes. J. Med. Microbiol. 1999, 48, 433–441. [Google Scholar] [CrossRef]
- Kowalczyk, P.; Madej, A.; Paprocki, D.; Szymczak, M.; Ostaszewski, R. Coumarin Derivatives as New Toxic Compounds to Selected K12, R1–R4 E. coli Strains. Materials 2020, 13, 2499. [Google Scholar] [CrossRef]
- Borkowski, A.; Kowalczyk, P.; Czerwonka, G.; Cieśla, J.; Cłapa, T.; Misiewicz, A.; Szala, M.; Drabik, M. Interaction of quaternary ammonium ionic liquids with bacterial membranes—Studies with Escherichia coli R1–R4-type lipopolysaccharides. J. Mol. Liq. 2017, 246, 282–289. [Google Scholar] [CrossRef]
- Thai, T.; Zito, P.M. Ciprofloxacin; StatPearls Publishing: Treasure Island, FL, USA, 2020. [Google Scholar]
- World Health Organization, Regional Office for the Western Pacific. Report of the Supreme Chamber of Control POST-CONTROL “Safety of Patients When Using Antibiotics in Hospitals”. In Western Pacific Country Health Information Profiles: 2008 Revision; WHO Regional Office for the Western Pacific: Manila, Philippines, 2008; ISBN 9789290613954. [Google Scholar]
- Kohanski, M.A.; Dwyer, D.J.; Collins, J.J. How antibiotics kill bacteria: From targets to networks. Nat. Rev. Microbiol. 2010, 8, 423–435. [Google Scholar] [CrossRef]
- Gedey, S.; Van Der Eycken, J.; Fülöp, F. Liquid-Phase Combinatorial Synthesis of Alicyclicβ-Lactams via Ugi Four-Component Reaction. Org. Lett. 2002, 4, 1967–1969. [Google Scholar] [CrossRef]
- Short, K.M.; Mjalli, A.M.M. A solid-phase combinatorial method for the synthesis of novel 5- and 6-membered ring lactams. Tetrahedron Lett. 1997, 38, 359–362. [Google Scholar] [CrossRef] [PubMed]
- Ugi, I. From isocyanides via four-component condensations to antibiotic syntheses. J. Ger. Chem. Soc. 1982, 21, 810–819. [Google Scholar]
- Mamat, C.; Pretze, M.; Gott, M.; Köckerling, M. Synthesis, dynamic NMR characterization and XRD studies of novel N,N’-substituted piperazines for bioorthogonal labeling. J. Org. Chem. 2016, 12, 2478–2489. [Google Scholar] [CrossRef] [PubMed]
- Greenberg, A.; Breneman, C.M.; Liebman, J.F. The Amide Linkage: Structural Significance in Chemistry, Biochemistry and Materials Science; John Wiley & Sons, Inc.: Hoboken, NJ, USA, 2003. [Google Scholar]
- Hartwig, A. Sensitive analysis of oxidative DNA damage in mammalian cells: Use of the bacterial Fpg protein in combination with alkaline unwinding. Toxicol. Lett. 1996, 88, 85–90. [Google Scholar] [CrossRef]
- Bailly, V.; Derydt, M.; Verly, W.G. Delta-elimination in the repair of AP (apurinic/apyrimidinic) sites in DNA. Biochem. J. 1989, 261, 707–713. [Google Scholar] [CrossRef]
- Jurado, J.; Saparbaev, M.; Matray, T.J.; Greenberg, M.M.; Laval, J. The Ring Fragmentation Product of Thymidine C5-Hydrate When Present in DNA Is Repaired by theEscherichia coliFpg and Nth Proteins. Biochemistry 1998, 37, 7757–7763. [Google Scholar] [CrossRef]
- Gilboa, R.; Zharkov, D.O.; Golan, G.; Fernandes, A.S.; Gerchman, S.E.; Matz, E.; Kycia, J.H.; Grollman, A.P.; Shoham, G. Structure of Formamidopyrimidine-DNA Glycosylase Covalently Complexed to DNA. J. Biol. Chem. 2002, 277, 19811–19816. [Google Scholar] [CrossRef]
- Graves, R.J.; Felzenszwalb, I.; Laval, J.; O’Connor, T.R. Excision of 5’-terminal deoxyribose phosphate from damaged DNA is catalyzed by the Fpg protein of Escherichia coli. J. Biol. Chem. 1992, 267, 14429–14435. [Google Scholar]
- Biela, A.; Coste, F.; Culard, F.; Guerin, M.; Goffinont, S.; Gasteiger, K.; Cieśla, J.; Winczura, A.; Kazimierczuk, Z.; Gasparutto, D.; et al. Zinc finger oxidation of Fpg/Nei DNA glycosylases by 2-thioxanthine: Biochemical and X-ray structural characterization. Nucleic Acids Res. 2014, 42, 10748–10761. [Google Scholar] [CrossRef]
- Saparbaev, M.; Sidorkina, O.M.; Jurado, J.; Privezentzev, C.V.; Greenberg, M.M.; Laval, J. Repair of oxidized purines and damaged pyrimidines by E. coli Fpg protein: Different roles of proline 2 and lysine 57 residues. Environ. Mol. Mutagen. 2002, 39, 10–17. [Google Scholar] [CrossRef]
- Tchou, J.; Kasai, H.; Shibutani, S.; Chung, M.H.; Laval, J.; Grollman, A.P.; Nishimura, S. 8-oxoguanine (8-hydroxyguanine) DNA glycosylase and its substrate specificity. Proc. Natl. Acad. Sci. USA 1991, 88, 4690–4694. [Google Scholar] [CrossRef]
- Tchou, J.; Michaels, M.L.; Miller, J.H.; Grollman, A.P. Function of the zinc finger in Escherichia coli Fpg protein. J. Biol. Chem. 1993, 268, 26738–26744. [Google Scholar] [PubMed]
- Tchou, J.; Bodepudi, V.; Shibutani, S.; Antoshechkin, I.; Miller, J.; Grollman, A.P.; Johnson, F. Substrate specificity of Fpg protein. Recognition and cleavage of oxidatively damaged DNA. J. Biol. Chem. 1994, 269, 15318–15324. [Google Scholar] [PubMed]
- Sidorkina, O.M.; Laval, J. Role of lysine-57 in catalytic activities of Escherichia coli. Formamidopirymidine-DNA glycosylase (Fpg protein). Nucleic Acid Res. 1998, 26, 5351–5357. [Google Scholar] [CrossRef]
- Sabourin, M.; Osheroff, N. Sensitivity of human type II topoisomerases to DNA damage: Stimulation of enzyme-mediated DNA cleavage by abasic, oxidized and alkylated lesions. Nucleic Acids Res. 2000, 28, 1947–1954. [Google Scholar] [CrossRef]
- Nair, J.; Barbin, A.; Velic, I.; Bartsch, H. Etheno DNA-base adducts from endogenous reactive species. Mutat. Res. Mol. Mech. Mutagen. 1999, 424, 59–69. [Google Scholar] [CrossRef]
- Rabow, L.E.; Kow, Y.W. Mechanism of Action of Base Release by Escherichia coli Fpg Protein: Role of Lysine 155 in Catalysis. Biochemistry 1997, 36, 5084–5096. [Google Scholar] [CrossRef]
- Cussac, C.; Laval, F. Reduction of the Toxicity and Mutagenicity of Aziridine in Mammalian Cells Harboring the Escherichia Coli fpg Gene. Nucleic Acids Res. 1996, 24, 1742–1746. [Google Scholar] [CrossRef]
- Czene, S.; Harms-Ringdahl, M. Detection of single-strand breaks and formamidopyrimidine-DNA glycosylase-sensitive sites in DNA of cultured human fibroblasts. Mutat. Res. Repair 1995, 336, 235–242. [Google Scholar] [CrossRef]
- Tchou, J.; Grollman, A. The catalytic mechanism of Fpg protein. Evidence for a Schiff base intermediate and amino terminus localization of the catalytic site. J. Biol. Chem. 1995, 270, 11671–11677. [Google Scholar] [CrossRef]
- Collins, A.; Duthie, S.; Dobson, V. Direct enzymatic detection of endogenous oxidative base damage in human lymphocyte DNA. Carcinogenesis 1993, 14, 1733–1735. [Google Scholar] [CrossRef]
- Boiteux, S.; O’Connor, T.R.; Laval, J. Formamidopyrimidine-DNA glycosylase of Escherichia coli: Cloning and sequencing of the fpg structural gene and overproduction of the protein. EMBO J. 1987, 6, 3177–3183. [Google Scholar] [CrossRef] [PubMed]
- Hurdle, J.; O’Neill, A.; Chopra, I.; Lee, R. Targeting bacterial membrane function: an underexploited mechanism for target in persistant infections. Nat. Rev. Microbiol. 2011, 9, 62–75. [Google Scholar] [CrossRef] [PubMed]

| No of Samples | 5a | 5b | 5c | 5d | 5e | 5f | 5g | 5h | 5i | 5j | 5k | 5l | 5m | 5n | 5o | 5p | 5r | 5s | 5t | 5u | Type of Test |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| K12 | ** | *** | *** | *** | ** | *** | ** | ** | *** | MIC | |||||||||||
| R2 | ** | *** | *** | *** | ** | *** | ** | ** | *** | MIC | |||||||||||
| R3 | ** | *** | *** | *** | ** | *** | ** | ** | *** | MIC | |||||||||||
| R4 | ** | *** | *** | *** | ** | *** | ** | ** | *** | MIC | |||||||||||
| K12 | ** | * | ** | ** | * | ** | * | *** | * | MBC | |||||||||||
| R2 | ** | * | ** | ** | * | ** | * | *** | * | MBC | |||||||||||
| R3 | ** | * | ** | ** | * | ** | * | *** | * | MBC | |||||||||||
| R4 | ** | * | ** | ** | * | ** | * | *** | * | MBC | |||||||||||
| K12 | * | ** | * | * | * | * | ** | * | ** | MBC/MIC | |||||||||||
| R2 | * | ** | * | * | * | * | ** | * | ** | MBC/MIC | |||||||||||
| R3 | * | ** | * | * | * | * | ** | * | ** | MBC/MIC | |||||||||||
| R4 | * | ** | * | * | * | * | ** | * | ** | MBC/MIC |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kowalczyk, P.; Madej, A.; Szymczak, M.; Ostaszewski, R. α-Amidoamids as New Replacements of Antibiotics—Research on the Chosen K12, R2–R4 E. coli Strains. Materials 2020, 13, 5169. https://doi.org/10.3390/ma13225169
Kowalczyk P, Madej A, Szymczak M, Ostaszewski R. α-Amidoamids as New Replacements of Antibiotics—Research on the Chosen K12, R2–R4 E. coli Strains. Materials. 2020; 13(22):5169. https://doi.org/10.3390/ma13225169
Chicago/Turabian StyleKowalczyk, Paweł, Arleta Madej, Mateusz Szymczak, and Ryszard Ostaszewski. 2020. "α-Amidoamids as New Replacements of Antibiotics—Research on the Chosen K12, R2–R4 E. coli Strains" Materials 13, no. 22: 5169. https://doi.org/10.3390/ma13225169
APA StyleKowalczyk, P., Madej, A., Szymczak, M., & Ostaszewski, R. (2020). α-Amidoamids as New Replacements of Antibiotics—Research on the Chosen K12, R2–R4 E. coli Strains. Materials, 13(22), 5169. https://doi.org/10.3390/ma13225169






