1. Introduction
The measurement of the Trans-Epithelial/Endothelial Electrical Resistance (TEER) is a widely accepted quantitative technique for in vitro assaying the barrier integrity of epithelial/endothelial tissues or monolayer epithelial cell cultures, as well as assessing ion transport and/or the quality (or operation) of the tight junctions. The measurement principle is straightforward; by applying an A.C. electrical signal across electrodes placed on both sides of the barrier, the TEER can be calculated from the applied voltage and resulting current established in the circuit. Higher TEER implies better barrier integrity.
In the case of monolayer cell cultures, measuring TEER is relatively simple: cells are plated and cultured in a Transwell insert whose bottom is a porous membrane permeable to media and ions. A pair of chopstick-shaped electrodes are inserted into the media to measure the TEER, with one leg in the insert and the other leg in the outer well. The interested reader can find more information on this topic in Wegener’s early work [
1], or in the reviews of Benson et al. [
2] and of Srinivasan et al. [
3], respectively.
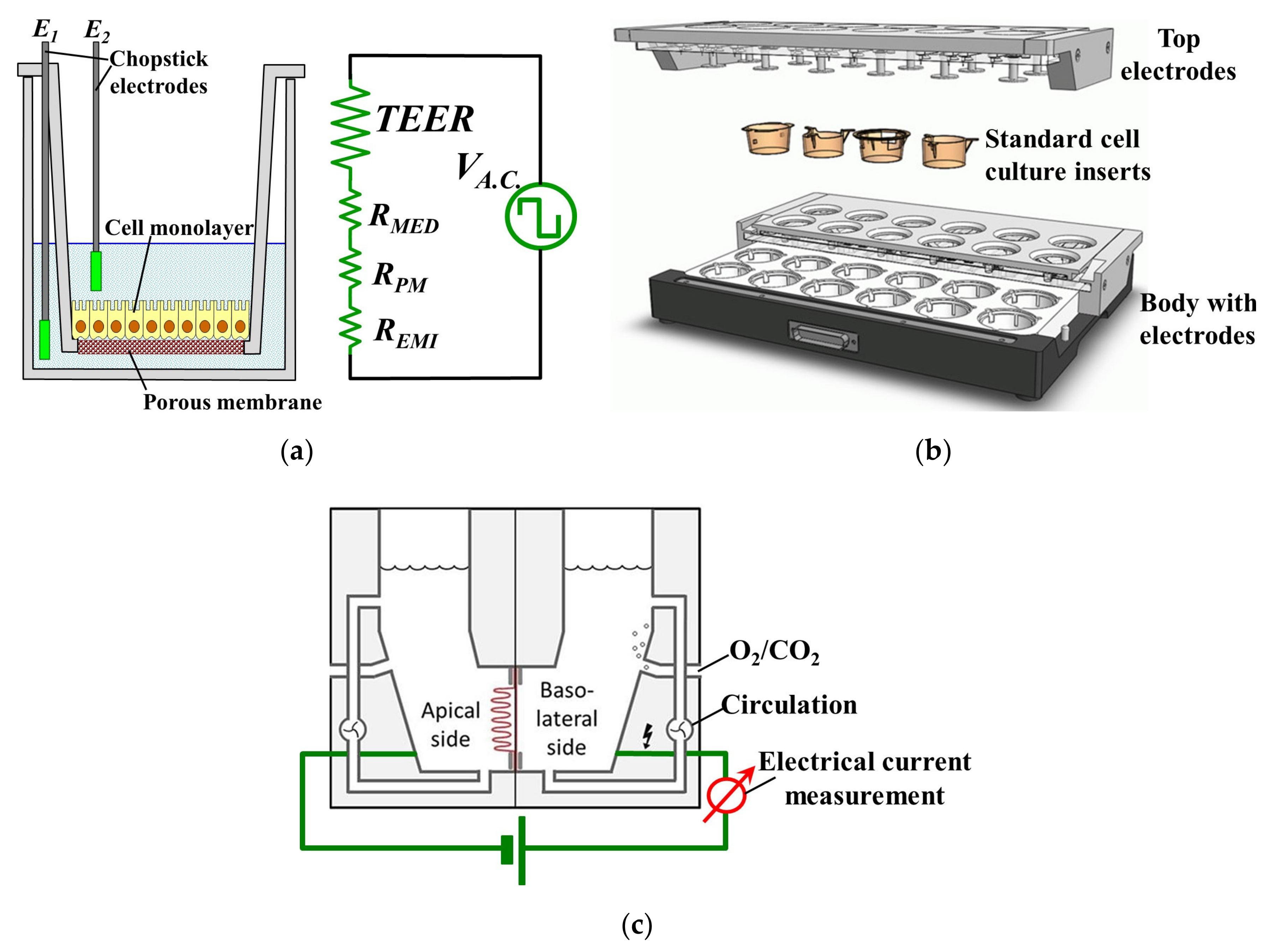
Figure 1a illustrates schematically the set-up, with a cellular monolayer cultured on a semipermeable filter insert that defines a partition for apical and basolateral compartments, each contacted by a chopstick-like electrode [
2]. However, the resulting TEER readings obtained with this macroscopic set-up inherently include a few sources of potentially significant errors [
2]. Thus, the measured TEER values depend strongly on the shape and position of the probing electrodes. Vibrations easily cause significant disturbances and the reproducibility of electrodes’ placements is also a serious concern, while the inherent inhomogeneity of the electric field across the cell layer typically leads to a systematic overestimation of the measured TEER [
2]. A further improvement was provided by the CellZscope
® machine (cellZscope, nanoAnalytics GmbH, Muenster, Germany) which eliminated most of the previous drawbacks. It comprises several wells which can all be cultured and measured simultaneously (e.g., in Transwell inserts with 6, 12 or 24 wells), each with its own electrodes that measure the A.C. impedance of barrier-forming cell cultures growing on the permeable membranes of standard cell culture inserts, providing more valuable information than TEER [
4]. It is also the only way with which information regarding the evolution of the cell culture can be obtained because light microscopy is no longer feasible for Transwell cultured cells due to the presence of the supports which hinders microscopic observation [
5]. The schematic structure of the CellZscope
® is shown in
Figure 1b. However, this impedance measurement is typically used just to signal when complete confluence of cells in a continuous monolayer was achieved, and thus the cells can be harvested and used for other purposes. Moreover, impedance measurement is more difficult and requires a more complex measurement set-up and data processing in order to derive the relevant parameters of interest from the measurements, after which correctly interpreting these parameters and extracting the right and meaningful conclusions is another non-trivial task.
This explains why TEER measurement is considered the easiest, fastest, and most convenient tool to monitor the quality and behavior of cells cultured on porous supports. Due to its simplicity and usefulness, the TEER method was also integrated with lab-on-a-chip technology to realize promising biodevices [
6,
7].
Unfortunately, such in vitro tests, although cheap and reasonably easy to do, have serious drawbacks. First, a simple monolayer of cells cannot capture the complex behavior of living tissue, and many cell types are not even identical with those present in the actual organ of interest but relatively similar cell lines are used instead. Second, such measurements are static and can study a limited number of variables and only for a very specific application. Last but not least, they are time-consuming and require a large volume of reagents and culture media.
A much more attractive alternative is to employ instead actual tissues biopsied from the organ of interest. Such tissues are much better biological models than monolayer cell cultures for in vitro experiments because they are more physiologically relevant to the environment in vivo that is being studied. However, much less research employed TEER of biopsied tissues due to its challenges. Unlike monolayer cell cultures, measuring the TEER of living tissues using the Transwell insert method is no longer feasible because a tissue specimen simply placed in the Transwell cup will not be adherent to the porous membrane. This means that there will be many gaps between the edge of the specimen and the wall/porous membrane of the Transwell insert. These gaps become channels for fluid and ion transportation which lead to incorrect and unreliable TEER measurements.
The solution for these problems is the Ussing chamber, shown in
Figure 1c, which has been used as the gold standard for assessing TEER of tissue specimens [
8]. Thus, TEER has been heavily involved in many epithelial tissue-related studies of various topics of interest, such as the permeability of the blood-brain barrier [
9,
10], the integrity and/or the metabolism of human intestinal mucosa [
11], intestinal drug delivery [
12], the factors affecting the development of tight junctions [
13], the response of tight junctions to proinflammatory cytokines [
14], the effect of absorption enhancers on the permeability of intestinal epithelial cells [
15] and investigating the pathways of ion transport across epithelial cells [
16].
As shown in
Figure 1c, an Ussing chamber system generally comprises two half-chambers with a perfusion system, and a piece of epithelial tissue is sandwiched by the two half-chambers in such a way that each side of the tissue is isolated and faces a separate half-chamber. Pathologists and gastroenterologists have been using the Ussing chamber to study the esophageal epithelial integrity in various diseases of the GastroIntestinal Tract (GIT). It was shown that esophageal biopsies had modified TEER values in patients with various GIT affections such as Barrett’s esophagus, gastro-esophageal reflux disease (GERD) [
17,
18] that also includes the non-erosive reflux disease (NERD) [
19,
20], celiac disease [
21], as well as microscopic colitis (MC), collagenous colitis and treated lymphocytic colitis [
22], all indicating disturbances in ion transport and barrier function, with typically a significantly decreased epithelial resistance.
Although the Ussing chamber system proved to be accurate and reliable, it has serious drawbacks. Most importantly, the number of specimens that can be measured at a time is limited by the number of chambers available, which greatly limits the type of experiments can be carried out. Other significant operational limitations include the lack of ability to preserve the specimen for relatively longer periods and difficulty in quickly introducing trace amounts of treatment molecules such as labeling dyes or therapeutic drugs. Finally, it is voluminous, cumbersome to use, and has a relatively high cost.
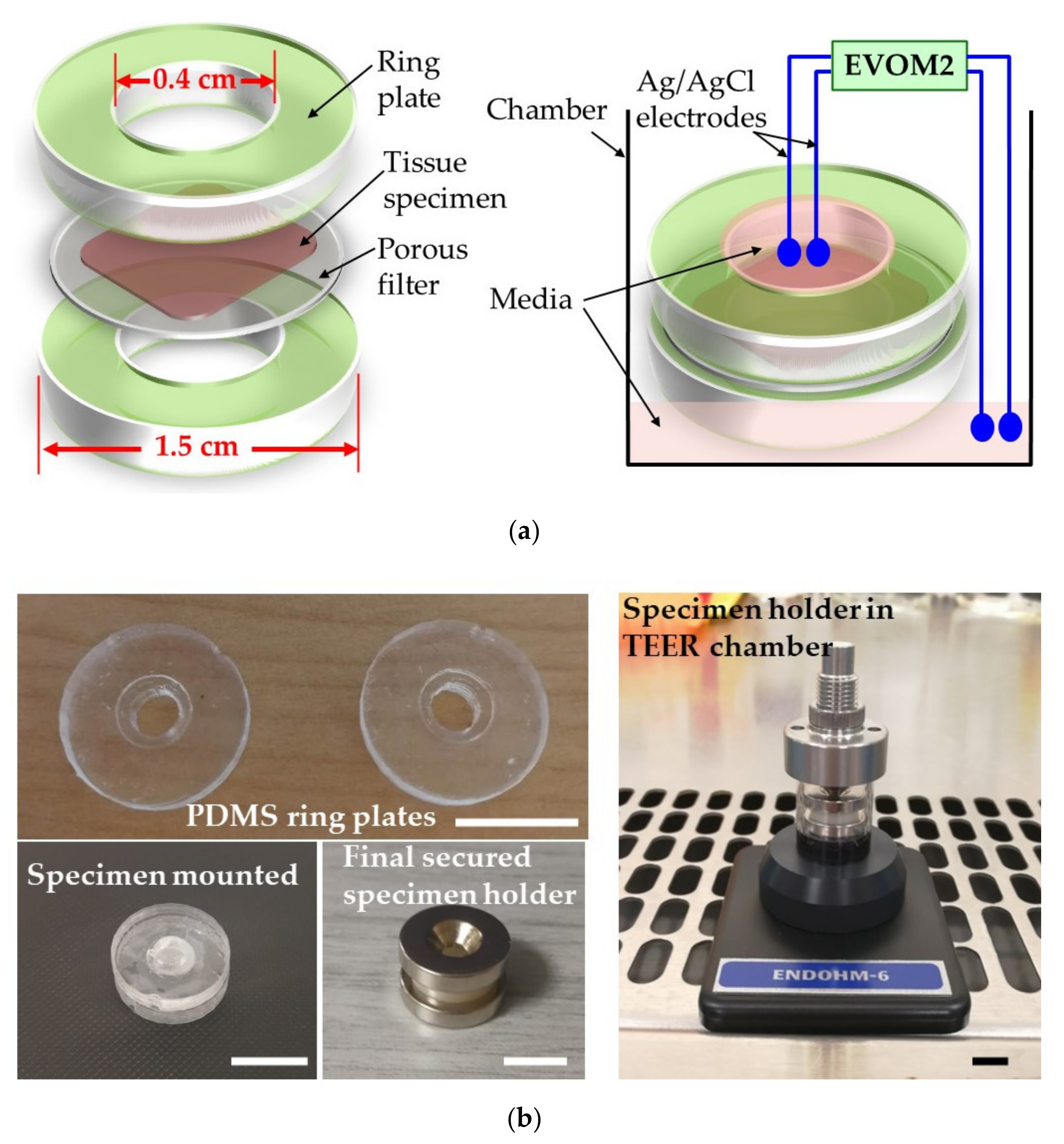
This paper first reports the development and successful usage of a low-cost home-made “sandwich ring” method for measuring TEER of epithelial tissues. The tissue can be mounted in a specimen holding set which consists of one pair of polydimethylsiloxane (PDMS) rings and one pair of ring magnets. The performance of the “sandwich ring” method was certified by performing TEER measurements on porcine esophageal epithelia. Next, we applied the same underlying idea of this “sandwich ring” method to develop a biochip made of PDMS and polymethyl methacrylate (PMMA) with which real-time continuous TEER measurements of tissue samples were performed. We believe this to be the first biochip being reported enabling both extended viability and TEER continuous measurements of biopsied tissue samples. The outstanding features of this biochip include the ability to allow temporarily culturing of the tissue on-chip, easiness in carrying out chemical treatments, re-usability, low fabrication cost and small size. Such a biochip could thus be used to easily and quickly assess quantitatively the type and severity of various gastro-esophageal deficiencies, to compare similar afflictions between different patients, or estimate the efficiency of treatment by comparing variations in time for the same patient. It will also be shown later in this paper that these TEER variations are directly and strongly correlated with variations in the tissue’s permeability. Both our “sandwich ring” method and the biochip can thus have very important applications in clinical practice with which such valuable assessments can be performed.
2. Materials and Methods
PDMS (SYLGARD 184) was purchased from Dow Corning and 1 mm thick PMMA sheets were bought from a local workshop. Small assembly components and tools including ring magnets, stainless steel screws, epoxy adhesives, surgical scissors and scalpel were purchased from RS Components. Fetal Bovine Serum (FBS) and phosphate-buffered saline (PBS) solutions were purchased online from ThermoFisher Scientific Singapore. All other bioreagents/chemicals such as Keratinocyte Growth Medium (KGM), Leibovitz’s L-15 medium, fluorescein isothiocyanate–dextran (FITC-Dextran), sodium alginate, acetic acid, dispase II enzyme, 0.25% trypsin-ethylenediaminetetraacetic acid (EDTA) and antibiotic antimycotic (100×) solutions were purchased from Sigma Aldrich Singapore (Sigma-Aldrich Pte Ltd, 2 Science Park Drive, #05-01/12 Ascent Building, Singapore 118222). Fresh porcine esophagus was harvested from pigs euthanized in Singapore General Hospital (SGH) and delivered to the lab for subsequent processing within three hours. During transportation, the harvested esophagus was kept in ice-cold Leibovitz’s L-15 medium.
2.1. Peeling Off the Mucosal/Epithelium Layer from the Porcine Esophagus
Prior to processing the esophagus sample, all surgical tools were rinsed with 70% ethanol followed by UV exposure for 1 h for sterilization and all the processing procedures were conducted in a biosafety cabinet. First the esophagus was cut open with a pair of surgical scissors and the muscular layer was stripped off from the mucosa and submucosa layer using tweezers. This reduced the thickness of sample, from an initial value of about 4 mm down to ~1.5 mm. Next, the harvested tissue was divided into shorter segments (about 5 cm long) using a scalpel. These tissue segments were subsequently washed with PBS solution three or four times until the solution remained clear. Then each segment was further cut into smaller pieces of 2 cm × 2 cm and immersed in 0.25% trypsin-EDTA solution for 5 min, followed by thorough rinsing with PBS. This latter step is necessary to remove blood cells as well as connective tissue cells present in the sample [
23]. After the trypsin treatment, the samples were kept in L-15 medium supplemented with 10 U/mL dispase II enzyme at 4 °C for 6 h in order to remove the bonding between the mucosal and subcostal layer without harming the integrity of the mucosal layer. Finally, the mucosal layer was carefully peeled off from the submucosa. This method was reported by Compton et al. [
24]. The harvested mucosal/epithelial samples, with a thickness between 400–600 µm, were either used immediately for measurements or temporarily maintained (in KGM culture media supplemented with antibiotic antimycotic solution) for later usage in an incubator at 37 °C with an ambient containing 5% CO
2. The thickness of the completely dried epithelial tissues was measured with a vernier caliper to be between 150–200 µm.
2.2. Fabrication of the “Sandwich Ring” and Biochip
All the PDMS components were fabricated using soft lithography, with molds 3D printed in polylactic acid (PLA) with a 3D printer model Ultimaker 3. Liquid PDMS was mixed with curing agent in a ratio of 10:1 and the mixture was poured into the mold, degassed in a vacuum desiccator for 1 h and cured in an oven at 60 °C for 6 to 8 h. The cured PDMS was then carefully peeled off from the mold using tweezers. One of the major issues of the 3D printed molds is that the mold surface is not perfectly smooth, leading to a relatively rough PMDS surface. These imperfections may prevent tight sealing with other surfaces and would thus result in leakages. This problem was addressed by a simple spin-coating technique. Basically, a thin layer of liquid PDMS was first created on a clean Si wafer 4 inch in diameter using a spin coater at 2000 rpm. The PDMS components were placed on the wafer with the non-smooth surface facing downwards to allow liquid PDMS to stick onto the non-smooth surface. Then the PDMS components were transferred onto another clean wafer, followed by degassing in a vacuum desiccator for 30 min. Lastly, the wafer with PDMS components was placed on a hot plate at 90 °C for 30 min. Once cured, the PDMS components can be peeled off from the wafer and the smoothness of the surface is greatly enhanced.
The PMMA perfusion channels which were used to assemble the biochip were fabricated by using a CO
2-laser (Universal Laser Systems M-360) to engrave a piece of PMMA sheet in order to realize the channel patterns. The laser power and scanning speed were first tested to make sure they were adequate the cut through the PMMA sheet. The optimal cutting recipe eventually adopted to cut two PMMA layers employed 70% laser power at 4% scanning speed. One of the PMMA layers contained the two-side open channel while the other PMMA layer serves as seal layer with some small holes as fluid inlet/outlet and for electrodes insertion. After placing the two layers together, the top and bottom fluidic channels were realized [
25]. The two PMMA layers were thermally bonded to prevent any fluid leakage. For this purpose, the two layers were clamped tightly and maintained in an oven at 95 °C for 20 min. [
26]. The Ag/AgCl electrodes used in the biochip were prepared by immersing sanded Ag wires of 0.3 mm diameter into chlorine bleach until the Ag wire turned black.
Both the PDMS rings used in the “sandwich ring” as well as the biochip can be re-used, if so desired. In such a case, every time before loading a new piece of tissue sample, all the biochip components (or the PDMS rings) are immersed in 70% ethanol for at least 15 min., then dried in a vacuum desiccator for one hour, followed by exposure to UV light in a biosafety cabinet for 1 h to achieve sterilization. Once all these operations were performed, the tissue sample was cut into 6 mm disks with a biopsy punch and loaded either between the rings (for the “sandwich ring”), or into the central groove between the central PDMS layers of the biochip. At last, the four PMMA layers and the two PDMS layers of the biochip are assembled together and secured with screws.
2.3. TEER Measurements Using the “Sandwich Ring” Tissue Holder
A piece of an epithelium sample (that had initially been prepared as mentioned above) was placed on a piece of membrane filter (WhatmanTM, 0.2 µm pore size) wetted with PBS. Next, disk-shaped specimens were prepared by cutting the epithelium sample using a disposable 6 mm biopsy punch. This disk specimen would fit exactly in the central groove of a PDMS ring. Then the other PDMS ring was placed above it to sandwich the specimen, and a pair of ring magnets purchased from Eclipse Magnetics Singapore (75 Rowell Road, Singapore 208011; product no.: N832RS; outer and inner hole diameters: 15.4 mm and 3.3 mm, respectively; 3.25 mm thick; pull force: 3.3 kg) were used to sandwich in-between them the PDMS rings, thus forming the complete specimen holder.
The TEER measurements were carried out using a voltohmmeter (EVOM2, World Precision Instruments) in a TEER chamber (Endohm-6, World Precision Instruments). Prior to measurement, 100 µL of KGM medium were added in the Endohm-6 chamber to wet the bottom surface. The bottom space of the specimen holder was filled with 60–70 µL medium and the specimen holder was mounted in Endohm-6. Then the top space of the Endohm’s specimen holder above the tissue was filled with 100 µL medium. Filling up the spaces of the specimen holder that are above and below the tissue with KGM culture media with a conductivity of ~1.4 S/m ensures a direct low resistance electrical connection between the electrodes and the tissue surface. The Endohm-6 specimen holder was then closed with its cap and the position of the electrode on the cap was carefully adjusted to make sure it was touching the medium in the top space. This ensured that an electrical circuit was formed between the two pairs of electrodes through the tissue sample, and TEER values could be measured and recorded after connecting the Endohm-6 chamber to EVOM-2.
The accuracy of the “sandwich ring” method was verified by measuring the TEER values of monolayer cell cultures and then compared with those obtained from the standard method (Transwell). In brief, Caco-2 cells (a colon epithelial cell line) were plated on the porous membranes of 4 inserts of Transwell dishes and cultured until full confluence. Then the TEER of these monolayer cell cultures was measured with a pair of chopstick electrodes and our EVOM2 voltohmmeter. The measured value was 1154 ± 97 Ω·cm2. Subsequently, these porous membranes with cells were carefully cut off from the Transwell inserts and their TEER was measured again using our “sandwich ring” method. The measured TEER was 1071 ± 65 Ω·cm2. The p-value for the two data sets is 0.161, indicating that the difference between these two values is statistically insignificant. Hence, the accuracy of our “sandwich ring” method was confirmed.
When sufficient data points could be collected for our measurements, the graphs illustrating the observed variations as a function of the modified parameters included both the data points and the least squares fitting curves which could be found for those data points. This is because such fitting curves would more clearly illustrate the overall variation trend, unlike a jagged multi-segmented line resulted from point-to-point connection that is less accurate and more prone to errors given the inherent spread in the statistical set of collected data. For brevity, the values of the coefficients of the fitting equations and of the measured data points are not mentioned here but are included in the
Supplementary Materials file.
Each data point in all the graphs presented in the paper is the mean value resulted from several separate measurements. The specific number n of measured values whose average provided each of the plotted data points is clearly specified in the caption of that respective Figure, and the accompanying error bars represent the standard deviation σ.
2.4. Preserving Tissue Specimens in the Biochips and Performing TEER Measurements
Prior to specimen mounting, all biochip components were rinsed with 70% ethanol followed by UV exposure for 1 h for sterilization. A 6 mm disk specimen was placed in the groove of one PDMS layer of the biochip and was then covered with the other PDMS layer to sandwich the specimen. Then the PDMS layers with the specimen were sandwiched themselves between the top and bottom PMMA channels which were stacked together and secured tightly with screws. The two perfusion flows were supplied using a peristaltic pump at a flow rate of 0.6 mL/h from a KGM medium reservoir which was maintained in an incubator at 37 °C and 5% CO2. To perform the TEER measurements, the four electrodes were connected to the EVOM2 using alligator clips.
Any outlier TEER readings (less than 100 or more than 700 Ω·cm2) obtained with the biochip were discarded. The low readings were probably caused by leakages in the experimental setup while the excessively high readings were most likely due to air bubbles trapped in the central hole.
2.5. Fluorescence Intensity Measurements to Determine the Tissue Permeability
The fluorescence intensity was measured using a plate reader (Tecan Infinite 200 Pro) with the software i-control™. The experimental procedure was as follows. Medium with added 5 μg/mL FITC-dextran was perfused into the upper (apical) channel of the biochip and the effluent from the lower (basal) channel was collected. Specifically, 50 µl of the same sample solution were added to each of the 4 wells out of the total of 8 wells of the same column of a 96-well (12 columns × 8 rows) Transwell plate, and each column held liquid from different samples. The Transwell plate was then inserted in the plate reader and the concentration of the FITC-dextran in the bottom effluent was quantified by measuring the fluorescence intensity at 530 nm, for an excitation wavelength of 480 nm. An increase in the FITC-dextran concentration in the lower channel effluent would imply an increased tissue permeability.
2.6. Tissue Sectioning and Haematoxylin & Eosin Staining for Histological Analysis
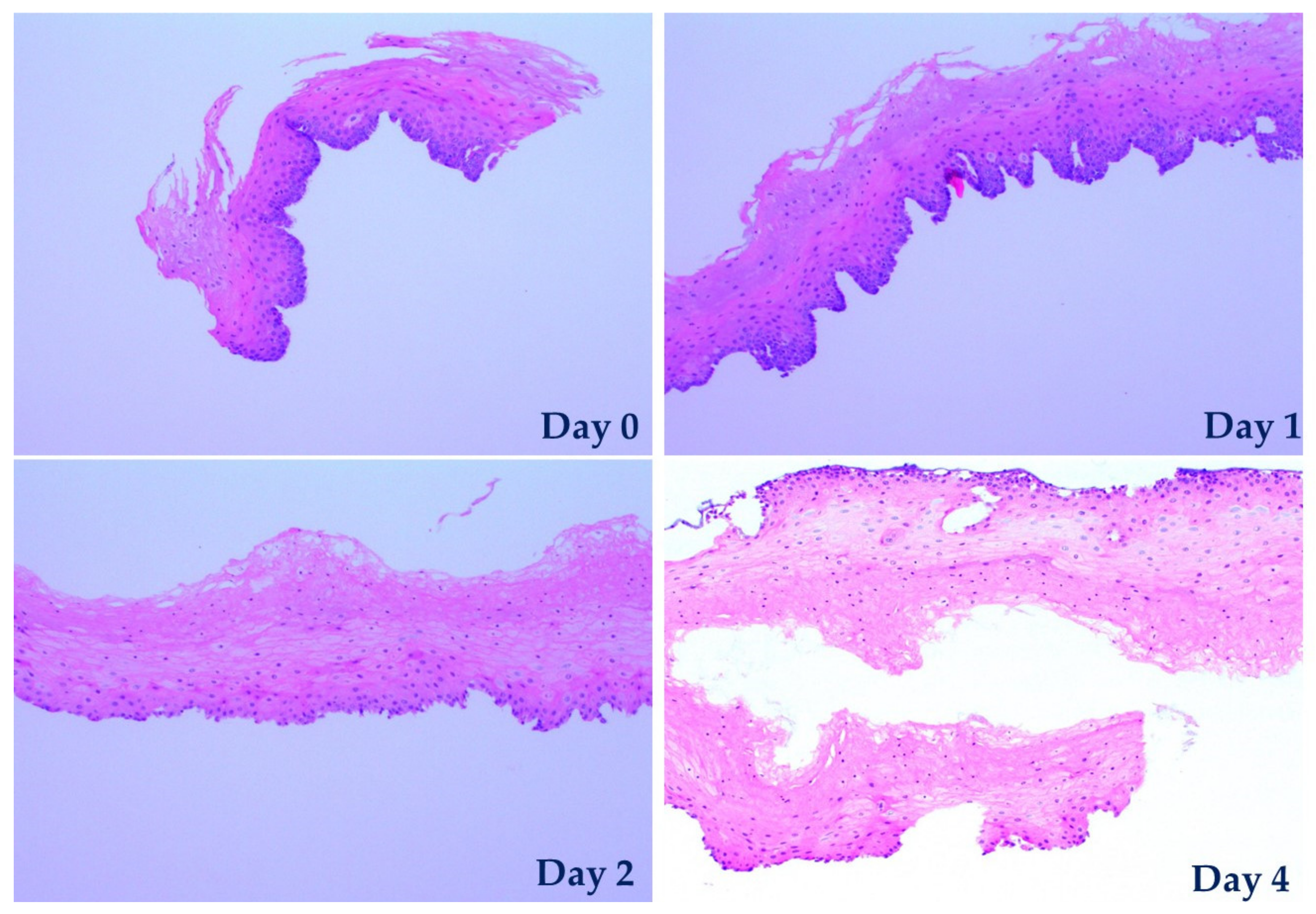
The tissue specimens were first fixed in a solution of 4% formalin, followed by dehydration with escalated concentrations of ethanol from 20% to 99%. Then the sample was embedded in paraffin blocks, followed by being sectioned vertically into slices 4 µm thick using a manual microtome (Leica RM2235) to reveal the cross-section of the specimens. The sliced samples were placed on glass slides and stained with haematoxylin and eosin in a fully automated stainer (Leica CV5030). Lastly, the stained samples were analyzed using a light microscope (Olympus BX43).
4. Conclusions
In this article, we demonstrated a “sandwich ring” method for faster and easier measuring the TEER of tissue specimens without using a Ussing chamber. The same concept was then applied to design and develop a dual-function tissue biochip for both temporary maintenance of tissue specimens as well as their continuous TEER monitoring. All the experimental results obtained using our tissue biochip clearly demonstrated that it is an effective substitute to an Ussing chamber. Moreover, we believe that our obtained results prove that this novel tissue biochip is much more practical, convenient and easier to use than an Ussing chamber due to its unique advantageous features, such as easy installation of the sample, re-usability, low-cost and rapid prototyping, small size, and enabling both long(er) term tissue viability and real-time monitoring of TEER.
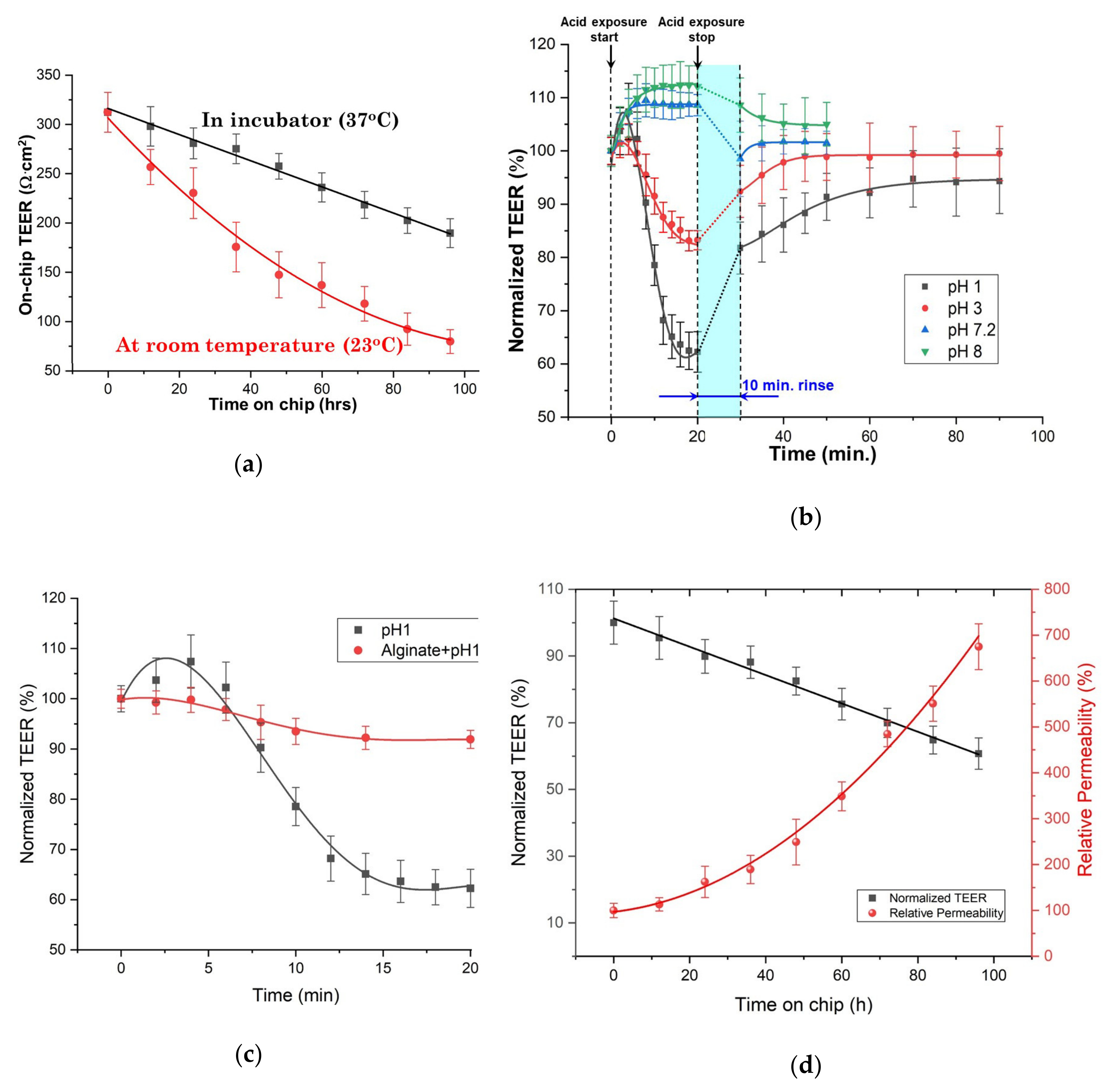
With the aid of this biochip, porcine esophageal tissues were maintained viable and their TEER was monitored for up to four days. Due to the biochip’s microfluidics which enables significant flexibility in its application for various purposes, tests in which tissues were exposed to media to various pH values were easily performed and in conjunction with a permeability test they effectively demonstrated that such a tissue biochip can be used in a large range of possible applications which can subject the sample to various biochemical treatments. Our experimental results obtained from both the exposure of tissues to media of various pH values and the permeability tests led to two crucial conclusions. Firstly, and most importantly, our results demonstrate a clear strong correlation between the measured on-chip TEER and the tissue permeability (the former decreases when latter increases) which was confirmed by histopathological observations. The current state-of-art methods for assessing tissue viability are all involving biochemical treatments of the samples to generate optical contrast or detectable signals such as fluorescence for observation under microscope. Hence, these assessing methods are usually costly, time-consuming, labor intensive and more significantly, destructive to the samples. Secondly, the reliable correlation found between the TEER values and tissue viability demonstrates that TEER is an excellent alternative method for assessing tissue viability, as it is easier, faster, label-free, and non-destructive. These findings suggest that our biochip may find important clinical applications that could e.g., determine the most efficient drugs or assess the efficiency of a given treatment.