Microorganism Assisted Synthesized Nanoparticles for Catalytic Applications
Abstract
1. Introduction
2. Biosynthesis of NPs
2.1. Elementary Substances
2.1.1. Single Metal
2.1.2. Metalloid Elements
2.2. Compounds
2.2.1. Metal Oxide
2.2.2. Metal Chalcogenides
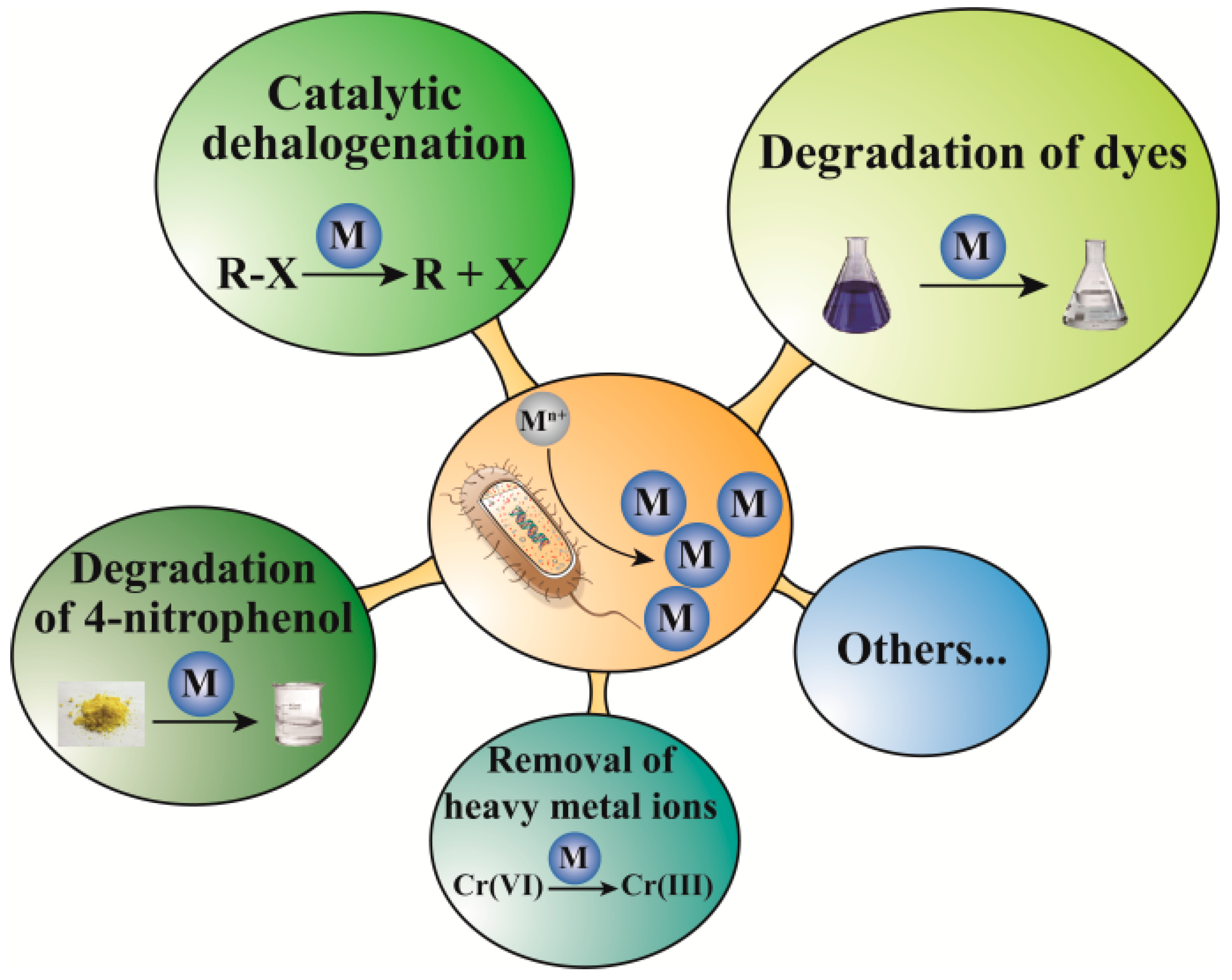
3. Catalytic Applications
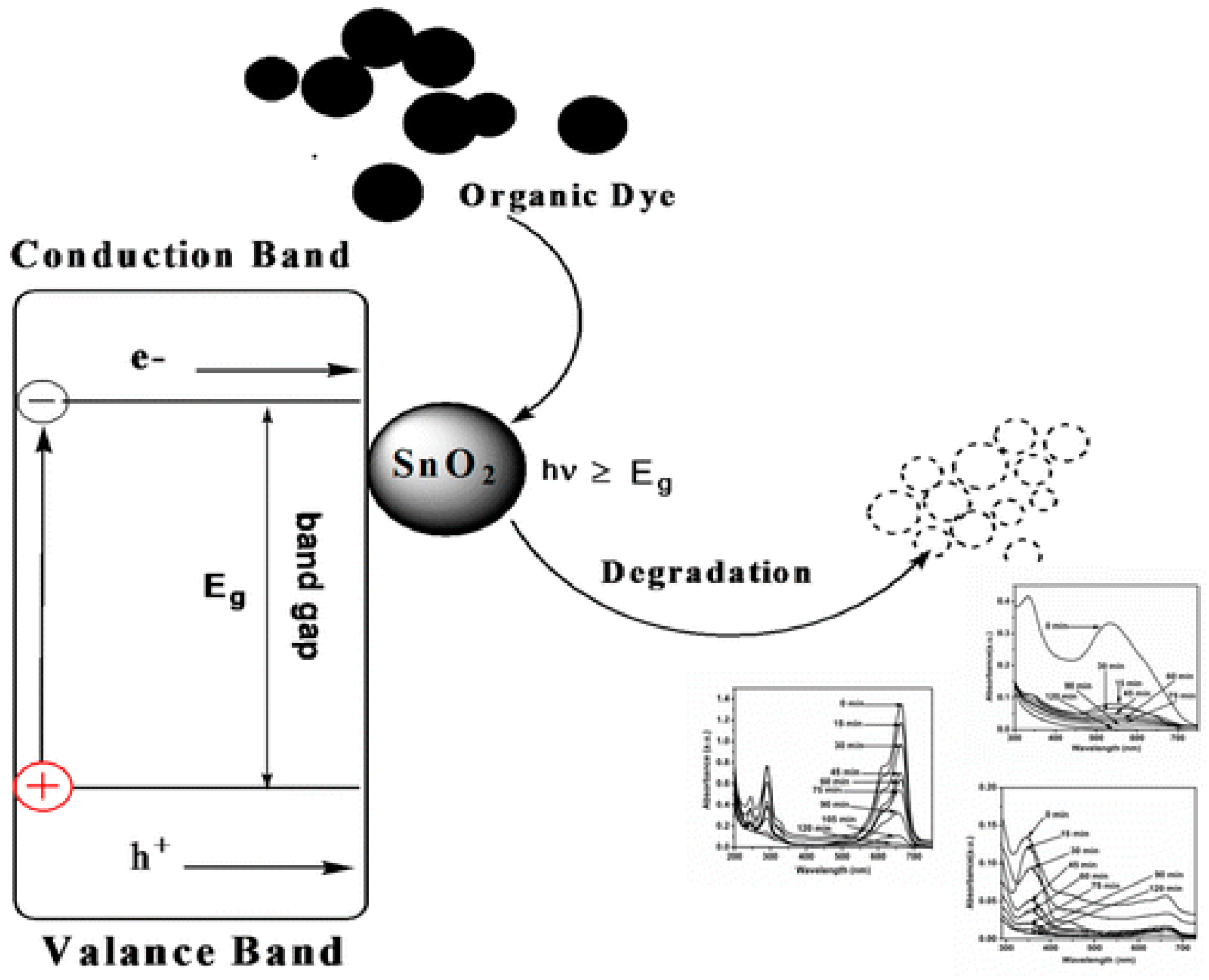
3.1. Degradation of Dyes
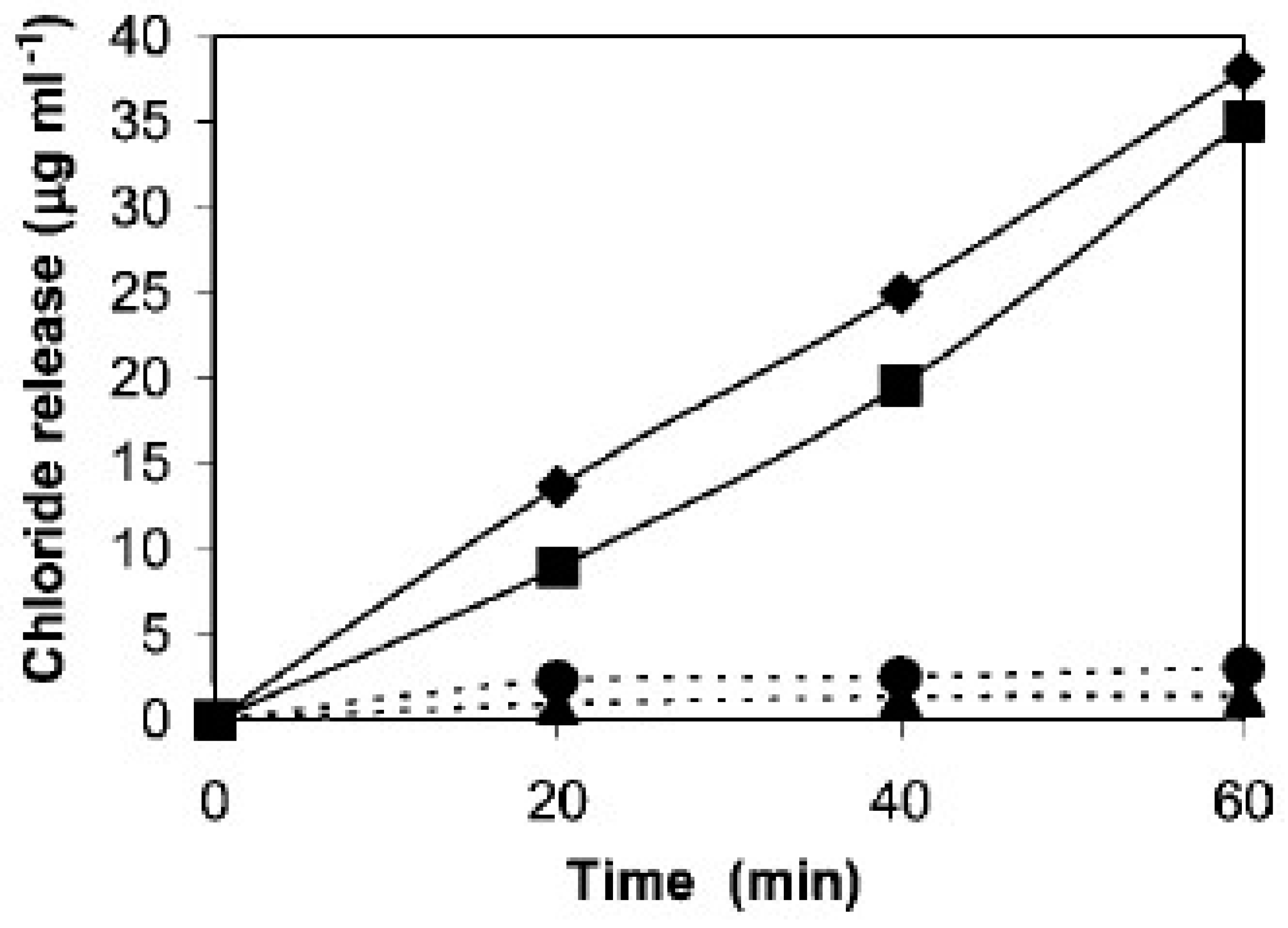
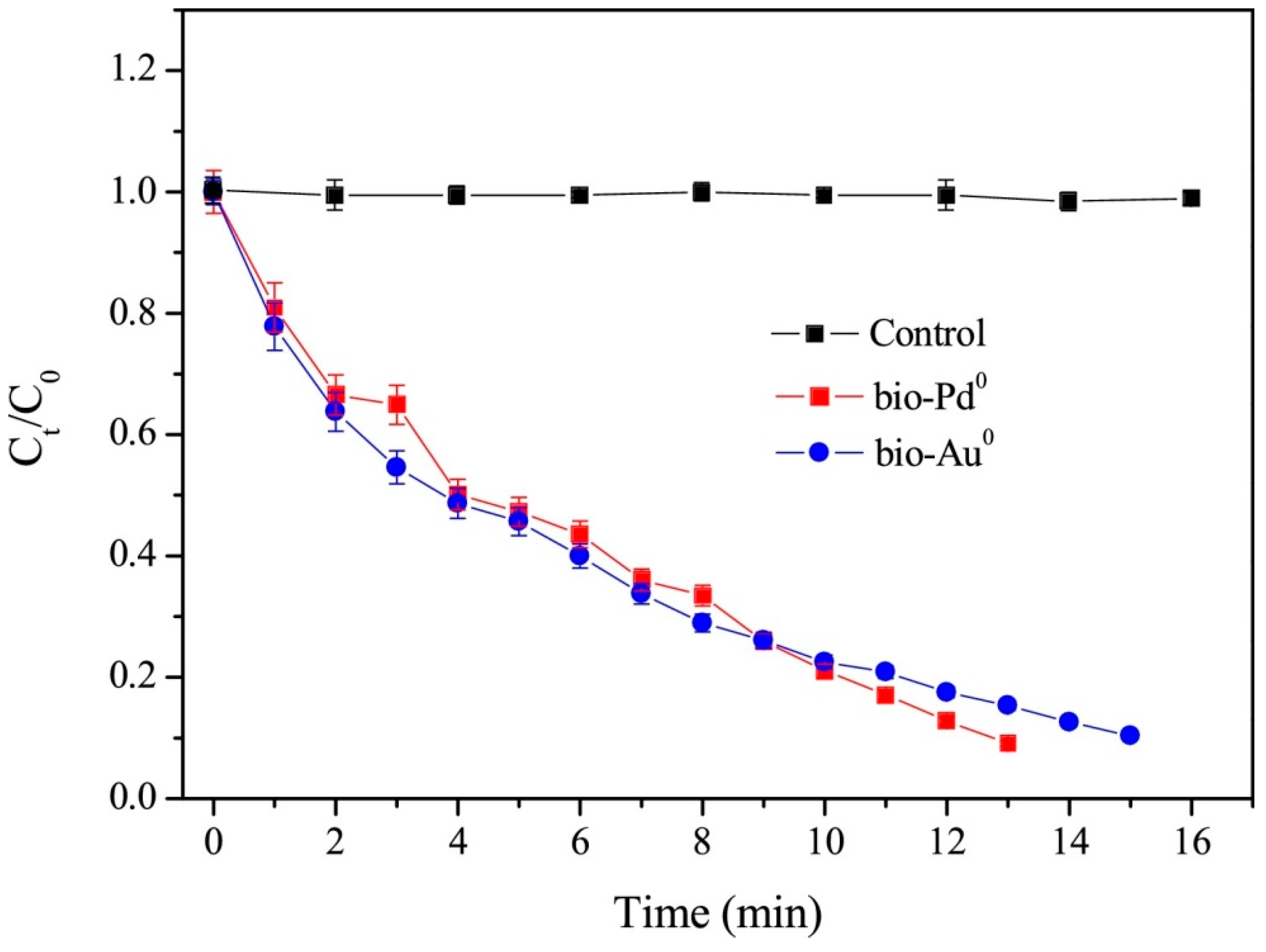
3.2. Catalytic Dehalogenation
3.3. Degradation of 4-Nitrophenol
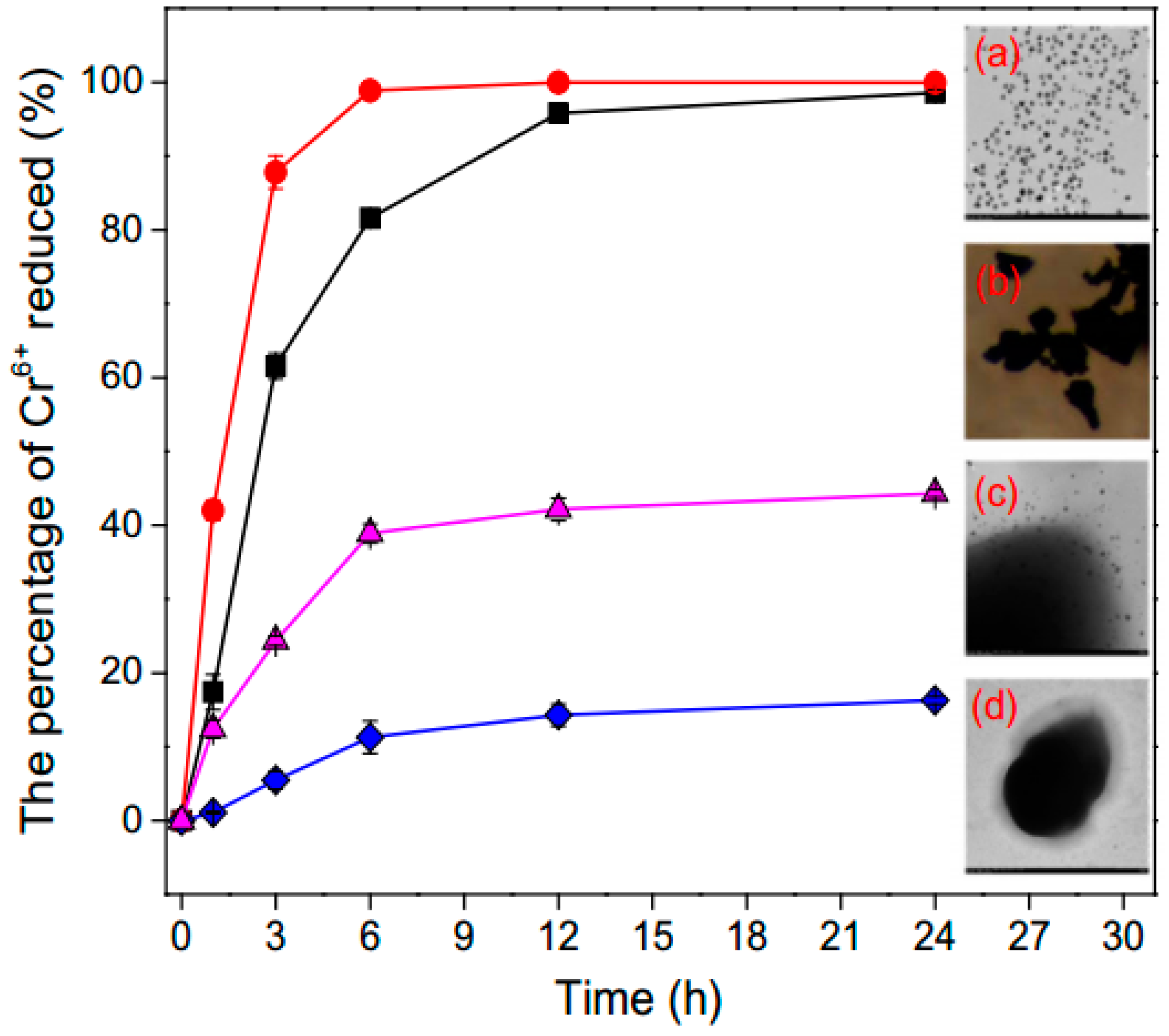
3.4. Removal of Heavy Metals Ions
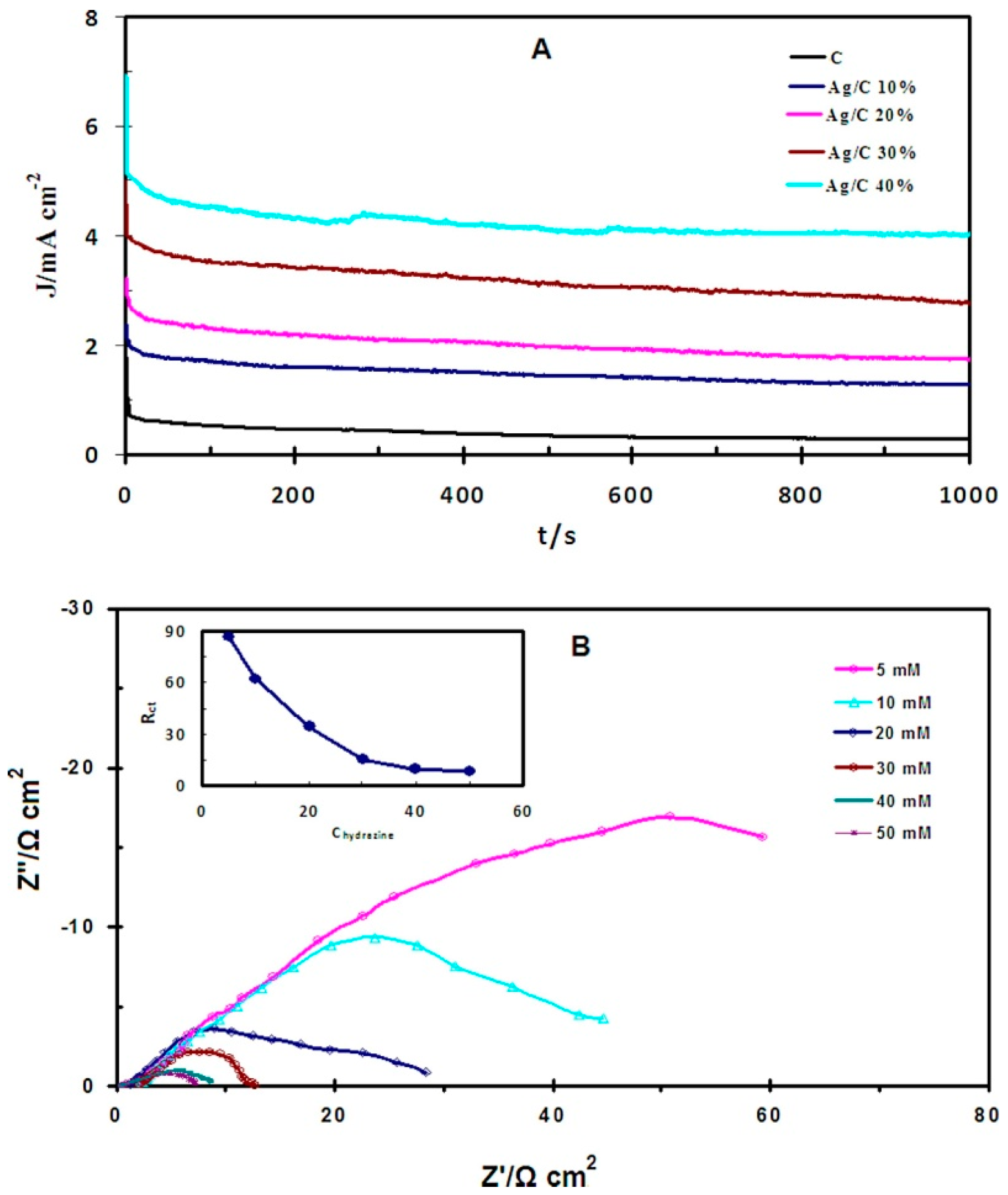
3.5. Other Catalytic Performance
4. Conclusion and Outlook
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Salunke, B.K.; Sawant, S.S.; Lee, S.I.; Kim, B.S. Microorganisms as efficient biosystem for the synthesis of metal nanoparticles: Current scenario and future possibilities. World J. Microbiol. Biotechnol. 2016, 32, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Bansal, V.; Rautaray, D.; Bharde, A.; Ahire, K.; Sanyal, A.; Ahmad, A.; Sastry, M. Fungus-mediated biosynthesis of silica and titania particles. J. Mater. Chem. 2005, 15, 2583–2589. [Google Scholar] [CrossRef]
- Bhushan, M.; Kumar, Y.; Periyasamy, L.; Viswanath, A.K. Facile synthesis of Fe/Zn oxide nanocomposites and study of their structural, magnetic, thermal, antibacterial and cytotoxic properties. Mater. Chem. Phys. 2018, 209, 233–248. [Google Scholar] [CrossRef]
- Labulo, A.H.; Adesuji, E.T.; Dedeke, O.A.; Bodede, O.S.; Oseghale, C.O.; Moodley, R.; Nyamori, V.O.; Dare, E.O.; Adegoke, O.A. A dual-purpose silver nanoparticles biosynthesized using aqueous leaf extract of Detarium microcarpum: An under-utilized species. Talanta 2016, 160, 735–744. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Li, Q.; Besenbacher, F.; Dong, M. Facile synthesis of single crystal PtSe2 nanosheets for nanoscale electronics. Adv. Mater. 2016, 28, 10224–10229. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Li, Q.; Xu, H.; Dahl-Petersen, C.; Yang, Q.; Cheng, D.; Cao, D.; Besenbacher, F.; Lauritsen, J.V.; Helveg, S.; et al. Controllable etching of MoS2 basal planes for enhanced hydrogen evolution through the formation of active edge sites. Nano Energy 2018, 49, 634–643. [Google Scholar] [CrossRef]
- Ma, L.; Su, W.; Liu, J.X.; Zeng, X.X.; Huang, Z.; Li, W.; Liu, Z.C.; Tang, J.X. Optimization for extracellular biosynthesis of silver nanoparticles by Penicillium aculeatum Su1 and their antimicrobial activity and cytotoxic effect compared with silver ions. Mater. Sci. Eng. C 2017, 77, 963–971. [Google Scholar] [CrossRef]
- Ovais, M.; Ahmad, I.; Khalil, A.T.; Mukherjee, S.; Javed, R.; Ayaz, M.; Raza, A.; Shinwari, Z.K. Wound healing applications of biogenic colloidal silver and gold nanoparticles: Recent trends and future prospects. Appl. Microbiol. Biotechnol. 2018, 102, 4305–4318. [Google Scholar] [CrossRef]
- Karunakaran, G.; Jagathambal, M.; Gusev, A.; Torres, J.A.L.; Kolesnikov, E.; Kuznetsov, D. Rapid Biosynthesis of AgNPs Using Soil Bacterium Azotobacter vinelandii With Promising Antioxidant and Antibacterial Activities for Biomedical Applications. JOM 2017, 69, 1206–1212. [Google Scholar] [CrossRef]
- Reverberi, A.P.; Kuznetsov, N.T.; Meshalkin, V.P.; Salerno, M.; Fabiano, B. Systematical analysis of chemical methods in metal nanoparticles synthesis. Theor. Found. Chem. Eng. 2016, 50, 59–66. [Google Scholar] [CrossRef]
- Gopinath, V.; Priyadarshini, S.; Loke, M.F.; Arunkumar, J.; Marsili, E.; MubarakAli, D.; Velusamy, P.; Vadivelu, J. Biogenic synthesis, characterization of antibacterial silver nanoparticles and its cell cytotoxicity. Arab. J. Chem. 2017, 10, 1107–1117. [Google Scholar] [CrossRef]
- Zhang, P.; Wang, Z.; Liu, L.; Klausen, L.H.; Wang, Y.; Mi, J.L.; Dong, M. Modulation the electronic property of 2D monolayer MoS2 by amino acid. Appl. Mater. Today 2019, 14, 151–158. [Google Scholar] [CrossRef]
- Wang, Z.; Li, Q.; Chen, Y.; Cui, B.; Li, Y.; Besenbacher, F.; Dong, M. The ambipolar transport behavior of WSe2 transistors and its analogue circuits. NPG Asia Mater. 2018, 10, 703–712. [Google Scholar] [CrossRef]
- Geng, J.; Jefferson, D.A.; Johnson, B.F.G. Exploring the structural complexities of metal-metalloid nanoparticles: The case of Ni·B as catalyst. Chemistry 2009, 15, 1134–1143. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Wu, H.H.; Li, Q.; Besenbacher, F.; Zeng, X.C.; Dong, M. Self-scrolling MoS2 metallic wires. Nanoscale 2018, 10, 18178–18185. [Google Scholar] [CrossRef] [PubMed]
- Ulug, B.; Haluk Turkdemir, M.; Cicek, A.; Mete, A. Role of irradiation in the green synthesis of silver nanoparticles mediated by fig (Ficus carica) leaf extract. Spectrochim. Acta Part A Mol. Biomol. Spectrosc. 2015, 135, 153–161. [Google Scholar] [CrossRef]
- Garole, D.J.; Choudhary, B.C.; Paul, D.; Borse, A.U. Sorption and recovery of platinum from simulated spent catalyst solution and refinery wastewater using chemically modified biomass as a novel sorbent. Environ. Sci. Pollut. Res. 2018, 25, 10911–10925. [Google Scholar] [CrossRef] [PubMed]
- Nazari, P.; Faramarzi, M.A.; Sepehrizadeh, Z.; Mofid, M.R.; Bazaz, R.D.; Shahverdi, A.R. Biosynthesis of bismuth nanoparticles using Serratia marcescens isolated from the Caspian Sea and their characterisation. IET Nanobiotechnol. 2012, 6, 58. [Google Scholar] [CrossRef]
- Reverberi, A.P.; Vocciante, M.; Lunghi, E.; Pietrelli, L.; Fabiano, B. New trends in the synthesis of nanoparticles by green methods. Chem. Eng. Trans. 2017, 61, 667–672. [Google Scholar] [CrossRef]
- Shukla, A.K.; Iravani, S. Metallic nanoparticles: Green synthesis and spectroscopic characterization. Environ. Chem. Lett. 2017, 15, 223–231. [Google Scholar] [CrossRef]
- Wen, X.; Du, C.; Zeng, G.; Huang, D.; Zhang, J.; Yin, L.; Tan, S.; Huang, L.; Chen, H.; Yu, G.; et al. A novel biosorbent prepared by immobilized Bacillus licheniformis for lead removal from wastewater. Chemosphere 2018, 200, 173–179. [Google Scholar] [CrossRef] [PubMed]
- Won, S.W.; Mao, J.; Kwak, I.S.; Sathishkumar, M.; Yun, Y.S. Platinum recovery from ICP wastewater by a combined method of biosorption and incineration. Bioresour. Technol. 2010, 101, 1135–1140. [Google Scholar] [CrossRef] [PubMed]
- Xiang, L.; Wei, J.; Jianbo, S.; Guili, W.; Feng, G.; Ying, L. Purified and sterilized magnetosomes from Magnetospirillum gryphiswaldense MSR-1 were not toxic to mouse fibroblasts in vitro. Lett. Appl. Microbiol. 2007, 45, 75–81. [Google Scholar] [CrossRef] [PubMed]
- Saravana Kumar, P.; Balachandran, C.; Duraipandiyan, V.; Ramasamy, D.; Ignacimuthu, S.; Al-Dhabi, N.A. Extracellular biosynthesis of silver nanoparticle using Streptomyces sp. 09 PBT 005 and its antibacterial and cytotoxic properties. Appl. Nanosci. 2015, 5, 169–180. [Google Scholar] [CrossRef]
- Verma, S.K.; Jha, E.; Panda, P.K.; Mishra, A.; Thirumurugan, A.; Das, B.; Parashar, S.K.S.; Suar, M. Rapid novel facile biosynthesized silver nanoparticles from bacterial release induce biogenicity and concentration dependent in vivo cytotoxicity with embryonic Zebrafish-A mechanistic insight. Toxicol. Sci. 2018, 161, 125–138. [Google Scholar] [CrossRef] [PubMed]
- Tamboli, D.P.; Lee, D.S. Mechanistic antimicrobial approach of extracellularly synthesized silver nanoparticles against gram positive and gram negative bacteria. J. Hazard. Mater. 2013, 260, 878–884. [Google Scholar] [CrossRef] [PubMed]
- Ma, B.; Kong, C.; Hu, X.; Liu, K.; Huang, Q.; Lv, J.; Lu, W.; Zhang, X.; Yang, Z.; Yang, S. A sensitive electrochemical nonenzymatic biosensor for the detection of H2O2 released from living cells based on ultrathin concave Ag nanosheets. Biosens. Bioelectron. 2018, 106, 29–36. [Google Scholar] [CrossRef]
- Naim, M.M.; El-Shafei, A.A.; Elewa, M.M.; Moneer, A.A. Application of silver-, iron-, and chitosan-nanoparticles in wastewater treatment. Int. Conf. Eur. Desalin. Soc. Desalin. Environ. Clean Water Energy 2016, 73, 268–280. [Google Scholar] [CrossRef]
- Liu, W.; Tian, S.; Zhao, X.; Xie, W.; Gong, Y.; Zhao, D. Application of Stabilized Nanoparticles for In Situ Remediation of Metal-Contaminated Soil and Groundwater: A Critical Review. Curr. Pollut. Rep. 2015, 1, 280–291. [Google Scholar] [CrossRef]
- Vanalakar, S.A.; Patil, P.S.; Kim, J.H. Recent advances in synthesis of Cu2FeSnS4materials for solar cell applications: A review. Sol. Energy Mater. Sol. Cells 2018, 182, 204–219. [Google Scholar] [CrossRef]
- Pantidos, N. Biological Synthesis of Metallic Nanoparticles by Bacteria, Fungi and Plants. J. Nanomed. Nanotechnol. 2014, 5, 233. [Google Scholar] [CrossRef]
- Ahmed, S.; Annu; Ikram, S.; Yudha, S. Biosynthesis of gold nanoparticles: A green approach. J. Photochem. Photobiol. B Biol. 2016, 161, 141–153. [Google Scholar] [CrossRef] [PubMed]
- Timoszyk, A.; Niedbach, J.; Śliżewska, P.; Mirończyk, A.; Kozioł, J.J. Eco-Friendly and Temperature Dependent Biosynthesis of Gold Nanoparticles Using the Bacterium Pseudomonas aeruginosa: Characterization and Antibacterial Activity. J. Nano Res. 2017, 48, 114–124. [Google Scholar] [CrossRef]
- Mekkawy, A.I.; El-Mokhtar, M.A.; Nafady, N.A.; Yousef, N.; Hamad, M.; El-Shanawany, S.M.; Ibrahim, E.H.; Elsabahy, M. In vitro and in vivo evaluation of biologically synthesized silver nanoparticles for topical applications: Effect of surface coating and loading into hydrogels. Int. J. Nanomed. 2017, 12, 759–777. [Google Scholar] [CrossRef] [PubMed]
- Siddiqi, K.S.; Husen, A.; Rao, R.A.K. A review on biosynthesis of silver nanoparticles and their biocidal properties. J. Nanobiotechnol. 2018, 16. [Google Scholar] [CrossRef] [PubMed]
- Klaus, T.; Joerger, R.; Olsson, E.; Granqvist, C.-G. Silver-based crystalline nanoparticles, microbially fabricated. Proc. Natl. Acad. Sci. USA 1999, 96, 13611–13614. [Google Scholar] [CrossRef] [PubMed]
- Pereira, L.; Mehboob, F.; Stams, A.J.M.; Mota, M.M.; Rijnaarts, H.H.M.; Alves, M.M. Metallic nanoparticles: Microbial synthesis and unique properties for biotechnological applications, bioavailability and biotransformation. Crit. Rev. Biotechnol. 2015, 35, 114–128. [Google Scholar] [CrossRef]
- Ghiuță, I.; Cristea, D.; Croitoru, C.; Kost, J.; Wenkert, R.; Vyrides, I.; Anayiotos, A.; Munteanu, D. Characterization and antimicrobial activity of silver nanoparticles, biosynthesized using Bacillus species. Appl. Surf. Sci. 2018, 438, 66–73. [Google Scholar] [CrossRef]
- Yurtluk, T.; Akçay, F.A.; Avcı, A. Biosynthesis of silver nanoparticles using novel Bacillus sp. SBT8. Prep. Biochem. Biotechnol. 2018, 48, 151–159. [Google Scholar] [CrossRef]
- Mishra, S.; Singh, B.R.; Naqvi, A.H.; Singh, H.B. Potential of biosynthesized silver nanoparticles using Stenotrophomonas sp. BHU-S7 (MTCC 5978) for management of soil-borne and foliar phytopathogens. Sci. Rep. 2017, 7, 1–15. [Google Scholar] [CrossRef]
- Li, J.; Tian, B.; Li, T.; Dai, S.; Weng, Y.; Lu, J.; Xu, X.; Jin, Y.; Pang, R.; Hua, Y. Biosynthesis of Au, Ag and Au–Ag bimetallic nanoparticles using protein extracts of Deinococcus radiodurans and evaluation of their cytotoxicity. Int. J. Nanomed. 2018, 13, 1411–1424. [Google Scholar] [CrossRef] [PubMed]
- Wadhwani, S.A.; Shedbalkar, U.U.; Singh, R.; Chopade, B.A. Biosynthesis of gold and selenium nanoparticles by purified protein from Acinetobacter sp. SW 30. Enzyme Microb. Technol. 2018, 111, 81–86. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Li, Q.; Ma, X.; Tian, B.; Li, T.; Yu, J.; Dai, S.; Weng, Y.; Hua, Y. Biosynthesis of gold nanoparticles by the extreme bacterium Deinococcus radiodurans and an evaluation of their antibacterial properties. Int. J. Nanomed. 2016, 11, 5931–5944. [Google Scholar] [CrossRef] [PubMed]
- Nair, B.; Pradeep, T. Coalescence of Nanoclusters and Formation of Submicron Crystallites Assisted by Lactobacillus Strains. Cryst. Growth Des. 2002, 2, 293–298. [Google Scholar] [CrossRef]
- Suresh, A.K.; Pelletier, D.A.; Wang, W.; Moon, J.-W.; Gu, B.; Mortensen, N.P.; Allison, D.P.; Joy, D.C.; Phelps, T.J.; Doktycz, M.J. Silver Nanocrystallites: Biofabrication using Shewanella oneidensis, and an Evaluation of Their Comparative Toxicity on Gram-negative and Gram-positive Bacteria. Environ. Sci. Technol. 2010, 44, 5210–5215. [Google Scholar] [CrossRef] [PubMed]
- Venil, C.K.; Sathishkumar, P.; Malathi, M.; Usha, R.; Jayakumar, R.; Yusoff, A.R.M.; Ahmad, W.A. Synthesis of flexirubin-mediated silver nanoparticles using Chryseobacterium artocarpi CECT 8497 and investigation of its anticancer activity. Mater. Sci. Eng. C 2016, 59, 228–234. [Google Scholar] [CrossRef] [PubMed]
- Lv, Q.; Zhang, B.; Xing, X.; Zhao, Y.; Cai, R.; Wang, W.; Gu, Q. Biosynthesis of copper nanoparticles using Shewanella loihica PV-4 with antibacterial activity: Novel approach and mechanisms investigation. J. Hazard. Mater. 2018, 347, 141–149. [Google Scholar] [CrossRef]
- Abirami, M.; Kannabiran, K. Streptomyces ghanaensis VITHM1 mediated green synthesis of silver nanoparticles: Mechanism and biological applications. Front. Chem. Sci. Eng. 2016, 10, 542–551. [Google Scholar] [CrossRef]
- Kimber, R.L.; Lewis, E.A.; Parmeggiani, F.; Smith, K.; Bagshaw, H.; Starborg, T.; Joshi, N.; Figueroa, A.I.; van der Laan, G.; Cibin, G.; et al. Biosynthesis and Characterization of Copper Nanoparticles Using Shewanella oneidensis: Application for Click Chemistry. Small 2018, 14, 1–8. [Google Scholar] [CrossRef]
- Ha, C.; Zhu, N.; Shang, R.; Shi, C.; Cui, J.; Sohoo, I.; Wu, P.; Cao, Y. Biorecovery of palladium as nanoparticles by Enterococcus faecalis and its catalysis for chromate reduction. Chem. Eng. J. 2016, 288, 246–254. [Google Scholar] [CrossRef]
- Dong, Z.Y.; Rao, M.P.N.; Xiao, M.; Wang, H.F.; Hozzein, W.N.; Chen, W.; Li, W.J. Antibacterial activity of silver nanoparticles against Staphylococcus warneri synthesized using endophytic bacteria by photo-irradiation. Front. Microbiol. 2017, 8, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Tian, L.-J.; Li, W.-W.; Zhu, T.-T.; Chen, J.-J.; Wang, W.-K.; An, P.-F.; Zhang, L.; Dong, J.-C.; Guan, Y.; Liu, D.-F.; et al. Directed Biofabrication of Nanoparticles through Regulating Extracellular Electron Transfer. J. Am. Chem. Soc. 2017, 139, 12149–12152. [Google Scholar] [CrossRef] [PubMed]
- Forootanfar, H.; Amirpour-Rostami, S.; Jafari, M.; Forootanfar, A.; Yousefizadeh, Z.; Shakibaie, M. Microbial-assisted synthesis and evaluation the cytotoxic effect of tellurium nanorods. Mater. Sci. Eng. C 2015, 49, 183–189. [Google Scholar] [CrossRef] [PubMed]
- Cheng, Y.; Xiao, X.; Li, X.; Song, D.; Lu, Z.; Wang, F.; Wang, Y. Characterization, antioxidant property and cytoprotection of exopolysaccharide-capped elemental selenium particles synthesized by Bacillus paralicheniformis SR14. Carbohydr. Polym. 2017, 178, 18–26. [Google Scholar] [CrossRef] [PubMed]
- Shoeibi, S.; Mozdziak, P.; Golkar-Narenji, A. Biogenesis of Selenium Nanoparticles Using Green Chemistry. Top. Curr. Chem. 2017, 375, 1–21. [Google Scholar] [CrossRef]
- Dias, D.A.; Kouremenos, K.A.; Beale, D.J.; Callahan, D.L.; Jones, O.A.H. Metal and metalloid containing natural products and a brief overview of their applications in biology, biotechnology and biomedicine. BioMetals 2016, 29, 1–13. [Google Scholar] [CrossRef]
- Presentato, A.; Piacenza, E.; Anikovskiy, M.; Cappelletti, M.; Zannoni, D.; Turner, R.J. Biosynthesis of selenium-nanoparticles and -nanorods as a product of selenite bioconversion by the aerobic bacterium Rhodococcus aetherivorans BCP1. New Biotechnol. 2018, 41, 1–8. [Google Scholar] [CrossRef]
- Song, D.; Li, X.; Cheng, Y.; Xiao, X.; Lu, Z.; Wang, Y.; Wang, F. Aerobic biogenesis of selenium nanoparticles by Enterobacter cloacae Z0206 as a consequence of fumarate reductase mediated selenite reduction. Sci. Rep. 2017, 7, 19–21. [Google Scholar] [CrossRef]
- Xu, C.; Qiao, L.; Guo, Y.; Ma, L.; Cheng, Y. Preparation, characteristics and antioxidant activity of polysaccharides and proteins-capped selenium nanoparticles synthesized by Lactobacillus casei ATCC 393. Carbohydr. Polym. 2018, 195, 576–585. [Google Scholar] [CrossRef]
- Wang, X.; Zhang, D.; Pan, X.; Lee, D.J.; Al-Misned, F.A.; Mortuza, M.G.; Gadd, G.M. Aerobic and anaerobic biosynthesis of nano-selenium for remediation of mercury contaminated soil. Chemosphere 2017, 170, 266–273. [Google Scholar] [CrossRef]
- Avendaño, R.; Chaves, N.; Fuentes, P.; Sánchez, E.; Jiménez, J.I.; Chavarría, M. Production of selenium nanoparticles in Pseudomonas putida KT2440. Sci. Rep. 2016, 6, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Zonaro, E.; Lampis, S.; Turner, R.J.; Junaid, S.; Vallini, G. Biogenic selenium and tellurium nanoparticles synthesized by environmental microbial isolates efficaciously inhibit bacterial planktonic cultures and biofilms. Front. Microbiol. 2015, 6, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Srivastava, P.; Nikhil, E.V.R.; Bragança, J.M.; Kowshik, M. Anti-bacterial TeNPs biosynthesized by haloarcheaon Halococcus salifodinae BK3. Extremophiles 2015, 19, 875–884. [Google Scholar] [CrossRef] [PubMed]
- Presentato, A.; Piacenza, E.; Anikovskiy, M.; Cappelletti, M.; Zannoni, D.; Turner, R.J. Rhodococcus aetherivorans BCP1 as cell factory for the production of intracellular tellurium nanorods under aerobic conditions. Microb. Cell Fact. 2016, 15, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Jeevanandam, J.; Barhoum, A.; Chan, Y.S.; Dufresne, A.; Danquah, M.K. Review on nanoparticles and nanostructured materials: History, sources, toxicity and regulations. Beilstein J. Nanotechnol. 2018, 9, 1050–1074. [Google Scholar] [CrossRef] [PubMed]
- Fabricius, A.L.; Duester, L.; Ecker, D.; Ternes, T.A. Metal and metalloid size-fractionation strategies in spatial high-resolution sediment pore water profiles. Environ. Sci. Technol. 2016, 50, 9506–9514. [Google Scholar] [CrossRef]
- Obayemi, J.D.; Dozie-Nwachukwu, S.; Danyuo, Y.; Odusanya, O.S.; Anuku, N.; Malatesta, K.; Soboyejo, W.O. Biosynthesis and the conjugation of magnetite nanoparticles with luteinizing hormone releasing hormone (LHRH). Mater. Sci. Eng. C 2015, 46, 482–496. [Google Scholar] [CrossRef]
- Rajendran, K.; Karunagaran, V.; Mahanty, B.; Sen, S. Biosynthesis of hematite nanoparticles and its cytotoxic effect on HepG2 cancer cells. Int. J. Biol. Macromol. 2015, 74, 376–381. [Google Scholar] [CrossRef]
- Saif, S.; Tahir, A.; Chen, Y. Green Synthesis of Iron Nanoparticles and Their Environmental Applications and Implications. Nanomaterials 2016, 6, 209. [Google Scholar] [CrossRef]
- Srivastava, N.; Mukhopadhyay, M. Biosynthesis of SnO2 Nanoparticles Using Bacterium Erwinia herbicola and Their Photocatalytic Activity for Degradation of Dyes. Ind. Eng. Chem. Res. 2014, 53, 13971–13979. [Google Scholar] [CrossRef]
- Ghasemi, N.; Jamali-Sheini, F.; Zekavati, R. CuO and Ag/CuO nanoparticles: Biosynthesis and antibacterial properties. Mater. Lett. 2017, 196, 78–82. [Google Scholar] [CrossRef]
- Bakhshi, M.; Hosseini, M.R. Synthesis of CdS nanoparticles from cadmium sulfate solutions using the extracellular polymeric substances of B. licheniformis as stabilizing agent. Enzyme Microb. Technol. 2016, 95, 209–216. [Google Scholar] [CrossRef] [PubMed]
- Yan, Z.Y.; Du, Q.Q.; Qian, J.; Wan, D.Y.; Wu, S.M. Eco-friendly intracellular biosynthesis of CdS quantum dots without changing Escherichia coli’s antibiotic resistance. Enzyme Microb. Technol. 2017, 96, 96–102. [Google Scholar] [CrossRef]
- Sakimoto, K.K.; Wong, A.B.; Yang, P. Self-photosensitization of nonphotosynthetic bacteria for solar-to-chemical production. Science 2016, 351, 74–77. [Google Scholar] [CrossRef] [PubMed]
- Qi, P.; Zhang, D.; Zeng, Y.; Wan, Y. Biosynthesis of CdS nanoparticles: A fluorescent sensor for sulfate-reducing bacteria detection. Talanta 2016, 147, 142–146. [Google Scholar] [CrossRef] [PubMed]
- Kominkova, M.; Milosavljevic, V.; Vitek, P.; Polanska, H.; Cihalova, K.; Dostalova, S.; Hynstova, V.; Guran, R.; Kopel, P.; Richtera, L.; et al. Comparative study on toxicity of extracellularly biosynthesized and laboratory synthesized CdTe quantum dots. J. Biotechnol. 2017, 241, 193–200. [Google Scholar] [CrossRef] [PubMed]
- Yue, L.; Wang, J.; Zhang, Y.; Qi, S.; Xin, B. Controllable biosynthesis of high-purity lead-sulfide (PbS) nanocrystals by regulating the concentration of polyethylene glycol in microbial system. Bioprocess Biosyst. Eng. 2016, 39, 1839–1846. [Google Scholar] [CrossRef]
- Srivastava, P.; Kowshik, M. Fluorescent Lead(IV) Sulfide Nanoparticles Synthesized by Idiomarina sp. Strain PR58-8 for Bioimaging Applications. Appl. Environ. Microbiol. 2017, 83, e03091-16. [Google Scholar] [CrossRef]
- Xiao, X.; Liu, Q.-Y.; Lu, X.-R.; Li, T.-T.; Feng, X.-L.; Li, Q.; Liu, Z.-Y.; Feng, Y.-J. Self-assembly of complex hollow CuS nano/micro shell by an electrochemically active bacterium Shewanella oneidensis MR-1. Int. Biodeterior. Biodegrad. 2017, 116, 10–16. [Google Scholar] [CrossRef]
- Zhou, N.Q.; Tian, L.J.; Wang, Y.C.; Li, D.B.; Li, P.P.; Zhang, X.; Yu, H.Q. Extracellular biosynthesis of copper sulfide nanoparticles by Shewanella oneidensis MR-1 as a photothermal agent. Enzym. Microb. Technol. 2016, 95, 230–235. [Google Scholar] [CrossRef]
- Xiao, X.; Ma, X.B.; Yuan, H.; Liu, P.C.; Lei, Y.B.; Xu, H.; Du, D.L.; Sun, J.F.; Feng, Y.J. Photocatalytic properties of zinc sulfide nanocrystals biofabricated by metal-reducing bacterium Shewanella oneidensis MR-1. J. Hazard. Mater. 2015, 288, 134–139. [Google Scholar] [CrossRef] [PubMed]
- Qi, S.; Zhang, M.; Guo, X.; Yue, L.; Wang, J.; Shao, Z.; Xin, B. Controlled extracellular biosynthesis of ZnS quantum dots by sulphate reduction bacteria in the presence of hydroxypropyl starch as a mediator. J. Nanopart. Res. 2017, 19. [Google Scholar] [CrossRef]
- Doshi, B.; Sillanpää, M.; Kalliola, S. A review of bio-based materials for oil spill treatment. Water Res. 2018, 135, 262–277. [Google Scholar] [CrossRef] [PubMed]
- Hu, X.; Ren, C.; Kang, W.; Mu, L.; Liu, X.; Li, X.; Wang, T.; Zhou, Q. Characterization and toxicity of nanoscale fragments in wastewater treatment plant effluent. Sci. Total Environ. 2018, 626, 1332–1341. [Google Scholar] [CrossRef] [PubMed]
- Schröfel, A.; Kratošová, G.; Šafařík, I.; Šafaříková, M.; Raška, I.; Shor, L.M. Applications of biosynthesized metallic nanoparticles—A review. Acta Biomater. 2014, 10, 4023–4042. [Google Scholar] [CrossRef] [PubMed]
- Choi, S.; Johnston, M.; Wang, G.S.; Huang, C.P. A seasonal observation on the distribution of engineered nanoparticles in municipal wastewater treatment systems exemplified by TiO2 and ZnO. Sci. Total Environ. 2018, 625, 1321–1329. [Google Scholar] [CrossRef] [PubMed]
- Seifan, M.; Ebrahiminezhad, A.; Ghasemi, Y.; Samani, A.K.; Berenjian, A. The role of magnetic iron oxide nanoparticles in the bacterially induced calcium carbonate precipitation. Appl. Microbiol. Biotechnol. 2018, 102, 3595–3606. [Google Scholar] [CrossRef] [PubMed]
- Huang, Z.; Zeng, Z.; Chen, A.; Zeng, G.; Xiao, R.; Xu, P.; He, K.; Song, Z.; Hu, L.; Peng, M.; et al. Differential behaviors of silver nanoparticles and silver ions towards cysteine: Bioremediation and toxicity to Phanerochaete chrysosporium. Chemosphere 2018, 203, 199–208. [Google Scholar] [CrossRef]
- Zhang, Y.; Qiang, L.; Yuan, Y.; Wu, W.; Sun, B.; Zhu, L. Impacts of titanium dioxide nanoparticles on transformation of silver nanoparticles in aquatic environments. Environ. Sci. Nano 2018, 5, 1191–1199. [Google Scholar] [CrossRef]
- Che, L.; Dong, Y.; Wu, M.; Zhao, Y.; Liu, L.; Zhou, H. Characterization of Selenite Reduction by Lysinibacillus sp. ZYM-1 and Photocatalytic Performance of Biogenic Selenium Nanospheres. ACS Sustain. Chem. Eng. 2017, 5, 2535–2543. [Google Scholar] [CrossRef]
- Baxter-Plant, V.S.; Mikheenko, I.P.; Macaskie, L.E. Sulphate-reducing bacteria, palladium and the reductive dehalogenation of chlorinated aromatic compounds. Biodegradation 2003, 14, 83–90. [Google Scholar] [CrossRef] [PubMed]
- Baxter-Plant, V.S.; Mikheenko, I.P.; Robson, M.; Harrad, S.J.; Macaskie, L.E. Dehalogenation of chlorinated aromatic compounds using a hybrid bioinorganic catalyst on cells of Desulfovibrio desulfuricans. Biotechnol. Lett. 2004, 26, 1885–1890. [Google Scholar] [CrossRef] [PubMed]
- De Windt, W.; Aelterman, P.; Verstraete, W. Bioreductive deposition of palladium (0) nanoparticles on Shewanella oneidensis with catalytic activity towards reductive dechlorination of polychlorinated biphenyls. Environ. Microbiol. 2005, 7, 314–325. [Google Scholar] [CrossRef] [PubMed]
- Hosseinpour, M.; Fatemi, S.; Ahmadi, S.J.; Morimoto, M.; Akizuki, M.; Oshima, Y.; Fumoto, E. The synergistic effect between supercritical waterand redox properties of iron oxide nanoparticlesfor in-situ catalyticupgrading heavy oil with formic acid. Isotopic study. Appl. Catal. B Environ. 2018, 230, 91–101. [Google Scholar] [CrossRef]
- Zhang, H.; Hu, X. Rapid production of Pd nanoparticle by a marine electrochemically active bacterium: Shewanella sp. CNZ-1 and its catalytic performance on 4-nitrophenol reduction. RSC Adv. 2017, 7, 41182–41189. [Google Scholar] [CrossRef]
- Zhang, H.; Hu, X. Biosynthesis of Pd and Au as nanoparticles by a marine bacterium Bacillus sp. GP and their enhanced catalytic performance using metal oxides for 4-nitrophenol reduction. Enzyme Microb. Technol. 2018, 113, 59–66. [Google Scholar] [CrossRef]
- Cumbal, L.; Greenleaf, J.; Leun, D.; SenGupta, A.K. Polymer supported inorganic nanoparticles: Characterization and environmental applications. React. Funct. Polym. 2003, 54, 167–180. [Google Scholar] [CrossRef]
- Dong, B.; Liu, G.; Zhou, J.; Wang, A.; Wang, J.; Jin, R.; Lv, H. Biogenic gold nanoparticles-reduced graphene oxide nanohybrid: Synthesis, characterization and application in chemical and biological reduction of nitroaromatics. RSC Adv. 2015, 5, 97798–97806. [Google Scholar] [CrossRef]
- Rüdel, H.; Díaz Muñiz, C.; Garelick, H.; Kandile, N.G.; Miller, B.W.; Pantoja Munoz, L.; Peijnenburg, W.J.G.M.; Purchase, D.; Shevah, Y.; van Sprang, P.; et al. Consideration of the bioavailability of metal/metalloid species in freshwaters: Experiences regarding the implementation of biotic ligand model-based approaches in risk assessment frameworks. Environ. Sci. Pollut. Res. 2015, 22, 7405–7421. [Google Scholar] [CrossRef]
- Vilela, P.; Liu, H.; Lee, S.C.; Hwangbo, S.; Nam, K.J.; Yoo, C.K. A systematic approach of removal mechanisms, control and optimization of silver nanoparticle in wastewater treatment plants. Sci. Total Environ. 2018, 633, 989–998. [Google Scholar] [CrossRef]
- Vilardi, G.; Mpouras, T.; Dermatas, D.; Verdone, N.; Polydera, A.; Di Palma, L. Nanomaterials application for heavy metals recovery from polluted water: The combination of nano zero-valent iron and carbon nanotubes. Competitive adsorption non-linear modeling. Chemosphere 2018, 201, 716–729. [Google Scholar] [CrossRef] [PubMed]
- Wang, G.; Zhang, B.; Li, S.; Yang, M.; Yin, C. Simultaneous microbial reduction of vanadium (V) and chromium (VI) by Shewanella loihica PV-4. Bioresour. Technol. 2017, 227, 353–358. [Google Scholar] [CrossRef] [PubMed]
- Bunge, M.; Søbjerg, L.S.; Rotaru, A.E.; Gauthier, D.; Lindhardt, A.T.; Hause, G.; Finster, K.; Kingshott, P.; Skrydstrup, T.; Meyer, R.L. Formation of palladium(0) nanoparticles at microbial surfaces. Biotechnol. Bioeng. 2010, 107, 206–215. [Google Scholar] [CrossRef] [PubMed]
- Rostami, H.; Khosravi, F.; Mohseni, M.; Rostami, A.A. Biosynthesis of Ag nanoparticles using isolated bacteria from contaminated sites and its application as an efficient catalyst for hydrazine electrooxidation. Int. J. Biol. Macromol. 2018, 107, 343–348. [Google Scholar] [CrossRef]
- Velmurugan, P.; Hong, S.C.; Aravinthan, A.; Jang, S.H.; Yi, P.I.; Song, Y.C.; Jung, E.S.; Park, J.S.; Sivakumar, S. Comparison of the Physical Characteristics of Green-Synthesized and Commercial Silver Nanoparticles: Evaluation of Antimicrobial and Cytotoxic Effects. Arab. J. Sci. Eng. 2017, 42, 201–208. [Google Scholar] [CrossRef]
- Pfeffer, C.; Larsen, S.; Song, J.; Dong, M.; Besenbacher, F.; Meyer, R.L.; Kjeldsen, K.U.; Schreiber, L.; Gorby, Y.A.; El-Naggar, M.Y.; et al. Filamentous bacteria transport electrons over centimetre distances. Nature 2012, 491, 218–221. [Google Scholar] [CrossRef]
- Jiang, Z.; Zhang, S.; Klausen, L.H.; Song, J.; Li, Q.; Wang, Z.; Stokke, B.T.; Huang, Y.; Besenbacher, F.; Nielsen, L.P.; et al. In vitro single-cell dissection revealing the interior structure of cable bacteria. Proc. Natl. Acad. Sci. USA 2018, 115, 8515–8522. [Google Scholar] [CrossRef]
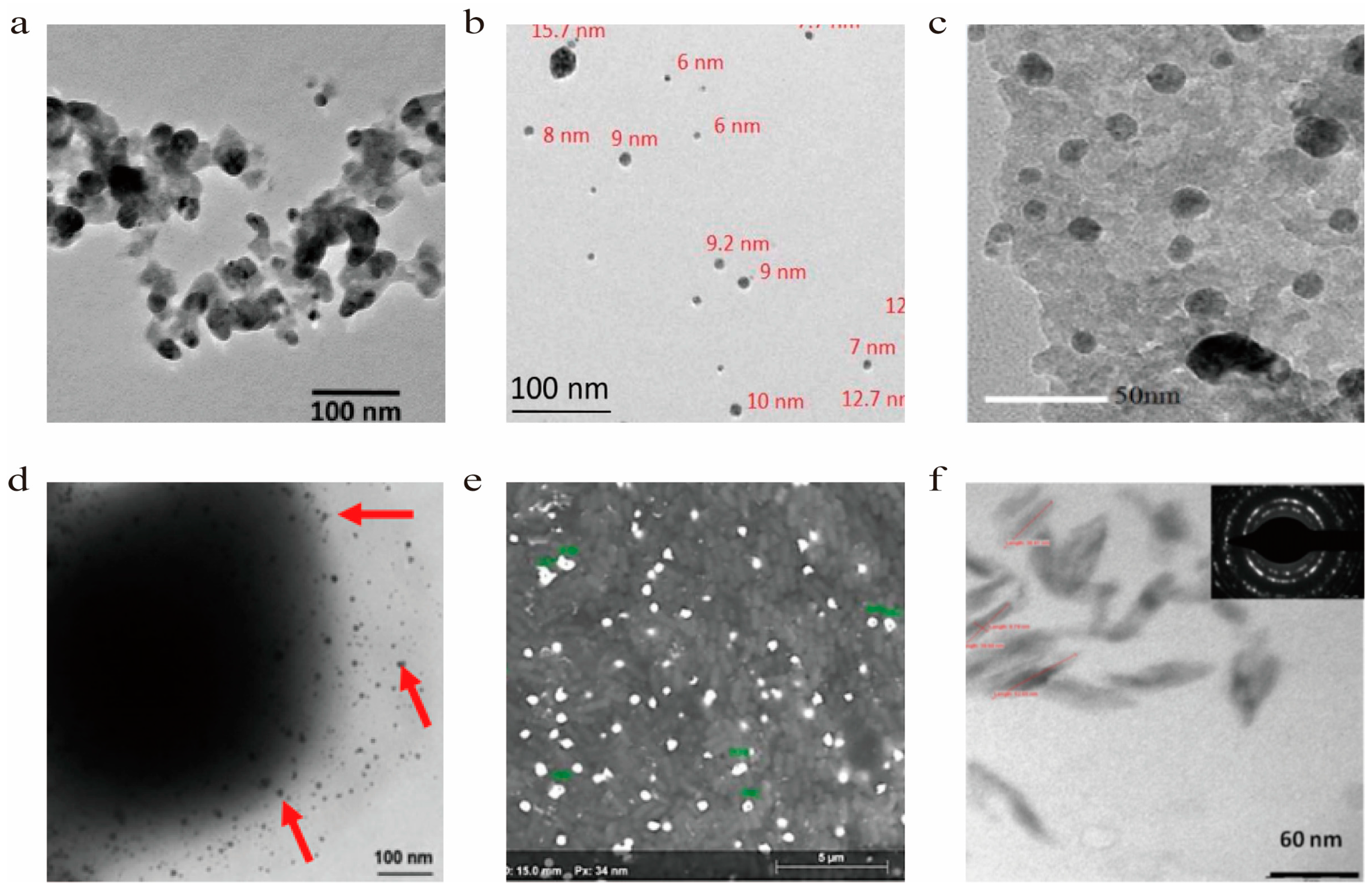
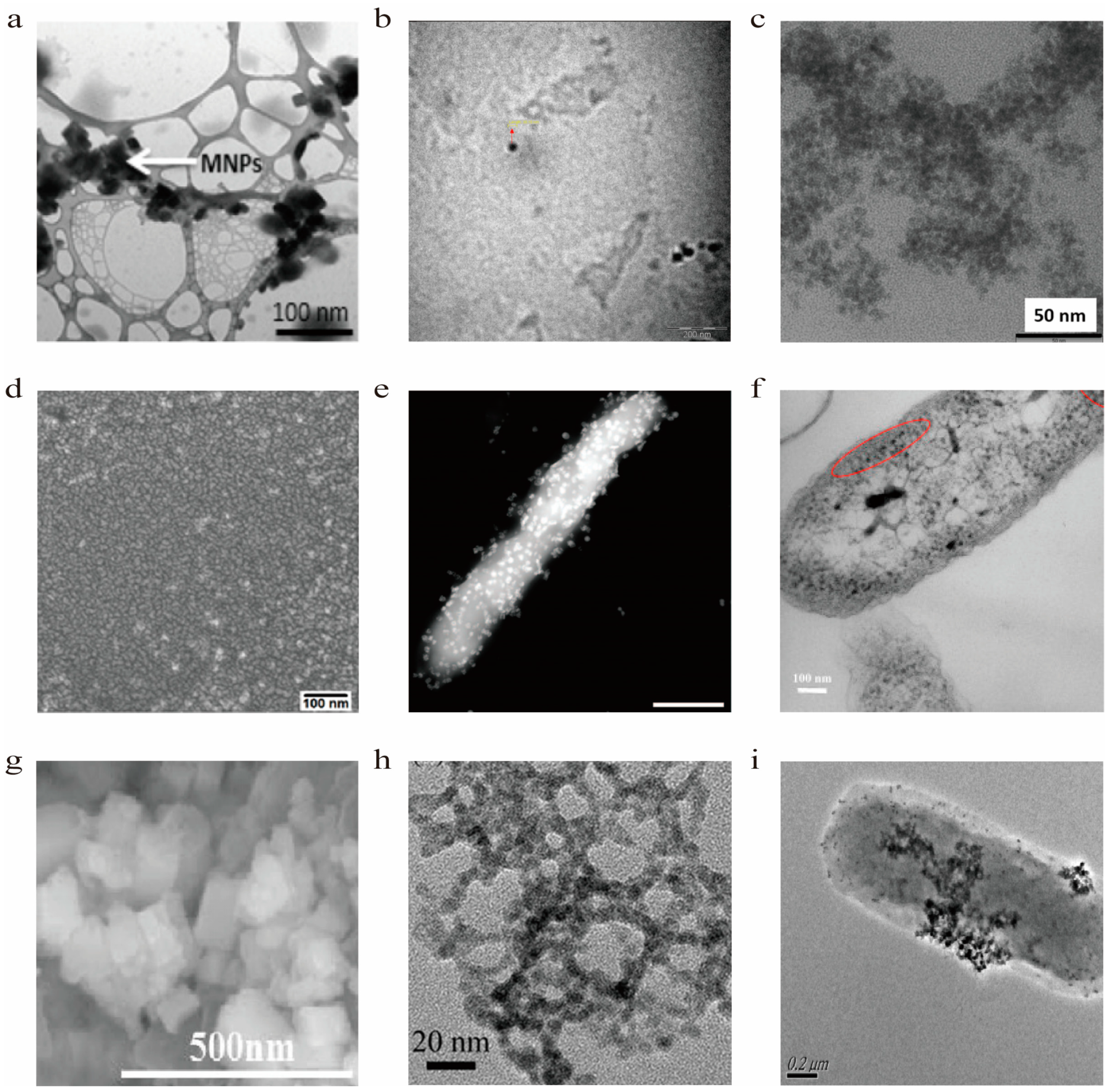


| NPs | Cellular | Shape | Size | Type of bacteria | Ref. |
|---|---|---|---|---|---|
| Ag | Extra- | Spherical NPs | 6~16 nm | Pseudomonas putida MVP2 | [12] |
| Extra- | Spherical NPs | 198~595 nm | Streptomyces sp. 09 PBT 005 | [24] | |
| Extra- | Spherical NPs | 4.4 nm | Exiguobacterium sp. KNU1 | [26] | |
| Extra- | Spherical NPs | 140 nm | Bacillus species | [38] | |
| Extra- | Spherical NPs | 12 nm | Stenotrophomonas sp. BHU-S7 | [40] | |
| Extra- | Spherical NPs | 30~50 nm | Streptomyces ghanaensis VITHM1 | [48] | |
| Intra- | Equilateral triangles, hexagons NPs | 200 nm | Pseudomonas stutzeri AG259 | [36] | |
| Intra- | NA | 15~500 nm | Lactobacillus | [44] | |
| NA | NA | 37.13 ± 5.97 nm | Deinococcusradiodurans | [41] | |
| NA | Spherical NPs | 2~11 nm | Shewanellaoneidensis MR-1 | [45] | |
| NA | Spherical and irregular NPs | 49 nm | Chryseobacteriumartocarpi CECT 8497 | [46] | |
| NA | Spherical NPs | 11~40 nm | Endophytic Bacteria | [51] | |
| Au | Intra- | NA | 20~50 nm | Lactobacillus | [44] |
| NA | Spherical NPs | 10 ± 2 nm | cinetobacter sp. SW 30 | [42] | |
| NA | NA | 51.72 ± 7.38 nm | Deinococcusradiodurans | [41] | |
| Au-Ag | Intra- | NA | 100~300 nm | Lactobacillus | [44] |
| Cu | Intra- | Spherical NPs | 20~40 nm | Shewanellaoneidensis MR1 | [49] |
| Extra- | Spherical NPs | 10~16 nm | Shewanellaloihica PV-4 | [47] | |
| Pd | Intra- & Extra- | NA | < 10 nm | Enterococcus faecalis | [50] |
| Pb | Intra- | Spherical NPs | 3~8 nm | Shewanella sp. KR-12 | [19] |
| Bi | Intra- | Irregular NPs | < 150 nm | Serratia marcescens | [18] |
| Se | Extra- | Spherical NPs | 50~500 nm | Actinomycete Rhodococcus aetherivorans | [57] |
| Extra- | Spherical NPs | 100~500 nm | Pseudomonas putida | [61] | |
| Intra- | Spherical NPs | 104.6 ± 8.4 nm | Shewanellaoneidensis MR-1 | [52] | |
| Intra- & Extra- | Rods | 100~300 nm | Enterobacter cloacae Z0206 | [58] | |
| Intra-& Extra- | Amorphous NPs | 580 ± 109 nm | Citrobacter freundii Y9 | [60] | |
| NA | Amorphous | 100 ± 10 nm | cinetobacter sp. SW 30 | [42] | |
| NA | NA | 6 h: 221.1 nm, 24 h: 345.2 nm, 48 h: 357.1 nm | Stenotrophomona smaltophilia SeITE02 | [62] | |
| Te | Intra- | Hexagonal needle-shaped NPs | diameter: 10 nm, length: 44 nm | Halococcus salifodinae BK3 | [64] |
| NA | NA | 6 h: 78.5 nm, 24h: 76.2 nm, 48 h: 76.1 nm | Stenotrophomona smaltophilia SeITE02 | [62] | |
| NA | Rods | diameter: 22 nm, length: 185 nm | Pseudomonas sp | [53] |
| NPs | Cellular | Shape | Size | Type of Bacteria | Ref. |
|---|---|---|---|---|---|
| Fe3O4 | Intra- | Cuboidal, rectangular and spherical NPs | 10~60 nm | Magnetospirillum magneticum | [67] |
| SnO2 | Extra- | Spherical and tetragonal NPs | 15~40 nm | Erwinia herbicola | [70] |
| Fe2O3 | NA | Spherical NPs | 30.2 nm | Bacillus cereus SVK1 | [68] |
| CuO | NA | Orthorhombic NPs | <10 nm | Morganelamorganii bacteria | [71] |
| CdS | Extra- | Cubic NPs | 2~10 nm | Bacillus licheniformis | [72] |
| Intra-& Extra- | Spherical NPs | 40~50 nm | Desulforibrio caledoiensis | [75] | |
| NA | Spherical NPs | 10 nm | Escherichia coli | [73] | |
| PbS | Extra- | Cubic NPs | 50 × 50 × 100 nm | Clostridiaceae sp | [77] |
| PbS2 | Intra- | Spherical NPs | 6 nm | Idiomarina sp. Strain PR58-8 | [78] |
| CuS | Extra- | Spherical NPs | 5 nm | Shewanella oneidensis MR-1 | [80] |
| ZnS | Extra- | Spherical NPs | 5 nm | Shewanella oneidensis MR-1 | [81] |
| CdSe | Intra- | NA | 3.3 ± 0.6 nm | Shewanella oneidensis MR-1 | [52] |
| CdTe | Extra- | NA | 62 nm | Escherichia coli | [76] |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Fang, X.; Wang, Y.; Wang, Z.; Jiang, Z.; Dong, M. Microorganism Assisted Synthesized Nanoparticles for Catalytic Applications. Energies 2019, 12, 190. https://doi.org/10.3390/en12010190
Fang X, Wang Y, Wang Z, Jiang Z, Dong M. Microorganism Assisted Synthesized Nanoparticles for Catalytic Applications. Energies. 2019; 12(1):190. https://doi.org/10.3390/en12010190
Chicago/Turabian StyleFang, Xiaojiao, Yin Wang, Zegao Wang, Zaixing Jiang, and Mingdong Dong. 2019. "Microorganism Assisted Synthesized Nanoparticles for Catalytic Applications" Energies 12, no. 1: 190. https://doi.org/10.3390/en12010190
APA StyleFang, X., Wang, Y., Wang, Z., Jiang, Z., & Dong, M. (2019). Microorganism Assisted Synthesized Nanoparticles for Catalytic Applications. Energies, 12(1), 190. https://doi.org/10.3390/en12010190






