Variation in the Distribution of Hydrogen Producers from the Clostridiales Order in Biogas Reactors Depending on Different Input Substrates
Abstract
1. Introduction
2. Materials and Methods
3. Results
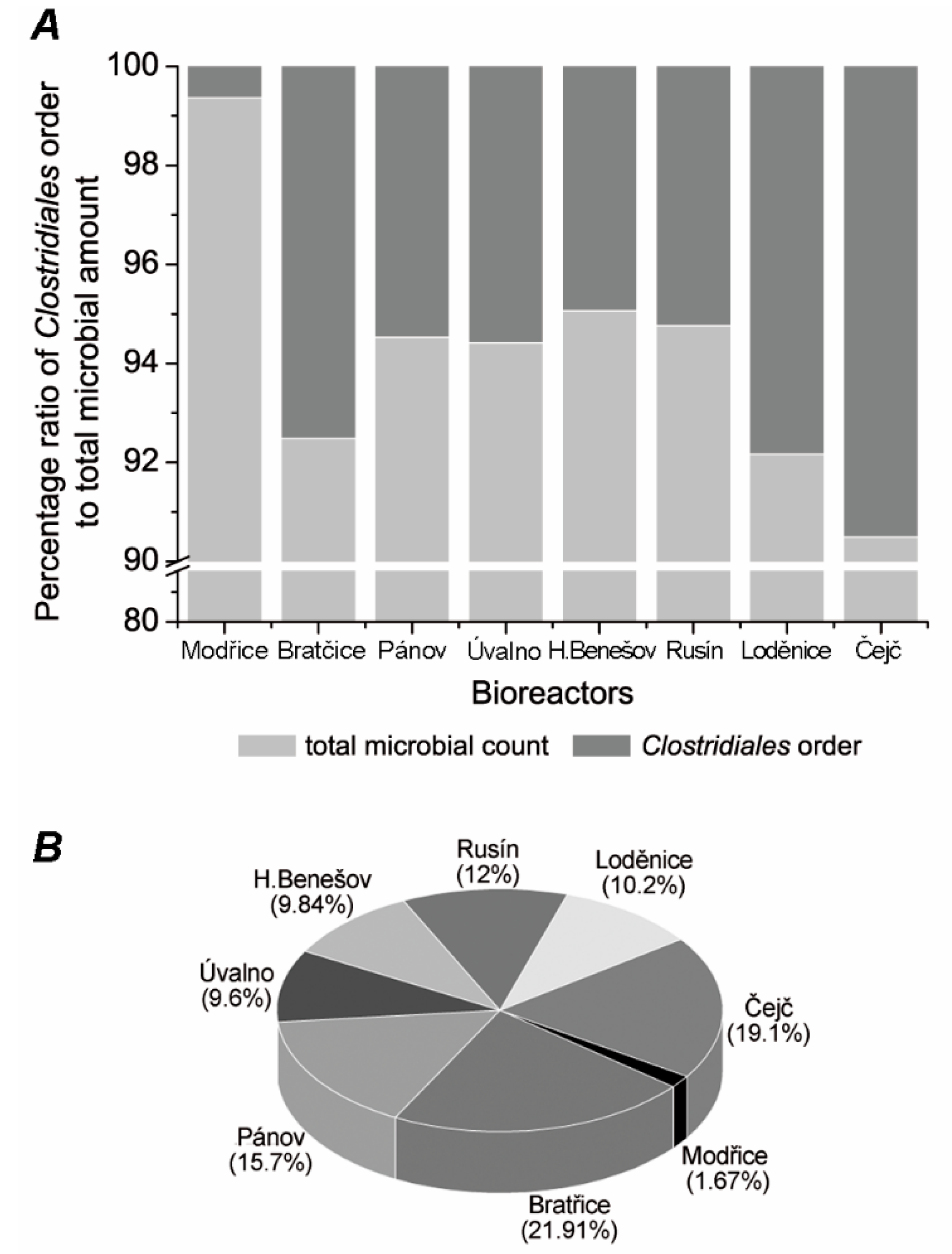
3.1. Commenting on the Control
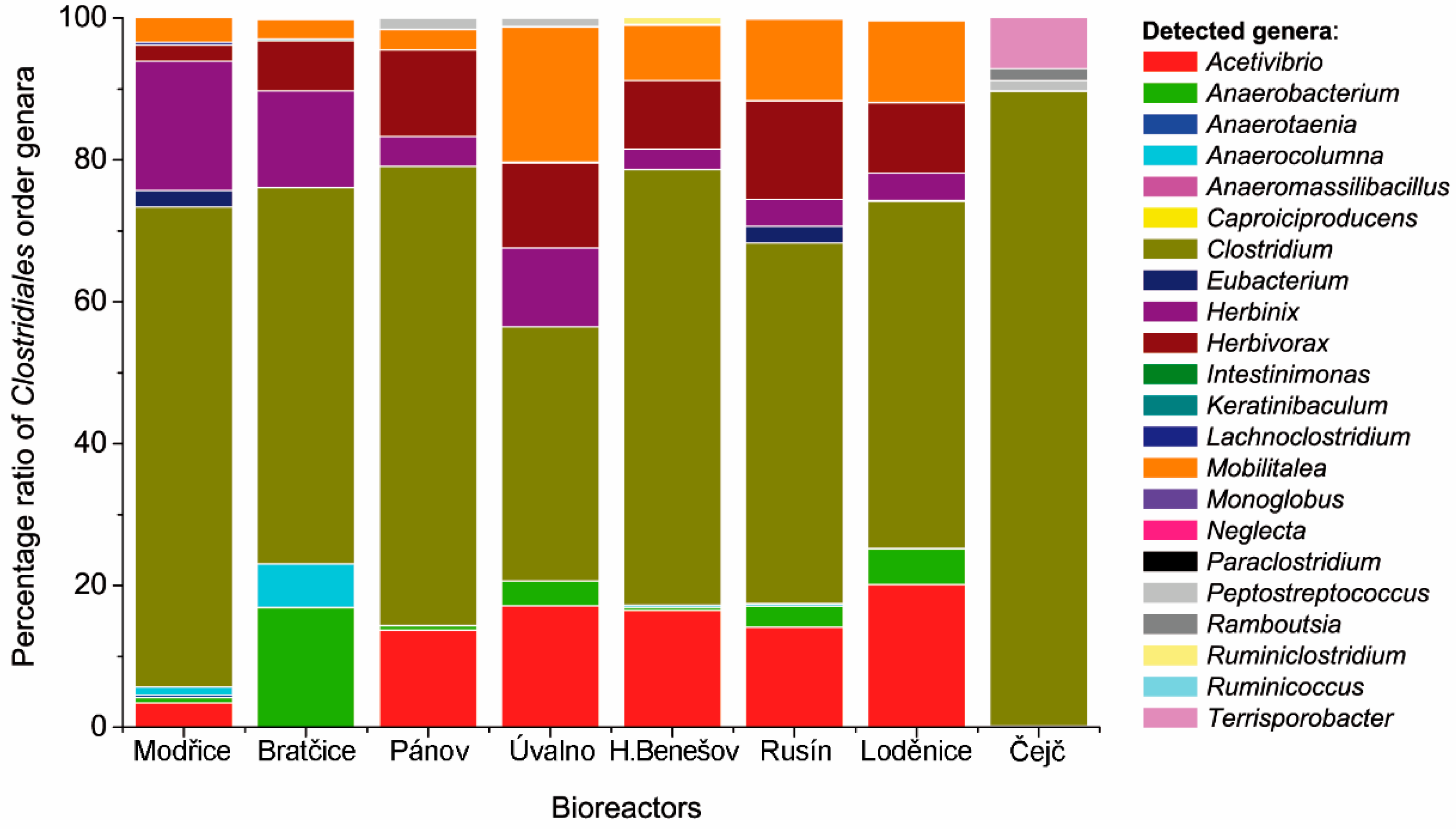
3.2. Comparison of the Clostridiales Order Communities
4. Discussion
Author Contributions
Funding
Conflicts of Interest
References
- Hobson, P.N. Biogas production from agricultural wastes. Experientia 1982, 38, 206–209. [Google Scholar] [CrossRef]
- Papurello, D.; Silvestri, S.; Tomasi, L.; Belcari, I.; Biasioli, F.; Santarelli, M. Biowaste for SOFCs. Energy Procedia 2016, 101, 424–431. [Google Scholar] [CrossRef]
- Oppermann, R.A.; Nelson, W.O.; Brown, R.E. In vivo studies of methanogenesis in the bovine rumen: Dissimilation of acetate. J. Gen. Microbiol. 1961, 25, 103–111. [Google Scholar] [CrossRef] [PubMed]
- Miller, T.L.; Wolin, M.J. Methanosphaera stadtmaniae gen. Nov., sp. nov.: A species that forms methane by reducing methanol with hydrogen. Arch. Microbiol. 1985, 141, 116–122. [Google Scholar] [CrossRef] [PubMed]
- Papurello, D.; Soukoulis, C.; Schuhfried, E.; Cappellin, L.; Gasperi, F.; Silvestri, S.; Santarelli, M.; Biasioli, F. Monitoring of volatile compound emissions during dry anaerobic digestion of the Organic Fraction of Municipal Solid Waste by Proton Transfer Reaction Time-of-Flight Mass Spectrometry. Bioresour. Technol. 2012, 126, 254–265. [Google Scholar] [CrossRef] [PubMed]
- Dridi, B.; Fardeau, M.-L.; Ollivier, B.; Raoult, D.; Drancourt, M. Methanomassiliicoccus luminyensis gen. nov., sp. nov., a methanogenic archaeon isolated from human faeces. Int. J. Syst. Evol. Microbiol. 2012, 62, 1902–1907. [Google Scholar] [CrossRef] [PubMed]
- Kushkevych, I.; Vítězová, M.; Vítěz, T.; Kováč, J.; Kaucká, P.; Jesionek, W.; Bartoš, M.; Barton, L. A new combination of substrates: Biogas production and diversity of the methanogenic microorganisms. Open Life Sci. 2018, 13, 119–128. [Google Scholar] [CrossRef]
- Kushkevych, I.; Vítězová, M.; Vítěz, T.; Bartoš, M. Production of biogas: Relationship between methanogenic and sulfate-reducing microorganisms. Open Life Sci. 2017, 12, 82–91. [Google Scholar] [CrossRef]
- Kushkevych, I.; Kováč, J.; Vítězová, M.; Vítěz, T.; Bartoš, M. The diversity of sulfate-reducing bacteria in the seven bioreactors. Arch. Microbiol. 2018. [Google Scholar] [CrossRef] [PubMed]
- Nath, K.; Das, D. Improvement of fermentative hydrogen production: Various approaches. Appl. Microbiol. Biotechnol. 2004, 65, 520–529. [Google Scholar] [CrossRef] [PubMed]
- Hawkes, F.R.; Dinsdale, R.; Hawkes, D.L.; Hussy, I. Sustainable fermentative hydrogen production: Challenges for process optimisation. Int. J. Hydrogen Energy. 2002, 27, 1339–1347. [Google Scholar] [CrossRef]
- Levin, D.B.; Pitt, L.; Love, M. Biohydrogen production: Prospects and limitations to practical application. Int. J. Hydrogen Energy. 2004, 29, 173–185. [Google Scholar] [CrossRef]
- Nanqi, R.; Wanqian, G.; Bingfeng, L.; Guangli, C.; Jie, D. Biological hydrogen production by dark fermentation: Challenges and prospects towards scaled-up production. Curr. Opin. Biotechnol. 2011, 22, 365–370. [Google Scholar] [CrossRef]
- Baek, G.; Kim, J.; Kim, J.; Lee, C. Role and potential of direct interspecies electron transfer in anaerobic digestion. Energies 2018, 11, 107. [Google Scholar] [CrossRef]
- Ozturk, S.S.; Palsson, B.O.; Thiele, J.H. Control of interspecies electron transfer flow during anaerobic digestion: Dynamic diffusion reaction models for hydrogen gas transfer in microbial flocs. Biotechnol. Bioeng. 1989, 33, 745–757. [Google Scholar] [CrossRef] [PubMed]
- Cord-Ruwisch, R.; Seitz, H.-J.; Conrad, R. The capacity of hydrogenotrophic anaerobic bacteria to compete for traces of hydrogen depends on the redox potential of the terminal electron acceptor. Arch. Microbiol. 1988, 149, 350–357. [Google Scholar] [CrossRef]
- Kong, X.; Yu, S.; Fang, W.; Liu, J.; Li, H. Enhancing syntrophic associations among Clostridium butyricum, Syntrophomonas and two types of methanogen by zero valent iron in an anaerobic assay with high organic loading. Bioresour. Technol. 2018, 257, 181–191. [Google Scholar] [CrossRef] [PubMed]
- Fang, H.H.P.; Zhang, T.; Liu, H. Microbial diversity of a mesophilic hydrogen-producing sludge. Appl. Microbiol. Biotechnol. 2002, 58, 112–118. [Google Scholar] [CrossRef] [PubMed]
- CSN EN 12176, Characterization of Sludge—Determination of pH-Value; Czech Standards Institute: Prague, Czech Republic, 1999.
- CSN EN 14346, Characterization of Waste—Calculation of Dry Matter by Determination of Dry Residue or Water Content; Czech Standards Institute: Prague, Czech Republic, 2007.
- CSN EN 15169, Characterization of Waste—Determination of Loss on Ignition in Waste, Sludge and Sediments; Czech Standards Institute: Prague, Czech Republic, 2007.
- Nossa, C.W.; Oberdorf, W.E.; Yang, L.; Aas, J.A.; Paster, B.J.; DeSantis, T.Z.; Brodie, E.L.; Malamud, D.; Poles, M.A.; Pei, Z. Design of 16S rRNA gene primers for 454 pyrosequencing of the human foregut microbiome. World. J. Gastroenterol. 2010, 16, 4135–4144. [Google Scholar] [CrossRef] [PubMed]
- Caporaso, J.G.; Kuczynski, J.; Stombaugh, J.; Bittinger, K.; Bushman, F.D.; Costello, E.K.; Fierer, N.; Peña, A.G.; Goodrich, J.K.; Gordon, J.I.; et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 2010, 7, 335–336. [Google Scholar] [CrossRef] [PubMed]
- Altschul, S.F.; Gish, W.; Mille, W.; Myers, E.W.; Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 1990, 215, 403–410. [Google Scholar] [CrossRef]
- Kearse, M.; Moir, R.; Wilson, A.; Stones-Havas, S.; Cheung, M.; Sturrock, S. Geneious Basic: An integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 2012, 28, 1647–1649. [Google Scholar] [CrossRef] [PubMed]
- Larkin, M.A.; Blackshields, G.; Brown, N.P.; Chenna, R.; McGettigan, P.A.; McWilliam, H. Clustal W and Clustal X version 2.0. Bioinformatics 2007, 23, 2947–2948. [Google Scholar] [CrossRef] [PubMed]
- Venkiteshwaran, K.; Bocher, B.; Maki, J.; Zitomer, D. Relating anaerobic digestion microbial community and process function. Microbiol. Insights 2015, 8, 37–44. [Google Scholar] [CrossRef] [PubMed]
- Moestedt, J.; Nilsson Påledal, S.; Schnürer, A. The effect of substrate and operational parameters on the abundance of sulphate-reducing bacteria in industrial anaerobic biogas digesters. Bioresour. Technol. 2013, 132, 327–332. [Google Scholar] [CrossRef] [PubMed]
- Shchegolkova, N.M.; Krasnov, G.S.; Belova, A.A.; Dmitriev, A.A.; Kharitonov, S.L.; Klimina, K.M.; Melnikova, N.V.; Kudryavtseva, A.V. Microbial community structure of activated sludge in treatment plants with different wastewater compositions. Front. Microbiol. 2016, 7, 90. [Google Scholar] [CrossRef] [PubMed]
- Kushkevych, I.V. Activity and kinetic properties of phosphotransacetylase from intestinal sulfate-reducing bacteria. Acta Biochim. Polonica 2015, 62, 103–108. [Google Scholar] [CrossRef]
- Kushkevych, I.; Kollar, P.; Suchy, P.; Parak, K.; Pauk, K.; Imramovsky, A. Activity of selected salicylamides against intestinal sulfate-reducing bacteria. Neuroendocrinol. Lett. 2015, 36, 106–113. [Google Scholar] [PubMed]
- Kushkevych, I.; Vítězová, M.; Kos, J.; Kollár, P.; Jampílek, J. Effect of selected 8-hydroxyquinoline-2-carboxanilides on viability and sulfate metabolism of Desulfovibrio piger. J. Appl. Biomed. 2018, 16, 241–246. [Google Scholar] [CrossRef]
- Rainey, F.A. Clostridiales. In Bergey’s Manual of Systematics of Archaea and Bacteria; John Wiley & Sons, Inc.: Hoboken, NJ, USA, 2015; pp. 1–5. [Google Scholar] [CrossRef]
- Kim, M.D.; Song, M.; Jo, M.; Shin, S.G.; Khim, J.H.; Hwang, S. Growth condition and bacterial community for maximum hydrolysis of suspended organic materials in anaerobic digestion of food waste-recycling wastewater. Appl. Microbiol. Biotechnol. 2010, 85, 1611–1618. [Google Scholar] [CrossRef] [PubMed]
- Fröschle, B.; Messelhäusser, U.; Höller, C.; Lebuhn, M. Fate of Clostridium botulinum and incidence of pathogenic clostridia in biogas processes. J. Appl. Microbiol. 2015, 119, 936–947. [Google Scholar] [CrossRef] [PubMed]
- Patel, G.B.; Khan, A.W.; Agnew, B.J.; Colvin, J.R. Isolation and characterization of an anaerobic, cellulolytic microorganism, Acetivibrio cellulolyticus gen. nov., sp. nov. Int. J. Syst. Bacteriol. 1980, 30, 179–185. [Google Scholar] [CrossRef]
- Koeck, D.E.; Hahnke, S.; Zverlov, V.V. Herbinix luporum sp. nov., a thermophilic cellulose-degrading bacterium isolated from thermophilic biogas reactor. Int. J. Syst. Evol. Microbiol. 2016, 66, 4132–4137. [Google Scholar] [CrossRef] [PubMed]
- Koeck, D.E.; Mechelke, M.; Zverlov, V.V.; Liebl, W.; Schwarz, W.H. Herbivorax saccincola gen. nov., sp. nov., a cellulolytic, anaerobic, thermophilic bacterium isolated via in sacco enrichments from a lab-scale biogas reactor. Int. J. Syst. Evol. Microbiol. 2016, 66, 4458–4463. [Google Scholar] [CrossRef] [PubMed]
- Podosokorskaya, O.A.; Bonch-Osmolovskaya, E.A.; Beskorovaynyy, A.V.; Toshchakov, S.V.; Kolganova, T.V.; Kublanov, I.V. Mobilitalea sibirica gen. nov., sp. nov., a halotolerant polysaccharide-degrading bacterium. Int. J. Syst. Evol. Microbiol. 2014, 64, 2657–2661. [Google Scholar] [CrossRef] [PubMed]
- Kushkevych, I.V. Kinetic Properties of Pyruvate Ferredoxin Oxidoreductase of Intestinal Sulfate-Reducing Bacteria Desulfovibrio piger Vib-7 and Desulfomicrobium sp. Rod-9. Polish J. Microbiol. 2015, 64, 107–114. [Google Scholar]
- Kushkevych, I.; Fafula, R.; Parak, T.; Bartos, M. Activity of Na+/K+-activated Mg2+-dependent ATP hydrolase in the cell-free extracts of the sulfate-reducing bacteria Desulfovibrio piger Vib-7 and Desulfomicrobium sp. Rod-9. Acta Vet Brno 2015, 84, 3–12. [Google Scholar] [CrossRef]
- Kushkevych, I.; Vítězová, M.; Fedrová, M.; Vochyanová, Z.; Paráková, L.; Hošek, J. Kinetic properties of growth of intestinal sulphate-reducing bacteria isolated from healthy mice and mice with ulcerative colitis. Acta Vet Brno 2017, 86, 405–411. [Google Scholar] [CrossRef]

| Geographic Location | Main Substrate | Input Ratio | Temperature (°C) | pH | ORP (mV) |
|---|---|---|---|---|---|
| Modřice | primary sludge, biological sludge | 50:50 | 34 | 7 | −3.1 |
| Bratčice | maize silage, whole crop silage, poultry litter | 63:31:6 | 43 | 8.3 | −75 |
| Pánov | maize silage, poultry litter | 92:8 | 49 | 8 | −58 |
| Úvalno | maize silage, sugar beet pulp, whole crop silage, cattle manure | 44:44:6:6 | 48 | 7.69 | −38.5 |
| Horní Benešov | maize silage, sugar beet pulp, whole crop silage, cattle manure, grass silage | 29:39:12:15:5 | 49 | 7.85 | −47.4 |
| Rusín | maize silage, sugar beet pulp | 70:30 | 48 | 7.63 | −34.7 |
| Loděnice | maize silage, sugar beet pulp | 75:25 | 44 | 7.65 | −36 |
| Čejč | pig manure, maize silage | 75:25 | – | – | – |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Černý, M.; Vítězová, M.; Vítěz, T.; Bartoš, M.; Kushkevych, I. Variation in the Distribution of Hydrogen Producers from the Clostridiales Order in Biogas Reactors Depending on Different Input Substrates. Energies 2018, 11, 3270. https://doi.org/10.3390/en11123270
Černý M, Vítězová M, Vítěz T, Bartoš M, Kushkevych I. Variation in the Distribution of Hydrogen Producers from the Clostridiales Order in Biogas Reactors Depending on Different Input Substrates. Energies. 2018; 11(12):3270. https://doi.org/10.3390/en11123270
Chicago/Turabian StyleČerný, Martin, Monika Vítězová, Tomáš Vítěz, Milan Bartoš, and Ivan Kushkevych. 2018. "Variation in the Distribution of Hydrogen Producers from the Clostridiales Order in Biogas Reactors Depending on Different Input Substrates" Energies 11, no. 12: 3270. https://doi.org/10.3390/en11123270
APA StyleČerný, M., Vítězová, M., Vítěz, T., Bartoš, M., & Kushkevych, I. (2018). Variation in the Distribution of Hydrogen Producers from the Clostridiales Order in Biogas Reactors Depending on Different Input Substrates. Energies, 11(12), 3270. https://doi.org/10.3390/en11123270






