The Use of DArTseq Technology to Identify Markers Linked to Genes Responsible for Seed Germination and Seed Vigor in Maize
Abstract
1. Introduction
2. Results
2.1. Analysis of the Seed Germination and Seed Vigor of Inbred Maize Lines
2.2. Association Mapping using GWAS Analysis
2.3. Physical Mapping of Gene Sequences
3. Discussion
4. Materials and Methods
4.1. Plant Material
4.2. Methods
4.2.1. Phenotyping
4.2.2. DNA Isolation
4.2.3. Genotyping
4.2.4. Statistical Analysis and Association Mapping using GWAS Analysis
4.2.5. Functional Analysis of Gene Sequences
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Conflicts of Interest
References
- Hilhorst, H.W.; Finch-Savage, W.E.; Buitink, J. Dormancy in Plant Seeds: Dormancy and Resistance in Harsh Environments; Springer: Berlin, Germany, 2010; pp. 43–67. [Google Scholar] [CrossRef]
- Delouche, J.C.; Baskin, C.C. Accelerated Aging Techniques for Predicting the Relative Storability of Seed Lots. Seed Sci. Technol. 2016, 1, 427–452. [Google Scholar]
- Gupta, M.L.; George, D.L.; Parwata, I.G. Effect of Harvest Time and Drying on Super Sweet Corn Seed Quality. Seed Sci. Technol. 2005, 33, 167–176. [Google Scholar] [CrossRef]
- Han, Z.; Ku, L.; Zhang, Z.; Zhang, J.; Guo, S.; Liu, H.; Zhao, R.; Ren, Z.; Zhang, L.; Su, H.; et al. QTLs for Seed Vigor-Related Traits Identified in Maize Seeds Germinated under Artificial Aging Conditions. PLoS ONE 2014, 9, e92535. [Google Scholar] [CrossRef]
- Rehman Arif, M.A.; Nagel, M.; Neumann, K.; Kobiljski, B.; Lohwasser, U.; Börner, A. Genetic Studies of Seed Longevity in Hexaploid Wheat using Segregation and Association Mapping Approaches. Euphytica 2012, 186, 1–13. [Google Scholar] [CrossRef]
- Xin, X.; Lin, X.H.; Zhou, Y.C.; Chen, X.L.; Liu, X.; Lu, X.X. Proteome analysis of maize seeds: The effect of artificial ageing. Physiol. Plant. 2011, 143, 126–138. [Google Scholar] [CrossRef]
- Nagel, M.; Vogel, H.; Landjeva, S.; Buck-Sorlin, G.; Lohwasser, U.; Scholz, U.; Boerner, A. Seed Conservation in ex Situ Genebanks Genetic Studies on Longevity in Barley. Euphytica 2009, 170, 5–14. [Google Scholar] [CrossRef]
- Zhu, C.; Gore, M.; Buckler, E.S.; Yu, J. Status and Prospects of Association Mapping in Plants. Plant Genome 2008, 1, 5–20. [Google Scholar] [CrossRef]
- Xue, Y.; Zhang, S.Q.; Yao, Q.H.; Peng, R.H.; Xiong, A.S.; Li, X.; Zhu, W.M.; Zhu, Y.Y.; Zha, D.S. Identification of Quantitative Trait Loci for Seed Storability in Rice (Oryza sativa L.). Euphytica 2008, 164, 739–744. [Google Scholar] [CrossRef]
- Xiao, Y.; Liu, H.; Wu, L.; Warburton, M.; Yan, J. Genome-wide Association Studies in Maize: Praise and Stargaze. Mol. Plant 2017, 10, 359–374. [Google Scholar] [CrossRef]
- Voss-Fels, K.; Snowdon, R.J. Understanding and Utilizing Crop Genome Diversity via High-Resolution Genotyping. Plant Biotechnol. J. 2016, 14, 1086–1094. [Google Scholar] [CrossRef]
- Sakhare, A.S.; Kota, S.; Rathod, S.; Parmar, B.; Chinnusamy, V. Genome-Wide Association Study. In Genotyping by Sequencing for Crop Improvement; Sonah, H., Goyal, V., Shivaraj, S., Deshmukh, R.K., Eds.; Wiley: Hoboken, NJ, USA, 2022. [Google Scholar] [CrossRef]
- Sanghamitra, P.; Nanda, N.; Barik, S.R.; Sahoo, S.; Pandit, E.; Bastia, R.; Bagchi, T.B.; Pradhan, S.K. Genetic structure and molecular markers-trait association for physiological traits related to seed vigour in rice. Plant Gene 2021, 28, 100338. [Google Scholar] [CrossRef]
- Sahoo, S.; Sanghamitra, P.; Nanda, N.; Pawar, S.; Pandit, E.; Bastia, R.; Muduli, K.C.H.; Pradhan, S.K. Association of molecular markers with physio-biochemical traits related to seed vigour in rice. Physiol. Mol. Biol. Plants 2020, 26, 1989–2003. [Google Scholar] [CrossRef]
- Mahender, A.; Anandan, A.; Pradhan, S.K. Early seedling vigour, an imperative trait for direct-seeded rice: An overview on physio-morphological parameters and molecular markers. Planta 2015, 241, 1027–1050. [Google Scholar] [CrossRef]
- Garza-Caligaris, L.E.; Avendaño-Vázquez, A.O.; Alvarado-López, S.; Zúñiga-Sánchez, E.; Orozco-Segovia, A.; Pérez-Ruíz, R.V.; Gamboa-de Buen, A. At3g08030 transcript: A molecular marker of seed ageing. Ann. Bot. 2012, 110, 1253–1260. [Google Scholar] [CrossRef]
- Froidure, S.; Roby, D.; Rivas, S. Expression of the Arabidopsis transcription factor AtMYB30 is post-transcriptionally regulated. Plant Physiol. Biochem. 2010, 48, 735–739. [Google Scholar] [CrossRef]
- Du, L.; Jiang, H.; Zhao, G.; Ren, J. Gene cloning of ZmMYB59 transcription factor in maize and its expression during seed germination in response to deep-sowing and exogenous hormones. Plant Breed. 2017, 136, 834–844. [Google Scholar] [CrossRef]
- Rodrigues, L.A.; Alves, C.Z.; Queiroz Rego, C.H.; Benetoli Da Silva, T.R.; Bispo Da Silva, J. Humic acid on germination and vigor of corn seeds. Agronomia Rev. Caatinga 2017, 30, 149–154. [Google Scholar] [CrossRef]
- Zing, J.; Ali, F.; Chen, G.; Li, H.; Mahuku, G.; Yang, N.; Narro, L.; Magorokosho, C.; Makumbi, D.; Yan, J. Genome-wide association mapping reveals novel sources of resistance to northern corn leaf blight in maize. BMC Plant Biol. 2015, 15, 206. [Google Scholar] [CrossRef]
- Tang, J.D.; Perkins, A.; Williams, W.P.; Warburton, M.L. Using genome-wide associations to identify metabolic pathways involved in maize aflatoxin accumulation resistance. BMC Genom. 2015, 16, 673. [Google Scholar] [CrossRef]
- Nannas, N.J.; Dawe, R.K. Genetic and genomic toolbox of Zea mays. Genetics 2015, 199, 655–669. [Google Scholar] [CrossRef]
- Huang, Y.F.; Poland, J.A.; Wight, C.P.; Jackson, E.W.; Tinker, N.A. Using Genotyping by Sequencing (GBS) for genomic discovery in cultivated oat. PLoS ONE 2014, 9, e102448. [Google Scholar] [CrossRef]
- Elshire, R.J.; Glaubitz, J.C.; Sun, Q.; Poland, J.A.; Kawamoto, K.; Buckler, E.S.; Mitchell, S.E.; Robust, A. Simple Genotyping-by-Sequencing (GBS) Approach for High Diversity Species. PLoS ONE 2011, 6, e19379. [Google Scholar] [CrossRef]
- Sansaloni, C.; Petroli, C.; Jaccoud, D.; Carling, J.; Detering, F.; Grattapaglia, D.; Kilian, A. Diversity Array Technology (DArT) and next-generation sequencing combined: Genome-wide, high throughput, highly informative genotyping for molecular breeding of Eucalyptus. BMC Proc. 2011, 5 (Suppl. 7), P54. [Google Scholar] [CrossRef]
- Kilian, B.; Graner, A. NGS technologies for analyzing germplasm diversity in genebanks. Briefinds Funct. Genom. 2012, 11, 38–50. [Google Scholar] [CrossRef]
- Tomkowiak, A.; Nowak, B.; Sobiech, A.; Bocianowski, J.; Wolko, Ł.; Spychała, J. The Use of DArTseq Technology to Identify New SNP and SilicoDArT Markers Related to the Yield-Related Traits Components in Maize. Genes 2022, 13, 848. [Google Scholar] [CrossRef]
- Sobiech, A.; Tomkowiak, A.; Nowak, B.; Bocianowski, J.; Wolko, Ł.; Spychała, J. Associative and Physical Mapping of Markers Related to Fusarium in Maize Resistance, Obtained by Next-Generation Sequencing (NGS). Int. J. Mol. Sci. 2022, 23, 6105. [Google Scholar] [CrossRef]
- Tomkowiak, A.; Bocianowski, J.; Spychała, J.; Grynia, J.; Sobiech, A.; Kowalczewski, P.Ł. DArTseq-Based High-Throughput SilicoDArT and SNP Markers Applied for Association Mapping of Genes Related to Maize Morphology. Int. J. Mol. Sci. 2021, 22, 5840. [Google Scholar] [CrossRef]
- Han, Z.P.; Bin, W.; Zhang, J.; Guo, S.L.; Zhang, H.C.; Xu, L.R.; Chen, Y.H. Mapping of QTLs Associated with Seed Vigor to Artificial Aging Using Two RIL Populations in Maize (Zea mays L.). Agric. Sci. 2018, 9, 397–415. [Google Scholar] [CrossRef]
- Zhang, H.W.; Ma, P.; Zhao, Z.N.; Zhao, G.W.; Tian, B.H.; Wang, J.H.; Wang, G.Y. Mapping QTL controlling maize deep-sowing tolerance-related traits and confirmation of a major QTL for mesocotyl length. Theor. Appl. Genet. 2012, 124, 223–232. [Google Scholar] [CrossRef]
- Zhao, X.; Niu, Y. The Combination of Conventional QTL Analysis, Bulked-Segregant Analysis, and RNA-Sequencing Provide New Genetic Insights into Maize Mesocotyl Elongation under Multiple Deep-Seeding Environments. Int. J. Mol. Sci. 2022, 23, 4223. [Google Scholar] [CrossRef]
- Li, S. Redox Modulation Matters: Emerging Functions for Glutaredoxins in Plant Development and Stress Responses. Plants 2014, 3, 559–582. [Google Scholar] [CrossRef]
- Stein, O.; Granot, D. An Overview of Sucrose Synthases in Plants. Front. Plant Sci. 2019, 10, 95. [Google Scholar] [CrossRef]
- Zhu, X.; Wang, M.; Li, X.; Jiu, S.; Wang, C.; Fang, J. Genome-Wide Analysis of the Sucrose Synthase Gene Family in Grape (Vitis vinifera): Structure, Evolution, and Expression Profiles. Genes 2017, 8, 111. [Google Scholar] [CrossRef]
- Mao, Y.; Tan, S. Functions and Mechanisms of SAC Phosphoinositide Phos-phatases in Plants. Front Plant Sci. 2021, 12, 803635. [Google Scholar] [CrossRef]
- Gagne, J.M.; Clark, S.E. The protein phosphatases POL and PLL1 are signaling intermediates for multiple pathways in Arabidopsis. Plant Signal. Behav. 2007, 2, 245–246. [Google Scholar] [CrossRef]
- Gagne, J.M.; Song, S.K.; Clark, S.E. POLTERGEIST and PLL1 are required for stem cell function with potential roles in cell asymmetry and auxin signaling. Commun. Integr. Biol. 2008, 1, 53–55. [Google Scholar] [CrossRef] [PubMed]
- Song, S.K.; Hofhuis, H.; Lee, M.M.; Clark, S.E. Key divisions in the early Arabidopsis embryo require POL and PLL1 phosphatases to establish the root stem cell organizer and vascular axis. Dev. Cell 2008, 15, 98–109. [Google Scholar] [CrossRef]
- Song, S.K.; Lee, M.M.; Clark, S.E. POL and PLL1 phosphatases are CLAVATA1 signaling intermediates required for Arabidopsis shoot and floral stem cells. Development 2006, 133, 4691–4698. [Google Scholar] [CrossRef]
- Thorstensen, T.; Grini, P.E.; Aalen, R.B. SET domain proteins in plant development. Biochim. Biophys. Acta Gene Regul. Mech. 2011, 1809, 407–420. [Google Scholar] [CrossRef]
- Ng, D.W.; Wang, T.; Chandrasekharan, M.B.; Aramayo, R.; Kertbundit, S.; Hall, T.C. Plant SET domain-containing proteins: Structure, function and regulation. Biochim. Biophys. Acta 2007, 1769, 316–329. [Google Scholar] [CrossRef]
- Li, T.; Zhang, Y.; Wang, D.; Liu, Y.; Dirk, L.M.A.; Goodman, J.; Downie, B.; Wang, J.; Wang, G.; Zhao, T. Regulation of seed vigor by manipulation of raffinose family oligosaccharides in maize and Arabidopsis thaliana. Mol. Plant 2017, 10, 1540–1555. [Google Scholar] [CrossRef] [PubMed]
- Mondal, S.; Singh, S.P. New insights on thioredoxins (Trxs) and glutaredoxins (Grxs) by in silico amino acid sequence, phylogenetic and comparative structural analyses in organisms of three domains of life. Heliyon 2022, 8, e10776. [Google Scholar] [CrossRef] [PubMed]
- Coppola, M.; Corrado, G.; Coppola, V.; Cascone, P.; Martinelli, R.; Digilio, M.C.; Pennacchio, F.; Rao, R. Prosystemin Overexpression in Tomato Enhances Resistance to Different Biotic Stresses by Activating Genes of Multiple Signaling Pathways. Plant Mol. Biol. Rep. 2015, 33, 1270–1285. [Google Scholar] [CrossRef] [PubMed]
- Bocianowski, J.; Liersch, A. Multidimensional Analysis of Diversity in Genotypes of Winter Oilseed Rape (Brassica napus L.). Agronomy 2022, 12, 633. [Google Scholar] [CrossRef]
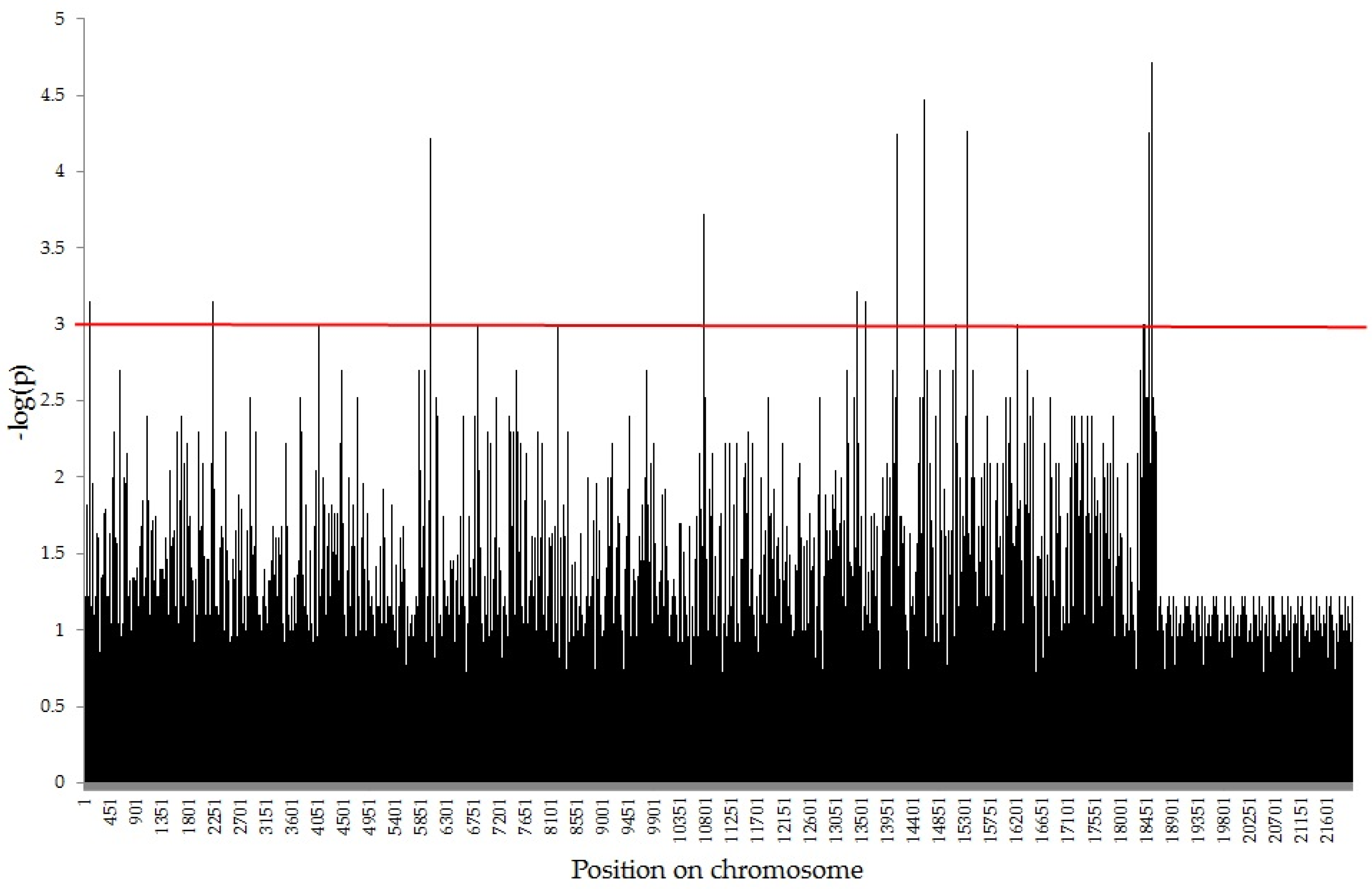
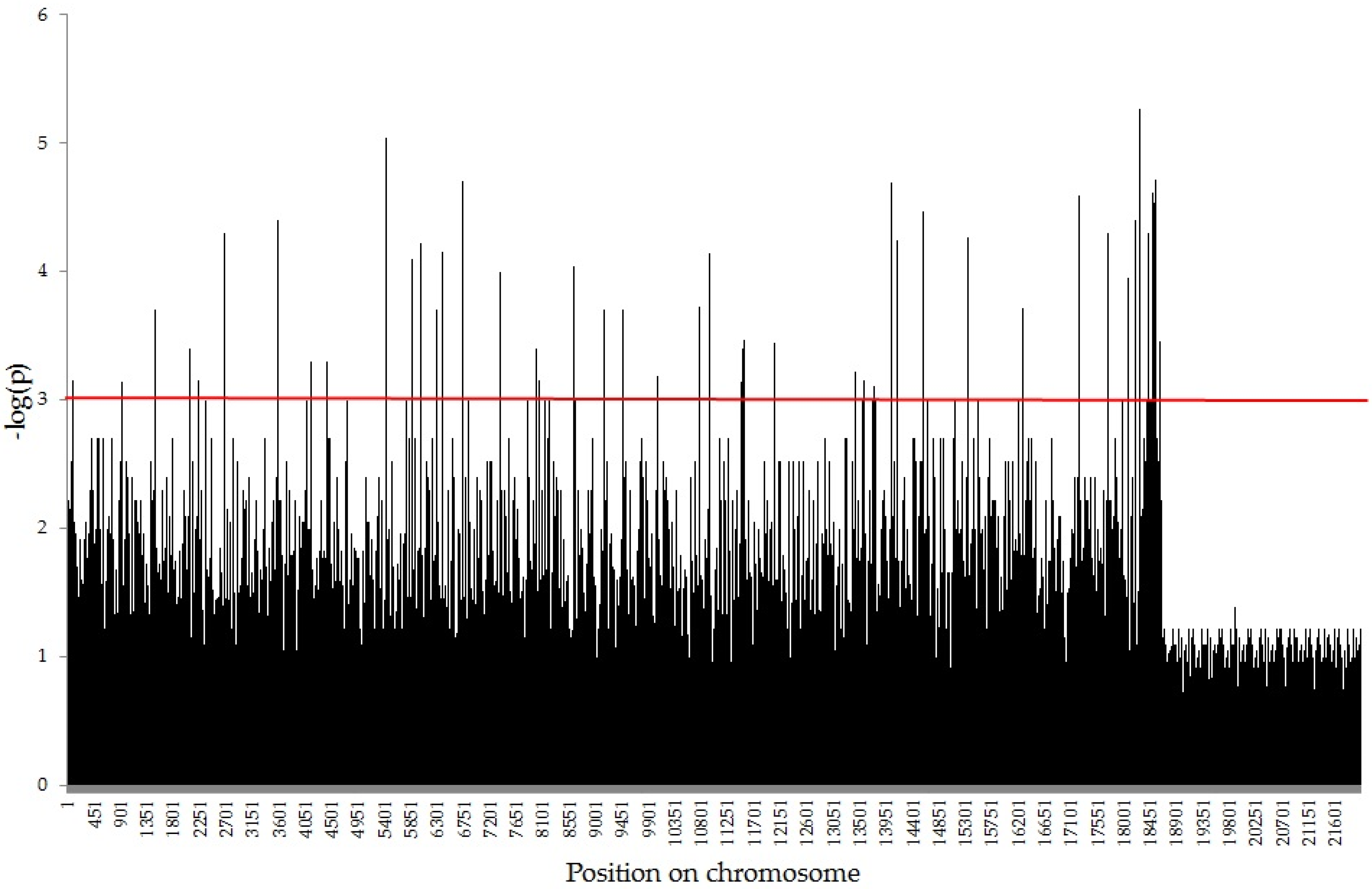
| Trait | Type of Markers | SilicoDArT | SNP | |
|---|---|---|---|---|
| Seed vigor | The number of significant markers | 758 | 132 | |
| Effect | min | −0.801 | −0.795 | |
| max | 0.78 | 0.714 | ||
| mean | 0.0015 | −0.036 | ||
| sum | 1.106 | −4.794 | ||
| Percentage variance accounted | min | 2 | 2 | |
| max | 7.6 | 6.6 | ||
| mean | 3.01 | 3.06 | ||
| Seed germination | The number of significant markers | 1115 | 208 | |
| Effect | min | −0.639 | −0.626 | |
| max | 0.621 | 0.637 | ||
| mean | −0.088 | −0.046 | ||
| sum | −98.092 | −9.591 | ||
| Percentage variance accounted | min | 1.9 | 1.9 | |
| max | 10.2 | 10 | ||
| mean | 3.14 | 3.3 | ||
| Type of Marker | Marker | Seed Vigor | Seed Germination | ||
|---|---|---|---|---|---|
| Estimate | Percentage Variance Accounted | Estimate | Percentage Variance Accounted | ||
| SNP | 2,383,357 | −0.423 | 2.0 | −0.37 | 3.0 |
| DArT | 2,384,346 | 0.431 | 2.1 | 0.332 | 2.3 |
| DArT | 2,394,856 | −0.479 | 2.4 | −0.337 | 2.1 |
| DArT | 2,427,730 | −0.477 | 2.7 | −0.316 | 2.0 |
| DArT | 2,435,414 | 0.485 | 2.3 | 0.379 | 2.6 |
| DArT | 2,435,784 | −0.677 | 5.5 | −0.373 | 2.7 |
| DArT | 2,459,805 | −0.478 | 2.3 | −0.341 | 2.0 |
| DArT | 2,480,018 | 0.459 | 2.2 | 0.331 | 2.0 |
| DArT | 2,496,196 | −0.466 | 2.6 | −0.371 | 3.0 |
| DArT | 2,500,039 | −0.462 | 2.5 | −0.42 | 4.0 |
| DArT | 2,512,711 | 0.542 | 2.8 | 0.357 | 2.0 |
| DArT | 2,517,355 | 0.525 | 2.9 | 0.402 | 3.1 |
| DArT | 2,523,102 | −0.592 | 4.3 | −0.36 | 2.6 |
| DArT | 2,524,206 | 0.618 | 4.4 | 0.373 | 2.7 |
| DArT | 2,526,601 | −0.536 | 3.4 | −0.388 | 3.2 |
| DArT | 2,558,379 | −0.471 | 2.6 | −0.355 | 2.7 |
| SNP | 2,565,188 | 0.487 | 2.2 | 0.378 | 2.5 |
| DArT | 4,576,613 | 0.663 | 5.1 | 0.357 | 2.3 |
| DArT | 4,578,890 | 0.695 | 4.8 | 0.386 | 2.4 |
| DArT | 4,584,107 | −0.533 | 3.6 | −0.373 | 3.1 |
| DArT | 4,584,433 | −0.483 | 2.7 | −0.318 | 1.9 |
| SNP | 4,584,461 | 0.566 | 3.8 | 0.398 | 3.3 |
| DArT | 4,589,607 | −0.737 | 6.6 | −0.343 | 2.2 |
| DArT | 4,592,681 | −0.449 | 2.2 | −0.336 | 2.3 |
| DArT | 4,765,935 | 0.441 | 2.1 | −0.54 | 6.7 |
| DArT | 4,768,602 | −0.44 | 2.0 | −0.325 | 1.9 |
| DArT | 4,772,587 | 0.509 | 2.9 | 0.455 | 4.5 |
| SNP | 4,772,814 | −0.427 | 2.0 | −0.318 | 1.9 |
| DArT | 4,772,944 | −0.46 | 2.3 | −0.371 | 2.8 |
| SNP | 4,773,324 | −0.607 | 4.8 | −0.362 | 2.8 |
| SNP | 4,773,863 | 0.663 | 5.1 | 0.357 | 2.3 |
| DArT | 4,774,632 | −0.518 | 2.7 | −0.342 | 2.0 |
| DArT | 4,775,276 | −0.455 | 2.4 | −0.353 | 2.7 |
| DArT | 4,776,334 | −0.651 | 5.2 | −0.348 | 2.4 |
| DArT | 4,777,053 | 0.562 | 4.0 | 0.326 | 2.2 |
| SNP | 4,777,566 | 0.441 | 2.2 | 0.323 | 2.1 |
| DArT | 5585,640 | −0.587 | 3.7 | −0.34 | 2.0 |
| DArT | 7,049,301 | −0.46 | 2.3 | −0.343 | 2.3 |
| DArT | 7,050,274 | −0.621 | 3.4 | −0.4 | 2.4 |
| DArT | 7,054,752 | −0.524 | 3.4 | −0.329 | 2.2 |
| DArT | 7,059,320 | 0.748 | 6.6 | 0.395 | 3.0 |
| DArT | 7,061,686 | 0.493 | 2.9 | 0.32 | 2.0 |
| DArT | 9,625,011 | −0.517 | 3.0 | −0.343 | 2.3 |
| SNP | 9,687,038 | −0.532 | 2.8 | −0.364 | 2.3 |
| DArT | 9,693,261 | −0.48 | 2.6 | −0.36 | 2.6 |
| DArT | 9,710,915 | 0.596 | 4.5 | 0.332 | 2.2 |
| DArT | 16,723,050 | 0.498 | 2.8 | 0.369 | 2.7 |
| DArT | 16,725,652 | 0.508 | 2.6 | 0.419 | 3.3 |
| DArT | 16,726,826 | −0.431 | 2.1 | −0.413 | 3.9 |
| DArT | 21,695,758 | 0.459 | 2.2 | 0.331 | 2.0 |
| SNP | 21,698,325 | −0.497 | 2.0 | −0.383 | 2.1 |
| DArT | 25,000,251 | −0.626 | 5.2 | −0.371 | 3.0 |
| DArT | 25,943,049 | 0.596 | 3.7 | 0.428 | 3.4 |
| DArT | 25,947,240 | 0.548 | 3.1 | −0.342 | 2.0 |
| DArT | 26,083,456 | 0.485 | 2.3 | 0.379 | 2.6 |
| DArT | 58,293,040 | −0.562 | 4.0 | −0.379 | 3.2 |
| Marker | Estimate | Percentage Variance Accounted | Marker | Estimate | Percentage Variance Accounted |
|---|---|---|---|---|---|
| 4,580,898 | −0.745 | 7.6 | 25,000,779 | −0.762 | 6.6 |
| 2,507,310 | 0.744 | 7.3 | 9,624,535 | −0.701 | 6.5 |
| 9,699,056 | −0.719 | 7.0 | 9,687,233 | 0.692 | 6.4 |
| 25,981,291 | −0.751 | 7.0 | 4,773,254 | −0.788 | 6.4 |
| 25,947,631 | −0.801 | 6.9 | 2,530,239 | 0.701 | 6.3 |
| 4,589,607 | −0.737 | 6.6 | 4,582,430 | −0.736 | 6.2 |
| 4,769,346 | 0.780 | 6.6 | 4,770,911 | −0.736 | 6.2 |
| 4,779,691 | −0.780 | 6.6 | 4,591,564 | −0.676 | 6.1 |
| 7,059,320 | 0.748 | 6.6 |
| Marker | Estimate | Percentage Variance Accounted | Marker | Estimate | Percentage Variance Accounted |
|---|---|---|---|---|---|
| 2,382,757 | −0.639 | 10.2 | 2,572,299 | −0.595 | 6.6 |
| 2,386,217 | 0.637 | 10.0 | 4,576,534 | −0.583 | 6.6 |
| 4,770,719 | −0.626 | 9.7 | 9,682,555 | −0.521 | 6.6 |
| 7,059,241 | 0.621 | 9.4 | 24,017,322 | −0.578 | 6.6 |
| 2,565,888 | 0.620 | 9.1 | 2,452,719 | 0.557 | 6.5 |
| 4,579,017 | 0.610 | 9.1 | 4,775,059 | 0.543 | 6.5 |
| 4,778,926 | 0.613 | 8.9 | 5,589,519 | −0.569 | 6.5 |
| 4,768,869 | 0.599 | 8.8 | 9,679,345 | −0.520 | 6.5 |
| 9,619,890 | −0.581 | 8.4 | 16,723,671 | −0.552 | 6.5 |
| 2,516,290 | −0.599 | 8.3 | 2,387,415 | 0.516 | 6.4 |
| 4,765,379 | −0.611 | 8.3 | 2,438,696 | −0.535 | 6.4 |
| 4,779,164 | 0.592 | 8.2 | 4,592,741 | 0.548 | 6.4 |
| 4,582,993 | 0.585 | 8.1 | 2,542,705 | −0.510 | 6.3 |
| 4,767,822 | −0.577 | 8.1 | 4,584,706 | −0.513 | 6.3 |
| 4,773,702 | 0.594 | 8.0 | 4,770,215 | −0.514 | 6.3 |
| 48,474,231 | −0.579 | 7.9 | 9,694,232 | −0.523 | 6.3 |
| 4,775,452 | −0.572 | 7.8 | 2,439,578 | −0.528 | 6.2 |
| 4,775,909 | −0.016 | 7.7 | 2,441,184 | −0.504 | 6.2 |
| 4,587,134 | 0.571 | 7.5 | 2,509,715 | −0.528 | 6.2 |
| 4,774,360 | −0.570 | 7.5 | 4,576,581 | 0.517 | 6.2 |
| 5,584,378 | −0.560 | 7.5 | 5,587,268 | 0.570 | 6.2 |
| 4,591,069 | −0.572 | 7.4 | 29,622,219 | −0.504 | 6.2 |
| 34,662,940 | −0.548 | 7.4 | 2,432,805 | 0.521 | 6.1 |
| 4,777,912 | −0.539 | 6.8 | 2,451,544 | 0.546 | 6.1 |
| 2,530,805 | −0.542 | 6.7 | 2,610,978 | 0.546 | 6.1 |
| 4,765,935 | −0.540 | 6.7 | 4,588,360 | −0.504 | 6.1 |
| 5,589,694 | −0.608 | 6.7 | 4,775,503 | −0.546 | 6.1 |
| 2,552,487 | −0.544 | 6.6 | 25,000,666 | 0.521 | 6.1 |
| Marker | Marker Type | Feature (Significant Level) | Chromosome | Marker Location | Candidate Genes |
|---|---|---|---|---|---|
| 4,589,607 | DArT | Seed vigor (0.001), Seed germination (0.05) | Chr2 | 168,930,481 | 128 bp at 5′ side: uncharacterized protein loc100272672 84,601 bp at 3′ side: diacylglycerol lipase-beta |
| 7,059,320 | DArT | Seed vigor (0.001), Seed germination (0.05) | Chr2 | 168,930,481 | 128 bp at 5′ side: uncharacterized protein loc100272672 84,601 bp at 3′ side: diacylglycerol lipase-beta |
| 2,435,784 | DArT | Seed vigor (0.001), Seed germination (0.05) | Ch1 | 56,888,568 | sucrose synthase 4 isoform ×2 sucrose synthase 4 isoform ×1 |
| 4,776,334 | DArT | Seed vigor (0.001), Seed germination (0.05) | Chr2 | 129,929,222 | phosphoinositide phosphatase sac7 isoform ×1 phosphoinositide phosphatase sac7 isoform ×2 |
| 25,000,251 | DArT | Seed vigor (0.001), Seed germination (0.05) | Chr1 | 89,047,678 | 110,109 bp at 5′ side: ndr1/hin1-like protein 10 552 bp at 3′ side: translocase of chloroplast 120, chloroplastic |
| 4,765,935 | DArT | Seed germination (0.001), Seed vigor (0.05) | Chr8 | 22,460,192 | 5781 bp at 5′ side: uncharacterized protein loc100279368 1404 bp at 3′ side: uncharacterized protein loc100280975 |
| 4,772,587 | DArT | Seed germination (0.001), Seed vigor (0.05) | Chr4 | 141,172,070 | putative set-domain containing protein family isoform ×1 putative set-domain containing protein family |
| 2,500,039 | DArT | Seed germination (0.001), Seed vigor (0.05) | Chr2 | 190,086,478 | 12,544 bp at 5′ side: atp-dependent dna helicase pif1-like 2947 bp at 3′ side: uncharacterized protein loc100217048 |
| 16,726,826 | DArT | Seed germination (0.001), Seed vigor (0.05) | Chr2 | 190,140,760 | 821 bp at 5′ side: uncharacterized protein loc100283853 4387 bp at 3′ side: alpha carbonic anhydrase 4 |
| 25,943,049 | DArT | Seed germination (0.001), Seed vigor (0.05) | Ch2 | 62,908,172 | 86,280 bp at 5’ side: uncharacterized protein loc100273593 isoform ×1 1726 bp at 3′ side: inhibitor of apoptosis-like protein |
| 4,580,898 | DArT | Seed vigor (0.001) | Ch1 | 159,125,630 | 30,617 bp at 5′ side: uncharacterized protein loc103644989 isoform ×1 161,136 bp at 3′ side: uncharacterized protein loc100274710 isoform ×2 |
| 2,507,310 | DArT | Seed vigor (0.001) | Chr2 | 69,438,881 | grx_c8–glutaredoxin subgroup iii |
| 9,699,056 | DArT | Seed vigor (0.001) | Chr1 | 159,575,935 | 228,802 bp at 5′ side: scarecrow-like protein 34 154,620 bp at 3′ side: uncharacterized protein loc103640060 |
| 25,981,291 | DArT | Seed vigor (0.001) | Chr1 | 86,637,007 | a-agglutinin anchorage subunit-like |
| 25,947,631 | DArT | Seed vigor (0.001) | Chr8 | 143,760,900 | 5439 bp at 5′ side: protein quirky 78,137 bp at 3′ side: uncharacterized protein loc100284991 |
| 2,382,757 | DArT | Seed germination (0.001) | Chr3 | 32,582,513 | 672 bp at 5′ side: probable mitochondrial import receptor subunit tom20 21,113 bp at 3′ side: uncharacterized protein loc100276184 |
| 2,386,217 | SNP | Seed germination (0.001) | Chr3 | 32,204,938 | probable 3-beta-hydroxysteroid-delta(8),delta(7)-isomerase |
| 4,770,719 | SNP | Seed germination (0.001) | Chr3 | 32,126,239 | uncharacterized protein loc100272990 uncharacterized protein loc100272990 isoform ×1 |
| 7,059,241 | DArT | Seed germination (0.001) | Chr3 | 32,125,561 | uncharacterized protein loc100272990 uncharacterized protein loc100272990 isoform ×1 |
| 2,565,888 | DArT | Seed germination (0.001) | Chr3 | 33,897,068 | 11,124 bp at 5′ side: gdsl esterase/lipase at1g33811 6976 bp at 3′ side: uncharacterized protein loc100273387 |
| Marker | Primer Sequences | Annual Temperature (°C) | Product Size (bp) | |
|---|---|---|---|---|
| Forward | Reverse | |||
| 4,589,607 | ACGGGAGAGGAACGCTGCAG | GCCTAAACACAAGCAAGTGGGC | 63 | 70 |
| 7,059,320 | ACGGGAGAGGAACGCTGCAG | TCTGAAGAGCCATGGCAAAAGC | 62 | 483 |
| 4,776,334 | AACATTTACATCATCTGCAG | AATTGATCACAAATGTTATT | 56 | 161 |
| 25,000,251 | GAGAGTGCAGAGTGCAG | TGGGCATGCTACTGAGTTTT | 54 | 207 |
| 4,765,935 | AACAGACAACTACTGTAG | TCGAAACAAATTAGGATCAAACTCT | 57 | 199 |
| 4,772,587 | TACCTTGTGAAACTGCAG | ACCTGCTCGGGTCATCAAAT | 52 | 149 |
| 2,500,039 | GCTCTGTTTTCGTGCTGCAG | ACAAGATCTGTGGTGCCGAG | 60 | 531 |
| 16,726,826 | AGCCAAGGGTAGCTGCAG | CGTAGCAGCTGCATTCAAGAC | 59 | 151 |
| 25,943,049 | ATTAATAAGTGCTGCTGCAG | CGACCATTTTCGATAGCAGTA | 54 | 74 |
| 4,580,898 | ACGGTAGCAACGAACTGCAG | TACAGGTTGCAGGCTTCCAG | 60 | 86 |
| 2,507,310 | TGATGATCGAAGGGCTGCAG | TAAAGCTACTTGCGCCCACA | 60 | 192 |
| 9,699,056 | CCATCGCCATTTCCCTGCAG | TTACCCACCCCAGTACACCA | 60 | 173 |
| 25,981,291 | CTCTGCGCCTCCGTCTGCAG | AGCGCAAGCAACGTGAGAGA | 62 | 197 |
| 25,947,631 | TATCAATGTAACATCTGCAG | CCTGTTCTACTTCGTCACCGCG | 60 | 185 |
| 2,382,757 | TGCATTGCCTACATCTGCAG | GCGCAAGTAGCCCAAATACG | 60 | 97 |
| 2,386,217 | CGTACGGCCACATCCTGCAG | GGTACGCGGTGACGAAGTAG | 60 | 78 |
| 4,770,719 | CCGTACGGCCACATCCTGCAG | TTGCCGACGAAATACGCCCA | 63 | 137 |
| 7,059,241 | ATAGTAGGTGATTGCTGCAG | GGCCTGTTTGCGATTCATTT | 57 | 153 |
| 2,565,888 | TCCCCACAGCACAGCTGCAC | CCGGTTCAGTTTTTCCGGCG | 62 | 88 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Nowak, B.; Tomkowiak, A.; Bocianowski, J.; Sobiech, A.; Bobrowska, R.; Kowalczewski, P.Ł.; Bocianowska, M. The Use of DArTseq Technology to Identify Markers Linked to Genes Responsible for Seed Germination and Seed Vigor in Maize. Int. J. Mol. Sci. 2022, 23, 14865. https://doi.org/10.3390/ijms232314865
Nowak B, Tomkowiak A, Bocianowski J, Sobiech A, Bobrowska R, Kowalczewski PŁ, Bocianowska M. The Use of DArTseq Technology to Identify Markers Linked to Genes Responsible for Seed Germination and Seed Vigor in Maize. International Journal of Molecular Sciences. 2022; 23(23):14865. https://doi.org/10.3390/ijms232314865
Chicago/Turabian StyleNowak, Bartosz, Agnieszka Tomkowiak, Jan Bocianowski, Aleksandra Sobiech, Roksana Bobrowska, Przemysław Łukasz Kowalczewski, and Marianna Bocianowska. 2022. "The Use of DArTseq Technology to Identify Markers Linked to Genes Responsible for Seed Germination and Seed Vigor in Maize" International Journal of Molecular Sciences 23, no. 23: 14865. https://doi.org/10.3390/ijms232314865
APA StyleNowak, B., Tomkowiak, A., Bocianowski, J., Sobiech, A., Bobrowska, R., Kowalczewski, P. Ł., & Bocianowska, M. (2022). The Use of DArTseq Technology to Identify Markers Linked to Genes Responsible for Seed Germination and Seed Vigor in Maize. International Journal of Molecular Sciences, 23(23), 14865. https://doi.org/10.3390/ijms232314865









