Predicting Breast Cancer Relapse from Histopathological Images with Ensemble Machine Learning Models
Abstract
1. Introduction
1.1. Existing Works
1.2. Research Gap and Motivation
1.3. Objective and Contribution
- The introduction of an ensemble ML approach for enhanced breast cancer relapse prediction of histopathological images.
- The creation of an ensemble model that reduces training time and optimizes performance.
- The consideration of various approaches for better image pre-processing and extracting more relevant features for prediction.
- An improvement in patient outcomes by assisting doctors in making better treatment decisions.
1.4. Paper Structure
2. Materials and Methods
2.1. Dataset Description
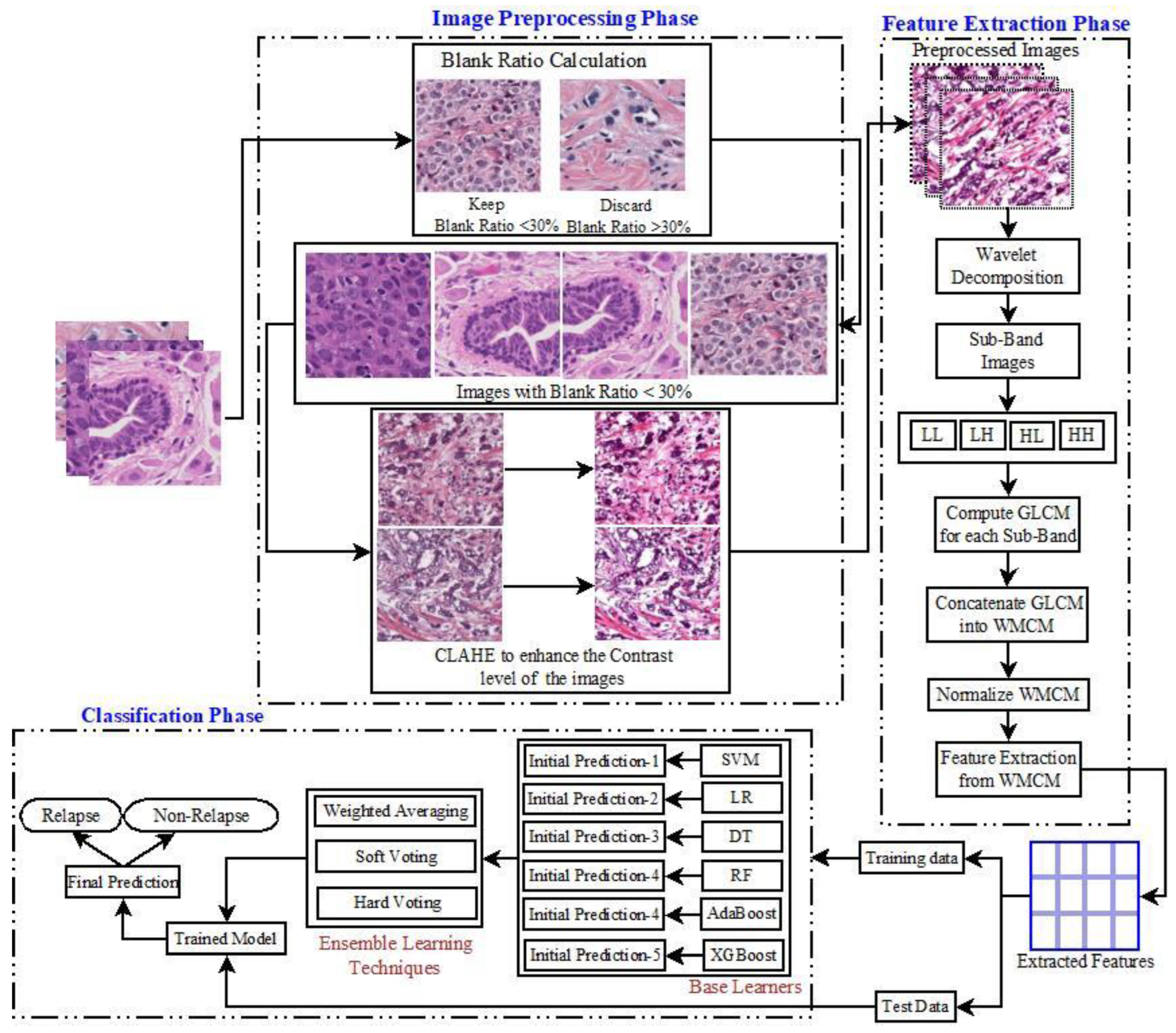
2.2. Methodology
2.2.1. Preprocessing: CLAHE
- Divide the original image into small, non-overlapping tiles.
- For each tile, compute the cumulative distribution function (C) as in Equation (1):where is the cumulative distribution function at intensity level i, is the histogram value at intensity level k, and is the dimension of the image. can be defined as Equation (2).
- Clip C to limit the contrast level for the current intensity level k as in Equation (3).where is the method used to limit the cumulative distribution function () in the range of [0, ]. is the controlling parameter for controlling the maximum contrast enhancement.
- Calculate the Histogram equalization Lookup Table (ELT) for the calculated by rounding the value to the nearest integer as in Equation (4)
- Apply to each individual pixel to find the enhanced pixel value.
2.2.2. Feature Extraction: 2D-WPT
- Step-1: Creating the WMCM
- Calculate the Gray-Level Co-Occurrence Matrix (GLCM) for each pixel (i,j) sub-bands (SBs) for HH, HL, LH, LL using Equation (9) with as the intensity value at any position (a, b). M and N are the dimensions of the image.where the is the distance from the initial position (a, b), and SB is the sub-band, which can be represented as .
- Combine the GLCM values to form the WMCM using Equation (12) with K as the total number of sub-bands available for the corresponding image.
- Step-2: WMCM-based Texture Feature Extraction Process
2.2.3. Employed ML Approaches
2.2.4. Employed EL Approaches
3. Proposed Model
4. Empirical Analysis
4.1. Critical Analysis
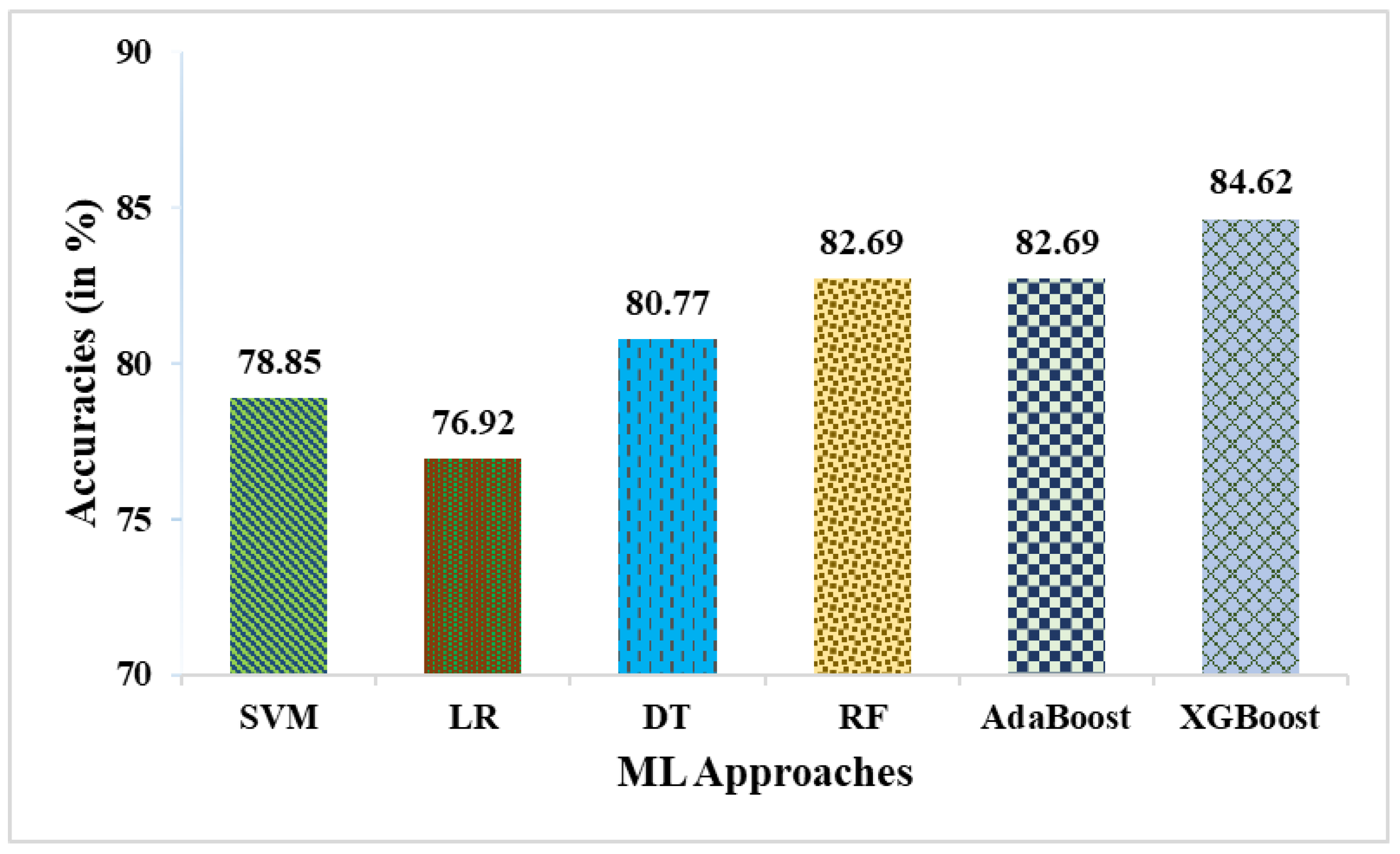
- While comparing the proposed weighted averaging of ML approaches in contrast to the ML classifiers, it outperforms the SVM, LR, DT, RF, AdaBoost, and XGBoost in terms of accuracies by ~12.19%, ~15%, ~9.52%, ~6.98%, ~6.98%, and ~4.54%, respectively.
- In the case of soft voting, it outperforms the considered ML classifiers, including SVM, LR, DT, RF, AdaBoost, and XGBoost, in terms of accuracies by ~7.32%, ~10.01%, ~4.77%, ~2.33%, ~2.33%, and ~0%, respectively.
- Comparing the hard voting ensemble technique with the ML classifiers, it outperforms SVM, LR, DT, RF, AdaBoost, and XGBoost in terms of accuracies by ~9.75 ~12.51%, ~7.14%, ~4.66%, ~4.66%, and ~2.27%, respectively.
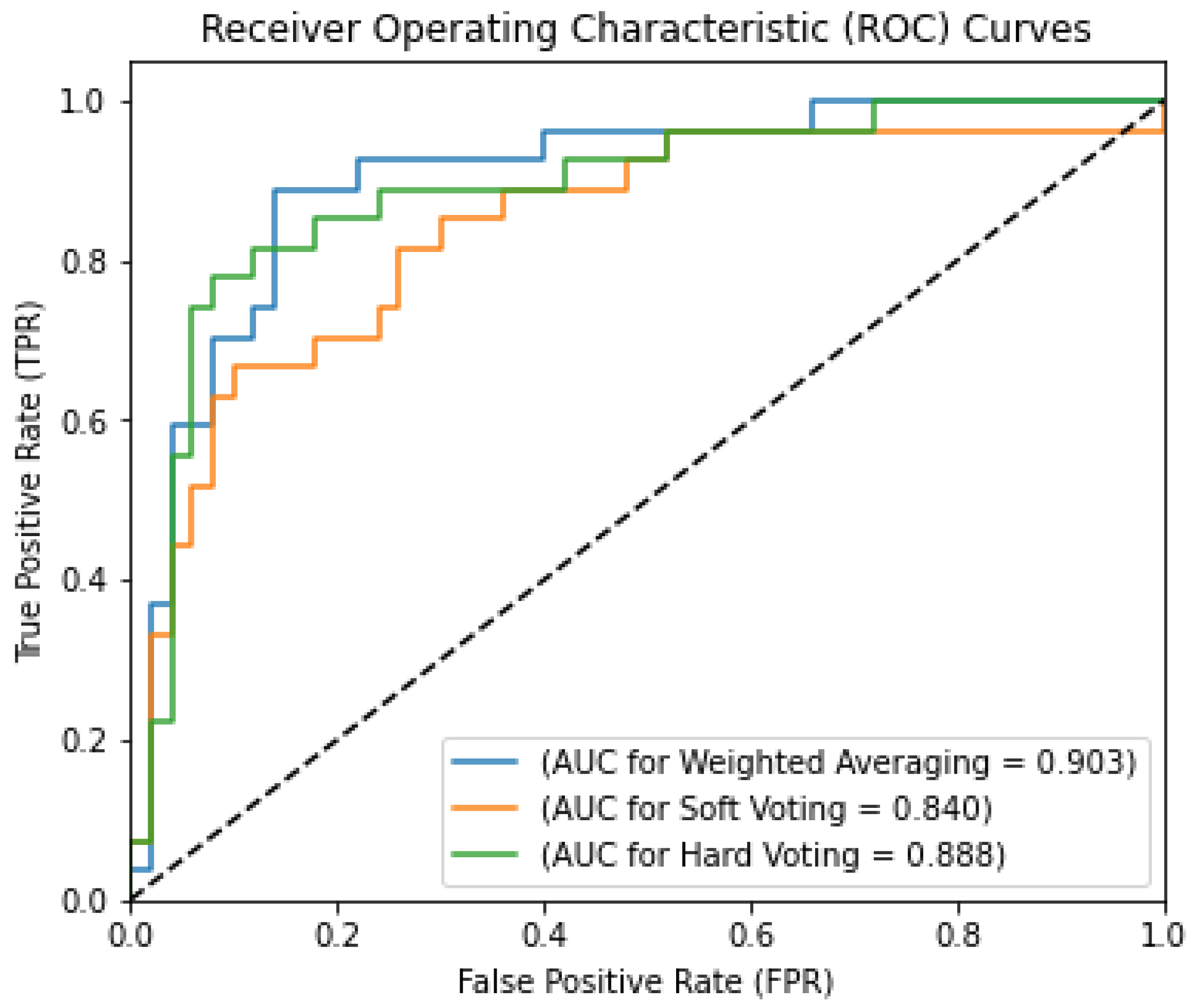
- From the above-stated comparison, it can be clearly observed that all ensemble techniques outperform the ML classifies more or less. However, three ensemble techniques were compared to find the best-fit ensemble technique for the current work. From the results obtained, it can be concluded that the weighted averaging outperforms soft voting and hard voting techniques by ~4.54% and ~2.22%, respectively, which states that the reported weighted averaging is the best-fitting ensemble technique for the current work.
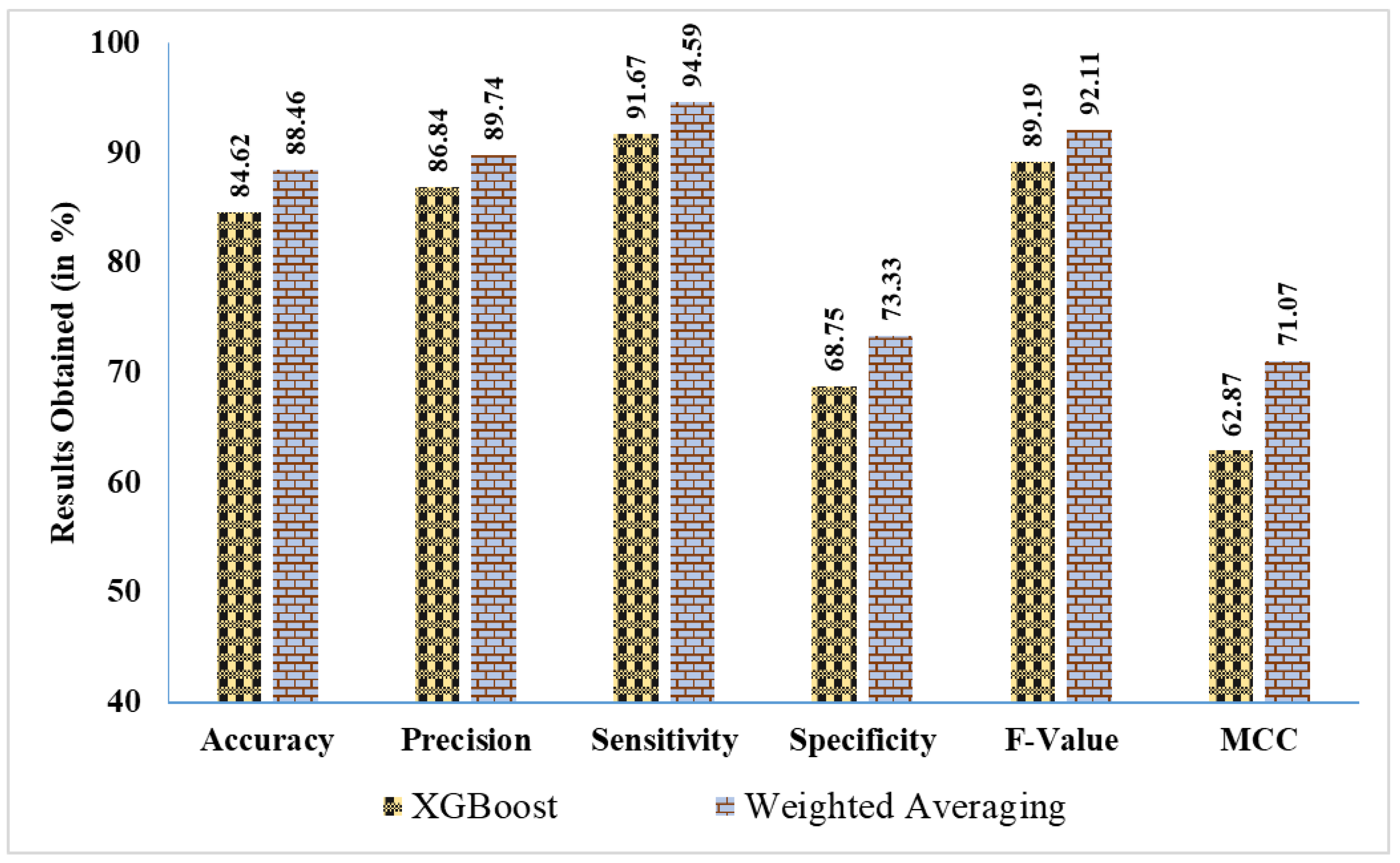
- Figure 10 presents a percentage-based comparison of results, illustrating the performance differences between the best EL approach and the best ML approach across various evaluation metrics:
- ⚬
- Weighted averaging achieves an accuracy of 88.46%, outperforming XGBoost, which obtained 84.62%. This highlights a notable improvement of ~3.84%.
- ⚬
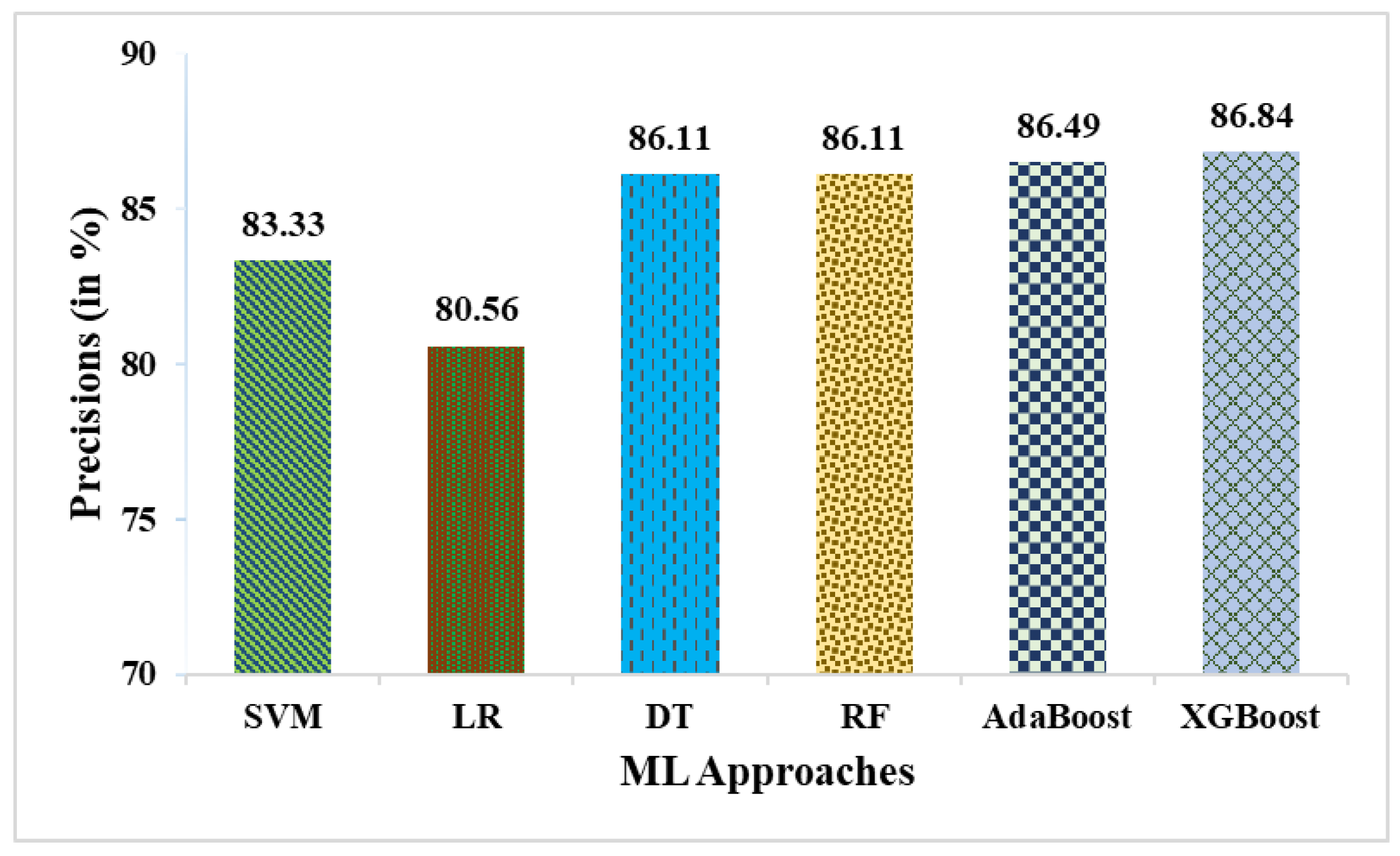
- The precision for weighted averaging is 89.74%, compared to 86.84% for XGBoost, indicating an increase of ~2.9% in precision.
- ⚬
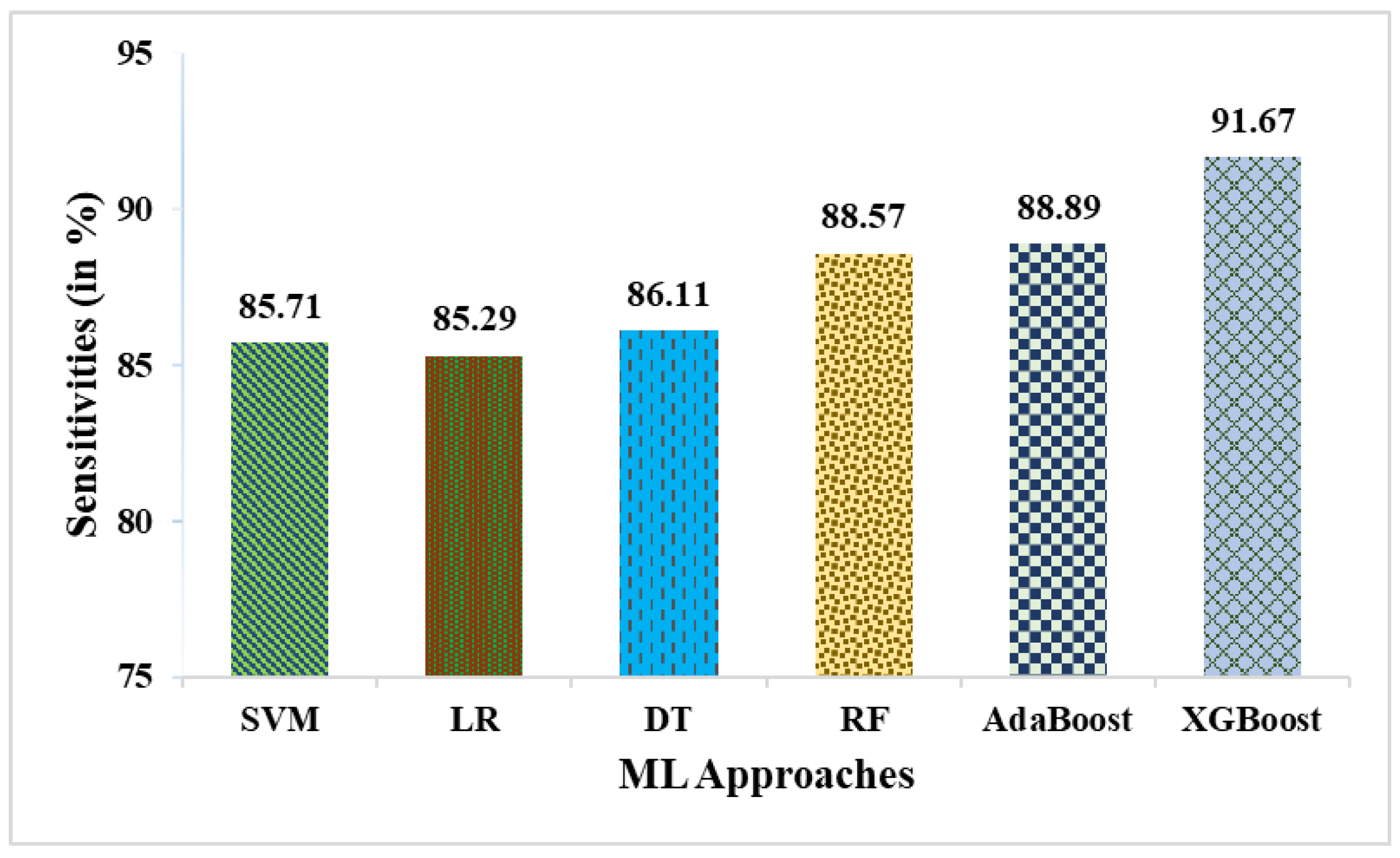
- In terms of sensitivity, weighted averaging reaches 94.59%, surpassing XGBoost’s sensitivity of 91.67% by ~2.92%. This suggests that weighted averaging is better at correctly identifying positive cases.
- ⚬
- Weighted averaging achieves 73.33% specificity, which is significantly higher than XGBoost’s 68.75%, resulting in an improvement of ~4.58% in identifying negative cases.
- ⚬
- The F-Value for weighted averaging is 92.11%, compared to 89.19% for XGBoost, showing an enhancement of ~2.92%.
- ⚬
- For MCC, weighted averaging scores 71.07%, significantly better than XGBoost’s 62.87%, demonstrating an improvement of ~8.2%, which reflects the overall quality of binary classifications.
4.2. Comparative Analysis
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Appendix A
| Algorithm A1: Pseudocode of the Proposed Model Working Principle. |
| Input: Input image set (D), total pixel (TP), Gray Level intensity (8/16/24), number of images (I) Output: Cancer Relapse, Cancer Non-Relapse Procedure: Initiate the image preprocessing phase-1 to calculate the blank ratio of an image from the dataset. For t 1←I Find the number of Blank Pixels (B) Find the TP Find the Blank Ratio (BR) as If (BR > 30%) Discard the image Else Keep the image EndIf Return D’ with a set of images having BR < 30% EndFor Initiate the preprocessing phase-2 (CLAHE) to D’ for enhancing the intensity level contrast Divide the original image into small, non-overlapping tiles (ST) For t 1←ST Calculate the Cumulative distribution function (C) using Equation (1). Clip the C to limit () the contrast level for the current intensity Return image i EndFor Form update the dataset D’ with enhanced Contrast level Apply WMCM to D’. For t ← 1 to I Divide the image into sub-bands For k ← 1 to |SB| Find GLCM for each intensity (i,j) of the image EndFor EndFor Initiate Texture feature extraction from WMCM For t ← 1 to I Calculate Calculate Texture Feature () for intensity (i,j) EndFor Find TF by combining calculated Split the TF with test size 0.2 Apply Base Learners SVM, LR, DT, RF, AdaBoost, and XGBoost to form an initial prediction. Apply Weighted Averaging, Hard Voting, and Soft Voting to form ensemble models. Evaluate the trained ensemble model over Test Data Cancer Classification. |
References
- Liu, H.; Qiu, C.; Wang, B.; Bing, P.; Tian, G.; Zhang, X.; Ma, J.; He, B.; Yang, J. Evaluating DNA methylation, gene expression, somatic mutation, and their combinations in inferring tumor tissue-of-origin. Front. Cell Dev. Biol. 2021, 9, 619330. [Google Scholar] [CrossRef]
- Ahmad, A. Current updates on trastuzumab resistance in HER2 overexpressing breast cancers. In Breast Cancer Metastasis and Drug Resistance: Challenges and Progress; Springer: Cham, Switzerland, 2019; pp. 217–228. [Google Scholar] [CrossRef]
- Sung, H.; Ferlay, J.; Siegel, R.L.; Laversanne, M.; Soerjomataram, I.; Jemal, A.; Bray, F. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 2021, 71, 209–249. [Google Scholar] [CrossRef] [PubMed]
- He, B.; Lang, J.; Wang, B.; Liu, X.; Lu, Q.; He, J.; Gao, W.; Bing, P.; Tian, G.; Yang, J. TOOme: A novel computational framework to infer cancer tissue-of-origin by integrating both gene mutation and expression. Front. Bioeng. Biotechnol. 2020, 8, 394. [Google Scholar] [CrossRef] [PubMed]
- Farahmand, S.; Fernandez, A.I.; Ahmed, F.S.; Rimm, D.L.; Chuang, J.H.; Reisenbichler, E.; Zarringhalam, K. Deep learning trained on hematoxylin and eosin tumor region of Interest predicts HER2 status and trastuzumab treatment response in HER2+ breast cancer. Mod. Pathol. 2022, 35, 44–51. [Google Scholar] [CrossRef] [PubMed]
- Ji, M.Y.; Yuan, L.; Jiang, X.D.; Zeng, Z.; Zhan, N.; Huang, P.X.; Lu, C.; Dong, W.G. Nuclear shape, architecture and orientation features from H&E images are able to predict recurrence in node-negative gastric adenocarcinoma. J. Transl. Med. 2019, 17, 92. [Google Scholar] [CrossRef] [PubMed]
- Dang, X.; Xiong, G.; Fan, C.; He, Y.; Sun, G.; Wang, S.; Liu, Y.; Zhang, L.; Bao, Y.; Xu, J.; et al. Systematic external evaluation of four preoperative risk prediction models for severe postpartum hemorrhage in patients with placenta previa: A multicenter retrospective study. J. Gynecol. Obstet. Hum. Reprod. 2022, 51, 102333. [Google Scholar] [CrossRef]
- Li, H.; Shi, W.; Shen, T.; Hui, S.; Hou, M.; Wei, Z.; Qin, S.; Bai, Z.; Cao, J. Network pharmacology-based strategy for predicting therapy targets of Ecliptae Herba on breast cancer. Medicine 2023, 102, e35384. [Google Scholar] [CrossRef]
- Sakri, S.B.; Rashid, N.B.; Zain, Z.M. Particle swarm optimization feature selection for breast cancer recurrence prediction. IEEE Access 2018, 6, 29637–29647. [Google Scholar] [CrossRef]
- Alom, M.Z.; Yakopcic, C.; Nasrin, M.S.; Taha, T.M.; Asari, V.K. Breast cancer classification from histopathological images with inception recurrent residual convolutional neural network. J. Digit. Imaging 2019, 32, 605–617. [Google Scholar] [CrossRef]
- Hong, H.C.; Chuang, C.H.; Huang, W.C.; Weng, S.L.; Chen, C.H.; Chang, K.H.; Liao, K.W.; Huang, H.D. A panel of eight microRNAs is a good predictive parameter for triple-negative breast cancer relapse. Theranostics 2020, 10, 8771. [Google Scholar] [CrossRef]
- Yan, S.; Wang, W.; Zhu, B.; Pan, X.; Wu, X.; Tao, W. Construction of nomograms for predicting pathological complete response and tumor shrinkage size in breast cancer. Cancer Manag. Res. 2020, 20, 8313–8323. [Google Scholar] [CrossRef] [PubMed]
- Mosayebi, A.; Mojaradi, B.; Bonyadi Naeini, A.; Khodadad Hosseini, S.H. Modeling and comparing data mining algorithms for prediction of recurrence of breast cancer. PLoS ONE 2020, 15, e0237658. [Google Scholar] [CrossRef]
- Comes, M.C.; La Forgia, D.; Didonna, V.; Fanizzi, A.; Giotta, F.; Latorre, A.; Martinelli, E.; Mencattini, A.; Paradiso, A.V.; Tamborra, P.; et al. Early prediction of breast cancer recurrence for patients treated with neoadjuvant chemotherapy: A transfer learning approach on DCE-MRIs. Cancers 2021, 13, 2298. [Google Scholar] [CrossRef] [PubMed]
- Sanyal, J.; Tariq, A.; Kurian, A.W.; Rubin, D.; Banerjee, I. Weakly supervised temporal model for prediction of breast cancer distant recurrence. Sci. Rep. 2021, 11, 9461. [Google Scholar] [CrossRef]
- Conde-Sousa, E.; Vale, J.; Feng, M.; Xu, K.; Wang, Y.; Della Mea, V.; La Barbera, D.; Montahaei, E.; Baghshah, M.; Turzynski, A.; et al. HEROHE challenge: Predicting HER2 status in breast cancer from hematoxylin–eosin whole-slide imaging. J. Imaging 2022, 8, 213. [Google Scholar] [CrossRef]
- Rabinovici-Cohen, S.; Fernández, X.M.; Grandal Rejo, B.; Hexter, E.; Hijano Cubelos, O.; Pajula, J.; Pölönen, H.; Reyal, F.; Rosen-Zvi, M. Multimodal prediction of five-year breast Cancer recurrence in women who receive Neoadjuvant chemotherapy. Cancers 2022, 14, 3848. [Google Scholar] [CrossRef]
- Liu, X.; Yuan, P.; Li, R.; Zhang, D.; An, J.; Ju, J.; Liu, C.; Ren, F.; Hou, R.; Li, Y.; et al. Predicting breast cancer recurrence and metastasis risk by integrating color and texture features of histopathological images and machine learning technologies. Comput. Biol. Med. 2022, 146, 105569. [Google Scholar] [CrossRef]
- Yang, J.; Ju, J.; Guo, L.; Ji, B.; Shi, S.; Yang, Z.; Gao, S.; Yuan, X.; Tian, G.; Liang, Y.; et al. Prediction of HER2-positive breast cancer recurrence and metastasis risk from histopathological images and clinical information via multimodal deep learning. Comput. Struct. Biotechnol. J. 2022, 20, 333–342. [Google Scholar] [CrossRef]
- Lu, W.; Toss, M.; Dawood, M.; Rakha, E.; Rajpoot, N.; Minhas, F. SlideGraph+: Whole slide image level graphs to predict HER2 status in breast cancer. Med. Image Anal. 2022, 80, 102486. [Google Scholar] [CrossRef]
- Su, Z.; Niazi, M.K.; Tavolara, T.E.; Niu, S.; Tozbikian, G.H.; Wesolowski, R.; Gurcan, M.N. BCR-Net: A deep learning framework to predict breast cancer recurrence from histopathology images. PLoS ONE 2023, 18, e0283562. [Google Scholar] [CrossRef]
- Liu, Y.; Shen, D.; Wang, H.Y.; Qi, M.Y.; Zeng, Q.Y. Development and validation to predict visual acuity and keratometry two years after corneal crosslinking with progressive keratoconus by machine learning. Front. Med. 2023, 10, 1146529. [Google Scholar] [CrossRef] [PubMed]
- Botlagunta, M.; Botlagunta, M.D.; Myneni, M.B.; Lakshmi, D.; Nayyar, A.; Gullapalli, J.S.; Shah, M.A. Classification and diagnostic prediction of breast cancer metastasis on clinical data using machine learning algorithms. Sci. Rep. 2023, 13, 485. [Google Scholar] [CrossRef] [PubMed]
- Dammu, H.; Ren, T.; Duong, T.Q. Deep learning prediction of pathological complete response, residual cancer burden, and progression-free survival in breast cancer patients. PLoS ONE 2023, 18, e0280148. [Google Scholar] [CrossRef]
- Bayramoglu, N.; Kannala, J.; Heikkilä, J. Deep learning for magnification independent breast cancer histopathology image classification. In Proceedings of the 2016 23rd International Conference on Pattern Recognition (ICPR), Cancun, Mexico, 4–8 December 2016; pp. 2440–2445. [Google Scholar] [CrossRef]
- Rawat, R.R.; Ortega, I.; Roy, P.; Sha, F.; Shibata, D.; Ruderman, D.; Agus, D.B. Deep learned tissue “fingerprints” classify breast cancers by ER/PR/Her2 status from H&E images. Sci. Rep. 2020, 10, 7275. [Google Scholar] [CrossRef]
- Pati, A.; Panigrahi, A.; Parhi, M.; Giri, J.; Qin, H.; Mallik, S.; Pattanayak, S.R.; Agrawal, U.K. Performance assessment of hybrid machine learning approaches for breast cancer and recurrence prediction. PLoS ONE 2024, 19, e0304768. [Google Scholar] [CrossRef]
- The Cancer Genome Atlas (TCGA). Available online: https://portal.gdc.cancer.gov/ (accessed on 22 July 2023).
- Alwazzan, M.J.; Ismael, M.A.; Ahmed, A.N. A hybrid algorithm to enhance colour retinal fundus images using a Wiener filter and CLAHE. J. Digit. Imaging 2021, 34, 750–759. [Google Scholar] [CrossRef]
- Hayati, M.; Muchtar, K.; Maulina, N.; Syamsuddin, I.; Elwirehardja, G.N.; Pardamean, B. Impact of CLAHE-based image enhancement for diabetic retinopathy classification through deep learning. Procedia Comput. Sci. 2023, 216, 57–66. [Google Scholar] [CrossRef]
- Srivastava, R.; Kumar, P. Deep-GAN: An improved model for thyroid nodule identification and classification. Neural Comput. Appl. 2024, 36, 7685–7704. [Google Scholar] [CrossRef]
- Pati, A.; Panigrahi, A.; Nayak, D.S.; Sahoo, G.; Singh, D. Predicting pediatric appendicitis using ensemble learning techniques. Procedia Comput. Sci. 2023, 218, 1166–1175. [Google Scholar] [CrossRef]
- Panigrahi, A.; Pati, A.; Sahu, B.; Das, M.N.; Nayak, D.S.; Sahoo, G.; Kant, S. En-MinWhale: An ensemble approach based on MRMR and Whale optimization for Cancer diagnosis. IEEE Access 2023, 11, 113526–113542. [Google Scholar] [CrossRef]
- Sahoo, G.; Nayak, A.K.; Tripathy, P.K.; Tripathy, J. A novel machine learning based hybrid approach for breast cancer relapse prediction. Indones. J. Electr. Eng. Comput. Sci. (IJEECS) 2023, 32, 1655–1663. [Google Scholar] [CrossRef]
- Pati, A.; Panigrahi, A.; Sahu, B.; Sahoo, G.; Dash, M.; Parhi, M.; Pattanayak, B.K. FOHC: Firefly Optimizer Enabled Hybrid approach for Cancer Classification. Int. J. Recent Innov. Trends Comput. Commun. 2023, 11, 118–125. [Google Scholar] [CrossRef]


| Reference | Methodologies | Dataset(s) | Outcomes |
|---|---|---|---|
| Sakri et al. [9] | REPTree, NB, and KNN-IBK with PSO | WPBC | Accuracy: 81.3%, Precision: 88.3%, Recall: 93.4%, F-Score: 87.7%, AUC: 0.820 |
| Alom et al. [10] | Inception-v4, ResNet, and RCNN | BreakHis and BCD | Accuracy: 100%, Sensitivity: 100%, Specificity: 100%, AUC: 1.0 |
| Hong et al. [11] | LR and Gaussian mixture | TCGA_TNBC, GEOD-40525, GSE40049 and GSE19783 | AUC (GSE40049): 0.89, AUC (GSE19783): 0.90 |
| Yan et al. [12] | LR | BCC at HMUCH | AUC (HER2-Positive): 0.820, AUC (TNBC): 0.785 |
| Mosayebi et al. [13] | RF, LVQ, NB, C5.0 DT, MLP, KPCA-SVM, and SVM | MHME-ICRC | Accuracy: 81.9%, Sensitivity: 86.9%, Specificity:77.7%, F-Value: 81.6%, AUC: 0.774 |
| Comes et al. [14] | CNN and SVM | I-SPY1 TRIAL and BREAST-MRI-NACT-Pilot | Accuracy: 85.2%, Sensitivity: 84.6%, AUC: 0.83 |
| Sanyal et al. [15] | LSTM and XGBoost | Manually and NLP-curated | Sensitivity: 89.0%, Specificity: 84.0%, AUC: 0.94 |
| Conde-Sousa et al. [16] | DL | HEROHE | Precision: 79%, Recall: 100%, F-Value: 79%, AUC: 0.88 |
| Rabinovici-Cohen et al. [17] | CNN | Real-World Retrospective Dataset | Specificity: 57%, F-Value: 56%, Balanced Accuracy: 72%, PPV: 41%, NPV: 96%, Sensitivity: 93%, AUC: 0.75 |
| Liu et al. [18] | RF, LR, and XGBoost | Clinical dataset | Accuracy: 80%, Precision: 40%, Recall: 75%, F-Value: 50%, AUC: 0.75 |
| TCGA | Accuracy: 73%, Precision: 33%, Recall: 60%, F-Value: 42%, AUC: 0.72 | ||
| Yang et al. [19] | CNN and ResNet50 | CAMS | AUC: 0.76 |
| TCGA | AUC: 0.72 | ||
| Lu et al. [20] | GNN-based Slide Graph | TCGA | AUC: 0.75 |
| HER2C and Nott-HER2 | AUC: 0.80 | ||
| Su et al. [21] | CNN | H&E and Ki67 | Accuracy: 80%, F-Value: 79.2%, AUC: 0.811 |
| Liu et al. [22] | XGBoost | AEHWU | R-Square (CVDA): 0.9993, R-Square (Kmax): 0.9888 |
| Botlagunta et al. [23] | LR, KNN, DT, RF, SVM, GB, and XGBoost | Medical data on MBC | Accuracy: 83%, Precision: 83%, Recall: 100%, F-Value: 85%, AUC: 0.87 |
| Dammu et al. [24] | CNN | I-SPY-1 TRIAL | Accuracy: 81%, Sensitivity: 68%, Specificity: 97%, F-Value: 76%, AUC: 0.83 |
| Dataset | Parameters | |||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Stages of Tumor | ER | PR | Lymph Node Status | Age (Years) | Outcome | Total | ||||||||
| I | II | III | ER− | ER+ | PR− | PR+ | LMN− | LMN+ | <50 | ≥50 | R | NR | ||
| TCGA | 14 | 77 | 32 | 33 | 90 | 50 | 73 | 58 | 65 | 34 | 89 | 5 | 118 | 123 |
| ML Approaches | Results Obtained (in %) | |||||
|---|---|---|---|---|---|---|
| Accuracy | Precision | Sensitivity | Specificity | F-Value | MCC | |
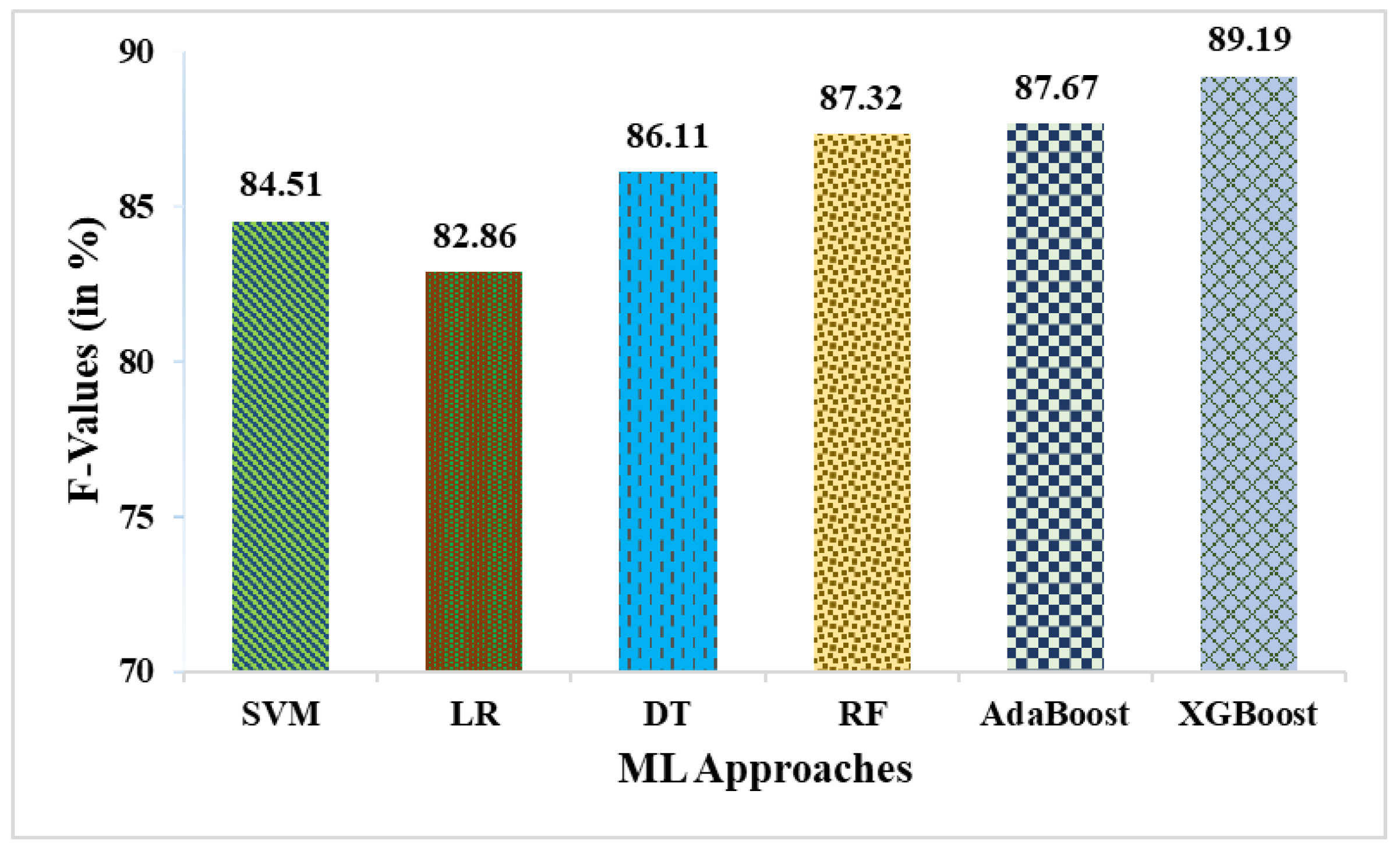
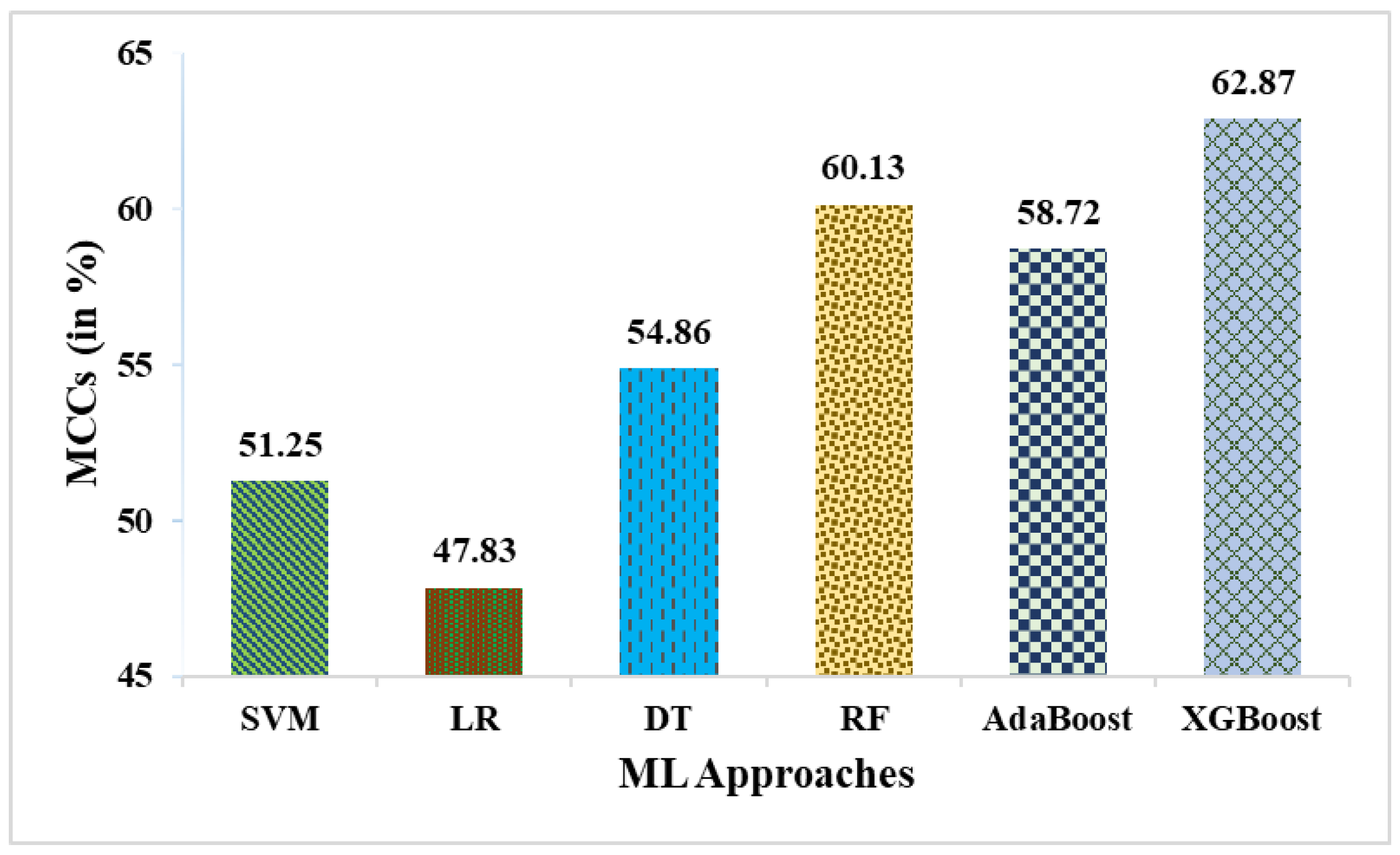
| SVM | 78.85 | 83.33 | 85.71 | 64.71 | 84.51 | 51.25 |
| LR | 76.92 | 80.56 | 85.29 | 61.11 | 82.86 | 47.83 |
| DT | 80.77 | 86.11 | 86.11 | 68.75 | 86.11 | 54.86 |
| RF | 82.69 | 86.11 | 88.57 | 70.59 | 87.32 | 60.13 |
| AdaBoost | 82.69 | 86.49 | 88.89 | 68.75 | 87.67 | 58.72 |
| XGBoost | 84.62 | 86.84 | 91.67 | 68.75 | 89.19 | 62.87 |
| EL Approaches | Results Obtained (in %) | |||||
|---|---|---|---|---|---|---|
| Accuracy | Precision | Sensitivity | Specificity | F-Value | MCC | |
| Weighted Averaging | 88.46 | 89.74 | 94.59 | 73.33 | 92.11 | 71.07 |
| Soft Voting | 84.62 | 86.11 | 91.18 | 72.22 | 88.57 | 65.35 |
| Hard Voting | 86.54 | 86.49 | 94.12 | 72.22 | 90.14 | 69.66 |
| Ref. | Dataset(s) | Comparison Parameters | ||||||
|---|---|---|---|---|---|---|---|---|
| Accuracy (%) | Precision (%) | Sensitivity (%) | Specificity (%) | F-Value (%) | MCC (%) | AUC | ||
| [9] | WPBC | 81.3 | 88.3 | 93.4 | - | 87.7 | - | 0.820 |
| [10] | BreakHis and BCD | 100 | - | 100 | 100 | - | - | 1.0 |
| [11] | GSE40049 | - | - | - | - | - | - | 0.89 |
| GSE19783 | - | - | - | - | - | - | 0.90 | |
| [12] | BCC at HMUCH | - | - | - | - | - | - | 0.820 |
| [13] | MHME-ICRC | 81.9 | - | 86.9 | 77.7 | 81.6 | - | 0.774 |
| [14] | I-SPY1 TRIAL, BREAST-MRI-NACT-Pilot | 85.2 | - | 84.6 | - | - | - | 0.83 |
| [15] | Manually and NLP-curated | - | - | 89.0 | 84.0 | - | - | 0.94 |
| [16] | HEROHE | - | 79 | 100 | - | 79 | - | 0.88 |
| [17] | Real-World Retrospective Dataset | - | - | 93 | 57 | 56 | - | 0.75 |
| [18] | Clinical dataset | 80 | 40 | 75 | - | 50 | - | 0.75 |
| TCGA | 73 | 33 | 60 | - | 42 | - | 0.72 | |
| [19] | CAMS | - | - | - | - | - | - | 0.76 |
| TCGA | - | - | - | - | - | - | 0.72 | |
| [20] | TCGA | - | - | - | - | - | - | 0.75 |
| HER2C and Nott-HER2 | - | - | - | - | - | - | 0.80 | |
| [21] | H&E and Ki67 | 80 | - | - | - | 79.2 | - | 0.811 |
| [22] | AEHWU | - | - | - | - | - | - | - |
| [23] | Medical data on MBC | 83 | 83 | 100 | - | 85 | - | 0.87 |
| [24] | I-SPY-1 TRIAL | 81 | - | 68 | 97 | 76 | - | 0.83 |
| [Proposed] | TCGA | 88.46 | 89.74 | 94.59 | 73.33 | 92.11 | 71.07 | 0.903 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sahoo, G.; Nayak, A.K.; Tripathy, P.K.; Panigrahi, A.; Pati, A.; Sahu, B.; Mahanty, C.; Mallik, S. Predicting Breast Cancer Relapse from Histopathological Images with Ensemble Machine Learning Models. Curr. Oncol. 2024, 31, 6577-6597. https://doi.org/10.3390/curroncol31110486
Sahoo G, Nayak AK, Tripathy PK, Panigrahi A, Pati A, Sahu B, Mahanty C, Mallik S. Predicting Breast Cancer Relapse from Histopathological Images with Ensemble Machine Learning Models. Current Oncology. 2024; 31(11):6577-6597. https://doi.org/10.3390/curroncol31110486
Chicago/Turabian StyleSahoo, Ghanashyam, Ajit Kumar Nayak, Pradyumna Kumar Tripathy, Amrutanshu Panigrahi, Abhilash Pati, Bibhuprasad Sahu, Chandrakanta Mahanty, and Saurav Mallik. 2024. "Predicting Breast Cancer Relapse from Histopathological Images with Ensemble Machine Learning Models" Current Oncology 31, no. 11: 6577-6597. https://doi.org/10.3390/curroncol31110486
APA StyleSahoo, G., Nayak, A. K., Tripathy, P. K., Panigrahi, A., Pati, A., Sahu, B., Mahanty, C., & Mallik, S. (2024). Predicting Breast Cancer Relapse from Histopathological Images with Ensemble Machine Learning Models. Current Oncology, 31(11), 6577-6597. https://doi.org/10.3390/curroncol31110486








