Role of Polygenic Risk Score in Cancer Precision Medicine of Non-European Populations: A Systematic Review
Abstract
:1. Introduction
2. Materials and Methods
2.1. Strategy of Research Question Definition
2.2. Search the Database
3. Results
4. PRS and the Risk of Breast Cancer
| Author, Year | Phenotype | Population/ Ethnicity | Subjects (n) | SNPs (n) | Significant SNPs (n) | Main Findings |
|---|---|---|---|---|---|---|
| Evans et al., 2021 [20] | Breast Cancer | Asian Black Jewish Mixed White others Unknown | 119 112 120 44 159 274 | 18-SNP 143-SNP | rs3803662 (TOX3) rs2981579 (FGFR2) | Not replicated |
| Allman et al., 2020 [32] | Breast Cancer | African American | Control: 7005 Case: 416 | 75-SNP | - | Replicated |
| Caucasian | Control: 405 Case: 750 | 77-SNP | ||||
| Hispanic | Control: 3210 Case: 147 | 71-SNP | ||||
| Starlard-Davenport et al., 2018 [33] | Breast Cancer | African-American | Control: 559 Case: 319 | 75-SNP | - | Replicated |
| Zhang et al., 2018 [34] | Breast Cancer | American | Control: 7874 Case: 4006 | 67-SNP | - | Replicated |
| Shi et al., 2020 [35] | Breast Cancer | Non-Hispanic | Control: 1120 Case: 1152 | 77-SNP | - | Replicated |
| Shieh et al., 2020 [36] | Breast Cancer | USA Latin and Latin American | Control: 7622 Cases: 4658 | 180-SNP | - | Replicated |
| Control: 7622 Cases: 4658 | 71-SNP | |||||
| Ho et al., 2020 [37] | Breast Cancer | Asia | Control: 16,483 Case: 15,755 | 287-SNP | - | Replicated |
| Hsieh et al., 2017 [38] | Breast Cancer | Asia | Control: 514 Case: 446 | 6-SNP | rs2981582 (FGFR2) rs981782 (HCN1) rs889312 (MAP3K1) rs3803662 (TOX3) rs10822013 (ZNF365) rs3784099 (RAD51B) | Replicated |
| Wen et al., 2016 [39] | Breast Cancer | Asia | Control: 11,612 Case: 11,760 | 44-SNP | rs2046210 (C6orf97) rs10822013 (ZNF365) rs2363956 (ANKLE1) | Replicated |
| Chan et al., 2018 [40] | Breast Cancer | Asia | Control: 885 Case: 1294 Control: 243 Case: 301 Model 1 Model 2 Model 3 | 51-SNP 46-SNP 11-SNP 9-SNP | rs16886165 (MAP3K1) rs3757318 (ESR1) rs11155804 (ESR1) rs12662670 (ESR1) rs2046210 (ERS1) rs10816625 (CHCHD4P2) rs704010 (ZMIZ1) rs2981579 (FGFR2) rs909116 (LSP1) rs7297051 (PTHLH) rs4784227 (TOX3) | Replicated |
| Coignet et al., 2017 [41] | Breast Cancer | African-American | Control: 744 Case: 621 | 53-SNP | rs2947411 (TMEM18) rs466639 (RXRG) | Not replicated |
| Wang et al., 2018 [42] | Breast Cancer | African | Control: 2029 Case: 1657 | 34-SNP | - | Not replicated |
| Wang et al., 2018 [43] | Pancreatic Cancer/Breast Cancer | African | Control: 2029 Case: 1657 | 23-SNP | rs31490 (CLPTM1L) rs40168 (CLPTM1L) | Not replicated |
5. PRS and the Risk of Prostate Cancer
6. PRS and the Risk of Pancreatic Cancer
7. PRS and the Risk of Melanoma
8. PRS and the Risk of Chronic Lymphocytic Leukemia
9. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Sung, H.; Ferlay, J.; Siegel, R.L.; Laversanne, M.; Soerjomataram, I.; Jemal, A.; Bray, F. Global Cancer Statistics 2020: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA Cancer J. Clin. 2021, 71, 209–249. [Google Scholar] [CrossRef]
- Siegel, R.L.; Miller, K.D.; Fuchs, H.E.; Jemal, A. Cancer Statistics, 2022. CA Cancer J. Clin. 2022, 72, 7–33. [Google Scholar] [CrossRef] [PubMed]
- Ward, E.; Jemal, A.; Cokkinides, V.; Singh, G.K.; Cardinez, C.; Ghafoor, A.; Thun, M. Cancer Disparities by Race/Ethnicity and Socioeconomic Status. CA Cancer J. Clin. 2004, 54, 78–93. [Google Scholar] [CrossRef] [PubMed]
- Jones, K.M.; Cook-Deegan, R.; Rotimi, C.N.; Callier, S.L.; Bentley, A.R.; Stevens, H.; Phillips, K.A.; Jansen, J.P.; Weyant, C.F.; Roberts, D.E.; et al. Complicated Legacies: The Human Genome at 20. Science 2021, 371, 564–569. [Google Scholar] [CrossRef]
- James, J.E.; Riddle, L.; Koenig, B.A.; Joseph, G. The Limits of Personalization in Precision Medicine: Polygenic Risk Scores and Racial Categorization in a Precision Breast Cancer Screening Trial. PLoS ONE 2021, 16, e0258571. [Google Scholar] [CrossRef]
- Hodson, R. Precision Medicine. Nature 2016, 537, S49. [Google Scholar] [CrossRef] [Green Version]
- Weinstein, J.N.; Collisson, E.A.; Mills, G.B.; Shaw, K.R.M.; Ozenberger, B.A.; Ellrott, K.; Shmulevich, I.; Sander, C.; Stuart, J.M. The Cancer Genome Atlas Pan-Cancer Analysis Project. Nat. Genet. 2013, 45, 1113–1120. [Google Scholar] [CrossRef]
- Hoadley, K.A.; Yau, C.; Hinoue, T.; Wolf, D.M.; Lazar, A.J.; Drill, E.; Shen, R.; Taylor, A.M.; Cherniack, A.D.; Thorsson, V.; et al. Cell-of-Origin Patterns Dominate the Molecular Classification of 10,000 Tumors from 33 Types of Cancer. Cell 2018, 173, 291–304. [Google Scholar] [CrossRef] [Green Version]
- Zhang, J.; Baran, J.; Cros, A.; Guberman, J.M.; Haider, S.; Hsu, J.; Liang, Y.; Rivkin, E.; Wang, J.; Whitty, B.; et al. International Cancer Genome Consortium Data Portal--a One-Stop Shop for Cancer Genomics Data. Database 2011, 2011, bar026. [Google Scholar] [CrossRef] [Green Version]
- Hudson, T.J.; Anderson, W.; Artez, A.; Barker, A.D.; Bell, C.; Bernabé, R.R.; Bhan, M.K.; Calvo, F.; Eerola, I.; Gerhard, D.S.; et al. International Network of Cancer Genome Projects. Nature 2010, 464, 993–998. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sudlow, C.; Gallacher, J.; Allen, N.; Beral, V.; Burton, P.; Danesh, J.; Downey, P.; Elliott, P.; Green, J.; Landray, M.; et al. UK Biobank: An Open Access Resource for Identifying the Causes of a Wide Range of Complex Diseases of Middle and Old Age. PLoS Med. 2015, 12, e1001779. [Google Scholar] [CrossRef] [Green Version]
- Ollier, W.; Sprosen, T.; Peakman, T. UK Biobank: From Concept to Reality. Pharmacogenomics 2005, 6, 639–646. [Google Scholar] [CrossRef]
- Janssens, A.; Cecile, J.W.; Ioannidis, J.P.A.; Bedrosian, S.; Boffetta, P.; Dolan, S.M.; Dowling, N.; Fortier, I.; Freedman, A.N.; Grimshaw, J.M.; et al. Strengthening the Reporting of Genetic Risk Prediction Studies (GRIPS): Explanation and Elaboration. Eur. J. Clin. Investig. 2011, 41, 1010–1035. [Google Scholar] [CrossRef] [Green Version]
- Pearson, T.A. How to Interpret a Genome-Wide Association Study. JAMA 2008, 299, 1335–1344. [Google Scholar] [CrossRef]
- Saulsberry, L.; Olopade, O.I. Precision Oncology: Directing Genomics and Pharmacogenomics toward Reducing Cancer Inequities. Cancer Cell 2021, 39, 730–733. [Google Scholar] [CrossRef]
- Rebbeck, T.R.; Mahal, B.; Maxwell, K.N.; Garraway, I.P.; Yamoah, K. The Distinct Impacts of Race and Genetic Ancestry on Health. Nat. Med. 2022, 28, 890–893. [Google Scholar] [CrossRef]
- Pereira, R.; Phillips, C.; Pinto, N.; Santos, C.; dos Santos, S.E.B.; Amorim, A.; Carracedo, Á.; Gusmão, L. Straightforward Inference of Ancestry and Admixture Proportions through Ancestry-Informative Insertion Deletion Multiplexing. PLoS ONE 2012, 7, e29684. [Google Scholar] [CrossRef] [Green Version]
- Norris, E.T.; Wang, L.; Conley, A.B.; Rishishwar, L.; Mariño-Ramírez, L.; Valderrama-Aguirre, A.; Jordan, I.K. Genetic Ancestry, Admixture and Health Determinants in Latin America. BMC Genom. 2018, 19, 861. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Buniello, A.; MacArthur, J.A.L.; Cerezo, M.; Harris, L.W.; Hayhurst, J.; Malangone, C.; McMahon, A.; Morales, J.; Mountjoy, E.; Sollis, E.; et al. The NHGRI-EBI GWAS Catalog of Published Genome-Wide Association Studies, Targeted Arrays and Summary Statistics 2019. Nucleic Acids Res. 2019, 47, D1005–D1012. [Google Scholar] [CrossRef] [Green Version]
- Evans, D.G.; Veen, E.M.; Byers, H.; Roberts, E.; Howell, A.; Howell, S.J.; Harkness, E.F.; Brentnall, A.; Cuzick, J.; Newman, W.G. The Importance of Ethnicity: Are Breast Cancer Polygenic Risk Scores Ready for Women Who Are Not of White European Origin? Int. J. Cancer 2021, 150, 73–79. [Google Scholar] [CrossRef]
- Kleinstern, G.; Weinberg, J.B.; Parikh, S.A.; Braggio, E.; Achenbach, S.J.; Robinson, D.P.; Norman, A.D.; Rabe, K.G.; Boddicker, N.J.; Vachon, C.M.; et al. Polygenic Risk Score and Risk of Monoclonal B-Cell Lymphocytosis in Caucasians and Risk of Chronic Lymphocytic Leukemia (CLL) in African Americans. Leukemia 2021, 36, 119–125. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.; Benyamin, B.; McEvoy, B.P.; Gordon, S.; Henders, A.K.; Nyholt, D.R.; Madden, P.A.; Heath, A.C.; Martin, N.G.; Montgomery, G.W.; et al. Common SNPs Explain a Large Proportion of the Heritability for Human Height. Nat. Genet. 2010, 42, 565–569. [Google Scholar] [CrossRef] [Green Version]
- Choi, S.W.; Mak, T.S.-H.; O’Reilly, P.F. Tutorial: A Guide to Performing Polygenic Risk Score Analyses. Nat. Protoc. 2020, 15, 1–14. [Google Scholar] [CrossRef]
- Wray, N.R.; Goddard, M.E.; Visscher, P.M. Prediction of Individual Genetic Risk to Disease from Genome-Wide Association Studies. Genome Res. 2007, 17, 1520–1528. [Google Scholar] [CrossRef] [Green Version]
- Fletcher, O.; Houlston, R.S. Architecture of Inherited Susceptibility to Common Cancer. Nat. Rev. Cancer 2010, 10, 353–361. [Google Scholar] [CrossRef]
- Mavaddat, N.; Michailidou, K.; Dennis, J.; Lush, M.; Fachal, L.; Lee, A.; Tyrer, J.P.; Chen, T.-H.; Wang, Q.; Bolla, M.K.; et al. Polygenic Risk Scores for Prediction of Breast Cancer and Breast Cancer Subtypes. Am. J. Hum. Genet. 2019, 104, 21–34. [Google Scholar] [CrossRef] [Green Version]
- Wand, H.; Lambert, S.A.; Tamburro, C.; Iacocca, M.A.; O’Sullivan, J.W.; Sillari, C.; Kullo, I.J.; Rowley, R.; Dron, J.S.; Brockman, D.; et al. Improving Reporting Standards for Polygenic Scores in Risk Prediction Studies. Nature 2021, 591, 211–219. [Google Scholar] [CrossRef]
- Lambert, S.A.; Gil, L.; Jupp, S.; Ritchie, S.C.; Xu, Y.; Buniello, A.; McMahon, A.; Abraham, G.; Chapman, M.; Parkinson, H.; et al. The Polygenic Score Catalog as an Open Database for Reproducibility and Systematic Evaluation. Nat. Genet. 2021, 53, 420–425. [Google Scholar] [CrossRef]
- Lockwood, C.; Munn, Z.; Porritt, K. Qualitative Research Synthesis: Methodological Guidance for Systematic Reviewers Utilizing Meta-Aggregation. Int. J. Evid.-Based Healthc. 2015, 13, 179–187. [Google Scholar] [CrossRef]
- Page, M.J.; McKenzie, J.E.; Bossuyt, P.M.; Boutron, I.; Hoffmann, T.C.; Mulrow, C.D.; Shamseer, L.; Tetzlaff, J.M.; Akl, E.A.; Brennan, S.E.; et al. The PRISMA 2020 Statement: An Updated Guideline for Reporting Systematic Reviews. BMJ 2021, 88, 105906. [Google Scholar] [CrossRef]
- Lilyquist, J.; Ruddy, K.J.; Vachon, C.M.; Couch, F.J. Common Genetic Variation and Breast Cancer Risk—Past, Present, and Future. Cancer Epidemiol. Biomark. Prev. 2018, 27, 380–394. [Google Scholar] [CrossRef] [Green Version]
- Allman, R.; Spaeth, E.; Lai, J.; Gross, S.J.; Hopper, J.L. A Streamlined Model for Use in Clinical Breast Cancer Risk Assessment Maintains Predictive Power and Is Further Improved with Inclusion of a Polygenic Risk Score. PLoS ONE 2021, 16, e0245375. [Google Scholar] [CrossRef]
- Starlard-Davenport, A.; Allman, R.; Dite, G.S.; Hopper, J.L.; Spaeth Tuff, E.; Macleod, S.; Kadlubar, S.; Preston, M.; Henry-Tillman, R. Validation of a Genetic Risk Score for Arkansas Women of Color. PLoS ONE 2018, 13, e0204834. [Google Scholar] [CrossRef]
- Zhang, X.; Rice, M.; Tworoger, S.S.; Rosner, B.A.; Eliassen, A.H.; Tamimi, R.M.; Joshi, A.D.; Lindstrom, S.; Qian, J.; Colditz, G.A.; et al. Addition of a Polygenic Risk Score, Mammographic Density, and Endogenous Hormones to Existing Breast Cancer Risk Prediction Models: A Nested Case–Control Study. PLoS Med. 2018, 15, e1002644. [Google Scholar] [CrossRef]
- Shi, M.; O’Brien, K.M.; Weinberg, C.R. Interactions between a Polygenic Risk Score and Non-Genetic Risk Factors in Young-Onset Breast Cancer. Sci. Rep. 2020, 10, 3242–3247. [Google Scholar] [CrossRef]
- Shieh, Y.; Fejerman, L.; Lott, P.C.; Marker, K.; Sawyer, S.D.; Hu, D.; Huntsman, S.; Torres, J.; Echeverry, M.; Bohórquez, M.E.; et al. A Polygenic Risk Score for Breast Cancer in US Latinas and Latin American Women. JNCI J. Natl. Cancer Inst. 2020, 112, 590–598. [Google Scholar] [CrossRef] [Green Version]
- Ho, W.-K.; Tan, M.-M.; Mavaddat, N.; Tai, M.-C.; Mariapun, S.; Li, J.; Ho, P.-J.; Dennis, J.; Tyrer, J.P.; Bolla, M.K.; et al. European Polygenic Risk Score for Prediction of Breast Cancer Shows Similar Performance in Asian Women. Nat. Commun. 2020, 11, 1–11. [Google Scholar] [CrossRef]
- Hsieh, Y.-C.; Tu, S.-H.; Su, C.-T.; Cho, E.-C.; Wu, C.-H.; Hsieh, M.-C.; Lin, S.-Y.; Liu, Y.-R.; Hung, C.-S.; Chiou, H.-Y. A Polygenic Risk Score for Breast Cancer Risk in a Taiwanese Population. Breast Cancer Res. Treat. 2017, 163, 131–138. [Google Scholar] [CrossRef]
- Wen, W.; Shu, X.; Guo, X.; Cai, Q.; Long, J.; Bolla, M.K.; Michailidou, K.; Dennis, J.; Wang, Q.; Gao, Y.-T.; et al. Prediction of Breast Cancer Risk Based on Common Genetic Variants in Women of East Asian Ancestry. Breast Cancer Res. 2016, 18, 124. [Google Scholar] [CrossRef] [Green Version]
- Chan, C.H.T.; Munusamy, P.; Loke, S.Y.; Koh, G.L.; Yang, A.Z.Y.; Law, H.Y.; Yoon, C.S.; Wong, C.Y.; Yong, W.S.; Wong, N.S.; et al. Evaluation of Three Polygenic Risk Score Models for the Prediction of Breast Cancer Risk in Singapore Chinese. Oncotarget 2018, 9, 12796–12804. [Google Scholar] [CrossRef] [Green Version]
- Coignet, M.v.; Zirpoli, G.R.; Roberts, M.R.; Khoury, T.; Bandera, E.v.; Zhu, Q.; Yao, S. Genetic Variations, Reproductive Aging, and Breast Cancer Risk in African American and European American Women: The Women’s Circle of Health Study. PLoS ONE 2017, 12, e0187205. [Google Scholar] [CrossRef] [Green Version]
- Wang, S.; Qian, F.; Zheng, Y.; Ogundiran, T.; Ojengbede, O.; Zheng, W.; Blot, W.; Nathanson, K.L.; Hennis, A.; Nemesure, B.; et al. Genetic Variants Demonstrating Flip-Flop Phenomenon and Breast Cancer Risk Prediction among Women of African Ancestry. Breast Cancer Res. Treat. 2018, 168, 703–712. [Google Scholar] [CrossRef]
- Wang, S.; Zheng, Y.; Ogundiran, T.O.; Ojengbede, O.; Zheng, W.; Nathanson, K.L.; Nemesure, B.; Ambs, S.; Olopade, O.I.; Huo, D. Association of Pancreatic Cancer Susceptibility Variants with Risk of Breast Cancer in Women of European and African Ancestry. Cancer Epidemiol. Biomark. Prev. 2018, 27, 116–118. [Google Scholar] [CrossRef] [Green Version]
- Siegel, R.L.; Miller, K.D.; Fuchs, H.E.; Jemal, A. Cancer Statistics, 2021. CA Cancer J. Clin. 2021, 71, 7–33. [Google Scholar] [CrossRef]
- Bancroft, E.K.; Raghallaigh, H.N.; Page, E.C.; Eeles, R.A. Updates in Prostate Cancer Research and Screening in Men at Genetically Higher Risk. Curr. Genet. Med. Rep. 2021, 9, 47–58. [Google Scholar] [CrossRef]
- Du, Z.; Hopp, H.; Ingles, S.A.; Huff, C.; Sheng, X.; Weaver, B.; Stern, M.; Hoffmann, T.J.; John, E.M.; van den Eeden, S.K.; et al. A Genome-wide Association Study of Prostate Cancer in Latinos. Int. J. Cancer 2020, 146, 1819–1826. [Google Scholar] [CrossRef]
- Harlemon, M.; Ajayi, O.; Kachambwa, P.; Kim, M.S.; Simonti, C.N.; Quiver, M.H.; Petersen, D.C.; Mittal, A.; Fernandez, P.W.; Hsing, A.W.; et al. A Custom Genotyping Array Reveals Population-Level Heterogeneity for the Genetic Risks of Prostate Cancer and Other Cancers in Africa. Cancer Res. 2020, 80, 2956–2966. [Google Scholar] [CrossRef]
- Du, Z.; Lubmawa, A.; Gundell, S.; Wan, P.; Nalukenge, C.; Muwanga, P.; Lutalo, M.; Nansereko, D.; Ndaruhutse, O.; Katuku, M.; et al. Genetic Risk of Prostate Cancer in Ugandan Men. Prostate 2018, 78, 370–376. [Google Scholar] [CrossRef]
- Bryc, K.; Durand, E.Y.; Macpherson, J.M.; Reich, D.; Mountain, J.L. The Genetic Ancestry of African Americans, Latinos, and European Americans across the United States. Am. J. Hum. Genet. 2015, 96, 37–53. [Google Scholar] [CrossRef] [Green Version]
- Zhang, M.; Wang, Z.; Obazee, O.; Jia, J.; Childs, E.J.; Hoskins, J.; Figlioli, G.; Mocci, E.; Collins, I.; Chung, C.C.; et al. Three New Pancreatic Cancer Susceptibility Signals Identified on Chromosomes 1q32.1, 5p15.33 and 8q24.21. Oncotarget 2016, 7, 66328–66343. [Google Scholar] [CrossRef] [Green Version]
- Klein, A.P.; Wolpin, B.M.; Risch, H.A.; Stolzenberg-Solomon, R.Z.; Mocci, E.; Zhang, M.; Canzian, F.; Childs, E.J.; Hoskins, J.W.; Jermusyk, A.; et al. Genome-Wide Meta-Analysis Identifies Five New Susceptibility Loci for Pancreatic Cancer. Nat. Commun. 2018, 9, 1–11. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Amundadottir, L.; Kraft, P.; Stolzenberg-Solomon, R.Z.; Fuchs, C.S.; Petersen, G.M.; Arslan, A.A.; Bueno-de-Mesquita, H.B.; Gross, M.; Helzlsouer, K.; Jacobs, E.J.; et al. Genome-Wide Association Study Identifies Variants in the ABO Locus Associated with Susceptibility to Pancreatic Cancer. Nat. Genet. 2009, 41, 986–990. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Petersen, G.M.; Amundadottir, L.; Fuchs, C.S.; Kraft, P.; Stolzenberg-Solomon, R.Z.; Jacobs, K.B.; Arslan, A.A.; Bueno-de-Mesquita, H.B.; Gallinger, S.; Gross, M.; et al. A Genome-Wide Association Study Identifies Pancreatic Cancer Susceptibility Loci on Chromosomes 13q22.1, 1q32.1 and 5p15.33. Nat. Genet. 2010, 42, 224–228. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wolpin, B.M.; Rizzato, C.; Kraft, P.; Kooperberg, C.; Petersen, G.M.; Wang, Z.; Arslan, A.A.; Beane-Freeman, L.; Bracci, P.M.; Buring, J.; et al. Genome-Wide Association Study Identifies Multiple Susceptibility Loci for Pancreatic Cancer. Nat. Genet. 2014, 46, 994–1000. [Google Scholar] [CrossRef] [Green Version]
- Childs, E.J.; Mocci, E.; Campa, D.; Bracci, P.M.; Gallinger, S.; Goggins, M.; Li, D.; Neale, R.E.; Olson, S.H.; Scelo, G.; et al. Common Variation at 2p13.3, 3q29, 7p13 and 17q25.1 Associated with Susceptibility to Pancreatic Cancer. Nat. Genet. 2015, 47, 911–916. [Google Scholar] [CrossRef] [PubMed]
- Nakatochi, M.; Lin, Y.; Ito, H.; Hara, K.; Kinoshita, F.; Kobayashi, Y.; Ishii, H.; Ozaka, M.; Sasaki, T.; Sasahira, N.; et al. Prediction Model for Pancreatic Cancer Risk in the General Japanese Population. PLoS ONE 2018, 13, e0203386. [Google Scholar] [CrossRef]
- Cust, A.E.; Drummond, M.; Kanetsky, P.A.; Goldstein, A.M.; Barrett, J.H.; MacGregor, S.; Law, M.H.; Iles, M.M.; Bui, M.; Hopper, J.L.; et al. Assessing the Incremental Contribution of Common Genomic Variants to Melanoma Risk Prediction in Two Population-Based Studies. J. Investig. Dermatol. 2018, 138, 2617–2624. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Olsen, C.M.; Carroll, H.J.; Whiteman, D.C. Familial Melanoma: A Meta-Analysis and Estimates of Attributable Fraction. Cancer Epidemiol. Biomark. Prev. 2010, 19, 65–73. [Google Scholar] [CrossRef] [Green Version]
- Cust, A.E.; Jenkins, M.A.; Goumas, C.; Armstrong, B.K.; Schmid, H.; Aitken, J.F.; Giles, G.G.; Kefford, R.F.; Hopper, J.L.; Mann, G.J. Early-Life Sun Exposure and Risk of Melanoma before Age 40 Years. Cancer Causes Control. 2011, 22, 885–897. [Google Scholar] [CrossRef]
- Gandini, S.; Sera, F.; Cattaruzza, M.S.; Pasquini, P.; Picconi, O.; Boyle, P.; Melchi, C.F. Meta-Analysis of Risk Factors for Cutaneous Melanoma: II. Sun Exposure. Eur. J. Cancer 2005, 41, 45–60. [Google Scholar] [CrossRef]
- Crowther-Swanepoel, D.; Broderick, P.; di Bernardo, M.C.; Dobbins, S.E.; Torres, M.; Mansouri, M.; Ruiz-Ponte, C.; Enjuanes, A.; Rosenquist, R.; Carracedo, A.; et al. Common Variants at 2q37.3, 8q24.21, 15q21.3 and 16q24.1 Influence Chronic Lymphocytic Leukemia Risk. Nat. Genet. 2010, 42, 132–136. [Google Scholar] [CrossRef] [Green Version]
- di Bernardo, M.C.; Crowther-Swanepoel, D.; Broderick, P.; Webb, E.; Sellick, G.; Wild, R.; Sullivan, K.; Vijayakrishnan, J.; Wang, Y.; Pittman, A.M.; et al. A Genome-Wide Association Study Identifies Six Susceptibility Loci for Chronic Lymphocytic Leukemia. Nat. Genet. 2008, 40, 1204–1210. [Google Scholar] [CrossRef] [PubMed]
- Berndt, S.I.; Skibola, C.F.; Joseph, V.; Camp, N.J.; Nieters, A.; Wang, Z.; Cozen, W.; Monnereau, A.; Wang, S.S.; Kelly, R.S.; et al. Genome-Wide Association Study Identifies Multiple Risk Loci for Chronic Lymphocytic Leukemia. Nat. Genet. 2013, 45, 868–876. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sava, G.P.; Speedy, H.E.; di Bernardo, M.C.; Dyer, M.J.S.; Holroyd, A.; Sunter, N.J.; Marr, H.; Mansouri, L.; Deaglio, S.; Karabon, L.; et al. Common Variation at 12q24.13 (OAS3) Influences Chronic Lymphocytic Leukemia Risk. Leukemia 2015, 29, 748–751. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Speedy, H.E.; di Bernardo, M.C.; Sava, G.P.; Dyer, M.J.S.; Holroyd, A.; Wang, Y.; Sunter, N.J.; Mansouri, L.; Juliusson, G.; Smedby, K.E.; et al. A Genome-Wide Association Study Identifies Multiple Susceptibility Loci for Chronic Lymphocytic Leukemia. Nat. Genet. 2014, 46, 56–60. [Google Scholar] [CrossRef] [PubMed]
- Berndt, S.I.; Camp, N.J.; Skibola, C.F.; Vijai, J.; Wang, Z.; Gu, J.; Nieters, A.; Kelly, R.S.; Smedby, K.E.; Monnereau, A.; et al. Meta-Analysis of Genome-Wide Association Studies Discovers Multiple Loci for Chronic Lymphocytic Leukemia. Nat. Commun. 2016, 7, 10933. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Law, P.J.; Berndt, S.I.; Speedy, H.E.; Camp, N.J.; Sava, G.P.; Skibola, C.F.; Holroyd, A.; Joseph, V.; Sunter, N.J.; Nieters, A.; et al. Genome-Wide Association Analysis Implicates Dysregulation of Immunity Genes in Chronic Lymphocytic Leukaemia. Nat. Commun. 2017, 8, 14175. [Google Scholar] [CrossRef]
- Slager, S.L.; Rabe, K.G.; Achenbach, S.J.; Vachon, C.M.; Goldin, L.R.; Strom, S.S.; Lanasa, M.C.; Spector, L.G.; Rassenti, L.Z.; Leis, J.F.; et al. Genome-Wide Association Study Identifies a Novel Susceptibility Locus at 6p21.3 among Familial CLL. Blood 2011, 117, 1911–1916. [Google Scholar] [CrossRef] [Green Version]
- Zavala, V.A.; Bracci, P.M.; Carethers, J.M.; Carvajal-Carmona, L.; Coggins, N.B.; Cruz-Correa, M.R.; Davis, M.; de Smith, A.J.; Dutil, J.; Figueiredo, J.C.; et al. Cancer Health Disparities in Racial/Ethnic Minorities in the United States. Br. J. Cancer 2021, 124, 315–332. [Google Scholar] [CrossRef]
- Davis, M.B. Genomics and Cancer Disparities: The Justice and Power of Inclusion. Cancer Discov. 2021, 11, 805–809. [Google Scholar] [CrossRef]
- Balogun, O.D.; Olopade, O.I. Addressing Health Disparities in Cancer with Genomics. Nat. Rev. Genet. 2021, 22, 621–622. [Google Scholar] [CrossRef]
- Fernandes, G.C.; Michelli, R.A.D.; Galvão, H.C.R.; Paula, A.E.; Pereira, R.; Andrade, C.E.; Felicio, P.S.; Souza, C.P.; Mendes, D.R.P.; Volc, S.; et al. Prevalence of BRCA1/BRCA2 Mutations in a Brazilian Population Sample at-Risk for Hereditary Breast Cancer and Characterization of Its Genetic Ancestry. Oncotarget 2016, 7, 80465–80481. [Google Scholar] [CrossRef] [Green Version]
- Leal, L.F.; de Paula, F.E.; de Marchi, P.; de Souza Viana, L.; Pinto, G.D.J.; Carlos, C.D.; Berardinelli, G.N.; Miziara, J.E.; da Silva, C.M.; Silva, E.C.A.; et al. Mutational Profile of Brazilian Lung Adenocarcinoma Unveils Association of EGFR Mutations with High Asian Ancestry and Independent Prognostic Role of KRAS Mutations. Sci. Rep. 2019, 9, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Durães, R.O.; Berardinelli, G.N.; da Costa, A.M.; Scapulatempo-Neto, C.; Pereira, R.; Oliveira, M.A.; Guimarães, D.P.; Reis, R.M. Role of Genetic Ancestry in 1,002 Brazilian Colorectal Cancer Patients From Barretos Cancer Hospital. Front. Oncol. 2020, 10, 145. [Google Scholar] [CrossRef] [PubMed] [Green Version]
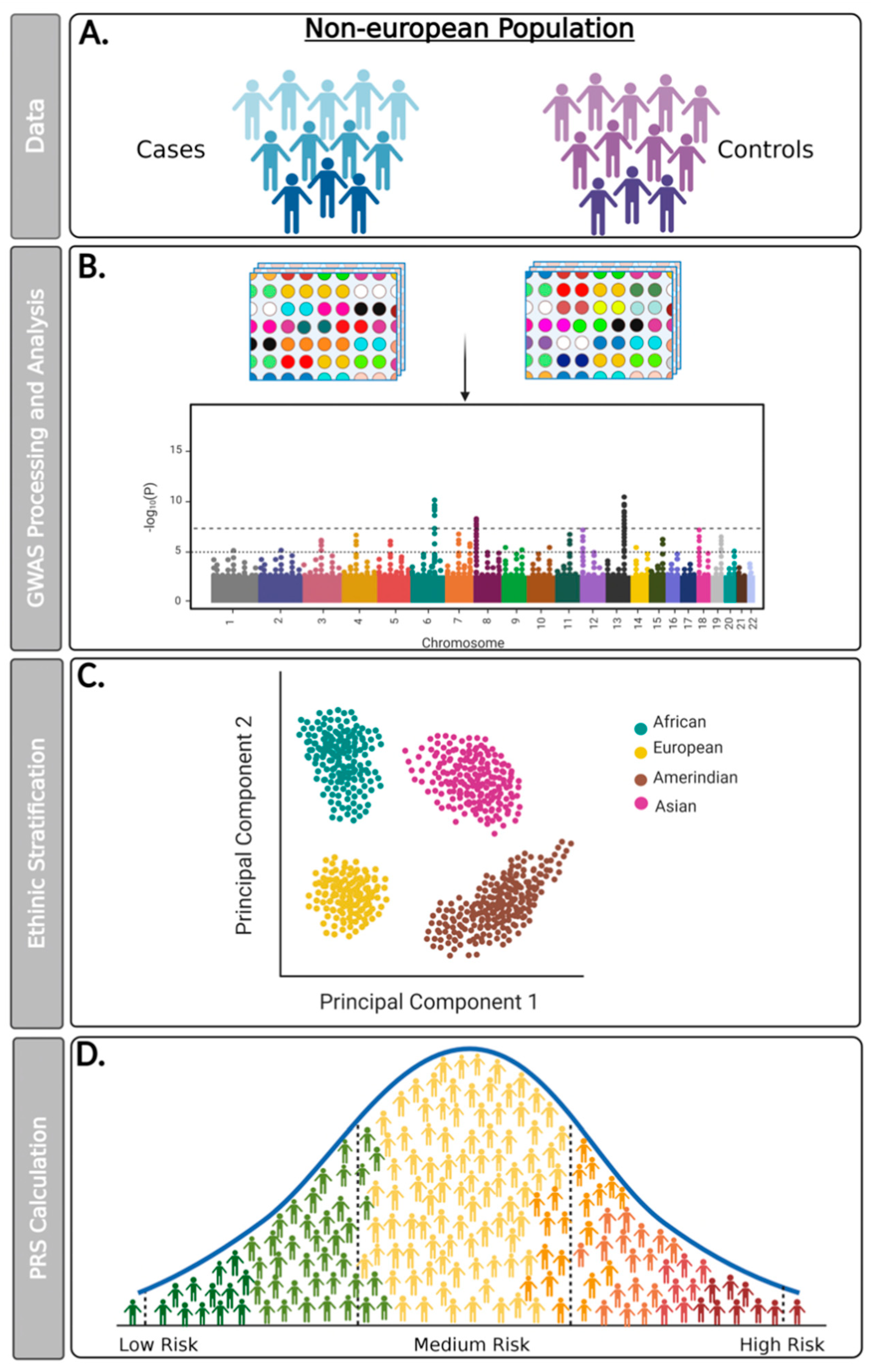
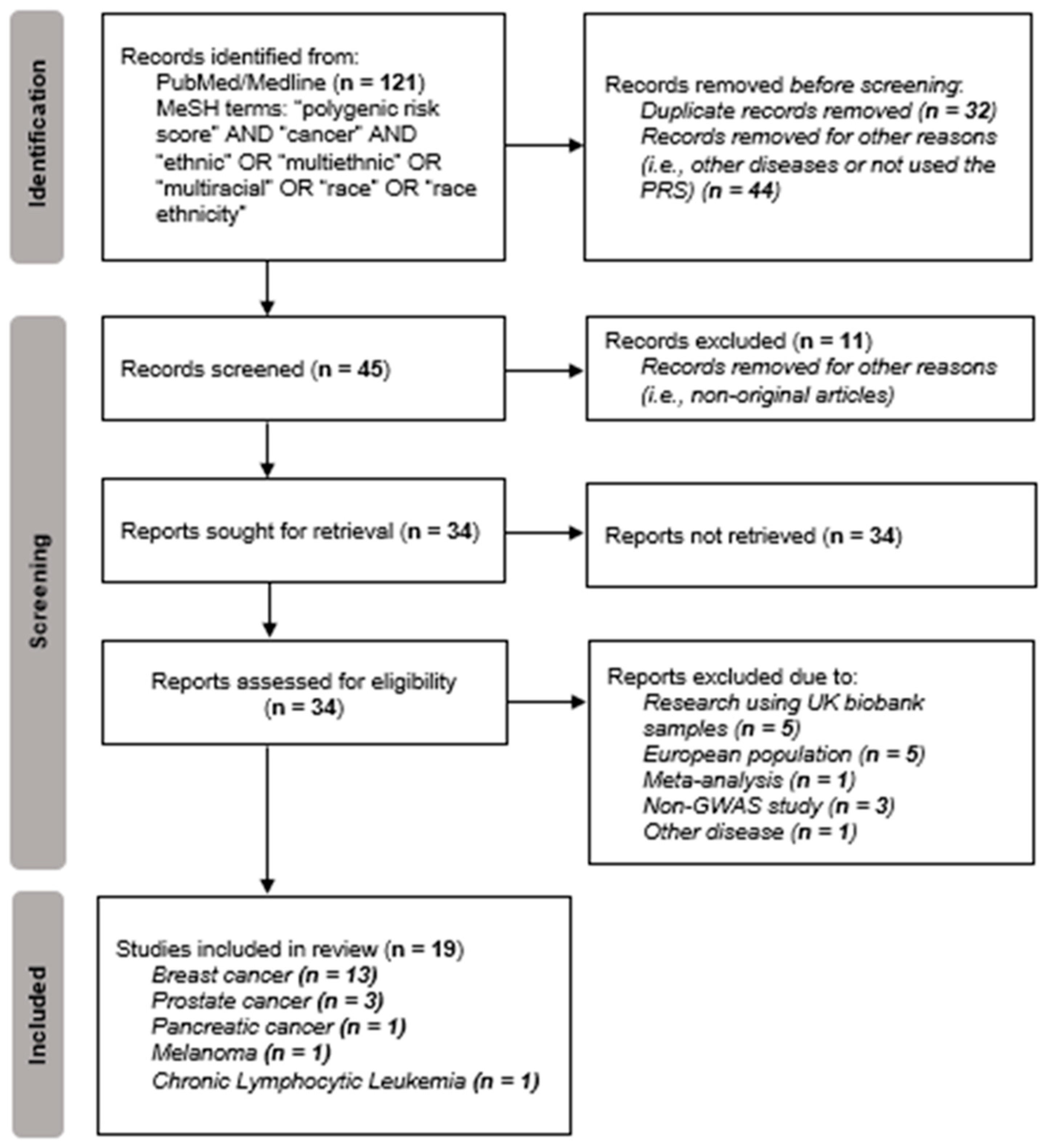
| Autor, Year | Phenotype | Population/ Ethnicity | Subjects (n) | SNPs (n) | Significant SNPs (n) | Main Finding * |
|---|---|---|---|---|---|---|
| Harlemon et al., 2020 [47] | Prostate Cancer | African | Control: 403 Case: 399 | 139-SNP | rs183373024 (PRNCR1) rs1447295 (CASC8) | Not replicated |
| Du et al., 2018 [48] | Prostate Cancer | African | Control: 485 Case: 571 | 97-SNP | rs72725854 (none) | Replicated |
| Du et al., 2020 [46] | Prostate Cancer | Latin | Control: 5293 Case: 2820 | 176-SNP | - | Replicated |
| Autor, Year | Phenotype | Population/ Ethnicity | Subjects (n) | SNPs (n) | Significant SNPs (n) | Main Finding |
|---|---|---|---|---|---|---|
| Nakatochi et al., 2018 [56] | Pancreatic Cancer | Asian | Control: 664 Case: 664 | 61-SNP | rs13303010 (NOC2L) rs12615966 (none) rs657152 (ABO) rs9564966 (none) rs16986825 (ZNRF3) | Replicated |
| Cust et al., 2018 [57] | Melanoma | Oceania and Europe | Australian: 1035 United Kingdom: 1406 | 21-SNP | - | Replicated |
| Kleinstern et al., 2021 [21] | Chronic Lymphocytic Leukemia | African-American | Control: 235 Case: 173 | 41-SNP | - | Not replicated |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Junior, H.L.R.; Novaes, L.A.C.; Datorre, J.G.; Moreno, D.A.; Reis, R.M. Role of Polygenic Risk Score in Cancer Precision Medicine of Non-European Populations: A Systematic Review. Curr. Oncol. 2022, 29, 5517-5530. https://doi.org/10.3390/curroncol29080436
Junior HLR, Novaes LAC, Datorre JG, Moreno DA, Reis RM. Role of Polygenic Risk Score in Cancer Precision Medicine of Non-European Populations: A Systematic Review. Current Oncology. 2022; 29(8):5517-5530. https://doi.org/10.3390/curroncol29080436
Chicago/Turabian StyleJunior, Howard Lopes Ribeiro, Lázaro Antônio Campanha Novaes, José Guilherme Datorre, Daniel Antunes Moreno, and Rui Manuel Reis. 2022. "Role of Polygenic Risk Score in Cancer Precision Medicine of Non-European Populations: A Systematic Review" Current Oncology 29, no. 8: 5517-5530. https://doi.org/10.3390/curroncol29080436
APA StyleJunior, H. L. R., Novaes, L. A. C., Datorre, J. G., Moreno, D. A., & Reis, R. M. (2022). Role of Polygenic Risk Score in Cancer Precision Medicine of Non-European Populations: A Systematic Review. Current Oncology, 29(8), 5517-5530. https://doi.org/10.3390/curroncol29080436






