Expression and Signaling Pathways of Nerve Growth Factor (NGF) and Pro-NGF in Breast Cancer: A Systematic Review
Abstract
1. Introduction
2. Breast Cancer: An Overview
2.1. Classification
2.1.1. Histological Type
2.1.2. Grading
2.1.3. Immunophenotype
2.1.4. TNM Classification
2.1.5. Molecular Subtype
2.2. Risk Factors
2.3. Diagnosis
2.4. Treatments and Prognosis
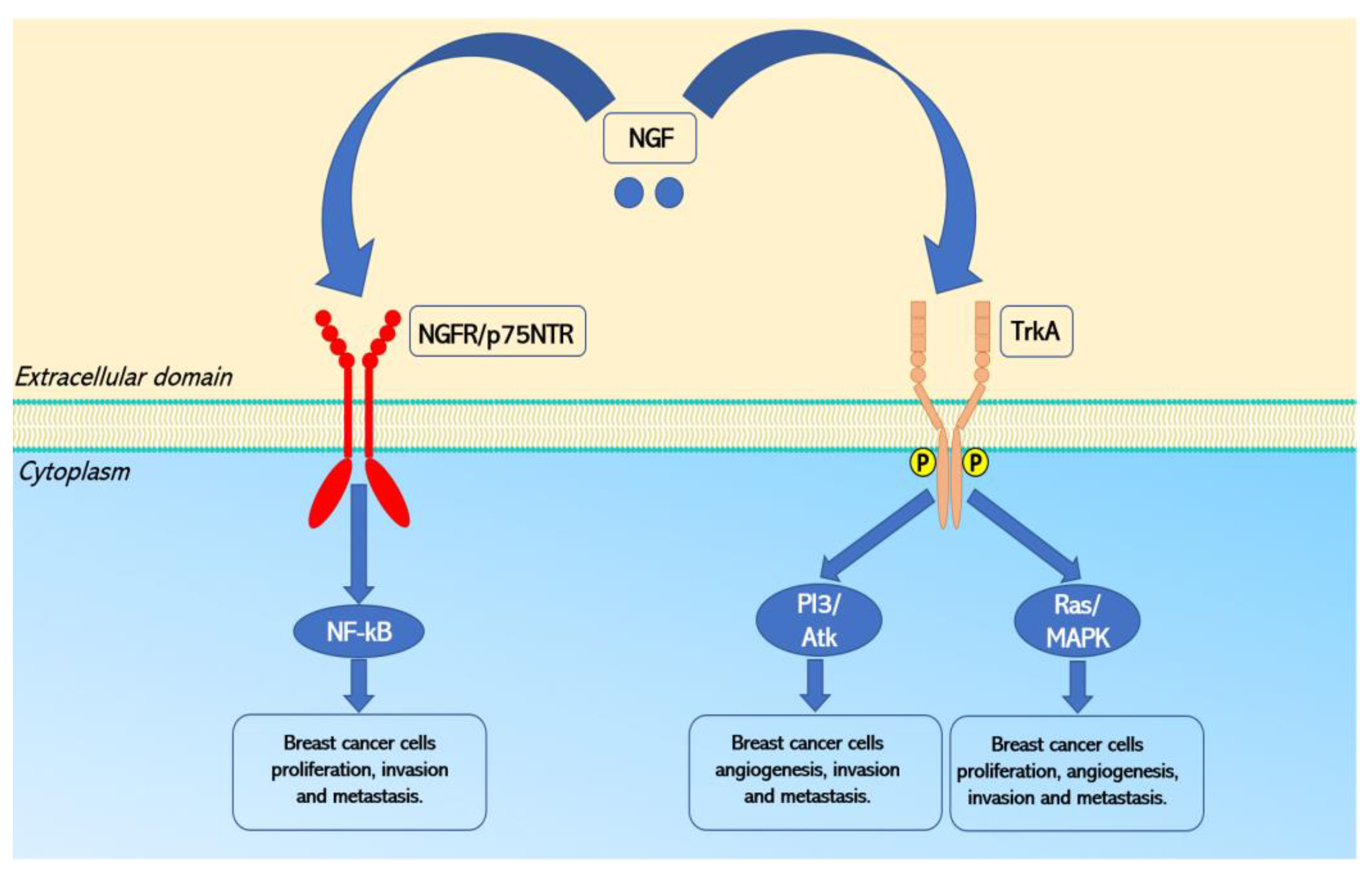
3. NGF Signaling Pathways
4. Methods
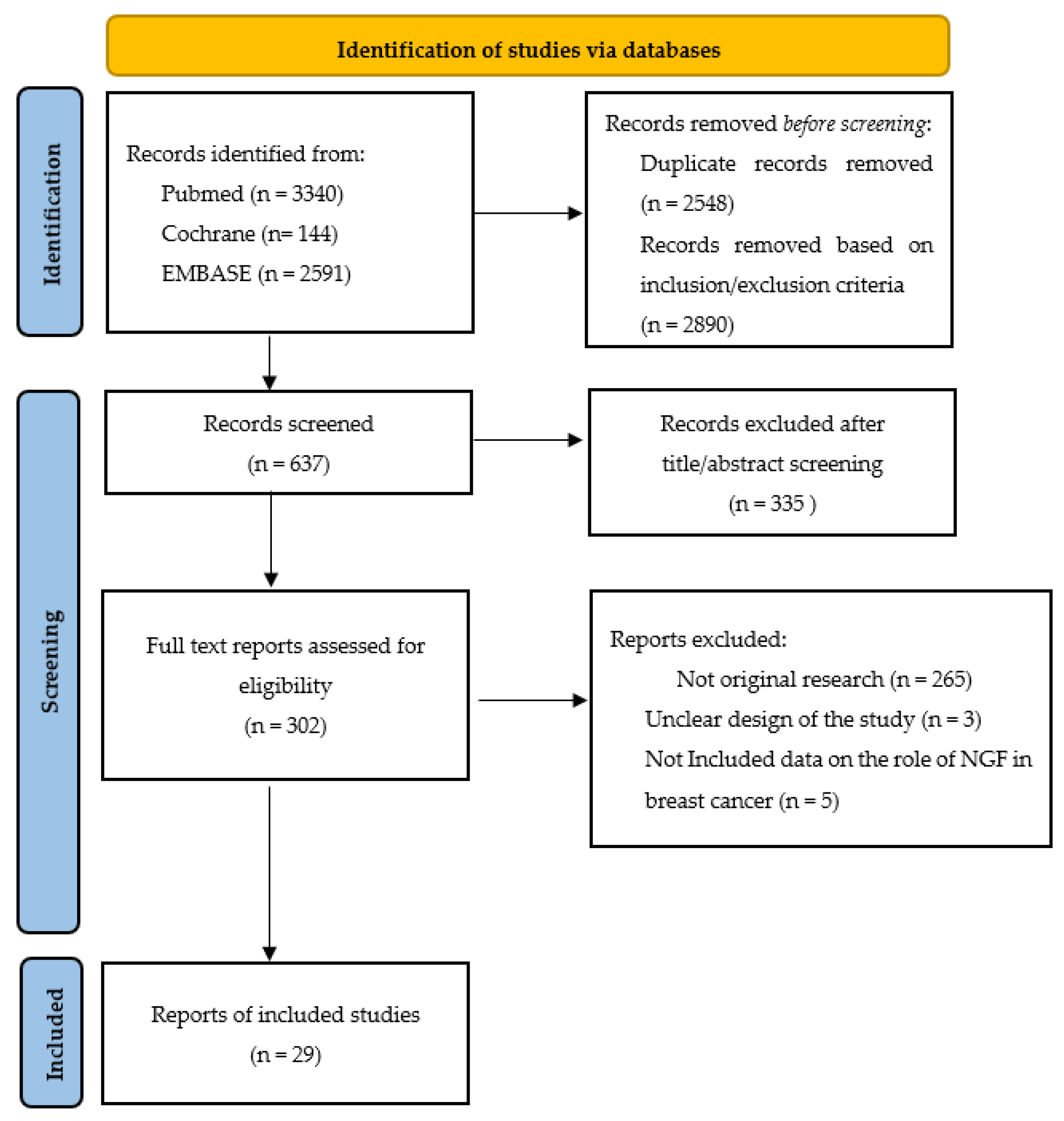
4.1. Study Search
4.2. Exclusion and Inclusion Criteria
5. Results
5.1. Included Studies
5.2. NGF and Its Receptors Expression in Breast Cancer
5.3. Mitogenesis of Breast Cancer Cells
5.4. Anti-Apoptosis and Survival of Breast Cancer Cells
5.5. Angiogenesis of Breast Cancer
5.6. Breast Cancer Invasion and Metastasis
5.7. NGF and Its Receptors as Diagnostic Markers for Breast Cancer
5.8. NGF and Its Receptors as Prognostic Markers for Breast Cancer
5.9. NGF Signaling Pathways as A Therapeutic Target for Breast Cancer
6. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Lukasiewicz, S.; Czeczelewski, M.; Forma, A.; Baj, J.; Sitarz, R.; Stanislawek, A. Breast Cancer-Epidemiology, Risk Factors, Classification, Prognostic Markers, and Current Treatment Strategies-An Updated Review. Cancers 2021, 13, 4287. [Google Scholar] [CrossRef]
- Sung, H.; Ferlay, J.; Siegel, R.L.; Laversanne, M.; Soerjomataram, I.; Jemal, A.; Bray, F. Global Cancer Statistics 2020: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA Cancer J. Clin. 2021, 71, 209–249. [Google Scholar] [CrossRef] [PubMed]
- Akram, M.; Iqbal, M.; Daniyal, M.; Khan, A.U. Awareness and current knowledge of breast cancer. Biol. Res. 2017, 50, 33. [Google Scholar] [CrossRef] [PubMed]
- Hortobagyi, G.N.; de la Garza Salazar, J.; Pritchard, K.; Amadori, D.; Haidinger, R.; Hudis, C.A.; Khaled, H.; Liu, M.C.; Martin, M.; Namer, M.; et al. The global breast cancer burden: Variations in epidemiology and survival. Clin. Breast Cancer 2005, 6, 391–401. [Google Scholar] [CrossRef] [PubMed]
- Harless, W.; Qiu, Y. Cancer: A medical emergency. Med. Hypotheses 2006, 67, 1054–1059. [Google Scholar] [CrossRef]
- Chiesi, F.; Vizza, D.; Valente, M.; Bruno, R.; Lau, C.; Campagna, M.R.; Lo Iacono, M.; Bruno, F. Positive personal resources and psychological distress during the COVID-19 pandemic: Resilience, optimism, hope, courage, trait mindfulness, and self-efficacy in breast cancer patients and survivors. Support. Care Cancer 2022, 30, 7005–7014. [Google Scholar] [CrossRef] [PubMed]
- Chirico, A.; Vizza, D.; Valente, M.; Iacono, M.L.; Campagna, M.R.; Palombi, T.; Alivernini, F.; Lucidi, F.; Bruno, F. Assessing the fear of recurrence using the Cancer Worry Scale in a sample of Italian breast cancer survivors. Support. Care Cancer 2022, 30, 2829–2837. [Google Scholar] [CrossRef] [PubMed]
- Bradshaw, R.A.; Pundavela, J.; Biarc, J.; Chalkley, R.J.; Burlingame, A.L.; Hondermarck, H. NGF and ProNGF: Regulation of neuronal and neoplastic responses through receptor signaling. Adv. Biol. Regul. 2015, 58, 16–27. [Google Scholar] [CrossRef] [PubMed]
- Dolle, L.; Adriaenssens, E.; El Yazidi-Belkoura, I.; Le Bourhis, X.; Nurcombe, V.; Hondermarck, H. Nerve growth factor receptors and signaling in breast cancer. Curr. Cancer Drug Targets 2004, 4, 463–470. [Google Scholar] [CrossRef]
- Molloy, N.H.; Read, D.E.; Gorman, A.M. Nerve growth factor in cancer cell death and survival. Cancers 2011, 3, 510–530. [Google Scholar] [CrossRef] [PubMed]
- Hondermarck, H. Neurotrophins and their receptors in breast cancer. Cytokine Growth Factor Rev. 2012, 23, 357–365. [Google Scholar] [CrossRef] [PubMed]
- Demir, I.E.; Tieftrunk, E.; Schorn, S.; Friess, H.; Ceyhan, G.O. Nerve growth factor & TrkA as novel therapeutic targets in cancer. Biochim. Biophys. Acta 2016, 1866, 37–50. [Google Scholar] [CrossRef] [PubMed]
- Griffin, N.; Faulkner, S.; Jobling, P.; Hondermarck, H. Targeting neurotrophin signaling in cancer: The renaissance. Pharmacol. Res. 2018, 135, 12–17. [Google Scholar] [CrossRef] [PubMed]
- Islam, T.; Majumder, M.; Bidkar, A.; Ghosh, S.S.; Mukhopadhyay, R.; Utkin, Y.; Mukherjee, A.K. Nerve growth factor from Indian Russell’s viper venom (RVV-NGFa) shows high affinity binding to TrkA receptor expressed in breast cancer cells: Application of fluorescence labeled RVV-NGFa in the clinical diagnosis of breast cancer. Biochimie 2020, 176, 31–44. [Google Scholar] [CrossRef] [PubMed]
- Wu, R.; Li, K.; Yuan, M.; Luo, K.Q. Nerve growth factor receptor increases the tumor growth and metastatic potential of triple-negative breast cancer cells. Oncogene 2021, 40, 2165–2181. [Google Scholar] [CrossRef]
- Noh, S.J.; Bae, J.S.; Jamiyandorj, U.; Park, H.S.; Kwon, K.S.; Jung, S.H.; Youn, H.J.; Lee, H.; Park, B.H.; Chung, M.J.; et al. Expression of nerve growth factor and heme oxygenase-1 predict poor survival of breast carcinoma patients. BMC Cancer 2013, 13, 516. [Google Scholar] [CrossRef]
- Chakravarthy, R.; Mnich, K.; Gorman, A.M. Nerve growth factor (NGF)-mediated regulation of p75(NTR) expression contributes to chemotherapeutic resistance in triple negative breast cancer cells. Biochem. Biophys. Res. Commun. 2016, 478, 1541–1547. [Google Scholar] [CrossRef]
- Demont, Y.; Corbet, C.; Page, A.; Ataman-Onal, Y.; Choquet-Kastylevsky, G.; Fliniaux, I.; Le Bourhis, X.; Toillon, R.A.; Bradshaw, R.A.; Hondermarck, H. Pro-nerve growth factor induces autocrine stimulation of breast cancer cell invasion through tropomyosin-related kinase A (TrkA) and sortilin protein. J. Biol. Chem. 2012, 287, 1923–1931. [Google Scholar] [CrossRef] [PubMed]
- Leveque, R.; Corbet, C.; Aubert, L.; Guilbert, M.; Lagadec, C.; Adriaenssens, E.; Duval, J.; Finetti, P.; Birnbaum, D.; Magne, N.; et al. ProNGF increases breast tumor aggressiveness through functional association of TrkA with EphA2. Cancer Lett. 2019, 449, 196–206. [Google Scholar] [CrossRef]
- Tsang, J.Y.S.; Tse, G.M. Molecular Classification of Breast Cancer. Adv. Anat. Pathol. 2020, 27, 27–35. [Google Scholar] [CrossRef]
- Malhotra, G.K.; Zhao, X.; Band, H.; Band, V. Histological, molecular and functional subtypes of breast cancers. Cancer Biol. Ther. 2010, 10, 955–960. [Google Scholar] [CrossRef]
- Association of Directors of Anatomic and Surgical Pathology. Recommendations for the reporting of breast carcinoma. Hum. Pathol. 1996, 27, 220–224. [Google Scholar] [CrossRef]
- Li, C.I.; Uribe, D.J.; Daling, J.R. Clinical characteristics of different histologic types of breast cancer. Br. J. Cancer 2005, 93, 1046–1052. [Google Scholar] [CrossRef] [PubMed]
- Lester, S.C.; Bose, S.; Chen, Y.Y.; Connolly, J.L.; de Baca, M.E.; Fitzgibbons, P.L.; Hayes, D.F.; Kleer, C.; O’Malley, F.P.; Page, D.L.; et al. Protocol for the examination of specimens from patients with invasive carcinoma of the breast. Arch. Pathol. Lab. Med. 2009, 133, 1515–1538. [Google Scholar] [CrossRef]
- Elston, E.W.; Ellis, I.O. Method for grading breast cancer. J. Clin. Pathol. 1993, 46, 189–190. [Google Scholar] [CrossRef]
- Bansal, C.; Pujani, M.; Sharma, K.L.; Srivastava, A.N.; Singh, U.S. Grading systems in the cytological diagnosis of breast cancer: A review. J. Cancer Res. Ther. 2014, 10, 839–845. [Google Scholar] [CrossRef]
- Ivshina, A.V.; George, J.; Senko, O.; Mow, B.; Putti, T.C.; Smeds, J.; Lindahl, T.; Pawitan, Y.; Hall, P.; Nordgren, H.; et al. Genetic reclassification of histologic grade delineates new clinical subtypes of breast cancer. Cancer Res. 2006, 66, 10292–10301. [Google Scholar] [CrossRef]
- Waks, A.G.; Winer, E.P. Breast Cancer Treatment: A Review. JAMA 2019, 321, 288–300. [Google Scholar] [CrossRef]
- Sawaki, M.; Shien, T.; Iwata, H. TNM classification of malignant tumors (Breast Cancer Study Group). Jpn. J. Clin. Oncol. 2019, 49, 228–231. [Google Scholar] [CrossRef]
- Fan, C.; Oh, D.S.; Wessels, L.; Weigelt, B.; Nuyten, D.S.; Nobel, A.B.; van’t Veer, L.J.; Perou, C.M. Concordance among gene-expression-based predictors for breast cancer. N. Engl. J. Med. 2006, 355, 560–569. [Google Scholar] [CrossRef]
- Hu, Z.; Fan, C.; Oh, D.S.; Marron, J.S.; He, X.; Qaqish, B.F.; Livasy, C.; Carey, L.A.; Reynolds, E.; Dressler, L.; et al. The molecular portraits of breast tumors are conserved across microarray platforms. BMC Genom. 2006, 7, 96. [Google Scholar] [CrossRef] [PubMed]
- Hashmi, A.A.; Aijaz, S.; Khan, S.M.; Mahboob, R.; Irfan, M.; Zafar, N.I.; Nisar, M.; Siddiqui, M.; Edhi, M.M.; Faridi, N.; et al. Prognostic parameters of luminal A and luminal B intrinsic breast cancer subtypes of Pakistani patients. World J. Surg. Oncol. 2018, 16, 1. [Google Scholar] [CrossRef] [PubMed]
- Teschendorff, A.E.; Miremadi, A.; Pinder, S.E.; Ellis, I.O.; Caldas, C. An immune response gene expression module identifies a good prognosis subtype in estrogen receptor negative breast cancer. Genome Biol. 2007, 8, R157. [Google Scholar] [CrossRef] [PubMed]
- Bertucci, F.; Finetti, P.; Cervera, N.; Esterni, B.; Hermitte, F.; Viens, P.; Birnbaum, D. How basal are triple-negative breast cancers? Int. J. Cancer 2008, 123, 236–240. [Google Scholar] [CrossRef] [PubMed]
- Kreike, B.; van Kouwenhove, M.; Horlings, H.; Weigelt, B.; Peterse, H.; Bartelink, H.; van de Vijver, M.J. Gene expression profiling and histopathological characterization of triple-negative/basal-like breast carcinomas. Breast Cancer Res. 2007, 9, R65. [Google Scholar] [CrossRef]
- Dias, K.; Dvorkin-Gheva, A.; Hallett, R.M.; Wu, Y.; Hassell, J.; Pond, G.R.; Levine, M.; Whelan, T.; Bane, A.L. Claudin-Low Breast Cancer; Clinical & Pathological Characteristics. PLoS ONE 2017, 12, e0168669. [Google Scholar] [CrossRef]
- Schettini, F.; Pascual, T.; Conte, B.; Chic, N.; Braso-Maristany, F.; Galvan, P.; Martinez, O.; Adamo, B.; Vidal, M.; Munoz, M.; et al. HER2-enriched subtype and pathological complete response in HER2-positive breast cancer: A systematic review and meta-analysis. Cancer Treat. Rev. 2020, 84, 101965. [Google Scholar] [CrossRef]
- Momenimovahed, Z.; Salehiniya, H. Epidemiological characteristics of and risk factors for breast cancer in the world. Breast Cancer 2019, 11, 151–164. [Google Scholar] [CrossRef]
- He, Z.; Chen, Z.; Tan, M.; Elingarami, S.; Liu, Y.; Li, T.; Deng, Y.; He, N.; Li, S.; Fu, J.; et al. A review on methods for diagnosis of breast cancer cells and tissues. Cell Prolif. 2020, 53, e12822. [Google Scholar] [CrossRef]
- Abdelrahman, L.; Al Ghamdi, M.; Collado-Mesa, F.; Abdel-Mottaleb, M. Convolutional neural networks for breast cancer detection in mammography: A survey. Comput. Biol. Med. 2021, 131, 104248. [Google Scholar] [CrossRef]
- Li, J.; Guan, X.; Fan, Z.; Ching, L.M.; Li, Y.; Wang, X.; Cao, W.M.; Liu, D.X. Non-Invasive Biomarkers for Early Detection of Breast Cancer. Cancers 2020, 12, 767. [Google Scholar] [CrossRef] [PubMed]
- BahadurSingh, S.; Maharaj, R.; Harnarayan, P.; Cawich, S.O.; Yearwood, M.; Naraynsingh, V. Mammographic screening: Is it relevant to developing countries? Curr. Med. Res. Pract. 2014, 4, 168–170. [Google Scholar] [CrossRef]
- Lumachi, F.; Luisetto, G.; Basso, S.M.; Basso, U.; Brunello, A.; Camozzi, V. Endocrine therapy of breast cancer. Curr. Med. Chem. 2011, 18, 513–522. [Google Scholar] [CrossRef] [PubMed]
- Tremont, A.; Lu, J.; Cole, J.T. Endocrine Therapy for Early Breast Cancer: Updated Review. Ochsner J. 2017, 17, 405–411. [Google Scholar]
- Tarantino, P.; Morganti, S.; Curigliano, G. Biologic therapy for advanced breast cancer: Recent advances and future directions. Expert Opin. Biol. Ther. 2020, 20, 1009–1024. [Google Scholar] [CrossRef]
- Ahmad, A. Breast Cancer Statistics: Recent Trends. Adv. Exp. Med. Biol. 2019, 1152, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Cao, J.; Zhang, M.; Wang, B.; Zhang, L.; Fang, M.; Zhou, F. Chemoresistance and Metastasis in Breast Cancer Molecular Mechanisms and Novel Clinical Strategies. Front. Oncol. 2021, 11, 658552. [Google Scholar] [CrossRef]
- Cipriano, E.; Mesquita, A. Emerging Therapeutic Drugs in Metastatic Triple-Negative Breast Cancer. Breast Cancer 2021, 15, 11782234211002491. [Google Scholar] [CrossRef]
- Harbeck, N.; Penault-Llorca, F.; Cortes, J.; Gnant, M.; Houssami, N.; Poortmans, P.; Ruddy, K.; Tsang, J.; Cardoso, F. Breast cancer. Nat. Rev. Dis. Primers 2019, 5, 66. [Google Scholar] [CrossRef]
- American Joint Committee on Cancer; Amin, M.B.; Edge, S.B. AJCC Cancer Staging Manual, 8th ed.; Springer: Cham, Switzerland, 2017; Volume xvii, 1024p. [Google Scholar]
- Nielsen, T.O.; Leung, S.C.Y.; Rimm, D.L.; Dodson, A.; Acs, B.; Badve, S.; Denkert, C.; Ellis, M.J.; Fineberg, S.; Flowers, M.; et al. Assessment of Ki67 in Breast Cancer: Updated Recommendations From the International Ki67 in Breast Cancer Working Group. J. Natl. Cancer Inst. 2021, 113, 808–819. [Google Scholar] [CrossRef]
- Taneja, P.; Maglic, D.; Kai, F.; Zhu, S.; Kendig, R.D.; Fry, E.A.; Inoue, K. Classical and Novel Prognostic Markers for Breast Cancer and their Clinical Significance. Clin. Med. Insights Oncol. 2010, 4, 15–34. [Google Scholar] [CrossRef] [PubMed]
- Skaper, S.D. The neurotrophin family of neurotrophic factors: An overview. Methods Mol. Biol. 2012, 846, 1–12. [Google Scholar] [CrossRef]
- Bibel, M.; Hoppe, E.; Barde, Y.A. Biochemical and functional interactions between the neurotrophin receptors trk and p75NTR. EMBO J. 1999, 18, 616–622. [Google Scholar] [CrossRef]
- Lu, B.; Pang, P.T.; Woo, N.H. The yin and yang of neurotrophin action. Nat. Rev. Neurosci. 2005, 6, 603–614. [Google Scholar] [CrossRef] [PubMed]
- Yamamoto, M.; Sobue, G.; Yamamoto, K.; Terao, S.; Mitsuma, T. Expression of mRNAs for neurotrophic factors (NGF, BDNF, NT-3, and GDNF) and their receptors (p75NGFR, trkA, trkB, and trkC) in the adult human peripheral nervous system and nonneural tissues. Neurochem. Res. 1996, 21, 929–938. [Google Scholar] [CrossRef] [PubMed]
- Ichim, G.; Tauszig-Delamasure, S.; Mehlen, P. Neurotrophins and cell death. Exp. Cell Res. 2012, 318, 1221–1228. [Google Scholar] [CrossRef] [PubMed]
- Reichardt, L.F. Neurotrophin-regulated signalling pathways. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2006, 361, 1545–1564. [Google Scholar] [CrossRef]
- Levi-Montalcini, R.; Hamburger, V. Selective growth stimulating effects of mouse sarcoma on the sensory and sympathetic nervous system of the chick embryo. J. Exp. Zool. 1951, 116, 321–361. [Google Scholar] [CrossRef]
- Cohen, S.; Levi-Montalcini, R.; Hamburger, V. A Nerve Growth-Stimulating Factor Isolated from Sarcom as 37 and 180. Proc. Natl. Acad. Sci. USA 1954, 40, 1014–1018. [Google Scholar] [CrossRef]
- Dawbarn, D.; Allen, S.J. Neurotrophins and neurodegeneration. Neuropathol. Appl. Neurobiol. 2003, 29, 211–230. [Google Scholar] [CrossRef]
- La Mendola, D. Nerve growth factor catches copper in neuronal inning. Neural Regen. Res. 2020, 15, 665–666. [Google Scholar] [CrossRef] [PubMed]
- Tessarollo, L. Pleiotropic functions of neurotrophins in development. Cytokine Growth Factor Rev. 1998, 9, 125–137. [Google Scholar] [CrossRef]
- Aloe, L.; Skaper, S.D.; Leon, A.; Levi-Montalcini, R. Nerve growth factor and autoimmune diseases. Autoimmunity 1994, 19, 141–150. [Google Scholar] [CrossRef] [PubMed]
- Chaldakov, G.N.; Fiore, M.; Stankulov, I.S.; Manni, L.; Hristova, M.G.; Antonelli, A.; Ghenev, P.I.; Aloe, L. Neurotrophin presence in human coronary atherosclerosis and metabolic syndrome: A role for NGF and BDNF in cardiovascular disease? Prog. Brain Res. 2004, 146, 279–289. [Google Scholar] [CrossRef]
- Fujimura, H.; Altar, C.A.; Chen, R.; Nakamura, T.; Nakahashi, T.; Kambayashi, J.; Sun, B.; Tandon, N.N. Brain-derived neurotrophic factor is stored in human platelets and released by agonist stimulation. Thromb. Haemost. 2002, 87, 728–734. [Google Scholar] [CrossRef]
- Nakahashi, T.; Fujimura, H.; Altar, C.A.; Li, J.; Kambayashi, J.; Tandon, N.N.; Sun, B. Vascular endothelial cells synthesize and secrete brain-derived neurotrophic factor. FEBS Lett. 2000, 470, 113–117. [Google Scholar] [CrossRef]
- Martin-Zanca, D.; Hughes, S.H.; Barbacid, M. A human oncogene formed by the fusion of truncated tropomyosin and protein tyrosine kinase sequences. Nature 1986, 319, 743–748. [Google Scholar] [CrossRef]
- Hsiao, S.J.; Zehir, A.; Sireci, A.N.; Aisner, D.L. Detection of Tumor NTRK Gene Fusions to Identify Patients Who May Benefit from Tyrosine Kinase (TRK) Inhibitor Therapy. J. Mol. Diagn. 2019, 21, 553–571. [Google Scholar] [CrossRef]
- Kyker-Snowman, K.; Hughes, R.M.; Yankaskas, C.L.; Cravero, K.; Karthikeyan, S.; Button, B.; Waters, I.; Rosen, D.M.; Dennison, L.; Hunter, N.; et al. TrkA overexpression in non-tumorigenic human breast cell lines confers oncogenic and metastatic properties. Breast Cancer Res. Treat. 2020, 179, 631–642. [Google Scholar] [CrossRef]
- Francke, U.; de Martinville, B.; Coussens, L.; Ullrich, A. The human gene for the beta subunit of nerve growth factor is located on the proximal short arm of chromosome 1. Science 1983, 222, 1248–1251. [Google Scholar] [CrossRef]
- Barker, P.A. p75NTR is positively promiscuous: Novel partners and new insights. Neuron 2004, 42, 529–533. [Google Scholar] [CrossRef] [PubMed]
- Krygier, S.; Djakiew, D. Molecular characterization of the loss of p75(NTR) expression in human prostate tumor cells. Mol. Carcinog. 2001, 31, 46–55. [Google Scholar] [CrossRef] [PubMed]
- Levy, Y.S.; Gilgun-Sherki, Y.; Melamed, E.; Offen, D. Therapeutic potential of neurotrophic factors in neurodegenerative diseases. BioDrugs 2005, 19, 97–127. [Google Scholar] [CrossRef] [PubMed]
- Moscatelli, I.; Pierantozzi, E.; Camaioni, A.; Siracusa, G.; Campagnolo, L. p75 neurotrophin receptor is involved in proliferation of undifferentiated mouse embryonic stem cells. Exp. Cell Res. 2009, 315, 3220–3232. [Google Scholar] [CrossRef]
- Roux, P.P.; Barker, P.A. Neurotrophin signaling through the p75 neurotrophin receptor. Prog. Neurobiol. 2002, 67, 203–233. [Google Scholar] [CrossRef]
- Chao, M.V. The p75 neurotrophin receptor. J. Neurobiol. 1994, 25, 1373–1385. [Google Scholar] [CrossRef]
- Nykjaer, A.; Lee, R.; Teng, K.K.; Jansen, P.; Madsen, P.; Nielsen, M.S.; Jacobsen, C.; Kliemannel, M.; Schwarz, E.; Willnow, T.E.; et al. Sortilin is essential for proNGF-induced neuronal cell death. Nature 2004, 427, 843–848. [Google Scholar] [CrossRef]
- Nykjaer, A.; Willnow, T.E. Sortilin: A receptor to regulate neuronal viability and function. Trends Neurosci. 2012, 35, 261–270. [Google Scholar] [CrossRef]
- Clewes, O.; Fahey, M.S.; Tyler, S.J.; Watson, J.J.; Seok, H.; Catania, C.; Cho, K.; Dawbarn, D.; Allen, S.J. Human ProNGF: Biological effects and binding profiles at TrkA, P75NTR and sortilin. J. Neurochem. 2008, 107, 1124–1135. [Google Scholar] [CrossRef]
- PRISMA. Available online: https://www.prisma-statement.org (accessed on 20 June 2022).
- Adriaenssens, E.; Vanhecke, E.; Saule, P.; Mougel, A.; Page, A.; Romon, R.; Nurcombe, V.; Le Bourhis, X.; Hondermarck, H. Nerve growth factor is a potential therapeutic target in breast cancer. Cancer Res. 2008, 68, 346–351. [Google Scholar] [CrossRef]
- Aragona, M.; Panetta, S.; Silipigni, A.M.; Romeo, D.L.; Pastura, G.; Mesiti, M.; Cascinu, S.; La Torre, F. Nerve growth factor receptor immunoreactivity in breast cancer patients. Cancer Investig. 2001, 19, 692–697. [Google Scholar] [CrossRef]
- Bashir, N.; Ishfaq, M.; Mazhar, K.; Khan, J.S.; Shahid, R. Upregulation of CD271 transcriptome in breast cancer promotes cell survival via NFkappaB pathway. Mol. Biol. Rep. 2022, 49, 487–495. [Google Scholar] [CrossRef] [PubMed]
- Chiarenza, A.; Lazarovici, P.; Lempereur, L.; Cantarella, G.; Bianchi, A.; Bernardini, R. Tamoxifen inhibits nerve growth factor-induced proliferation of the human breast cancerous cell line MCF-7. Cancer Res. 2001, 61, 3002–3008. [Google Scholar] [PubMed]
- Com, E.; Lagadec, C.; Page, A.; El Yazidi-Belkoura, I.; Slomianny, C.; Spencer, A.; Hammache, D.; Rudkin, B.B.; Hondermarck, H. Nerve growth factor receptor TrkA signaling in breast cancer cells involves Ku70 to prevent apoptosis. Mol. Cell. Proteom. 2007, 6, 1842–1854. [Google Scholar] [CrossRef]
- Davidson, B.; Reich, R.; Lazarovici, P.; Ann Florenes, V.; Nielsen, S.; Nesland, J.M. Altered expression and activation of the nerve growth factor receptors TrkA and p75 provide the first evidence of tumor progression to effusion in breast carcinoma. Breast Cancer Res. Treat. 2004, 83, 119–128. [Google Scholar] [CrossRef]
- Descamps, S.; Lebourhis, X.; Delehedde, M.; Boilly, B.; Hondermarck, H. Nerve growth factor is mitogenic for cancerous but not normal human breast epithelial cells. J. Biol. Chem. 1998, 273, 16659–16662. [Google Scholar] [CrossRef]
- Descamps, S.; Pawlowski, V.; Revillion, F.; Hornez, L.; Hebbar, M.; Boilly, B.; Hondermarck, H.; Peyrat, J.P. Expression of nerve growth factor receptors and their prognostic value in human breast cancer. Cancer Res. 2001, 61, 4337–4340. [Google Scholar]
- Descamps, S.; Toillon, R.A.; Adriaenssens, E.; Pawlowski, V.; Cool, S.M.; Nurcombe, V.; Le Bourhis, X.; Boilly, B.; Peyrat, J.P.; Hondermarck, H. Nerve growth factor stimulates proliferation and survival of human breast cancer cells through two distinct signaling pathways. J. Biol. Chem. 2001, 276, 17864–17870. [Google Scholar] [CrossRef]
- Di Donato, M.; Galasso, G.; Giovannelli, P.; Sinisi, A.A.; Migliaccio, A.; Castoria, G. Targeting the Nerve Growth Factor Signaling Impairs the Proliferative and Migratory Phenotype of Triple-Negative Breast Cancer Cells. Front. Cell Dev. Biol. 2021, 9, 676568. [Google Scholar] [CrossRef]
- Dolle, L.; El Yazidi-Belkoura, I.; Adriaenssens, E.; Nurcombe, V.; Hondermarck, H. Nerve growth factor overexpression and autocrine loop in breast cancer cells. Oncogene 2003, 22, 5592–5601. [Google Scholar] [CrossRef]
- Jung, H.H.; Kim, J.Y.; Cho, E.Y.; Oh, J.M.; Lee, J.E.; Kim, S.W.; Nam, S.J.; Park, Y.H.; Ahn, J.S.; Im, Y.H. Elevated Level of Nerve Growth Factor (NGF) in Serum-Derived Exosomes Predicts Poor Survival in Patients with Breast Cancer Undergoing Neoadjuvant Chemotherapy. Cancers 2021, 13, 5260. [Google Scholar] [CrossRef] [PubMed]
- Kumar, A.; Raza, K.; Nag, T.C.; Srivastava, A.; Sehgal, R. Localization and hypersecretion of nerve growth factor in breast phyllodes tumors: Evidence from a preliminary study. Cancer Rep. 2021, 4, e1300. [Google Scholar] [CrossRef]
- Lagadec, C.; Meignan, S.; Adriaenssens, E.; Foveau, B.; Vanhecke, E.; Romon, R.; Toillon, R.A.; Oxombre, B.; Hondermarck, H.; Le Bourhis, X. TrkA overexpression enhances growth and metastasis of breast cancer cells. Oncogene 2009, 28, 1960–1970. [Google Scholar] [CrossRef] [PubMed]
- Melck, D.; De Petrocellis, L.; Orlando, P.; Bisogno, T.; Laezza, C.; Bifulco, M.; Di Marzo, V. Suppression of nerve growth factor Trk receptors and prolactin receptors by endocannabinoids leads to inhibition of human breast and prostate cancer cell proliferation. Endocrinology 2000, 141, 118–126. [Google Scholar] [CrossRef]
- Naderi, A.; Teschendorff, A.E.; Beigel, J.; Cariati, M.; Ellis, I.O.; Brenton, J.D.; Caldas, C. BEX2 is overexpressed in a subset of primary breast cancers and mediates nerve growth factor/nuclear factor-kappaB inhibition of apoptosis in breast cancer cell lines. Cancer Res. 2007, 67, 6725–6736. [Google Scholar] [CrossRef] [PubMed]
- Romon, R.; Adriaenssens, E.; Lagadec, C.; Germain, E.; Hondermarck, H.; Le Bourhis, X. Nerve growth factor promotes breast cancer angiogenesis by activating multiple pathways. Mol. Cancer 2010, 9, 157. [Google Scholar] [CrossRef]
- Sakamoto, Y.; Kitajima, Y.; Edakuni, G.; Hamamoto, T.; Miyazaki, K. Combined evaluation of NGF and p75NGFR expression is a biomarker for predicting prognosis in human invasive ductal breast carcinoma. Oncol. Rep. 2001, 8, 973–980. [Google Scholar] [CrossRef] [PubMed]
- Tagliabue, E.; Castiglioni, F.; Ghirelli, C.; Modugno, M.; Asnaghi, L.; Somenzi, G.; Melani, C.; Menard, S. Nerve growth factor cooperates with p185(HER2) in activating growth of human breast carcinoma cells. J. Biol. Chem. 2000, 275, 5388–5394. [Google Scholar] [CrossRef]
- Trouvilliez, S.; Cicero, J.; Leveque, R.; Aubert, L.; Corbet, C.; Van Outryve, A.; Streule, K.; Angrand, P.O.; Volkel, P.; Magnez, R.; et al. Direct interaction of TrkA/CD44v3 is essential for NGF-promoted aggressiveness of breast cancer cells. J. Exp. Clin. Cancer Res. 2022, 41, 110. [Google Scholar] [CrossRef]
- Tsang, J.Y.; Wong, K.H.; Lai, M.W.; Lacambra, M.D.; Ko, C.W.; Chan, S.K.; Lam, C.C.; Yu, A.M.; Tan, P.H.; Tse, G.M. Nerve growth factor receptor (NGFR): A potential marker for specific molecular subtypes of breast cancer. J. Clin. Pathol. 2013, 66, 291–296. [Google Scholar] [CrossRef]
- Zhang, J.; Wang, L.S.; Ye, S.L.; Luo, P.; Wang, B.L. Blockage of tropomyosin receptor kinase a (TrkA) enhances chemo-sensitivity in breast cancer cells and inhibits metastasis in vivo. Int. J. Clin. Exp. Med. 2015, 8, 634–641. [Google Scholar] [PubMed]
- Slamon, D.J.; Clark, G.M.; Wong, S.G.; Levin, W.J.; Ullrich, A.; McGuire, W.L. Human breast cancer: Correlation of relapse and survival with amplification of the HER-2/neu oncogene. Science 1987, 235, 177–182. [Google Scholar] [CrossRef] [PubMed]
- Kaplan, D.R.; Miller, F.D. Signal transduction by the neurotrophin receptors. Curr. Opin. Cell Biol. 1997, 9, 213–221. [Google Scholar] [CrossRef]
- Janes, P.W.; Daly, R.J.; deFazio, A.; Sutherland, R.L. Activation of the Ras signalling pathway in human breast cancer cells overexpressing erbB-2. Oncogene 1994, 9, 3601–3608. [Google Scholar] [PubMed]
- Carmeliet, P. Mechanisms of angiogenesis and arteriogenesis. Nat. Med. 2000, 6, 389–395. [Google Scholar] [CrossRef]
- Gasparini, G. Prognostic value of vascular endothelial growth factor in breast cancer. Oncologist 2000, 5 (Suppl. S1), 37–44. [Google Scholar] [CrossRef]
- Linderholm, B.; Tavelin, B.; Grankvist, K.; Henriksson, R. Vascular endothelial growth factor is of high prognostic value in node-negative breast carcinoma. J. Clin. Oncol. 1998, 16, 3121–3128. [Google Scholar] [CrossRef]
- Hanahan, D.; Weinberg, R.A. The hallmarks of cancer. Cell 2000, 100, 57–70. [Google Scholar] [CrossRef]
- Hanahan, D.; Weinberg, R.A. Hallmarks of cancer: The next generation. Cell 2011, 144, 646–674. [Google Scholar] [CrossRef]
| Breast Cancer Classification | |||
|---|---|---|---|
| General classification | Type | Sub-type | |
| Histological Type | Carcinoma in situ | Ductal | Comedo Cribiform Micropalillary Papillary Solid |
| Lobular | |||
| Invasive or infiltrating Carcinoma | Tubular | ||
| Ductal lobular | |||
| Invasive lobular | |||
| Infiltrating ductal | |||
| Mucinous | |||
| Medullary | |||
| Grade | Differentiation | Elston-Ellis Score | |
| Grading | Grade 1 (G1) | Well | 3–5 |
| Grade 2 (G2) | Moderate | 6–7 | |
| Grade 3 (G3) | Poor | 8–9 | |
| Molecular markers | Cases | ||
| Immunophenotype | HR positive/HER2 negative | 70% | |
| HER2 positive | 15–20% | ||
| Triple-negative | ~15% | ||
| Stage | TNM | Category | |
| TNM classification | 0 | Tis N0 M0 | Carcinoma in situ |
| I | T1, N0, M0 | Early breast cancer | |
| II | T1, N1, M0 T2, NO1, M0 | Early breast cancer | |
| III | Any T, N2–3, M0, T3 Any N, M0 | Locally advanced | |
| IV | Any T, any N M1 | Metastatic | |
| Subtype | Cases | ||
| Molecular subtype | Luminal A | ~40% | |
| Luminal B | ~20% | ||
| Basal-like | 15–20% | ||
| Claudin-low | 12–14% | ||
| HER2-enriched | 10–15% |
| Authors | Title | Year of Publication | Type of Study | Reference |
|---|---|---|---|---|
| Adriaenssens et al. | Nerve Growth Factor Is a Potential Therapeutic Target in Breast Cancer | 2008 | ex vivo and in vivo | [82] |
| Aragona et al. | Nerve Growth Factor Receptor Immunoreactivity in Breast Cancer Patients | 2001 | ex vivo | [83] |
| Bashir et al. | Upregulation of CD271 transcriptome in breast cancer promotes cell survival via NFκB pathway | 2022 | ex vivo and in vitro | [84] |
| Chakravarthy et al. | Nerve growth factor (NGF)-mediated regulation of p75NTR expression contributes to chemotherapeutic resistance in triple negative breast cancer cells | 2016 | ex vivo and in vitro | [17] |
| Chiarenza et al. | Tamoxifen Inhibits Nerve Growth Factor-induced Proliferation of the Human Breast Cancerous Cell Line MCF-7 | 2001 | in vitro | [85] |
| Com et al. | Nerve Growth Factor Receptor TrkA Signaling in Breast Cancer Cells Involves Ku70 to Prevent Apoptosis | 2007 | in vitro | [86] |
| Davidson et al. | Altered expression and activation of the nerve growth factor receptors TrkA and p75 provide the first evidence of tumor progression to effusion in breast carcinoma | 2004 | ex vivo and in vitro | [87] |
| Demont et al. | Pro-nerve Growth Factor Induces Autocrine Stimulation of Breast Cancer Cell Invasion through Tropomyosin-related Kinase A (TrkA) and Sortilin Protein | 2012 | ex vivo and in vitro | [18] |
| Descamps et al. | Nerve Growth Factor Is Mitogenic for Cancerous but Not Normal Human Breast Epithelial Cells | 1998 | in vitro | [88] |
| Descamps et al. | Expression of Nerve Growth Factor Receptors and Their Prognostic Value in Human Breast Cancer | 2001a | ex vivo | [89] |
| Descamps et al. | Nerve Growth Factor Stimulates Proliferation and Survival of Human Breast Cancer Cells through Two Distinct Signaling Pathways | 2001b | in vitro | [90] |
| Di Donato et al. | Targeting the Nerve Growth Factor Signaling Impairs the Proliferative and Migratory Phenotype of Triple-Negative Breast Cancer Cells | 2021 | in vitro | [91] |
| Dolle’ et al. | Nerve growth factor overexpression and autocrine loop in breast cancer cells | 2003 | ex vivo and in vitro | [92] |
| Islam et al. | Nerve growth factor from Indian Russell’s viper venom (RVV-NGFa) shows high affinity binding to TrkA receptor expressed in breast cancer cells: Application of fluorescence labeled RVV-NGFa in the clinical diagnosis of breast cancer | 2020 | ex vivo and in vitro | [14] |
| Jung et al. | Elevated Level of Nerve Growth Factor (NGF) in Serum-Derived Exosomes Predicts Poor Survival in Patients with Breast Cancer Undergoing Neoadjuvant Chemotherapy | 2021 | in vivo | [93] |
| Kumar et al. | Localization and hypersecretion of nerve growth factor in breast phyllodes tumors: Evidence from a preliminary study | 2020 | ex vivo | [94] |
| Kyker-Snowman et al. | TrkA overexpression in non-tumorigenic human breast cell lines confers oncogenic and metastatic properties | 2020 | in vitro and in vivo | [70] |
| Lagadec et al. | TrkA overexpression enhances growth and metastasis of breast cancer cells | 2009 | in vitro and in vivo | [95] |
| Lévêque et al. | ProNGF increases breast tumor aggressiveness through functional association of TrkA with EphA2 | 2019 | in vitro and in vivo | [19] |
| Melck et al. | Suppression of Nerve Growth Factor Trk Receptors and Prolactin Receptors by Endocannabinoids Leads to Inhibition of Human Breast and Prostate Cancer Cell Proliferation | 2000 | in vitro | [96] |
| Naderi et al. | BEX2 Is Overexpressed in a Subset of Primary Breast Cancers and Mediates Nerve Growth Factor/Nuclear Factor-KB Inhibition of Apoptosis in Breast Cancer Cell Lines | 2007 | ex vivo and in vitro | [97] |
| Noh et al. | Expression of nerve growth factor and heme oxygenase-1 predict poor survival of breast carcinoma patients | 2013 | ex vivo | [16] |
| Romon et al. | Nerve growth factor promotes breast cancer angiogenesis by activating multiple pathways | 2010 | in vitro and in vivo | [98] |
| Sakamoto et al. | Combined evaluation of NGF and p75NGFR expression is a biomarker for predicting prognosis in human invasive ductal breast carcinoma | 2001 | ex vivo | [99] |
| Tagliabue et al. | Nerve Growth Factor Cooperates with p185HER2 in Activating Growth of Human Breast Carcinoma Cells | 2000 | ex vivo and in vitro | [100] |
| Trouvilliez et al. | Direct interaction of TrkA/CD44v3 is essential for NGF-promoted aggressiveness of breast cancer cells | 2022 | in vitro and in vivo | [101] |
| Tsang et al. | Nerve growth factor receptor (NGFR): a potential marker for specific molecular subtypes of breast cancer | 2013 | ex vivo | [102] |
| Wu et al. | Nerve growth factor receptor increases the tumor growth and metastatic potential of triple-negative breast cancer cells | 2021 | in vitro and in vivo | [15] |
| Zhang et al. | Blockage of tropomyosin receptor kinase a (TrkA) enhances chemo-sensitivity in breast cancer cells and inhibits metastasis in vivo | 2015 | in vitro and in vivo | [103] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bruno, F.; Arcuri, D.; Vozzo, F.; Malvaso, A.; Montesanto, A.; Maletta, R. Expression and Signaling Pathways of Nerve Growth Factor (NGF) and Pro-NGF in Breast Cancer: A Systematic Review. Curr. Oncol. 2022, 29, 8103-8120. https://doi.org/10.3390/curroncol29110640
Bruno F, Arcuri D, Vozzo F, Malvaso A, Montesanto A, Maletta R. Expression and Signaling Pathways of Nerve Growth Factor (NGF) and Pro-NGF in Breast Cancer: A Systematic Review. Current Oncology. 2022; 29(11):8103-8120. https://doi.org/10.3390/curroncol29110640
Chicago/Turabian StyleBruno, Francesco, Domenico Arcuri, Francesca Vozzo, Antonio Malvaso, Alberto Montesanto, and Raffaele Maletta. 2022. "Expression and Signaling Pathways of Nerve Growth Factor (NGF) and Pro-NGF in Breast Cancer: A Systematic Review" Current Oncology 29, no. 11: 8103-8120. https://doi.org/10.3390/curroncol29110640
APA StyleBruno, F., Arcuri, D., Vozzo, F., Malvaso, A., Montesanto, A., & Maletta, R. (2022). Expression and Signaling Pathways of Nerve Growth Factor (NGF) and Pro-NGF in Breast Cancer: A Systematic Review. Current Oncology, 29(11), 8103-8120. https://doi.org/10.3390/curroncol29110640






