The Gut Microbiota of Peruvian Children Under the Age of Two During the Complementary Feeding Period
Abstract
1. Introduction
2. Materials and Methods
2.1. Sample and Data Collection
2.2. Sample Processing
2.3. Study Groups
2.4. Diversity and Taxonomy Analyses
2.5. Metadata Analysis
2.6. Multivariable Association Analysis
3. Results
3.1. Alpha Diversity
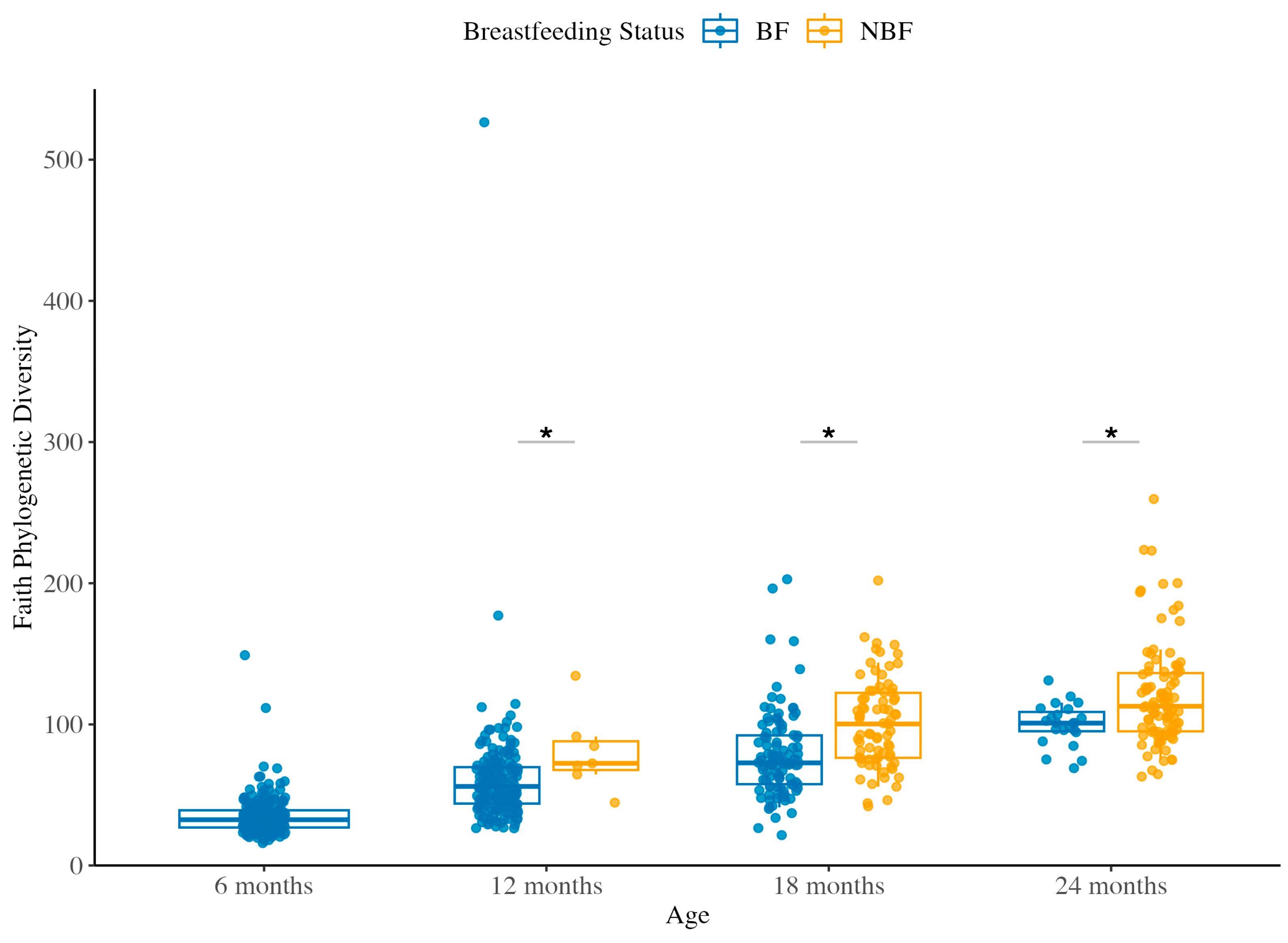
3.1.1. Species Richness Across Ages and Between BF and NBF Groups
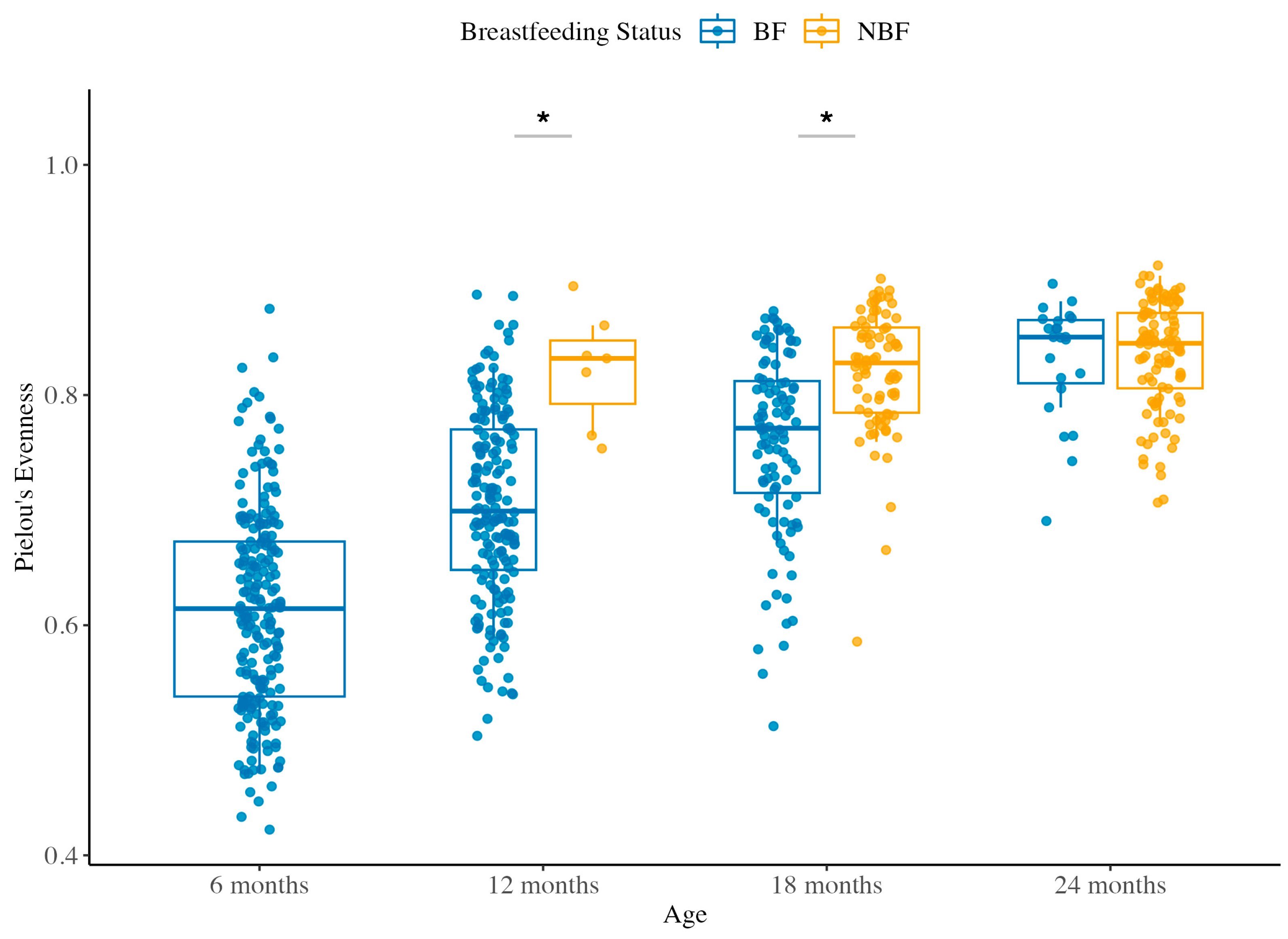
3.1.2. Species Evenness Across Ages and Between BF and NBF Groups
3.2. Beta Diversity Across Ages and Between BF and NBF
3.3. Taxonomy
3.3.1. Microbial Composition at Phylum Level
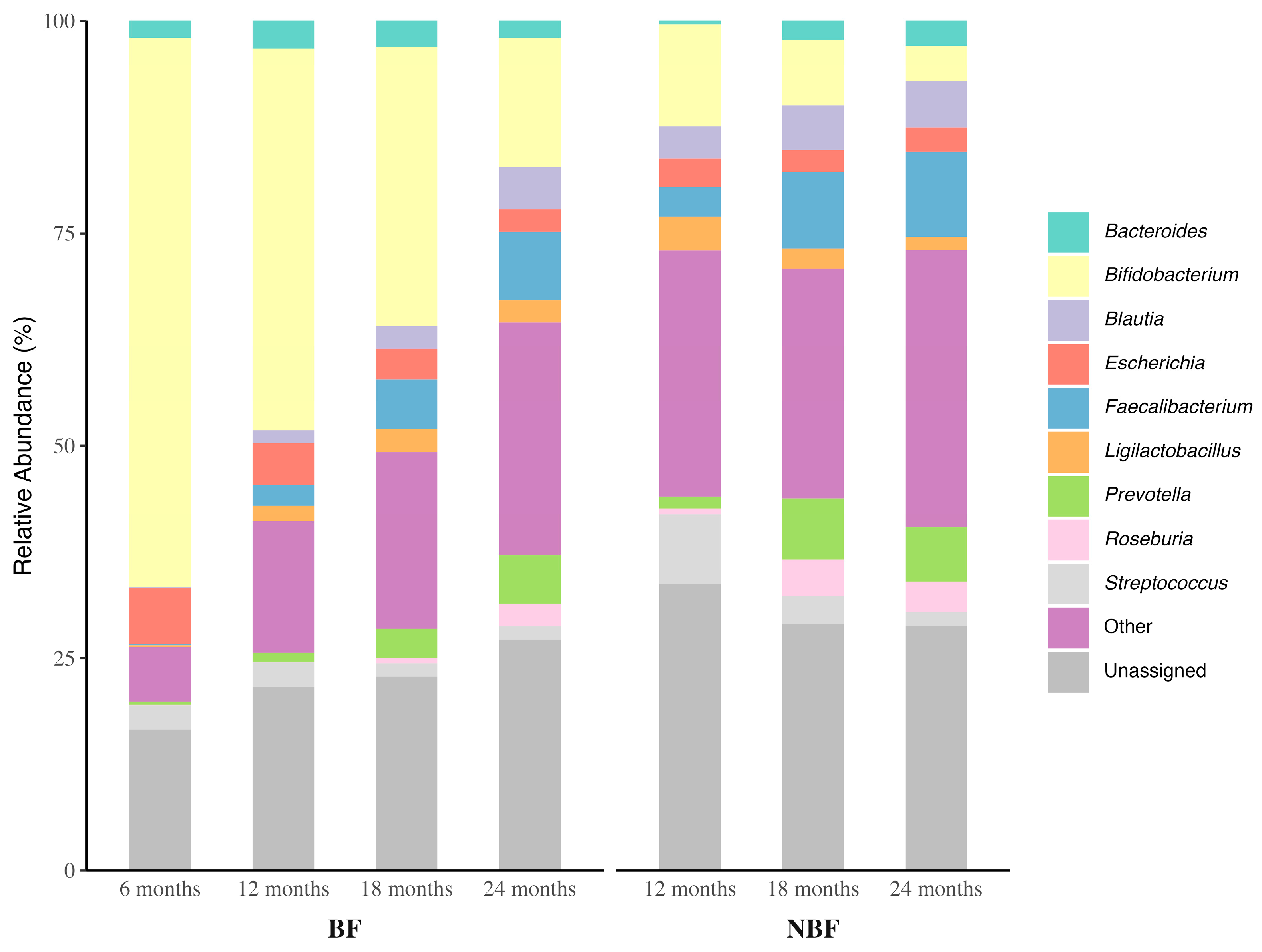
3.3.2. Microbial Composition at Genus Level
3.4. Comparison of Potential Confounding Factors
3.5. Effect Size of Age and Breast Milk Exposure
4. Discussion
4.1. Summary of Study Findings About Impact of Prolonged Breastmilk on Gut Microbiota
4.2. Impact of Human Milk on Infant and Toddler Gut Microbiota
4.3. Implications for Health and Development
4.4. Implications for Feeding Practices and Nutritional Guidance
4.5. Limitations
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| BF | Breastfeeding group (Breastmilk + Complementary Foods) |
| NBF | Non-Breastfeeding group (Breastmilk Weaning Completed) |
| 6M | 6 months group |
| 12M | 12 months group |
| 18M | 18 months group |
| 24M | 24 months group |
| MAL-ED | Etiology, Risk Factors and Interactions of Enteric Infections and Malnutrition and the Consequences for Child Health and Development |
| ENA | European Nucleotide Archive |
| ClinEpiDB | Clinical Epidemiology Resources |
| rRNA | ribosomal RNA |
| ASVs | Amplicon Sequence Variants |
| OTUs | Operational Taxonomic Units |
| FDR | False Discovery Rate |
| PCoA | Principal Coordinates Analysis |
| PERMANOVA | Principal Coordinates Analysis |
| PERMDISP | Multivariate Dispersions |
| HMOs | Human Milk Oligosaccharides |
| SCFAs | Short-Chain Fatty Acids |
| WHO | World Health Organization |
References
- Lin, L.; Zhang, J. Role of Intestinal Microbiota and Metabolites on Gut Homeostasis and Human Diseases. BMC Immunol. 2017, 18, 2. [Google Scholar] [CrossRef]
- Schoultz, I.; Claesson, M.J.; Dominguez-Bello, M.G.; Fåk Hållenius, F.; Konturek, P.; Korpela, K.; Laursen, M.F.; Penders, J.; Roager, H.; Vatanen, T.; et al. Gut Microbiota Development across the Lifespan: Disease Links and Health-promoting Interventions. J. Intern. Med. 2025, 297, 560–583. [Google Scholar] [CrossRef] [PubMed]
- Chibuye, M.; Mende, D.R.; Spijker, R.; Simuyandi, M.; Luchen, C.C.; Bosomprah, S.; Chilengi, R.; Schultsz, C.; Harris, V.C. Systematic Review of Associations Between Gut Microbiome Composition and Stunting in Under-Five Children. npj Biofilms Microbiomes 2024, 10, 46. [Google Scholar] [CrossRef] [PubMed]
- Tang, M.H.; Ligthart, I.; Varga, S.; Lebeer, S.; Van Overveld, F.J.; Rijkers, G.T. Mutual Interactions Between Microbiota and the Human Immune System During the First 1000 Days of Life. Biology 2025, 14, 299. [Google Scholar] [CrossRef]
- Sarkar, A.; Yoo, J.Y.; Valeria Ozorio Dutra, S.; Morgan, K.H.; Groer, M. The Association between Early-Life Gut Microbiota and Long-Term Health and Diseases. J. Clin. Med. 2021, 10, 459. [Google Scholar] [CrossRef]
- Indrio, F.; Martini, S.; Francavilla, R.; Corvaglia, L.; Cristofori, F.; Mastrolia, S.A.; Neu, J.; Rautava, S.; Russo Spena, G.; Raimondi, F.; et al. Epigenetic Matters: The Link between Early Nutrition, Microbiome, and Long-Term Health Development. Front. Pediatr. 2017, 5, 178. [Google Scholar] [CrossRef]
- Stokholm, J.; Blaser, M.J.; Thorsen, J.; Rasmussen, M.A.; Waage, J.; Vinding, R.K.; Schoos, A.-M.M.; Kunøe, A.; Fink, N.R.; Chawes, B.L.; et al. Maturation of the Gut Microbiome and Risk of Asthma in Childhood. Nat. Commun. 2018, 9, 141. [Google Scholar] [CrossRef]
- Xu, H.; Wang, X.; Feng, W.; Liu, Q.; Zhou, S.; Liu, Q.; Cai, L. The Gut Microbiota and Its Interactions with Cardiovascular Disease. Microb. Biotechnol. 2020, 13, 637–656. [Google Scholar] [CrossRef]
- Hashimoto, K. Emerging Role of the Host Microbiome in Neuropsychiatric Disorders: Overview and Future Directions. Mol. Psychiatry 2023, 28, 3625–3637. [Google Scholar] [CrossRef]
- Houghteling, P.D.; Walker, W.A. Why Is Initial Bacterial Colonization of the Intestine Important to Infants’ and Children’s Health? J. Pediatr. Gastroenterol. Nutr. 2015, 60, 294–307. [Google Scholar] [CrossRef]
- Milani, C.; Duranti, S.; Bottacini, F.; Casey, E.; Turroni, F.; Mahony, J.; Belzer, C.; Delgado Palacio, S.; Arboleya Montes, S.; Mancabelli, L.; et al. The First Microbial Colonizers of the Human Gut: Composition, Activities, and Health Implications of the Infant Gut Microbiota. Microbiol. Mol. Biol. Rev. 2017, 81, e00036-17. [Google Scholar] [CrossRef]
- Suárez-Martínez, C.; Santaella-Pascual, M.; Yagüe-Guirao, G.; Martínez-Graciá, C. Infant Gut Microbiota Colonization: Influence of Prenatal and Postnatal Factors, Focusing on Diet. Front. Microbiol. 2023, 14, 1236254. [Google Scholar] [CrossRef] [PubMed]
- Robertson, R.C.; Manges, A.R.; Finlay, B.B.; Prendergast, A.J. The Human Microbiome and Child Growth—First 1000 Days and Beyond. Trends Microbiol. 2019, 27, 131–147. [Google Scholar] [CrossRef] [PubMed]
- Gritz, E.C.; Bhandari, V. The Human Neonatal Gut Microbiome: A Brief Review. Front. Pediatr. 2015, 3, 17. [Google Scholar] [CrossRef] [PubMed]
- Mao, S.; Zhao, A.; Jiang, H.; Yan, J.; Zhong, W.; Xun, Y.; Zhang, Y. Patterns of Human Milk Oligosaccharides in Mature Milk Are Associated with Certain Gut Microbiota in Infants. Nutrients 2024, 16, 1287. [Google Scholar] [CrossRef]
- Biagioli, V.; Sortino, V.; Falsaperla, R.; Striano, P. Role of Human Milk Microbiota in Infant Neurodevelopment: Mechanisms and Clinical Implications. Children 2024, 11, 1476. [Google Scholar] [CrossRef]
- Ho, N.T.; Li, F.; Lee-Sarwar, K.A.; Tun, H.M.; Brown, B.P.; Pannaraj, P.S.; Bender, J.M.; Azad, M.B.; Thompson, A.L.; Weiss, S.T.; et al. Meta-Analysis of Effects of Exclusive Breastfeeding on Infant Gut Microbiota across Populations. Nat. Commun. 2018, 9, 4169. [Google Scholar] [CrossRef]
- Davis, E.C.; Castagna, V.P.; Sela, D.A.; Hillard, M.A.; Lindberg, S.; Mantis, N.J.; Seppo, A.E.; Järvinen, K.M. Gut Microbiome and Breast-Feeding: Implications for Early Immune Development. J. Allergy Clin. Immunol. 2022, 150, 523–534. [Google Scholar] [CrossRef]
- Munyaka, P.M.; Khafipour, E.; Ghia, J.-E. External Influence of Early Childhood Establishment of Gut Microbiota and Subsequent Health Implications. Front. Pediatr. 2014, 2, 109. [Google Scholar] [CrossRef]
- Noles, D.L.; Matzeller, K.L.; Frank, D.N.; Krebs, N.F.; Tang, M. Complementary Feeding and Infant Gut Microbiota: A Narrative Review. Nutrients 2025, 17, 743. [Google Scholar] [CrossRef]
- McKeen, S.; Roy, N.C.; Mullaney, J.A.; Eriksen, H.; Lovell, A.; Kussman, M.; Young, W.; Fraser, K.; Wall, C.R.; McNabb, W.C. Adaptation of the Infant Gut Microbiome during the Complementary Feeding Transition. PLoS ONE 2022, 17, e0270213. [Google Scholar] [CrossRef]
- Shi, Y.; Yin, R.; Pang, J.; Chen, Y.; Li, Z.; Su, S.; Wen, Y. Impact of Complementary Feeding on Infant Gut Microbiome, Metabolites and Early Development. Food Funct. 2024, 15, 10663–10678. [Google Scholar] [CrossRef] [PubMed]
- Flores, J.N.; Lubin, J.-B.; Silverman, M.A. The Case for Microbial Intervention at Weaning. Gut Microbes 2024, 16, 2414798. [Google Scholar] [CrossRef] [PubMed]
- Gensollen, T.; Iyer, S.S.; Kasper, D.L.; Blumberg, R.S. How Colonization by Microbiota in Early Life Shapes the Immune System. Science 2016, 352, 539–544. [Google Scholar] [CrossRef] [PubMed]
- World Health Organization. Breastfeeding. WHO. Available online: https://www.who.int/health-topics/breastfeeding?utm_source=chatgpt.com#tab=tab_2 (accessed on 27 August 2025).
- Stewart, C.J.; Ajami, N.J.; O’Brien, J.L.; Hutchinson, D.S.; Smith, D.P.; Wong, M.C.; Ross, M.C.; Lloyd, R.E.; Doddapaneni, H.; Metcalf, G.A.; et al. Temporal Development of the Gut Microbiome in Early Childhood from the TEDDY Study. Nature 2018, 562, 583–588. [Google Scholar] [CrossRef]
- Eshriqui, I.; Viljakainen, H.T.; Ferreira, S.R.G.; Raju, S.C.; Weiderpass, E.; Figueiredo, R.A.O. Breastfeeding May Have a Long-Term Effect on Oral Microbiota: Results from the Fin-HIT Cohort. Int. Breastfeed. J. 2020, 15, 42. [Google Scholar] [CrossRef]
- Fehr, K.; Moossavi, S.; Sbihi, H.; Boutin, R.C.T.; Bode, L.; Robertson, B.; Yonemitsu, C.; Field, C.J.; Becker, A.B.; Mandhane, P.J.; et al. Breastmilk Feeding Practices Are Associated with the Co-Occurrence of Bacteria in Mothers’ Milk and the Infant Gut: The CHILD Cohort Study. Cell Host Microbe 2020, 28, 285–297.e4. [Google Scholar] [CrossRef]
- Geniselli Da Silva, V.; Tonkie, J.N.; Roy, N.C.; Smith, N.W.; Wall, C.; Kruger, M.C.; Mullaney, J.A.; McNabb, W.C. The Effect of Complementary Foods on the Colonic Microbiota of Weaning Infants: A Systematic Review. Crit. Rev. Food Sci. Nutr. 2024, 1–16. [Google Scholar] [CrossRef]
- Parkin, K.; Palmer, D.J.; Verhasselt, V.; Amenyogbe, N.; Cooper, M.N.; Christophersen, C.T.; Prescott, S.L.; Silva, D.; Martino, D. Metagenomic Characterisation of the Gut Microbiome and Effect of Complementary Feeding on Bifidobacterium Spp. in Australian Infants. Microorganisms 2024, 12, 228. [Google Scholar] [CrossRef]
- Roche, M.L.; Creed-Kanashiro, H.M.; Tuesta, I.; Kuhnlein, H.V. Infant and Young Child Feeding in the Peruvian Amazon: The Need to Promote Exclusive Breastfeeding and Nutrient-dense Traditional Complementary Foods. Matern. Child. Nutr. 2011, 7, 284–294. [Google Scholar] [CrossRef]
- The MAL-ED Network Investigators; Acosta, A.M.; Chavez, C.B.; Flores, J.T.; Olotegui, M.P.; Pinedo, S.R.; Trigoso, D.R.; Vasquez, A.O.; Ahmed, I.; Alam, D.; et al. The MAL-ED Study: A Multinational and Multidisciplinary Approach to Understand the Relationship Between Enteric Pathogens, Malnutrition, Gut Physiology, Physical Growth, Cognitive Development, and Immune Responses in Infants and Children Up to 2 Years of Age in Resource-Poor Environments. Clin. Infect. Dis. 2014, 59 (Suppl. S4), S193–S206. [Google Scholar] [CrossRef]
- MAL-ED Network Investigators. Early Childhood Cognitive Development Is Affected by Interactions among Illness, Diet, Enteropathogens and the Home Environment: Findings from the MAL-ED Birth Cohort Study. BMJ Glob. Health 2018, 3, e000752. [Google Scholar] [CrossRef]
- Richard, S.A.; Barrett, L.J.; Guerrant, R.L.; Checkley, W.; Miller, M.A. Disease Surveillance Methods Used in the 8-Site MAL-ED Cohort Study. Clin. Infect. Dis. 2014, 59 (Suppl. S4), S220–S224. [Google Scholar] [CrossRef] [PubMed]
- Murray-Kolb, L.E.; Rasmussen, Z.A.; Scharf, R.J.; Rasheed, M.A.; Svensen, E.; Seidman, J.C.; Tofail, F.; Koshy, B.; Shrestha, R.; Maphula, A.; et al. The MAL-ED Network Investigators. The MAL-ED Cohort Study: Methods and Lessons Learned When Assessing Early Child Development and Caregiving Mediators in Infants and Young Children in 8 Low- and Middle-Income Countries. Clin. Infect. Dis. 2014, 59 (Suppl. S4), S261–S272. [Google Scholar] [CrossRef]
- Yori, P.P.; Lee, G.; Olórtegui, M.P.; Chávez, C.B.; Flores, J.T.; Vasquez, A.O.; Burga, R.; Pinedo, S.R.; Asayag, C.R.; Black, R.E.; et al. Santa Clara de Nanay: The MAL-ED Cohort in Peru. Clin. Infect. Dis. 2014, 59 (Suppl. S4), S310–S316. [Google Scholar] [CrossRef]
- Rouhani, S.; Griffin, N.W.; Yori, P.P.; Gehrig, J.L.; Olortegui, M.P.; Salas, M.S.; Trigoso, D.R.; Moulton, L.H.; Houpt, E.R.; Barratt, M.J.; et al. Diarrhea as a Potential Cause and Consequence of Reduced Gut Microbial Diversity Among Undernourished Children in Peru. Clin. Infect. Dis. 2020, 71, 989–999. [Google Scholar] [CrossRef]
- Callahan, B.J.; McMurdie, P.J.; Rosen, M.J.; Han, A.W.; Johnson, A.J.A.; Holmes, S.P. DADA2: High-Resolution Sample Inference from Illumina Amplicon Data. Nat. Methods 2016, 13, 581–583. [Google Scholar] [CrossRef]
- Bolyen, E.; Rideout, J.R.; Dillon, M.R.; Bokulich, N.A.; Abnet, C.C.; Al-Ghalith, G.A.; Alexander, H.; Alm, E.J.; Arumugam, M.; Asnicar, F.; et al. Reproducible, Interactive, Scalable and Extensible Microbiome Data Science Using QIIME 2. Nat. Biotechnol. 2019, 37, 852–857. [Google Scholar] [CrossRef]
- Caporaso, J.G.; Kuczynski, J.; Stombaugh, J.; Bittinger, K.; Bushman, F.D.; Costello, E.K.; Fierer, N.; Peña, A.G.; Goodrich, J.K.; Gordon, J.I.; et al. QIIME Allows Analysis of High-Throughput Community Sequencing Data. Nat. Methods 2010, 7, 335–336. [Google Scholar] [CrossRef]
- Instituto Nacional de Estadística e Informática. Encuesta Demográfica y de Salud Familiar 2011; INEI: Lima, Peru, 2011; Available online: https://proyectos.inei.gob.pe/web/biblioineipub/bancopub/Est/Lib1027/Libro.pdf (accessed on 1 July 2025).
- Instituto Nacional de Estadística e Informática. Encuesta Demográfica y de Salud Familiar-ENDES Continua, 2010; INEI: Lima, Peru, 2010; Available online: https://proyectos.inei.gob.pe/endes/2010/endes02/libro.pdf (accessed on 1 July 2025).
- Victora, C.G.; De Onis, M.; Hallal, P.C.; Blössner, M.; Shrimpton, R. Worldwide Timing of Growth Faltering: Revisiting Implications for Interventions. Pediatrics 2010, 125, e473–e480. [Google Scholar] [CrossRef]
- McDonald, D.; Jiang, Y.; Balaban, M.; Cantrell, K.; Zhu, Q.; Gonzalez, A.; Morton, J.T.; Nicolaou, G.; Parks, D.H.; Karst, S.M.; et al. Greengenes2 Unifies Microbial Data in a Single Reference Tree. Nat. Biotechnol. 2024, 42, 715–718. [Google Scholar] [CrossRef]
- Robeson, M.S.; O’Rourke, D.R.; Kaehler, B.D.; Ziemski, M.; Dillon, M.R.; Foster, J.T.; Bokulich, N.A. RESCRIPt: Reproducible Sequence Taxonomy Reference Database Management. PLoS Comput. Biol. 2021, 17, e1009581. [Google Scholar] [CrossRef] [PubMed]
- Bokulich, N.A.; Kaehler, B.D.; Rideout, J.R.; Dillon, M.; Bolyen, E.; Knight, R.; Huttley, G.A.; Gregory Caporaso, J. Optimizing Taxonomic Classification of Marker-Gene Amplicon Sequences with QIIME 2′s Q2-Feature-Classifier Plugin. Microbiome 2018, 6, 90. [Google Scholar] [CrossRef] [PubMed]
- Janssen, S.; McDonald, D.; Gonzalez, A.; Navas-Molina, J.A.; Jiang, L.; Xu, Z.Z.; Winker, K.; Kado, D.M.; Orwoll, E.; Manary, M.; et al. Phylogenetic Placement of Exact Amplicon Sequences Improves Associations with Clinical Information. mSystems 2018, 3, e00021-18. [Google Scholar] [CrossRef] [PubMed]
- Yokono, M.; Satoh, S.; Tanaka, A. Comparative Analyses of Whole-Genome Protein Sequences from Multiple Organisms. Sci. Rep. 2018, 8, 6800. [Google Scholar] [CrossRef]
- Armstrong, G.; Cantrell, K.; Huang, S.; McDonald, D.; Haiminen, N.; Carrieri, A.P.; Zhu, Q.; Gonzalez, A.; McGrath, I.; Beck, K.L.; et al. Efficient Computation of Faith’s Phylogenetic Diversity with Applications in Characterizing Microbiomes. Genome Res. 2021, 31, 2131–2137. [Google Scholar] [CrossRef]
- Chao, A.; Gotelli, N.J.; Hsieh, T.C.; Sander, E.L.; Ma, K.H.; Colwell, R.K.; Ellison, A.M. Rarefaction and Extrapolation with Hill Numbers: A Framework for Sampling and Estimation in Species Diversity Studies. Ecol. Monogr. 2014, 84, 45–67. [Google Scholar] [CrossRef]
- Faith, D.P.; Baker, A.M. Phylogenetic Diversity (PD) and Biodiversity Conservation: Some Bioinformatics Challenges. Evol. Bioinform. 2007, 2, 121–128. [Google Scholar] [CrossRef]
- Faith, D.P. Conservation Evaluation and Phylogenetic Diversity. Biol. Conserv. 1992, 61, 1–10. [Google Scholar] [CrossRef]
- Pielou, E.C. The Measurement of Diversity in Different Types of Biological Collections. J. Theor. Biol. 1966, 13, 131–144. [Google Scholar] [CrossRef]
- Kruskal, W.H.; Wallis, W.A. Use of Ranks in One-Criterion Variance Analysis. J. Am. Stat. Assoc. 1952, 47, 583–621. [Google Scholar] [CrossRef]
- Dunn, O.J. Multiple Comparisons Using Rank Sums. Technometrics 1964, 6, 241–252. [Google Scholar] [CrossRef]
- Dunn, O.J. Multiple Comparisons among Means. J. Am. Stat. Assoc. 1961, 56, 52–64. [Google Scholar] [CrossRef]
- Benjamini, Y.; Hochberg, Y. Controlling the False Discovery Rate: A Practical and Powerful Approach to Multiple Testing. J. R. Stat. Soc. Ser. B Stat. Methodol. 1995, 57, 289–300. [Google Scholar] [CrossRef]
- Lozupone, C.A.; Hamady, M.; Kelley, S.T.; Knight, R. Quantitative and Qualitative β Diversity Measures Lead to Different Insights into Factors That Structure Microbial Communities. Appl. Environ. Microbiol. 2007, 73, 1576–1585. [Google Scholar] [CrossRef] [PubMed]
- McDonald, D.; Vázquez-Baeza, Y.; Koslicki, D.; McClelland, J.; Reeve, N.; Xu, Z.; Gonzalez, A.; Knight, R. Striped UniFrac: Enabling Microbiome Analysis at Unprecedented Scale. Nat. Methods 2018, 15, 847–848. [Google Scholar] [CrossRef] [PubMed]
- Lozupone, C.; Lladser, M.E.; Knights, D.; Stombaugh, J.; Knight, R. UniFrac: An Effective Distance Metric for Microbial Community Comparison. ISME J. 2011, 5, 169–172. [Google Scholar] [CrossRef]
- Lozupone, C.; Knight, R. UniFrac: A New Phylogenetic Method for Comparing Microbial Communities. Appl. Environ. Microbiol. 2005, 71, 8228–8235. [Google Scholar] [CrossRef]
- Hamady, M.; Lozupone, C.; Knight, R. Fast UniFrac: Facilitating High-Throughput Phylogenetic Analyses of Microbial Communities Including Analysis of Pyrosequencing and PhyloChip Data. ISME J. 2010, 4, 17–27. [Google Scholar] [CrossRef]
- Anderson, M.J. Permutational Multivariate Analysis of Variance (PERMANOVA). In Wiley StatsRef: Statistics Reference Online; Kenett, R.S., Longford, N.T., Piegorsch, W.W., Ruggeri, F., Eds.; John Wiley &Sons, Ltd.: Hoboken, NJ, USA, 2017; pp. 1–15. [Google Scholar] [CrossRef]
- Anderson, M.J. A New Method for Non-parametric Multivariate Analysis of Variance. Austral Ecol. 2001, 26, 32–46. [Google Scholar] [CrossRef]
- Lin, H.; Peddada, S.D. Analysis of Microbial Compositions: A Review of Normalization and Differential Abundance Analysis. npj Biofilms Microbiomes 2020, 6, 60. [Google Scholar] [CrossRef]
- Shapiro, S.S.; Wilk, M.B. An Analysis of Variance Test for Normality (Complete Samples). Biometrika 1965, 52, 591–611. [Google Scholar] [CrossRef]
- Fox, K.A. Intermediate Economic Statistics, 2nd ed.; R. E. Krieger: Huntington, NY, USA, 1980. [Google Scholar]
- Student. The Probable Error of a Mean. Biometrika 1908, 6, 1–25. [Google Scholar] [CrossRef]
- Fisher, R.A. Statistical Methods for Research Workers, 7th ed.; rev.enl.; Biological Monographs and Manuals; Oliver and Boyd: Edinburgh, UK, 1938. [Google Scholar]
- Fisher, R.A. On the Interpretation of χ2 from Contingency Tables, and the Calculation of P. J. R. Stat. Soc. 1922, 85, 87. [Google Scholar] [CrossRef]
- Mallick, H.; Rahnavard, A.; McIver, L.J.; Ma, S.; Zhang, Y.; Nguyen, L.H.; Tickle, T.L.; Weingart, G.; Ren, B.; Schwager, E.H.; et al. Multivariable Association Discovery in Population-Scale Meta-Omics Studies. PLoS Comput. Biol. 2021, 17, e1009442. [Google Scholar] [CrossRef] [PubMed]
- Timmerman, H.M.; Rutten, N.B.M.M.; Boekhorst, J.; Saulnier, D.M.; Kortman, G.A.M.; Contractor, N.; Kullen, M.; Floris, E.; Harmsen, H.J.M.; Vlieger, A.M.; et al. Intestinal Colonisation Patterns in Breastfed and Formula-Fed Infants during the First 12 Weeks of Life Reveal Sequential Microbiota Signatures. Sci. Rep. 2017, 7, 8327. [Google Scholar] [CrossRef]
- Laursen, M.F.; Pekmez, C.T.; Larsson, M.W.; Lind, M.V.; Yonemitsu, C.; Larnkjær, A.; Mølgaard, C.; Bode, L.; Dragsted, L.O.; Michaelsen, K.F.; et al. Maternal Milk Microbiota and Oligosaccharides Contribute to the Infant Gut Microbiota Assembly. ISME Commun. 2021, 1, 21. [Google Scholar] [CrossRef]
- Fouhy, F.; Watkins, C.; Hill, C.J.; O’Shea, C.-A.; Nagle, B.; Dempsey, E.M.; O’Toole, P.W.; Ross, R.P.; Ryan, C.A.; Stanton, C. Perinatal Factors Affect the Gut Microbiota up to Four Years after Birth. Nat. Commun. 2019, 10, 1517. [Google Scholar] [CrossRef]
- Jeong, S. Factors Influencing Development of the Infant Microbiota: From Prenatal Period to Early Infancy. Clin. Exp. Pediatr. 2022, 65, 438–447. [Google Scholar] [CrossRef]
- Oki, K.; Akiyama, T.; Matsuda, K.; Gawad, A.; Makino, H.; Ishikawa, E.; Oishi, K.; Kushiro, A.; Fujimoto, J. Long-Term Colonization Exceeding Six Years from Early Infancy of Bifidobacterium longum Subsp. Longum in Human Gut. BMC Microbiol. 2018, 18, 209. [Google Scholar] [CrossRef]
- Chichlowski, M.; Shah, N.; Wampler, J.L.; Wu, S.S.; Vanderhoof, J.A. Bifidobacterium longum Subspecies Infantis (B. Infantis) in Pediatric Nutrition: Current State of Knowledge. Nutrients 2020, 12, 1581. [Google Scholar] [CrossRef]
- Laursen, M.F.; Andersen, L.B.B.; Michaelsen, K.F.; Mølgaard, C.; Trolle, E.; Bahl, M.I.; Licht, T.R. Infant Gut Microbiota Development Is Driven by Transition to Family Foods Independent of Maternal Obesity. mSphere 2016, 1, e00069-15. [Google Scholar] [CrossRef] [PubMed]
- Bergström, A.; Skov, T.H.; Bahl, M.I.; Roager, H.M.; Christensen, L.B.; Ejlerskov, K.T.; Mølgaard, C.; Michaelsen, K.F.; Licht, T.R. Establishment of Intestinal Microbiota during Early Life: A Longitudinal, Explorative Study of a Large Cohort of Danish Infants. Appl. Environ. Microbiol. 2014, 80, 2889–2900. [Google Scholar] [CrossRef] [PubMed]
- De Filippo, C.; Cavalieri, D.; Di Paola, M.; Ramazzotti, M.; Poullet, J.B.; Massart, S.; Collini, S.; Pieraccini, G.; Lionetti, P. Impact of Diet in Shaping Gut Microbiota Revealed by a Comparative Study in Children from Europe and Rural Africa. Proc. Natl. Acad. Sci. USA 2010, 107, 14691–14696. [Google Scholar] [CrossRef]
- Wu, G.D.; Chen, J.; Hoffmann, C.; Bittinger, K.; Chen, Y.-Y.; Keilbaugh, S.A.; Bewtra, M.; Knights, D.; Walters, W.A.; Knight, R.; et al. Linking Long-Term Dietary Patterns with Gut Microbial Enterotypes. Science 2011, 334, 105–108. [Google Scholar] [CrossRef]
- Inchingolo, F.; Inchingolo, A.M.; Latini, G.; Ferrante, L.; De Ruvo, E.; Campanelli, M.; Longo, M.; Palermo, A.; Inchingolo, A.D.; Dipalma, G. Difference in the Intestinal Microbiota between Breastfeed Infants and Infants Fed with Artificial Milk: A Systematic Review. Pathogens 2024, 13, 533. [Google Scholar] [CrossRef]
- Zhang, X.; Mushajiang, S.; Luo, B.; Tian, F.; Ni, Y.; Yan, W. The Composition and Concordance of Lactobacillus Populations of Infant Gut and the Corresponding Breast-Milk and Maternal Gut. Front. Microbiol. 2020, 11, 597911. [Google Scholar] [CrossRef]
- Czosnykowska-Łukacka, M.; Królak-Olejnik, B.; Orczyk-Pawiłowicz, M. Breast Milk Macronutrient Components in Prolonged Lactation. Nutrients 2018, 10, 1893. [Google Scholar] [CrossRef]
- Czosnykowska-Łukacka, M.; Lis-Kuberka, J.; Królak-Olejnik, B.; Orczyk-Pawiłowicz, M. Changes in Human Milk Immunoglobulin Profile During Prolonged Lactation. Front. Pediatr. 2020, 8, 428. [Google Scholar] [CrossRef]
- Kalkan, A.E.; BinMowyna, M.N.; Raposo, A.; Ahmad, M.F.; Ahmed, F.; Otayf, A.Y.; Carrascosa, C.; Saraiva, A.; Karav, S. Beyond the Gut: Unveiling Butyrate’s Global Health Impact Through Gut Health and Dysbiosis-Related Conditions: A Narrative Review. Nutrients 2025, 17, 1305. [Google Scholar] [CrossRef]
- Yao, Y.; Cai, X.; Ye, Y.; Wang, F.; Chen, F.; Zheng, C. The Role of Microbiota in Infant Health: From Early Life to Adulthood. Front. Immunol. 2021, 12, 708472. [Google Scholar] [CrossRef]
- Davis, E.C.; Dinsmoor, A.M.; Wang, M.; Donovan, S.M. Microbiome Composition in Pediatric Populations from Birth to Adolescence: Impact of Diet and Prebiotic and Probiotic Interventions. Dig. Dis. Sci. 2020, 65, 706–722. [Google Scholar] [CrossRef] [PubMed]
- Bäckhed, F.; Roswall, J.; Peng, Y.; Feng, Q.; Jia, H.; Kovatcheva-Datchary, P.; Li, Y.; Xia, Y.; Xie, H.; Zhong, H.; et al. Dynamics and Stabilization of the Human Gut Microbiome during the First Year of Life. Cell Host Microbe 2015, 17, 690–703. [Google Scholar] [CrossRef] [PubMed]
- Forbes, J.D.; Azad, M.B.; Vehling, L.; Tun, H.M.; Konya, T.B.; Guttman, D.S.; Field, C.J.; Lefebvre, D.; Sears, M.R.; Becker, A.B.; et al. Association of Exposure to Formula in the Hospital and Subsequent Infant Feeding Practices with Gut Microbiota and Risk of Overweight in the First Year of Life. JAMA Pediatr. 2018, 172, e181161. [Google Scholar] [CrossRef] [PubMed]
- Marcobal, A.; Barboza, M.; Sonnenburg, E.D.; Pudlo, N.; Martens, E.C.; Desai, P.; Lebrilla, C.B.; Weimer, B.C.; Mills, D.A.; German, J.B.; et al. Bacteroides in the Infant Gut Consume Milk Oligosaccharides via Mucus-Utilization Pathways. Cell Host Microbe 2011, 10, 507–514. [Google Scholar] [CrossRef]
- De Filippis, F.; Pasolli, E.; Ercolini, D. Newly Explored Faecalibacterium Diversity Is Connected to Age, Lifestyle, Geography, and Disease. Curr. Biol. 2020, 30, 4932–4943.e4. [Google Scholar] [CrossRef]
- Nie, K.; Ma, K.; Luo, W.; Shen, Z.; Yang, Z.; Xiao, M.; Tong, T.; Yang, Y.; Wang, X. Roseburia intestinalis: A Beneficial Gut Organism from the Discoveries in Genus and Species. Front. Cell. Infect. Microbiol. 2021, 11, 757718. [Google Scholar] [CrossRef]
- La Rosa, S.L.; Leth, M.L.; Michalak, L.; Hansen, M.E.; Pudlo, N.A.; Glowacki, R.; Pereira, G.; Workman, C.T.; Arntzen, M.Ø.; Pope, P.B.; et al. The Human Gut Firmicute Roseburia Intestinalis Is a Primary Degrader of Dietary β-Mannans. Nat. Commun. 2019, 10, 905. [Google Scholar] [CrossRef]
- Fehlner-Peach, H.; Magnabosco, C.; Raghavan, V.; Scher, J.U.; Tett, A.; Cox, L.M.; Gottsegen, C.; Watters, A.; Wiltshire-Gordon, J.D.; Segata, N.; et al. Distinct Polysaccharide Utilization Profiles of Human Intestinal Prevotella Copri Isolates. Cell Host Microbe 2019, 26, 680–690.e5. [Google Scholar] [CrossRef]
- Laursen, M.F.; Sakanaka, M.; Von Burg, N.; Mörbe, U.; Andersen, D.; Moll, J.M.; Pekmez, C.T.; Rivollier, A.; Michaelsen, K.F.; Mølgaard, C.; et al. Bifidobacterium Species Associated with Breastfeeding Produce Aromatic Lactic Acids in the Infant Gut. Nat. Microbiol. 2021, 6, 1367–1382. [Google Scholar] [CrossRef]
- Freitas, R.G.B.D.O.N.; Vasques, A.C.J.; Fernandes, G.D.R.; Ribeiro, F.B.; Solar, I.; Barbosa, M.G.; de Almeida-Pititto, B.; Geloneze, B.; Ferreira, S.R.G. Associations of Blautia genus with Early-Life Events and Later Phenotype in the NutriHS. Front. Cell. Infect. Microbiol. 2022, 12, 838750. [Google Scholar] [CrossRef]
- Horigome, A.; Hashikura, N.; Yoshida, K.; Xiao, J.; Odamaki, T. 2′-Fucosyllactose Increases the Abundance of Blautia in the Presence of Extracellular Fucosidase-Possessing Bacteria. Front. Microbiol. 2022, 13, 913624. [Google Scholar] [CrossRef]
- Gorvitovskaia, A.; Holmes, S.P.; Huse, S.M. Interpreting Prevotella and Bacteroides as Biomarkers of Diet and Lifestyle. Microbiome 2016, 4, 15. [Google Scholar] [CrossRef]
- Dempsey, E.; Corr, S.C. Lactobacillus spp. for Gastrointestinal Health: Current and Future Perspectives. Front. Immunol. 2022, 13, 840245. [Google Scholar] [CrossRef]
- Singh, V.; Lee, G.; Son, H.; Koh, H.; Kim, E.S.; Unno, T.; Shin, J.-H. Butyrate Producers, “The Sentinel of Gut”: Their Intestinal Significance with and beyond Butyrate, and Prospective Use as Microbial Therapeutics. Front. Microbiol. 2023, 13, 1103836. [Google Scholar] [CrossRef] [PubMed]
- Belfort, M.B.; Rifas-Shiman, S.L.; Kleinman, K.P.; Guthrie, L.B.; Bellinger, D.C.; Taveras, E.M.; Gillman, M.W.; Oken, E. Infant Feeding and Childhood Cognition at Ages 3 and 7 Years: Effects of Breastfeeding Duration and Exclusivity. JAMA Pediatr. 2013, 167, 836. [Google Scholar] [CrossRef] [PubMed]
- Victora, C.G.; Horta, B.L.; De Mola, C.L.; Quevedo, L.; Pinheiro, R.T.; Gigante, D.P.; Gonçalves, H.; Barros, F.C. Association between Breastfeeding and Intelligence, Educational Attainment, and Income at 30 Years of Age: A Prospective Birth Cohort Study from Brazil. Lancet Glob. Health 2015, 3, e199–e205. [Google Scholar] [CrossRef] [PubMed]
- Przyrembel, H. Timing of Introduction of Complementary Food: Short- and Long-Term Health Consequences. Ann. Nutr. Metab. 2012, 60 (Suppl. S2), 8–20. [Google Scholar] [CrossRef] [PubMed]
- Krawiec, M.; Fisher, H.R.; Du Toit, G.; Bahnson, H.T.; Lack, G. Overview of Oral Tolerance Induction for Prevention of Food Allergy—Where Are We Now? Allergy 2021, 76, 2684–2698. [Google Scholar] [CrossRef]


Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kutbi, A.; Gong, J.; Dluzen, D.; Pop, M.; Li, Y. The Gut Microbiota of Peruvian Children Under the Age of Two During the Complementary Feeding Period. Int. J. Environ. Res. Public Health 2025, 22, 1369. https://doi.org/10.3390/ijerph22091369
Kutbi A, Gong J, Dluzen D, Pop M, Li Y. The Gut Microbiota of Peruvian Children Under the Age of Two During the Complementary Feeding Period. International Journal of Environmental Research and Public Health. 2025; 22(9):1369. https://doi.org/10.3390/ijerph22091369
Chicago/Turabian StyleKutbi, Ayat, Junming Gong, Douglas Dluzen, Mihai Pop, and Yuejin Li. 2025. "The Gut Microbiota of Peruvian Children Under the Age of Two During the Complementary Feeding Period" International Journal of Environmental Research and Public Health 22, no. 9: 1369. https://doi.org/10.3390/ijerph22091369
APA StyleKutbi, A., Gong, J., Dluzen, D., Pop, M., & Li, Y. (2025). The Gut Microbiota of Peruvian Children Under the Age of Two During the Complementary Feeding Period. International Journal of Environmental Research and Public Health, 22(9), 1369. https://doi.org/10.3390/ijerph22091369





