Machine Learning Techniques, Applications, and Potential Future Opportunities in Pressure Injuries (Bedsores) Management: A Systematic Review
Abstract
1. Introduction
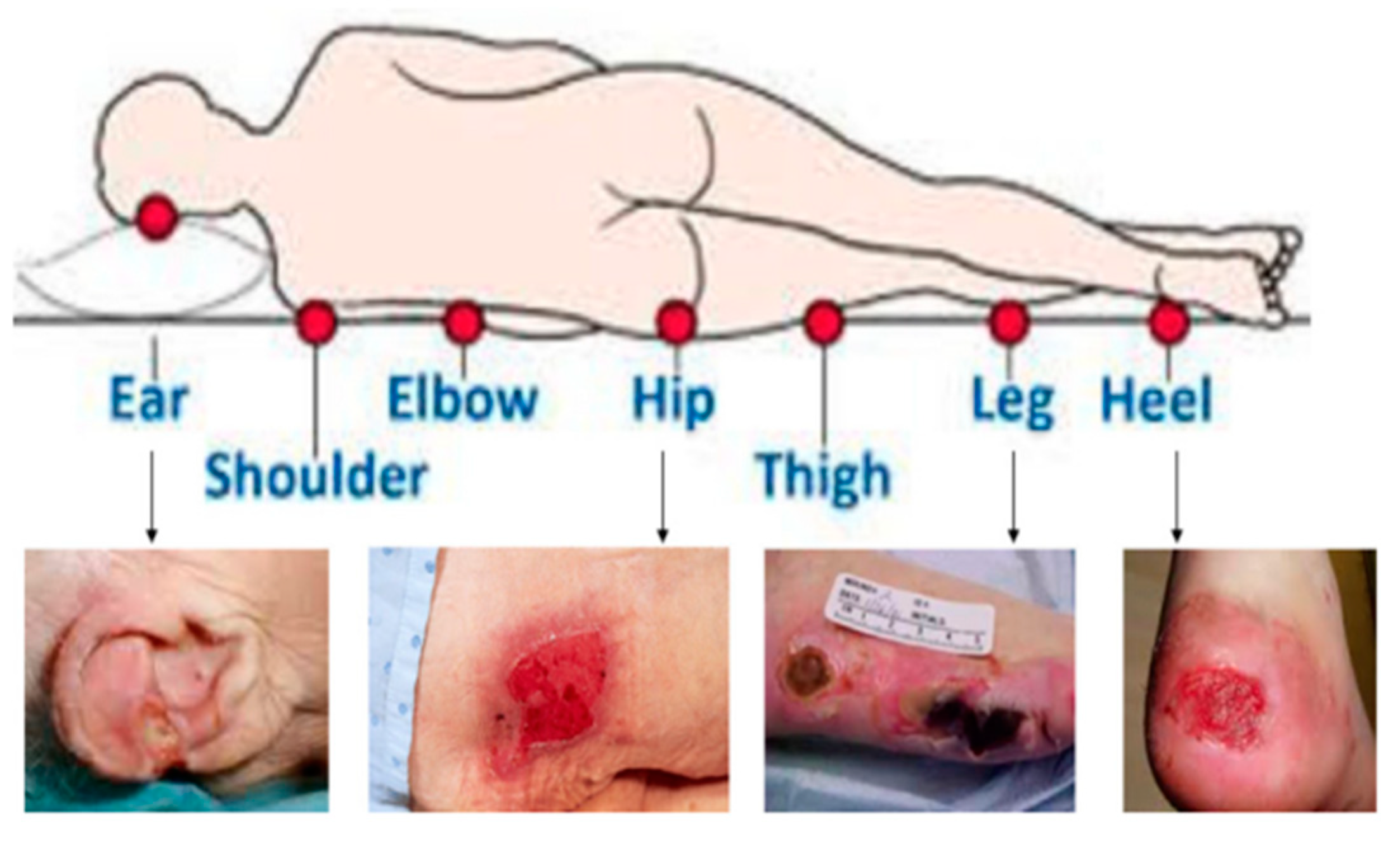
1.1. Pressure Injuries Formation
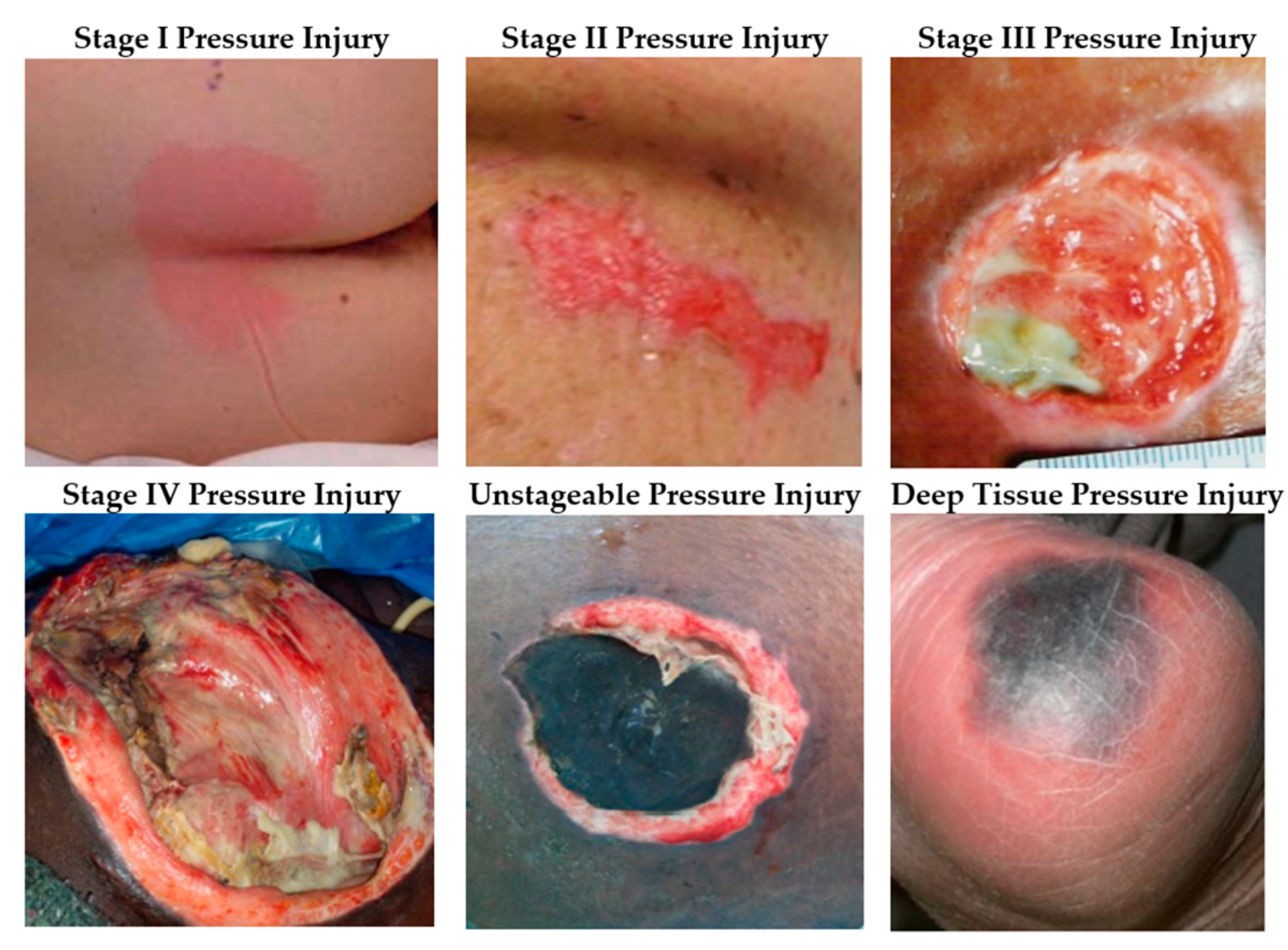
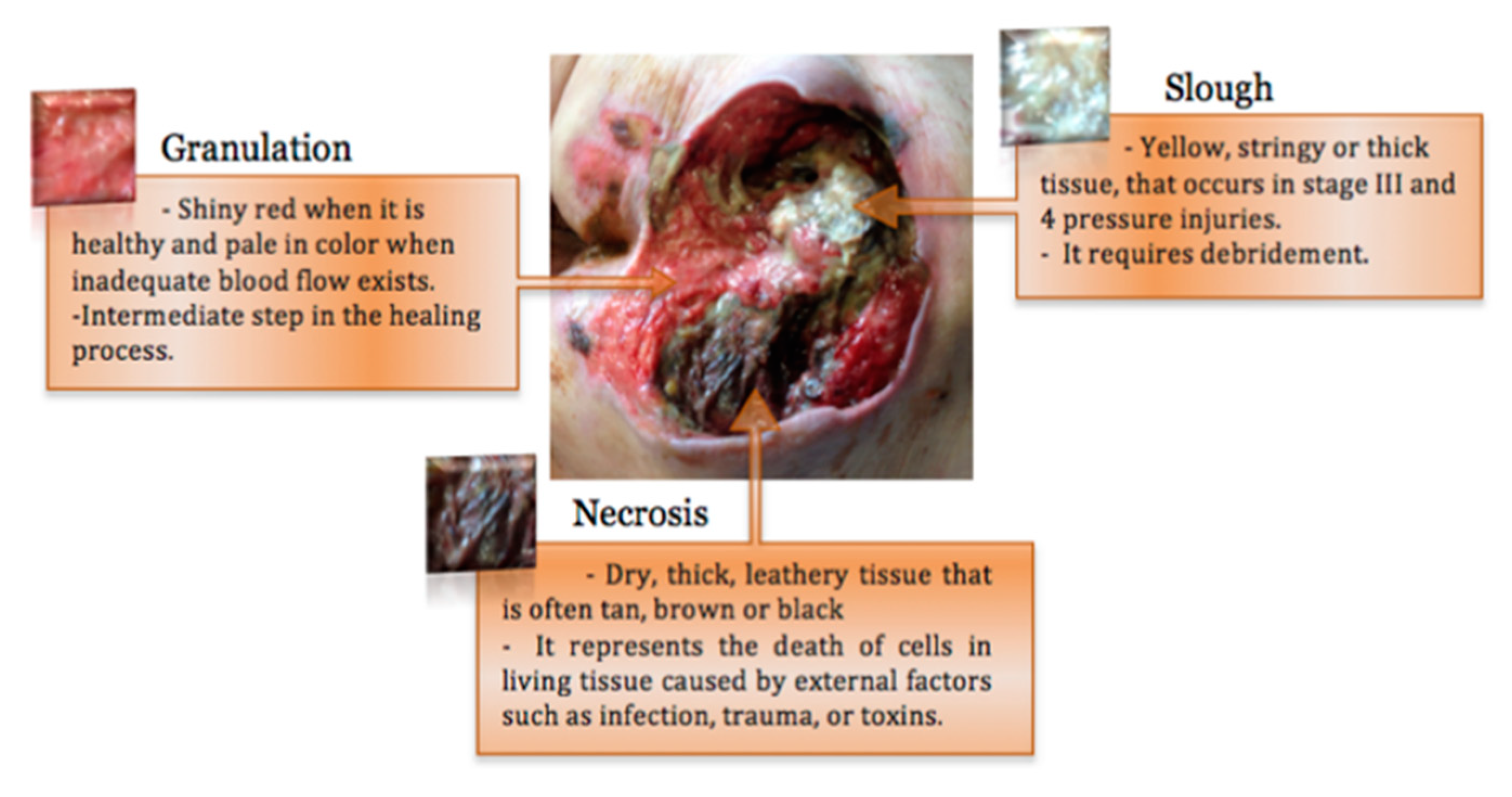
1.2. Pressure Injuries Stages
1.3. Pressure Injuries Consequences
1.4. Pressure Injuries Prevention
1.5. Standardized Risk Assessment Tools and Risk Factors
1.6. Research Objectives
1.7. Research Structure
2. Methods
2.1. Reporting Method
2.2. Search Strategy
2.3. Inclusion Criteria
2.4. Exclusion Criteria
2.5. Study Selection Methods
2.6. Data Extraction
3. Results
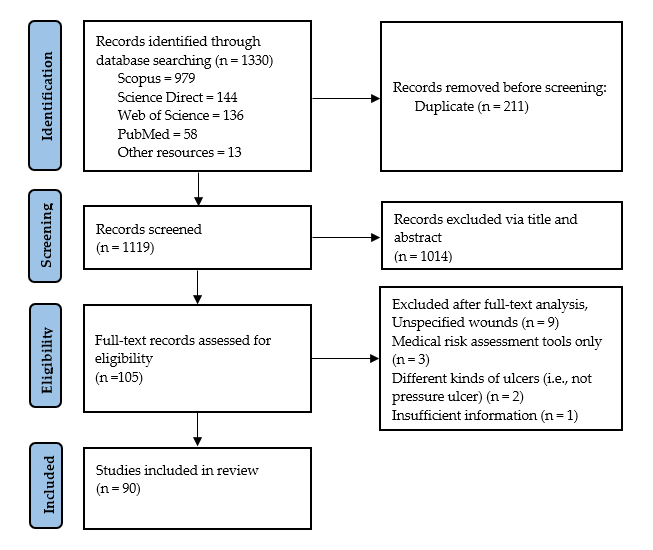
3.1. Study Process
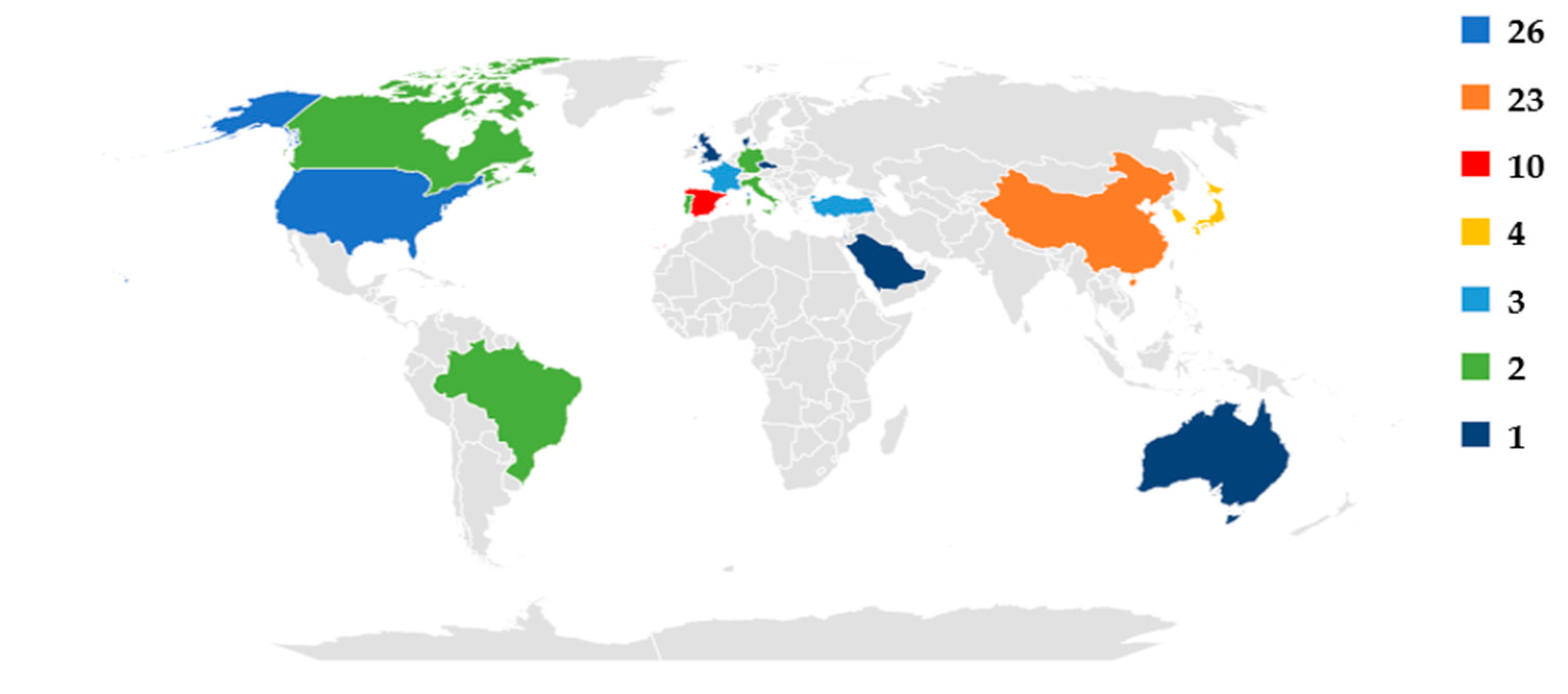
3.2. Characteristics of Included Studies
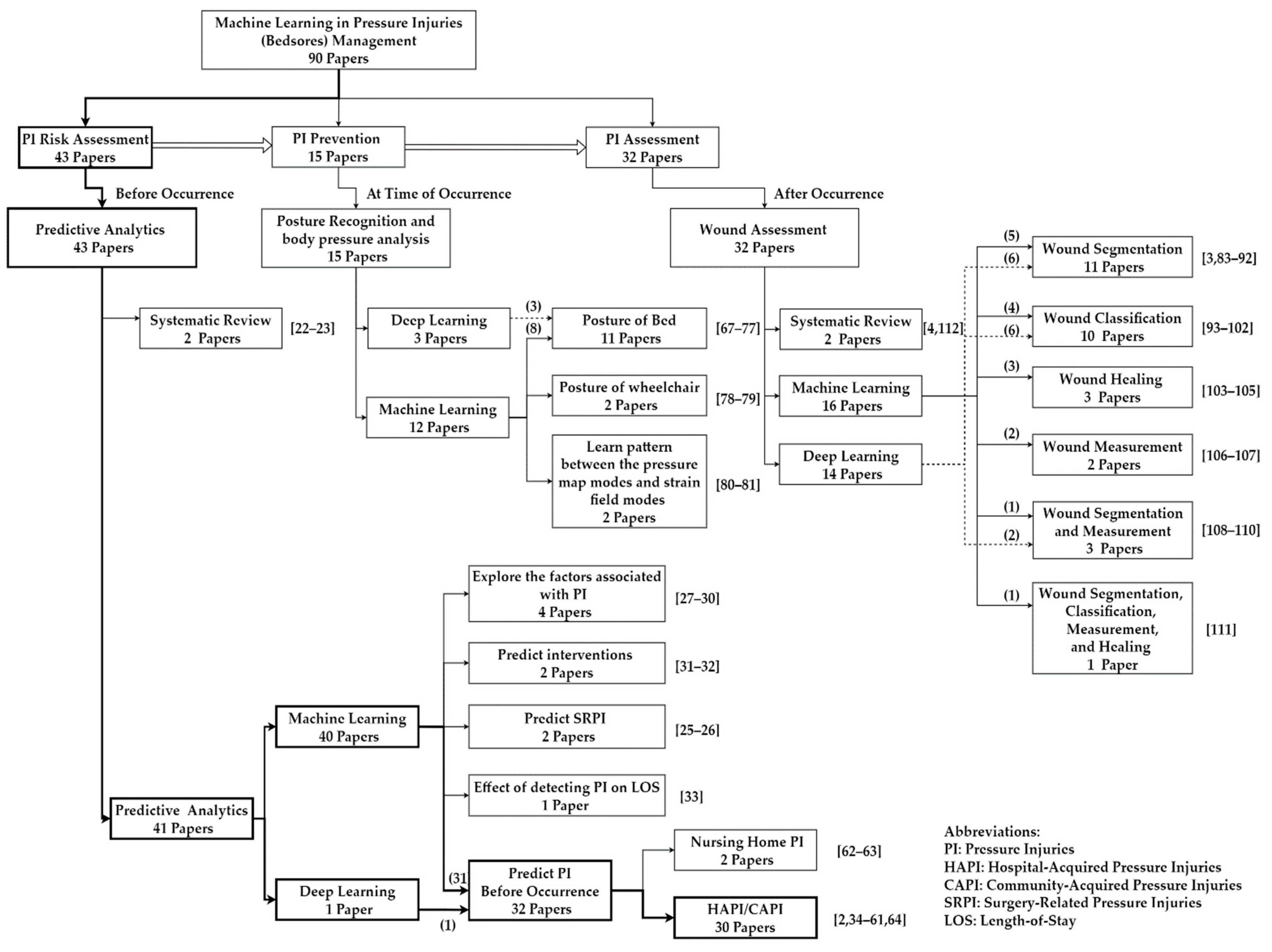
3.3. Categorization of the Studies with High-Level Analysis
4. Discussion
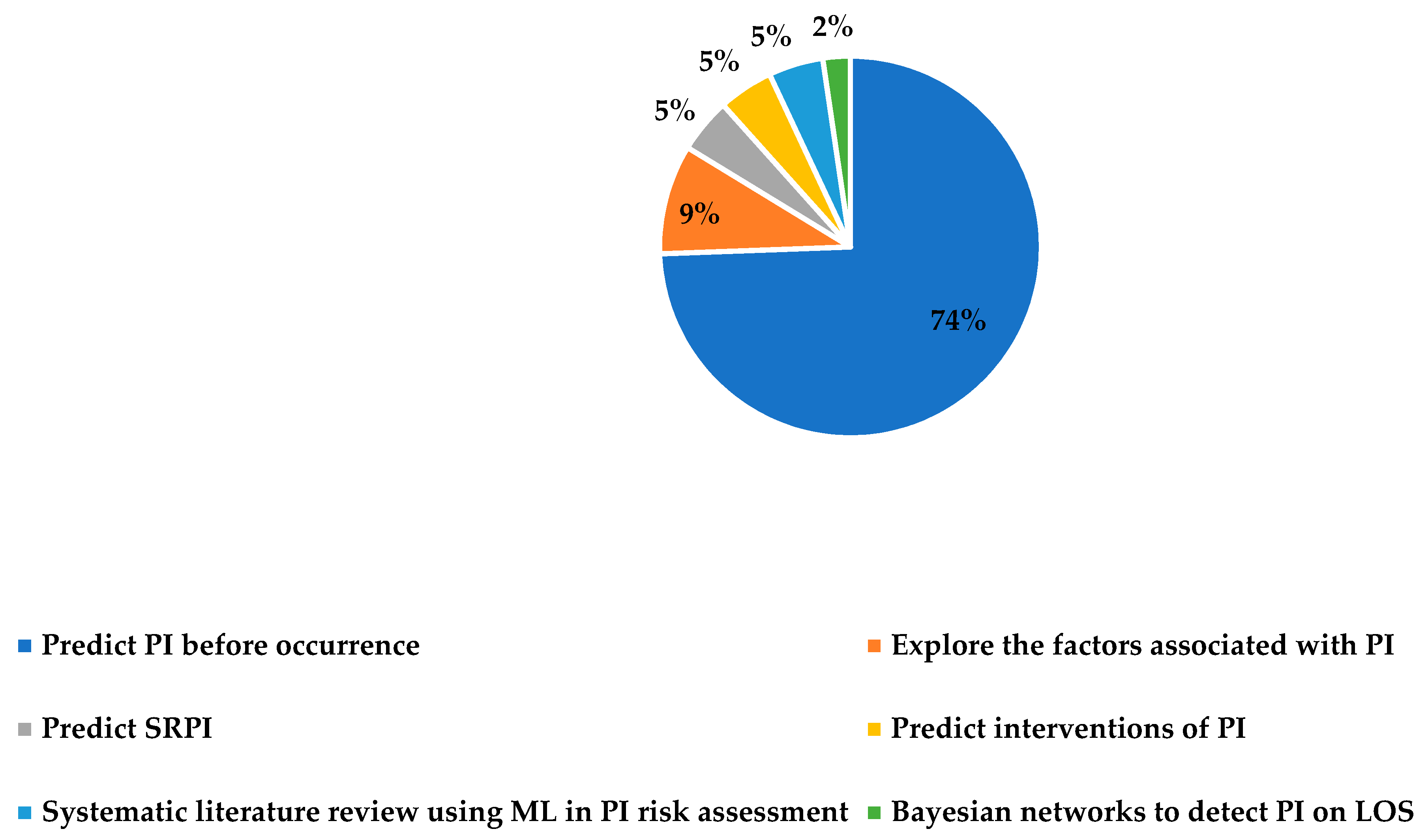
4.1. PI Risk Assessment (Before Occurrence)
4.2. PI Prevention (At Time of Occurrence)
4.3. PI Assessment (After Occurrence)
5. Potential Future Opportunities
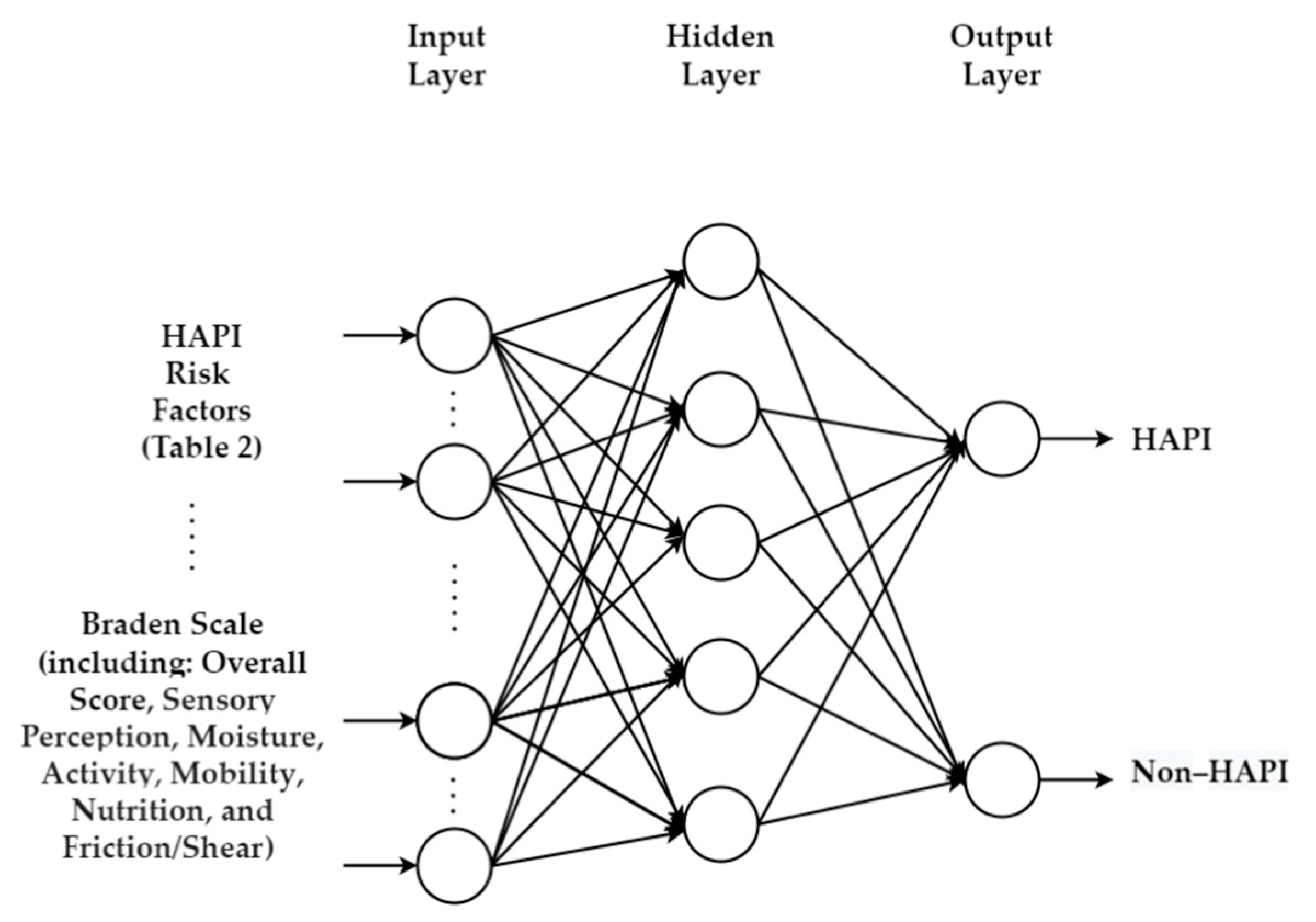
5.1. Integrating Braden Scale with Machine Learning
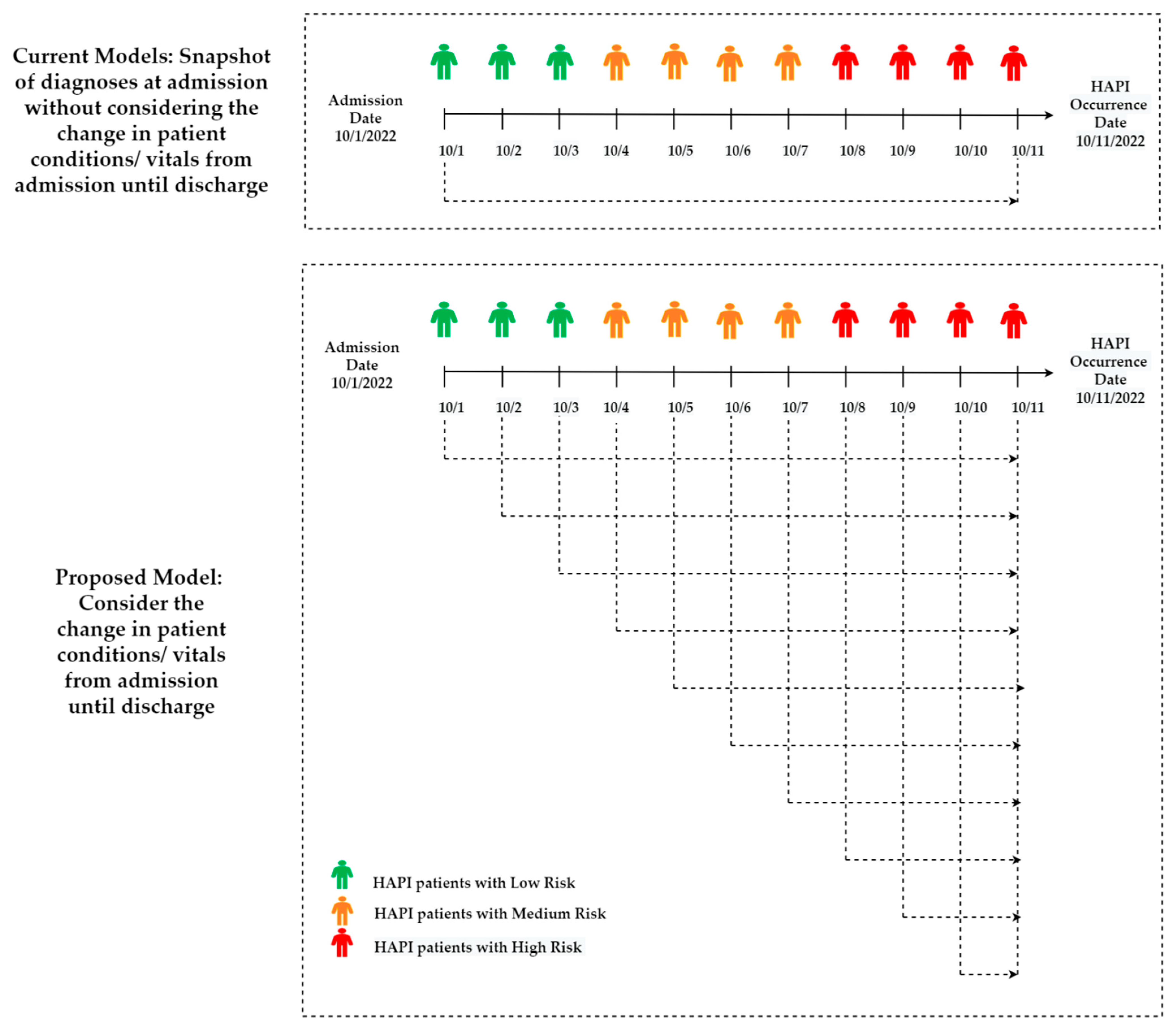
5.2. Utilizing Real-Time/Daily Electronic Health Records
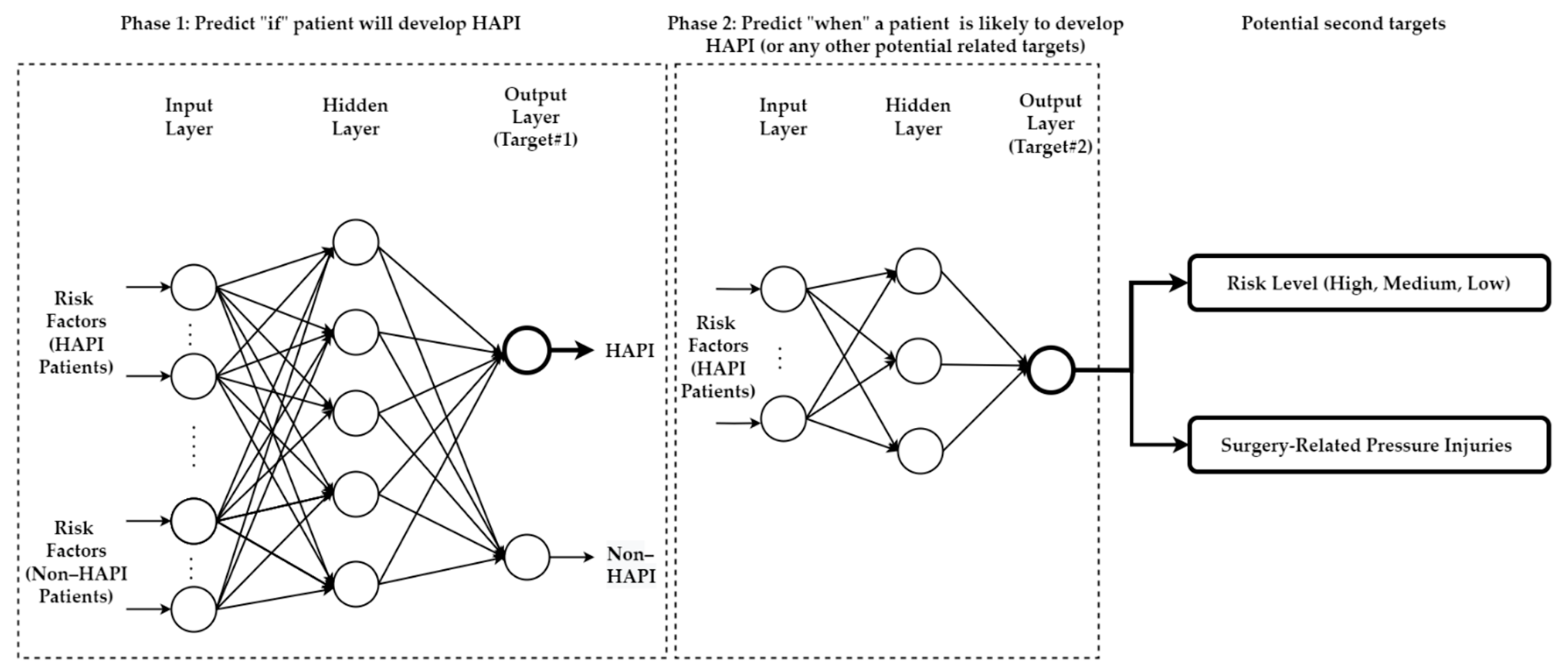
5.3. Predicting Multiple Targets/Outputs
5.4. Hybrid Models
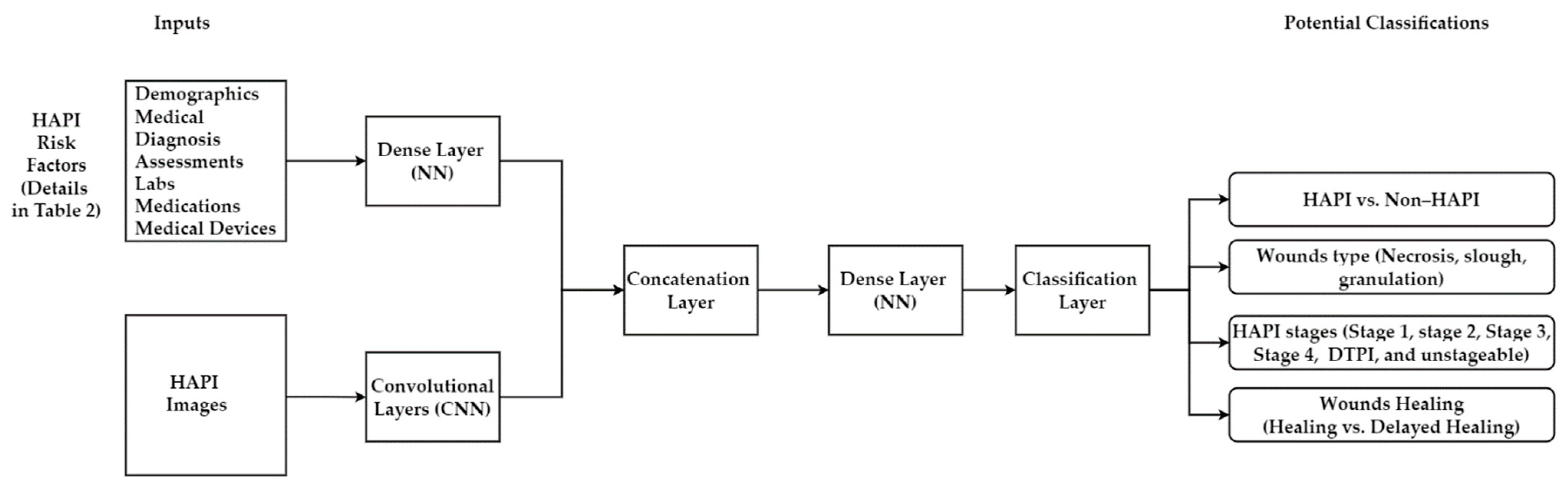
5.4.1. Multimodal Machine Learning
5.4.2. Deep Learning-Machine Learning Hybrid Model
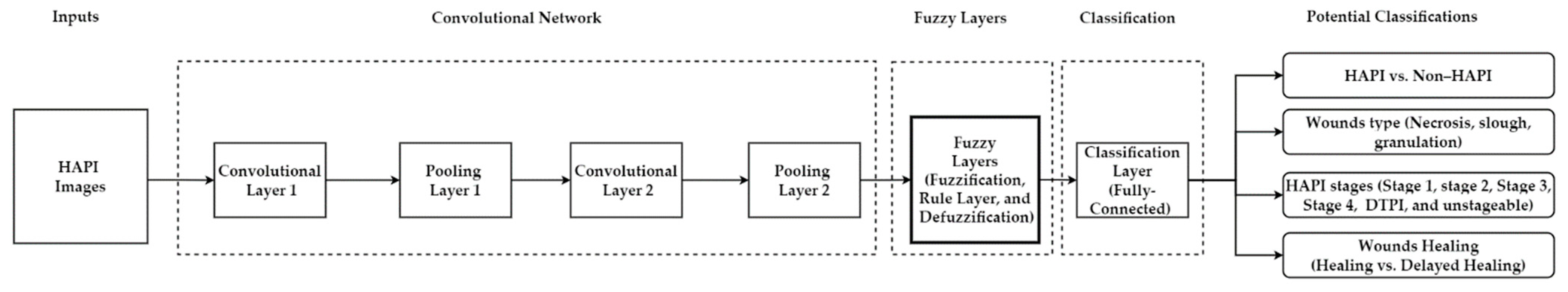
5.4.3. Deep Learning-Fuzzy Logic Hybrid Model
5.4.4. Deep Learning-Transfer Learning Hybrid Model
5.4.5. Deep Learning-Metaheuristics Hybrid Model
5.5. Limitations of Research Method Adopted in This Review
6. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Kottner, J.; Cuddigan, J.; Carville, K.; Balzer, K.; Berlowitz, D.; Law, S.; Litchford, M.; Mitchell, P.; Moore, Z.; Pittman, J.; et al. Prevention and treatment of pressure ulcers/injuries: The protocol for the second update of the international Clinical Practice Guideline 2019. J. Tissue Viability 2019, 28, 51–58. [Google Scholar] [CrossRef] [PubMed]
- Song, W.; Kang, M.-J.; Zhang, L.; Jung, W.; Song, J.; Bates, D.W.; Dykes, P.C. Predicting pressure injury using nursing assessment phenotypes and machine learning methods. J. Am. Med. Inform. Assoc. 2021, 28, 759–765. [Google Scholar] [CrossRef]
- Zahia, S.; Sierra-Sosa, D.; Garcia-Zapirain, B.; Elmaghraby, A. Tissue classification and segmentation of pressure injuries using convolutional neural networks. Comput. Methods Programs Biomed. 2018, 159, 51–58. [Google Scholar] [CrossRef] [PubMed]
- Zahia, S.; Zapirain, M.B.G.; Sevillano, X.; González, A.; Kim, P.J.; Elmaghraby, A. Pressure injury image analysis with machine learning techniques: A systematic review on previous and possible future methods. Artif. Intell. Med. 2020, 102, 101742. [Google Scholar] [CrossRef]
- Mervis, J.S.; Phillips, T.J. Pressure ulcers: Prevention and management. J. Am. Acad. Dermatol. 2019, 81, 893–902. [Google Scholar] [CrossRef]
- Mengist, S.T.; Geletie, H.A.; Zewudie, B.T.; Mewahegn, A.A.; Terefe, T.F.; Amlak, B.T.; Tadesse, B.; GebreEyesus, F.A.; Tsehay, T.; Solomon, M.; et al. Pressure ulcer prevention knowledge, practices, and their associated factors among nurses in Gurage Zone Hospitals, South Ethiopia, 2021. SAGE Open Med. 2022, 10, 205031212211055. [Google Scholar] [CrossRef]
- Anthony, D.M.; Parboteeah, S.; Saleh, M.; Papanikolaou, P. Norton, Waterlow and Braden scores: A review of the literature and a comparison between the scores and clinical judgement. J. Clin. Nurs. 2008, 17, 646–653. [Google Scholar] [CrossRef]
- Flett, H.M.; Delparte, J.J.; Scovil, C.Y.; Higgins, J.; Laramée, M.-T.; Burns, A.S. Determining Pressure Injury Risk on Admission to Inpatient Spinal Cord Injury Rehabilitation: A Comparison of the FIM, Spinal Cord Injury Pressure Ulcer Scale, and Braden Scale. Arch. Phys. Med. Rehabil. 2019, 100, 1881–1887. [Google Scholar] [CrossRef] [PubMed]
- Kim, E.; Choi, M.; Lee, J.; Kim, Y.A. Reusability of EMR Data for Applying Cubbin and Jackson Pressure Ulcer Risk Assessment Scale in Critical Care Patients. Healthc. Inform. Res. 2013, 19, 261–270. [Google Scholar] [CrossRef]
- Kwong, E.; Pang, S.; Wong, T.; Ho, J.; Shao-Ling, X.; Li-Jun, T. Predicting pressure ulcer risk with the modified Braden, Braden, and Norton scales in acute care hospitals in Mainland China. Appl. Nurs. Res. 2005, 18, 122–128. [Google Scholar] [CrossRef]
- Liao, Y.; Gao, G.; Mo, L. Predictive Accuracy of the Braden Q Scale in Risk Assessment for Paediatric Pressure Ulcer: A Me-ta-Analysis. Int. J. Nurs. Sci. 2018, 5, 419–426. [Google Scholar] [CrossRef] [PubMed]
- Riccioni, N.; Berlanga, R.; Hagan, J.; Schier, R.; Gordon, M. Interrater Reliability of the Braden and Braden Q by Skin Champion Nurses. J. Pediatr. Nurs. 2019, 44, 9–15. [Google Scholar] [CrossRef] [PubMed]
- Šáteková, L.; Žiaková, K.; Zeleníková, R. Predictive validity of the Braden scale, Norton scale and Waterlow scale in Slovak Republic. Central Eur. J. Nurs. Midwifery 2015, 6, 283–290. [Google Scholar] [CrossRef]
- Serpa, L.F.; Santos, V.L.; Peres, G.R.; Cavicchioli, M.G.; Hermida, M.M. Validity of the Braden and Waterlow subscales in predicting pressure ulcer risk in hospitalized patients. Appl. Nurs. Res. 2011, 24, e23–e28. [Google Scholar] [CrossRef] [PubMed]
- Wei, M.; Wu, L.; Chen, Y.; Fu, Q.; Chen, W.; Yang, D. Predictive Validity of the Braden Scale for Pressure Ulcer Risk in Critical Care: A Meta-Analysis. Nurs. Crit. Care 2020, 25, 165–170. [Google Scholar] [CrossRef] [PubMed]
- Papanikolaou, P.; Lyne, P.; Anthony, D. Risk assessment scales for pressure ulcers: A methodological review. Int. J. Nurs. Stud. 2007, 44, 285–296. [Google Scholar] [CrossRef] [PubMed]
- Jansen, R.C.S.; Silva, K.B.D.A.; Moura, M.E.S. Braden Scale in pressure ulcer risk assessment. Rev. Bras. de Enferm. 2020, 73, e20190413. [Google Scholar] [CrossRef]
- Watkins, A.A.; Castillo-Angeles, M.; Calvillo-Ortiz, R.; Guetter, C.R.; Eskander, M.F.; Ghaffarpasand, E.; Anguiano-Landa, L.; Tseng, J.F.; Moser, A.J.; Callery, M.P.; et al. Braden scale for pressure ulcer risk predicts rehabilitation placement after pancreatic resection. Hpb 2019, 21, 923–927. [Google Scholar] [CrossRef] [PubMed]
- Gaspar, S.; Peralta, M.; Budri, A.; Ferreira, C.; de Matos, M.G. Pressure ulcer risk profiles of hospitalized patients based on the Braden Scale: A cluster analysis. Int. J. Nurs. Pr. 2022, 28, e13038. [Google Scholar] [CrossRef] [PubMed]
- Grewal, R.; Sklar, M.C.; Almeida, J.R.; Xu, W.; Su, J.; Thomas, C.M.; Alibhai, S.M.; Goldstein, D.P. Evaluation of the Braden scale in predicting surgical outcomes in older patients undergoing major head and neck surgery. Laryngoscope 2021, 6, 103–108. [Google Scholar] [CrossRef] [PubMed]
- Huang, C.; Ma, Y.; Wang, C.; Jiang, M.; Foon, L.Y.; Lv, L.; Han, L. Predictive validity of the braden scale for pressure injury risk assessment in adults: A systematic review and meta-analysis. Nurs. Open 2021, 8, 2194–2207. [Google Scholar] [CrossRef]
- Ribeiro, F.; Fidalgo, F.; Silva, A.; Metrôlho, J.; Santos, O.; Dionisio, R. Literature Review of Machine-Learning Algorithms for Pressure Ulcer Prevention: Challenges and Opportunities. Informatics 2021, 8, 76. [Google Scholar] [CrossRef]
- Jiang, M.; Ma, Y.; Guo, S.; Jin, L.; Lv, L.; Han, L.; An, N. Using Machine Learning Technologies in Pressure Injury Management: Systematic Review. JMIR Med. Inform. 2021, 9, e25704. [Google Scholar] [CrossRef]
- Moher, D.; Liberati, A.; Tetzlaff, J.; Altman, D.G.; PRISMA Group. Preferred reporting items for systematic reviews and meta-analyses: The PRISMA statement. PLoS Med. 2009, 6, e1000097. [Google Scholar] [CrossRef]
- Cai, J.-Y.; Zha, M.-L.; Song, Y.-P.; Chen, H.-L. Predicting the Development of Surgery-Related Pressure Injury Using a Machine Learning Algorithm Model. J. Nurs. Res. 2020, 29, e135. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.-L.; Yu, S.-J.; Xu, Y.; Yu, S.-Q.; Zhang, J.-Q.; Zhao, J.-Y.; Liu, P.; Zhu, B. Artificial Neural Network. J. Wound Ostomy Cont. Nurs. 2018, 45, 26–30. [Google Scholar] [CrossRef] [PubMed]
- Aloweni, F.; Ang, S.Y.; Fook-Chong, S.; Agus, N.; Yong, P.; Goh, M.M.; Tucker-Kellogg, L.; Soh, R.C. A prediction tool for hospital-acquired pressure ulcers among surgical patients: Surgical pressure ulcer risk score. Int. Wound J. 2019, 16, 164–175. [Google Scholar] [CrossRef]
- Moon, M.; Lee, S.-K. Applying of Decision Tree Analysis to Risk Factors Associated with Pressure Ulcers in Long-Term Care Facilities. Healthc. Inform. Res. 2017, 23, 43–52. [Google Scholar] [CrossRef]
- Lu, C.-X.; Chen, H.-L.; Shen, W.-Q.; Feng, L.-P. A new nomogram score for predicting surgery-related pressure ulcers in cardiovascular surgical patients. Int. Wound J. 2016, 14, 226–232. [Google Scholar] [CrossRef]
- Raju, D.; Su, X.; Patrician, P.A.; Loan, L.A.; McCarthy, M.S. Exploring factors associated with pressure ulcers: A data mining approach. Int. J. Nurs. Stud. 2015, 52, 102–111. [Google Scholar] [CrossRef]
- Jin, L.; Pan, Y.; Yang, J.; Han, L.; Lv, L.; Raviv, M.; An, N. Intervention Prediction for Patients with Pressure Injury Using Random Forest. In Proceedings of the Proceedings—12th IEEE International Conference on Big Knowledge, ICBK 2021, Auckland, New Zealand, 7–8 December 2021; Institute of Electrical and Electronics Engineers Inc.: New York City, NY, USA, 2021; pp. 490–496. [Google Scholar]
- Mota, F.; Abreu, N.; Guimarães, T.; Santos, M.F. A Data Mining Study on Pressure Ulcers. In Proceedings of the DATA 2019—Proceedings of the 8th International Conference on Data Science, Technology and Applications, Prague, Czech Republic, 26–28 July 2019; SciTePress: Setúbal, Portugal, 2019; pp. 251–258. [Google Scholar]
- Cho, I.; Park, I.; Kim, E.; Lee, E.; Bates, D.W. Using EHR data to predict hospital-acquired pressure ulcers: A prospective study of a Bayesian Network model. Int. J. Med. Inform. 2013, 82, 1059–1067. [Google Scholar] [CrossRef] [PubMed]
- Nakagami, G.; Yokota, S.; Kitamura, A.; Takahashi, T.; Morita, K.; Noguchi, H.; Ohe, K.; Sanada, H. Supervised machine learning-based prediction for in-hospital pressure injury development using electronic health records: A retrospective observational cohort study in a university hospital in Japan. Int. J. Nurs. Stud. 2021, 119, 103932. [Google Scholar] [CrossRef] [PubMed]
- Walther, F.; Heinrich, L.; Schmitt, J.; Eberlein-Gonska, M.; Roessler, M. Prediction of inpatient pressure ulcers based on routine healthcare data using machine learning methodology. Sci. Rep. 2021, 12, 5044. [Google Scholar] [CrossRef]
- Alderden, J.; Drake, K.P.; Wilson, A.; Dimas, J.; Cummins, M.R.; Yap, T.L. Hospital acquired pressure injury prediction in surgical critical care patients. BMC Med. Inform. Decis. Mak. 2021, 21, 12. [Google Scholar] [CrossRef]
- Anderson, C.; Bekele, Z.; Qiu, Y.; Tschannen, D.; Dinov, I.D. Modeling and prediction of pressure injury in hospitalized patients using artificial intelligence. BMC Med. Inform.Decis. Mak. 2021, 21, 253. [Google Scholar] [CrossRef] [PubMed]
- Ossai, C.I.; O’Connor, L.; Wickramasighe, N. Real-Time Inpatients Risk Profiling in Acute Care: A Comparative Study of Falls and Pressure Injuries Vulnerabilities; University of Maribor: Maribor, Slovenia, 2021; pp. 35–50. [Google Scholar]
- Hu, Y.-H.; Lee, Y.-L.; Kang, M.-F.; Lee, P.-J. Constructing Inpatient Pressure Injury Prediction Models Using Machine Learning Techniques. CIN Comput. Inform. Nurs. 2020, 38, 415–423. [Google Scholar] [CrossRef]
- Vyas, K.; Samadani, A.; Milosevic, M.; Ostadabbas, S.; Parvaneh, S. Additional Value of Augmenting Current Subscales in Braden Scale with Advanced Machine Learning Technique for Pressure Injury Risk Assessment. In Proceedings of the Proceedings—2020 IEEE International Conference on Bioinformatics and Biomedicine, BIBM 2020, Seoul, Republic of Korea, 16–19 December 2020; Institute of Electrical and Electronics Engineers Inc.: New York City, NY, USA, 2020; pp. 2993–2995. [Google Scholar]
- Jin, Y.; Jin, T.; Lee, S.-M. Automated Pressure Injury Risk Assessment System Incorporated into an Electronic Health Record System. Nurs. Res. 2017, 66, 462–472. [Google Scholar] [CrossRef] [PubMed]
- Alderden, J.; Pepper, G.A.; Wilson, A.; Whitney, J.D.; Richardson, S.; Butcher, R.; Jo, Y.; Cummins, M.R. Predicting Pressure Injury in Critical Care Patients: A Machine-Learning Model. Am. J. Crit. Care 2018, 27, 461–468. [Google Scholar] [CrossRef] [PubMed]
- Deng, X.; Yu, T.; Hu, A. Predicting the Risk for Hospital-Acquired Pressure Ulcers in Critical Care Patients. Crit. Care Nurse 2017, 37, e1–e11. [Google Scholar] [CrossRef]
- Ladios-Martin, M.M.; Fernández-De-Maya, J.; Ballesta-López, M.F.-J.; Belso-Garzas, M.A.; Mas-Asencio, M.M.; Cabañero-Martínez, M.J. Predictive Modeling of Pressure Injury Risk in Patients Admitted to an Intensive Care Unit. Am. J. Crit. Care 2020, 29, e70–e80. [Google Scholar] [CrossRef]
- Cramer, E.M.; Seneviratne, M.G.; Sharifi, H.; Ozturk, A.; Hernandez-Boussard, T. Predicting the Incidence of Pressure Ulcers in the Intensive Care Unit Using Machine Learning. eGEMs 2019, 7, 49. [Google Scholar] [CrossRef] [PubMed]
- Gao, L.; Yang, L.; Li, X.; Chen, J.; Du, J.; Bai, X.; Yang, X. The use of a logistic regression model to develop a risk assessment of intraoperatively acquired pressure ulcer. J. Clin. Nurs. 2018, 27, 2984–2992. [Google Scholar] [CrossRef] [PubMed]
- Ahmad, M.A.; Larson, B.; Overman, S.; Kumar, V.; Xie, J.; Rossington, A.; Patel, A.; Teredesai, A. Machine Learning Approaches for Pressure Injury Prediction. In Proceedings of the 2021 IEEE 9th International Conference on Healthcare Informatics, ISCHI 2021, Victoria, BC, Canada, 9–12 August 2021; Institute of Electrical and Electronics Engineers Inc.: New York City, NY, USA, 2021; pp. 427–431. [Google Scholar]
- Song, J.; Gao, Y.; Yin, P.; Li, Y.; Li, Y.; Zhang, J.; Su, Q.; Fu, X.; Pi, H. The Random Forest Model Has the Best Accuracy Among the Four Pressure Ulcer Prediction Models Using Machine Learning Algorithms. Risk Manag. Healthc. Policy 2021, 14, 1175–1187. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.-C.; Wang, P.-C.; Su, C.-T. Pressure Ulcers Prediction Using Support Vector Machines. In Proceedings of the 2008 4th International Conference on Wireless Communications, Networking and Mobile Computing, Dalian, China, 12–14 October 2008; IEEE: New York, NY, USA; pp. 1–4. [Google Scholar]
- Su, C.-T.; Wang, P.-C.; Chen, Y.-C.; Chen, L.-F. Data Mining Techniques for Assisting the Diagnosis of Pressure Ulcer Development in Surgical Patients. J. Med. Syst. 2012, 36, 2387–2399. [Google Scholar] [CrossRef] [PubMed]
- Li, H.-L.; Lin, S.-W.; Hwang, Y.-T. Using Nursing Information and Data Mining to Explore the Factors That Predict Pressure Injuries for Patients at the End of Life. CIN: Comput. Inform. Nurs. 2019, 37, 133–141. [Google Scholar] [CrossRef] [PubMed]
- Cichosz, S.L.; Voelsang, A.-B.; Tarnow, L.; Hasenkam, J.M.; Fleischer, J. Prediction of In-Hospital Pressure Ulcer Development. Adv. Wound Care 2019, 8, 1–6. [Google Scholar] [CrossRef] [PubMed]
- Šín, P.; Hokynková, A.; Marie, N.; Andrea, P.; Krč, R.; Podroužek, J. Machine Learning-Based Pressure Ulcer Prediction in Modular Critical Care Data. Diagnostics 2022, 12, 850. [Google Scholar] [CrossRef]
- Xu, J.; Chen, D.; Deng, X.; Pan, X.; Chen, Y.; Zhuang, X.; Sun, C. Development and validation of a machine learning algorithm–based risk prediction model of pressure injury in the intensive care unit. Int. Wound J. 2022. [Google Scholar] [CrossRef]
- Do, Q.; Lipatov, K.; Ramar, K.; Rasmusson, J.; Pickering, B.W.; Herasevich, V. Pressure Injury Prediction Model Using Advanced Analytics for At-Risk Hospitalized Patients. J. Patient Saf. 2022, 18, e1083–e1089. [Google Scholar] [CrossRef]
- Kaewprag, P.; Newton, C.; Vermillion, B.; Hyun, S.; Huang, K.; Machiraju, R. Predictive models for pressure ulcers from intensive care unit electronic health records using Bayesian networks. BMC Med. Inform. Decis. Mak. 2017, 17, 81–91. [Google Scholar] [CrossRef] [PubMed]
- Hyun, S.; Moffatt-Bruce, S.; Cooper, C.; Hixon, B.; Kaewprag, P. Prediction Model for Hospital-Acquired Pressure Ulcer Development: Retrospective Cohort Study. JMIR Med. Inform. 2019, 7, e13785. [Google Scholar] [CrossRef]
- Borlawsky, T.; Hripcsak, G. Evaluation of an Automated Pressure Ulcer Risk Assessment Model. Home Healthc. Care Manag. Pr. 2007, 19, 272–284. [Google Scholar] [CrossRef]
- Levy, J.J.; Lima, J.F.; Miller, M.W.; Freed, G.L.; James O’malley, A.; Emeny, R.T. Investigating the Potential for Machine Learning Prediction of Patient Outcomes: A Retrospective Study of Hospital Acquired Pressure Injuries. medRxiv 2020. [Google Scholar] [CrossRef]
- Setoguchi, Y.; Ghaibeh, A.A.; Mitani, K.; Abe, Y.; Hashimoto, I.; Moriguchi, H. Predictability of Pressure Ulcers Based on Operation Duration, Transfer Activity, and Body Mass Index Through the Use of an Alternating Decision Tree. J. Med. Investig. 2016, 63, 248–255. [Google Scholar] [CrossRef][Green Version]
- Cheng, F.M.; Jin, Y.J.; Chien, C.W.; Chuang, Y.C.; Tung, T.H. The Application of Braden Scale and Rough Set Theory for Pressure Injury Risk in Elderly Male Population. J. Mens. Health 2021, 17, 156–165. [Google Scholar] [CrossRef]
- Charon, C.; Wuillemin, P.-H.; Belmin, J. Learning Bayesian Networks for the Prediction of Unfavorable Health Events in Nursing Homes. Stud. Health Technol. Inform. 2022, 294, 147–148. [Google Scholar] [CrossRef]
- Lee, S.-K.; Shin, J.; Ahn, J.; Lee, J.; Jang, D. Identifying the Risk Factors Associated with Nursing Home Residents’ Pressure Ulcers Using Machine Learning Methods. Int. J. Environ. Res. Public Healthc. 2021, 18, 2954. [Google Scholar] [CrossRef]
- Wang, Y.; Jiang, X.; Yu, K.; Shi, F.; Qin, L.; Zhou, H.; Cai, F. Infrared Thermal Images Classification for Pressure Injury Prevention Incorporating the Convolutional Neural Networks. IEEE Access 2021, 9, 15181–15190. [Google Scholar] [CrossRef]
- Iranmehr, A.; Masnadi-Shirazi, H.; Vasconcelos, N. Cost-sensitive support vector machines. Neurocomputing 2019, 343, 50–64. [Google Scholar] [CrossRef]
- Johnson, A.E.W.; Pollard, T.J.; Shen, L.; Lehman, L.-W.H.; Feng, M.; Ghassemi, M.; Moody, B.; Szolovits, P.; Celi, L.A.; Mark, R.G. MIMIC-III, a freely accessible critical care database. Sci. Data 2016, 3, 160035. [Google Scholar] [CrossRef] [PubMed]
- Chiang, J.-C.; Lie, W.-N.; Huang, H.-C.; Chen, K.-T.; Liang, J.-Y.; Lo, Y.-C.; Huang, W.-H. Posture Monitoring for Health Care of Bedridden Elderly Patients Using 3D Human Skeleton Analysis via Machine Learning Approach. Appl. Sci. 2022, 12, 3087. [Google Scholar] [CrossRef]
- Cicceri, G.; De Vita, F.; Bruneo, D.; Merlino, G.; Puliafito, A. A deep learning approach for pressure ulcer prevention using wearable computing. Human-Centric Comput. Inf. Sci. 2020, 10, 5. [Google Scholar] [CrossRef]
- Heydarzadeh, M.; Nourani, M.; Ostadabbas, S. In-Bed Posture Classification Using Deep Autoencoders. In Proceedings of the Proceedings of the Annual International Conference of the IEEE Engineering in Medicine and Biology Society, EMBS; Montreal, QC, Canada, 16–20 August 2016, Institute of Electrical and Electronics Engineers Inc.: New York City, NY, USA, 2016; pp. 3839–3842. [Google Scholar]
- Matar, G.; Lina, J.-M.; Kaddoum, G. Artificial Neural Network for in-Bed Posture Classification Using Bed-Sheet Pressure Sensors. IEEE J. Biomed. Health Inform. 2020, 24, 101–110. [Google Scholar] [CrossRef] [PubMed]
- Duvall, J.; Karg, P.; Brienza, D.; Pearlman, J. Detection and classification methodology for movements in the bed that supports continuous pressure injury risk assessment and repositioning compliance. J. Tissue Viability 2019, 28, 7–13. [Google Scholar] [CrossRef] [PubMed]
- Enayati, M.; Skubic, M.; Keller, J.M.; Popescu, M.; Zanjirani Farahani, N. Sleep Posture Classification Using Bed Sensor Data and Neural Networks. In Proceedings of the 2018 40th Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC), Honolulu, HI, USA, 18–21 July 2018; ISBN 9781538636466. [Google Scholar]
- Xu, X.; Lin, F.; Wang, A.; Hu, Y.; Huang, M.-C.; Xu, W. Body-Earth Mover’s Distance: A Matching-Based Approach for Sleep Posture Recognition. IEEE Trans. Biomed. Circuits Syst. 2016, 10, 1023–1035. [Google Scholar] [CrossRef]
- Pouyan, M.B.; Birjandtalab, J.; Nourani, M.; Pompeo, M.M. Automatic limb identification and sleeping parameters assessment for pressure ulcer prevention. Comput. Biol. Med. 2016, 75, 98–108. [Google Scholar] [CrossRef] [PubMed]
- Hsiao, R.-S.; Mi, Z.; Yang, B.-R.; Kau, L.-J.; Bitew, M.A.; Li, T.-Y. Body posture recognition and turning recording system for the care of bed bound patients. Technol. Health Care 2016, 24, S307–S312. [Google Scholar] [CrossRef] [PubMed]
- Pouyan, M.B.; Ostadabbas, S.; Nourani, M.; Pompeo, M. Classifying Bed Inclination Using Pressure Images. In Proceedings of the 2014 36th Annual International Conference of the IEEE Engineering in Medicine and Biology Society, EMBC 2014, Chicago, IL, USA, 26–30 August 2014; Institute of Electrical and Electronics Engineers Inc.: New York City, NY, USA, 2014; pp. 4663–4666. [Google Scholar]
- Barsocchi, P. Position Recognition to Support Bedsores Prevention. IEEE J. Biomed. Healthc. Inform. 2013, 17, 53–59. [Google Scholar] [CrossRef]
- Jaffery, M.H.; Ashraf, M.A.; Almogren, A.; Asim, H.M.; Arshad, J.; Khan, J.; Rehman, A.U.; Hussen, S. FSR-Based Smart System for Detection of Wheelchair Sitting Postures Using Machine Learning Algorithms and Techniques. J. Sens. 2022, 2022, 1901058. [Google Scholar] [CrossRef]
- Ma, C.; Li, W.; Gravina, R.; Fortino, G. Posture Detection Based on Smart Cushion for Wheelchair Users. Sensors 2017, 17, 719. [Google Scholar] [CrossRef] [PubMed]
- Grunerbel, L.; Heinrich, F.; Diebolder, D.; Richter, M. Wearable Decubitus Prophylaxis Tool Based on Machine Learning Methods; Institute of Electrical and Electronics Engineers (IEEE): New York City, NY, USA, 2022; pp. 730–734. [Google Scholar]
- Luboz, V.; Bailet, M.; Grivot, C.B.; Rochette, M.; Diot, B.; Bucki, M.; Payan, Y. Personalized modeling for real-time pressure ulcer prevention in sitting posture. J. Tissue Viability 2018, 27, 54–58. [Google Scholar] [CrossRef]
- McNichol, L.L.; Ratliff, C.R.; Yates, S.S. Wound, Ostomy and Continence Nurses Society Core Curriculum: Wound Management, 2nd ed.; Lippincott Williams & Wilkins: Philadelphia, PA, USA, 2021. [Google Scholar]
- Ramachandram, D.; Ramirez-GarciaLuna, J.L.; Fraser, R.D.J.; Martínez-Jiménez, M.A.; E Arriaga-Caballero, J.; Allport, J. Fully Automated Wound Tissue Segmentation Using Deep Learning on Mobile Devices: Cohort Study. JMIR mHealth uHealth 2022, 10, e36977. [Google Scholar] [CrossRef] [PubMed]
- Chang, C.W.; Christian, M.; Chang, D.H.; Lai, F.; Liu, T.J.; Chen, Y.S.; Chen, W.J. Deep learning approach based on superpixel segmentation assisted labeling for automatic pressure ulcer diagnosis. PLoS ONE 2022, 17, e0264139. [Google Scholar] [CrossRef]
- Howell, R.S.; Liu, H.H.; Khan, A.A.; Woods, J.S.; Lin, L.J.; Saxena, M.; Saxena, H.; Castellano, M.; Petrone, P.; Slone, E.; et al. Development of a Method for Clinical Evaluation of Artificial Intelligence–Based Digital Wound Assessment Tools. JAMA Netw. Open 2021, 4, e217234. [Google Scholar] [CrossRef]
- Wang, C.; Anisuzzaman, D.M.; Williamson, V.; Dhar, M.K.; Rostami, B.; Niezgoda, J.; Gopalakrishnan, S.; Yu, Z. Fully automatic wound segmentation with deep convolutional neural networks. Sci. Rep. 2020, 10, 21897. [Google Scholar] [CrossRef] [PubMed]
- Ohura, N.; Mitsuno, R.; Sakisaka, M.; Terabe, Y.; Morishige, Y.; Uchiyama, A.; Okoshi, T.; Shinji, I.; Takushima, A. Convolutional neural networks for wound detection: The role of artificial intelligence in wound care. J. Wound Care 2019, 28, S13–S24. [Google Scholar] [CrossRef]
- García-Zapirain, B.; Elmogy, M.; El-Baz, A.; Elmaghraby, A.S. Classification of pressure ulcer tissues with 3D convolutional neural network. Med. Biol. Eng. Comput. 2018, 56, 2245–2258. [Google Scholar] [CrossRef] [PubMed]
- Garcia-Zapirain, B.; Shalaby, A.; El-Baz, A.; Elmaghraby, A. Automated framework for accurate segmentation of pressure ulcer images. Comput. Biol. Med. 2017, 90, 137–145. [Google Scholar] [CrossRef]
- Veredas, F.J.; Luque-Baena, R.M.; Martín-Santos, F.J.; Morilla-Herrera, J.C.; Morente, L. Wound image evaluation with machine learning. Neurocomputing 2015, 164, 112–122. [Google Scholar] [CrossRef]
- Wannous, H.; Treuillet, S.; Lucas, Y. Supervised Tissue Classification from Color Images for a Complete Wound Assessment Tool. Annu. Int. Conf. IEEE Eng. Med. Biol. Soc. 2007, 2007, 6031–6034. [Google Scholar] [CrossRef]
- Veredas, F.; Mesa, H.; Morente, L. Binary Tissue Classification on Wound Images with Neural Networks and Bayesian Classifiers. IEEE Trans. Med. Imaging 2010, 29, 410–427. [Google Scholar] [CrossRef] [PubMed]
- Ay, B.; Tasar, B.; Utlu, Z.; Ay, K.; Aydin, G. Deep transfer learning-based visual classification of pressure injuries stages. Neural Comput. Appl. 2022, 34, 16157–16168. [Google Scholar] [CrossRef]
- Fergus, P.; Chalmers, C.; Henderson, W.; Roberts, D.; Waraich, A. Pressure Ulcer Categorisation Using Deep Learning: A Clinical Trial to Evaluate Model Performance. arXiv 2022, arXiv:2203.06248. [Google Scholar]
- Liu, T.J.; Christian, M.; Chu, Y.-C.; Chen, Y.-C.; Chang, C.-W.; Lai, F.; Tai, H.-C. A pressure ulcers assessment system for diagnosis and decision making using convolutional neural networks. J. Formos. Med. Assoc. 2022, 121, 2227–2236. [Google Scholar] [CrossRef] [PubMed]
- Anisuzzaman, D.M.; Patel, Y.; Rostami, B.; Niezgoda, J.; Gopalakrishnan, S.; Yu, Z. Multi-modal wound classification using wound image and location by deep neural network. Sci. Rep. 2022, 12, 20057. [Google Scholar] [CrossRef] [PubMed]
- Matsumoto, M.; Karube, M.; Nakagami, G.; Kitamura, A.; Tamai, N.; Miura, Y.; Kawamoto, A.; Kurita, M.; Miyake, T.; Hayashi, C.; et al. Development of an Automatic Ultrasound Image Classification System for Pressure Injury Based on Deep Learning. Appl. Sci. 2021, 11, 7817. [Google Scholar] [CrossRef]
- Yilmaz, A.; Kızıl, H.; Kaya, U.; Çakır, R.; Demiral, M. Prediction and classification of pressure injuries by deep learning. Healthc. Probl. Civiliz. 2021, 15, 328–335. [Google Scholar] [CrossRef]
- Yilmaz, B.; Atagun, E.; Demircan, F.O.; Yucedag, I. Classification of Pressure Ulcer Images with Logistic Regression. In Proceedings of the 2021 International Conference on INnovations in Intelligent SysTems and Applications, INISTA 2021—Proceedings, Kocaeli, Turkey, 25–27 August 2021; Institute of Electrical and Electronics Engineers Inc.: New York City, NY, USA, 2021. [Google Scholar]
- Mombini, H.; Tulu, B.; Strong, D.; Agu, E.; Nguyen, H.; Lindsay, C.; Loretz, L.; Pedersen, P. Design of a Machine Learning System for Prediction of Chronic Wound Management Decisions. In International Conference on Design Science Research in Information Systems and Technology; Springer: Cham, Switzerland, 2020; pp. 15–27. [Google Scholar]
- Chang, D.H.; Chu, P.J.; Li, Y.J.; Ning, C.K.; Chien, T.Y. A Clinical Decision Support System of Pressure Ulcers Tissue Classification. In ACM International Conference Proceeding Series; Association for Computing Machinery: New York City, NY, USA, 2021; pp. 338–343. [Google Scholar]
- Kavitha, I.; Suganthi, S.S.; Ramakrishnan, S. Analysis of Chronic Wound Images Using Factorization Based Segmentation and Machine Learning Methods. In ACM International Conference Proceeding Series; Association for Computing Machinery: New York City, NY, USA, 2017; pp. 74–78. [Google Scholar]
- Lustig, M.; Schwartz, D.; Bryant, R.; Gefen, A. A machine learning algorithm for early detection of heel deep tissue injuries based on a daily history of sub-epidermal moisture measurements. Int. Wound J. 2022, 19, 1339–1348. [Google Scholar] [CrossRef] [PubMed]
- Chun, X.; Pan, L.; Lin, Y.; Ye, L.; Liang, H.; Tao, J.; Luo, Y. A model for predicting 7-day pressure injury outcomes in paediatric patients: A machine learning approach. J. Adv. Nurs. 2021, 77, 1304–1314. [Google Scholar] [CrossRef] [PubMed]
- Veredas, F.J.; Mesa, H.; Morilla, J.C.; Morente, L. Predicting the Evolution of Pressure Ulcers. In Proceedings of the HEALTHINF 2010—3rd International Conference on Health Informatics, Valencia, Spain, 20–23 January 2010; pp. 5–12. [Google Scholar]
- e Silva, R.H.L.; Machado, A.M.C. Automatic measurement of pressure ulcers using Support Vector Machines and GrabCut. Comput. Methods Programs Biomed. 2021, 200, 105867. [Google Scholar] [CrossRef] [PubMed]
- Li, D.; Mathews, C. Automated measurement of pressure injury through image processing. J. Clin. Nurs. 2017, 26, 3564–3575. [Google Scholar] [CrossRef]
- Zahia, S.; Garcia-Zapirain, B.; Elmaghraby, A. Integrating 3D Model Representation for an Accurate Non-Invasive Assessment of Pressure Injuries with Deep Learning. Sensors 2020, 20, 2933. [Google Scholar] [CrossRef]
- Chino, D.Y.; Scabora, L.C.; Cazzolato, M.T.; Jorge, A.E.; Traina, C., Jr.; Traina, A.J. Segmenting skin ulcers and measuring the wound area using deep convolutional networks. Comput. Methods Programs Biomed. 2020, 191, 105376. [Google Scholar] [CrossRef]
- Veredas, F.J.; Mesa, H.; Morente, L. Tissue Recognition Approach to Pressure Ulcer Area Estimation with Neural Networks. In Proceedings of the International Work-Conference on Artificial Neural Networks, Salamanca, Spain, 10–12 June 2009; pp. 1045–1052. [Google Scholar] [CrossRef]
- Chang, M.-C.; Yu, T.; Luo, J.; Duan, K.; Tu, P.; Zhao, Y.; Nagraj, N.; Rajiv, V.; Priebe, M.; Wood, E.A.; et al. Multimodal Sensor System for Pressure Ulcer Wound Assessment and Care. IEEE Trans. Ind. Inform. 2018, 14, 1186–1196. [Google Scholar] [CrossRef]
- Kaswan, K.E.; Jamil, N.; Roslan, R. Deep Learning on Wound Segmentation and Classification: A Short Review and Evaluation of Methods Used. Eur. J. Soft Comput. 2020, 9, 6–10. [Google Scholar]
- Metz, L.; Brain, G.; Poole, B.; Pfau, D.; Deepmind, G.; Sohl-Dickstein, J. Unrolled Generative Adversarial Networks. arXiv 2016, arXiv:1611.02163. [Google Scholar]
- Pan, Z.; Yu, W.; Yi, X.; Khan, A.; Yuan, F.; Zheng, Y. Recent Progress on Generative Adversarial Networks (GANs): A Survey. IEEE Access 2019, 7, 36322–36333. [Google Scholar] [CrossRef]
- National Pressure Injury Advisory Panel (NPIAP) Pressure Injury Fact Sheet. Available online: https://npiap.com/store/viewproduct.aspx?id=14427618 (accessed on 12 November 2022).
- Jena, B.; Saxena, S.; Nayak, G.K.; Saba, L.; Sharma, N.; Suri, J.S. Artificial intelligence-based hybrid deep learning models for image classification: The first narrative review. Comput. Biol. Med. 2021, 137, 104803. [Google Scholar] [CrossRef] [PubMed]
- Yin, Q.; Zhang, R.; Shao, X. CNN and RNN mixed model for image classification. MATEC Web Conf. 2019, 277, 02001. [Google Scholar] [CrossRef]
- Bharati, S.; Podder, P.; Mondal, M.R.H. Hybrid deep learning for detecting lung diseases from X-ray images. Inform. Med. Unlocked 2020, 20, 100391. [Google Scholar] [CrossRef]
- Rezaee, K.; Badiei, A.; Meshgini, S. A Hybrid Deep Transfer Learning Based Approach for COVID-19 Classification in Chest X-Ray Images; A Hybrid Deep Transfer Learning Based Approach for COVID-19 Classification in Chest X-Ray Images. In Proceedings of the 2020 27th National and 5th International Iranian Conference on Biomedical Engineering (ICBME), Tehran, Iran, 26–27 November 2020. [Google Scholar] [CrossRef]
- Deepak, S.; Ameer, P. Brain tumor classification using deep CNN features via transfer learning. Comput. Biol. Med. 2019, 111, 103345. [Google Scholar] [CrossRef]
- Shakarami, A.; Tarrah, H.; Mahdavi-Hormat, A. A CAD system for diagnosing Alzheimer’s disease using 2D slices and an improved AlexNet-SVM method. Optik 2020, 212, 164237. [Google Scholar] [CrossRef]
- Elkorany, A.S.; Elsharkawy, Z.F. COVIDetection-Net: A tailored COVID-19 detection from chest radiography images using deep learning. Optik 2021, 231, 166405. [Google Scholar] [CrossRef] [PubMed]
- Carvalho, E.D.; Silva, R.R.; Araújo, F.H.; Rabelo, R.D.A.; Filho, A.O.D.C. An approach to the classification of COVID-19 based on CT scans using convolutional features and genetic algorithms. Comput. Biol. Med. 2021, 136, 104744. [Google Scholar] [CrossRef]
- Dweekat, O.Y.; Bazrafshan, N.; Lu, S.S. Heart Disease Prediction and Factor Identification Using Machine Learning Algorithms and Association Rule Mining. In Proceedings of the IISE Annual Conference & Expo 2022, Seattle, WA, USA, 21–24 May 2022; Institute of Industrial & Systems Engineers (IISE): Seattle, WA, USA, 2022; Volume 1, pp. 79–84. [Google Scholar]
- Gessert, N.; Nielsen, M.; Shaikh, M.; Werner, R.; Schlaefer, A. Skin lesion classification using ensembles of multi-resolution EfficientNets with meta data. Methodsx 2020, 7, 100864. [Google Scholar] [CrossRef] [PubMed]
- Sun, Q.; Huang, C.; Chen, M.; Xu, H.; Yang, Y. Skin Lesion Classification Using Additional Patient Information. BioMed Res. Int. 2021, 2021, 1–6. [Google Scholar] [CrossRef]
- Yap, J.; Yolland, W.; Tschandl, P. Multimodal skin lesion classification using deep learning. Exp. Dermatol. 2018, 27, 1261–1267. [Google Scholar] [CrossRef]
- Thomas, S.A. Combining Image Features and Patient Metadata to Enhance Transfer Learning. Annu. Int. Conf. IEEE Eng. Med. Biol. Soc. 2021, 2021, 2660–2663. [Google Scholar] [CrossRef]
- Chieregato, M.; Frangiamore, F.; Morassi, M.; Baresi, C.; Nici, S.; Bassetti, C.; Bnà, C.; Galelli, M. A hybrid machine learning/deep learning COVID-19 severity predictive model from CT images and clinical data. Sci. Rep. 2022, 12, 1–15. [Google Scholar] [CrossRef]
- Zhang, F.; Li, Z.; Zhang, B.; Du, H.; Wang, B.; Zhang, X. Multi-modal deep learning model for auxiliary diagnosis of Alzheimer’s disease. Neurocomputing 2019, 361, 185–195. [Google Scholar] [CrossRef]
- Rambhatla, S.; Huang, S.; Trinh, L.; Eng, M.; Zhang, M.; Long, B.; Dong, M.; Unadkat, V.; Haig, A.Y.; Gillenwater, J.; et al. DL4Burn: Burn Surgical Candidacy Prediction Using Multimodal Deep Learning. In MIA Annual Symposium Proceedings; American Medical Informatics Association: Bethesda, MD, USA, 2022. [Google Scholar]
- Polat, H.; Danaei Mehr, H. Classification of Pulmonary CT Images by Using Hybrid 3D-Deep Convolutional Neural Network Architecture. Appl. Sci. 2019, 9, 940. [Google Scholar] [CrossRef]
- Mahum, R.; Rehman, S.U.; Okon, O.D.; Alabrah, A.; Meraj, T.; Rauf, H.T. A Novel Hybrid Approach Based on Deep CNN to Detect Glaucoma Using Fundus Imaging. Electronics 2021, 11, 26. [Google Scholar] [CrossRef]
- Khairandish, M.; Sharma, M.; Jain, V.; Chatterjee, J.; Jhanjhi, N. A Hybrid CNN-SVM Threshold Segmentation Approach for Tumor Detection and Classification of MRI Brain Images. IRBM 2021, 43, 290–299. [Google Scholar] [CrossRef]
- Zheng, Y.; Xu, Z.; Wang, X. The Fusion of Deep Learning and Fuzzy Systems: A State-of-the-Art Survey. IEEE Trans. Fuzzy Syst. 2021, 30, 2783–2799. [Google Scholar] [CrossRef]
- Averkin, A. Hybrid Intelligent Systems Based on Fuzzy Logic and Deep Learning. In Artificial Intelligence; Lecture Notes in Computer Science (including subseries Lecture Notes in Artificial Intelligence and Lecture Notes in Bioinformatics); Springer: Cham, Switzerland, 2019; pp. 3–12. [Google Scholar] [CrossRef]
- Das, R.; Sen, S.; Maulik, U. A Survey on Fuzzy Deep Neural Networks. ACM Comput. Surv. 2020, 53, 1–25. [Google Scholar] [CrossRef]
- Price, S.R.; Price, S.R.; Anderson, D.T. Introducing Fuzzy Layers for Deep Learning. In Proceedings of the IEEE International Conference on Fuzzy Systems 2019, New Orleans, LA, USA, 23–26 June 2019. [Google Scholar] [CrossRef]
- Korshunova, K.P. A Convolutional Fuzzy Neural Network for Image Classification. In Proceedings of the RPC 2018—3rd Russian-Pacific Conference on Computer Technology and Applications 2018, Vladivostok, Russia, 18–25 August 2018. [Google Scholar] [CrossRef]
- Molaei, S.; Ghorbani, N.; Dashtiahangar, F.; Peivandi, M.; Pourasad, Y.; Esmaeili, M. FDCNet: Presentation of the Fuzzy CNN and Fractal Feature Extraction for Detection and Classification of Tumors. Comput. Intell. Neurosci. 2022, 2022, 7543429. [Google Scholar] [CrossRef]
- Dey, S.; Bhattacharya, R.; Malakar, S.; Schwenker, F.; Sarkar, R. CovidConvLSTM: A fuzzy ensemble model for COVID-19 detection from chest X-rays. Expert Syst. Appl. 2022, 206, 117812. [Google Scholar] [CrossRef]
- Ieracitano, C.; Mammone, N.; Versaci, M.; Varone, G.; Ali, A.-R.; Armentano, A.; Calabrese, G.; Ferrarelli, A.; Turano, L.; Tebala, C.; et al. A fuzzy-enhanced deep learning approach for early detection of Covid-19 pneumonia from portable chest X-ray images. Neurocomputing 2022, 481, 202–215. [Google Scholar] [CrossRef]
- Banerjee, S.; Singh, S.K.; Chakraborty, A.; Das, A.; Bag, R. Melanoma Diagnosis Using Deep Learning and Fuzzy Logic. Diagnostics 2020, 10, 577. [Google Scholar] [CrossRef] [PubMed]
- Nida, N.; Irtaza, A.; Javed, A.; Yousaf, M.H.; Mahmood, M.T. Melanoma lesion detection and segmentation using deep region based convolutional neural network and fuzzy C-means clustering. Int. J. Med. Inf. 2019, 124, 37–48. [Google Scholar] [CrossRef] [PubMed]
- Bhimavarapu, U.; Battineni, G. Skin Lesion Analysis for Melanoma Detection Using the Novel Deep Learning Model Fuzzy GC-SCNN. Healthcare 2022, 10, 962. [Google Scholar] [CrossRef]
- Badawy, S.M.; Mohamed, A.E.-N.A.; Hefnawy, A.A.; Zidan, H.E.; GadAllah, M.T.; El-Banby, G.M. Automatic semantic segmentation of breast tumors in ultrasound images based on combining fuzzy logic and deep learning—A feasibility study. PLoS ONE 2021, 16, e0251899. [Google Scholar] [CrossRef] [PubMed]
- Huang, K.; Zhang, Y.; Cheng, H.; Xing, P.; Zhang, B. Semantic segmentation of breast ultrasound image with fuzzy deep learning network and breast anatomy constraints. Neurocomputing 2021, 450, 319–335. [Google Scholar] [CrossRef]
- Liang, H.; Fu, W.; Yi, F. A Survey of Recent Advances in Transfer Learning. In Proceedings of the 2019 IEEE 19th International Conference on Communication Technology (ICCT), Xi’an, China, 16–19 October 2019. [Google Scholar]
- Hosny, K.M.; Kassem, M.A.; Fouad, M.M. Classification of Skin Lesions into Seven Classes Using Transfer Learning with AlexNet. J. Digit. Imaging 2020, 33, 1325–1334. [Google Scholar] [CrossRef] [PubMed]
- Lu, S.; Lu, Z.; Zhang, Y.-D. Pathological brain detection based on AlexNet and transfer learning. J. Comput. Sci. 2019, 30, 41–47. [Google Scholar] [CrossRef]
- Ferreira, C.A.; Melo, T.; Sousa, P.; Meyer, M.I.; Shakibapour, E.; Costa, P.; Campilho, A. Classification of Breast Cancer Histology Images Through Transfer Learning Using a Pre-Trained Inception Resnet V2. In Proceedings of the Lecture Notes in Computer Science (Including Subseries Lecture Notes in Artificial Intelligence and Lecture Notes in Bioinformatics); Springer: Berlin/Heidelberg, Germany, 2018; Volume 10882 LNCS, pp. 763–770. [Google Scholar]
- Liu, Y.; Du, B.; Ni, F. Adversarial strategy for transductive zero-shot learning. Inf. Sci. 2021, 578, 750–761. [Google Scholar] [CrossRef]
- Akay, B.; Karaboga, D.; Akay, R. A comprehensive survey on optimizing deep learning models by metaheuristics. Artif. Intell. Rev. 2021, 55, 829–894. [Google Scholar] [CrossRef]
- Dweekat, O.Y.; Lam, S.S. Cervical Cancer Diagnosis Using an Integrated System of Principal Component Analysis, Genetic Algorithm, and Multilayer Perceptron. Healthcare 2022, 10, 2002. [Google Scholar] [CrossRef]
- Devi, M.A.; Ravi, S.; Vaishnavi, J.; Punitha, S. Classification of Cervical Cancer Using Artificial Neural Networks. Procedia Comput. Sci. 2016, 89, 465–472. [Google Scholar] [CrossRef]
- Jamshidi, M.; Roshani, S.; Daneshfar, F.; Lalbakhsh, A.; Roshani, S.; Parandin, F.; Malek, Z.; Talla, J.; Peroutka, Z.; Jamshidi, A.; et al. Hybrid Deep Learning Techniques for Predicting Complex Phenomena: A Review on COVID-19. AI 2022, 3, 416–433. [Google Scholar] [CrossRef]
- Kaur, M.; Kumar, V.; Yadav, V.; Singh, D.; Kumar, N.; Das, N.N. Metaheuristic-based Deep COVID-19 Screening Model from Chest X-Ray Images. J. Healthc. Eng. 2021, 2021, 8829829. [Google Scholar] [CrossRef] [PubMed]
- Padmakala, S.; Revathy, S.; Vijayalakshmi, K.; Mathankumar, M. CNN supported automated recognition of Covid-19 infection in chest X-ray images. Mater. Today Proc. 2022. [Google Scholar] [CrossRef] [PubMed]
- Kilicarslan, S.; Celik, M.; Sahin. Hybrid models based on genetic algorithm and deep learning algorithms for nutritional Anemia disease classification. Biomed. Signal Process Control 2021, 63, 102231. [Google Scholar] [CrossRef]
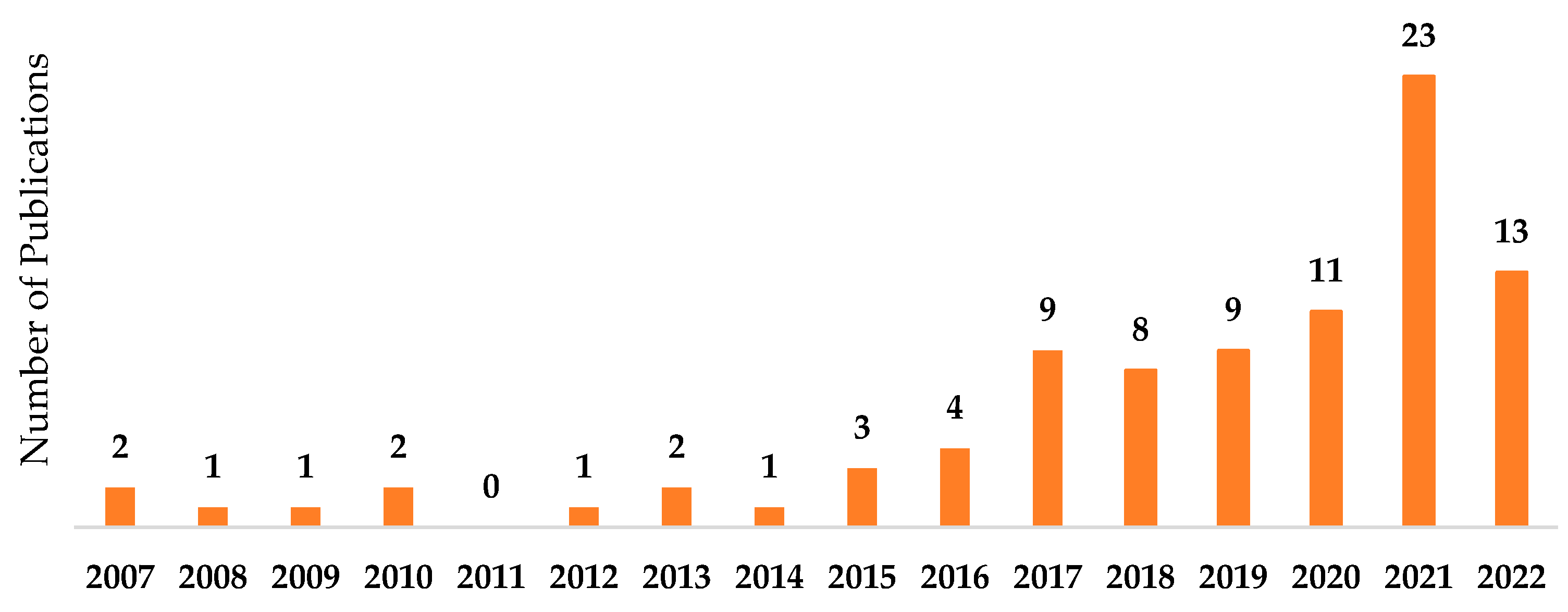




| The Most Common Risk Assessment Tools | Braden Scale | Waterlow Scale | Norton Scale | Cubbin-Jackson Scale | Braden Q Scale | SCIPUS | |
|---|---|---|---|---|---|---|---|
| Specialty | General | General/Orthopedics | General/Elderly Patients | ICU Patients | Pediatric Patients | Spinal Cord | |
| Strengths | High Sensitivity | ✕ | ✕ | ✕ | ✕ | ✕ | ✕ |
| Generalizability | ✕ | ✕ | |||||
| Drawbacks | Limited Number of Risk Factors | ✕ | ✕ | ✕ | ✕ | ✕ | ✕ |
| High False Positive Rate | ✕ | ✕ | ✕ | ✕ | ✕ | ✕ | |
| Risk Factors | Skin Status | ✕ | ✕ | ||||
| Mobility | ✕ | ✕ | ✕ | ✕ | ✕ | ✕ | |
| Friction Shear | ✕ | ✕ | |||||
| Blood Glucose Levels | ✕ | ||||||
| Hygiene | ✕ | ||||||
| Activity Status | ✕ | ✕ | ✕ | ||||
| Hemodynamics | ✕ | ||||||
| Perfusion | ✕ | ✕ | |||||
| Cardiac Disease | ✕ | ||||||
| Oxygenation | ✕ | ✕ | |||||
| Tobacco Use | ✕ | ||||||
| Gender | ✕ | ||||||
| Neurological Deficit | ✕ | ||||||
| Poor Nutrition Status | ✕ | ✕ | ✕ ✓ | ✕ | ✕ | ||
| Age | ✕ | ✕ | ✕ | ||||
| Sensory Perception | ✕ | ✕ | |||||
| Incontinence/Continence | ✕ | ✕ | ✕ | ✕ | |||
| Respiration | ✕ | ||||||
| Increase Skin Moisture | ✕ | ✕ | |||||
| Abnormal Lab Blood Results | ✕ # | ||||||
| Medications | ✕ | ||||||
| Renal Disease | ✕ | ||||||
| Mental Condition | ✕ | ✕ | |||||
| Respiratory Disease | ✕ | ||||||
| Weight for Height | ✕ | ||||||
| Physical Condition | ✕ | ||||||
| Past Medical Condition | ✕ | ||||||
| Major Surgery/Trauma | ✕ | ✕ | |||||
| Demographics | Medical | Diagnosis | Assessments | Labs | Medications | Medical Devices |
|---|---|---|---|---|---|---|
| Age | Admission Source | Comorbidity | Blood Pressure Systolic | Albumin | Opioids | Artificial Air Management |
| Ethnic Group | American Society of Anesthesiologists (ASA) Score | Depression | Blood Pressure Diastolic | Blood Urea Nitrogen (BUN) | Steroid Use | Face Mask |
| Race | Length-of-Stay at Emergency Department | Diabetes | Body Mass Index (BMI) | C-reactive Protein | Stimuli Anesthesia | Nasal Cannula |
| Sex | Visiting ICU during Hospitalization | Pressure Injury on Admission | Count of Glasgow Coma Score (GCS) Comment | Creatine Serum | Stimuli Paralytics | Noninvasive Ventilation |
| Number of Surgeries | Renal Failure | Glasgow Coma Score | Hemoglobin | Stimuli Sedation | Pharyngeal | |
| Number of Pressure Injuries | Sepsis Diagnosis | Weight Loss | High Mean Arterial Pressure (MAP) | Stimuli Tracheostomy | Room Air | |
| Palliative Orders | Stroke History | Patient Refusal to Change Position | Lactate | Vasopressor | Ventilator | |
| Prior Year Inpatient Visit Counter | Pulse Oximetry | Sodium | Feeding Tube | |||
| Steroid History | Skin Abnormality on Admission | |||||
| Visiting Transitional Unit during Hospitalization | Body Temperature |
| * Area | Reference | Author | Year | Country | Type | Dataset Size | Algorithm/ Method | Validation | Input of the Model | Output of the Model | ||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Prediction of SRPI | [25] | Cai et al. | 2020 | China | Journal | 149 patients | XGBoost | N/A | Risk factors for those who underwent cardiovascular surgery | Predict patients who will develop SRPI | ||||
| [26] | Chen et al. | 2018 | China | Journal | 149 patients | ANN | N/A | Risk factors for those who underwent cardiovascular surgery | Predict patients who will develop SRPI | |||||
| Exploration of the factors associated with PI | [27] | Aloweni et al. | 2019 | US | Journal | 269 patients | Multivariate LR | N/A | Potential risk factors for those who underwent operations | Identify risk factors associated with patients who will develop SRPI | ||||
| [28] | Moon and Lee | 2017 | South Korea | Journal | 15,856 patients | DT, univariate analysis | 10-fold CV | Records from the Health Insurance Review and Assessment (HIRA Service), and National Inpatient Sample (NIS) | Identify risk factors associated with elderly patients who will develop PI | |||||
| [29] | Lu et al. | 2017 | China | Journal | 149 patients | LR, univariate analysis, and multivariate analysis | N/A | Potential risk factors for those who underwent operations | Identify significant risk factors associated with patients who will develop SRPI | |||||
| [30] | Raju et al. | 2015 | US | Journal | 1653 patients | LR, DT, RF, Multivariate adaptive regression splines | 10-fold CV | Potential risk factors for PI patients that include Braden Scale, lab values, and demographics | Identify risk factors associated with patients who will develop PI | |||||
| Prediction of types of interventions based on the conditions of the patients | [31] | Jin et al. | 2021 | China | Conference | 1483 patients | PSO-RF, KNN, SVM DT | 10-fold CV | Different risk factors associated with PI patients/ patient status and physical characteristics | Predict the treatment/action needs for patients with PI based on risk factors (i.e., guidelines for actions) | ||||
| [32] | Mota et al. | 2019 | Portugal | Conference | 1339 patients | DT, NB | 66.00% Training vs. 34.00% testing | Different risk factors associated with PI patients/ patient status | Predict the treatment/action needs for patients with PI based on risk factors (i.e., guidelines for actions) | |||||
| Outcome of adopting Bayesian networks to detect PI on LOS | [33] | Cho et al. | 2013 | South Korea | Journal | 1214 Patients | Bayesian Networks | N/A | EHR of patients with PI | Evaluate the impact of the development of ML model to predict patients with PI (i.e., improvement before adopting a predictive model and after) | ||||
| Systematic literature PI management | [23] | Jiang et al. | 2021 | China | Journal | 32 Studies | A systematic review was conducted by analyzing 32 studies related to ML in PI management (PRISMA) | |||||||
| Predicting PI before occurrence | Systematic literature | [22] | Ribeiro et al. | 2021 | Portugal | Journal | 7 Studies | A systematic review was conducted by analyzing seven studies (PRISMA) to discover the most related algorithms for PI prevention | ||||||
| Predicting PI for nursing home residents | [62] | Charon et al. | 2022 | France | Conference | 3000 Patients | Bayesian networks, RF | N/A | Nursing home patients’ variables | Predict who will develop PI in nursing home | ||||
| [63] | Lee et al. | 2021 | South Korea | Journal | 60 Patients | RF, LR, SVM | N/A | Nursing home patients’ variables | Predict factors for PI related to nursing home residents and predict PI | |||||
| Predicting HAPI/CAPI | ML | [2,34,35,36,37,38,39,40,41,42,43,44,45,46,47,48,49,50,51,52,53,54,55,56,57,58,59,60,61] | Table 4 and Table 5 summarize the 30 studies that had the same inputs (risk factors) and same outputs (i.e., predicting PI before occurrence (HAPI/CAPI)) | |||||||||||
| DL | [64] | |||||||||||||
| Study | Author(s) | Year | Country | Type | Dataset Size | Validation Method | Feature Importance | Feature Selection | Cost-Sensitive Learning | Hyperparameter Tuning (Grid Search) | Balancing Method | HAPI % | Retrospective Study | Comparing Results to Risk Assessment Tools |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| [53] | Šín et al. | 2022 | Czech Republic | Journal | 9304 | 80% Training vs. 20% testing | ✓ | RO | 50.00 | |||||
| [54] | J. Xu et al. | 2022 | China | Journal | 618 | 70% Training (5-fold CV) vs. 30% testing | RO | 33.33 | ✓ | ✓ | ||||
| [55] | Do et al. | 2022 | US | Journal | 6742 | 70% Training (5-fold CV) vs. 30% testing | ✓ | RO | 31.92 | ✓ | ✓ | |||
| [35] | Walther et al. | 2021 | Germany | Journal | 149,006 | 10-fold CV | ✓ | ✓ | RO | 3.10 | ✓ | |||
| [34] | Nakagami et al. | 2021 | Japan | Journal | 75,353 | 70% Training (5-fold CV) vs. 30% testing | ✓ | ✓ | ✓ | RO | 0.52 | ✓ | ||
| [2] | W. Song et al. | 2021 | US | Journal | 188,512 | 80% Training (5-fold CV) vs. 20% testing | ✓ | ✓ | RO | 3.27 | ✓ | |||
| [36] | Alderden et al. | 2021 | US | Journal | 5101 | 80% Training vs. 20% testing | SMOTE | 6.50 | ✓ | ✓ | ||||
| [37] | Anderson et al. | 2021 | US | Journal | 23,000 | 80% Training vs. 20% testing | SMOTE | N/A | ✓ | ✓ | ||||
| [47] | Ahmad et al. | 2021 | US | Conference | 713 | 10-fold CV | ✓ | RO | 52.31 | |||||
| [38] | Ossai et al. | 2021 | Australia | Conference | 1014 | 10-fold CV | ✓ | SMOTE | N/A | ✓ | ||||
| [48] | J. Song et al. | 2021 | China | Journal | 5814 | 50% Training (10-fold CV) vs. 50% testing | ✓ | US and RO | 28.78 | ✓ | ||||
| [64] | Y. Wang et al. | 2021 | China | Journal | 246 | 67% Training vs. 33% testing | ✓ | Image rotation and image dilation | 50.00 | ✓ | ||||
| [61] | Cheng et al. | 2021 | China | Journal | 245 | 5-fold CV | ✓ | US | 80.00 | ✓ | ||||
| [39] | Hu et al. | 2020 | China | Journal | 11,838 | 50% Training (10-fold CV) vs. 50% testing | ✓ | ✓ | US and RO | 1.36 | ✓ | |||
| [40] | Vyas et al. | 2020 | US | Conference | 13,282 | 80% Training vs. 20% testing | RO | 16.80 | ✓ | |||||
| [44] | Ladios-Martin et al. | 2020 | Spain | Journal | 6694 | 64% Training vs. 36% testing | SMOTE | 4.12 | ✓ | ✓ | ||||
| [59] | Levy et al. | 2020 | US | Journal | 57,227 | 80% Training (5-fold CV) vs. 20% testing | ✓ | ✓ | SMOTE, RO | 0.42 | ✓ | |||
| [45] | Cramer et al. | 2019 | US | Journal | 50,851 | 80% Training vs. 20% testing | ✓ | ✓ | ✓ | SMOTE, RO | 7.80 | ✓ | ||
| [57] | Hyun et al. | 2019 | US | Journal | 12,654 | N/A | RO | 5.81 | ✓ | ✓ | ||||
| [51] | H. L. Li et al. | 2019 | China | Journal | 2062 | K-fold CV (k is not specified) | RO | 50.00 | ✓ | |||||
| [52] | Cichosz et al. | 2019 | Denmark | Journal | 383 | 65% Training (5-fold CV) vs. 35% testing | ✓ | RO | 28.10 | ✓ | ✓ | |||
| [42] | Alderden et al. | 2018 | US | Journal | 6376 | 67% Training vs. 33% testing | RO | 8.10 | ✓ | |||||
| [46] | Gao et al. | 2018 | China | Journal | 1963 | N/A | RO | 2.38 | ✓ | |||||
| [56] | Kaewprag et al. | 2017 | US | Journal | 7717 | 67% Training vs. 33% testing | ✓ | ✓ | RO | 7.65 | ✓ | ✓ | ||
| [41] | Y. Jin et al. | 2017 | South Korea | Journal | 11,191 | 80% Training vs. 20% testing | ✓ | RO | 20.00 | ✓ | ✓ | |||
| [43] | Deng et al. | 2017 | China | Journal | 468 | 10-fold CV | ✓ | RO | 20.10 | ✓ | ✓ | |||
| [60] | Setoguchi et al. | 2016 | Japan | Journal | 8286 | 10-fold CV | ✓ | ✓ | N/A | 0.62 | ✓ | |||
| [50] | Su et al. | 2012 | China | Journal | 168 | 4-fold CV | ✓ | RO | 4.80 | ✓ | ||||
| [49] | Y.-C. Chen et al. | 2008 | China | Conference | 168 | 3-fold CV | RO | 4.80 | ✓ | |||||
| [58] | Borlawsky and Hripcsak | 2007 | US | Journal | 3151 | 4-fold CV | ✓ | RO | 8.10 | ✓ |
| Study | Performance Metrics | Algorithm Adopted | ||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Accuracy | Sensitivity | Specificity | Precision | FI-score | AUC | False Positive | Other(s) | LR | RF | DT | SVM | MLP | KNN | LDA | Other(s) | |
| [53] | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | |||
| [54] | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ||||||||
| [55] | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ||||||||
| [35] | ✓ | ✓ | ✓ | |||||||||||||
| [34] | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | |||||||
| [2] | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | |||||||
| [36] | ✓ | ✓ | ✓ | ✓ | ||||||||||||
| [37] | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ||||||||
| [47] | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | |||||||||
| [38] | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ||||||
| [48] | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | |||||||
| [64] | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | |||||||||
| [61] | ✓ | ✓ | ✓ | ✓ | ✓ | |||||||||||
| [39] | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | |||||||
| [40] | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ||||||||
| [44] | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | |||||||
| [59] | ✓ | ✓ | ✓ | ✓ | ✓ | |||||||||||
| [45] | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | |||||||||
| [57] | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ||||||||||
| [51] | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ||||||||||
| [52] | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ||||||||||
| [42] | ✓ | ✓ | ||||||||||||||
| [46] | ✓ | ✓ | ✓ | |||||||||||||
| [56] | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ||||||||||
| [41] | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | |||||||||
| [43] | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ||||||||||
| [60] | ✓ | ✓ | ✓ | ✓ | ||||||||||||
| [50] | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ||||||||
| [49] | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | |||||||||
| [58] | ✓ | ✓ | ✓ | ✓ | ✓ | |||||||||||
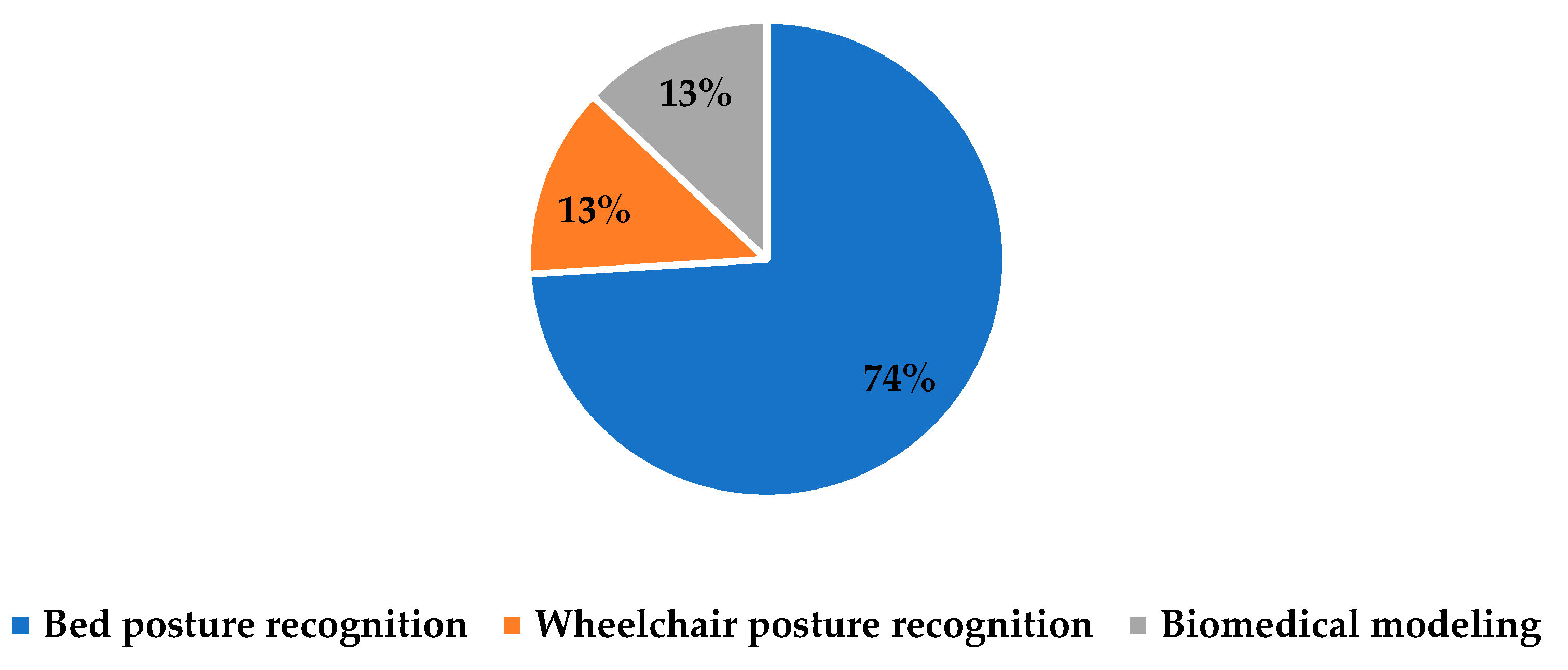
| Area | Reference | Author(s) | Year | Country | Type | Dataset Size | Algorithms | Input of the Model | Output of the Model | |
|---|---|---|---|---|---|---|---|---|---|---|
| Posture bed recognition | DL | [67] | Chiang et al. | 2022 | China | Journal | Seven different sets of samples that have been trained and tested; each set has its own number of images (Table 3 in [67]) | CNN | 3D skeleton of PI patients that has articulated joints | Skeleton-based posture classification of elderly patients with PI |
| DL | [68] | Cicceri et al. | 2020 | Italy | Journal | N/A | DNN, SVM, RF | Data was collected through internal sensors that estimate the position of PI patients | Classify the position of PI patients based on sensors and send a notification to change the patient’s body position when a patient remains in the same position for a while | |
| DL | [69] | Heydarzadeh et al. | 2016 | US | Conference | 60,000 PI images | Deep autoencoder neural network- Histogram of Gradient (HoG), PCA-SVM, Bayesian inference, Kurtosis-Skewness, Gaussian Mixture Model (GMM) | A commercial pressure mapping system was used to gather the data | Classify in-bed into posture: right foetus, right yearner, supine, left yearner, and left foetus | |
| ML | [70] | Matar et al. | 2020 | Canada | Journal | 1728 sensors | MLP | Bed-sheet pressure sensors were used to collect the signals of body pressure/ pressure image | Autonomous approach to classify bed posture: spine, right, left, and prone | |
| ML | [71] | Duvall et al. | 2019 | US | Journal | 4 sensors | KNN | Data were collected through e-scale positioned under the bed to measure the weights of each leg on the bed | Classify types of movement in bed: turn in place, roll, extremity movements, and assisted turn | |
| ML | [72] | Enayati et al. | 2018 | US | Conference | 4 sensors | PCA, NN | Pressure sensors were used to collect signals of body pressure from 58 patients | Classify the most common four sleeping postures of the patients: left lateral, right lateral, supine, and prone | |
| ML | [73] | X. Xu et al. | 2016 | China | Journal | a 3 × 3 pressure sensor array | Developed skew-based sleep posture classifier based on KNN | Pressure sensors were used to collect signals of body pressure/pressure image | Predict sleep posture recognition based on Body-Earth Mover’s Distance (BEMD) | |
| ML | [74] | Baran Pouyan et al. | 2016 | US | Journal | 1728 sensors | KNN | Commercial pressure map model, which has sensors, were used to capture pressure data continuously/ pressure image | Clustering model to extract body limbs from pressure data gathered by a commercial pressure map device | |
| ML | [75] | Hsiao et al. | 2015 | China | Journal | 5 Force Sensing Resistor (FSR) | Fuzzy theory, KNN, SVM | A pressure sensing pad was developed and used to collect signals of body pressure/pressure image | Classify the position of the PI patients in nursing homes, then send a notification when a patient remains in the same position for a while to change the patient’s body position | |
| ML | [76] | Pouyan et al. | 2014 | US | Conference | 2048 pressure sensors, the model was tested on 15 different patients | KNN, NB, DT | Pressure image of bed inclination | Classify the most common three-bed inclination: B0 degree, B30 degree, and B60 degree | |
| ML | [77] | Barsocchi | 2013 | Italy | Conference | 3 sensors | SVM, KNN | Receive signals of the body through signal strength, and transmit signals from a wireless appliance to a server | Classify the position of the PI elderly patients based on sensors, and observe the activities of patients who cannot move their bodies the way they should | |
| Posture wheel-chair recognition | ML | [78] | Jaffery et al. | 2022 | Saudi Arabia | Journal | Matrix configuration (9 sensors), and cross configuration (5 sensors) | KNN, LR, DT, SVM, LightGBM | Capture a real-time posture/signals of a patient on a wheelchair seat using sensors (two configuration systems) | Recognize the sitting posture of wheelchair users to prevent PI. The five positions are ideal, left-leaning, forward-leaning, right-leaning, and backward-leaning |
| ML | [79] | Ma et al. | 2017 | China | Journal | 12 sensors | DT, SVM, MLP, NB, KNN | Collect the posture of patients through sensor configuration | Capture cushion-of posture of wheelchair | |
| Biomedical modeling (i.e., ML model to learn the pattern between pressure map modes and strain field modes) | ML | [80] | Grunerbel et al. | 2022 | Germany | Conference | A pressure sensor and a vital parameter sensor node: collect data for 17 nights | A multivariate subsequence clustering algorithm | Collect signals of skin temperature, SpO2, and heart rate | Measure skin temperature and blood oxygen saturation around potential wound sites in addition to pressure loads. Then send a medical alarm based on the status of the patients |
| ML | [81] | Luboz et al. | 2018 | France | Journal | 19 pressure modes | N/A | Collect signals of skin, fats, and muscles in real-time | Design a 3D buttock model to provide PI prevention (skin detection) | |
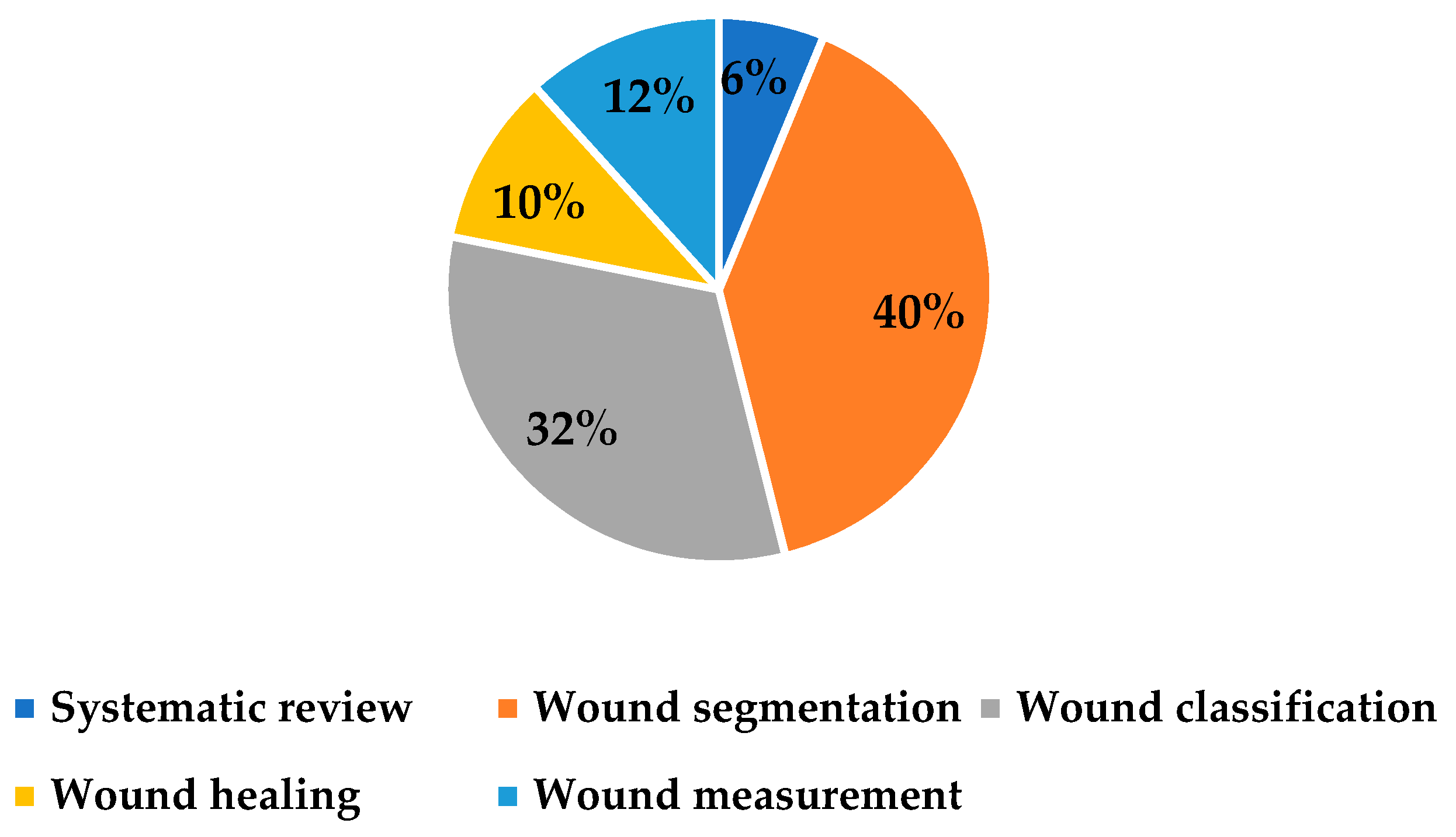
| Area | Reference | Author(s) | Year | Country | Type | Dataset Size | Algorithms | Input of the Model | Output of the Model | |
|---|---|---|---|---|---|---|---|---|---|---|
| Systematic Review | Approach | [112] | Kaswan et al. | 2020 | Malaysia | Journal | A brief review of wound classification and wound segmentation, most of the literature was imported from the dataset of national pressure ulcer advisory panel (10 studies) | |||
| [4] | Zahia et al. | 2019 | Spain | Journal | A systematic review was conducted by analyzing 114 studies related to wound analysis in general, using image processing | |||||
| Wound Segmentation | DL | [83] | Ramachandram et al. | 2022 | Canada | Journal | 58 images | CNN | Wound images of PI, arterial ulcers, and venous ulcers | Wound tissue segmentation |
| DL | [84] | C. W. Chang et al. | 2022 | China | Journal | 2893 images | U-Net, DeeplabV3, PsPNet, FPN, and Mask R-CNN | Wound images of PI | Wound tissue segmentation | |
| ML | [85] | Howell et al. | 2021 | US | Journal | 199 images | Droice Labs wound analytics service | Wound images of PI | Wound area and granulation tissue tracing/wound boundary detection | |
| DL | [86] | C. Wang et al. | 2020 | US | Journal | 1109 images | CNN based on MobileNetV2 | PI foot ulcer images | Segment wound regions/boundary detection | |
| DL | [87] | Ohura et al. | 2019 | Japan | Journal | 440 images | CNN and TL | Wound detection/ boundary detection | Wound detection/ boundary detection | |
| DL | [3] | Zahia et al. | 2018 | Spain | Journal | 22 high-resolution images per class, and then cut into 5*5 sub- images, ending with 380,000 small images | CNN | PI wound images | Tissue segmentation: granulation, slough, and necrotic tissues | |
| DL | [88] | García-Zapirain et al. | 2018 | Spain | Journal | 193 images | 3D CNN | PI wound images | Tissue segmentation: granulation, slough, and necrotic tissues | |
| ML | [89] | Garcia-Zapirain et al. | 2017 | Spain | Journal | 48 images | Developed their own framework | PI wound images | Design a segmentation software for image segmentation and wound detection | |
| ML | [90] | F. J. Veredas et al. | 2015 | Spain | Journal | 113 images | k-means, NN, SVM, RF | Wound images of PI patients with home-care assistance | Tissue identification, then classify wound tissue types: necrosis, slough, granulation, and healing skin | |
| ML | [92] | F. Veredas et al. | 2010 | Spain | Journal | 113 images | PCA, MLP, NB | PI wound images | Tissue identification in wound image/ recognition for different types of tissue: necrosis, slough, granulation, healing, skin, and global | |
| ML | [91] | Wannous et al. | 2007 | France | Conference | 905 images/regions (granulation: 302, slough: 243, necrosis: 73, and healthy: 287) | SVM | PI wound images | Segment wound regions/boundary detection (granulation, slough, and necrosis) | |
| Wound Classification | DL | [93] | Ay et al. | 2022 | Turkey | Journal | 1091 images | Deep TL, CNN | PI wound images | Classification of four stages of PI: stages 1–4 |
| ML | [94] | Fergus et al. | 2022 | UK | Conference | 216 images | NN | PI wound images | Classification of six stages of PI: stage 1–4, unstageable PI, and DTPI | |
| DL | [95] | Liu et al. | 2022 | China | Journal | 327 images | ResNet-v2 model (CNN) | PI wound images | Wound classification in two phases: phase 1: erythema or non- erythema; phase 2: “extensive necrosis or moderate necrosis | |
| DL | [96] | Anisuzzaman et al. | 2021 | US | Conference | 2176 images collected from three datasets (730, 358, 1088) | TL and MMML | Wound images and corresponding locations | Wound classification: diabetic, pressure, venous ulcers, and surgical | |
| DL | [97] | Matsumoto et al. | 2021 | Japan | Journal | 860 images | CNN | Ultrasound images of PI | Classification of types of DTPI: unclear layer structure, cobblestone-like pattern, cloud-like pattern, and anechoic pattern | |
| DL | [98] | A. Yilmaz et al. | 2021 | Turkey | Conference | 175 images | CNN | PI wound images | Classification of six stages of PI: stage 1–4, unstageable PI, and DTPI | |
| ML | [99] | B. Yilmaz et al. | 2021 | Turkey | Conference | 142 images | LR, NN | PI wound images | Classification of six stages of PI: stage 1–4, unstageable PI, and DTPI | |
| ML | [100] | Mombini et al. | 2021 | US | Conference | 2056 images | XGboost, DT, RF, SVM | PI chronic images | Classifying the status of the wound into maintaining current treatment, referring the patient to a specialist, changing the current treatment | |
| DL | [101] | D. H. Chang et al. | 2021 | China | Conference | 210 images | U-net CNN | PI wound images | Tissue classification and severity evaluation of wound condition and severity: granulation > 90%, granulation 70–90, granulation < 30, necrosis < 50, necrosis > 50 | |
| ML | [102] | Kavitha et al. | 2017 | US | Conference | 59 images | MLP, SVM, RF, NB | wound images (leg ulcers, venous and arterial, and pressure) | Classify images into pressure ulcer vs. leg ulcers | |
| Wound Healing | ML | [103] | Lustig et al. | 2022 | Israel | Journal | 173 images | Developed their own algorithm | Subepidermal moisture delta | Predict heel deep tissue injuries for ICU patients (heal or not within seven days) |
| ML | [104] | Chun et al. | 2021 | China | Journal | 152 images | RF, XGBoost | EHR variables | Classifying pediatric patients into healing or delayed healing | |
| ML | [105] | F. J. Veredas et al. | 2010 | Spain | Conference | 743 images | SVM, NB, NN, DT | PI wound images (sacrum and hip) | Predict the wound status: improvement or improvement delayed | |
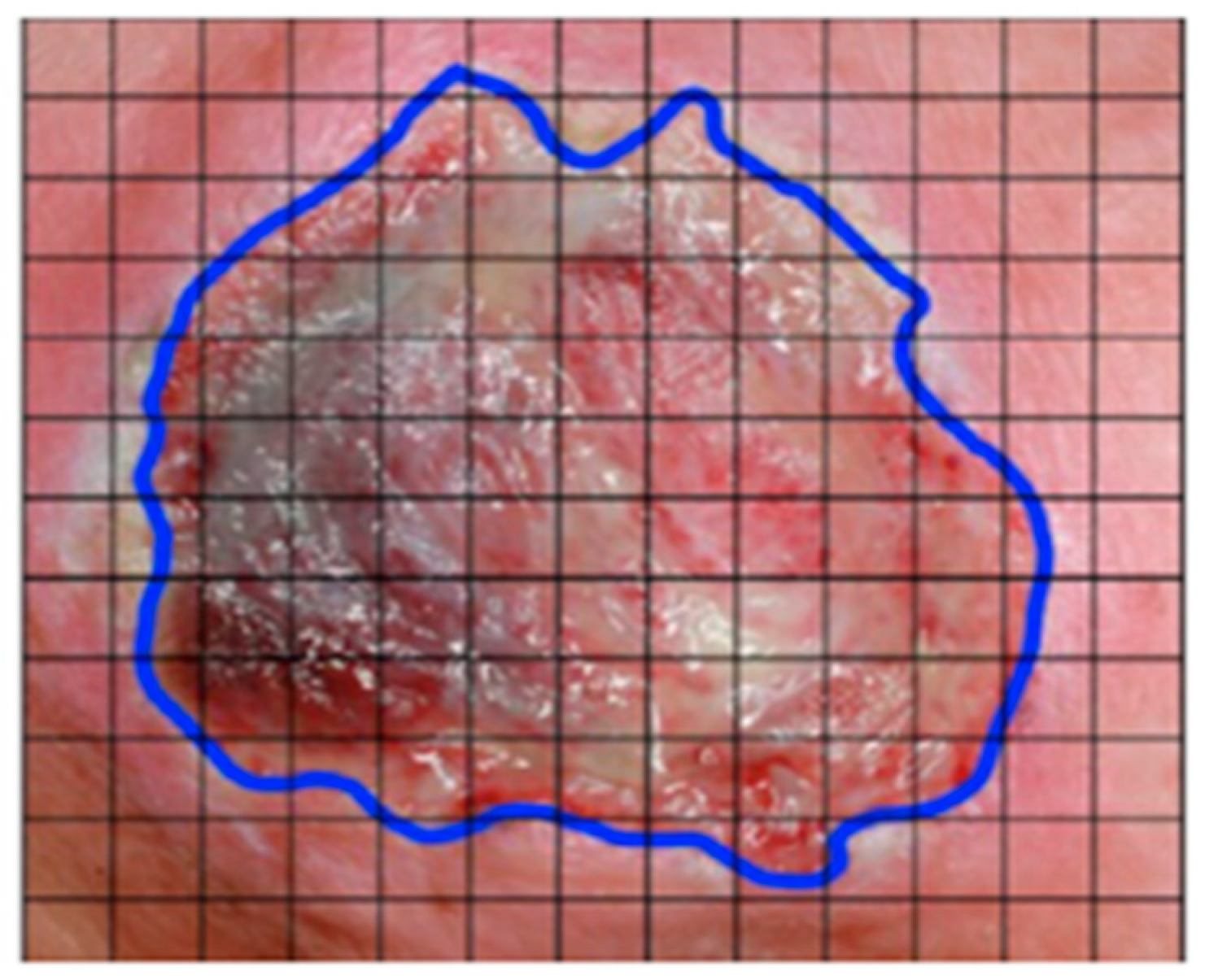
| Wound Measurement | ML | [106] | Silva and Machado | 2021 | Brazil | Journal | 105 images | SVM-Grabcut | PI wound images | Measurement of the “area” affected by PI |
| ML | [107] | D. Li and Mathews | 2017 | US | Journal | 32 images | Developed their own model using SVM and Gaussian model | 3D wound images | Measure size of PI | |
| Wound Segmentation and Wound Measurement | DL | [108] | Zahia et al. | 2020 | Spain | Journal | 210 images | Mask RCNN | 3D mesh and 2D wound images | Measurement of depth, area, volume, major axis, and minor axis (external segmentation of the wound) |
| DL | [109] | Chino et al. | 2020 | Brazil | Journal | 446 images | CNN | Wound images (two datasets) | Wound tissue segmentation, and measurement size of PI | |
| ML | [110] | F. J. Veredas et al. | 2009 | Spain | Conference | 50 images | A hybrid model: NN and Bayesian classifiers | Wound images of PI | Wound segmentation into wound regions (separate wounds from healing areas) then measure area of wounds | |
| Wound Segmentation, Classification, Measurement, and Healing | ML | [111] | M. C. Chang et al. | 2018 | US | Journal | 133 scanning sessions from 23 enrolled subjects | Developed their own algorithm | Multimodal PI images: 3D depth, RGB, chemical sensing, thermal, and multispectral | Tissue classification (granulation vs. slough), 3D depth: 3D wound size measurement (length, width, depth, surface, and volume), thermal profiling, and chemical sensing: heal trend analysis |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Dweekat, O.Y.; Lam, S.S.; McGrath, L. Machine Learning Techniques, Applications, and Potential Future Opportunities in Pressure Injuries (Bedsores) Management: A Systematic Review. Int. J. Environ. Res. Public Health 2023, 20, 796. https://doi.org/10.3390/ijerph20010796
Dweekat OY, Lam SS, McGrath L. Machine Learning Techniques, Applications, and Potential Future Opportunities in Pressure Injuries (Bedsores) Management: A Systematic Review. International Journal of Environmental Research and Public Health. 2023; 20(1):796. https://doi.org/10.3390/ijerph20010796
Chicago/Turabian StyleDweekat, Odai Y., Sarah S. Lam, and Lindsay McGrath. 2023. "Machine Learning Techniques, Applications, and Potential Future Opportunities in Pressure Injuries (Bedsores) Management: A Systematic Review" International Journal of Environmental Research and Public Health 20, no. 1: 796. https://doi.org/10.3390/ijerph20010796
APA StyleDweekat, O. Y., Lam, S. S., & McGrath, L. (2023). Machine Learning Techniques, Applications, and Potential Future Opportunities in Pressure Injuries (Bedsores) Management: A Systematic Review. International Journal of Environmental Research and Public Health, 20(1), 796. https://doi.org/10.3390/ijerph20010796





