An Overview of Antibiotic Resistance and Abiotic Stresses Affecting Antimicrobial Resistance in Agricultural Soils
Abstract
:1. Introduction
2. Abiotic Stresses
2.1. Soil Pollutants
2.1.1. Fertilizers
2.1.2. Heavy Metals
2.1.3. Hydrocarbons
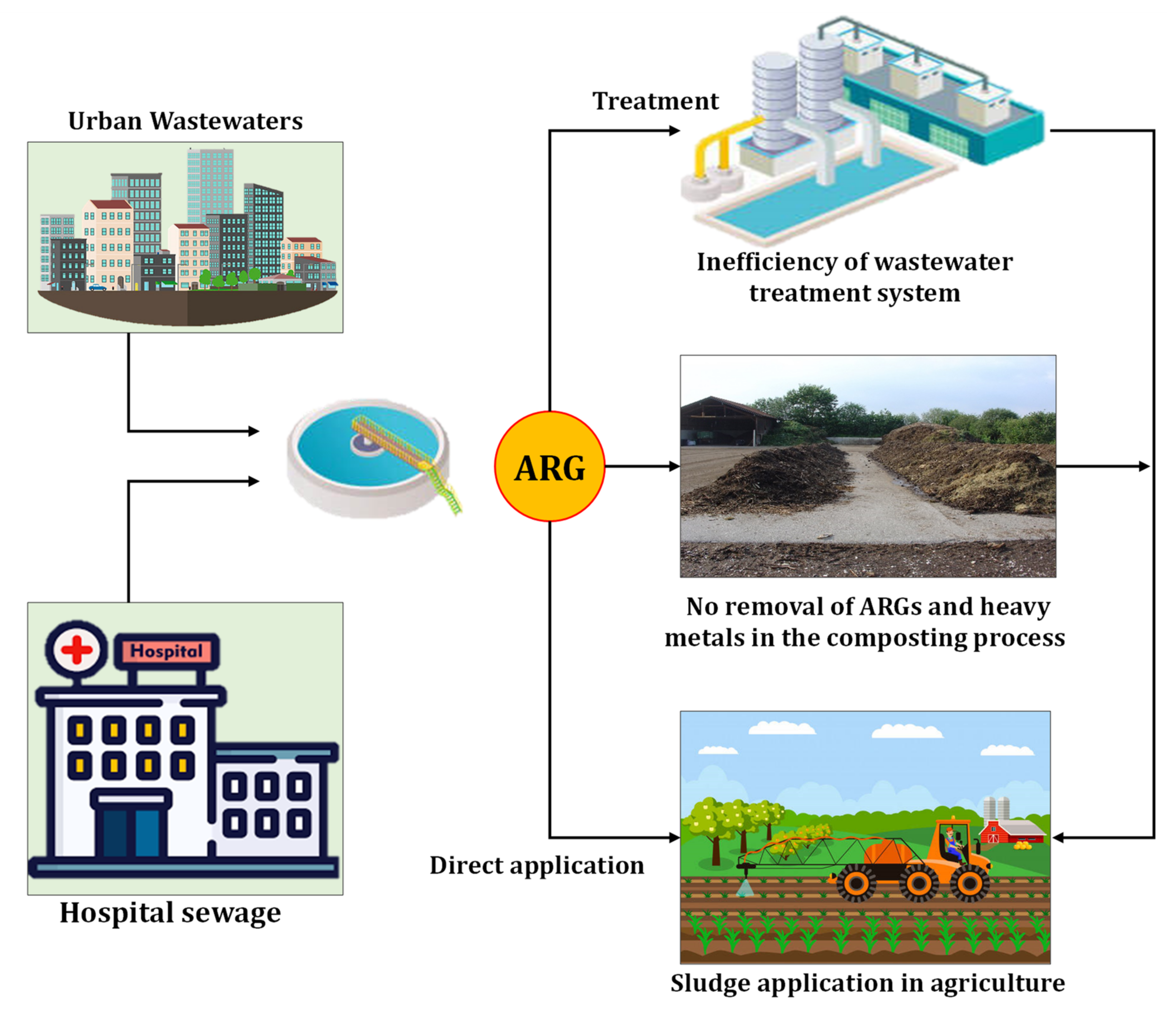
2.1.4. Sewage Sludge
2.2. Salinity
3. Combating Antibiotic Resistance Prevalence
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Zhu, Y.G.; Zhao, Y.; Zhu, D.; Gillings, M.; Penuelas, J.; Ok, Y.S.; Capon, A.; Banwart, S. Soil biota, antimicrobial resistance and planetary health. Environ. Int. 2019, 131, 105059. [Google Scholar] [CrossRef] [PubMed]
- Armalyte, J.; Skerniskyte, J.; Bakiene, E.; Krasauskas, R.; Siugzdiniene, R.; Kareiviene, V.; Kerziene, S.; Klimiene, I.; Suziedeliene, E.; Ruzauskas, M. Microbial Diversity and Antimicrobial Resistance Profile in Microbiota from Soils of Conventional and Organic Farming Systems. Front. Microbiol. 2019, 10, 892. [Google Scholar] [CrossRef] [PubMed]
- Bahram, M.; Hildebrand, F.; Forslund, S.K.; Anderson, J.L.; Soudzilovskaia, N.A.; Bodegom, P.M.; Bengtsson-Palme, J.; Anslan, S.; Coelho, L.P.; Harend, H.; et al. Structure and function of the global topsoil microbiome. Nature 2018, 560, 233–237. [Google Scholar] [CrossRef] [PubMed]
- Nelkner, J.; Henke, C.; Lin, T.W.; Patzold, W.; Hassa, J.; Jaenicke, S.; Grosch, R.; Puhler, A.; Sczyrba, A.; Schluter, A. Effect of Long-Term Farming Practices on Agricultural Soil Microbiome Members Represented by Metagenomically Assembled Genomes (MAGs) and Their Predicted Plant-Beneficial Genes. Genes 2019, 10, 424. [Google Scholar] [CrossRef] [Green Version]
- Nguyen, B.T.; Chen, Q.L.; He, J.Z.; Hu, H.W. Microbial regulation of natural antibiotic resistance: Understanding the protist-bacteria interactions for evolution of soil resistome. Sci. Total Environ. 2020, 705, 135882. [Google Scholar] [CrossRef] [PubMed]
- Foo, J.L.; Ling, H.; Lee, Y.S.; Chang, M.W. Microbiome engineering: Current applications and its future. Biotechnol. J. 2017, 12, 1600099. [Google Scholar] [CrossRef] [Green Version]
- Ochoa-Hueso, R. Global Change and the Soil Microbiome: A Human-Health Perspective. Front. Ecol. Evol. 2017, 5, 71. [Google Scholar] [CrossRef] [Green Version]
- Kemper, N. Veterinary antibiotics in the aquatic and terrestrial environment. Ecol. Indic. 2008, 8, 1–13. [Google Scholar] [CrossRef]
- Bilal, M.; Mehmood, S.; Rasheed, T.; Iqbal, H.M.N. Antibiotics traces in the aquatic environment: Persistence and adverse environmental impact. Curr. Opin. Environ. Sci. Health 2020, 13, 68–74. [Google Scholar] [CrossRef]
- Larsson, D.G. Antibiotics in the environment. Ups. J. Med. Sci. 2014, 119, 108–112. [Google Scholar] [CrossRef]
- Durand, G.A.; Raoult, D.; Dubourg, G. Antibiotic discovery: History, methods and perspectives. Int. J. Antimicrob. Agents 2019, 53, 371–382. [Google Scholar] [CrossRef] [PubMed]
- Baguer, A.J.; Jensen, J.; Krogh, P.H. Effects of the antibiotics oxytetracycline and tylosin on soil fauna. Chemosphere 2000, 40, 751–757. [Google Scholar] [CrossRef]
- Isaacson, R.E.; Torrence, M.E. The Role of Antibiotics in Agriculture: This Report Is Based on a Colloquium Sponsored by the American Academy of Microbiology Held November 2–4, 2001, in Santa Fe, New Mexico; American Society for Microbiology: Washington, DC, USA, 2002. [Google Scholar] [CrossRef]
- Callaway, T.R.; Lillehoj, H.; Chuanchuen, R.; Gay, C.G. Alternatives to Antibiotics: A Symposium on the Challenges and Solutions for Animal Health and Production. Antibiotics 2021, 10, 471. [Google Scholar] [CrossRef] [PubMed]
- Akram, R.; Amin, A.; Hashmi, M.Z.; Wahid, A.; Mubeen, M.; Hammad, H.M.; Fahad, S.; Nasim, W. Fate of Antibiotics in Soil. In Antibiotics and Antibiotics Resistance Genes in Soils; Springer: Cham, Switzerland, 2017; pp. 207–220. [Google Scholar] [CrossRef]
- Sarmah, A.K.; Meyer, M.T.; Boxall, A.B. A global perspective on the use, sales, exposure pathways, occurrence, fate and effects of veterinary antibiotics (VAs) in the environment. Chemosphere 2006, 65, 725–759. [Google Scholar] [CrossRef]
- Kumar, K.; Gupta, S.; Chander, Y.; Singh, A.K. Antibiotic Use in Agriculture and Its Impact on the Terrestrial Environment. Adv. Agron. 2005, 87, 1–54. [Google Scholar] [CrossRef]
- Shen, Y.; Stedtfeld, R.D.; Guo, X.; Bhalsod, G.D.; Jeon, S.; Tiedje, J.M.; Li, H.; Zhang, W. Pharmaceutical exposure changed antibiotic resistance genes and bacterial communities in soil-surface- and overhead-irrigated greenhouse lettuce. Environ. Int. 2019, 131, 105031. [Google Scholar] [CrossRef]
- Zalewska, M.; Blazejewska, A.; Czapko, A.; Popowska, M. Antibiotics and Antibiotic Resistance Genes in Animal Manure—Consequences of Its Application in Agriculture. Front. Microbiol. 2021, 12, 610656. [Google Scholar] [CrossRef]
- CDC. About Antibiotic Resistance, 2021, Last Updated: 13 December 2021. Available online: https://www.cdc.gov/drugresistance/about.html (accessed on 2 January 2022).
- WHO. Antibiotic Resistance. 2020. Available online: https://www.who.int/news-room/fact-sheets/detail/antibiotic-resistance (accessed on 31 July 2020).
- World Bank. Antimicrobial Resistance (AMR). 2021. Available online: https://www.worldbank.org/en/topic/health/brief/antimicrobial-resistance-amr (accessed on 13 May 2021).
- CDC. 2019 AR Threats Report. 2021. Available online: https://www.cdc.gov/drugresistance/biggest-threats.html (accessed on 23 November 2021).
- WHO. Antimicrobial Resistance. 2021. Available online: https://www.who.int/news-room/fact-sheets/detail/antimicrobial-resistance (accessed on 17 November 2021).
- WHO. Infographics: Antibiotic Resistance. 2015. Available online: https://apps.who.int/mediacentre/events/2015/world-antibiotic-awareness-week/infographics/en/index.html (accessed on 22 November 2015).
- He, Y.; Yuan, Q.; Mathieu, J.; Stadler, L.; Senehi, N.; Sun, R.; Alvarez, P.J.J. Antibiotic resistance genes from livestock waste: Occurrence, dissemination, and treatment. NPJ Clean Water 2020, 3, 1–11. [Google Scholar] [CrossRef] [Green Version]
- Laconi, A.; Mughini-Gras, L.; Tolosi, R.; Grilli, G.; Trocino, A.; Carraro, L.; Di Cesare, F.; Cagnardi, P.; Piccirillo, A. Microbial community composition and antimicrobial resistance in agricultural soils fertilized with livestock manure from conventional farming in Northern Italy. Sci. Total Environ. 2021, 760, 143404. [Google Scholar] [CrossRef]
- Yukgehnaish, K.; Kumar, P.; Sivachandran, P.; Marimuthu, K.; Arshad, A.; Paray, B.A.; Arockiaraj, J. Gut microbiota metagenomics in aquaculture: Factors influencing gut microbiome and its physiological role in fish. Rev. Aquac. 2020, 12, 1903–1927. [Google Scholar] [CrossRef]
- Xie, W.Y.; Shen, Q.; Zhao, F.J. Antibiotics and antibiotic resistance from animal manures to soil: A review. Eur. J. Soil Sci. 2017, 69, 181–195. [Google Scholar] [CrossRef] [Green Version]
- Ter Kuile, B.H.; Kraupner, N.; Brul, S. The risk of low concentrations of antibiotics in agriculture for resistance in human health care. FEMS Microbiol. Lett. 2016, 363, fnw210. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- ECDC. Country Overview of Antimicrobial Consumption. 2020. Available online: https://www.ecdc.europa.eu/en/antimicrobial-consumption/database/country-overview (accessed on 20 November 2021).
- Chang, Q.; Wang, W.; Regev-Yochay, G.; Lipsitch, M.; Hanage, W.P. Antibiotics in agriculture and the risk to human health: How worried should we be? Evol. Appl. 2015, 8, 240–247. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hassan, M.M.; El Zowalaty, M.E.; Lundkvist, A.; Jarhult, J.D.; Khan Nayem, M.R.; Tanzin, A.Z.; Badsha, M.R.; Khan, S.A.; Ashour, H.M. Residual antimicrobial agents in food originating from animals. Trends Food Sci. Technol. 2021, 111, 141–150. [Google Scholar] [CrossRef] [PubMed]
- Treiber, F.M.; Beranek-Knauer, H. Antimicrobial Residues in Food from Animal Origin-A Review of the Literature Focusing on Products Collected in Stores and Markets Worldwide. Antibiotics 2021, 10, 534. [Google Scholar] [CrossRef] [PubMed]
- Martinez, J.L. Effect of antibiotics on bacterial populations: A multi-hierachical selection process. F1000Research 2017, 6, 51. [Google Scholar] [CrossRef] [Green Version]
- Quaik, S.; Embrandiri, A.; Ravindran, B.; Hossain, K.; Al-Dhabi, N.A.; Arasu, M.V.; Ignacimuthu, S.; Ismail, N. Veterinary antibiotics in animal manure and manure laden soil: Scenario and challenges in Asian countries. J. King Saud Univ. Sci. 2020, 32, 1300–1305. [Google Scholar] [CrossRef]
- Grenni, P.; Ancona, V.; Barra Caracciolo, A. Ecological effects of antibiotics on natural ecosystems: A review. Microchem. J. 2018, 136, 25–39. [Google Scholar] [CrossRef]
- Lucas, J.M.; Sone, B.M.; Whitmore, D.; Strickland, M.S. Antibiotics and temperature interact to disrupt soil communities and nutrient cycling. Soil Biol. Biochem. 2021, 163, 108437. [Google Scholar] [CrossRef]
- Toth, J.D.; Feng, Y.; Dou, Z. Veterinary antibiotics at environmentally relevant concentrations inhibit soil iron reduction and nitrification. Soil Biol. Biochem. 2011, 43, 2470–2472. [Google Scholar] [CrossRef]
- Kong, W.D.; Zhu, Y.G.; Fu, B.J.; Marschner, P.; He, J.Z. The veterinary antibiotic oxytetracycline and Cu influence functional diversity of the soil microbial community. Environ. Pollut. 2006, 143, 129–137. [Google Scholar] [CrossRef] [PubMed]
- Uddin, M.; Chen, J.; Qiao, X.; Tian, R.; Arafat, Y.; Yang, X. Bacterial community variations in paddy soils induced by application of veterinary antibiotics in plant-soil systems. Ecotoxicol. Environ. Saf. 2019, 167, 44–53. [Google Scholar] [CrossRef]
- Keesing, F.; Belden, L.K.; Daszak, P.; Dobson, A.; Harvell, C.D.; Holt, R.D.; Hudson, P.; Jolles, A.; Jones, K.E.; Mitchell, C.E.; et al. Impacts of biodiversity on the emergence and transmission of infectious diseases. Nature 2010, 468, 647–652. [Google Scholar] [CrossRef] [PubMed]
- Wall, D.H.; Nielsen, U.N.; Six, J. Soil biodiversity and human health. Nature 2015, 528, 69–76. [Google Scholar] [CrossRef]
- Cycon, M.; Mrozik, A.; Piotrowska-Seget, Z. Antibiotics in the Soil Environment-Degradation and Their Impact on Microbial Activity and Diversity. Front. Microbiol. 2019, 10, 338. [Google Scholar] [CrossRef] [PubMed]
- Gao, F.; Cui, S.; Li, P.; Wang, X.; Li, M.; Song, J.; Li, J.; Song, Y. Ecological toxicological effect of antibiotics in soil. IOP Conf. Ser. Earth Environ. Sci. 2018, 186, 012082. [Google Scholar] [CrossRef] [Green Version]
- Zhi, D.; Yang, D.; Zheng, Y.; Yang, Y.; He, Y.; Luo, L.; Zhou, Y. Current progress in the adsorption, transport and biodegradation of antibiotics in soil. J. Environ. Manage. 2019, 251, 109598. [Google Scholar] [CrossRef] [PubMed]
- Chander, Y.; Kumar, K.; Goyal, S.M.; Gupta, S.C. Antibacterial activity of soil-bound antibiotics. J. Environ. Qual. 2005, 34, 1952–1957. [Google Scholar] [CrossRef]
- Merino, D.; Tomadoni, B.; Salcedo, M.F.; Mansilla, A.Y.; Casalongué, C.A.; Alvarez, V.A. Nanoclay as Carriers of Bioactive Molecules Applied to Agriculture. Handb. Nanomater. Nanocomposites Energy Environ. Appl. 2020, 2020, 1–12. [Google Scholar] [CrossRef]
- Sanchez-Cid, C.; Guironnet, A.; Wiest, L.; Vulliet, E.; Vogel, T.M. Gentamicin Adsorption onto Soil Particles Prevents Overall Short-Term Effects on the Soil Microbiome and Resistome. Antibiotics 2021, 10, 191. [Google Scholar] [CrossRef]
- Wang, F.; Tiedje, J.M. Antibiotic Resistance in Soil. In Antibiotic Resistance in the Environment; Springer: New York, NY, USA, 2020; pp. 267–293. [Google Scholar] [CrossRef]
- Wang, F.; Xu, M.; Stedtfeld, R.D.; Sheng, H.; Fan, J.; Liu, M.; Chai, B.; Soares de Carvalho, T.; Li, H.; Li, Z.; et al. Long-Term Effect of Different Fertilization and Cropping Systems on the Soil Antibiotic Resistome. Environ. Sci. Technol. 2018, 52, 13037–13046. [Google Scholar] [CrossRef] [PubMed]
- Soucy, S.M.; Huang, J.; Gogarten, J.P. Horizontal gene transfer: Building the web of life. Nat. Rev. Genet. 2015, 16, 472–482. [Google Scholar] [CrossRef]
- Keeling, P.J.; Palmer, J.D. Horizontal gene transfer in eukaryotic evolution. Nat. Rev. Genet. 2008, 9, 605–618. [Google Scholar] [CrossRef]
- Cerqueira, F.; Matamoros, V.; Bayona, J.; Pina, B. Antibiotic resistance genes distribution in microbiomes from the soil-plant-fruit continuum in commercial Lycopersicon esculentum fields under different agricultural practices. Sci. Total Environ. 2019, 652, 660–670. [Google Scholar] [CrossRef] [PubMed]
- Tasho, R.P.; Cho, J.Y. Veterinary antibiotics in animal waste, its distribution in soil and uptake by plants: A review. Sci. Total Environ. 2016, 563, 366–376. [Google Scholar] [CrossRef] [PubMed]
- Hirt, H. Healthy soils for healthy plants for healthy humans: How beneficial microbes in the soil, food and gut are interconnected and how agriculture can contribute to human health. EMBO Rep. 2020, 21, e51069. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.; Hu, R.; Strong, P.J.; Zhuang, W.; Huang, W.; Luo, Z.; Yan, Q.; He, Z.; Shu, L. Prevalence of antibiotic resistance genes and bacterial pathogens along the soil-mangrove root continuum. J. Hazard. Mater. 2021, 408, 124985. [Google Scholar] [CrossRef] [PubMed]
- Liu, F.; Ying, G.G.; Tao, R.; Zhao, J.L.; Yang, J.F.; Zhao, L.F. Effects of six selected antibiotics on plant growth and soil microbial and enzymatic activities. Environ. Pollut. 2009, 157, 1636–1642. [Google Scholar] [CrossRef]
- Yu, X.; Liu, X.; Liu, H.; Chen, J.; Sun, Y. The accumulation and distribution of five antibiotics from soil in 12 cultivars of pak choi. Environ. Pollut. 2019, 254, 113115. [Google Scholar] [CrossRef]
- Tadic, D.; Bleda Hernandez, M.J.; Cerqueira, F.; Matamoros, V.; Pina, B.; Bayona, J.M. Occurrence and human health risk assessment of antibiotics and their metabolites in vegetables grown in field-scale agricultural systems. J. Hazard. Mater. 2021, 401, 123424. [Google Scholar] [CrossRef]
- Kumar, K.; Gupta, S.C.; Baidoo, S.K.; Chander, Y.; Rosen, C.J. Antibiotic uptake by plants from soil fertilized with animal manure. J. Environ. Qual. 2005, 34, 2082–2085. [Google Scholar] [CrossRef] [Green Version]
- Gudda, F.O.; Waigi, M.G.; Odinga, E.S.; Yang, B.; Carter, L.; Gao, Y. Antibiotic-contaminated wastewater irrigated vegetables pose resistance selection risks to the gut microbiome. Environ. Pollut. 2020, 264, 114752. [Google Scholar] [CrossRef]
- Li, M.Y.; Chen, X.Q.; Wang, J.Y.; Wang, H.T.; Xue, X.M.; Ding, J.; Juhasz, A.L.; Zhu, Y.G.; Li, H.B.; Ma, L.Q. Antibiotic exposure decreases soil arsenic oral bioavailability in mice by disrupting ileal microbiota and metabolic profile. Environ. Int. 2021, 151, 106444. [Google Scholar] [CrossRef] [PubMed]
- Dong, L.; Gao, J.; Xie, X.; Zhou, Q. DNA damage and biochemical toxicity of antibiotics in soil on the earthworm Eisenia fetida. Chemosphere 2012, 89, 44–51. [Google Scholar] [CrossRef] [PubMed]
- Yuan, J.; Wei, H.; Zeng, B.; Tang, H.; Li, W.; Zhang, Z. Impact of neonatal antibiotic treatment on the biodiversity of the murine intestinal Lactobacillus community. Curr. Microbiol. 2010, 60, 6–11. [Google Scholar] [CrossRef]
- Baquero, F.; Martinez, J.L.; Canton, R. Antibiotics and antibiotic resistance in water environments. Curr. Opin. Biotechnol. 2008, 19, 260–265. [Google Scholar] [CrossRef] [PubMed]
- Xue, X.; Jia, J.; Yue, X.; Guan, Y.; Zhu, L.; Wang, Z. River contamination shapes the microbiome and antibiotic resistance in sharpbelly (Hemiculter leucisculus). Environ. Pollut. 2021, 268, 115796. [Google Scholar] [CrossRef]
- Qian, M.; Wang, J.; Ji, X.; Yang, H.; Tang, B.; Zhang, H.; Yang, G.; Bao, Z.; Jin, Y. Sub-chronic exposure to antibiotics doxycycline, oxytetracycline or florfenicol impacts gut barrier and induces gut microbiota dysbiosis in adult zebrafish (Daino rerio). Ecotoxicol. Environ. Saf. 2021, 221, 112464. [Google Scholar] [CrossRef]
- Goncuoglu, M.; Bilir Ormanci, F.S.; Ayaz, N.D.; Erol, I. Antibiotic resistance of Escherichia coli O157:H7 isolated from cattle and sheep. Ann. Microbiol. 2010, 60, 489–494. [Google Scholar] [CrossRef]
- Askari, N.; Momtaz, H.; Tajbakhsh, E. Acinetobacter baumannii in sheep, goat, and camel raw meat: Virulence and antibiotic resistance pattern. AIMS Microbiol. 2019, 5, 272–284. [Google Scholar] [CrossRef]
- Guo, K.; Zhao, Y.; Cui, L.; Cao, Z.; Zhang, F.; Wang, X.; Peng, Z.; Feng, J.; Hu, T.; Dai, M. Longitudinal Surveillance and Risk Assessment of Resistance in Escherichia coli to Enrofloxacin from A Large-Scale Chicken Farm in Hebei, China. Antibiotics 2021, 10, 1222. [Google Scholar] [CrossRef]
- Han, T.; Zhang, Q.; Liu, N.; Wang, J.; Li, Y.; Huang, X.; Liu, J.; Wang, J.; Qu, Z.; Qi, K. Changes in antibiotic resistance of Escherichia coli during the broiler feeding cycle. Poult. Sci. 2020, 99, 6983–6989. [Google Scholar] [CrossRef] [PubMed]
- Stepien-Pysniak, D.; Hauschild, T.; Dec, M.; Marek, A.; Urban-Chmiel, R. Clonal Structure and Antibiotic Resistance of Enterococcus spp. from Wild Birds in Poland. Microb. Drug Resist. 2019, 25, 1227–1237. [Google Scholar] [CrossRef] [PubMed]
- Bhat, M.D.; Vijay Kumar, D.; Kumar, B.; Shetty, V.; Chakraborty, A.; Karunasagar, I. Occurrence of antibiotic resistance among Gram negative bacteria isolated from effluents of fish processing plants in and around Mangalore. Int. J. Environ. Health Res. 2019, 30, 1–8. [Google Scholar] [CrossRef]
- Fernandez Marquez, M.L.; Burgos, M.J.; Pulido, R.P.; Galvez, A.; Lopez, R.L. Biocide Tolerance and Antibiotic Resistance in Salmonella Isolates from Hen Eggshells. Foodborne Pathog. Dis. 2017, 14, 89–95. [Google Scholar] [CrossRef]
- Lim, S.K.; Kim, D.; Moon, D.C.; Cho, Y.; Rho, M. Antibiotic resistomes discovered in the gut microbiomes of Korean swine and cattle. Gigascience 2020, 9, giaa043. [Google Scholar] [CrossRef] [PubMed]
- Pexara, A.; Solomakos, N.; Govaris, A. Occurrence, antibiotic resistance and enteroxigenicity of Staphylococcus spp. in tonsils of slaughtered pigs in Greece. Lett. Appl. Microbiol. 2020, 71, 394–399. [Google Scholar] [CrossRef] [PubMed]
- Eltai, N.; Al Thani, A.A.; Al-Hadidi, S.H.; Abdfarag, E.A.; Al-Romaihi, H.; Mahmoud, M.H.; Alawad, O.K.; Yassine, H.M. Antibiotic resistance profile of commensal Escherichia coli isolated from healthy sheep in Qatar. J. Infect. Dev. Ctries 2020, 14, 138–145. [Google Scholar] [CrossRef] [Green Version]
- Adinortey, C.A.; Aheto, D.W.; Boateng, A.A.; Agbeko, R. Multiple Antibiotic Resistance-Coliform Bacteria in Some Selected Fish Farms of the Central Region of Ghana. Scientifica 2020, 2020, 6641461. [Google Scholar] [CrossRef]
- Wang, J.; Wang, L.; Zhu, L.; Wang, J.; Xing, B. Antibiotic resistance in agricultural soils: Source, fate, mechanism and attenuation strategy. Crit. Rev. Environ. Sci. Technol. 2020, 43, 1835438. [Google Scholar] [CrossRef]
- Tyrrell, C.; Burgess, C.M.; Brennan, F.P.; Walsh, F. Antibiotic resistance in grass and soil. Biochem. Soc. Trans. 2019, 47, 477–486. [Google Scholar] [CrossRef] [PubMed]
- Smalla, K.; Cook, K.; Djordjevic, S.P.; Klumper, U.; Gillings, M. Environmental dimensions of antibiotic resistance: Assessment of basic science gaps. FEMS Microbiol. Ecol. 2018, 94, fiy195. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jiao, S.; Chen, W.; Wei, G. Resilience and Assemblage of Soil Microbiome in Response to Chemical Contamination Combined with Plant Growth. Appl. Environ. Microbiol. 2019, 85, e02523-18. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, Y.J.; Hu, H.W.; Yan, H.; Wang, J.T.; Lam, S.K.; Chen, Q.L.; Chen, D.; He, J.Z. Salinity as a predominant factor modulating the distribution patterns of antibiotic resistance genes in ocean and river beach soils. Sci. Total Environ. 2019, 668, 193–203. [Google Scholar] [CrossRef]
- Hill, K.E.; Top, E.M. Gene transfer in soil systems using microcosms. FEMS Microbiol. Ecol. 1998, 25, 319–329. [Google Scholar] [CrossRef]
- Beceiro, A.; Tomas, M.; Bou, G. Antimicrobial resistance and virulence: A successful or deleterious association in the bacterial world? Clin. Microbiol. Rev. 2013, 26, 185–230. [Google Scholar] [CrossRef] [Green Version]
- Calanca, P.P. Effects of Abiotic Stress in Crop Production. In Quantification of Climate Variability, Adaptation and Mitigation for Agricultural Sustainability; Springer: Cham, Switzerland, 2017; pp. 165–180. [Google Scholar] [CrossRef]
- Francini, A.; Sebastiani, L. Abiotic Stress Effects on Performance of Horticultural Crops. Horticulturae 2019, 5, 67. [Google Scholar] [CrossRef] [Green Version]
- Nyamangara, J.; Kodzwa, J.; Masvaya, E.N.; Soropa, G. The role of synthetic fertilizers in enhancing ecosystem services in crop production systems in developing countries. In The Role of Ecosystem Services in Sustainable Food Systems; Academic Press: Cambridge, MA, USA, 2020; pp. 95–117. [Google Scholar] [CrossRef]
- Bhattacharya, A. Global Climate Change and Its Impact on Agriculture. In Changing Climate and Resource Use Efficiency in Plants; Academic Press: Cambridge, MA, USA, 2019; pp. 1–50. [Google Scholar] [CrossRef]
- Cerqueira, F.; Matamoros, V.; Bayona, J.M.; Berendonk, T.U.; Elsinga, G.; Hornstra, L.M.; Pina, B. Antibiotic resistance gene distribution in agricultural fields and crops. A soil-to-food analysis. Environ. Res. 2019, 177, 108608. [Google Scholar] [CrossRef]
- Sun, Y.; Qiu, T.; Gao, M.; Shi, M.; Zhang, H.; Wang, X. Inorganic and organic fertilizers application enhanced antibiotic resistome in greenhouse soils growing vegetables. Ecotoxicol. Environ. Saf. 2019, 179, 24–30. [Google Scholar] [CrossRef]
- Wang, F.; Han, W.; Chen, S.; Dong, W.; Qiao, M.; Hu, C.; Liu, B. Fifteen-Year Application of Manure and Chemical Fertilizers Differently Impacts Soil ARGs and Microbial Community Structure. Front. Microbiol. 2020, 11, 62. [Google Scholar] [CrossRef]
- Wang, F.; Chen, S.; Wang, Y.; Zhang, Y.; Hu, C.; Liu, B. Long-Term Nitrogen Fertilization Elevates the Activity and Abundance of Nitrifying and Denitrifying Microbial Communities in an Upland Soil: Implications for Nitrogen Loss from Intensive Agricultural Systems. Front. Microbiol. 2018, 9, 2424. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Curutiu, C.; Lazar, V.; Chifiriuc, M.C. Pesticides and antimicrobial resistance: From environmental compartments to animal and human infections. In New Pesticides and Soil Sensors; Academic Press: Cambridge, MA, USA, 2017; pp. 373–392. [Google Scholar] [CrossRef]
- Malagon-Rojas, J.N.; Parra Barrera, E.L.; Lagos, L. From environment to clinic: The role of pesticides in antimicrobial resistance. Rev. Panam. Salud. Publica 2020, 44, e44. [Google Scholar] [CrossRef] [PubMed]
- Kang, Y.; Hao, Y.; Shen, M.; Zhao, Q.; Li, Q.; Hu, J. Impacts of supplementing chemical fertilizers with organic fertilizers manufactured using pig manure as a substrate on the spread of tetracycline resistance genes in soil. Ecotoxicol. Environ. Saf. 2016, 130, 279–288. [Google Scholar] [CrossRef] [PubMed]
- Xie, W.-Y.; Yuan, S.-T.; Xu, M.-G.; Yang, X.-P.; Shen, Q.-R.; Zhang, W.-W.; Su, J.-Q.; Zhao, F.-J. Long-term effects of manure and chemical fertilizers on soil antibiotic resistome. Soil Biol. Biochem. 2018, 122, 111–119. [Google Scholar] [CrossRef]
- McKinney, C.W.; Dungan, R.S.; Moore, A.; Leytem, A.B. Occurrence and abundance of antibiotic resistance genes in agricultural soil receiving dairy manure. FEMS Microbiol. Ecol. 2018, 94, fiy010. [Google Scholar] [CrossRef] [PubMed]
- Zhao, F.; Chen, L.; Yen, H.; Sun, L.; Li, S.; Li, M.; Feng, Q.; Yang, L. Multimedia mass balance approach to characterizing the transport potential of antibiotics in soil-plant systems following manure application. J. Hazard. Mater. 2020, 393, 122363. [Google Scholar] [CrossRef]
- Qian, M.; Wu, H.; Wang, J.; Zhang, H.; Zhang, Z.; Zhang, Y.; Lin, H.; Ma, J. Occurrence of trace elements and antibiotics in manure-based fertilizers from the Zhejiang Province of China. Sci. Total Environ. 2016, 559, 174–181. [Google Scholar] [CrossRef]
- Martinez-Carballo, E.; Gonzalez-Barreiro, C.; Scharf, S.; Gans, O. Environmental monitoring study of selected veterinary antibiotics in animal manure and soils in Austria. Environ. Pollut. 2007, 148, 570–579. [Google Scholar] [CrossRef]
- Ruuskanen, M.; Muurinen, J.; Meierjohan, A.; Parnanen, K.; Tamminen, M.; Lyra, C.; Kronberg, L.; Virta, M. Fertilizing with Animal Manure Disseminates Antibiotic Resistance Genes to the Farm Environment. J. Environ. Qual. 2016, 45, 488–493. [Google Scholar] [CrossRef]
- Liu, P.; Jia, S.; He, X.; Zhang, X.; Ye, L. Different impacts of manure and chemical fertilizers on bacterial community structure and antibiotic resistance genes in arable soils. Chemosphere 2017, 188, 455–464. [Google Scholar] [CrossRef]
- Heuer, H.; Kopmann, C.; Binh, C.T.; Top, E.M.; Smalla, K. Spreading antibiotic resistance through spread manure: Characteristics of a novel plasmid type with low %G + C content. Environ. Microbiol. 2009, 11, 937–949. [Google Scholar] [CrossRef] [PubMed]
- Wei, R.; He, T.; Zhang, S.; Zhu, L.; Shang, B.; Li, Z.; Wang, R. Occurrence of seventeen veterinary antibiotics and resistant bacterias in manure-fertilized vegetable farm soil in four provinces of China. Chemosphere 2019, 215, 234–240. [Google Scholar] [CrossRef] [PubMed]
- Han, L.; Cai, L.; Zhang, H.; Long, Z.; Yu, Y.; Fang, H. Development of antibiotic resistance genes in soils with ten successive treatments of chlortetracycline and ciprofloxacin. Environ. Pollut. 2019, 253, 152–160. [Google Scholar] [CrossRef]
- Ma, J.; Zhu, D.; Chen, Q.L.; Ding, J.; Zhu, Y.G.; Sheng, G.D.; Qiu, Y.P. Exposure to tetracycline perturbs the microbiome of soil oligochaete Enchytraeus crypticus. Sci. Total Environ. 2019, 654, 643–650. [Google Scholar] [CrossRef]
- Liu, F.; Wu, J.; Ying, G.G.; Luo, Z.; Feng, H. Changes in functional diversity of soil microbial community with addition of antibiotics sulfamethoxazole and chlortetracycline. Appl. Microbiol. Biotechnol. 2012, 95, 1615–1623. [Google Scholar] [CrossRef] [PubMed]
- Du, S.; Shen, J.-P.; Sun, Y.-F.; Bai, Y.-F.; Pan, H.; Li, Y.; Wang, Z.-W.; Han, G.-D.; Zhang, L.-M.; He, J.-Z. Grazing does not increase soil antibiotic resistome in two types of grasslands in Inner Mongolia, China. Appl. Soil Ecol. 2020, 155, 103644. [Google Scholar] [CrossRef]
- Zhou, X.; Qiao, M.; Wang, F.H.; Zhu, Y.G. Use of commercial organic fertilizer increases the abundance of antibiotic resistance genes and antibiotics in soil. Environ. Sci. Pollut. Res. Int. 2017, 24, 701–710. [Google Scholar] [CrossRef]
- Liu, W.; Ling, N.; Guo, J.; Ruan, Y.; Wang, M.; Shen, Q.; Guo, S. Dynamics of the antibiotic resistome in agricultural soils amended with different sources of animal manures over three consecutive years. J. Hazard. Mater. 2021, 401, 123399. [Google Scholar] [CrossRef]
- Frost, L.S.; Leplae, R.; Summers, A.O.; Toussaint, A. Mobile genetic elements: The agents of open source evolution. Nat. Rev. Microbiol. 2005, 3, 722–732. [Google Scholar] [CrossRef]
- Solliec, M.; Roy-Lachapelle, A.; Sauve, S. Development of a suspect and non-target screening approach to detect veterinary antibiotic residues in a complex biological matrix using liquid chromatography/high-resolution mass spectrometry. Rapid Commun. Mass Spectrom. 2015, 29, 2361–2373. [Google Scholar] [CrossRef]
- Yan, Q.; Li, X.; Ma, B.; Zou, Y.; Wang, Y.; Liao, X.; Liang, J.; Mi, J.; Wu, Y. Different Concentrations of Doxycycline in Swine Manure Affect the Microbiome and Degradation of Doxycycline Residue in Soil. Front. Microbiol. 2018, 9, 3129. [Google Scholar] [CrossRef] [PubMed]
- Chen, Z.; Zhang, W.; Yang, L.; Stedtfeld, R.D.; Peng, A.; Gu, C.; Boyd, S.A.; Li, H. Antibiotic resistance genes and bacterial communities in cornfield and pasture soils receiving swine and dairy manures. Environ. Pollut. 2019, 248, 947–957. [Google Scholar] [CrossRef] [PubMed]
- Yang, Q.; Ren, S.; Niu, T.; Guo, Y.; Qi, S.; Han, X.; Liu, D.; Pan, F. Distribution of antibiotic-resistant bacteria in chicken manure and manure-fertilized vegetables. Environ. Sci. Pollut. Res. Int. 2014, 21, 1231–1241. [Google Scholar] [CrossRef] [PubMed]
- Munir, M.; Xagoraraki, I. Levels of antibiotic resistance genes in manure, biosolids, and fertilized soil. J. Environ. Qual. 2011, 40, 248–255. [Google Scholar] [CrossRef] [Green Version]
- Eurostat. Archive: Agri-Environmental Indicator—Manure Application. 2013. Available online: https://ec.europa.eu/eurostat/statistics-explained/index.php?title=Archive:Agri-environmental_indicator_-_manure_application#Further_Eurostat_information (accessed on 20 November 2021).
- Zhang, Y.; Gu, A.; Cen, T.; Li, X.; He, M.; Li, D.; Chen, J.-M. Sub-inhibitory concentrations of heavy metals facilitate the horizontal transfer of plasmid-mediated antibiotic resistance genes in water environment. Environ. Pollut. 2018, 237, 74–82. [Google Scholar] [CrossRef]
- Kang, C.-H.; So, J.-S. Heavy metal and antibiotic resistance of ureolytic bacteria and their immobilization of heavy metals. Ecol. Eng. 2016, 97, 304–312. [Google Scholar] [CrossRef]
- Gorovtsov, A.V.; Sazykin, I.S.; Sazykina, M.A. The influence of heavy metals, polyaromatic hydrocarbons, and polychlorinated biphenyls pollution on the development of antibiotic resistance in soils. Environ. Sci. Pollut. Res. Int. 2018, 25, 9283–9292. [Google Scholar] [CrossRef]
- Zhao, Z.; Wang, J.; Han, Y.; Chen, J.; Liu, G.; Lu, H.; Yan, B.; Chen, S. Nutrients, heavy metals and microbial communities co-driven distribution of antibiotic resistance genes in adjacent environment of mariculture. Environ. Pollut. 2017, 220, 909–918. [Google Scholar] [CrossRef]
- Chen, Q.L.; Zhu, D.; An, X.L.; Ding, J.; Zhu, Y.G.; Cui, L. Does nano silver promote the selection of antibiotic resistance genes in soil and plant? Environ. Int. 2019, 128, 399–406. [Google Scholar] [CrossRef]
- Yang, Y.; Xu, C.; Cao, X.; Lin, H.; Wang, J. Antibiotic resistance genes in surface water of eutrophic urban lakes are related to heavy metals, antibiotics, lake morphology and anthropic impact. Ecotoxicology 2017, 26, 831–840. [Google Scholar] [CrossRef]
- Ji, X.; Shen, Q.; Liu, F.; Ma, J.; Xu, G.; Wang, Y.; Wu, M. Antibiotic resistance gene abundances associated with antibiotics and heavy metals in animal manures and agricultural soils adjacent to feedlots in Shanghai, China. J. Hazard. Mater. 2012, 235, 178–185. [Google Scholar] [CrossRef]
- Lu, L.; Liu, J.; Li, Z.; Zou, X.; Guo, J.; Liu, Z.; Yang, J.; Zhou, Y. Antibiotic resistance gene abundances associated with heavy metals and antibiotics in the sediments of Changshou Lake in the three Gorges Reservoir area, China. Ecol. Indic. 2020, 113, 106275. [Google Scholar] [CrossRef]
- Zhou, B.; Wang, C.; Zhao, Q.; Wang, Y.; Huo, M.; Wang, J.; Wang, S. Prevalence and dissemination of antibiotic resistance genes and coselection of heavy metals in Chinese dairy farms. J. Hazard. Mater. 2016, 320, 10–17. [Google Scholar] [CrossRef]
- Li, H.; Tian, Y.; Liu, W.; Long, Y.; Ye, J.; Li, B.; Li, N.; Yan, M.; Zhu, C. Impact of electrokinetic remediation of heavy metal contamination on antibiotic resistance in soil. Chem. Eng. J. 2020, 400, 125866. [Google Scholar] [CrossRef]
- Mazhar, S.H.; Li, X.; Rashid, A.; Su, J.; Xu, J.; Brejnrod, A.D.; Su, J.Q.; Wu, Y.; Zhu, Y.G.; Zhou, S.G.; et al. Co-selection of antibiotic resistance genes, and mobile genetic elements in the presence of heavy metals in poultry farm environments. Sci. Total Environ. 2021, 755, 142702. [Google Scholar] [CrossRef]
- Nguyen, C.C.; Hugie, C.N.; Kile, M.L.; Navab-Daneshmand, T. Association between heavy metals and antibiotic-resistant human pathogens in environmental reservoirs: A review. Front. Environ. Sci. Eng. 2019, 13, 1–17. [Google Scholar] [CrossRef]
- Seiler, C.; Berendonk, T.U. Heavy metal driven co-selection of antibiotic resistance in soil and water bodies impacted by agriculture and aquaculture. Front. Microbiol. 2012, 3, 399. [Google Scholar] [CrossRef] [Green Version]
- Safari Sinegani, A.A.; Younessi, N. Antibiotic resistance of bacteria isolated from heavy metal-polluted soils with different land uses. J. Glob. Antimicrob. Resist. 2017, 10, 247–255. [Google Scholar] [CrossRef] [PubMed]
- Yamamura, S.; Watanabe, K.; Suda, W.; Tsuboi, S.; Watanabe, M. Effect of antibiotics on redox transformations of arsenic and diversity of arsenite-oxidizing bacteria in sediment microbial communities. Environ. Sci. Technol. 2014, 48, 350–357. [Google Scholar] [CrossRef]
- Chen, S.; Li, X.; Sun, G.; Zhang, Y.; Su, J.; Ye, J. Heavy Metal Induced Antibiotic Resistance in Bacterium LSJC7. Int. J. Mol. Sci. 2015, 16, 23390–23404. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, X.; Gu, A.Z.; Zhang, Y.; Xie, B.; Li, D.; Chen, J. Sub-lethal concentrations of heavy metals induce antibiotic resistance via mutagenesis. J. Hazard. Mater. 2019, 369, 9–16. [Google Scholar] [CrossRef] [PubMed]
- Zhang, F.; Zhao, X.; Li, Q.; Liu, J.; Ding, J.; Wu, H.; Zhao, Z.; Ba, Y.; Cheng, X.; Cui, L.; et al. Bacterial community structure and abundances of antibiotic resistance genes in heavy metals contaminated agricultural soil. Environ. Sci. Pollut. Res. Int. 2018, 25, 9547–9555. [Google Scholar] [CrossRef]
- Xing, B.S.; Jin, R.C. Inhibitory effects of heavy metals and antibiotics on nitrifying bacterial activities in mature partial nitritation. Chemosphere 2018, 200, 437–445. [Google Scholar] [CrossRef]
- Benghait, Y.; Blaghen, M. Heavy metals and antibiotics resistance of bacteria isolated from Marchica lagoon: Biodegradation of anthracene on submerged aerated fixed bed reactor. Environ. Technol. 2020, 10, 1839133. [Google Scholar] [CrossRef] [PubMed]
- Nourhene, S.; Rihab, L.; Fethi, B.A.; Karima, B.R.; Amina, B. Slime producing, heavy metals and antibiotics resistance in Aeromonas hydrophila isolated in Tunisia. Afr. J. Microbiol. Res. 2013, 7, 5697–5708. [Google Scholar] [CrossRef] [Green Version]
- Oyetibo, G.O.; Ilori, M.O.; Adebusoye, S.A.; Obayori, O.S.; Amund, O.O. Bacteria with dual resistance to elevated concentrations of heavy metals and antibiotics in Nigerian contaminated systems. Environ. Monit. Assess. 2010, 168, 305–314. [Google Scholar] [CrossRef]
- Yamina, B.; Tahar, B.; Marie Laure, F. Isolation and screening of heavy metal resistant bacteria from wastewater: A study of heavy metal co-resistance and antibiotics resistance. Water Sci. Technol. 2012, 66, 2041–2048. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.; Xu, Y.B.; Xu, J.X.; Zhang, X.H.; Xu, S.H.; Du, Q.P. Combined toxic effects of heavy metals and antibiotics on a Pseudomonas fluorescens strain ZY2 isolated from swine wastewater. Int. J. Mol. Sci. 2015, 16, 2839–2850. [Google Scholar] [CrossRef]
- Abdelrehim, K.; Soltan, E.-S.; Abu-Garbia, M.; El-Zien, F. Heavy Metals and Antibiotics Resistance of Halophilic Bacteria Isolated from Different Areas in Red Sea, Egypt. Egypt. Acad. J. Biol. Sci. G Microbiol. 2014, 6, 77–89. [Google Scholar] [CrossRef]
- Bhardwaj, R.; Gupta, A.; Garg, J.K. Impact of heavy metals on inhibitory concentration of Escherichia coli-a case study of river Yamuna system, Delhi, India. Environ. Monit. Assess. 2018, 190, 674. [Google Scholar] [CrossRef]
- Sharifi, Y.; Abedzadeh, A.; Salighe, A.; Kalhor, N. Antibiotics and heavy metals resistance patterns of Enterococcus faecalis and faecium bacteria isolated from the human and the livestock sources. Environ. Health Eng. Manag. J. 2015, 2, 199–202. [Google Scholar]
- Adekanmbi, A.O.; Falodun, O.I. Heavy Metals and Antibiotics Susceptibility Profiles of Staphylococcus aureus; Isolated from Several Points Receiving Daily Input from the Bodija Abattoir in Ibadan, Oyo State, Nigeria. Adv. Microbiol. 2015, 5, 871–880. [Google Scholar] [CrossRef] [Green Version]
- Li, G.; Sun, G.X.; Ren, Y.; Luo, X.S.; Zhu, Y.G. Urban soil and human health: A review. Eur. J. Soil Sci. 2018, 69, 196–215. [Google Scholar] [CrossRef] [Green Version]
- Zhou, C.; Pan, Y.; Ge, S.; Coulon, F.; Yang, Z. Rapid methods for antimicrobial resistance diagnosis in contaminated soils for effective remediation strategy. TrAC Trends Anal. Chem. 2021, 137, 116203. [Google Scholar] [CrossRef]
- Wang, J.; Wang, J.; Zhao, Z.; Chen, J.; Lu, H.; Liu, G.; Zhou, J.; Guan, X. PAHs accelerate the propagation of antibiotic resistance genes in coastal water microbial community. Environ. Pollut. 2017, 231, 1145–1152. [Google Scholar] [CrossRef] [PubMed]
- Chen, B.; He, R.; Yuan, K.; Chen, E.; Lin, L.; Chen, X.; Sha, R.; Zhong, J.; Lin, L.; Yang, L.; et al. Polycyclic aromatic hydrocarbons (PAHs) enriching antibiotic resistance genes (ARGs) in the soils. Environ. Pollut. 2016, 220, 1005–1013. [Google Scholar] [CrossRef] [Green Version]
- Amala, S.E.; Agwor, N.O.; Vivian Agi, N.; Monsi, T.P. Evaluation of the Impact of Hydrocarbon-Generated Soot on Antibiotics Susceptibility of Staphylococcus aureus and Escherichia coli Isolates. Adv. Microbiol. 2021, 11, 444–452. [Google Scholar] [CrossRef]
- Cunningham, C.; Kuyukina, M.; Ivshina, I.; Konev, A.; Peshkur, T.; Knapp, C. Potential risks of antibiotic resistant bacteria and genes in bioremediation of petroleum hydrocarbon contaminated soils. Environ. Sci. Processes Impacts 2020, 22, 1110–1124. [Google Scholar] [CrossRef]
- Das, N.; Kotoky, R.; Maurya, A.P.; Bhuyan, B.; Pandey, P. Paradigm shift in antibiotic-resistome of petroleum hydrocarbon contaminated soil. Sci. Total Environ. 2021, 757, 143777. [Google Scholar] [CrossRef]
- Hemala, L.; Zhang, D.; Margesin, R. Cold-active antibacterial and antifungal activities and antibiotic resistance of bacteria isolated from an alpine hydrocarbon-contaminated industrial site. Res. Microbiol. 2014, 165, 447–456. [Google Scholar] [CrossRef]
- Wetherill, Y.B.; Akingbemi, B.T.; Kanno, J.; McLachlan, J.A.; Nadal, A.; Sonnenschein, C.; Watson, C.S.; Zoeller, R.T.; Belcher, S.M. In vitro molecular mechanisms of bisphenol A action. Reprod. Toxicol. 2007, 24, 178–198. [Google Scholar] [CrossRef]
- Vandenberg, L.N.; Hauser, R.; Marcus, M.; Olea, N.; Welshons, W.V. Human exposure to bisphenol A (BPA). Reprod. Toxicol. 2007, 24, 139–177. [Google Scholar] [CrossRef] [PubMed]
- Rochester, J.R. Bisphenol A and human health: A review of the literature. Reprod. Toxicol. 2013, 42, 132–155. [Google Scholar] [CrossRef] [PubMed]
- Pop, C.-E.; Draga, S.; Măciucă, R.; Niță, R.; Crăciun, N.; Wolff, R. Bisphenol a Effects in Aqueous Environment on Lemna minor. Processes 2021, 9, 1512. [Google Scholar] [CrossRef]
- Eladak, S.; Grisin, T.; Moison, D.; Guerquin, M.J.; N’Tumba-Byn, T.; Pozzi-Gaudin, S.; Benachi, A.; Livera, G.; Rouiller-Fabre, V.; Habert, R. A new chapter in the bisphenol A story: Bisphenol S and bisphenol F are not safe alternatives to this compound. Fertil. Steril. 2015, 103, 11–21. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Russell, A.D. Do Biocides Select for Antibiotic Resistance? J. Pharm. Pharmacol. 2000, 52, 227–233. [Google Scholar] [CrossRef] [PubMed]
- Hartmann, E.M.; Hickey, R.; Hsu, T.; Betancourt Roman, C.M.; Chen, J.; Schwager, R.; Kline, J.; Brown, G.Z.; Halden, R.U.; Huttenhower, C.; et al. Antimicrobial Chemicals Are Associated with Elevated Antibiotic Resistance Genes in the Indoor Dust Microbiome. Environ. Sci. Technol. 2016, 50, 9807–9815. [Google Scholar] [CrossRef]
- Reichert, G.; Hilgert, S.; Fuchs, S.; Azevedo, J.C.R. Emerging contaminants and antibiotic resistance in the different environmental matrices of Latin America. Environ. Pollut. 2019, 255, 113140. [Google Scholar] [CrossRef]
- Maurya, A.P.; Rajkumari, J.; Pandey, P. Enrichment of antibiotic resistance genes (ARGs) in polyaromatic hydrocarbon-contaminated soils: A major challenge for environmental health. Environ. Sci. Pollut. Res. Int. 2021, 28, 12178–12189. [Google Scholar] [CrossRef]
- European Environment Agency. Overview of Contaminants Affecting Soil and Groundwater in Europe. Available online: https://www.eea.europa.eu/data-and-maps/figures/overview-of-contaminants-affecting-soil-and-groundwater-in-europe (accessed on 2 January 2022).
- Ya, H.; Jiang, B.; Xing, Y.; Zhang, T.; Lv, M.; Wang, X. Recent advances on ecological effects of microplastics on soil environment. Sci. Total Environ. 2021, 798, 149338. [Google Scholar] [CrossRef]
- Zhang, Y.; Lu, J.; Wu, J.; Wang, J.; Luo, Y. Potential risks of microplastics combined with superbugs: Enrichment of antibiotic resistant bacteria on the surface of microplastics in mariculture system. Ecotoxicol. Environ. Saf. 2020, 187, 109852. [Google Scholar] [CrossRef]
- Liu, Y.; Liu, W.; Yang, X.; Wang, J.; Lin, H.; Yang, Y. Microplastics are a hotspot for antibiotic resistance genes: Progress and perspective. Sci. Total Environ. 2021, 773, 145643. [Google Scholar] [CrossRef] [PubMed]
- Kumar, M.; Xiong, X.; He, M.; Tsang, D.C.W.; Gupta, J.; Khan, E.; Harrad, S.; Hou, D.; Ok, Y.S.; Bolan, N.S. Microplastics as pollutants in agricultural soils. Environ. Pollut. 2020, 265, 114980. [Google Scholar] [CrossRef] [PubMed]
- Ma, J.; Sheng, G.D.; O’Connor, P. Microplastics combined with tetracycline in soils facilitate the formation of antibiotic resistance in the Enchytraeus crypticus microbiome. Environ. Pollut. 2020, 264, 114689. [Google Scholar] [CrossRef] [PubMed]
- Yan, X.; Yang, X.; Tang, Z.; Fu, J.; Chen, F.; Zhao, Y.; Ruan, L.; Yang, Y. Downward transport of naturally-aged light microplastics in natural loamy sand and the implication to the dissemination of antibiotic resistance genes. Environ. Pollut. 2020, 262, 114270. [Google Scholar] [CrossRef]
- Huang, F.Y.; Yang, K.; Zhang, Z.X.; Su, J.Q.; Zhu, Y.G.; Zhang, X. Effects of Microplastics on Antibiotic Resistance Genes in Estuarine Sediments. Huan Jing Ke Xue 2019, 40, 2234–2239. [Google Scholar] [CrossRef] [PubMed]
- Peng, C.; Zhang, X.; Zhang, X.; Liu, C.; Chen, Z.; Sun, H.; Wang, L. Bacterial Community under the Influence of Microplastics in Indoor Environment and the Health Hazards Associated with Antibiotic Resistance Genes. Environ. Sci. Technol. 2021, 56, 422–432. [Google Scholar] [CrossRef]
- Wang, Y.; Wang, X.; Li, Y.; Liu, Y.; Sun, Y.; Xia, S.; Zhao, J. Effects of coexistence of tetracycline, copper and microplastics on the fate of antibiotic resistance genes in manured soil. Sci. Total Environ. 2021, 790, 148087. [Google Scholar] [CrossRef]
- Shi, J.; Wu, D.; Su, Y.; Xie, B. Selective enrichment of antibiotic resistance genes and pathogens on polystyrene microplastics in landfill leachate. Sci. Total Environ. 2021, 765, 142775. [Google Scholar] [CrossRef]
- Sucato, A.; Vecchioni, L.; Savoca, D.; Presentato, A.; Arculeo, M.; Alduina, R. A Comparative Analysis of Aquatic and Polyethylene-Associated Antibiotic-Resistant Microbiota in the Mediterranean Sea. Biology 2021, 10, 200. [Google Scholar] [CrossRef]
- Zhu, D.; Ma, J.; Li, G.; Rillig, M.C.; Zhu, Y.G. Soil plastispheres as hotpots of antibiotic resistance genes and potential pathogens. ISME J. 2021, 2021, 1–12. [Google Scholar] [CrossRef]
- Sathicq, M.B.; Sabatino, R.; Corno, G.; Di Cesare, A. Are microplastic particles a hotspot for the spread and the persistence of antibiotic resistance in aquatic systems? Environ. Pollut. 2021, 279, 116896. [Google Scholar] [CrossRef]
- Imran, M.; Das, K.R.; Naik, M.M. Co-selection of multi-antibiotic resistance in bacterial pathogens in metal and microplastic contaminated environments: An emerging health threat. Chemosphere 2019, 215, 846–857. [Google Scholar] [CrossRef] [PubMed]
- Gao, D.; Li, X.Y.; Liu, H.T. Source, occurrence, migration and potential environmental risk of microplastics in sewage sludge and during sludge amendment to soil. Sci. Total Environ. 2020, 742, 140355. [Google Scholar] [CrossRef] [PubMed]
- Zhu, F.; Zhu, C.; Wang, C.; Gu, C. Occurrence and Ecological Impacts of Microplastics in Soil Systems: A Review. Bull. Environ. Contam. Toxicol. 2019, 102, 741–749. [Google Scholar] [CrossRef] [PubMed]
- Zhang, B.; Yang, X.; Chen, L.; Chao, J.; Teng, J.; Wang, Q. Microplastics in soils: A review of possible sources, analytical methods and ecological impacts. J. Chem. Technol. Biotechnol. 2020, 95, 2052–2068. [Google Scholar] [CrossRef]
- Cerqueira, F.; Christou, A.; Fatta-Kassinos, D.; Vila-Costa, M.; Bayona, J.M.; Pina, B. Effects of prescription antibiotics on soil- and root-associated microbiomes and resistomes in an agricultural context. J. Hazard. Mater. 2020, 400, 123208. [Google Scholar] [CrossRef]
- Urra, J.; Alkorta, I.; Mijangos, I.; Epelde, L.; Garbisu, C. Application of sewage sludge to agricultural soil increases the abundance of antibiotic resistance genes without altering the composition of prokaryotic communities. Sci. Total Environ. 2019, 647, 1410–1420. [Google Scholar] [CrossRef]
- Xu, S.; Lu, W.; Qasim, M.Z. High-throughput characterization of the expressed antibiotic resistance genes in sewage sludge with transcriptional analysis. Ecotoxicol. Environ. Saf. 2020, 205, 111377. [Google Scholar] [CrossRef]
- Venkatesan, A.K.; Done, H.Y.; Halden, R.U. United States National Sewage Sludge Repository at Arizona State University—A new resource and research tool for environmental scientists, engineers, and epidemiologists. Environ. Sci. Pollut. Res. Int. 2015, 22, 1577–1586. [Google Scholar] [CrossRef] [Green Version]
- Poszytek, K.; Karczewska-Golec, J.; Ciok, A.; Decewicz, P.; Dziurzynski, M.; Gorecki, A.; Jakusz, G.; Krucon, T.; Lomza, P.; Romaniuk, K.; et al. Genome-Guided Characterization of Ochrobactrum sp. POC9 Enhancing Sewage Sludge Utilization-Biotechnological Potential and Biosafety Considerations. Int. J. Environ. Res. Public Health 2018, 15, 1501. [Google Scholar] [CrossRef] [Green Version]
- Urra, J.; Alkorta, I.; Mijangos, I.; Garbisu, C. Data on links between structural and functional prokaryotic diversity in long-term sewage sludge amended soil. Data Brief. 2018, 20, 1787–1796. [Google Scholar] [CrossRef]
- Karkman, A.; Do, T.T.; Walsh, F.; Virta, M.P.J. Antibiotic-Resistance Genes in Waste Water. Trends Microbiol. 2018, 26, 220–228. [Google Scholar] [CrossRef] [Green Version]
- Yang, Y.; Li, B.; Zou, S.; Fang, H.H.P.; Zhang, T. Fate of antibiotic resistance genes in sewage treatment plant revealed by metagenomic approach. Water Res. 2014, 62, 97–106. [Google Scholar] [CrossRef]
- Rahube, T.O.; Marti, R.; Scott, A.; Tien, Y.C.; Murray, R.; Sabourin, L.; Duenk, P.; Lapen, D.R.; Topp, E. Persistence of antibiotic resistance and plasmid-associated genes in soil following application of sewage sludge and abundance on vegetables at harvest. Can. J. Microbiol. 2016, 62, 600–607. [Google Scholar] [CrossRef] [PubMed]
- Lyu, J.; Yang, L.; Zhang, L.; Ye, B.; Wang, L. Antibiotics in soil and water in China-a systematic review and source analysis. Environ. Pollut. 2020, 266, 115147. [Google Scholar] [CrossRef] [PubMed]
- Gatica, J.; Cytryn, E. Impact of treated wastewater irrigation on antibiotic resistance in the soil microbiome. Environ. Sci. Pollut. Res. Int. 2013, 20, 3529–3538. [Google Scholar] [CrossRef] [Green Version]
- Kummerer, K. Significance of antibiotics in the environment. J. Antimicrob. Chemother. 2003, 52, 5–7. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Khan, N.A.; Ahmed, S.; Farooqi, I.H.; Ali, I.; Vambol, V.; Changani, F.; Yousefi, M.; Vambol, S.; Khan, S.U.; Khan, A.H. Occurrence, sources and conventional treatment techniques for various antibiotics present in hospital wastewaters: A critical review. TrAC Trends Anal. Chem. 2020, 129, 115921. [Google Scholar] [CrossRef]
- Hubeny, J.; Harnisz, M.; Korzeniewska, E.; Buta, M.; Zielinski, W.; Rolbiecki, D.; Giebultowicz, J.; Nalecz-Jawecki, G.; Plaza, G. Industrialization as a source of heavy metals and antibiotics which can enhance the antibiotic resistance in wastewater, sewage sludge and river water. PLoS ONE 2021, 16, e0252691. [Google Scholar] [CrossRef]
- Rahube, T.O.; Marti, R.; Scott, A.; Tien, Y.C.; Murray, R.; Sabourin, L.; Zhang, Y.; Duenk, P.; Lapen, D.R.; Topp, E. Impact of fertilizing with raw or anaerobically digested sewage sludge on the abundance of antibiotic-resistant coliforms, antibiotic resistance genes, and pathogenic bacteria in soil and on vegetables at harvest. Appl. Environ. Microbiol. 2014, 80, 6898–6907. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zarfel, G.; Galler, H.; Feierl, G.; Haas, D.; Kittinger, C.; Leitner, E.; Grisold, A.J.; Mascher, F.; Posch, J.; Pertschy, B.; et al. Comparison of extended-spectrum-beta-lactamase (ESBL) carrying Escherichia coli from sewage sludge and human urinary tract infection. Environ. Pollut. 2013, 173, 192–199. [Google Scholar] [CrossRef]
- Sahlstrom, L.; Rehbinder, V.; Albihn, A.; Aspan, A.; Bengtsson, B. Vancomycin resistant enterococci (VRE) in Swedish sewage sludge. Acta Vet. Scand. 2009, 51, 24. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Markowicz, A.; Bondarczuk, K.; Wiekiera, A.; Sułowicz, S. Is sewage sludge a valuable fertilizer? A soil microbiome and resistome study under field conditions. J. Soils Sediments 2021, 21, 2882–2895. [Google Scholar] [CrossRef]
- Buta, M.; Hubeny, J.; Zielinski, W.; Harnisz, M.; Korzeniewska, E. Sewage sludge in agriculture—The effects of selected chemical pollutants and emerging genetic resistance determinants on the quality of soil and crops—A review. Ecotoxicol. Environ. Saf. 2021, 214, 112070. [Google Scholar] [CrossRef]
- You, R.; Margenat, A.; Lanzas, C.S.; Canameras, N.; Carazo, N.; Navarro-Martin, L.; Matamoros, V.; Bayona, J.M.; Diez, S. Dose effect of Zn and Cu in sludge-amended soils on vegetable uptake of trace elements, antibiotics, and antibiotic resistance genes: Human health implications. Environ. Res. 2020, 191, 109879. [Google Scholar] [CrossRef] [PubMed]
- Cucina, M.; Ricci, A.; Zadra, C.; Pezzolla, D.; Tacconi, C.; Sordi, S.; Gigliotti, G. Benefits and risks of long-term recycling of pharmaceutical sewage sludge on agricultural soil. Sci. Total Environ. 2019, 695, 133762. [Google Scholar] [CrossRef]
- Bondarczuk, K.; Markowicz, A.; Piotrowska-Seget, Z. The urgent need for risk assessment on the antibiotic resistance spread via sewage sludge land application. Environ. Int. 2016, 87, 49–55. [Google Scholar] [CrossRef]
- Collivignarelli, M.; Abbà, A.; Frattarola, A.; Carnevale Miino, M.; Padovani, S.; Katsoyiannis, I.; Torretta, V. Legislation for the Reuse of Biosolids on Agricultural Land in Europe: Overview. Sustainability 2019, 11, 6015. [Google Scholar] [CrossRef] [Green Version]
- Fijalkowski, K.; Rorat, A.; Grobelak, A.; Kacprzak, M.J. The presence of contaminations in sewage sludge—The current situation. J. Environ. Manage. 2017, 203, 1126–1136. [Google Scholar] [CrossRef]
- Rad, A.K.; Shamshiri, R.R.; Azarm, H.; Balasundram, S.K.; Sultan, M. Effects of the COVID-19 Pandemic on Food Security and Agriculture in Iran: A Survey. Sustainability 2021, 13, 10103. [Google Scholar] [CrossRef]
- Yan, W.; Bai, R.; Wang, S.; Tian, X.; Li, Y.; Wang, S.; Yang, F.; Xiao, Y.; Lu, X.; Zhao, F. Antibiotic resistance genes are increased by combined exposure to sulfamethoxazole and naproxen but relieved by low-salinity. Environ. Int. 2020, 139, 105742. [Google Scholar] [CrossRef] [PubMed]
- Tan, L.; Wang, F.; Liang, M.; Wang, X.; Das, R.; Mao, D.; Luo, Y. Antibiotic resistance genes attenuated with salt accumulation in saline soil. J. Hazard. Mater. 2019, 374, 35–42. [Google Scholar] [CrossRef] [PubMed]
- Sepulveda-Correa, A.; Daza-Giraldo, L.V.; Polania, J.; Arenas, N.E.; Munoz-Garcia, A.; Sandoval-Figueredo, A.V.; Vanegas, J. Genes associated with antibiotic tolerance and synthesis of antimicrobial compounds in a mangrove with contrasting salinities. Mar. Pollut. Bull. 2021, 171, 112740. [Google Scholar] [CrossRef]
- Liu, M.; Li, Q.; Sun, H.; Jia, S.; He, X.; Li, M.; Zhang, X.-X.; Ye, L. Impact of salinity on antibiotic resistance genes in wastewater treatment bioreactors. Chem. Eng. J. 2018, 338, 557–563. [Google Scholar] [CrossRef]
- Chen, Y.; Zhou, J.L.; Cheng, L.; Zheng, Y.Y.; Xu, J. Sediment and salinity effects on the bioaccumulation of sulfamethoxazole in zebrafish (Danio rerio). Chemosphere 2017, 180, 467–475. [Google Scholar] [CrossRef] [PubMed]
- Yang, C.-C.; Huang, C.-L.; Cheng, T.-C.; Lai, H.-T. Inhibitory effect of salinity on the photocatalytic degradation of three sulfonamide antibiotics. Int. Biodeterior. Biodegrad. 2015, 102, 116–125. [Google Scholar] [CrossRef]
- Nayiga, S.; Kayendeke, M.; Nabirye, C.; Willis, L.D.; Chandler, C.I.R.; Staedke, S.G. Use of antibiotics to treat humans and animals in Uganda: A cross-sectional survey of households and farmers in rural, urban and peri-urban settings. JAC Antimicrob. Resist. 2020, 2, dlaa082. [Google Scholar] [CrossRef]
- Madaras-Kelly, K. Optimizing antibiotic use in hospitals: The role of population-based antibiotic surveillance in limiting antibiotic resistance. Insights from the society of infectious diseases pharmacists. Pharmacotherapy 2003, 23, 1627–1633. [Google Scholar] [CrossRef]
- Kromker, V.; Leimbach, S. Mastitis Treatment-Reduction in antibiotic usage in dairy cows. Reprod. Domest. Anim. 2017, 52, 21–29. [Google Scholar] [CrossRef] [Green Version]
- Levy, S. Reduced antibiotic use in livestock: How Denmark tackled resistance. Environ. Health Perspect. 2014, 122, A160–A165. [Google Scholar] [CrossRef]
- Mevius, D.; Heederik, D. Reduction of antibiotic use in animals “let’s go Dutch”. J. Verbrauch. Lebensm. 2014, 9, 177–181. [Google Scholar] [CrossRef] [Green Version]
- Sneeringer, S.; Short, G.; MacLachlan, M.; Bowman, M. Impacts on Livestock Producers and Veterinarians of FDA Policies on Use of Medically Important Antibiotics in Food Animal Production. Appl. Econ. Perspect. Policy 2020, 42, 674–694. [Google Scholar] [CrossRef]
- Founou, L.L.; Founou, R.C.; Essack, S.Y. Antibiotic Resistance in the Food Chain: A Developing Country-Perspective. Front. Microbiol. 2016, 7, 1881. [Google Scholar] [CrossRef] [PubMed]
- Lhermie, G.; Grohn, Y.T.; Raboisson, D. Addressing Antimicrobial Resistance: An Overview of Priority Actions to Prevent Suboptimal Antimicrobial Use in Food-Animal Production. Front. Microbiol. 2016, 7, 2114. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Samanta, I.; Bandyopadhyay, S. The emergence of antimicrobial-resistant bacteria in livestock, poultry and agriculture. In Antimicrobial Resistance in Agriculture; Academic Press: Cambridge, MA, USA, 2020; p. 1927. [Google Scholar] [CrossRef]
- Mshana, S.E.; Sindato, C.; Matee, M.I.; Mboera, L.E.G. Antimicrobial Use and Resistance in Agriculture and Food Production Systems in Africa: A Systematic Review. Antibiotics 2021, 10, 976. [Google Scholar] [CrossRef]
- Robert, P.C. Precision agriculture: A challenge for crop nutrition management. Plant Soil 2002, 247, 143–149. [Google Scholar] [CrossRef]
- Bongiovanni, R.; Lowenberg-Deboer, J. Precision Agriculture and Sustainability. Precis. Agric. 2004, 5, 359–387. [Google Scholar] [CrossRef]
- Zarei, M.; Kaviani Rad, A. Covid-19, Challenges and Recommendations in Agriculture. J. Bot. Res. 2020, 2, 1841. [Google Scholar] [CrossRef]
- Du, L.; Liu, W. Occurrence, fate, and ecotoxicity of antibiotics in agro-ecosystems. A review. Agron. Sustain. Dev. 2011, 32, 309–327. [Google Scholar] [CrossRef] [Green Version]
- Sodhi, K.K.; Kumar, M.; Singh, D.K. Insight into the amoxicillin resistance, ecotoxicity, and remediation strategies. J. Water Process. Eng. 2021, 39, 101858. [Google Scholar] [CrossRef]
- Gu, S.; Kang, X.; Wang, L.; Lichtfouse, E.; Wang, C. Clay mineral adsorbents for heavy metal removal from wastewater: A review. Environ. Chem. Lett. 2018, 17, 629–654. [Google Scholar] [CrossRef]
- Han, H.; Rafiq, M.K.; Zhou, T.; Xu, R.; Masek, O.; Li, X. A critical review of clay-based composites with enhanced adsorption performance for metal and organic pollutants. J. Hazard. Mater. 2019, 369, 780–796. [Google Scholar] [CrossRef] [PubMed]
- Adeyemo, A.A.; Adeoye, I.O.; Bello, O.S. Adsorption of dyes using different types of clay: A review. Appl. Water Sci. 2015, 7, 543–568. [Google Scholar] [CrossRef] [Green Version]
- Mustapha, S.; Ndamitso, M.M.; Abdulkareem, A.S.; Tijani, J.O.; Mohammed, A.K.; Shuaib, D.T. Potential of using kaolin as a natural adsorbent for the removal of pollutants from tannery wastewater. Heliyon 2019, 5, e02923. [Google Scholar] [CrossRef]
- Yang, W.; Wu, Y.; Zhang, L.; Jiang, J.; Feng, L. Removal of five selected pharmaceuticals by coagulation in the presence of dissolved humic acids and kaolin. Desalination Water Treat. 2014, 54, 1134–1140. [Google Scholar] [CrossRef]
- Ugwu, I.M.; Igbokwe, O.A. Sorption of heavy metals on clay minerals and oxides: A review. Adv. Sorpt. Process Appl. 2019, 23, 80989. [Google Scholar] [CrossRef] [Green Version]
- Al-Mashhadany, D.A. Detection of antibiotic residues among raw beef in Erbil City (Iraq) and impact of temperature on antibiotic remains. Ital. J. Food Saf. 2019, 8, 7897. [Google Scholar] [CrossRef] [Green Version]
- Zhang, Y.; Liu, H.; Dai, X.; Cai, C.; Wang, J.; Wang, M.; Shen, Y.; Wang, P. Impact of application of heat-activated persulfate oxidation treated erythromycin fermentation residue as a soil amendment: Soil chemical properties and antibiotic resistance. Sci. Total Environ. 2020, 736, 139668. [Google Scholar] [CrossRef]
- Xu, R.; Zhang, Y.; Xiong, W.; Sun, W.; Fan, Q.; Zhaohui, Y. Metagenomic approach reveals the fate of antibiotic resistance genes in a temperature-raising anaerobic digester treating municipal sewage sludge. J. Clean. Prod. 2020, 277, 123504. [Google Scholar] [CrossRef]
- Sun, C.; Li, W.; Chen, Z.; Qin, W.; Wen, X. Responses of antibiotics, antibiotic resistance genes, and mobile genetic elements in sewage sludge to thermal hydrolysis pre-treatment and various anaerobic digestion conditions. Environ. Int. 2019, 133, 105156. [Google Scholar] [CrossRef] [PubMed]
- Liao, H.; Lu, X.; Rensing, C.; Friman, V.P.; Geisen, S.; Chen, Z.; Yu, Z.; Wei, Z.; Zhou, S.; Zhu, Y. Hyperthermophilic Composting Accelerates the Removal of Antibiotic Resistance Genes and Mobile Genetic Elements in Sewage Sludge. Environ. Sci. Technol. 2018, 52, 266–276. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Deng, W.; Zhang, A.; Chen, S.; He, X.; Jin, L.; Yu, X.; Yang, S.; Li, B.; Fan, L.; Ji, L.; et al. Heavy metals, antibiotics and nutrients affect the bacterial community and resistance genes in chicken manure composting and fertilized soil. J. Environ. Manag. 2020, 257, 109980. [Google Scholar] [CrossRef]
- Gou, M.; Hu, H.W.; Zhang, Y.J.; Wang, J.T.; Hayden, H.; Tang, Y.Q.; He, J.Z. Aerobic composting reduces antibiotic resistance genes in cattle manure and the resistome dissemination in agricultural soils. Sci. Total Environ. 2018, 612, 1300–1310. [Google Scholar] [CrossRef]
- Youngquist, C.P.; Mitchell, S.M.; Cogger, C.G. Fate of Antibiotics and Antibiotic Resistance during Digestion and Composting: A Review. J. Environ. Qual. 2016, 45, 537–545. [Google Scholar] [CrossRef]
- Keenum, I.; Williams, R.K.; Ray, P.; Garner, E.D.; Knowlton, K.F.; Pruden, A. Combined effects of composting and antibiotic administration on cattle manure-borne antibiotic resistance genes. Microbiome 2021, 9, 81. [Google Scholar] [CrossRef]
- Sardar, M.F.; Zhu, C.; Geng, B.; Ahmad, H.R.; Song, T.; Li, H. The fate of antibiotic resistance genes in cow manure composting: Shaped by temperature-controlled composting stages. Bioresour. Technol. 2021, 320, 124403. [Google Scholar] [CrossRef] [PubMed]
- Gao, M.; Qiu, T.; Sun, Y.; Wang, X. The abundance and diversity of antibiotic resistance genes in the atmospheric environment of composting plants. Environ. Int. 2018, 116, 229–238. [Google Scholar] [CrossRef] [PubMed]
- Guerin, T.F. The differential removal of aged polycyclic aromatic hydrocarbons from soil during bioremediation. Environ. Sci. Pollut. Res. 2000, 7, 19–26. [Google Scholar] [CrossRef]
- Imam, A.; Suman, S.K.; Ghosh, D.; Kanaujia, P.K. Analytical approaches used in monitoring the bioremediation of hydrocarbons in petroleum-contaminated soil and sludge. TrAC Trends Anal. Chem. 2019, 118, 50–64. [Google Scholar] [CrossRef]
- Stroud, J.L.; Paton, G.I.; Semple, K.T. Microbe-aliphatic hydrocarbon interactions in soil: Implications for biodegradation and bioremediation. J. Appl. Microbiol. 2007, 102, 1239–1253. [Google Scholar] [CrossRef]
- Diplock, E.E.; Mardlin, D.P.; Killham, K.S.; Paton, G.I. Predicting bioremediation of hydrocarbons: Laboratory to field scale. Environ. Pollut. 2009, 157, 1831–1840. [Google Scholar] [CrossRef]
- Ghazali, F.M.; Rahman, R.N.Z.A.; Salleh, A.B.; Basri, M. Biodegradation of hydrocarbons in soil by microbial consortium. Int. Biodeterior. Biodegrad. 2004, 54, 61–67. [Google Scholar] [CrossRef]
- Gargouri, B.; Karray, F.; Mhiri, N.; Aloui, F.; Sayadi, S. Bioremediation of petroleum hydrocarbons-contaminated soil by bacterial consortium isolated from an industrial wastewater treatment plant. J. Chem. Technol. Biotechnol. 2014, 89, 978–987. [Google Scholar] [CrossRef]
- Guarino, C.; Spada, V.; Sciarrillo, R. Assessment of three approaches of bioremediation (Natural Attenuation, Landfarming and Bioagumentation—Assistited Landfarming) for a petroleum hydrocarbons contaminated soil. Chemosphere 2017, 170, 10–16. [Google Scholar] [CrossRef] [PubMed]
- Muyzer, G.; Stams, A.J. The ecology and biotechnology of sulphate-reducing bacteria. Nat. Rev. Microbiol. 2008, 6, 441–454. [Google Scholar] [CrossRef] [PubMed]
- Castro, H.F.; Williams, N.H.; Ogram, A. Phylogeny of sulfate-reducing bacteria1. FEMS Microbiol. Ecol. 2000, 31, 1–9. [Google Scholar] [CrossRef]
- Xu, Y.N.; Chen, Y. Advances in heavy metal removal by sulfate-reducing bacteria. Water Sci. Technol. 2020, 81, 1797–1827. [Google Scholar] [CrossRef]
- Jong, T.; Parry, D.L. Removal of sulfate and heavy metals by sulfate reducing bacteria in short-term bench scale upflow anaerobic packed bed reactor runs. Water Res. 2003, 37, 3379–3389. [Google Scholar] [CrossRef]
- Zhao, Q.; Li, X.; Xiao, S.; Peng, W.; Fan, W. Integrated remediation of sulfate reducing bacteria and nano zero valent iron on cadmium contaminated sediments. J. Hazard. Mater. 2021, 406, 124680. [Google Scholar] [CrossRef]
- Barton, L.L.; Fauque, G.D. Biochemistry, Physiology and Biotechnology of Sulfate-Reducing Bacteria. Adv. Appl. Microbiol. 2009, 68, 41–98. [Google Scholar] [CrossRef] [PubMed]
- Fang, H.; Oberoi, A.S.; He, Z.; Khanal, S.K.; Lu, H. Ciprofloxacin-degrading Paraclostridium sp. isolated from sulfate-reducing bacteria-enriched sludge: Optimization and mechanism. Water Res. 2021, 191, 116808. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Song, S.; Jia, Y.; Wu, D.; Lu, H. Stress-responses of activated sludge and anaerobic sulfate-reducing bacteria sludge under long-term ciprofloxacin exposure. Water Res. 2019, 164, 114964. [Google Scholar] [CrossRef]
- Zhang, H.; Jia, Y.; Khanal, S.K.; Lu, H.; Fang, H.; Zhao, Q. Understanding the Role of Extracellular Polymeric Substances on Ciprofloxacin Adsorption in Aerobic Sludge, Anaerobic Sludge, and Sulfate-Reducing Bacteria Sludge Systems. Environ. Sci. Technol. 2018, 52, 6476–6486. [Google Scholar] [CrossRef] [PubMed]
- Jia, Y.; Khanal, S.K.; Shu, H.; Zhang, H.; Chen, G.H.; Lu, H. Ciprofloxacin degradation in anaerobic sulfate-reducing bacteria (SRB) sludge system: Mechanism and pathways. Water Res. 2018, 136, 64–74. [Google Scholar] [CrossRef]
- Zhang, X.; Zhu, R.; Li, W.; Ma, J.; Lin, H. Genomic insights into the antibiotic resistance pattern of the tetracycline-degrading bacterium, Arthrobacter nicotianae OTC-16. Sci. Rep. 2021, 11, 15638. [Google Scholar] [CrossRef]
- Maki, T.; Hasegawa, H.; Kitami, H.; Fumoto, K.; Munekage, Y.; Ueda, K. Bacterial degradation of antibiotic residues in marine fish farm sediments of Uranouchi Bay and phylogenetic analysis of antibiotic-degrading bacteria using 16S rDNA sequences. Fish. Sci. 2006, 72, 811–820. [Google Scholar] [CrossRef]
- Hirth, N.; Topp, E.; Dörfler, U.; Stupperich, E.; Munch, J.C.; Schroll, R. An effective bioremediation approach for enhanced microbial degradation of the veterinary antibiotic sulfamethazine in an agricultural soil. Chem. Biol. Technol. Agric. 2016, 3, 1–11. [Google Scholar] [CrossRef] [Green Version]
- Mojiri, A.; Baharlooeian, M.; Zahed, M.A. The Potential of Chaetoceros muelleri in Bioremediation of Antibiotics: Performance and Optimization. Int. J. Environ. Res. Public Health 2021, 18, 977. [Google Scholar] [CrossRef]
- Wu, Y.; Feng, P.; Li, R.; Chen, X.; Li, X.; Sumpradit, T.; Liu, P. Progress in microbial remediation of antibiotic-residue contaminated environment. Sheng Wu Gong Cheng Xue Bao 2019, 35, 2133–2150. [Google Scholar] [CrossRef]
- Xue, W.; Zhou, Q.; Li, F. Bacterial community changes and antibiotic resistance gene quantification in microbial electrolysis cells during long-term sulfamethoxazole treatment. Bioresour. Technol. 2019, 294, 122170. [Google Scholar] [CrossRef] [PubMed]
- Hua, T.; Li, S.; Li, F.; Ondon, B.S.; Liu, Y.; Wang, H. Degradation performance and microbial community analysis of microbial electrolysis cells for erythromycin wastewater treatment. Biochem. Eng. J. 2019, 146, 1–9. [Google Scholar] [CrossRef]
- Zhang, X.; Li, R. Variation and distribution of antibiotic resistance genes and their potential hosts in microbial electrolysis cells treating sewage sludge. Bioresour. Technol. 2020, 315, 123838. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Li, R. Electrodes bioaugmentation promotes the removal of antibiotics from concentrated sludge in microbial electrolysis cells. Sci. Total Environ. 2020, 715, 136997. [Google Scholar] [CrossRef] [PubMed]
- Yan, W.; Xiao, Y.; Yan, W.; Ding, R.; Wang, S.; Zhao, F. The effect of bioelectrochemical systems on antibiotics removal and antibiotic resistance genes: A review. Chem. Eng. J. 2019, 358, 1421–1437. [Google Scholar] [CrossRef]
- Dong, X.; Rao, D.; Tian, L.; Wang, Q.; Yang, K. A slurry microcosm study on the interaction between antibiotics and soil bacterial community. Heliyon 2020, 6, e03348. [Google Scholar] [CrossRef]
- Naqvi, S.A.R.; Nadeem, S.; Komal, S.; Naqvi, S.A.A.; Mubarik, M.S.; Qureshi, S.Y.; Ahmad, S.; Abbas, A.; Zahid, M.; Raza, S.S. Antioxidants: Natural Antibiotics. In Antioxidants; IntechOpen: London, UK, 2019. [Google Scholar] [CrossRef] [Green Version]
- Abd El-Ghany, W.A. Paraprobiotics and postbiotics: Contemporary and promising natural antibiotics alternatives and their applications in the poultry field. Open Vet. J. 2020, 10, 323–330. [Google Scholar] [CrossRef]
- Gupta, R.; Sharma, S. Herbal antibiotics: A Review. Bull. Env. Pharmacol. Life Sci. 2020, 9, 136–142. [Google Scholar]
- Ionescu, M.I. Are Herbal Products an Alternative to Antibiotics. In Bacterial Pathogenesis and Antibacterial Control; IntechOpen: London, UK, 2018. [Google Scholar] [CrossRef] [Green Version]
- Fit, N.; Gheorghe, R.; Rapuntean, S.; Flore, C.; Nadas, G. Antibacterial Effect of Essential Vegetal Extracts on Staphylococcus aureus Compared to Antibiotics. Not. Bot. Horti Agrobot. Cluj Napoca 2009, 37, 117–123. [Google Scholar] [CrossRef]
- Awan, U.; Andleeb, D.S.; Kiyani, A.; Zafar, A.; Shafique, I.; Riaz, N.; Azhar, M.T.; Uddin, H. Antibacterial screening of traditional herbal plants and standard antibiotics against some human bacterial pathogens. Pak. J. Pharm. Sci. 2013, 26, 1109–1116. [Google Scholar]
- Saquib, S.A.; AlQahtani, N.A.; Ahmad, I.; Kader, M.A.; Al Shahrani, S.S.; Asiri, E.A. Evaluation and Comparison of Antibacterial Efficacy of Herbal Extracts in Combination with Antibiotics on Periodontal pathobionts: An in vitro Microbiological Study. Antibiotics 2019, 8, 89. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nweze, E.I.; Eze, E.E. Justification for the use of Ocimum gratissimum L in herbal medicine and its interaction with disc antibiotics. BMC Complement. Altern. Med. 2009, 9, 37. [Google Scholar] [CrossRef] [Green Version]
- Sharifi-Rad, J. Herbal Antibiotics: Moving back into the mainstream as an alternative for “Superbugs”. Cell Mol. Biol. 2016, 62, 1–2. [Google Scholar] [CrossRef] [PubMed]
- Martin, K.W.; Ernst, E. Herbal medicines for treatment of bacterial infections: A review of controlled clinical trials. Antimicrob. Chemother. 2003, 51, 241–246. [Google Scholar] [CrossRef] [Green Version]
- Zasloff, M. Antimicrobial peptides of multicellular organisms. Nature 2002, 415, 389–395. [Google Scholar] [CrossRef]
- Koczulla, R.; Bals, R. Antimicrobial Peptides: Current Status and Therapeutic Potential. Drugs 2003, 63, 389–406. [Google Scholar] [CrossRef] [PubMed]
- Reddy, K.V.; Yedery, R.D.; Aranha, C. Antimicrobial peptides: Premises and promises. Int. J. Antimicrob. Agents 2004, 24, 536–547. [Google Scholar] [CrossRef]
- Sang, Y.; Blecha, F. Antimicrobial peptides and bacteriocins: Alternatives to traditional antibiotics. Anim. Health Res. Rev. 2008, 9, 227–235. [Google Scholar] [CrossRef] [Green Version]
- Leontiadou, H.; Mark, A.E.; Marrink, S.J. Antimicrobial Peptides in Action. J. Am. Chem. Soc. 2006, 128, 12156–12161. [Google Scholar] [CrossRef] [Green Version]
- Fjell, C.D.; Hiss, J.A.; Hancock, R.E.; Schneider, G. Designing antimicrobial peptides: Form follows function. Nat. Rev. Drug Discov. 2011, 11, 37–51. [Google Scholar] [CrossRef]
- Malmsten, M. Antimicrobial peptides. Ups. J. Med. Sci. 2014, 119, 199–204. [Google Scholar] [CrossRef]
- Wang, G.; Mishra, B.; Lau, K.; Lushnikova, T.; Golla, R.; Wang, X. Antimicrobial peptides in 2014. Pharmaceuticals 2015, 8, 123–150. [Google Scholar] [CrossRef]
- Bahar, A.A.; Ren, D. Antimicrobial peptides. Pharmaceuticals 2013, 6, 1543–1575. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kosciuczuk, E.M.; Lisowski, P.; Jarczak, J.; Strzalkowska, N.; Jozwik, A.; Horbanczuk, J.; Krzyzewski, J.; Zwierzchowski, L.; Bagnicka, E. Cathelicidins: Family of antimicrobial peptides. A review. Mol. Biol. Rep. 2012, 39, 10957–10970. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tamhankar, A.J.; Stalsby Lundborg, C. Antimicrobials and Antimicrobial Resistance in the Environment and Its Remediation: A Global One Health Perspective. Int. J. Environ. Res. Public Health 2019, 16, 4614. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rad, A.; Zarei, M.; Pourghasemi, H.; Tiefenbacher, J. The COVID-19 crisis and its consequences for global warming and climate change. In Computers in Earth and Environmental Sciences; Elsevier: Amsterdam, The Netherlands, 2022; pp. 377–385. [Google Scholar] [CrossRef]
- Manyi-Loh, C.; Mamphweli, S.; Meyer, E.; Okoh, A. Antibiotic Use in Agriculture and Its Consequential Resistance in Environmental Sources: Potential Public Health Implications. Molecules 2018, 23, 795. [Google Scholar] [CrossRef] [Green Version]
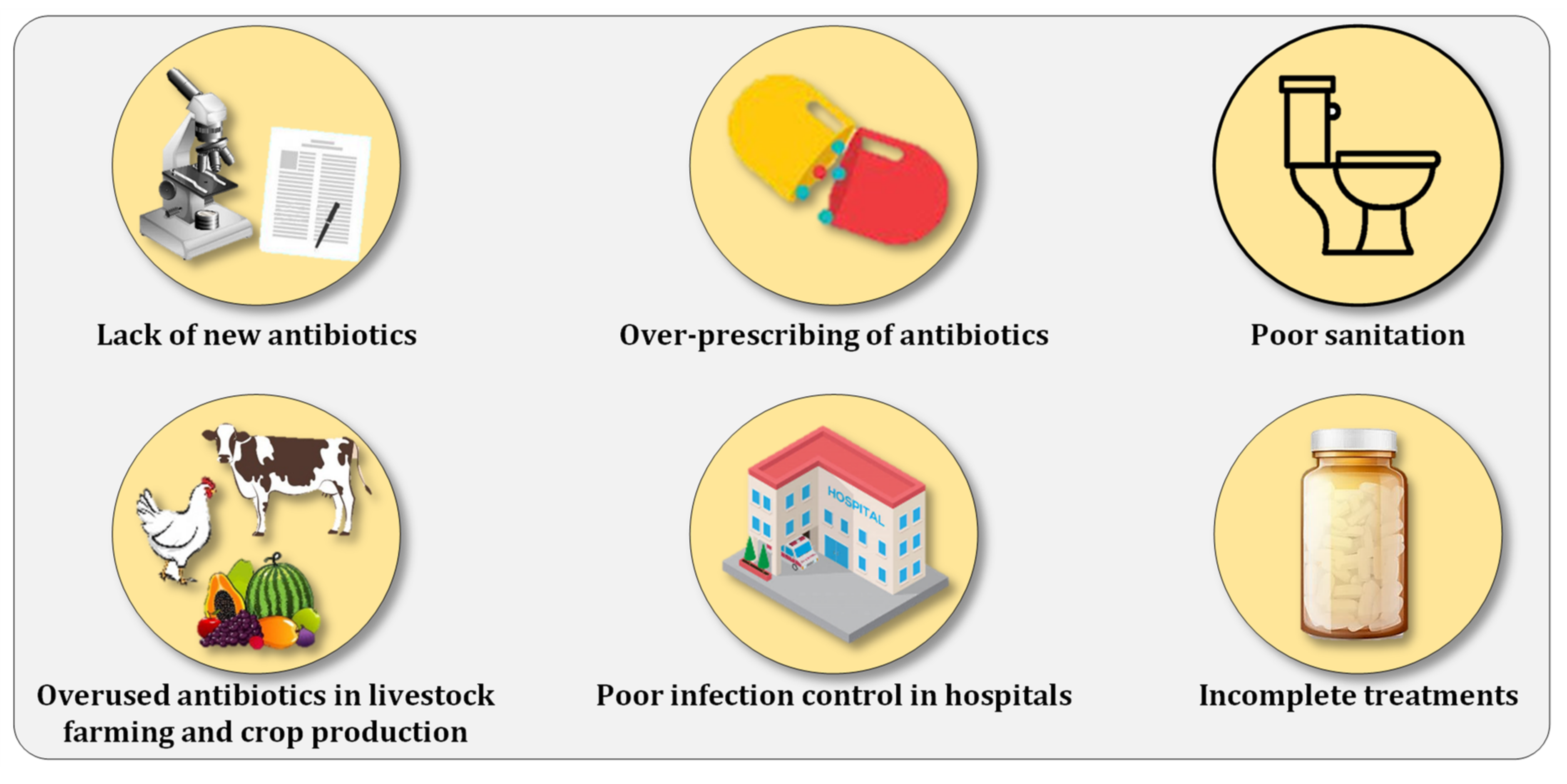
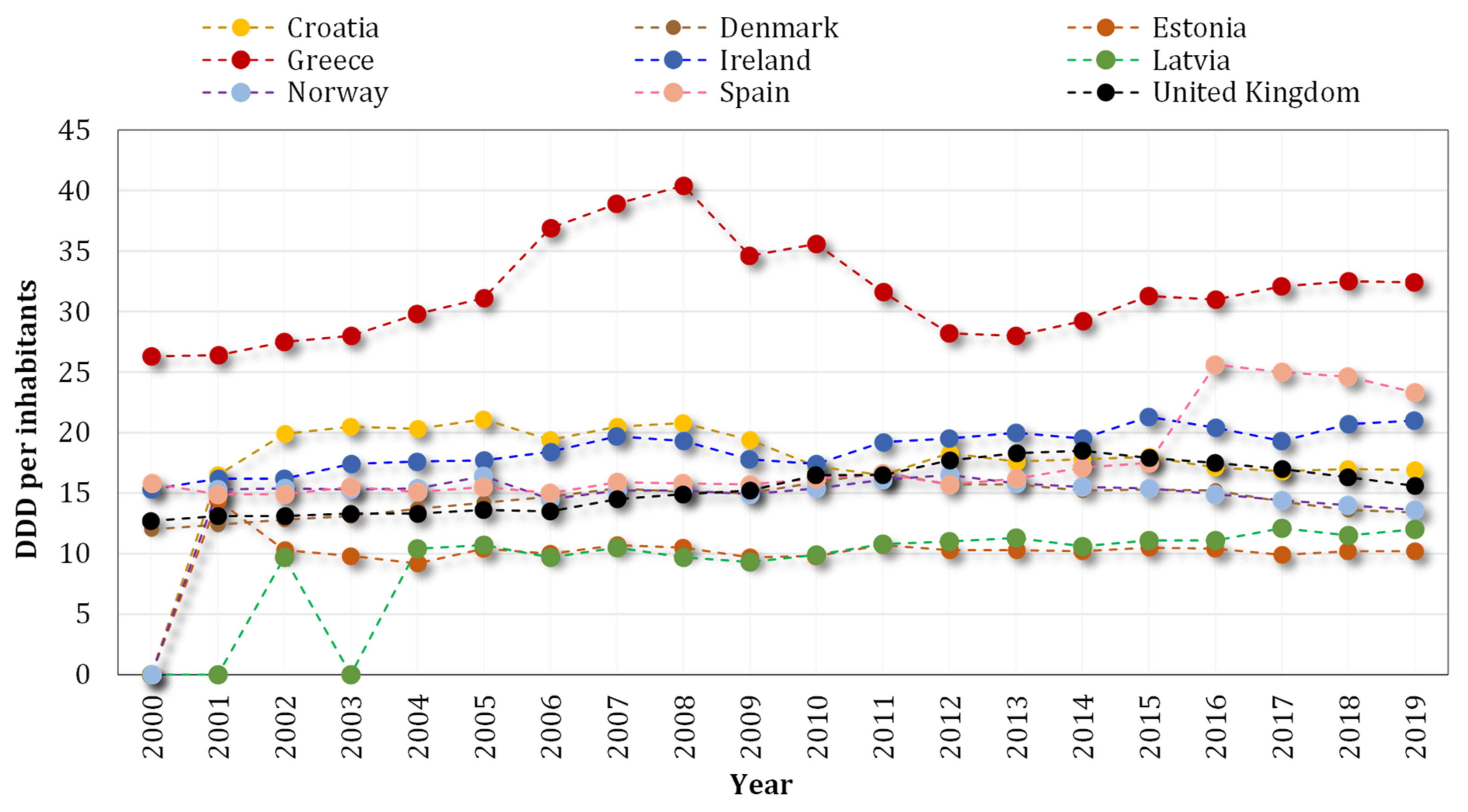
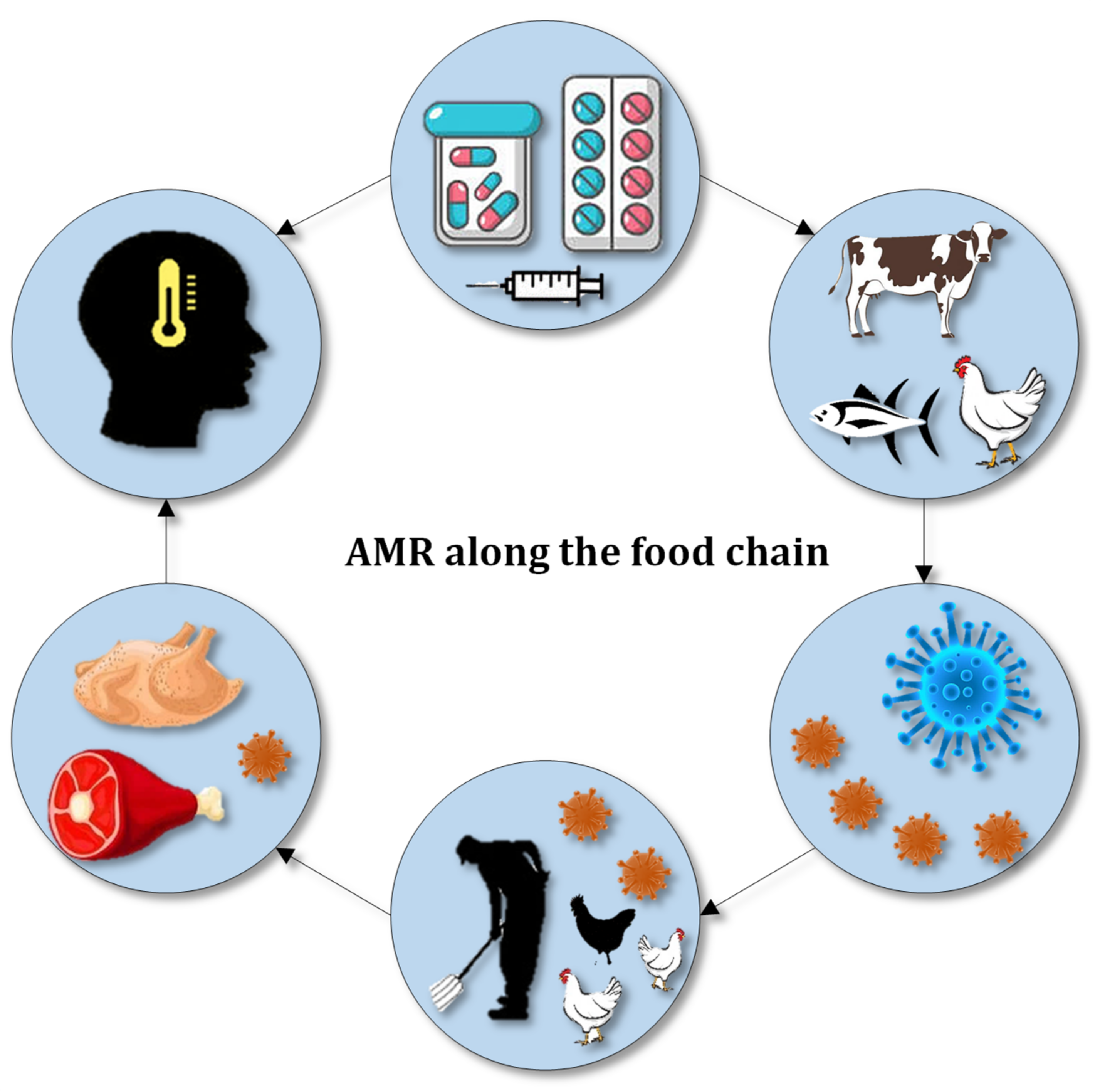
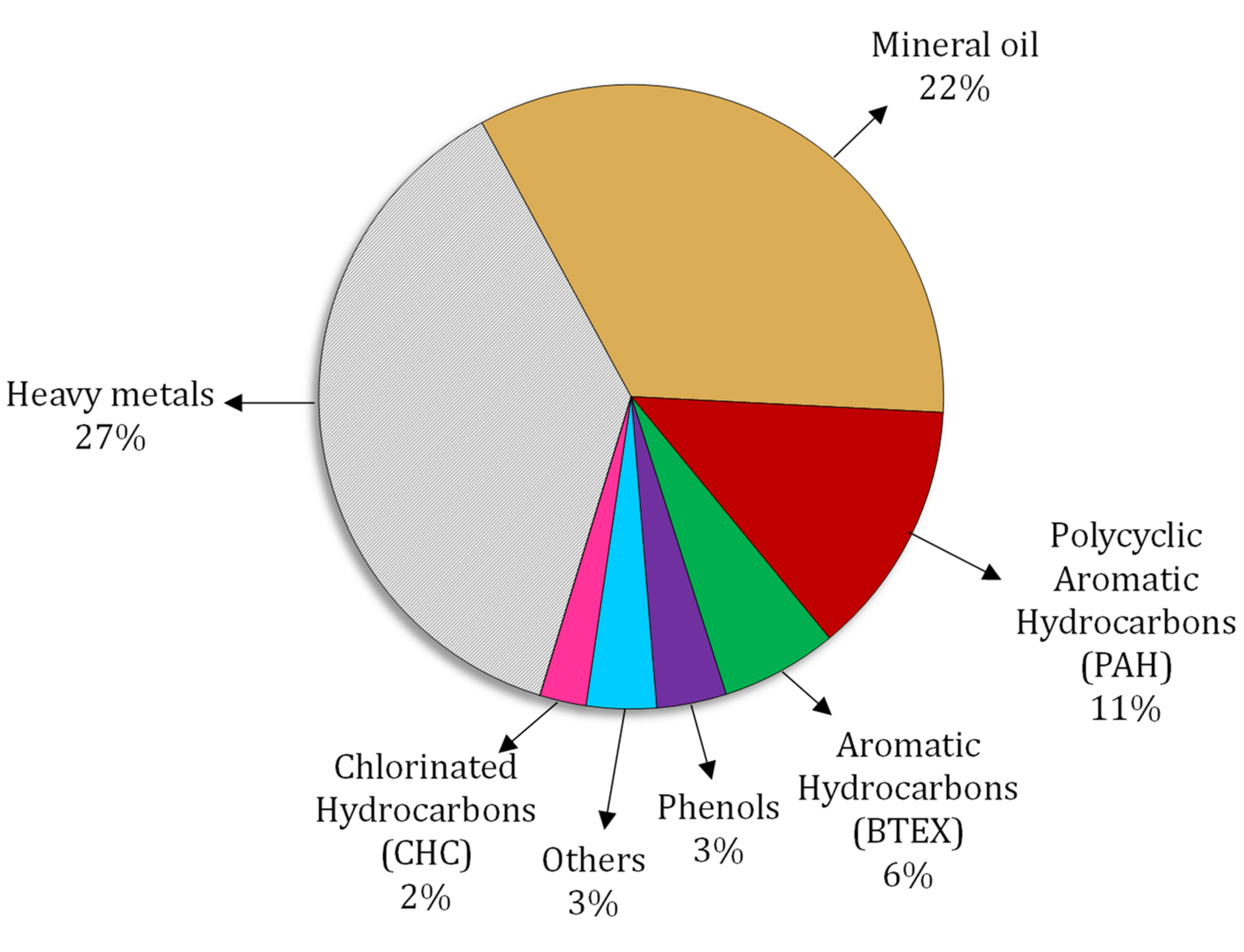
| Antibiotic | Animal | Resistant Bacteria | Result | Ref |
|---|---|---|---|---|
| Cefalotin, streptomycin, and sulfamethoxazole | Cattle and sheep | Escherichia coli | Most isolates were moderately resistant to antibiotics. | [69] |
| Streptomycin, gentamycin, tetracycline, and trimethoprim | Sheep, goat, camel | Acinetobacter baumannii | Antibiotic resistance was observed in more than half of the strains isolated from sheep samples. | [70] |
| Norfloxacin and Doxycycline | Fowl | Escherichia coli | An increasing resistance rate of E. coli toward norfloxacin in chickens was detected. | [71] |
| Ampicillin, tetracycline, and sulfamethoxazole | Broiler | Escherichia coli | Isolated strains were resistant to antibiotics. | [72] |
| Lincomycin, erythromycin, ciprofloxacin, and tetracycline | Wild bird | Enterococcus faecium, Enterococcus hirae, Enterococcus durans, Enterococcus casseliflavus | The highest resistance was recorded for lincomycin. | [73] |
| Ampicillin, tetracycline, and nitrofurantoin | Fish | Gram-negative bacteria | Maximum resistance was recognized for ampicillin and tetracycline. | [74] |
| Ampicillin, tetracycline, and chloramphenicol | Hen eggshells | Salmonella enterica | Most isolates were resistant to ampicillin. | [75] |
| Tetracycline | Cattle | Gut microbiomes | Resistance to tetracycline was highly prevalent in cattle. | [76] |
| Tetracycline and clindamycin | Swine | Staphylococcusaureus | High antibiotic resistance was observed for tetracycline or clindamycin. | [77] |
| Ciprofloxacin, nitrofurantoin, trimethoprim, and cefalotin | Sheep | Escherichia coli | The highest AMR was recorded toward ciprofloxacin (69.4%). | [78] |
| Ampicillin and tetracycline | Catfish (Clarias gariepinus) | Klebsiella pneumoniae | All coliform bacteria were resistant to antibiotics. | [79] |
| Strain | Heavy Metal | Antibiotic | Location | Result | Ref |
|---|---|---|---|---|---|
| Pseudomonas putida, Staphylococcus epidermidis, Serratia ficaria, and Bacillus anthracis | Cu, Cd, Cr, Ag, and Hg | Amoxicillin, gentamycin, vancomycin, tetracycline, and ciprofloxacin | Marchica, Morocco | Simultaneous resistance to heavy metals and antibiotics | [139] |
| Aeromonas hydrophila | Cu, Co, Zn, and Hg | Sulfamide, oxytetracycline, and trimethoprim | Tunisia | Relationship between antibiotic resistance and resistance to heavy metals | [140] |
| Pseudomonas aeruginosa, Actinomyces turicensis, and Micrococcus sp. | Hg, Cd, Co, Ni, and Cr | Chloramphenicol, streptomycin, erythromycin, and metronidazole | Nigeria | 22 out of 270 strains of isolated bacteria had simultaneous resistance to antibiotics and heavy metals | [141] |
| Staphylococcus aureus, Alcaligenes sp., Bacillus sp. and Klebsiella sp. | Pb, Cr, Zn, and Cd | Ampicillin, cefalotin, gentamycin, and doxycyclin | Algeria | Eighty-five percent of heavy metal isolates were similarly resistant to several antibiotics | [142] |
| Pseudomonas fluorescens | Pb, Cu, Cr, Zn, and Hg | Amoxicillin, cefradine, norfloxacin, and tetracycline | Guangzhou, China | Correlation between the antibiotic type and the concentration of heavy metals | [143] |
| 138 halophilic bacterial isolates | Cd, Zn, Pb, Cu, and Co | Cefalexin, vancomycin, cefalotin, and ampicillin | Red Sea, Egypt | Simultaneous resistance to heavy metals and antibiotics | [144] |
| Escherichia coli | Ni, Cr, Cu, Pb, and Cd | - | Yamuna, India | A higher level of metal resistance was recognized by increasing the average concentration of metals | [145] |
| Enterococci faecalis | Zn, Ni, Cu, and Co | Penicillin, ampicillin, ciprofloxacin, and sulfamethoxazole | Iran | Simultaneous resistance to antibiotics and metals in the most strains | [146] |
| Staphylococcus aureus | Pb, Cu, Zn, Cr, Cd, and Ni | Tetracycline, ceftazidime, ciprofloxacin, and vancomycin | Nigeria | Multiple resistance to antibiotics and heavy metals in the strains | [147] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kaviani Rad, A.; Astaykina, A.; Streletskii, R.; Afsharyzad, Y.; Etesami, H.; Zarei, M.; Balasundram, S.K. An Overview of Antibiotic Resistance and Abiotic Stresses Affecting Antimicrobial Resistance in Agricultural Soils. Int. J. Environ. Res. Public Health 2022, 19, 4666. https://doi.org/10.3390/ijerph19084666
Kaviani Rad A, Astaykina A, Streletskii R, Afsharyzad Y, Etesami H, Zarei M, Balasundram SK. An Overview of Antibiotic Resistance and Abiotic Stresses Affecting Antimicrobial Resistance in Agricultural Soils. International Journal of Environmental Research and Public Health. 2022; 19(8):4666. https://doi.org/10.3390/ijerph19084666
Chicago/Turabian StyleKaviani Rad, Abdullah, Angelika Astaykina, Rostislav Streletskii, Yeganeh Afsharyzad, Hassan Etesami, Mehdi Zarei, and Siva K. Balasundram. 2022. "An Overview of Antibiotic Resistance and Abiotic Stresses Affecting Antimicrobial Resistance in Agricultural Soils" International Journal of Environmental Research and Public Health 19, no. 8: 4666. https://doi.org/10.3390/ijerph19084666
APA StyleKaviani Rad, A., Astaykina, A., Streletskii, R., Afsharyzad, Y., Etesami, H., Zarei, M., & Balasundram, S. K. (2022). An Overview of Antibiotic Resistance and Abiotic Stresses Affecting Antimicrobial Resistance in Agricultural Soils. International Journal of Environmental Research and Public Health, 19(8), 4666. https://doi.org/10.3390/ijerph19084666









