Dynamics of Microbial Communities during the Removal of Copper and Zinc in a Sulfate-Reducing Bioreactor with a Limestone Pre-Column System
Abstract
:1. Introduction
2. Materials and Methods
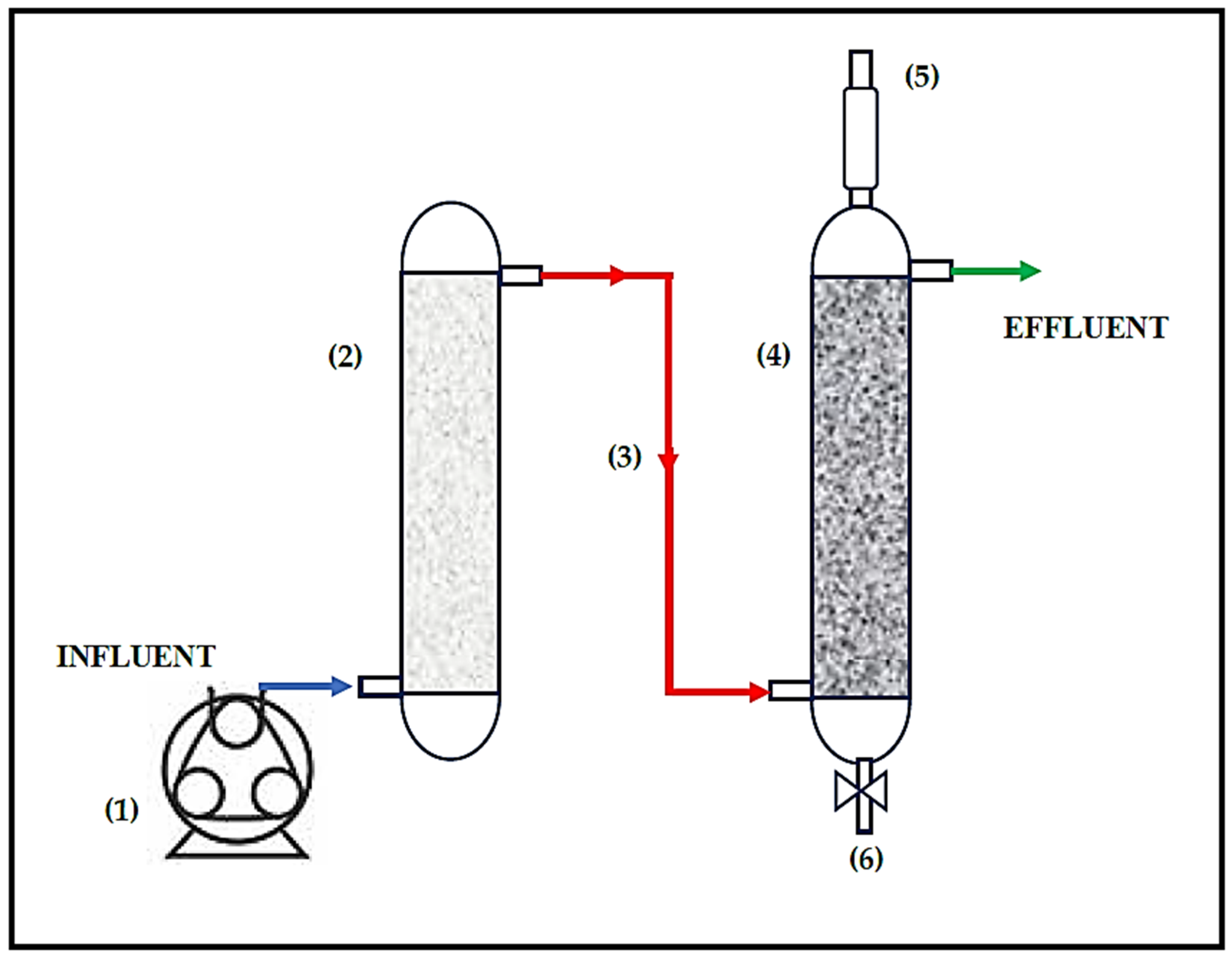
2.1. Configuration and Operation of the Treatment System
2.2. Analytical Methods
2.3. Sludge Sample Collection
2.4. DNA Extraction
2.5. rRNA Analysis
2.6. Microbial Diversity and Statistical Analysis
3. Results
3.1. Performance of the Treatment System
3.2. Microbial Community Analysis
4. Discussion
4.1. Bioreactor Performance: Sulfate Removal, Sulfide Production and Metals Removal
4.2. Microbial Diversity and Community Structure
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Simate, G.S.; Ndlovu, S. Acid Mine Drainage: Challenges and Opportunities. J. Environ. Chem. Eng. 2014, 2, 1785–1803. [Google Scholar] [CrossRef]
- Nieto, J.M.; Sarmiento, A.M.; Olías, M.; Canovas, C.R.; Riba, I.; Kalman, J.; Delvalls, T.A. Acid Mine Drainage Pollution in the Tinto and Odiel Rivers (Iberian Pyrite Belt, SW Spain) and Bioavailability of the Transported Metals to the Huelva Estuary. Environ. Int. 2007, 33, 445–455. [Google Scholar] [CrossRef]
- Zipper, C.; Skounsen, J. Acid Mine Drainage, Rock Drainage, and Acid Sulfate Soils; Jacobs, J.A., Lehr, J.H., Testa, S.M., Eds.; John Wiley & Sons, Inc.: Hoboken, NJ, USA, 2014; ISBN 978-1-118-74919-7. [Google Scholar]
- Dar, S.A.; Yao, L.; van Dongen, U.; Kuenen, J.G.; Muyzer, G. Analysis of Diversity and Activity of Sulfate-Reducing Bacterial Communities in Sulfidogenic Bioreactors Using 16S RRNA and DsrB Genes as Molecular Markers. Appl. Environ. Microbiol. 2007, 73, 594–604. [Google Scholar] [CrossRef] [Green Version]
- Muyzer, G.; Stams, A.J.M. The Ecology and Biotechnology of Sulphate-Reducing Bacteria. Nat. Rev. Microbiol. 2008, 6, 441–454. [Google Scholar] [CrossRef]
- Kaksonen, A.H.; Puhakka, J.A. Sulfate Reduction Based Bioprocesses for the Treatment of Acid Mine Drainage and the Recovery of Metals. Eng. Life Sci. 2007, 7, 541–564. [Google Scholar] [CrossRef]
- Freeman, S.A.; Sierra-Alvarez, R.; Altinbas, M.; Hollingsworth, J.; Stams, A.J.M.; Smidt, H. Molecular Characterization of Mesophilic and Thermophilic Sulfate Reducing Microbial Communities in Expanded Granular Sludge Bed (EGSB) Reactors. Biodegradation 2008, 19, 161–177. [Google Scholar] [CrossRef]
- Hulshoff, L.W.; Lens, P.N.L.; Weijma, J.; Stams, A.J.M. New Developments in Reactor and Process Technology for Sulfate Reduction. Water Sci. Technol. 2001, 44, 67–76. [Google Scholar] [CrossRef]
- Xu, Y.-N.; Chen, Y. Advances in Heavy Metal Removal by Sulfate-Reducing Bacteria. Water Sci. Technol. 2020, 81, 1797–1827. [Google Scholar] [CrossRef]
- Hui, C.; Guo, Y.; Liu, L.; Yi, J. Recent Advances in Bacterial Biosensing and Bioremediation of Cadmium Pollution: A Mini-Review. World J. Microbiol. Biotechnol. 2022, 38, 9. [Google Scholar] [CrossRef]
- McCauley, C. Assessment of Passive Treatment and Biochemical Reactors for Ameliorating Acid Mine Drainage at Stockton Coal Mine. Ph.D Thesis, Univesity of Canterbury, Christchurch, New Zealand, 2011; pp. 1–366. [Google Scholar]
- Angai, J.U.; Ptacek, C.J.; Pakostova, E.; Bain, J.G.; Verbuyst, B.R.; Blowes, D.W. Removal of Arsenic and Metals from Groundwater Impacted by Mine Waste Using Zero-Valent Iron and Organic Carbon: Laboratory Column Experiments. J. Hazard. Mater. 2022, 424, 127295. [Google Scholar] [CrossRef]
- Jørgensen, B.B.; Findlay, A.J.; Pellerin, A. The Biogeochemical Sulfur Cycle of Marine Sediments. Front. Microbiol. 2019, 10, 849. [Google Scholar] [CrossRef] [PubMed]
- Wu, J.; Niu, Q.; Li, L.; Hu, Y.; Mribet, C.; Hojo, T.; Li, Y.-Y. A Gradual Change between Methanogenesis and Sulfidogenesis during a Long-Term UASB Treatment of Sulfate-Rich Chemical Wastewater. Sci. Total Environ. 2018, 636, 168–176. [Google Scholar] [CrossRef] [PubMed]
- St. James, A.R.; Richardson, R.E. Ecogenomics Reveals Community Interactions in a Long-Term Methanogenic Bioreactor and a Rapid Switch to Sulfate-Reducing Conditions. FEMS Microbiol. Ecol. 2020, 96, fiaa050. [Google Scholar] [CrossRef]
- Yu, N.; Guo, B.; Liu, Y. Shaping Biofilm Microbiomes by Changing GAC Location during Wastewater Anaerobic Digestion. Sci. Total Environ. 2021, 780, 146488. [Google Scholar] [CrossRef]
- Schmidtova, J. Microbial Processes and Carbon Utilization in High Sulfate Waters and Sediments. Ph.D. Thesis, University of British Columbia, Kelowna, BC, Canada, 2010; pp. 1–174. [Google Scholar]
- Satoh, H.; Oshima, K.; Suda, W.; Ranasinghe, P.; Li, N.; Gunawardana, E.G.W.; Hattori, M.; Mino, T. Bacterial Population Dynamics in a Laboratory Activated Sludge Reactor Monitored by Pyrosequencing of 16S RRNA. Microbes Environ. 2013, 28, 65–70. [Google Scholar] [CrossRef] [Green Version]
- Dodamani, S.; Hattiholi, A.; Kurjogi, M. Microbial Communities: An Effective Tool for Cleaning Environment. In An Integration of Phycoremediation Processes in Wastewater Treatment; Elsevier: Amsterdam, The Netherlands, 2022; pp. 231–248. ISBN 978-0-12-823499-0. [Google Scholar]
- Sierra-Alvarez, R.; Hollingsworth, J.; Zhou, M.S. Removal of Copper in an Integrated Sulfate Reducing Bioreactor−Crystallization Reactor System. Environ. Sci. Technol. 2007, 41, 1426–1431. [Google Scholar] [CrossRef]
- Chen, J.L.; Ortiz, R.; Steele, T.W.J.; Stuckey, D.C. Toxicants Inhibiting Anaerobic Digestion: A Review. Biotechnol. Adv. 2014, 32, 1523–1534. [Google Scholar] [CrossRef]
- Paulo, L.M.; Stams, A.J.M.; Sousa, D.Z. Methanogens, Sulphate and Heavy Metals: A Complex System. Rev. Environ. Sci. Bio/Technol. 2015, 14, 537–553. [Google Scholar] [CrossRef] [Green Version]
- Loreto, C.D.; Monge, O.; Martin, A.R.; Ochoa-Herrera, V.; Sierra-Alvarez, R.; Almendariz, F.J. Effect of Carbon Source and Metal Toxicity for Potential Acid Mine Drainage (AMD) Treatment with an Anaerobic Sludge Using Sulfate-Reduction. Water Sci. Technol. 2021, 83, 2669–2677. [Google Scholar] [CrossRef]
- Lin, Y.; Grembi, J.A.; Goots, S.S.; Sebastian, A.; Albert, I.; Brennan, R.A. Advantageous Microbial Community Development and Improved Performance of Pilot-Scale Field Systems Treating High-Risk Acid Mine Drainage with Crab Shell. J. Hazard. Mater. 2021, 420, 126665. [Google Scholar] [CrossRef]
- Méndez-García, C.; Pelaez, A.I.; Mesa, V.; Sanchez, J.; Golyshina, O.V.; Ferrer, M. Microbial Diversity and Metabolic Networks in Acid Mine Drainage Habitats. Front. Microbiol. 2015, 6, 475. [Google Scholar] [CrossRef] [Green Version]
- Grettenberger, C.L.; Hamilton, T.L. Metagenome-Assembled Genomes of Novel Taxa from an Acid Mine Drainage Environment. Appl. Environ. Microbiol. 2021, 87. [Google Scholar] [CrossRef]
- Konopka, A.; Lindemann, S.; Fredrickson, J. Dynamics in Microbial Communities: Unraveling Mechanisms to Identify Principles. ISME J. 2015, 9, 1488–1495. [Google Scholar] [CrossRef]
- Nemergut, D.R.; Schmidt, S.K.; Fukami, T.; O’Neill, S.P.; Bilinski, T.M.; Stanish, L.F.; Knelman, J.E.; Darcy, J.L.; Lynch, R.C.; Wickey, P.; et al. Patterns and Processes of Microbial Community Assembly. Microbiol. Mol. Biol. Rev. 2013, 77, 342–356. [Google Scholar] [CrossRef] [Green Version]
- Aguinaga, O.E.; White, K.N.; Dean, A.P.; Pittman, J.K. Addition of Organic Acids to Acid Mine Drainage Polluted Wetland Sediment Leads to Microbial Community Structure and Functional Changes and Improved Water Quality. Environ. Pollut. 2021, 290, 118064. [Google Scholar] [CrossRef]
- Méndez, G.; Trueba, G.; Sierra-Alvarez, R.; Ochoa-Herrera, V. Treatment of Acid Rock Drainage Using a Sulphate-Reducing Bioreactor with a Limestone Precolumn. Environ. Technol. 2021, 1–12. [Google Scholar] [CrossRef]
- Hedin, R.S.; Watzlaf, G.R.; Nairn, R.W. Passive Treatment of Acid Mine Drainage with Limestone. J. Environ. Qual. 1994, 23, 1338–1345. [Google Scholar] [CrossRef] [Green Version]
- Rice, E.W.; Baird, R.B.; Eaton, A.D.; Clesceri, L.S. Standard Methods for the Examination of Water and Wastewater, 22nd ed.; American Public Health Association (APHA); American Water Works Association (AWWA); Water Environment Federation (WEF): Washington, WA, USA, 2012. [Google Scholar]
- Trüper, H.G.; Schlegel, H.G. Sulphur Metabolism in Thiorhodaceae I. Quantitative Measurements on Growing Cells OfChromatium Okenii. Antonie Leeuwenhoek 1964, 30, 225–238. [Google Scholar] [CrossRef]
- Barua, V.B.; Kalamdhad, A.S. Biochemical Methane Potential Test of Untreated and Hot Air Oven Pretreated Water Hyacinth: A Comparative Study. J. Clean. Prod. 2017, 166, 273–284. [Google Scholar] [CrossRef]
- Zambrano-Romero, A. Changes in Microbial Composition during the Removal of Copper and Zinc in a Bioreactor with a Limestone Pre-Column System. Master’s Thesis, Universidad San Francisco de Quito, Quito, Ecuador, 2018. [Google Scholar]
- Callahan, B.J.; McMurdie, P.J.; Rosen, M.J.; Han, A.W.; Johnson, A.J.A.; Holmes, S.P. DADA2: High-Resolution Sample Inference from Illumina Amplicon Data. Nat. Methods 2016, 13, 581–583. [Google Scholar] [CrossRef] [Green Version]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Copenhagen, Denmark, 2018. [Google Scholar]
- Schloss, P.D.; Westcott, S.L.; Ryabin, T.; Hall, J.R.; Hartmann, M.; Hollister, E.B.; Lesniewski, R.A.; Oakley, B.B.; Parks, D.H.; Robinson, C.J.; et al. Introducing Mothur: Open-Source, Platform-Independent, Community-Supported Software for Describing and Comparing Microbial Communities. Appl. Environ. Microbiol. 2009, 75, 7537–7541. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Quast, C.; Pruesse, E.; Yilmaz, P.; Gerken, J.; Schweer, T.; Yarza, P.; Peplies, J.; Glöckner, F.O. The SILVA Ribosomal RNA Gene Database Project: Improved Data Processing and Web-Based Tools. Nucleic Acids Res. 2012, 41, D590–D596. [Google Scholar] [CrossRef] [PubMed]
- Oksanen, J.; Blanchet, F.G.; Kindt, R.; Legendre, P.; Minchin, P.R.; O’Hara, R.B.; Simpson, G.L.; Solymos, P.; Stevens, M.H.H.; Wagner, H. Vegan: Community Ecology Package; R Package, 2015; Available online: https://www.worldagroforestry.org/publication/vegan-community-ecology-package-r-package-vegan-vers-22-1 (accessed on 14 December 2021).
- Kim, B.-R.; Shin, J.; Guevarra, R.B.; Lee, J.H.; Kim, D.W.; Seol, K.-H.; Lee, J.-H.; Kim, H.B.; Isaacson, R.E. Deciphering Diversity Indices for a Better Understanding of Microbial Communities. J. Microbiol. Biotechnol. 2017, 27, 2089–2093. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jost, L.; Chao, A.; Chazdon, R. Compositional Similarity and β (Beta) Diversity. In Biological Diversity Frontiers in Measurement and Assesment. Oxford University Press: Oxford, UK, 2010. [Google Scholar]
- Wolda, H. Similarity Indices, Sample Size and Diversity. Oecologia 1981, 50, 296–302. [Google Scholar] [CrossRef]
- Vítěz, T.; Novák, D.; Lochman, J.; Vítězová, M. Methanogens Diversity during Anaerobic Sewage Sludge Stabilization and the Effect of Temperature. Processes 2020, 8, 822. [Google Scholar] [CrossRef]
- Rabus, R.; Venceslau, S.S.; Wöhlbrand, L.; Voordouw, G.; Wall, J.D.; Pereira, I.A.C. Chapter Two—A Post-Genomic View of the Ecophysiology, Catabolism and Biotechnological Relevance of Sulphate-Reducing Prokaryotes. Adv. Microb. Physiol. 2015, 66, 55–321. [Google Scholar]
- Hattori, S. Syntrophic Acetate-Oxidizing Microbes in Methanogenic Environments. Microbes Environ. 2008, 23, 118–127. [Google Scholar] [CrossRef] [Green Version]
- Kolde, R. Pheatmap: Pretty Heatmaps. 2015. Available online: https://mran.microsoft.com/snapshot/2015-09-04/web/packages/pheatmap/pheatmap.pdf (accessed on 14 December 2021).
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular Evolutionary Genetics Analysis across Computing Platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef]
- Haroon, M.F.; Hu, S.; Shi, Y.; Imelfort, M.; Keller, J.; Hugenholtz, P.; Yuan, Z.; Tyson, G.W. Anaerobic Oxidation of Methane Coupled to Nitrate Reduction in a Novel Archaeal Lineage. Nature 2013, 500, 567–570. [Google Scholar] [CrossRef]
- Thompson, J.D.; Higgins, D.G.; Gibson, T.J. CLUSTAL W: Improving the Sensitivity of Progressive Multiple Sequence Alignment through Sequence Weighting, Position-Specific Gap Penalties and Weight Matrix Choice. Nucleic Acids Res. 1994, 22, 4673–4680. [Google Scholar] [CrossRef] [Green Version]
- Letunic, I.; Bork, P. Interactive Tree Of Life (ITOL) v4: Recent Updates and New Developments. Nucleic Acids Res. 2019, 47, W256–W259. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chen, Y.; Cheng, J.J.; Creamer, K.S. Inhibition of Anaerobic Digestion Process: A Review. Bioresour. Technol. 2008, 99, 4044–4064. [Google Scholar] [CrossRef] [PubMed]
- Ayala-Parra, P.; Sierra-Alvarez, R.; Field, J.A. Treatment of Acid Rock Drainage Using a Sulfate-Reducing Bioreactor with Zero-Valent Iron. J. Hazard. Mater. 2016, 308, 97–105. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dinh, H.T.; Kuever, J.; Mußmann, M.; Hassel, A.W.; Stratmann, M.; Widdel, F. Iron Corrosion by Novel Anaerobic Microorganisms. Nature 2004, 427, 829–832. [Google Scholar] [CrossRef] [PubMed]
- Kieu, H.T.Q.; Müller, E.; Horn, H. Heavy Metal Removal in Anaerobic Semi-Continuous Stirred Tank Reactors by a Consortium of Sulfate-Reducing Bacteria. Water Res. 2011, 45, 3863–3870. [Google Scholar] [CrossRef]
- Bao, Y.; Jin, X.; Guo, C.; Lu, G.; Dang, Z. Sulfate-Reducing Bacterial Community Shifts in Response to Acid Mine Drainage in the Sediment of the Hengshi Watershed, South China. Environ. Sci. Pollut. Res. 2021, 28, 2822–2834. [Google Scholar] [CrossRef]
- Calderón, D.; Ochoa-Herrera, V. Evaluación de Consumo de Sustrato y Toxicidad Microbiana de Cobre (Cu (II)) y Zinc (Zn (II)) En Bacterias Sulfato-Reductoras Presentes En Sedimentos Anaerobios. Ph.D Thesis, Universidad San Francisco de Quito, Quito, Ecuador, 2016; pp. 1–54. [Google Scholar]
- Jarrell, K.F.; Saulnier, M.; Ley, A. Inhibition of Methanogenesis in Pure Cultures by Ammonia, Fatty Acids, and Heavy Metals, and Protection against Heavy Metal Toxicity by Sewage Sludge. Can. J. Microbiol. 1987, 33, 551–554. [Google Scholar] [CrossRef]
- Moosa, S.; Harrison, S.T.L. Product Inhibition by Sulphide Species on Biological Sulphate Reduction for the Treatment of Acid Mine Drainage. Hydrometallurgy 2006, 83, 214–222. [Google Scholar] [CrossRef]
- Gonzalez-Estrella, J.; Puyol, D.; Sierra-Alvarez, R.; Field, J.A. Role of Biogenic Sulfide in Attenuating Zinc Oxide and Copper Nanoparticle Toxicity to Acetoclastic Methanogenesis. J. Hazard. Mater. 2015, 283, 755–763. [Google Scholar] [CrossRef]
- Yue, Z.; Yu, H.; Wang, Z. Anaerobic Digestion of Cattail with Rumen Culture in the Presence of Heavy Metals. Bioresour. Technol. 2007, 98, 781–786. [Google Scholar] [CrossRef]
- Mudhoo, A.; Kumar, S. Effects of Heavy Metals as Stress Factors on Anaerobic Digestion Processes and Biogas Production from Biomass. Int. J. Environ. Sci. Technol. 2013, 10, 1383–1398. [Google Scholar] [CrossRef] [Green Version]
- Alrawashdeh, K.A.b.; Gul, E.; Yang, Q.; Yang, H.; Bartocci, P.; Fantozzi, F. Effect of Heavy Metals in the Performance of Anaerobic Digestion of Olive Mill Waste. Processes 2020, 8, 1146. [Google Scholar] [CrossRef]
- Medírcio, S.N.; Leão, V.A.; Teixeira, M.C. Specific Growth Rate of Sulfate Reducing Bacteria in the Presence of Manganese and Cadmium. J. Hazard. Mater. 2007, 143, 593–596. [Google Scholar] [CrossRef]
- Neculita, C.-M.; Zagury, G.J.; Bussière, B. Passive Treatment of Acid Mine Drainage in Bioreactors Using Sulfate-Reducing Bacteria. J. Environ. Qual. 2007, 36, 1–16. [Google Scholar] [CrossRef]
- Iakovleva, E.; Mäkilä, E.; Salonen, J.; Sitarz, M.; Wang, S.; Sillanpää, M. Acid Mine Drainage (AMD) Treatment: Neutralization and Toxic Elements Removal with Unmodified and Modified Limestone. Ecol. Eng. 2015, 81, 30–40. [Google Scholar] [CrossRef]
- Tuomisto, H. A Diversity of Beta Diversities: Straightening up a Concept Gone Awry. Part 1. Defining Beta Diversity as a Function of Alpha and Gamma Diversity. Ecography 2010, 33, 2–22. [Google Scholar] [CrossRef]
- Wagner, B.D.; Grunwald, G.K.; Zerbe, G.O.; Mikulich-Gilbertson, S.K.; Robertson, C.E.; Zemanick, E.T.; Harris, J.K. On the Use of Diversity Measures in Longitudinal Sequencing Studies of Microbial Communities. Front. Microbiol. 2018, 9, 1037. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Walters, K.E.; Martiny, J.B.H. Alpha-, Beta-, and Gamma-Diversity of Bacteria Varies across Habitats. PLoS ONE 2020, 15, e0233872. [Google Scholar] [CrossRef] [PubMed]
- ElNaker, N.A.; Elektorowicz, M.; Naddeo, V.; Hasan, S.W.; Yousef, A.F. Assessment of Microbial Community Structure and Function in Serially Passaged Wastewater Electro-Bioreactor Sludge: An Approach to Enhance Sludge Settleability. Sci. Rep. 2018, 8, 7013. [Google Scholar] [CrossRef]
- Pane, C.; Sorrentino, R.; Scotti, R.; Molisso, M.; Di Matteo, A.; Celano, G.; Zaccardelli, M. Alpha and Beta-Diversity of Microbial Communities Associated to Plant Disease Suppressive Functions of On-Farm Green Composts. Agriculture 2020, 10, 113. [Google Scholar] [CrossRef] [Green Version]
- Zhou, L.; Zhou, Y.; Yao, X.; Cai, J.; Liu, X.; Tang, X.; Zhang, Y.; Jang, K.-S.; Jeppesen, E. Decreasing Diversity of Rare Bacterial Subcommunities Relates to Dissolved Organic Matter along Permafrost Thawing Gradients. Environ. Int. 2020, 134, 105330. [Google Scholar] [CrossRef] [PubMed]
- Sánchez-Andrea, I.; Sanz, J.L.; Bijmans, M.F.M.; Stams, A.J.M. Sulfate Reduction at Low PH to Remediate Acid Mine Drainage. J. Hazard. Mater. 2014, 269, 98–109. [Google Scholar] [CrossRef] [PubMed]
- De Vrieze, J.; Hennebel, T.; Boon, N.; Verstraete, W. Methanosarcina: The Rediscovered Methanogen for Heavy Duty Biomethanation. Bioresour. Technol. 2012, 112, 1–9. [Google Scholar] [CrossRef]
- Lackner, N.; Hintersonnleitner, A.; Wagner, A.O.; Illmer, P. Hydrogenotrophic Methanogenesis and Autotrophic Growth of Methanosarcina Thermophila. Archaea 2018, 2018, 4712608. [Google Scholar] [CrossRef] [Green Version]
- Serrano-Silva, N.; Sarria-Guzmán, Y.; Dendooven, L.; Luna-Guido, M. Methanogenesis and Methanotrophy in Soil: A Review. Pedosphere 2014, 24, 291–307. [Google Scholar] [CrossRef]
- Tate, K.R. Soil Methane Oxidation and Land-Use Change—From Process to Mitigation. Soil Biol. Biochem. 2015, 80, 260–272. [Google Scholar] [CrossRef]
- Ferry, J.G. Methanosarcina Acetivorans: A Model for Mechanistic Understanding of Aceticlastic and Reverse Methanogenesis. Front. Microbiol. 2020, 11, 1806. [Google Scholar] [CrossRef]
- Ochoa-Herrera, V.; Field, J.A.; Luna-Velasco, A.; Sierra-Alvarez, R. Microbial Toxicity and Biodegradability of Perfluorooctane Sulfonate (PFOS) and Shorter Chain Perfluoroalkyl and Polyfluoroalkyl Substances (PFASs). Environ. Sci. Processes Impacts 2016, 18, 1236–1246. [Google Scholar] [CrossRef]
- Waite, D.W.; Chuvochina, M.; Pelikan, C.; Parks, D.H.; Yilmaz, P.; Wagner, M.; Loy, A.; Naganuma, T.; Nakai, R.; Whitman, W.B.; et al. Proposal to Reclassify the Proteobacterial Classes Deltaproteobacteria and Oligoflexia, and the Phylum Thermodesulfobacteria into Four Phyla Reflecting Major Functional Capabilities. Int. J. Syst. Evol. Microbiol. 2020, 70, 5972–6016. [Google Scholar] [CrossRef]
- Cruz-Morales, P.; Orellana, C.A.; Moutafis, G.; Moonen, G.; Rincon, G.; Nielsen, L.K.; Marcellin, E. Revisiting the Evolution and Taxonomy of Clostridia, a Phylogenomic Update. Genome Biol. Evol. 2019, 11, 2035–2044. [Google Scholar] [CrossRef] [Green Version]
- Besaury, L.; Bodilis, J.; Delgas, F.; Andrade, S.; De la Iglesia, R.; Ouddane, B.; Quillet, L. Abundance and Diversity of Copper Resistance Genes CusA and CopA in Microbial Communities in Relation to the Impact of Copper on Chilean Marine Sediments. Mar. Pollut. Bull. 2013, 67, 16–25. [Google Scholar] [CrossRef] [PubMed]
- Gillan, D.C. Metal Resistance Systems in Cultivated Bacteria: Are They Found in Complex Communities? Curr. Opin. Biotechnol. 2016, 38, 123–130. [Google Scholar] [CrossRef] [PubMed]
- Giordani, A.; Rodriguez, R.P.; Sancinetti, G.P.; Hayashi, E.A.; Beli, E.; Brucha, G. Effect of Low PH and Metal Content on Microbial Community Structure in an Anaerobic Sequencing Batch Reactor Treating Acid Mine Drainage. Miner. Eng. 2019, 141, 105860. [Google Scholar] [CrossRef]
- Müller, A.L.; Kjeldsen, K.U.; Rattei, T.; Pester, M.; Loy, A. Phylogenetic and Environmental Diversity of DsrAB-Type Dissimilatory (Bi)Sulfite Reductases. ISME J. 2015, 9, 1152–1165. [Google Scholar] [CrossRef] [Green Version]
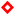
 ), and bioreactor effluent (
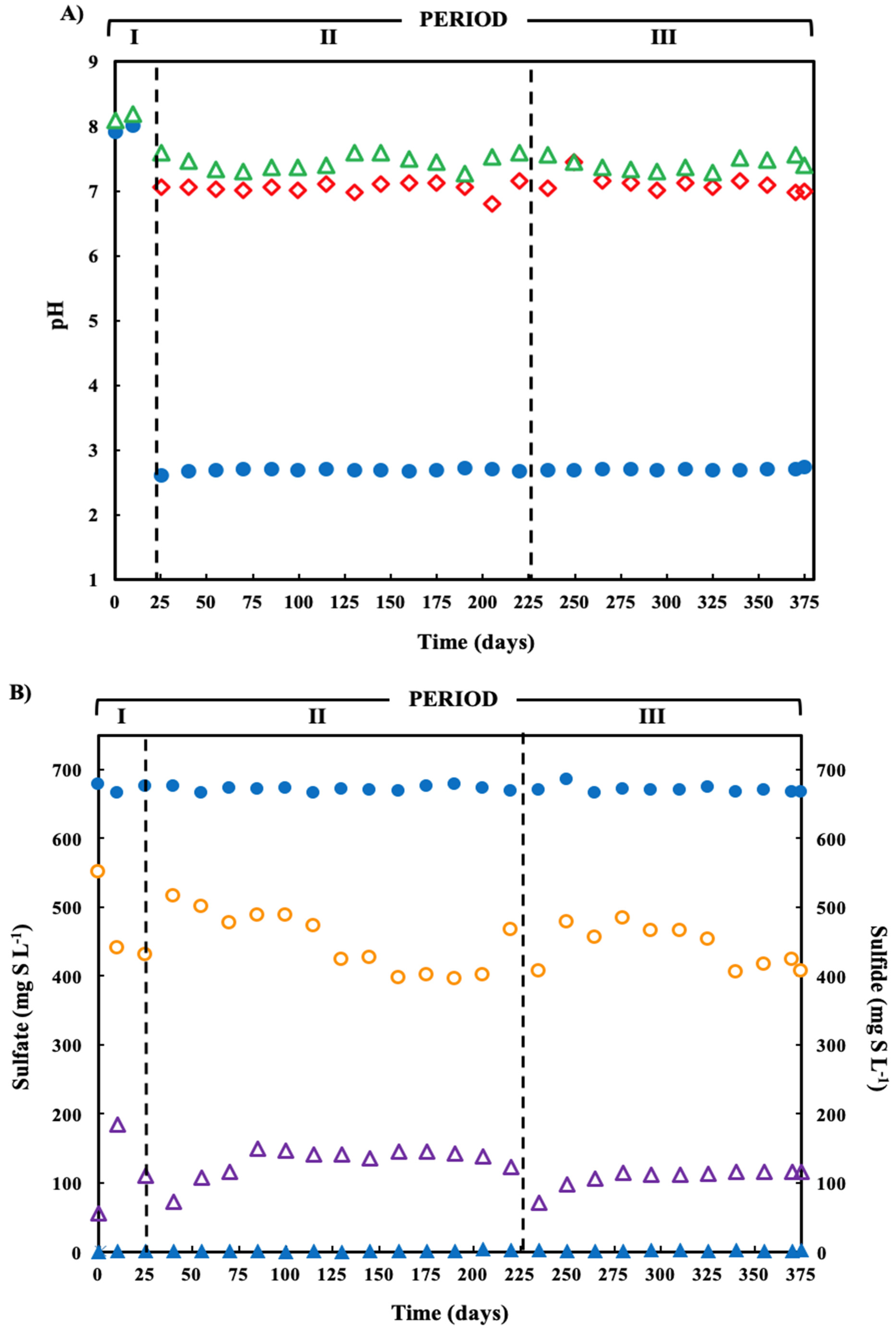
), and bioreactor effluent (  ). (B) Sulfate reduction (primary axis-left) and sulfide production (secondary axis-right) in the treatment system fed with a synthetic ARD containing sulfate (2000 mg L−1), acetate as electron donor (2.5 g COD L−1), Cu(II) (15 mg L−1 during periods II and III) and Zn(II) (15 mg L−1 during period III): sulfate (●) and sulfide (▲) in the influent and sulfate (
). (B) Sulfate reduction (primary axis-left) and sulfide production (secondary axis-right) in the treatment system fed with a synthetic ARD containing sulfate (2000 mg L−1), acetate as electron donor (2.5 g COD L−1), Cu(II) (15 mg L−1 during periods II and III) and Zn(II) (15 mg L−1 during period III): sulfate (●) and sulfide (▲) in the influent and sulfate (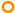  ) and sulfide (
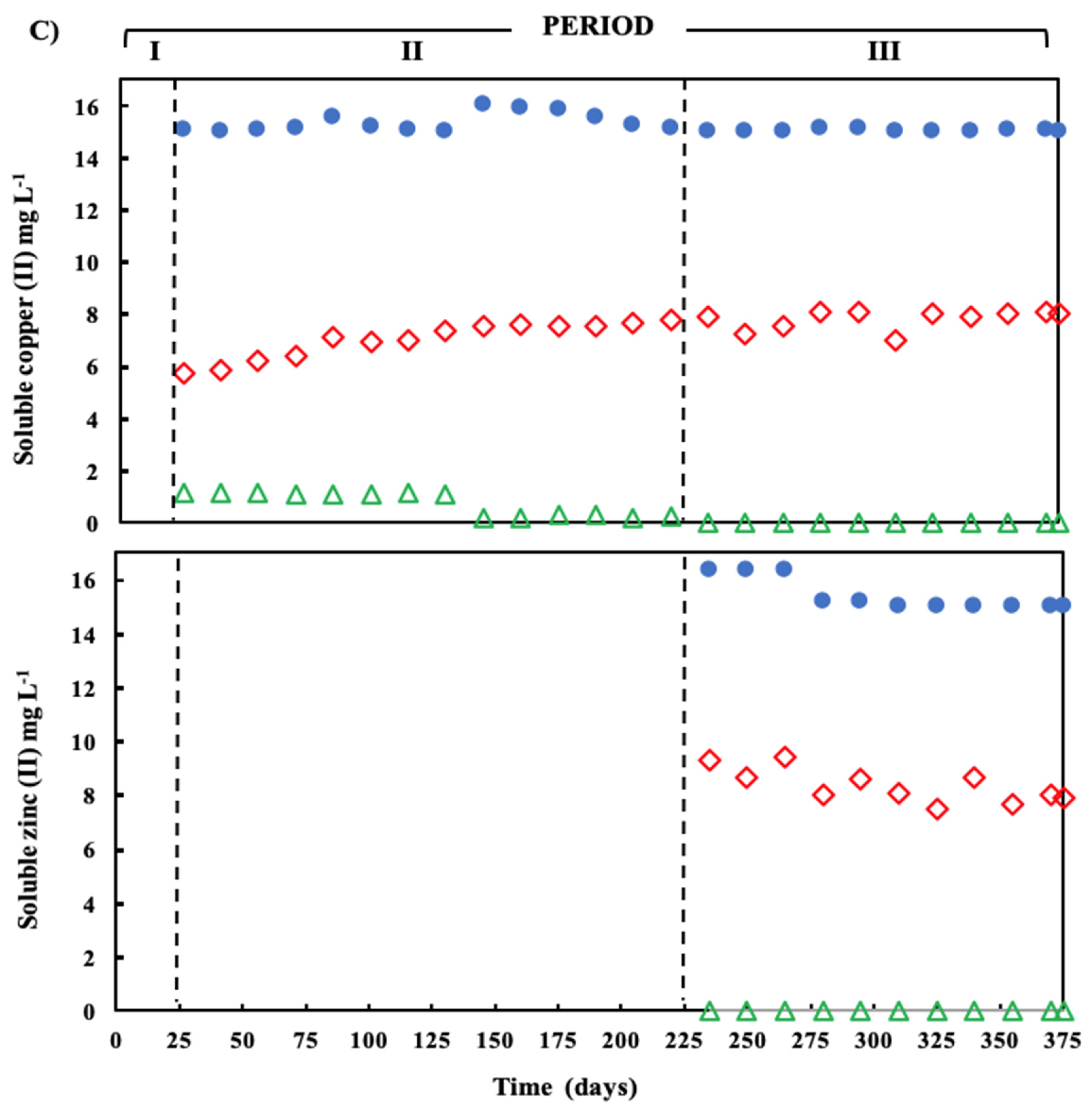
) and sulfide (  ) in the effluent. (C) Concentration of soluble Cu(II) and soluble Zn(II) during the operation of the treatment system fed with a synthetic ARD containing sulfate (2000 mg L−1), acetate as electron donor (2.5 g COD L−1), Cu (II) (15 mg L−1 during periods II and III) and Zn(II) (15 mg L−1 during period III): limestone pre-column influent (●), limestone pre-column effluent/bioreactor influent (
) in the effluent. (C) Concentration of soluble Cu(II) and soluble Zn(II) during the operation of the treatment system fed with a synthetic ARD containing sulfate (2000 mg L−1), acetate as electron donor (2.5 g COD L−1), Cu (II) (15 mg L−1 during periods II and III) and Zn(II) (15 mg L−1 during period III): limestone pre-column influent (●), limestone pre-column effluent/bioreactor influent (  ), and bioreactor effluent (
), and bioreactor effluent (  ).
).
 ), and bioreactor effluent (
), and bioreactor effluent (  ). (B) Sulfate reduction (primary axis-left) and sulfide production (secondary axis-right) in the treatment system fed with a synthetic ARD containing sulfate (2000 mg L−1), acetate as electron donor (2.5 g COD L−1), Cu(II) (15 mg L−1 during periods II and III) and Zn(II) (15 mg L−1 during period III): sulfate (●) and sulfide (▲) in the influent and sulfate (
). (B) Sulfate reduction (primary axis-left) and sulfide production (secondary axis-right) in the treatment system fed with a synthetic ARD containing sulfate (2000 mg L−1), acetate as electron donor (2.5 g COD L−1), Cu(II) (15 mg L−1 during periods II and III) and Zn(II) (15 mg L−1 during period III): sulfate (●) and sulfide (▲) in the influent and sulfate (  ) and sulfide (
) and sulfide (  ) in the effluent. (C) Concentration of soluble Cu(II) and soluble Zn(II) during the operation of the treatment system fed with a synthetic ARD containing sulfate (2000 mg L−1), acetate as electron donor (2.5 g COD L−1), Cu (II) (15 mg L−1 during periods II and III) and Zn(II) (15 mg L−1 during period III): limestone pre-column influent (●), limestone pre-column effluent/bioreactor influent (
) in the effluent. (C) Concentration of soluble Cu(II) and soluble Zn(II) during the operation of the treatment system fed with a synthetic ARD containing sulfate (2000 mg L−1), acetate as electron donor (2.5 g COD L−1), Cu (II) (15 mg L−1 during periods II and III) and Zn(II) (15 mg L−1 during period III): limestone pre-column influent (●), limestone pre-column effluent/bioreactor influent (  ), and bioreactor effluent (
), and bioreactor effluent (  ).
).
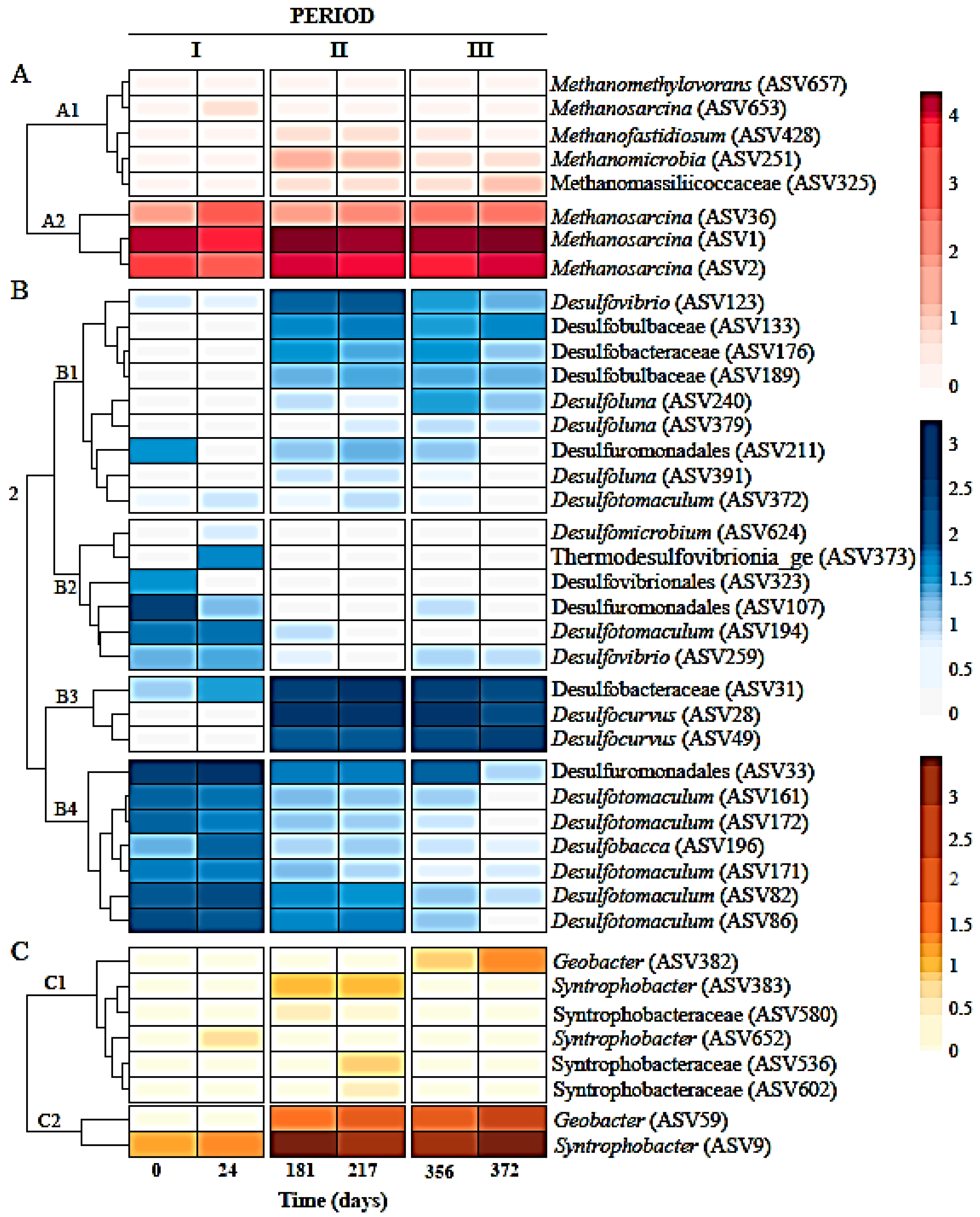
 ). (b) Relative abundances of most abundant amplicon sequence variants (ASVs) at genus level in six sludge samples during the three different operation periods of the sulfate-reducing bioreactor with the limestone pre-column system. Two samples or biological replicates were collected in each period.
). (b) Relative abundances of most abundant amplicon sequence variants (ASVs) at genus level in six sludge samples during the three different operation periods of the sulfate-reducing bioreactor with the limestone pre-column system. Two samples or biological replicates were collected in each period.
 ). (b) Relative abundances of most abundant amplicon sequence variants (ASVs) at genus level in six sludge samples during the three different operation periods of the sulfate-reducing bioreactor with the limestone pre-column system. Two samples or biological replicates were collected in each period.
). (b) Relative abundances of most abundant amplicon sequence variants (ASVs) at genus level in six sludge samples during the three different operation periods of the sulfate-reducing bioreactor with the limestone pre-column system. Two samples or biological replicates were collected in each period.
| Period a | Time of Operation b (d) | Influent | Effluent pH | Sulfate Removal (%) | Effluent Sulfide (mg S L−1) | %CODin c | ||
|---|---|---|---|---|---|---|---|---|
| pH | H2S Formed | CH4 | Organic COD Removal | |||||
| I d | 24 | 8.04 ± 0.36 | 8.04 ± 0.36 | 37.7 ± 4.8 | 151.0 ± 20.1 | 49.4 ± 7.1 | 0.0 ± 0.0 | 68.1 ± 7.48 |
| II | 196 | 7.45 ± 0.15 | 7.45 ± 0.15 | 38.0 ± 12.3 | 179.8 ± 19.6 | 51.3 ± 12.6 | 0.0 ± 0.0 | 74.0 ± 5.09 |
| III | 155 | 7.43 ± 0.21 | 7.43 ± 0.21 | 48.8 ± 5.1 | 149.1 ± 18.0 | 42.4 ± 10.1 | 0.0 ± 0.0 | 73.0 ± 4.82 |
| Period a | Metal in the Influent | Removal of Soluble Metal (%) | ||||||
|---|---|---|---|---|---|---|---|---|
| Limestone Pre-Column | Bioreactor | Complete System | ||||||
| (mg Cu(II) L−1) | (mg Zn(II) L−1) | Cu(II) | Zn(II) | Cu(II) | Zn(II) | Cu(II) | Zn(II) | |
| II | 15.33 ± 0.37 | - | 54.5 ± 0.8 | - | 96.8 ± 0.8 | - | 98.5 ± 0.6 | - |
| III | 15.17 ± 0.35 | 15.54 ± 0.60 | 50.3 ± 1.9 | 47.1 ± 0.7 | 99.8 ± 0.9 | 99.9 ± 1.0 | 99.2 ± 0.4 | >99.9 ± 1.0 |
| Period | Time (d) | Shannon | Inverse Simpson | Richness | Chao1 | Evenness |
|---|---|---|---|---|---|---|
| I | 0 | 3.64 | 13.24 | 306.00 | 306.86 | 0.64 |
| 24 | 3.90 | 23.11 | 250.00 | 250.00 | 0.71 | |
| II | 181 | 2.88 | 7.28 | 272.00 | 278.11 | 0.51 |
| 217 | 2.99 | 8.07 | 277.00 | 278.56 | 0.53 | |
| III | 356 | 2.93 | 7.72 | 246.00 | 246.43 | 0.53 |
| 372 | 2.73 | 6.77 | 217.00 | 219.57 | 0.51 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zambrano-Romero, A.; Ramirez-Villacis, D.X.; Trueba, G.; Sierra-Alvarez, R.; Leon-Reyes, A.; Cardenas, P.; Ochoa-Herrera, V. Dynamics of Microbial Communities during the Removal of Copper and Zinc in a Sulfate-Reducing Bioreactor with a Limestone Pre-Column System. Int. J. Environ. Res. Public Health 2022, 19, 1484. https://doi.org/10.3390/ijerph19031484
Zambrano-Romero A, Ramirez-Villacis DX, Trueba G, Sierra-Alvarez R, Leon-Reyes A, Cardenas P, Ochoa-Herrera V. Dynamics of Microbial Communities during the Removal of Copper and Zinc in a Sulfate-Reducing Bioreactor with a Limestone Pre-Column System. International Journal of Environmental Research and Public Health. 2022; 19(3):1484. https://doi.org/10.3390/ijerph19031484
Chicago/Turabian StyleZambrano-Romero, Aracely, Dario X. Ramirez-Villacis, Gabriel Trueba, Reyes Sierra-Alvarez, Antonio Leon-Reyes, Paul Cardenas, and Valeria Ochoa-Herrera. 2022. "Dynamics of Microbial Communities during the Removal of Copper and Zinc in a Sulfate-Reducing Bioreactor with a Limestone Pre-Column System" International Journal of Environmental Research and Public Health 19, no. 3: 1484. https://doi.org/10.3390/ijerph19031484
APA StyleZambrano-Romero, A., Ramirez-Villacis, D. X., Trueba, G., Sierra-Alvarez, R., Leon-Reyes, A., Cardenas, P., & Ochoa-Herrera, V. (2022). Dynamics of Microbial Communities during the Removal of Copper and Zinc in a Sulfate-Reducing Bioreactor with a Limestone Pre-Column System. International Journal of Environmental Research and Public Health, 19(3), 1484. https://doi.org/10.3390/ijerph19031484








