Exome Sequencing in an ADSHE Family: VUS Identification and Limits
Abstract
1. Introduction
2. Materials and Methods
2.1. Sample Composition
2.2. Whole Exome Sequencing and Data Processing
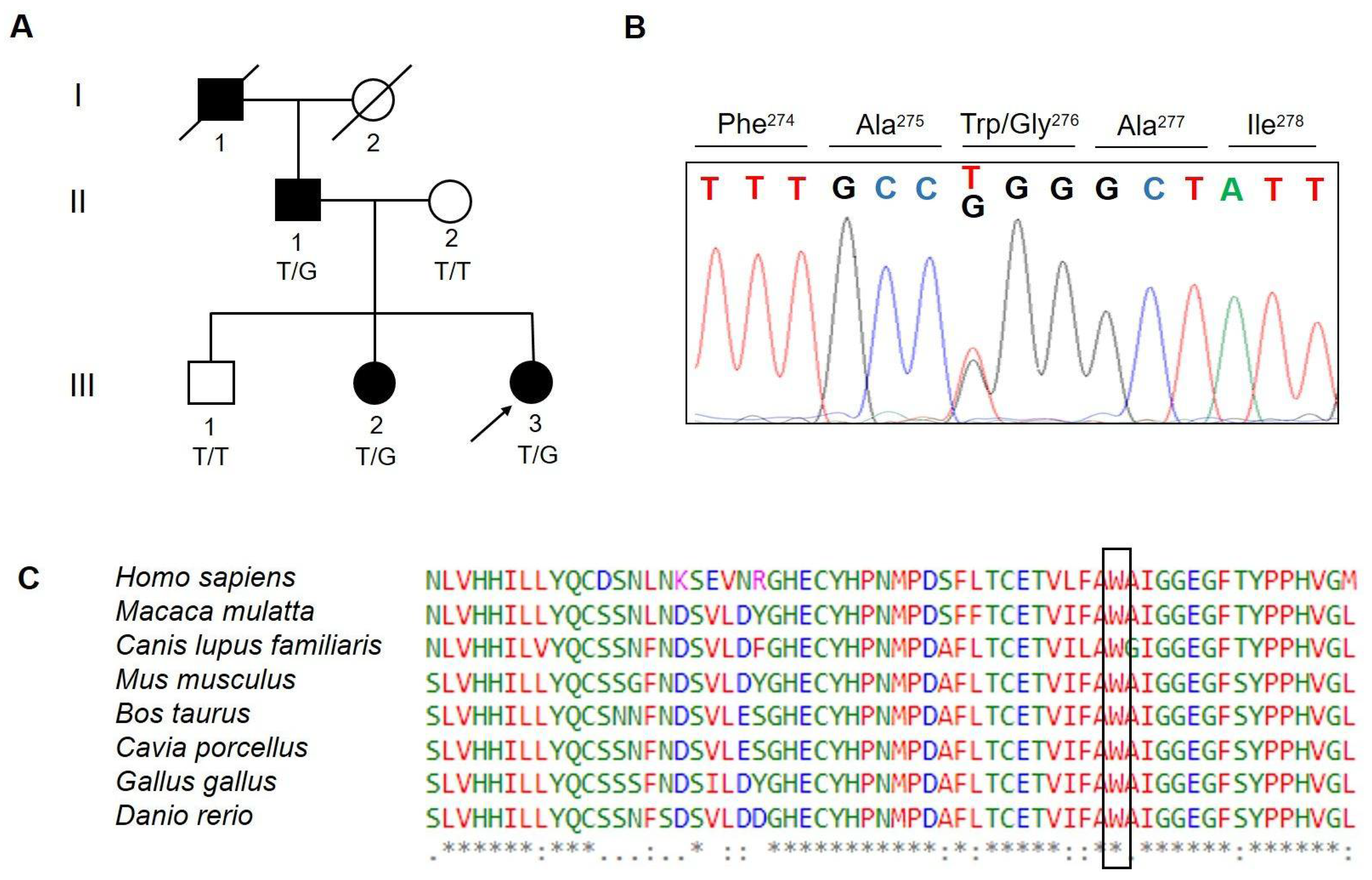
2.3. Variant Confirmation and Familial Segregation Analysis
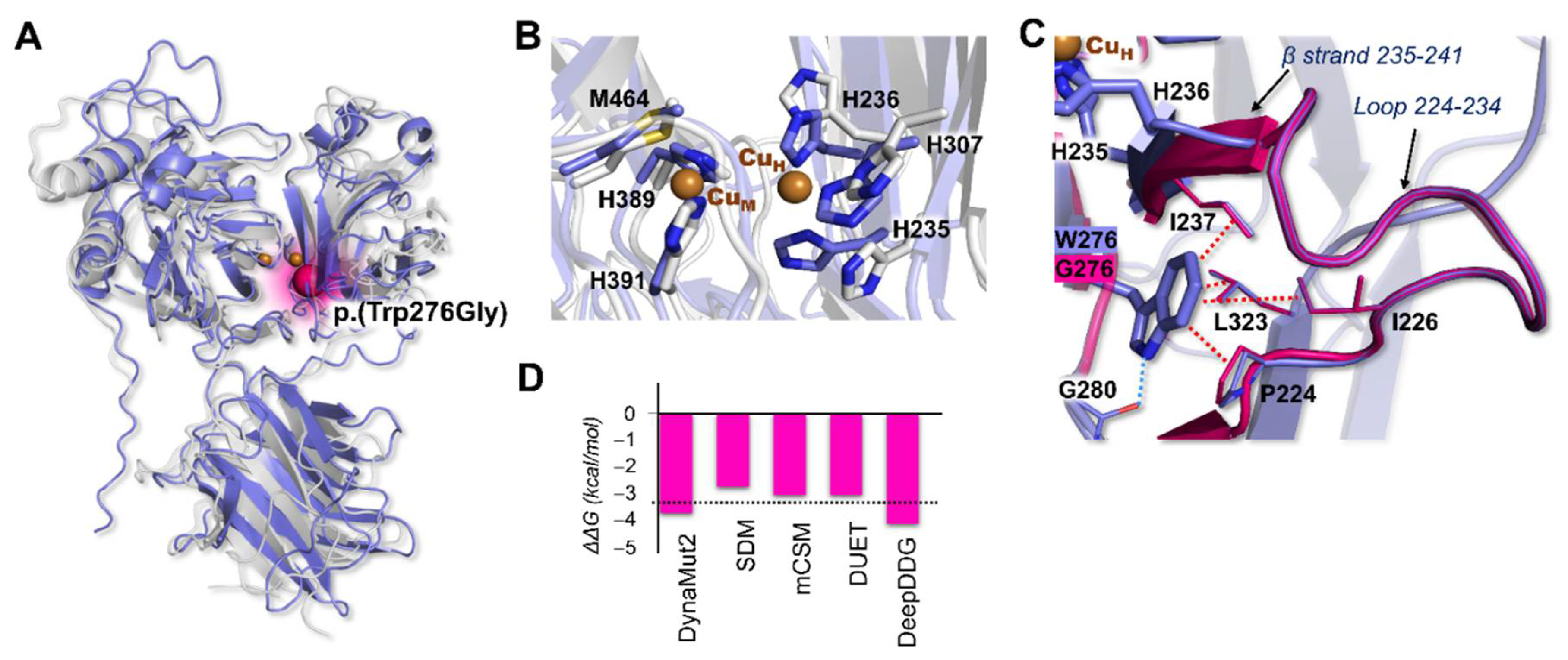
2.4. Computational Analysis of MOXD1 and Mutant Structural Features
3. Results
3.1. Variant Filtration and Prioritization
3.2. Modeling
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Tinuper, P.; Bisulli, F.; Cross, J.H.; Hesdorffer, D.; Kahane, P.; Nobili, L.; Provini, F.; Scheffer, I.E.; Tassi, L.; Vignatelli, L.; et al. Definition and diagnostic criteria of sleep-related hypermotor epilepsy. Neurology 2016, 86, 1834–1842. [Google Scholar] [CrossRef] [PubMed]
- Ryvlin, P.; Minotti, L.; Demarquay, G.; Hirsch, E.; Arzimanoglou, A.; Hoffman, D.; Guénot, M.; Picard, F.; Rheims, S.; Kahane, P. Nocturnal hypermotor seizures, suggesting frontal lobe epilepsy, can originate in the insula. Epilepsia 2006, 47, 755–765. [Google Scholar] [CrossRef] [PubMed]
- Gibbs, S.A.; Figorilli, M.; Casaceli, G.; Proserpio, P.; Nobili, L. Sleep Related Hypermotor Seizures with a Right Parietal Onset. J. Clin. Sleep Med. 2015, 11, 953–955. [Google Scholar] [CrossRef]
- Steinlein, O.K.; Kaneko, S.; Hirose, S. Nicotinic acetylcholine receptor mutations. In Jasper’s Basic Mechanisms of the Epilepsies; Noebels, J.L., Avoli, M., Rogawski, M.A., Olsen, R.W., Delgado-Escueta, A.V., Eds.; National Center for Biotechnology Information (US): Bethesda, MD, USA, 2012. [Google Scholar]
- Menghi, V.; Bisulli, F.; Tinuper, P.; Nobili, L. Sleep-related hypermotor epilepsy: Prevalence, impact and management strategies. Nat. Sci. Sleep 2018, 10, 317–326. [Google Scholar] [CrossRef] [PubMed]
- Oyrer, J.; Maljevic, S.; Scheffer, I.E.; Berkovic, S.F.; Petrou, S.; Reid, C.A. Ion Channels in Genetic Epilepsy: From Genes and Mechanisms to Disease-Targeted Therapies. Pharmacol. Rev. 2018, 70, 142–173. [Google Scholar] [CrossRef]
- Asioli, G.M.; Rossi, S.; Bisulli, F.; Licchetta, L.; Tinuper, P.; Provini, F. Therapy in Sleep-Related Hypermotor Epilepsy (SHE). Curr. Treat Options Neurol. 2020, 22, 1. [Google Scholar] [CrossRef]
- Steinlein, O.K.; Hoda, J.C.; Bertrand, S.; Bertrand, D. Mutations in familial nocturnal frontal lobe epilepsy might be associated with distinct neurological phenotypes. Seizure 2012, 21, 118–123. [Google Scholar] [CrossRef][Green Version]
- Nobili, L.; Proserpio, P.; Combi, R.; Provini, F.; Plazzi, G.; Bisulli, F.; Tassi, L.; Tinuper, P. Nocturnal frontal lobe epilepsy. Curr. Neurol. Neurosci. Rep. 2014, 14, 424. [Google Scholar] [CrossRef]
- Ferini-Strambi, L.; Sansoni, V.; Combi, R. Nocturnal frontal lobe epilepsy and the acetylcholine receptor. Neurologist 2012, 18, 343–349. [Google Scholar] [CrossRef]
- Combi, R.; Dalprà, L.; Ferini-Strambi, L.; Tenchini, M.L. Frontal lobe epilepsy and mutations of the corticotropin-releasing hormone gene. Ann. Neurol. 2005, 58, 899–904. [Google Scholar] [CrossRef]
- Heron, S.E.; Smith, K.R.; Bahlo, M.; Nobili, L.; Kahana, E.; Licchetta, L.; Oliver, K.L.; Mazarib, A.; Afawi, Z.; Korczyn, A.; et al. Missense mutations in the sodium-gated potassium channel gene KCNT1 cause severe autosomal dominant nocturnal frontal lobe epilepsy. Nat. Genet. 2012, 44, 1188–1190. [Google Scholar] [CrossRef] [PubMed]
- Ishida, S.; Picard, F.; Rudolf, G.; Noé, E.; Achaz, G.; Thomas, P.; Genton, P.; Mundwiller, E.; Wolff, M.; Marescaux, C.; et al. Mutations of DEPDC5 cause autosomal dominant focal epilepsies. Nat. Genet. 2013, 45, 552–555. [Google Scholar] [CrossRef] [PubMed]
- Korenke, G.C.; Eggert, M.; Thiele, H.; Nürnberg, P.; Sander, T.; Steinlein, O.K. Nocturnal frontal lobe epilepsy caused by a mutation in the GATOR1 complex gene NPRL3. Epilepsia 2016, 57, e60–e63. [Google Scholar] [CrossRef] [PubMed]
- Ricos, M.G.; Hodgson, B.L.; Pippucci, T.; Saidin, A.; Ong, Y.S.; Heron, S.E.; Licchetta, L.; Bisulli, F.; Bayly, M.A.; Hughes, J.; et al. Mutations in the mammalian target of rapamycin pathway regulators NPRL2 and NPRL3 cause focal epilepsy. Ann. Neurol. 2016, 79, 120–131. [Google Scholar] [CrossRef]
- Tinuper, P.; Bisulli, F. From nocturnal frontal lobe epilepsy to Sleep-Related Hypermotor Epilepsy: A 35-year diagnostic challenge. Seizure 2017, 44, 87–92. [Google Scholar] [CrossRef]
- Li, H.; Durbin, R. Fast and accurate long-read alignment with Burrows-Wheeler transform. Bioinformatics 2010, 26, 589–595. [Google Scholar] [CrossRef]
- Danecek, P.; Bonfield, J.K.; Liddle, J.; Marshall, J.; Ohan, V.; Pollard, M.O.; Whitwham, A.; Keane, T.; McCarthy, S.A.; Davies, R.M.; et al. Twelve years of SAMtools and BCFtools. Gigascience 2021, 10, giab008. [Google Scholar] [CrossRef]
- Varadi, M.; Anyango, S.; Deshpande, M.; Nair, S.; Natassia, C.; Yordanova, G.; Yuan, D.; Stroe, O.; Wood, G.; Laydon, A.; et al. AlphaFold Protein Structure Database: Massively expanding the structural coverage of protein-sequence space with high-accuracy models. Nucleic Acids Res. 2022, 50, D439–D444. [Google Scholar] [CrossRef]
- Rodrigues, C.H.M.; Pires, D.E.V.; Ascher, D.B. DynaMut2: Assessing changes in stability and flexibility upon single and multiple point missense mutations. Protein Sci. 2021, 30, 60–69. [Google Scholar] [CrossRef]
- Pandurangan, A.P.; Ochoa-Montaño, B.; Ascher, D.B.; Blundell, T.L. SDM: A server for predicting effects of mutations on protein stability. Nucleic Acids Res. 2017, 45, W229–W235. [Google Scholar] [CrossRef]
- Pires, D.E.; Ascher, D.B.; Blundell, T.L. mCSM: Predicting the effects of mutations in proteins using graph-based signatures. Bioinformatics 2014, 30, 335–342. [Google Scholar] [CrossRef] [PubMed]
- Pires, D.E.; Ascher, D.B.; Blundell, T.L. DUET: A server for predicting effects of mutations on protein stability using an integrated computational approach. Nucleic Acids Res. 2014, 42, W314–W319. [Google Scholar] [CrossRef] [PubMed]
- Cao, H.; Wang, J.; He, L.; Qi, Y.; Zhang, J.Z. DeepDDG: Predicting the Stability Change of Protein Point Mutations Using Neural Networks. J. Chem. Inf. Model 2019, 59, 1508–1514. [Google Scholar] [CrossRef] [PubMed]
- Ittisoponpisan, S.; Islam, S.A.; Khanna, T.; Alhuzimi, E.; David, A.; Sternberg, M.J.E. Can Predicted Protein 3D Structures Provide Reliable Insights into whether Missense Variants Are Disease Associated? J. Mol. Biol. 2019, 431, 2197–2212. [Google Scholar] [CrossRef]
- Sastry, G.M.; Adzhigirey, M.; Day, T.; Annabhimoju, R.; Sherman, W. Protein and ligand preparation: Parameters, protocols, and influence on virtual screening enrichments. J. Comput. Aided Mol. Des. 2013, 27, 221–234. [Google Scholar] [CrossRef]
- Schrödinger. Schrödinger Release 2021-4; Maestro; Schrödinger LLC: New York, NY, USA, 2021. [Google Scholar]
- Zhu, K.; Day, T.; Warshaviak, D.; Murrett, C.; Friesner, R.; Pearlman, D. Antibody structure determination using a combination of homology modeling, energy-based refinement, and loop prediction. Proteins 2014, 82, 1646–1655. [Google Scholar] [CrossRef]
- Geer, L.Y.; Domrachev, M.; Lipman, D.J.; Bryant, S.H. CDART: Protein homology by domain architecture. Genome Res. 2002, 12, 1619–1623. [Google Scholar] [CrossRef]
- Schrödinger. The PyMOL Molecular Graphics System; Version 2.0; Schrödinger LLC: New York, NY, USA, 2020. [Google Scholar]
- Vendelboe, T.V.; Harris, P.; Zhao, Y.; Walter, T.S.; Harlos, K.; El Omari, K.; Christensen, H.E. The crystal structure of human dopamine β-hydroxylase at 2.9 Å resolution. Sci. Adv. 2016, 2, e1500980. [Google Scholar] [CrossRef]
- Iyer, L.M.; Anantharaman, V.; Aravind, L. The DOMON domains are involved in heme and sugar recognition. Bioinformatics 2007, 23, 2660–2664. [Google Scholar] [CrossRef]
- Solomon, E.I.; Heppner, D.E.; Johnston, E.M.; Ginsbach, J.W.; Cirera, J.; Qayyum, M.; Kieber-Emmons, M.T.; Kjaergaard, C.H.; Hadt, R.G.; Tian, L. Copper active sites in biology. Chem. Rev. 2014, 114, 3659–3853. [Google Scholar] [CrossRef]
- Xin, X.; Mains, R.E.; Eipper, B.A. Monooxygenase X, a member of the copper-dependent monooxygenase family localized to the endoplasmic reticulum. J. Biol. Chem. 2004, 279, 48159–48167. [Google Scholar] [CrossRef] [PubMed]
- Shi, P.; Xu, J.; Xia, F.; Wang, Y.; Ren, J.; Liang, P.; Cui, H. MOXD1 knockdown suppresses the proliferation and tumor growth of glioblastoma cells via ER stress-inducing apoptosis. Cell Death Discov. 2022, 8, 174. [Google Scholar] [CrossRef] [PubMed]
- Lauterborn, J.C.; Ribak, C.E. Differences in dopamine beta-hydroxylase immunoreactivity between the brains of genetically epilepsy-prone and Sprague-Dawley rats. Epilepsy Res. 1989, 4, 161–176. [Google Scholar] [CrossRef]
- Szot, P.; Weinshenker, D.; White, S.S.; Robbins, C.A.; Rust, N.C.; Schwartzkroin, P.A.; Palmiter, R.D. Norepinephrine-deficient mice have increased susceptibility to seizure-inducing stimuli. J. Neurosci. 1999, 19, 10985–10992. [Google Scholar] [CrossRef]
- Fontenot, M.R.; Berto, S.; Liu, Y.; Werthmann, G.; Douglas, C.; Usui, N.; Gleason, K.; Tamminga, C.A.; Takahashi, J.S.; Konopka, G. Novel transcriptional networks regulated by CLOCK in human neurons. Genes Dev. 2017, 31, 2121–2135. [Google Scholar] [CrossRef]
- Tsuneoka, Y.; Tsukahara, S.; Yoshida, S.; Takase, K.; Oda, S.; Kuroda, M.; Funato, H. Moxd1 Is a Marker for Sexual Dimorphism in the Medial Preoptic Area, Bed Nucleus of the Stria Terminalis and Medial Amygdala. Front. Neuroanat. 2017, 11, 26. [Google Scholar] [CrossRef]
- Hoerder-Suabedissen, A.; Wang, W.Z.; Lee, S.; Davies, K.E.; Goffinet, A.M.; Rakić, S.; Parnavelas, J.; Reim, K.; Nicolić, M.; Paulsen, O.; et al. Novel markers reveal subpopulations of subplate neurons in the murine cerebral cortex. Cereb. Cortex 2009, 19, 1738–1750. [Google Scholar] [CrossRef]
- Douka, K.; Birds, I.; Wang, D.; Kosteletos, A.; Clayton, S.; Byford, A.; Vasconcelos, E.J.R.; O’Connell, M.J.; Deuchars, J.; Whitehouse, A.; et al. Cytoplasmic long noncoding RNAs are differentially regulated and translated during human neuronal differentiation. RNA 2021, 27, 1082–1101. [Google Scholar] [CrossRef]
- Enterría-Morales, D.; Del Rey, N.L.; Blesa, J.; López-López, I.; Gallet, S.; Prévot, V.; López-Barneo, J.; d’Anglemont de Tassigny, X. Molecular targets for endogenous glial cell line-derived neurotrophic factor modulation in striatal parvalbumin interneurons. Brain Commun. 2020, 2, fcaa105. [Google Scholar] [CrossRef]
| Chr | Position (GRCh37) | Ref | Alt | Gene ID | Effect | Feature_ID:HGVS.c:HGVS.p | CADD PHRED Score |
|---|---|---|---|---|---|---|---|
| Chr6 | 132649571 | A | C | MOXD1 | Nonsynonymous SNV | MOXD1:NM_015529:exon5: c.T826G:p.Trp276Gly | 29.9 |
| Chr11 | 2290998 | A | T | ASCL2 | Nonsynonymous SNV | ASCL2:NM_005170:exon1: c.T565A:p.Trp189Arg | 33.0 |
| Chr14 | 64655368 | A | C | SYNE2 | Nonsynonymous SNV | SYNE2:NM_015180:exon98: c.A17813C:p.Asn5938Thr | 25.5 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Villa, C.; Arrigoni, F.; Rivellini, E.; Lavitrano, M.; De Gioia, L.; Ferini-Strambi, L.; Combi, R. Exome Sequencing in an ADSHE Family: VUS Identification and Limits. Int. J. Environ. Res. Public Health 2022, 19, 12548. https://doi.org/10.3390/ijerph191912548
Villa C, Arrigoni F, Rivellini E, Lavitrano M, De Gioia L, Ferini-Strambi L, Combi R. Exome Sequencing in an ADSHE Family: VUS Identification and Limits. International Journal of Environmental Research and Public Health. 2022; 19(19):12548. https://doi.org/10.3390/ijerph191912548
Chicago/Turabian StyleVilla, Chiara, Federica Arrigoni, Eleonora Rivellini, Marialuisa Lavitrano, Luca De Gioia, Luigi Ferini-Strambi, and Romina Combi. 2022. "Exome Sequencing in an ADSHE Family: VUS Identification and Limits" International Journal of Environmental Research and Public Health 19, no. 19: 12548. https://doi.org/10.3390/ijerph191912548
APA StyleVilla, C., Arrigoni, F., Rivellini, E., Lavitrano, M., De Gioia, L., Ferini-Strambi, L., & Combi, R. (2022). Exome Sequencing in an ADSHE Family: VUS Identification and Limits. International Journal of Environmental Research and Public Health, 19(19), 12548. https://doi.org/10.3390/ijerph191912548








