Epidemiology of Δ8THC-Related Carcinogenesis in USA: A Panel Regression and Causal Inferential Study
Abstract
:1. Introduction
2. Methods
3. Results
3.1. Continuous Bivariate Analysis
3.2. Categorical Bivariate Analysis
3.3. Multivariable Panel Regression
3.4. Temporally Lagged Panel Models
3.4.1. Two Years Temporal Lag
3.4.2. Four Years Temporal Lag
3.4.3. Marginal Effects
4. Discussion
4.1. Main Results
4.2. Mechanisms
4.3. Mechanisms of Cannabinoid Carcinogenesis
4.4. Recent DNA Methylation Studies
4.5. Chromosomal Structural Observations
4.5.1. Shedding of Chromosomal Arms and Possible Breakage–Fusion–Bridge Cycles
4.5.2. Errors of Meiosis and Mitosis
4.5.3. Epigenomic Control of Chromosomal Centromeric Function
4.5.4. Epigenomic Impacts on DNA Breakage Sites
4.5.5. Cannabinoids Deliver Multiple Carcinogenic Insults
4.6. Causal Inference
4.7. Generalizability
4.8. Strengths and Limitations
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Babalonis, S.; Raup-Konsavage, W.M.; Akpunonu, P.D.; Balla, A.; Vrana, K.E. Δ(8)-THC: Legal Status, Widespread Availability, and Safety Concerns. Cannabis Cannabinoid Res. 2021, 6, 362–365. [Google Scholar] [CrossRef] [PubMed]
- National Library of Medicine. PubChem: Delta8-Tetrahydrocannabinol (Compound). Available online: https://pubchem.ncbi.nlm.nih.gov/compound/delta8-Tetrahydrocannabinol (accessed on 10 January 2022).
- Centers for Disease Control and Prevention. Increases in Availability of Cannabis Products Containing Delta-8 THC and Reported Cases of Adverse Events. Available online: https://emergency.cdc.gov/han/2021/han00451.asp (accessed on 10 January 2022).
- U.S. Food and Drug Administration. 5 Things to Know about Delta-8 Tetrahydrocannabinol—Delta-8 THC. Available online: https://www.fda.gov/consumers/consumer-updates/5-things-know-about-delta-8-tetrahydrocannabinol-delta-8-thc (accessed on 10 January 2022).
- Ross, R.A.; Gibson, T.M.; Stevenson, L.A.; Saha, B.; Crocker, P.; Razdan, R.K.; Pertwee, R.G. Structural determinants of the partial agonist-inverse agonist properties of 6′-azidohex-2′-yne-delta8-tetrahydrocannabinol at cannabinoid receptors. Br. J. Pharmacol. 1999, 128, 735–743. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Thompson, G.R.; Rosenkrantz, H.; Schaeppi, U.H.; Braude, M.C. Comparison of acute oral toxicity of cannabinoids in rats, dogs and monkeys. Toxicol. Appl. Pharmacol. 1973, 25, 363–372. [Google Scholar] [CrossRef]
- Leuchtenberger, C.; Leuchtenberger, R. Morphological and cytochemical effects of marijuana cigarette smoke on epithelioid cells of lung explants from mice. Nature 1971, 234, 227–229. [Google Scholar] [CrossRef]
- Leuchtenberger, C.; Leuchtenberger, R.; Schneider, A. Effects of marijuana and tobacco smoke on human lung physiology. Nature 1973, 241, 137–139. [Google Scholar] [CrossRef]
- Stenchever, M.A.; Kunysz, T.J.; Allen, M.A. Chromosome breakage in users of marihuana. Am. J. Obstet. Gynecol. 1974, 118, 106–113. [Google Scholar] [CrossRef]
- Zimmerman, A.M.; Zimmerman, S.; Raj, A.Y. Effects of Cannabinoids on Spermatogensis in Mice. In Marijuana and Medicine, 1st ed.; Nahas, G.G., Sutin, K.M., Harvey, D.J., Agurell, S., Eds.; Humana Press: Totowa, NJ, USA, 1999; Volume 1, pp. 347–358. [Google Scholar]
- Morishima, A.; Henrich, R.T.; Jayaraman, J.; Nahas, G.G. Hypoploid metaphases in cultured lymphocytes of marihuana smokers. Adv. Biosci. 1978, 22–23, 371–376. [Google Scholar]
- Henrich, R.T.; Nogawa, T.; Morishima, A. In vitro induction of segregational errors of chromosomes by natural cannabinoids in normal human lymphocytes. Environ. Mutagenesis 1980, 2, 139–147. [Google Scholar] [CrossRef]
- Russo, C.; Ferk, F.; Mišík, M.; Ropek, N.; Nersesyan, A.; Mejri, D.; Holzmann, K.; Lavorgna, M.; Isidori, M.; Knasmüller, S. Low doses of widely consumed cannabinoids (cannabidiol and cannabidivarin) cause DNA damage and chromosomal aberrations in human-derived cells. Arch. Toxicol. 2019, 93, 179–188. [Google Scholar] [CrossRef] [Green Version]
- Szutorisz, H.; DiNieri, J.A.; Sweet, E.; Egervari, G.; Michaelides, M.; Carter, J.M.; Ren, Y.; Miller, M.L.; Blitzer, R.D.; Hurd, Y.L. Parental THC exposure leads to compulsive heroin-seeking and altered striatal synaptic plasticity in the subsequent generation. Neuropsychopharmacology 2014, 39, 1315–1323. [Google Scholar] [CrossRef] [Green Version]
- Szutorisz, H.; Hurd, Y.L. Epigenetic Effects of Cannabis Exposure. Biol. Psychiatry 2016, 79, 586–594. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Watson, C.T.; Szutorisz, H.; Garg, P.; Martin, Q.; Landry, J.A.; Sharp, A.J.; Hurd, Y.L. Genome-Wide DNA Methylation Profiling Reveals Epigenetic Changes in the Rat Nucleus Accumbens Associated with Cross-Generational Effects of Adolescent THC Exposure. Neuropsychopharmacology 2015, 40, 2993–3005. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bartova, A.; Birmingham, M.K. Effect of delta9-tetrahydrocannabinol on mitochondrial NADH-oxidase activity. J. Biol. Chem. 1976, 251, 5002–5006. [Google Scholar] [CrossRef]
- Benard, G.; Massa, F.; Puente, N.; Lourenco, J.; Bellocchio, L.; Soria-Gomez, E.; Matias, I.; Delamarre, A.; Metna-Laurent, M.; Cannich, A.; et al. Mitochondrial CB(1) receptors regulate neuronal energy metabolism. Nat. Neurosci. 2012, 15, 558–564. [Google Scholar] [CrossRef] [PubMed]
- Hebert-Chatelain, E.; Desprez, T.; Serrat, R.; Bellocchio, L.; Soria-Gomez, E.; Busquets-Garcia, A.; Pagano Zottola, A.C.; Delamarre, A.; Cannich, A.; Vincent, P.; et al. A cannabinoid link between mitochondria and memory. Nature 2016, 539, 555–559. [Google Scholar] [CrossRef]
- Hebert-Chatelain, E.; Reguero, L.; Puente, N.; Lutz, B.; Chaouloff, F.; Rossignol, R.; Piazza, P.V.; Benard, G.; Grandes, P.; Marsicano, G. Cannabinoid control of brain bioenergetics: Exploring the subcellular localization of the CB1 receptor. Mol. Metab. 2014, 3, 495–504. [Google Scholar] [CrossRef]
- Mahoney, J.M.; Harris, R.A. Effect of 9-tetrahydrocannabinol on mitochondrial processes. Biochem. Pharmacol. 1972, 21, 1217–1226. [Google Scholar] [CrossRef]
- Callaghan, R.C.; Allebeck, P.; Akre, O.; McGlynn, K.A.; Sidorchuk, A. Cannabis Use and Incidence of Testicular Cancer: A 42-Year Follow-up of Swedish Men between 1970 and 2011. Cancer Epidemiol. Biomark. Prev. 2017, 26, 1644–1652. [Google Scholar] [CrossRef] [Green Version]
- Daling, J.R.; Doody, D.R.; Sun, X.; Trabert, B.L.; Weiss, N.S.; Chen, C.; Biggs, M.L.; Starr, J.R.; Dey, S.K.; Schwartz, S.M. Association of marijuana use and the incidence of testicular germ cell tumors. Cancer 2009, 115, 1215–1223. [Google Scholar] [CrossRef] [Green Version]
- Lacson, J.C.; Carroll, J.D.; Tuazon, E.; Castelao, E.J.; Bernstein, L.; Cortessis, V.K. Population-based case-control study of recreational drug use and testis cancer risk confirms an association between marijuana use and nonseminoma risk. Cancer 2012, 118, 5374–5383. [Google Scholar] [CrossRef] [Green Version]
- Trabert, B.; Sigurdson, A.J.; Sweeney, A.M.; Strom, S.S.; McGlynn, K.A. Marijuana use and testicular germ cell tumors. Cancer 2011, 117, 848–853. [Google Scholar] [CrossRef] [PubMed]
- Reece, A.S. Chronic toxicology of cannabis. Clin. Toxicol. 2009, 47, 517–524. [Google Scholar] [CrossRef] [PubMed]
- Reece, A.S.; Hulse, G.K. Epidemiological Overview of Multidimensional Chromosomal and Genome Toxicity of Cannabis Exposure in Congenital Anomalies and Cancer Development. Sci. Rep. 2021, 11, 13892–13912. [Google Scholar] [CrossRef] [PubMed]
- Reece, A.S.; Hulse, G.K. Cannabinoid exposure as a major driver of pediatric acute lymphoid Leukaemia rates across the USA: Combined geospatial, multiple imputation and causal inference study. BMC Cancer 2021, 21, 984–1017. [Google Scholar] [CrossRef]
- Grobner, S.N.; Worst, B.C.; Weischenfeldt, J.; Buchhalter, I.; Kleinheinz, K.; Rudneva, V.A.; Johann, P.D.; Balasubramanian, G.P.; Segura-Wang, M.; Brabetz, S.; et al. The landscape of genomic alterations across childhood cancers. Nature 2018, 555, 321–327. [Google Scholar] [CrossRef] [Green Version]
- Ma, X.; Liu, Y.; Liu, Y.; Alexandrov, L.B.; Edmonson, M.N.; Gawad, C.; Zhou, X.; Li, Y.; Rusch, M.C.; Easton, J.; et al. Pan-cancer genome and transcriptome analyses of 1699 paediatric leukaemias and solid tumours. Nature 2018, 555, 371–376. [Google Scholar] [CrossRef]
- Nahas, G.G.; Morishima, A.; Desoize, B. Effects of cannabinoids on macromolecular synthesis and replication of cultured lymphocytes. Fed. Proc. 1977, 36, 1748–1752. [Google Scholar]
- McClean, D.K.; Zimmerman, A.M. Action of delta 9-tetrahydrocannabinol on cell division and macromolecular synthesis in division-synchronized protozoa. Pharmacology 1976, 14, 307–321. [Google Scholar] [CrossRef]
- Zimmerman, A.M.; Stich, H.; San, R. Nonmutagenic action of cannabinoids in vitro. Pharmacology 1978, 16, 333–343. [Google Scholar] [CrossRef]
- Thomas, J.; Tilak, S.; Zimmerman, S.; Zimmerman, A.M. Action of delta 9-tetrahydrocannabinol on the pool of acid soluble nucleotides. Cytobios 1984, 40, 71–85. [Google Scholar]
- Tahir, S.K.; Zimmerman, A.M. Influence of marihuana on cellular structures and biochemical activities. Pharmacol. Biochem. Behav. 1991, 40, 617–623. [Google Scholar] [CrossRef]
- Carchman, R.A.; Harris, L.S.; Munson, A.E. The inhibition of DNA synthesis by cannabinoids. Cancer Res. 1976, 36, 95–100. [Google Scholar] [PubMed]
- Friedman, M.A. In vivo effects of cannabinoids on macromolecular biosynthesis in Lewis lung carcinomas. Cancer Biochem. Biophys. 1977, 2, 51–54. [Google Scholar] [PubMed]
- Tucker, A.N.; Friedman, M.A. Effects of cannabinoids on L1210 murine leukemia. 1. Inhibition of DNA synthesis. Res. Commun. Chem. Pathol. Pharmacol. 1977, 17, 703–714. [Google Scholar]
- Krishnamurthy, M.; Gurley, S.; Moore, B.M., 2nd. Exploring the substituent effects on a novel series of C1′-dimethyl-aryl Delta8-tetrahydrocannabinol analogs. Bioorganic Med. Chem. 2008, 16, 6489–6500. [Google Scholar] [CrossRef]
- Semlali, A.; Beji, S.; Ajala, I.; Rouabhia, M. Effects of tetrahydrocannabinols on human oral cancer cell proliferation, apoptosis, autophagy, oxidative stress, and DNA damage. Arch. Oral Biol. 2021, 129, 105200. [Google Scholar] [CrossRef]
- Nahas, G.G. Cannabis Physiopathology Epidemiology Detection; CRC Press Revivals: Boca Raton, FL, USA, 1990; Volume 1. [Google Scholar]
- Nahas, G.G. Keep off the Grass; Eriksson, P.S., Ed.; Elsevier: Middlebury, VT, USA, 1990; Volume 1. [Google Scholar]
- Huang, H.F.S.; Nahas, G.G.; Hembree, W.C. Effects of Marijuana Inhalation on Spermatogenesis of the Rat. In Marijuana in Medicine; Nahas, G.G., Sutin, K.M., Harvey, D.J., Agurell, S., Eds.; Human Press: Totowa, NJ, USA, 1999; Volume 1, pp. 359–366. [Google Scholar]
- Reece, A.S.; Hulse, G.K. Cannabis Teratology Explains Current Patterns of Coloradan Congenital Defects: The Contribution of Increased Cannabinoid Exposure to Rising Teratological Trends. Clin. Pediatrics 2019, 58, 1085–1123. [Google Scholar] [CrossRef]
- Reece, A.S.; Hulse, G.K. Cannabis in Pregnancy—Rejoinder, Exposition and Cautionary Tales. Psychiatr. Times 2020, 37. [Google Scholar]
- Reece, A.S.; Hulse, G.K. Broad Spectrum epidemiological contribution of cannabis and other substances to the teratological profile of northern New South Wales: Geospatial and causal inference analysis. BMC Pharmacol. Toxicol. 2020, 21, 75–103. [Google Scholar] [CrossRef]
- Reece, A.S.; Hulse, G.K. Canadian Cannabis Consumption and Patterns of Congenital Anomalies: An Ecological Geospatial Analysis. J. Addict. Med. 2020, 14, e195–e210. [Google Scholar] [CrossRef]
- Lu, Y.; Brommer, B.; Tian, X.; Krishnan, A.; Meer, M.; Wang, C.; Vera, D.L.; Zeng, Q.; Yu, D.; Bonkowski, M.S.; et al. Reprogramming to recover youthful epigenetic information and restore vision. Nature 2020, 588, 124–129. [Google Scholar] [CrossRef] [PubMed]
- National Program of Cancer Registries and Surveillance, Epidemiology, and End Results SEER*Stat Database: NPCR and SEER Incidence—U.S. Cancer Statistics Public Use Research Database, 2019 Submission (2001–2017), United States Department of Health and Human Services, Centers for Disease Control and Prevention and National Cancer Institute. Released June 2020. Available online: www.cdc.gov/cancer/public-use (accessed on 10 January 2022).
- National Survey of Drug Use and Health 2018, NSDUH. Available online: https://www.samhsa.gov/data/all-reports (accessed on 10 January 2022).
- Tidycensus: Load US Census Boundary and Attribute Data as ‘Tidyverse’ and ‘sf’-Ready Data Frames. Available online: https://www.r-pkg.org/pkg/tidycensus (accessed on 10 January 2022).
- ElSohly, M.A.; Mehmedic, Z.; Foster, S.; Gon, C.; Chandra, S.; Church, J.C. Changes in Cannabis Potency over the Last 2 Decades (1995–2014): Analysis of Current Data in the United States. Biol. Psychiatry 2016, 79, 613–619. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chandra, S.; Radwan, M.M.; Majumdar, C.G.; Church, J.C.; Freeman, T.P.; ElSohly, M.A. New trends in cannabis potency in USA and Europe during the last decade (2008–2017). Eur. Arch. Psychiatry Clin. Neurosci. 2019, 269, 5–15. [Google Scholar] [CrossRef] [PubMed]
- ElSohly, M.A.; Ross, S.A.; Mehmedic, Z.; Arafat, R.; Yi, B.; Banahan, B.F., 3rd. Potency trends of delta9-THC and other cannabinoids in confiscated marijuana from 1980–1997. J. Forensic Sci. 2000, 45, 24–30. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wickham, H.; Averick, M.; Bryan, J.; Chang, W.; McGowan, L.D.; Francios, R.; Groelmund, G.; Hayes, A.; Henry, L.; Hester, J.; et al. Welcome to the Tidyverse. J. Open Source Softw. 2019, 4, 1686–1691. [Google Scholar] [CrossRef]
- Pebesma, E. Simple Features for R: Standardized Support for Spatial Vector Data. R J. 2018, 10, 439–446. [Google Scholar] [CrossRef] [Green Version]
- Warnes, G.R.; Bolker, B.; Bonebakker, L.; Gentleman, R.; Huber, W.; Liaw, A.; Lumley, T.; Maechler, M.; Magnusson, A.; Moeller, S.; et al. gplots: Various R Programming Tools for Plotting Data; R Package Version 3.1.1; R Foundation: Vienna, Austria, 2020. [Google Scholar]
- Stevenson, M.; Sergeant, E.; Nunes, T.; Heuer, C.; Marshall, J.; Sanchez, J.; Thornton, R.; Reiczigel, J.; Robison-Cox, J.; Sebastiani, P.; et al. epiR: Tools for the Analysis of Epidemiological Data. Available online: https://fvas.unimelb.edu.au/research/groups/veterinary-epidemiology-melbourne (accessed on 10 January 2022).
- Croissant, Y.; Millo, G.; Tappe, K.; Toomet, O.; Kleiber, C.; Zeileis, A.; Henningsen, A.; Andronic, L.; Schoenfelder, N. Package ‘plm’. Available online: https://cran.r-project.org/web/packages/plm/plm.pdf (accessed on 10 January 2022).
- Wal, W.; Geskus, R. ipw: An R Package for Inverse Probability Weighting. J. Stat. Softw. 2011, 43, 1–23. [Google Scholar] [CrossRef] [Green Version]
- VanderWeele, T.J.; Ding, P. Sensitivity Analysis in Observational Research: Introducing the E-Value. Ann. Intern. Med. 2017, 167, 268–274. [Google Scholar] [CrossRef]
- VanderWeele, T.J.; Mathur, M.B. Commentary: Developing best-practice guidelines for the reporting of E-values. Int. J. Epidemiol. 2020, 49, 1495–1497. [Google Scholar] [CrossRef]
- Mathur, M. Package ‘EValue’. Available online: https://cran.r-project.org/web/packages/EValue/index.html (accessed on 10 January 2022).
- VanderWeele, T.J.; Ding, P.; Mathur, M. Technical Considerations in the Use of the E-Value. J. Causal Inference 2019, 7, 1–11. [Google Scholar] [CrossRef] [Green Version]
- Pearl, J.; Mackaenzie, D. The Book of Why: The New Science of Cause and Effect; Basic Books: New York, NY, USA, 2019; Volume 1. [Google Scholar]
- Leeper, T.J. Margins: Marginal Effects for Model Objects; R Package Version 0.3.26; R Foundation: Vienna, Austria, 2021; pp. 1–36. [Google Scholar]
- Robinson, D.; Hayes, A.; Couch, S. Broom: Convert Statistical Objects into Tidy Tibbles. Available online: https://CRAN.R-project.org/package=broom (accessed on 10 January 2022).
- Hill, A.B. The Environment and Disease: Association or Causation? Proc. R Soc. Med. 1965, 58, 295–300. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jakubovič, A.; McGeer, P.L.; Fitzsimmons, R.C. Effects of Δ9-tetrahydrocannabinol and ethanol on body weight protein and nucleic acid synthesis in chick embryos. J. Toxicol. Environ. Health 1976, 1, 441–447. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Yuan, W.; Li, M.D. Genes and pathways co-associated with the exposure to multiple drugs of abuse, including alcohol, amphetamine/methamphetamine, cocaine, marijuana, morphine, and/or nicotine: A review of proteomics analyses. Mol. Neurobiol. 2011, 44, 269–286. [Google Scholar] [CrossRef] [PubMed]
- Murphy, S.K.; Itchon-Ramos, N.; Visco, Z.; Huang, Z.; Grenier, C.; Schrott, R.; Acharya, K.; Boudreau, M.H.; Price, T.M.; Raburn, D.J.; et al. Cannabinoid exposure and altered DNA methylation in rat and human sperm. Epigenetics 2018, 13, 1208–1221. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- DiNieri, J.A.; Wang, X.; Szutorisz, H.; Spano, S.M.; Kaur, J.; Casaccia, P.; Dow-Edwards, D.; Hurd, Y.L. Maternal cannabis use alters ventral striatal dopamine D2 gene regulation in the offspring. Biol. Psychiatry 2011, 70, 763–769. [Google Scholar] [CrossRef] [Green Version]
- Levin, E.D.; Hawkey, A.B.; Hall, B.J.; Cauley, M.; Slade, S.; Yazdani, E.; Kenou, B.; White, H.; Wells, C.; Rezvani, A.H.; et al. Paternal THC exposure in rats causes long-lasting neurobehavioral effects in the offspring. Neurotoxicol. Teratol. 2019, 74, 106806. [Google Scholar] [CrossRef]
- Holloway, Z.R.; Hawkey, A.B.; Pippin, E.; White, H.; Wells, C.; Kenou, B.; Rezvani, A.H.; Murphy, S.K.; Levin, E.D. Paternal factors in neurodevelopmental toxicology: THC exposure of male rats causes long-lasting neurobehavioral effects in their offspring. Neurotoxicology 2020, 78, 57–63. [Google Scholar] [CrossRef]
- Schrott, R.; Murphy, S.K.; Modliszewski, J.L.; King, D.E.; Hill, B.; Itchon-Ramos, N.; Raburn, D.; Price, T.; Levin, E.D.; Vandrey, R.; et al. Refraining from use diminishes cannabis-associated epigenetic changes in human sperm. Environ. Epigenetics 2021, 7, dvab009. [Google Scholar] [CrossRef]
- Shen, H.; Shih, J.; Hollern, D.P.; Wang, L.; Bowlby, R.; Tickoo, S.K.; Thorsson, V.; Mungall, A.J.; Newton, Y.; Hegde, A.M.; et al. Integrated Molecular Characterization of Testicular Germ Cell Tumors. Cell Rep. 2018, 23, 3392–3406. [Google Scholar] [CrossRef]
- Malouf, C.; Ottersbach, K. Molecular processes involved in B cell acute lymphoblastic leukaemia. Cell Mol. Life Sci. 2018, 75, 417–446. [Google Scholar] [CrossRef] [Green Version]
- Reece, A.S.; Hulse, G.K. Geotemporospatial and Causal Inferential Epidemiological Overview and Survey of USA Cannabis, Cannabidiol and Cannabinoid Genotoxicity Expressed in Cancer Incidence 2003–2017: Part 1—Continuous Bivariate Analysis. Arch. Public Health 2022, 80, 99–133. [Google Scholar] [CrossRef] [PubMed]
- Reece, A.S.; Hulse, G.K. Geotemporospatial and Causal Inferential Epidemiological Overview and Survey of USA Cannabis, Cannabidiol and Cannabinoid Genotoxicity Expressed in Cancer Incidence 2003–2017: Part 2—Categorical Bivariate Analysis and Attributable Fractions. Arch. Public Health 2022, 80, 100–135. [Google Scholar] [CrossRef] [PubMed]
- Reece, A.S.; Hulse, G.K. Geotemporospatial and Causal Inferential Epidemiological Overview and Survey of USA Cannabis, Cannabidiol and Cannabinoid Genotoxicity Expressed in Cancer Incidence 2003–2017: Part 3—Spatiotemporal, Multivariable and Causal Inferential Pathfinding and Exploratory Analyses of Prostate and Ovarian Cancers. Arch. Public Health 2022, 80, 100–136. [Google Scholar] [CrossRef] [PubMed]
- Reece, A.S.; Hulse, G.K. Geotemporospatial and causal inference epidemiological analysis of US survey and overview of cannabis, cannabidiol and cannabinoid genotoxicity in relation to congenital anomalies 2001–2015. BMC Pediatrics 2022, 22, 47–124. [Google Scholar] [CrossRef] [PubMed]
- Szutorisz, H.; Hurd, Y.L. High times for cannabis: Epigenetic imprint and its legacy on brain and behavior. Neurosci. Biobehav. Rev. 2018, 85, 93–101. [Google Scholar] [CrossRef] [PubMed]
- Ellis, R.J.; Bara, A.; Vargas, C.A.; Frick, A.L.; Loh, E.; Landry, J.; Uzamere, T.O.; Callens, J.E.; Martin, Q.; Rajarajan, P.; et al. Prenatal Δ(9)-Tetrahydrocannabinol Exposure in Males Leads to Motivational Disturbances Related to Striatal Epigenetic Dysregulation. Biol. Psychiatry 2021, in press. [CrossRef]
- Schrott, R.; Acharya, K.; Itchon-Ramos, N.; Hawkey, A.B.; Pippen, E.; Mitchell, J.T.; Kollins, S.H.; Levin, E.D.; Murphy, S.K. Cannabis use is associated with potentially heritable widespread changes in autism candidate gene DLGAP2 DNA methylation in sperm. Epigenetics 2020, 15, 161–173. [Google Scholar] [CrossRef] [Green Version]
- Zimmerman, A.M.; Raj, A.Y. Influence of cannabinoids on somatic cells in vivo. Pharmacology 1980, 21, 277–287. [Google Scholar] [CrossRef]
- Zimmerman, A.M.; Zimmerman, S. Cytogenetic Studies of Cannabinoid Effects. In Genetic and Perinatal Effects of Abused Substances; Braude, M.C., Zimmerman, A.M., Eds.; Academic Press Inc., Harcourt, Brace Jovanovich: New York, NY, USA, 1987; Volume 1, pp. 95–112. [Google Scholar]
- Zimmerman, S.; Zimmerman, A.M. Genetic effects of marijuana. Int. J. Addict. 1990, 25, 19–33. [Google Scholar] [CrossRef]
- Tahir, S.K.; Trogadis, J.E.; Stevens, J.K.; Zimmerman, A.M. Cytoskeletal organization following cannabinoid treatment in undifferentiated and differentiated PC12 cells. Biochem. Cell Biol. 1992, 70, 1159–1173. [Google Scholar] [CrossRef]
- Parker, S.J.; Zuckerman, B.S.; Zimmermann, A.M. The Effects of Maternal Marijuana Use during Pregnancy on Fetal Growth. In Marijuana in Medicine; Nahas, G.G., Sutin, K.M., Harvey, D.J., Agurell, S., Eds.; Humana Press: Totowa, NJ, USA, 1999; Volume 1, pp. 461–468. [Google Scholar]
- Cozens, D.D.; Clark, R.; Palmer, A.K.; Hardy, N.; Nahas, G.G.; Harvey, D.J. The effect of a crude marihuana extract on embryonic and foetal development of the rabbit. Adv. Biosci. 1978, 22–23, 469–477. [Google Scholar]
- Cozens, D.D.; Nahas, G.G.; Harvey, D. Prenatal Exposure to Cannabis and Fetal Development. In Marijuana in Medicine; Nahas, G.G., Sutin, K.M., Harvey, D.J., Agurell, S., Eds.; Humana Press: Totowa, NJ, USA, 1999; Volume 1, pp. 431–440. [Google Scholar]
- Khwaja, A.; Bjorkholm, M.; Gale, R.E.; Levine, R.L.; Jordan, C.T.; Ehninger, G.; Bloomfield, C.D.; Estey, E.; Burnett, A.; Cornelissen, J.J.; et al. Acute myeloid leukaemia. Nat. Rev. Dis. Primers 2016, 2, 16010. [Google Scholar] [CrossRef] [PubMed]
- Gurney, J.; Shaw, C.; Stanley, J.; Signal, V.; Sarfati, D. Cannabis exposure and risk of testicular cancer: A systematic review and meta-analysis. BMC Cancer 2015, 15, 897–906. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Reece, A.S.; Hulse, G.K. Causal inference multiple imputation investigation of the impact of cannabinoids and other substances on ethnic differentials in US testicular cancer incidence. BMC Pharmacol. Toxicol. 2021, 22, 40–71. [Google Scholar] [CrossRef]
- Reece, A.S.; Hulse, G.K. Cannabinoid- and Substance-Relationships of European Congenital Anomaly Patterns: A Space-Time Panel Regression and Causal Inferential Study. Environ. Epigenetics 2022, 8, dvab015. [Google Scholar] [CrossRef]
- Reece, A.S.; Hulse, G.K. Cannabis Genotoxicity and Cancer Incidence: A Highly Concordant Synthesis of European and USA Datasets. In Cannabis, Cannabinoids and Endocannabinoids; Preedy, V., Patel, V., Eds.; Elsevier: London, UK, 2022; Volume 1, in press. [Google Scholar]
- Reece, A.S.; Hulse, G.K. Cannabinoid Genotoxicity and Congenital Anomalies: A Convergent Synthesis of European and USA Datasets. In Cannabis, Cannabinoids and Endocannabinoids; Preedy, V., Patel, V., Eds.; Elsevier: London, UK, 2022; Volume 1, in press. [Google Scholar]
- Morishima, A. Effects of cannabis and natural cannabinoids on chromosomes and ova. NIDA Res. Monogr. 1984, 44, 25–45. [Google Scholar]
- McClintock, B. The Production of Homozygous Deficient Tissues with Mutant Characteristics by Means of the Aberrant Mitotic Behavior of Ring-Shaped Chromosomes. Genetics 1938, 23, 315–376. [Google Scholar] [CrossRef]
- McClintock, B. The Stability of Broken Ends of Chromosomes in Zea Mays. Genetics 1941, 26, 234–282. [Google Scholar] [CrossRef]
- Forrester, M.B.; Merz, R.D. Risk of selected birth defects with prenatal illicit drug use, Hawaii, 1986–2002. J. Toxicol. Environ. Health 2007, 70, 7–18. [Google Scholar] [CrossRef]
- Leuchtenberger, C.; Leuchtenberger, R.; Ritter, U.; Inui, N. Effects of marijuana and tobacco smoke on DNA and chromosomal complement in human lung explants. Nature 1973, 242, 403–404. [Google Scholar] [CrossRef]
- Hall, W.; Degenhardt, L. Adverse health effects of non-medical cannabis use. Lancet 2009, 374, 1383–1391. [Google Scholar] [CrossRef]
- Reece, A.S.; Hulse, G.K. Chromothripsis and epigenomics complete causality criteria for cannabis- and addiction-connected carcinogenicity, congenital toxicity and heritable genotoxicity. Mutat. Res. 2016, 789, 15–25. [Google Scholar] [CrossRef] [PubMed]
- Zhang, C.Z.; Spektor, A.; Cornils, H.; Francis, J.M.; Jackson, E.K.; Liu, S.; Meyerson, M.; Pellman, D. Chromothripsis from DNA damage in micronuclei. Nature 2015, 522, 179–184. [Google Scholar] [CrossRef] [Green Version]
- Kuhn, J.; Dumont, S. Mammalian kinetochores count attached microtubules in a sensitive and switch-like manner. J. Cell Biol. 2019, 218, 3583–3596. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Beh, T.T.; Kalitsis, P. Centromeres and Kinetochores. In Centromeres and Kinetochores; Black, B.E., Ed.; Springer: Philadelphia, PA, USA, 2017. [Google Scholar]
- Beh, T.T.; Kalitsis, P. The Role of Centromere Defects in Cancer. In Centromeres and Kinetochores; Black, B.E., Ed.; Springer: Philadelphia, PA, USA, 2017; Volume 1, pp. 1–554. [Google Scholar]
- Black, B.E. Preface to: Centromeres and Kinetochores. In Centromeres and Kinetochores; Black, B.E., Ed.; Springer: Cham, Switzerland, 2017; Volume 1, pp. v–viii. [Google Scholar]
- Corbett, K.D. Molecular Mechanisms of Spindle Assembly Checkpoint Activation and Silencing. In Centromeres and Kinetochores; Black, B.E., Ed.; Springer: Philadelphia, PA, USA, 2017; Volume 1, pp. 1–554. [Google Scholar]
- French, B.T.; Straight, A.F. The Power of Xenopus Egg Extract for Reconstitution of Centromere and Kinetochore Function. In Centromeres and Kinetochores; Black, B.E., Ed.; Springer: Philadelphia, PA, USA, 2017; Volume 1, pp. 1–554. [Google Scholar]
- Hara, M.; Fukagawa, T. Critical Foundation of the Kinetochore: The Constitutive Centromere—Associated Network (CCAN). In Centromeres and Kinetochores; Black, B.E., Ed.; Springer: Philadelphia, PA, USA, 2017; Volume 1, pp. 1–554. [Google Scholar]
- Dong, Q.; Liu, X.L.; Wang, X.H.; Zhao, Y.; Chen, Y.H.; Li, F. Ccp1-Ndc80 switch at the N terminus of CENP-T regulates kinetochore assembly. Proc. Natl. Acad. Sci. USA 2021, 118, e2104459118. [Google Scholar] [CrossRef] [PubMed]
- Mellone, B.G.; Fachinetti, D. Diverse mechanisms of centromere specification. Curr. Biol. 2021, 31, R1491–R1504. [Google Scholar] [CrossRef]
- Ryu, H.Y.; Hochstrasser, M. Histone sumoylation and chromatin dynamics. Nucleic Acids Res. 2021, 49, 6043–6052. [Google Scholar] [CrossRef]
- Gowran, A.; Murphy, C.E.; Campbell, V.A. Delta(9)-tetrahydrocannabinol regulates the p53 post-translational modifiers Murine double minute 2 and the Small Ubiquitin MOdifier protein in the rat brain. FEBS Lett. 2009, 583, 3412–3418. [Google Scholar] [CrossRef] [Green Version]
- Attia, S.M.; Al-Khalifa, M.K.; Al-Hamamah, M.A.; Alotaibi, M.R.; Attia, M.S.M.; Ahmad, S.F.; Ansari, M.A.; Nadeem, A.; Bakheet, S.A. Vorinostat is genotoxic and epigenotoxic in the mouse bone marrow cells at the human equivalent doses. Toxicology 2020, 441, 152507. [Google Scholar] [CrossRef]
- Ben Yamin, B.; Ahmed-Seghir, S.; Tomida, J.; Despras, E.; Pouvelle, C.; Yurchenko, A.; Goulas, J.; Corre, R.; Delacour, Q.; Droin, N.; et al. DNA polymerase zeta contributes to heterochromatin replication to prevent genome instability. EMBO J. 2021, 40, e104543. [Google Scholar] [CrossRef]
- Cheloshkina, K.; Poptsova, M. Comprehensive analysis of cancer breakpoints reveals signatures of genetic and epigenetic contribution to cancer genome rearrangements. PLoS Comput. Biol. 2021, 17, e1008749. [Google Scholar] [CrossRef] [PubMed]
- García Fernández, F.; Lemos, B.; Khalil, Y.; Batrin, R.; Haber, J.E.; Fabre, E. Modified chromosome structure caused by phosphomimetic H2A modulates the DNA damage response by increasing chromatin mobility in yeast. J. Cell Sci. 2021, 134, jcs258500. [Google Scholar] [CrossRef] [PubMed]
- George, J.; Lim, J.S.; Jang, S.J.; Cun, Y.; Ozretić, L.; Kong, G.; Leenders, F.; Lu, X.; Fernández-Cuesta, L.; Bosco, G.; et al. Comprehensive genomic profiles of small cell lung cancer. Nature 2015, 524, 47–53. [Google Scholar] [CrossRef] [PubMed]
- Harrod, A.; Lane, K.A.; Downs, J.A. The role of the SWI/SNF chromatin remodelling complex in the response to DNA double strand breaks. DNA Repair 2020, 93, 102919. [Google Scholar] [CrossRef] [PubMed]
- He, Y.; Ren, J.; Xu, X.; Ni, K.; Schwader, A.; Finney, R.; Wang, C.; Sun, L.; Klarmann, K.; Keller, J.; et al. Lsh/HELLS is required for B lymphocyte development and immunoglobulin class switch recombination. Proc. Natl. Acad. Sci. USA 2020, 117, 20100–20108. [Google Scholar] [CrossRef]
- Jamsen, J.A.; Sassa, A.; Perera, L.; Shock, D.D.; Beard, W.A.; Wilson, S.H. Structural basis for proficient oxidized ribonucleotide insertion in double strand break repair. Nat. Commun. 2021, 12, 5055. [Google Scholar] [CrossRef]
- Llorens-Agost, M.; Ensminger, M.; Le, H.P.; Gawai, A.; Liu, J.; Cruz-García, A.; Bhetawal, S.; Wood, R.D.; Heyer, W.D.; Löbrich, M. POLθ-mediated end joining is restricted by RAD52 and BRCA2 until the onset of mitosis. Nat. Cell Biol. 2021, 23, 1095–1104. [Google Scholar] [CrossRef]
- Mir, U.S.; Bhat, A.; Mushtaq, A.; Pandita, S.; Altaf, M.; Pandita, T.K. Role of histone acetyltransferases MOF and Tip60 in genome stability. DNA Repair 2021, 107, 103205. [Google Scholar] [CrossRef]
- Shoeb, M.; Meier, H.C.S.; Antonini, J.M. Telomeres in toxicology: Occupational health. Pharmacol. Ther. 2021, 220, 107742. [Google Scholar] [CrossRef]
- Tsai, L.J.; Lopezcolorado, F.W.; Bhargava, R.; Mendez-Dorantes, C.; Jahanshir, E.; Stark, J.M. RNF8 has both KU-dependent and independent roles in chromosomal break repair. Nucleic Acids Res. 2020, 48, 6032–6052. [Google Scholar] [CrossRef]
- Vasilyeva, T.A.; Marakhonov, A.V.; Sukhanova, N.V.; Kutsev, S.I.; Zinchenko, R.A. Preferentially Paternal Origin of De Novo 11p13 Chromosome Deletions Revealed in Patients with Congenital Aniridia and WAGR Syndrome. Genes 2020, 11, 812. [Google Scholar] [CrossRef] [PubMed]
- Yamazoe, T.; Mori, T.; Yoshio, S.; Kanto, T. Hepatocyte ploidy and pathological mutations in hepatocellular carcinoma: Impact on oncogenesis and therapeutics. Glob. Health Med. 2020, 2, 273–281. [Google Scholar] [CrossRef] [PubMed]
- Ali, S.; Zhang, T.; Lambing, C.; Wang, W.; Zhang, P.; Xie, L.; Wang, J.; Khan, N.; Zhang, Q. Loss of chromatin remodeler DDM1 causes segregation distortion in Arabidopsis thaliana. Planta 2021, 254, 107. [Google Scholar] [CrossRef]
- Alonso-Ramos, P.; Álvarez-Melo, D.; Strouhalova, K.; Pascual-Silva, C.; Garside, G.B.; Arter, M.; Bermejo, T.; Grigaitis, R.; Wettstein, R.; Fernández-Díaz, M.; et al. The Cdc14 Phosphatase Controls Resolution of Recombination Intermediates and Crossover Formation during Meiosis. Int. J. Mol. Sci. 2021, 22, 9811. [Google Scholar] [CrossRef] [PubMed]
- Arrieta, M.; Macaulay, M.; Colas, I.; Schreiber, M.; Shaw, P.D.; Waugh, R.; Ramsay, L. An Induced Mutation in HvRECQL4 Increases the Overall Recombination and Restores Fertility in a Barley HvMLH3 Mutant Background. Front. Plant Sci. 2021, 12, 706560. [Google Scholar] [CrossRef] [PubMed]
- Gutiérrez Pinzón, Y.; González Kise, J.K.; Rueda, P.; Ronceret, A. The Formation of Bivalents and the Control of Plant Meiotic Recombination. Front. Plant Sci. 2021, 12, 717423. [Google Scholar] [CrossRef]
- Malinovskaya, L.P.; Tishakova, K.V.; Bikchurina, T.I.; Slobodchikova, A.Y.; Torgunakov, N.Y.; Torgasheva, A.A.; Tsepilov, Y.A.; Volkova, N.A.; Borodin, P.M. Negative heterosis for meiotic recombination rate in spermatocytes of the domestic chicken Gallus gallus. Vavilov J. Genet. Breed. 2021, 25, 661–668. [Google Scholar] [CrossRef] [PubMed]
- Maroilley, T.; Li, X.; Oldach, M.; Jean, F.; Stasiuk, S.J.; Tarailo-Graovac, M. Deciphering complex genome rearrangements in C. elegans using short-read whole genome sequencing. Sci. Rep. 2021, 11, 18258. [Google Scholar] [CrossRef] [PubMed]
- Torgasheva, A.; Malinovskaya, L.; Zadesenets, K.; Shnaider, E.; Rubtsov, N.; Borodin, P. Germline-Restricted Chromosome (GRC) in Female and Male Meiosis of the Great Tit (Parus major, Linnaeus, 1758). Front. Genet. 2021, 12, 768056. [Google Scholar] [CrossRef] [PubMed]
- Ur, S.N.; Corbett, K.D. Architecture and Dynamics of Meiotic Chromosomes. Annu. Rev. Genet. 2021, 55, 497–526. [Google Scholar] [CrossRef]
- Ajamian, L.; Melnychuk, L.; Jean-Pierre, P.; Zaharatos, G.J. DNA Vaccine-Encoded Flagellin Can Be Used as an Adjuvant Scaffold to Augment HIV-1 gp41 Membrane Proximal External Region Immunogenicity. Viruses 2018, 10, 100. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dos Santos Rocha, A.; de Amorim, I.S.S.; Simão, T.A.; da Fonseca, A.S.; Garrido, R.G.; Mencalha, A.L. High-Resolution Melting (HRM) of Hypervariable Mitochondrial DNA Regions for Forensic Science. J. Forensic Sci. 2018, 63, 536–540. [Google Scholar] [CrossRef] [PubMed]
- Krebs, S.J.; McBurney, S.P.; Kovarik, D.N.; Waddell, C.D.; Jaworski, J.P.; Sutton, W.F.; Gomes, M.M.; Trovato, M.; Waagmeester, G.; Barnett, S.J.; et al. Multimeric scaffolds displaying the HIV-1 envelope MPER induce MPER-specific antibodies and cross-neutralizing antibodies when co-immunized with gp160 DNA. PLoS ONE 2014, 9, e113463. [Google Scholar] [CrossRef]
- Li, D.; Yu, A.Q.; Li, X.J.; Zhu, Y.T.; Jin, X.K.; Li, W.W.; Wang, Q. Antimicrobial activity of a novel hypervariable immunoglobulin domain-containing receptor Dscam in Cherax quadricarinatus. Fish Shellfish Immunol. 2015, 47, 766–776. [Google Scholar] [CrossRef]
- Pogorelyy, M.V.; Minervina, A.A.; Shugay, M.; Chudakov, D.M.; Lebedev, Y.B.; Mora, T.; Walczak, A.M. Detecting T cell receptors involved in immune responses from single repertoire snapshots. PLoS Biol. 2019, 17, e3000314. [Google Scholar] [CrossRef] [Green Version]
- Richter, A.; Eggenstein, E.; Skerra, A. Anticalins: Exploiting a non-Ig scaffold with hypervariable loops for the engineering of binding proteins. FEBS Lett. 2014, 588, 213–218. [Google Scholar] [CrossRef]
- Wang, C.Y.; Fang, Y.X.; Chen, G.H.; Jia, H.J.; Zeng, S.; He, X.B.; Feng, Y.; Li, S.J.; Jin, Q.W.; Cheng, W.Y.; et al. Analysis of the CDR3 length repertoire and the diversity of T cell receptor α and β chains in swine CD4+ and CD8+ T lymphocytes. Mol. Med. Rep. 2017, 16, 75–86. [Google Scholar] [CrossRef] [Green Version]
- Al-Hamamah, M.A.; Alotaibi, M.R.; Ahmad, S.F.; Ansari, M.A.; Attia, M.S.M.; Nadeem, A.; Bakheet, S.A.; As Sobeai, H.M.; Attia, S.M. Genetic and epigenetic alterations induced by the small-molecule panobinostat: A mechanistic study at the chromosome and gene levels. DNA Repair 2019, 78, 70–80. [Google Scholar] [CrossRef]
- Bhargava, R.; Lopezcolorado, F.W.; Tsai, L.J.; Stark, J.M. The canonical non-homologous end joining factor XLF promotes chromosomal deletion rearrangements in human cells. J. Biol. Chem. 2020, 295, 125–137. [Google Scholar] [CrossRef]
- Kelso, A.A.; Lopezcolorado, F.W.; Bhargava, R.; Stark, J.M. Distinct roles of RAD52 and POLQ in chromosomal break repair and replication stress response. PLoS Genet. 2019, 15, e1008319. [Google Scholar] [CrossRef] [Green Version]
- Liu, X.; Jiang, Y.; Takata, K.I.; Nowak, B.; Liu, C.; Wood, R.D.; Hittelman, W.N.; Plunkett, W. CNDAC-Induced DNA Double-Strand Breaks Cause Aberrant Mitosis Prior to Cell Death. Mol. Cancer Ther. 2019, 18, 2283–2295. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Srinivas, N.; Rachakonda, S.; Kumar, R. Telomeres and Telomere Length: A General Overview. Cancers 2020, 12, 558. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Young, N.L.; Dere, R. Mechanistic insights into KDM4A driven genomic instability. Biochem. Soc. Trans. 2021, 49, 93–105. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Shi, Y.; Ramesh, K.H.; Naeem, R.; Wang, Y. Karyotypic complexity, TP53 pathogenic variants, and increased number of variants on Next-Generation Sequencing are associated with disease progression in a North American Adult T-Cell Leukemia/Lymphoma cohort. Int. J. Lab. Hematol. 2021, 43, 651–657. [Google Scholar] [CrossRef]
- Zhao, S.; Klattenhoff, A.W.; Thakur, M.; Sebastian, M.; Kidane, D. Mutation in DNA Polymerase Beta Causes Spontaneous Chromosomal Instability and Inflammation-Associated Carcinogenesis in Mice. Cancers 2019, 11, 1160. [Google Scholar] [CrossRef] [Green Version]
- Bernard, E.; Nannya, Y.; Hasserjian, R.P.; Devlin, S.M.; Tuechler, H.; Medina-Martinez, J.S.; Yoshizato, T.; Shiozawa, Y.; Saiki, R.; Malcovati, L.; et al. Implications of TP53 allelic state for genome stability, clinical presentation and outcomes in myelodysplastic syndromes. Nat. Med. 2020, 26, 1549–1556. [Google Scholar] [CrossRef]
- Brieghel, C.; Aarup, K.; Torp, M.H.; Andersen, M.A.; Yde, C.W.; Tian, X.; Wiestner, A.; Ahn, I.E.; Niemann, C.U. Clinical Outcomes in Patients with Multi-Hit TP53 Chronic Lymphocytic Leukemia Treated with Ibrutinib. Clin. Cancer Res. 2021, 27, 4531–4538. [Google Scholar] [CrossRef]
- Fornalski, K.W.; Dobrzyński, L. Modeling of single cell cancer transformation using phase transition theory: Application of the Avrami equation. Radiat. Environ. Biophys. 2022, 61, 169–175. [Google Scholar] [CrossRef]
- Mitchell, S.R.; Gopakumar, J.; Jaiswal, S. Insights into clonal hematopoiesis and its relation to cancer risk. Curr. Opin. Genet. Dev. 2021, 66, 63–69. [Google Scholar] [CrossRef]
- Tang, S.; Lu, Y.; Zhang, P.; Chen, D.; Liu, X.; Du, X.; Cao, J.; Ye, P.; Chen, L.; Li, S.; et al. Lenalidomide, bortezomib and dexamethasone followed by tandem-autologous stem cell transplantation is an effective treatment modality for multi-hit multiple myeloma. Leuk. Res. 2021, 110, 106710. [Google Scholar] [CrossRef]
- Reece, A.S.; Hulse, G.K. A geospatiotemporal and causal inference epidemiological exploration of substance and cannabinoid exposure as drivers of rising US pediatric cancer rates. BMC Cancer 2021, 21, 197–230. [Google Scholar] [CrossRef] [PubMed]

| Cancer | Cases in Quintile 5 | Non-Cases in Quintile 5 | Cases in Quintile 1 | Non-Cases in Quintile 1 | R.R. | R.R. (Lower C.I.) | R.R. (Upper C.I.) | A.F.E. | A.F.E. (Lower C.I.) | A.F.E. (Upper C.I.) | P.A.R. | P.A.R. (Lower C.I.) | P.A.R. (Upper C.I.) | Chi. Squared | p-Value | E-Value Estimate | E-Value (Lower C.I.) |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Melanoma | 100,207 | 384,314,333 | 115,471 | 540,452,122 | 1.2203 | 1.2100 | 1.2307 | 0.1805 | 0.1736 | 0.1874 | 0.0839 | 0.0803 | 0.0875 | 2134.601 | 0 | 1.74 | 1.71 |
| Liver | 49,147 | 384,365,393 | 62,900 | 540,504,693 | 1.0987 | 1.0859 | 1.1118 | 0.0899 | 0.0791 | 0.1005 | 0.0394 | 0.0344 | 0.0444 | 244.865 | 1.71 × 10−55 | 1.43 | 1.39 |
| Corpus_Uteri | 78,333 | 384,336,207 | 100,741 | 540,466,852 | 1.0934 | 1.0833 | 1.1037 | 0.0854 | 0.0769 | 0.0939 | 0.0374 | 0.0334 | 0.0413 | 351.830 | 8.46 × 10−79 | 1.41 | 1.38 |
| Anorectum | 10,462 | 384,404,078 | 13,537 | 540,554,056 | 1.0868 | 1.0594 | 1.1149 | 0.0799 | 0.0561 | 0.1030 | 0.0348 | 0.0240 | 0.0455 | 40.895 | 8.03 × 10−11 | 1.39 | 1.31 |
| Thyroid | 67,875 | 384,346,665 | 89,788 | 540,477,805 | 1.0630 | 1.0525 | 1.0737 | 0.0593 | 0.0499 | 0.0686 | 0.0255 | 0.0213 | 0.0297 | 144.443 | 1.42 × 10−33 | 1.32 | 1.29 |
| Gastric_Cardia | 144,970 | 384,269,570 | 193,373 | 540,374,220 | 1.0542 | 1.0471 | 1.0614 | 0.0514 | 0.0450 | 0.0579 | 0.0220 | 0.0192 | 0.0249 | 231.159 | 1.67 × 10−52 | 1.29 | 1.27 |
| AML | 23,937 | 384,390,603 | 31,708 | 540,535,885 | 1.0616 | 1.0439 | 1.0795 | 0.0580 | 0.0421 | 0.0737 | 0.0250 | 0.0179 | 0.0320 | 48.720 | 1.48 × 10−12 | 1.32 | 1.26 |
| Pancreas | 67,490 | 384,347,050 | 90,446 | 540,477,147 | 1.0493 | 1.0389 | 1.0598 | 0.0470 | 0.0374 | 0.0564 | 0.0201 | 0.0159 | 0.0242 | 89.545 | 1.50 × 10−21 | 1.28 | 1.24 |
| Postmenopausal Breast Cancer | 249,834 | 384,164,706 | 337,900 | 540,229,693 | 1.0397 | 1.0344 | 1.0451 | 0.0382 | 0.0332 | 0.0432 | 0.0162 | 0.0141 | 0.0184 | 218.017 | 1.22 × 10−49 | 1.24 | 1.22 |
| Breast | 337,544 | 384,076,996 | 462,078 | 540,105,515 | 1.0272 | 1.0227 | 1.0318 | 0.0265 | 0.0222 | 0.0308 | 0.0112 | 0.0093 | 0.0130 | 140.859 | 8.63 × 10−33 | 1.19 | 1.17 |
| Testis | 52,627 | 1,726,611,593 | 19,713 | 670,166,199 | 1.0362 | 1.0194 | 1.0533 | 0.0349 | 0.0190 | 0.0506 | 0.0254 | 0.0137 | 0.0369 | 18.136 | 1.03 × 10−5 | 1.23 | 1.16 |
| Oropharynx | 63,178 | 384,351,362 | 87,440 | 540,480,153 | 1.0160 | 1.0057 | 1.0265 | 0.0158 | 0.0057 | 0.0258 | 0.0066 | 0.0023 | 0.0109 | 9.276 | 0.0012 | 1.14 | 1.08 |
| Myeloma | 31,335 | 384,383,205 | 44,787 | 540,522,806 | 0.9838 | 0.9697 | 0.9982 | −0.0164 | −0.0312 | −0.0019 | −0.0068 | −0.0128 | −0.0008 | 4.889 | 0.0135 | 1.15 | - |
| Bladder | 103,627 | 384,310,913 | 148,415 | 540,419,178 | 0.9819 | 0.9741 | 0.9897 | −0.0185 | −0.0266 | −0.0104 | −0.0076 | −0.0109 | −0.0043 | 20.478 | 3.02 × 10−6 | 1.16 | - |
| Vulva.&.Vagina | 375,122,812 | 20,981 | 536,846,206 | 0.9803 | 0.9597 | 1.0013 | −0.0201 | −0.0420 | 0.0013 | −0.0082 | −0.0169 | 0.0005 | 3.370 | 0.0332 | 1.16 | - | |
| CML | 9482 | 373,925,516 | 13,840 | 532,699,861 | 0.9760 | 0.9509 | 1.0019 | −0.0246 | −0.0517 | 0.0019 | −0.0100 | −0.0208 | 0.0007 | 3.313 | 0.0344 | 1.18 | - |
| Penis | 2283 | 312,646,824 | 3380 | 451,235,667 | 0.9749 | 0.9244 | 1.0280 | −0.0258 | −0.0817 | 0.0272 | −0.0104 | −0.0323 | 0.0110 | 0.884 | 0.1736 | 1.19 | - |
| Esophagus | 23,815 | 382,573,677 | 34,228 | 535,139,818 | 0.9732 | 0.9573 | 0.9895 | −0.0275 | −0.0446 | −0.0106 | −0.0113 | −0.0182 | −0.0044 | 10.330 | 6.54 × 10−4 | 1.20 | - |
| ALL | 6865 | 344,103,689 | 9149 | 445,794,343 | 0.9721 | 0.9422 | 1.0030 | −0.0287 | −0.0614 | 0.0030 | −0.0123 | −0.0260 | 0.0012 | 3.140 | 0.0382 | 1.20 | - |
| Brain | 29,472 | 384,385,068 | 42,767 | 540,524,826 | 0.9691 | 0.9548 | 0.9835 | −0.0319 | −0.0474 | −0.0167 | −0.0130 | −0.0192 | −0.0069 | 17.236 | 1.65 × 10−5 | 1.21 | - |
| Kidney | 82,655 | 384,331,885 | 120,241 | 540,447,352 | 0.9666 | 0.9581 | 0.9752 | −0.0345 | −0.0437 | −0.0254 | −0.0141 | −0.0177 | −0.0104 | 56.391 | 2.97 × 10−14 | 1.22 | - |
| NH_Lymphoma | 93,104 | 384,321,436 | 137,276 | 540,430,317 | 0.9537 | 0.9458 | 0.9617 | −0.0485 | −0.0573 | −0.0398 | −0.0196 | −0.0230 | −0.0162 | 124.585 | 3.14 × 10−29 | 1.27 | - |
| Gall_Bladder | 14,123 | 370,777,042 | 21,633 | 534,521,894 | 0.9412 | 0.9214 | 0.9613 | −0.0625 | −0.0853 | −0.0402 | −0.0247 | −0.0333 | −0.0161 | 31.433 | 1.03 × 10−8 | 1.32 | - |
| Stomach | 32,140 | 384,382,400 | 48,205 | 540,519,388 | 0.9376 | 0.9244 | 0.9509 | −0.0666 | −0.0817 | −0.0516 | −0.0266 | −0.0324 | −0.0209 | 80.166 | 1.72 × 10−19 | 1.33 | - |
| All_Cancer | 2,151,597 | 382,262,943 | 3,244,348 | 537,323,245 | 0.9326 | 0.9310 | 0.9342 | −0.0723 | −0.0741 | −0.0705 | −0.0288 | −0.0295 | −0.0281 | 6343.397 | 0 | 1.35 | - |
| Hodgkins | 10,210 | 384,404,330 | 15,405 | 540,552,188 | 0.9320 | 0.9090 | 0.9556 | −0.0730 | −0.1001 | −0.0465 | −0.0291 | −0.0394 | −0.0189 | 30.469 | 1.70 × 10−8 | 1.35 | - |
| CLL | 26,057 | 384,388,483 | 39,705 | 540,527,888 | 0.9228 | 0.9085 | 0.9374 | −0.0836 | −0.1007 | −0.0668 | −0.0331 | −0.0395 | −0.0267 | 101.487 | 3.60 × 10−24 | 1.38 | - |
| Ovary | 26,688 | 384,387,852 | 43,321 | 540,524,272 | 0.8663 | 0.8532 | 0.8796 | −0.1543 | −0.1721 | −0.1369 | −0.0588 | −0.0650 | −0.0527 | 340.797 | 2.14 × 10−76 | 1.58 | - |
| Lung | 272,701 | 384,141,839 | 452,339 | 540,115,254 | 0.8478 | 0.8437 | 0.8518 | −0.1796 | −0.1852 | −0.1740 | −0.0675 | −0.0695 | −0.0656 | 4654.942 | 0 | 1.64 | - |
| Cervix | 14,961 | 384,399,579 | 25,037 | 540,542,556 | 0.8403 | 0.8234 | 0.8575 | −0.1901 | −0.2144 | −0.1662 | −0.0711 | −0.0792 | −0.0630 | 284.293 | 4.36 × 10−64 | 1.67 | - |
| Colorectal | 171,103 | 384,243,437 | 286,861 | 540,280,732 | 0.8388 | 0.8338 | 0.8438 | −0.1922 | −0.1994 | −0.1851 | −0.0718 | −0.0742 | −0.0694 | 3323.813 | 0 | 1.67 | - |
| Larynx | 14,027 | 384,400,513 | 26,349 | 540,541,244 | 0.7486 | 0.7334 | 0.7641 | −0.3358 | −0.3635 | −0.3087 | −0.1167 | −0.1246 | −0.1087 | 772.855 | 2.15 × 10−170 | 2.01 | - |
| Prostate | 218,368 | 384,196,172 | 413,423 | 540,154,170 | 0.7428 | 0.7389 | 0.7466 | −0.3463 | −0.3533 | −0.3394 | −0.1197 | −0.1217 | −0.1177 | 12739.787 | 0 | 2.03 | - |
| Kaposi | 1270 | 177,333,272 | 1589 | 143,565,460 | 0.6471 | 0.6010 | 0.6966 | −0.5455 | −0.6638 | −0.4356 | −0.2423 | −0.2837 | −0.2023 | 135.891 | 1.05 × 10−31 | 2.46 | - |
| Cancer | Term | Estimate | Std. Error | t-Statistic | S.D. | Adj. R. Squared | p-Value | E-Value Estimate | E-Value (Lower C.I.) |
|---|---|---|---|---|---|---|---|---|---|
| All_Cancer | cigmon: Δ8THC | 798.48 | 80.06 | 9.97 | 0.24 | 0.27 | 3.15 × 10−21 | Infinity | Infinity |
| Lung | cigmon: Δ8THC | 2052.34 | 228.44 | 8.98 | 0.68 | 0.31 | 8.11 × 10−18 | Infinity | Infinity |
| Stomach | cigmon: Δ8THC | 3221.86 | 395.94 | 8.14 | 1.17 | 0.02 | 4.33 × 10−15 | Infinity | Infinity |
| Gall_Bladder | cigmon: Δ8THC | 2700.28 | 340.42 | 7.93 | 0.99 | −0.04 | 2.32 × 10−14 | Infinity | Infinity |
| NH_Lymphoma | cigmon: Δ8THC | 1326.92 | 170.89 | 7.76 | 0.51 | 0.01 | 5.96 × 10−14 | Infinity | Infinity |
| Kidney | cigmon: Δ8THC | 1253.70 | 162.80 | 7.70 | 0.48 | 0.09 | 9.27 × 10−14 | Infinity | Infinity |
| CML | cigmon: Δ8THC | 2204.88 | 308.65 | 7.14 | 0.88 | 0.00 | 4.42 × 10−12 | Infinity | Infinity |
| Thyroid | cigmon: Δ8THC | 1681.77 | 242.80 | 6.93 | 0.72 | −0.01 | 1.57 × 10−11 | Infinity | Infinity |
| Vulva.&.Vagina | cigmon: Δ8THC | 2190.07 | 328.80 | 6.66 | 0.97 | 0.04 | 8.29 × 10−11 | Infinity | Infinity |
| Bladder | cigmon: Δ8THC | 1224.02 | 190.07 | 6.44 | 0.56 | 0.22 | 3.18 × 10−10 | Infinity | Infinity |
| Pancreas | cigmon: Δ8THC | 1553.31 | 245.48 | 6.33 | 0.73 | 0.06 | 6.22 × 10−10 | Infinity | Infinity |
| Cervix | cigmon: Δ8THC | 1859.44 | 320.08 | 5.81 | 0.95 | 0.08 | 1.22 × 10−8 | Infinity | Infinity |
| NH_Lymphoma | Δ8THC: AUD | 2282.42 | 393.82 | 5.80 | 0.51 | 0.01 | 1.31 × 10−8 | Infinity | Infinity |
| Stomach | Δ8THC: AUD | 5230.42 | 912.47 | 5.73 | 1.17 | 0.02 | 1.86 × 10−8 | Infinity | Infinity |
| Prostate | cigmon: Δ8THC: AUD | 8926.85 | 1562.45 | 5.71 | 0.39 | 0.26 | 2.06 × 10−8 | Infinity | Infinity |
| All_Cancer | Δ8THC: AUD | 1052.81 | 184.49 | 5.71 | 0.24 | 0.27 | 2.14 × 10−8 | Infinity | Infinity |
| Lung | Δ8THC: AUD | 2939.06 | 526.46 | 5.58 | 0.68 | 0.31 | 4.19 × 10−8 | Infinity | Infinity |
| Corpus_Uteri | cigmon: Δ8THC | 1158.96 | 213.51 | 5.43 | 0.63 | 0.13 | 9.50 × 10−8 | Infinity | Infinity |
| Melanoma | cigmon: Δ8THC: AUD | 56,176.95 | 10656.53 | 5.27 | 2.64 | 0.04 | 2.14 × 10−7 | Infinity | Infinity |
| ALL | cigmon: Δ8THC: AUD | 34,385.14 | 6634.60 | 5.18 | 1.53 | −0.04 | 3.67 × 10−7 | Infinity | Infinity |
| Colorectal | Δ8THC: AUD | 1353.86 | 266.99 | 5.07 | 0.34 | 0.15 | 5.88 × 10−7 | Infinity | Infinity |
| CML | Δ8THC: AUD | 3523.42 | 702.47 | 5.02 | 0.88 | 0.00 | 8.01 × 10−7 | Infinity | Infinity |
| Colorectal | cigmon: Δ8THC | 553.21 | 115.85 | 4.78 | 0.34 | 0.15 | 2.46 × 10−6 | Infinity | Infinity |
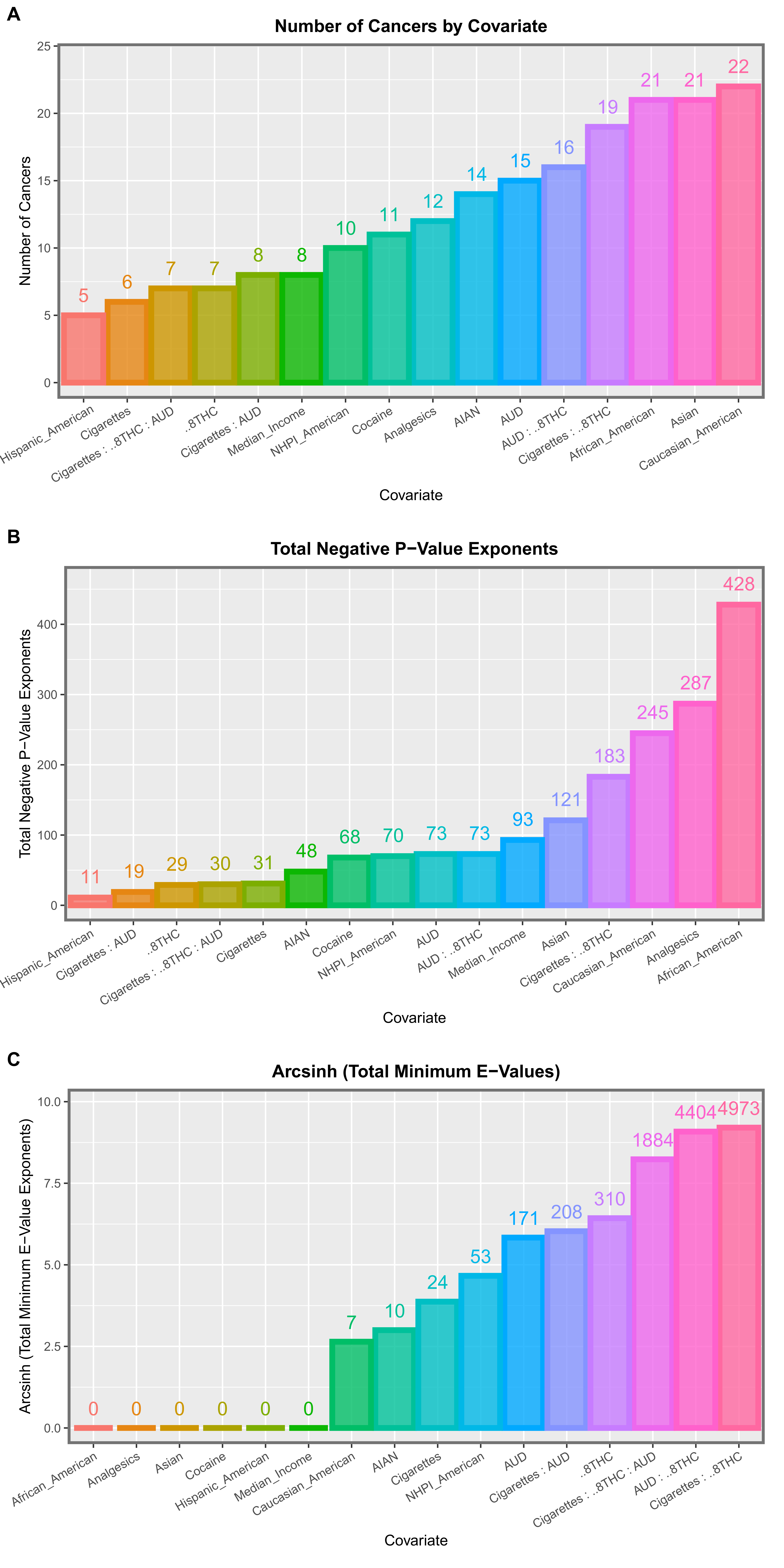
| Covariate | Number of Cancers | Total Negative Exponent of p-Value | Total Negative Exponent of Lower E-Value Bound |
|---|---|---|---|
| AIAN American | 14 | 48 | 10 |
| AUD | 15 | 73 | 171 |
| Cigarettes | 6 | 31 | 24 |
| Cigarettes: AUD | 8 | 19 | 208 |
| Cigarettes: Δ8THC | 19 | 183 | 4973 |
| Cigarettes: Δ8THC: AUD | 7 | 30 | 1884 |
| Δ8THC | 7 | 29 | 310 |
| AUD: Δ8THC | 16 | 73 | 4404 |
| Analgesics | 12 | 287 | 0 |
| Asian American | 21 | 121 | 0 |
| African_American | 21 | 428 | 0 |
| Cocaine | 11 | 68 | 0 |
| Hispanic_American | 5 | 11 | 0 |
| Median Income | 8 | 93 | 0 |
| NHPI_American | 10 | 70 | 53 |
| Caucasian American | 22 | 245 | 7 |
| Cancer | Term | Estimate | Std. Error | t-Statistic | S.D. | Adj. R. Squared | p-Value | E-Value Estimate | E-Value (Lower C.I.) |
|---|---|---|---|---|---|---|---|---|---|
| Kidney | lag(cigmon, 4): lag(Δ8THC, 4): lag(AUD, 4) | 20,775.83 | 729.67 | 28.47 | 0.35 | −0.03 | 8.04 × 10−78 | Infinity | Infinity |
| AML | lag(Δ8THC, 4): lag(AUD, 4) | 37,051.70 | 1513.53 | 24.48 | 0.72 | −0.06 | 2.36 × 10−66 | Infinity | Infinity |
| Pancreas | lag(Δ8THC, 4): lag(AUD, 4) | 30,354.71 | 1244.74 | 24.39 | 0.60 | −0.06 | 4.51 × 10−66 | Infinity | Infinity |
| Kidney | lag(cigmon, 4): lag(Δ8THC, 4) | 6008.51 | 278.08 | 21.61 | 0.35 | −0.03 | 1.44 × 10−57 | Infinity | Infinity |
| AML | lag(cigmon, 4): lag(Δ8THC, 4) | 11,523.47 | 576.82 | 19.98 | 0.72 | −0.06 | 2.09 × 10−52 | Infinity | Infinity |
| Larynx | lag(Δ8THC, 4): lag(AUD, 4) | 42,033.01 | 2140.10 | 19.64 | 1.02 | 0.12 | 2.54 × 10−51 | Infinity | Infinity |
| Brain | lag(cigmon, 4): lag(Δ8THC, 4): lag(AUD, 4) | 103,777.35 | 5628.18 | 18.44 | 0.63 | −0.06 | 2.01 × 10−47 | Infinity | Infinity |
| Pancreas | lag(cigmon, 4): lag(Δ8THC, 4) | 8433.65 | 474.38 | 17.78 | 0.60 | −0.06 | 2.93 × 10−45 | Infinity | Infinity |
| Vulva.&.Vagina | lag(cigmon, 4): lag(Δ8THC, 4) | 13,904.77 | 815.61 | 17.05 | 1.02 | 0.12 | 7.47 × 10−43 | Infinity | Infinity |
| All_Cancer | lag(Δ8THC, 4): lag(AUD, 4) | 4404.42 | 263.84 | 16.69 | 0.13 | 0.11 | 1.12 × 10−41 | Infinity | Infinity |
| Brain | lag(Δ8THC, 4) | 1766.26 | 115.19 | 15.33 | 0.63 | −0.06 | 3.70 × 10−37 | Infinity | Infinity |
| Liver | lag(Δ8THC, 4): lag(AUD, 4) | 15,683.99 | 1078.13 | 14.55 | 0.52 | 0.22 | 1.52 × 10−34 | Infinity | Infinity |
| ALL | lag(cigmon, 4): lag(Δ8THC, 4): lag(AUD, 4) | 80,386.15 | 6115.36 | 13.14 | 0.66 | −0.08 | 3.84 × 10−28 | Infinity | Infinity |
| Liver | lag(cigmon, 4): lag(Δ8THC, 4) | 4887.56 | 410.89 | 11.90 | 0.52 | 0.22 | 8.25 × 10−26 | Infinity | Infinity |
| ALL | lag(Δ8THC, 4) | 1491.45 | 125.55 | 11.88 | 0.66 | −0.08 | 1.96 × 10−24 | Infinity | Infinity |
| Oropharynx | lag(cigmon, 4): lag(Δ8THC, 4): lag(AUD, 4) | 42,596.04 | 3835.11 | 11.11 | 0.43 | 0.27 | 2.79 × 10−23 | Infinity | Infinity |
| EsophLagus | lag(cigmon, 4): lag(Δ8THC, 4): lag(AUD, 4) | 79,727.71 | 7246.10 | 11.00 | 0.80 | −0.06 | 8.94 × 10−23 | Infinity | Infinity |
| All_Cancer | lag(cigmon, 4): lag(Δ8THC, 4) | 1092.50 | 100.55 | 10.87 | 0.13 | 0.11 | 1.63 × 10−22 | Infinity | Infinity |
| CLL | lag(Δ8THC, 4): lag(AUD, 4) | 16,144.24 | 1490.78 | 10.83 | 0.71 | 0.09 | 2.11 × 10−22 | Infinity | Infinity |
| NH_Lymphoma | lag(Δ8THC, 4): lag(AUD, 4) | 11,559.70 | 1096.50 | 10.54 | 0.52 | −0.03 | 1.68 × 10−21 | Infinity | Infinity |
| EsophLagus | lag(Δ8THC, 4) | 1551.04 | 148.10 | 10.47 | 0.80 | −0.06 | 3.93 × 10−21 | Infinity | Infinity |
| Oropharynx | lag(Δ8THC, 4) | 765.23 | 78.49 | 9.75 | 0.43 | 0.27 | 4.72 × 10−19 | Infinity | Infinity |
| Myeloma | lag(cigmon, 4): lag(Δ8THC, 4): lag(AUD, 4) | 51,756.76 | 5401.86 | 9.58 | 0.61 | 0.02 | 1.52 × 10−18 | Infinity | Infinity |
| Postmenopausal Breast Cancer | lag(cigmon, 4): lag(Δ8THC, 4): lag(AUD, 4) | 23,566.70 | 2616.35 | 9.01 | 0.29 | 0.10 | 7.76 × 10−17 | Infinity | Infinity |
| Postmenopausal Breast Cancer | lag(Δ8THC, 4) | 461.33 | 53.55 | 8.61 | 0.29 | 0.10 | 1.07 × 10−15 | Infinity | Infinity |
| Bladder | lag(Δ8THC, 4): lag(AUD, 4) | 11,273.58 | 1317.42 | 8.56 | 0.63 | 0.06 | 1.57 × 10−15 | Infinity | Infinity |
| Anorectum | lag(Δ8THC, 4): lag(AUD, 4) | 16,569.38 | 1977.38 | 8.38 | 0.95 | −0.02 | 5.04 × 10−15 | Infinity | Infinity |
| CLL | lag(cigmon, 4): lag(Δ8THC, 4) | 4683.01 | 568.15 | 8.24 | 0.71 | 0.09 | 1.23 × 10−14 | Infinity | Infinity |
| Lung | lag(Δ8THC, 4): lag(AUD, 4) | 9636.37 | 1178.24 | 8.18 | 0.56 | 0.10 | 1.85 × 10−14 | Infinity | Infinity |
| Myeloma | lag(Δ8THC, 4) | 856.96 | 110.56 | 7.75 | 0.61 | 0.02 | 2.81 × 10−13 | Infinity | Infinity |
| Thyroid | lag(Δ8THC, 4): lag(AUD, 4) | 6736.53 | 885.15 | 7.61 | 0.42 | 0.18 | 6.73 × 10−13 | Infinity | Infinity |
| Bladder | lag(cigmon, 4): lag(Δ8THC, 4) | 3651.37 | 502.08 | 7.27 | 0.63 | 0.06 | 5.33 × 10−12 | Infinity | Infinity |
| Anorectum | lag(cigmon, 4): lag(Δ8THC, 4) | 4954.17 | 753.59 | 6.57 | 0.95 | −0.02 | 3.18 × 10−10 | Infinity | Infinity |
| Prostate | lag(cigmon, 4): lag(Δ8THC, 4): lag(AUD, 4) | 16,430.44 | 2513.72 | 6.54 | 0.28 | 0.15 | 3.94 × 10−10 | Infinity | Infinity |
| NH_Lymphoma | lag(cigmon, 4): lag(Δ8THC, 4) | 2661.93 | 417.88 | 6.37 | 0.52 | −0.03 | 9.98 × 10−10 | Infinity | Infinity |
| Ovary | lag(cigmon, 4): lag(Δ8THC, 4): lag(AUD, 4) | 41,343.79 | 6533.78 | 6.33 | 0.73 | −0.02 | 1.26 × 10−9 | Infinity | Infinity |
| Ovary | lag(Δ8THC, 4) | 806.38 | 133.73 | 6.03 | 0.73 | −0.02 | 6.38 × 10−9 | Infinity | Infinity |
| Thyroid | lag(cigmon, 4): lag(Δ8THC, 4) | 1993.67 | 337.34 | 5.91 | 0.42 | 0.18 | 1.21 × 10−8 | Infinity | Infinity |
| Breast | lag(cigmon, 4): lag(Δ8THC, 4): lag(AUD, 4) | 10,735.82 | 1828.95 | 5.87 | 0.21 | 0.30 | 1.49 × 10−8 | Infinity | Infinity |
| Prostate | lag(Δ8THC, 4) | 299.18 | 51.45 | 5.82 | 0.28 | 0.15 | 1.99 × 10−8 | Infinity | Infinity |
| Breast | lag(Δ8THC, 4) | 210.27 | 37.43 | 5.62 | 0.21 | 0.30 | 5.51 × 10−8 | Infinity | Infinity |
| Hodgkins | lag(Δ8THC, 4) | 1216.18 | 217.41 | 5.59 | 1.19 | 0.03 | 6.20 × 10−8 | Infinity | Infinity |
| Lung | lag(cigmon, 4): lag(Δ8THC, 4) | 2417.46 | 449.04 | 5.38 | 0.56 | 0.10 | 1.78 × 10−7 | Infinity | Infinity |
| Hodgkins | lag(cigmon, 4): lag(Δ8THC, 4): lag(AUD, 4) | 55,559.09 | 10,622.37 | 5.23 | 1.19 | 0.03 | 3.76 × 10−7 | Infinity | Infinity |
| Vulva.&.Vagina | lag(cigmon, 4): lag(Δ8THC, 4): lag(AUD, 4) | 61,495.75 | 13,454.10 | 4.57 | 1.51 | −0.01 | 8.25 × 10−6 | Infinity | Infinity |
| Stomach | lag(cigmon, 4): lag(Δ8THC, 4): lag(AUD, 4) | 33,237.10 | 7879.98 | 4.22 | 0.89 | −0.03 | 3.53 × 10−5 | Infinity | Infinity |
| Stomach | lag(Δ8THC, 4) | 669.75 | 161.28 | 4.15 | 0.89 | −0.03 | 4.61 × 10−5 | Infinity | Infinity |
| Corpus_Uteri | Caucasian American | 2187.58 | 1013.65 | 2.16 | 0.48 | −0.01 | 3.19 × 10−2 | Infinity | Infinity |
| Vulva.&.Vagina | lag(Δ8THC, 4) | 1006.92 | 275.18 | 3.66 | 1.51 | −0.01 | 3.19 × 10−4 | Infinity | 2.31 × 10123 |
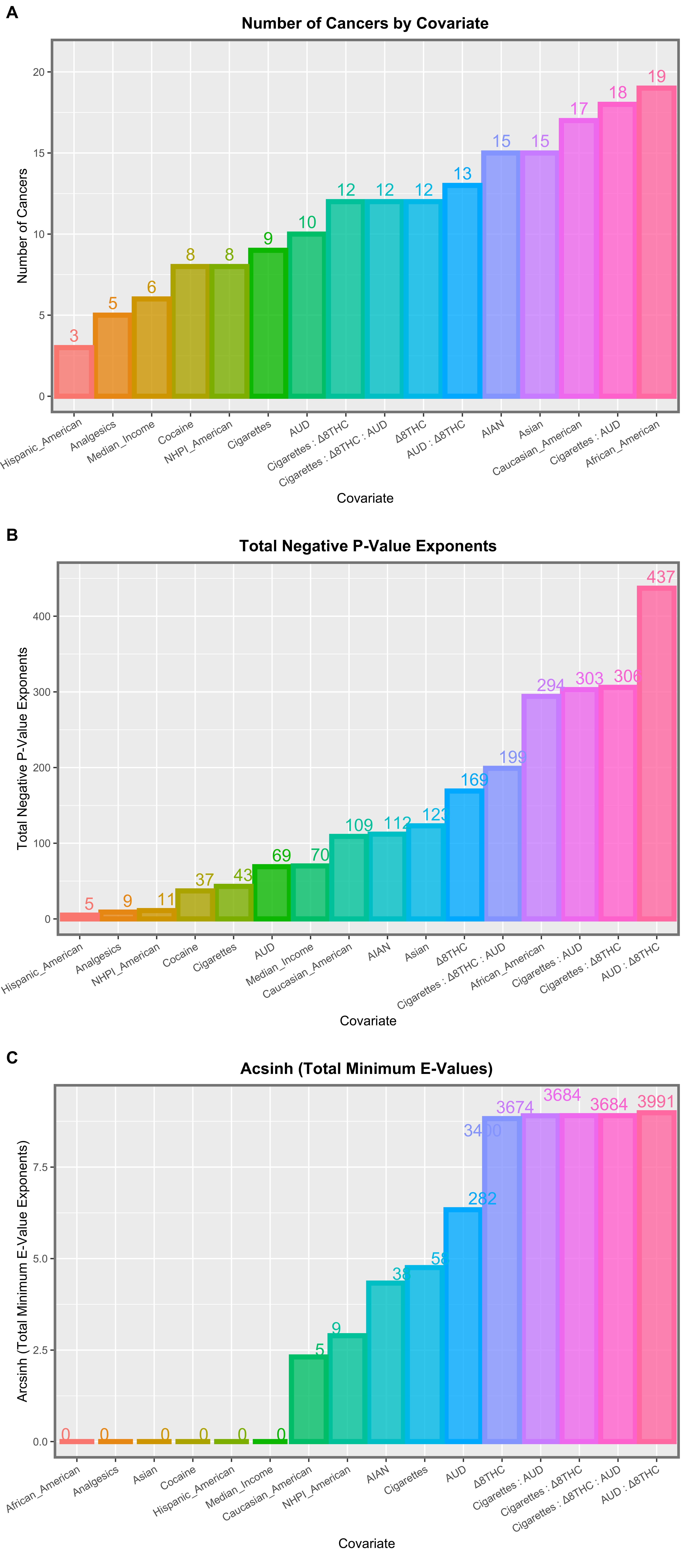
| Covariate | Number of Cancers | Total Negative Exponent of p-Value | Total Negative Exponent of Lower E-Value Bound |
|---|---|---|---|
| AIAN American | 15 | 112 | 38 |
| AUD | 10 | 69 | 282 |
| Cigarettes | 9 | 43 | 58 |
| Cigarettes: AUD | 18 | 303 | 3674 |
| Cigarettes: Δ8THC | 12 | 306 | 3684 |
| Cigarettes: Δ8THC: AUD | 12 | 199 | 3684 |
| Δ8THC | 12 | 169 | 3400 |
| AUD: Δ8THC | 13 | 437 | 3991 |
| Analgesics | 5 | 9 | 0 |
| Asian American | 15 | 123 | 0 |
| African_American | 19 | 294 | 0 |
| Cocaine | 8 | 37 | 0 |
| Hispanic_American | 3 | 5 | 0 |
| Median Income | 6 | 70 | 0 |
| NHPI_American | 8 | 11 | 9 |
| Caucasian American | 17 | 109 | 5 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Reece, A.S.; Hulse, G.K. Epidemiology of Δ8THC-Related Carcinogenesis in USA: A Panel Regression and Causal Inferential Study. Int. J. Environ. Res. Public Health 2022, 19, 7726. https://doi.org/10.3390/ijerph19137726
Reece AS, Hulse GK. Epidemiology of Δ8THC-Related Carcinogenesis in USA: A Panel Regression and Causal Inferential Study. International Journal of Environmental Research and Public Health. 2022; 19(13):7726. https://doi.org/10.3390/ijerph19137726
Chicago/Turabian StyleReece, Albert Stuart, and Gary Kenneth Hulse. 2022. "Epidemiology of Δ8THC-Related Carcinogenesis in USA: A Panel Regression and Causal Inferential Study" International Journal of Environmental Research and Public Health 19, no. 13: 7726. https://doi.org/10.3390/ijerph19137726
APA StyleReece, A. S., & Hulse, G. K. (2022). Epidemiology of Δ8THC-Related Carcinogenesis in USA: A Panel Regression and Causal Inferential Study. International Journal of Environmental Research and Public Health, 19(13), 7726. https://doi.org/10.3390/ijerph19137726







