A New Compartment Model of COVID-19 Transmission: The Broken-Link Model
Abstract
:1. Introduction
2. Materials and Methods
3. Results
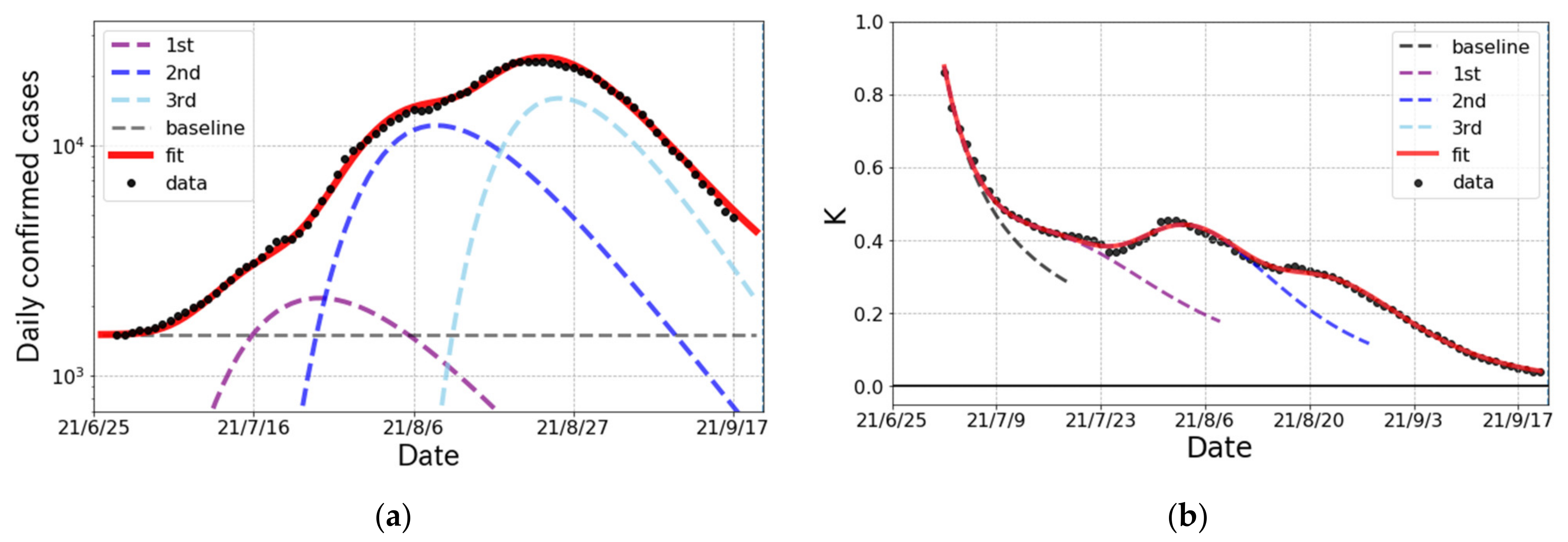
3.1. The Epidemic Surge in Japan
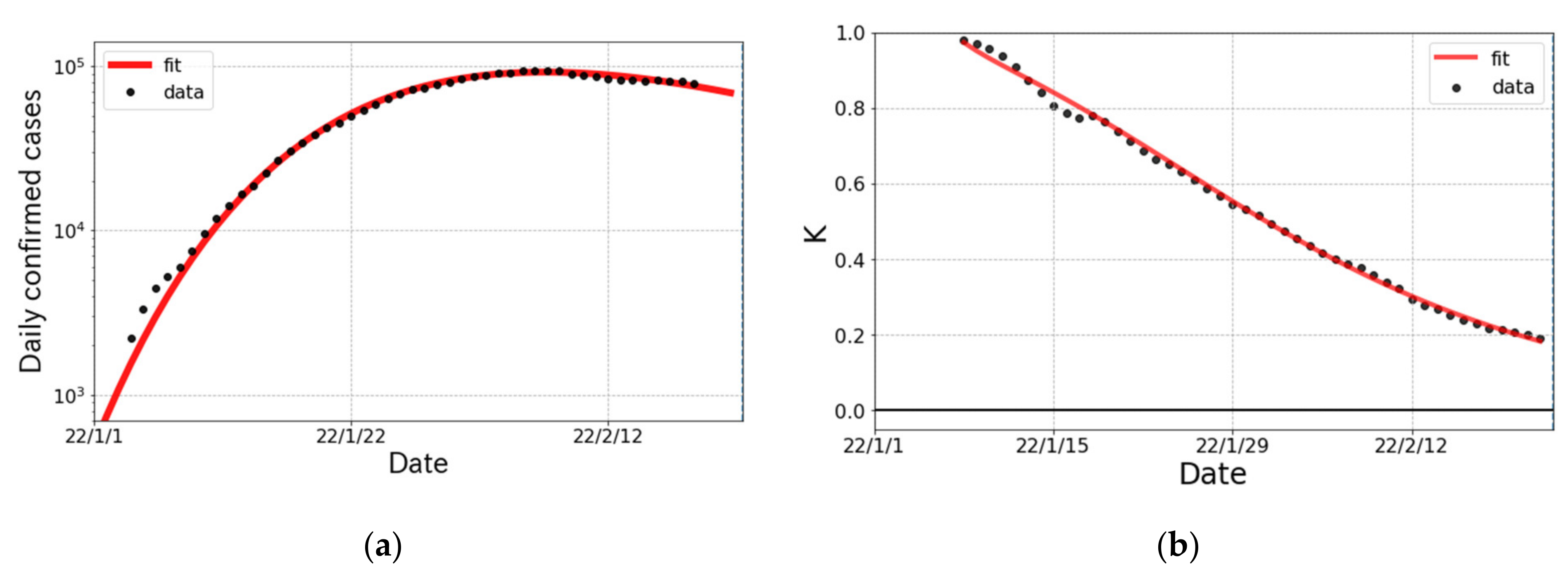
3.2. The Epidemic Surge in Japan
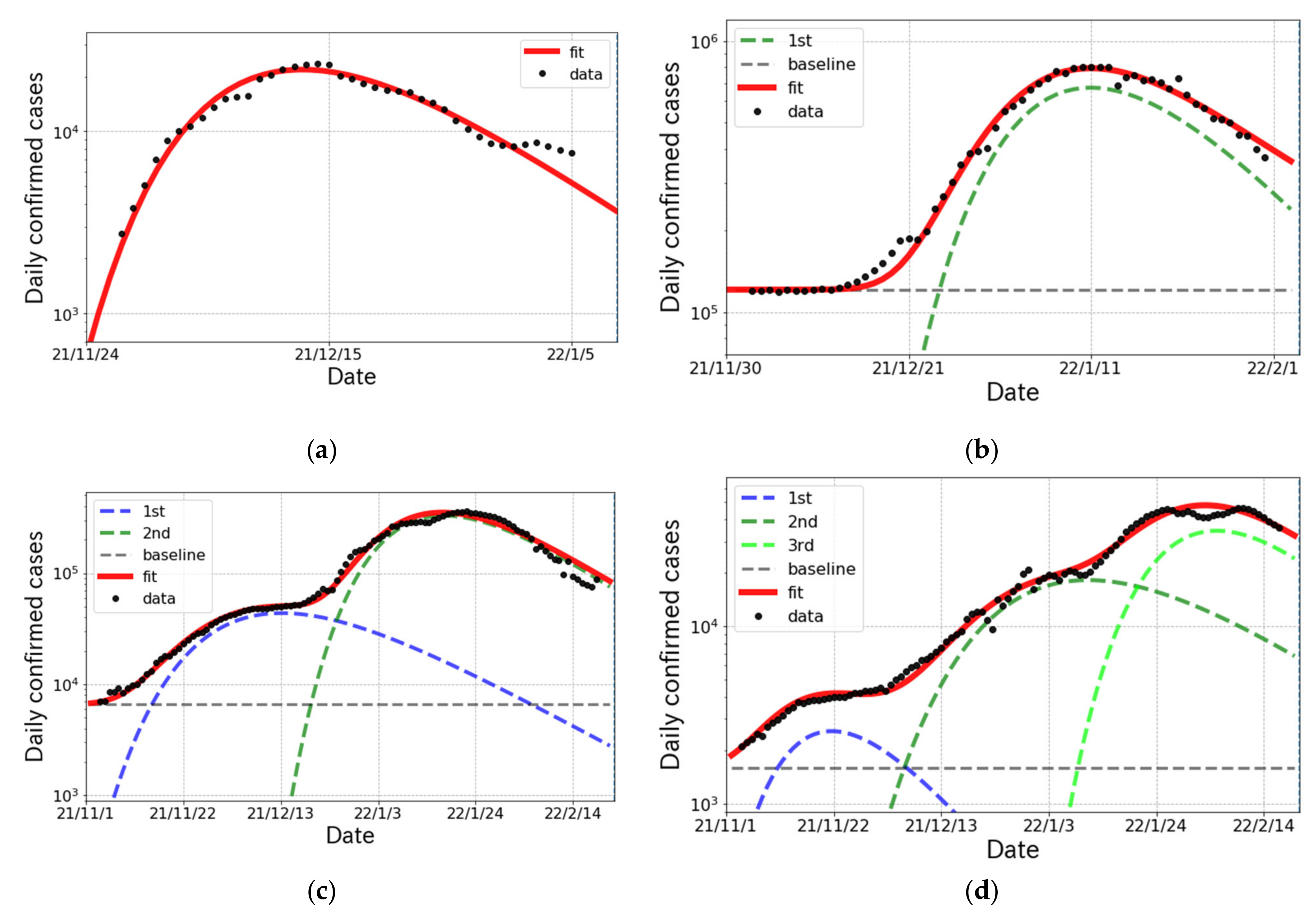
3.3. The Status of the and Surges in Other Countries
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Kermack, W.O.; McKendrick, A.G. A Contribution to the Mathematical Theory of Epidemics. Proc. R. Soc. A 1927, 115, 700–721. [Google Scholar]
- Hethcote, H.W. Qualitative analyses of communicable disease models. Math Biosci. 1976, 28, 335–356. [Google Scholar] [CrossRef]
- Flaxman, S.; Mishra, S.; Gandy, A.; Unwin, H.J.; Mellan, T.A.; Coupland, H.; Whittaker, C.; Zhu, H.; Berah, T.; Eaton, J.W.; et al. Estimating the effects of non-pharmaceutical interventions on COVID-19 in Europe. Nature 2020, 584, 257–261. [Google Scholar] [CrossRef] [PubMed]
- Gatto, M.; Bertuzzo, E.; Mari, L.; Miccoli, S.; Carraro, L.; Casagrandi, R.; Rinaldo, A. Spread and dynamics of the COVID-19 epidemic in Italy: Effects of emergency containment measures. Proc. Natl. Acad. Sci. USA 2020, 117, 10484–10491. [Google Scholar] [CrossRef] [Green Version]
- Tuite, A.R.; Fisman, D.N.; Greer, A.L. Mathematical modelling of COVID-19 transmission and mitigation strategies in the population of Ontario, Canada. CMAJ 2020, 192, E497–E505. [Google Scholar] [CrossRef] [Green Version]
- Davies, N.G.; Klepac, P.; Liu, Y.; Prem, K.; Jit, M.; Eggo, R.M. Age-dependent effects in the transmission and control of COVID-19 epidemics. Nat. Med. 2020, 26, 1205–1211. [Google Scholar] [CrossRef]
- Courtemanche, C.; Garuccio, J.; Le, A.; Pinkston, J.; Yelowitz, A. Strong Social Distancing Measures In The United States Reduced The COVID-19 Growth Rate. Health Aff. 2020, 39, 1237–1246. [Google Scholar] [CrossRef]
- Britton, T.; Ball, F.; and Trapman, P. A mathematical model reveals the influence of population heterogeneity on herd immunity to SARS-CoV-2. Science 2020, 369, 846–849. [Google Scholar] [CrossRef]
- Gola, A.; Arya, R.K.; Dugh, A.R. Review of Forecasting Models for Coronavirus (COVID-19) Pandemic in India during Country-wise Lockdowns. medRxiv 2020. [Google Scholar] [CrossRef]
- Van Wees, J.D.; van der Kuip, M.; Osinga, S.; Keijser, B.; van Westerloo, D.; Hanegraaf, M.; Pluymaekers, M.; Leeuwenburgh, O.; Brunner, L.; van Furth, M.T. SIR model for assessing the impact of the advent of Omicron and mitigating measures on infection pressure and hospitalization needs. medRxiv 2021. [Google Scholar] [CrossRef]
- Ngonghala, C.N.; Iboi, E.; Eikenberry, S.; Scotch, M.; MacIntyre, C.R.; Bonds, M.H.; Gumel, A.B. Mathematical assessment of the impact of non-pharmaceutical interventions on curtailing the 2019 novel Coronavirus. Math. Biosci. 2020, 325, 108364. [Google Scholar] [CrossRef] [PubMed]
- Wood, S.N.; Wit, E.C. Was R <1 before the English lockdowns? On modelling mechanistic detail, causality and inference about COVID-19. PLoS ONE 2021, 16, e0257455. [Google Scholar] [CrossRef]
- Schlickeiser, R.; Kröger, M. Forecast of Omicron Wave Time Evolution. COVID 2022, 2, 17. [Google Scholar] [CrossRef]
- Kamiya, T.; Alvarez-Iglesias, A.; Ferguson, J.; Murphy, S.; Sofonea, M.T.; Fitz-Simon, N. Estimating time-dependent infectious contact: A multi-strain epidemiological model of SARS-CoV-2 on the island of Ireland. medRxiv 2022. [Google Scholar] [CrossRef]
- Roda, W.C.; Varughese, M.B.; Han, D.; Li, M.Y. Why is it difficult to accurately predict the COVID-19 epidemic? Infect. Dis. Model. 2020, 5, 271–281. [Google Scholar] [CrossRef] [PubMed]
- Ferguson, N.M.; Laydon, D.; Nedjati-Gilani, G.; Imai, N.; Ainslie, K.; Baguelin, M.; Bhatia, S.; Boonyasiri, A.; Cucunubá, Z.; Cuomo-Dannenburg, G. Impact of Non-pharmaceutical Interventions (NPIs) to Reduce COVID-19 Mortality and Healthcare Demand. Bull. Math. Biol. 2020, 82, 52. [Google Scholar] [CrossRef] [Green Version]
- Fu, X.; Ying, Q.; Zeng, T.; Long, T.; Wang, Y. Simulating and forecasting the cumulative confirmed cases of SARSCoV-2 in China by Boltzmann function-based regression analyses. J. Infect. Lett. Ed. 2020, 80, 602–605. [Google Scholar]
- Derevich, I.V.; Panova, A.A. Estimation of COVID-19 Infection Growth Rate Based on the Imbedding Method. J. Eng. Phys. 2021, 94, 18–29. [Google Scholar] [CrossRef]
- Nakano, T.; Ikeda, Y. Novel Indicator to Ascertain the Status and Trend of COVID-19 Spread: Modeling Study. J. Med. Internet Res. 2020, 22, e20144. [Google Scholar] [CrossRef]
- Akiyama, Y. The K Indicator Epidemic Model Follows the Gompertz Curve. Available online: http://www.bi.cs.titech.ac.jp/COVID-19/The_K_indicator_epidemic_model_follows_the_Gompertz_curve.html (accessed on 9 February 2022).
- Ohnishi, A.; Namekawa, Y.; Fukui, T. Universality in COVID-19 spread in view of the Gompertz function. Prog. Theor. Exp. Phys. 2020, 2020, 123J01. [Google Scholar] [CrossRef]
- Levitt, M.; Scaiewicz, A.; Zonta, F. Predicting the trajectory of any COVID-19 epidemic from the best straight line. medRxiv 2020. [Google Scholar] [CrossRef]
- Nishiura, H.; Oshitani, H.; Kobayashi, T.; Saito, T.; Sunagawa, T.; Matsui, T.; Wakita, T.; MHLW COVID-19 Response Team; Suzuki, M. Closed environments facilitate secondary transmission of coronavirus disease 2019 (COVID-19). medRxiv 2020. [Google Scholar] [CrossRef] [Green Version]
- Dong, E.; Du, H.; Gardner, L. An interactive web-based dashboard to track COVID-19 in real time. Lancet Inf. Dis. 2020, 20, 533–534. Available online: https://github.com/CSSEGISandData/COVID-19 (accessed on 20 February 2022). [CrossRef]
- Shimizu, K.; Iyoda, T.; Sanpei, A.; Nakazato, H.; Okada, M.; Ueda, S.; Kato-Murayama, M.; Murayama, K.; Shirouzu, M.; Harada, N.; et al. Identification of TCR repertoires in functionally competent cytotoxic T cells cross-reactive to SARS-CoV-2. Commun. Biol. 2021, 4, 1365. [Google Scholar] [CrossRef]
- Ministry of Health, Lavour and Welfare (MHLW) of Japan. Situation Report for COVID-19. Available online: https://www.mhlw.go.jp/stf/covid-19/kokunainohasseijoukyou_00006.html (accessed on 11 February 2022).
- Rambaut, A.; Holmes, E.C.; O’Toole, Á.; Hill, V.; McCrone, J.T.; Ruis, C.; du Plessis, L.; Pybus, O.G. A dynamic nomenclature proposal for SARS-CoV-2 lineages to assist genomic epidemiology. Nat. Microbiol. 2020, 7, 1403–1407. [Google Scholar] [CrossRef]
- PANGO Lineages: Latest Epidemiological Lineages of SARS-CoV-2. Available online: https://cov-lineages.org/index.html (accessed on 11 February 2022).
- Liu, Y.; Rocklöv, J. The reproductive number of the delta variant of SARS-CoV-2 is far higher compared to the ancestral SARS-CoV-2 virus. J. Travel Med. 2021, 28, taab124. [Google Scholar] [CrossRef]
- Burki, T.K. Omicron variant and booster COVID-19 vaccines. Lancet Respir. Med. 2022, 10, e17. [Google Scholar] [CrossRef]
- Mullen, J.L.; Tsueng, G.; Latif, A.A.; Alkuzweny, M.; Cano, M.; Haag, E.; Zhou, J.; Zeller, M.; Hufbauer, E.; Matteson, N.; et al. The Center for Viral Systems Biology. 2020. Available online: https://outbreak.info/ (accessed on 11 February 2022).

| Parameters/Functions | Descriptions |
|---|---|
| day | |
| cumulative number of confirmed cases | |
| K-value (indicator) | |
| geometric progression | |
| constant attenuation factor |
| Parameters/Functions | Descriptions |
|---|---|
| time | |
| number of susceptible people | |
| number of infected people | |
| cumulative number of confirmed cases | |
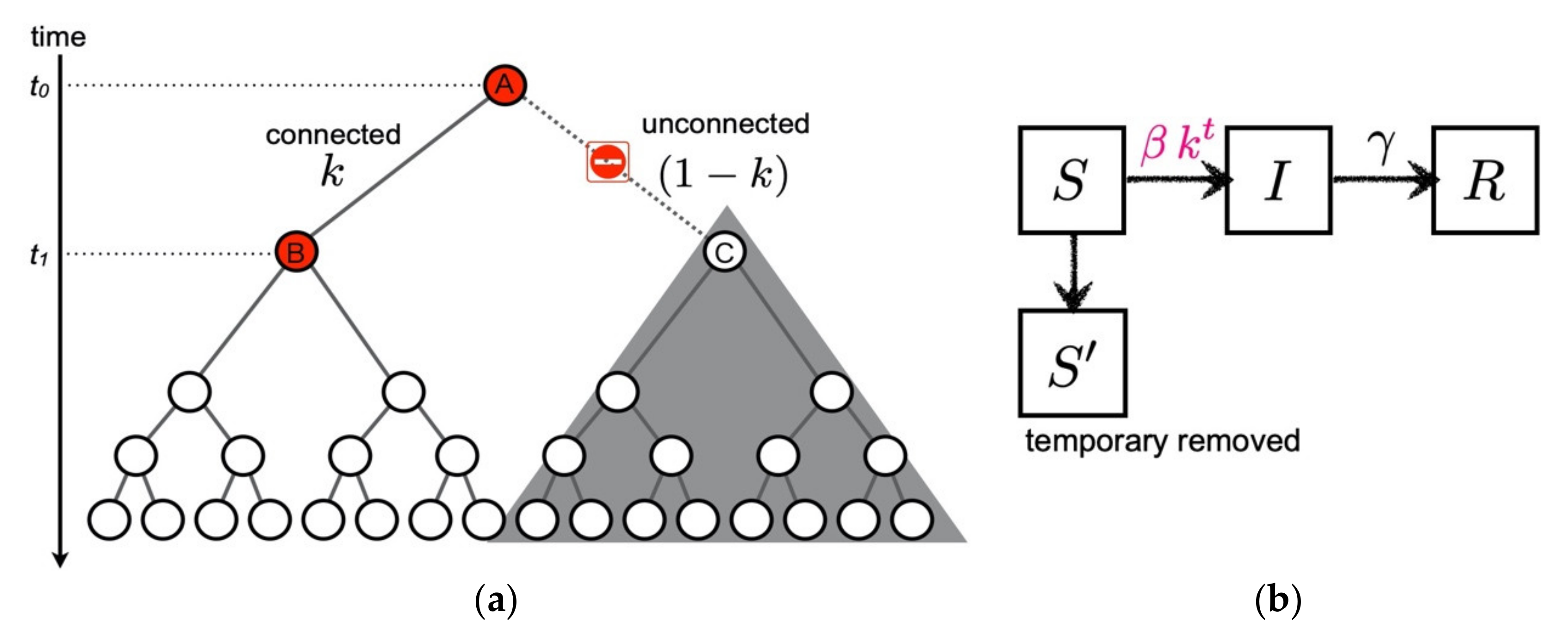
| probability of connected transmission links | |
| basic reproduction number | |
| cumulative number of infected people in each infection wave generated by | |
| removal rate from transmission trees |
| Partial Wave | Shift (Days) | |||
|---|---|---|---|---|
| 1st | 75 (12) k | 6.49 (20) | 0.918 (4) | 7.2 (3) |
| 2nd | 340 (23) k | 6.98 (16) | 0.907 (3) | 24.5 (3) |
| 3rd | 375 (2) k | 4.40 (15) | 0.892 (1) | 47.7 (3) |
| Partial Wave | Shift (Days) | |||
|---|---|---|---|---|
| 1st | 4332 (6) k | 10.4 (1) | 0.944 (1) | −4.3 (1) |
| Region | Partial Wave | Shift (Days) | |||
|---|---|---|---|---|---|
| South Africa | 1st | 592 (1) k | 8.98 (11) | 0.905 (1) | −3.7 (2) |
| United States | 1st | 22,411 (269) k | 9.68 (6) | 0.922 (1) | 13.6 (4) |
| France | 1st | 2233 (235) k | 7.18 (22) | 0.949 (2) | 4.3 (6) |
| 2nd | 13,330 (16) k | 9.92 (1) | 0.935 (1) | 42.4 (1) | |
| Denmark | 1st | 84 (1) k | 6.16 (64) | 0.924 (1) | −3.0 (6) |
| 2nd | 1094 (124) k | 9.93 (28) | 0.956 (2) | 19.4 (1.0) | |
| 3rd | 1401 (6) k | 8.00 (25) | 0.937 (1) | 63.3 (5) |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ikeda, Y.; Sasaki, K.; Nakano, T. A New Compartment Model of COVID-19 Transmission: The Broken-Link Model. Int. J. Environ. Res. Public Health 2022, 19, 6864. https://doi.org/10.3390/ijerph19116864
Ikeda Y, Sasaki K, Nakano T. A New Compartment Model of COVID-19 Transmission: The Broken-Link Model. International Journal of Environmental Research and Public Health. 2022; 19(11):6864. https://doi.org/10.3390/ijerph19116864
Chicago/Turabian StyleIkeda, Yoichi, Kenji Sasaki, and Takashi Nakano. 2022. "A New Compartment Model of COVID-19 Transmission: The Broken-Link Model" International Journal of Environmental Research and Public Health 19, no. 11: 6864. https://doi.org/10.3390/ijerph19116864
APA StyleIkeda, Y., Sasaki, K., & Nakano, T. (2022). A New Compartment Model of COVID-19 Transmission: The Broken-Link Model. International Journal of Environmental Research and Public Health, 19(11), 6864. https://doi.org/10.3390/ijerph19116864






