Microbial Biofilms in the Food Industry—A Comprehensive Review
Abstract
1. Introduction
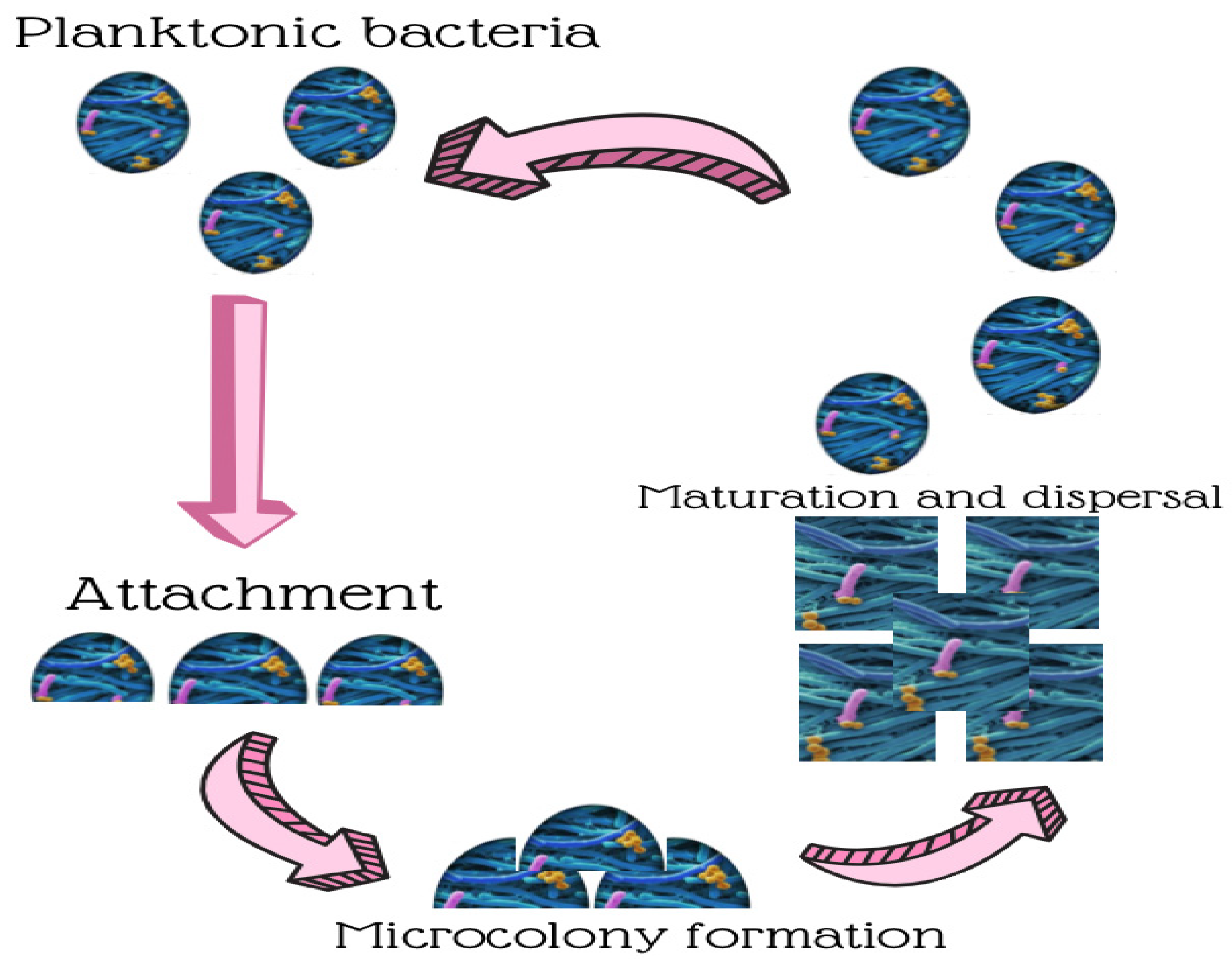
2. Biofilm Development in Food Processing Environments
3. Examples of the Most Relevant Biofilms in the Food Industry
3.1. Bacillus Cereus
3.2. Campylobacter Jejuni
3.3. Enterohaemorrhagic Escherichia coli (EHEC)
3.4. Listeria Monocytogenes
3.5. Salmonella Enterica
3.6. Staphylococcus Aureus
3.7. Pseudomonas spp.
3.8. Geobacillus stearothermophilus
3.9. Anoxybacillus flavithermus
3.10. Pectinatus spp.
3.11. Synergistic Pathogens
4. Biofilm Control and Elimination
4.1. Biofilm Elimination in the Food Industry
4.2. Factors Associated with Bacteria
- Surface charge: the adhesion sequence can be influenced by the particle surface charge in combination with the electrode’s surface charge.
- Hydrophobicity: adsorption of surface-active organics influences the surface’s hydrophobic or hydrophilic character, and changes surface tension [98].
- Temperature: temperature and contact time impact the bacterial adhesion and biofilm formation process, and increasing the formulation of mathematical models is necessary to assess how both factors and their interactions influence the process [99].
- Presence of substrates: adsorption of electrolyte components and the particle surface are equally important. Adsorption of surface-active organics impacts the surface’s hydrophobic or hydrophilic character, and also changes surface tension [98].
- Fimbria, pili, and flagella: fimbriae, non-flagellar appendages other than those implicated in the transfer of viral or bacterial nucleic acids (known as pili) are responsible for cell surface hydrophobicity. Most investigated fimbriae contain a high proportion of hydrophobic amino acid residues [100]. Fimbriae include adhesins that attach to some sort of substratum so that bacteria can withstand shear forces and obtain nutrients. Therefore, fimbriae play a role in cell surface hydrophobicity and attachment, presumably by overcoming the initial electrostatic repulsion barrier between the cell and the substratum [101].
4.3. Disinfectants and Biofilm Resistance
- Sodium hypochlorite (NaOCI):
- 2.
- Quaternary ammonium:
- 3.
- Peracetic acid:
Resistance to Disinfectants
4.4. Alternative Methods to Eliminate Biofilms
5. Biofilm Identification Techniques and Methods to Visualise Biofilms In Situ
5.1. General Aspects of Biofilm Study Techniques
5.2. Colorimetric Methods
5.2.1. Evaluating the Biofilm Matrix
5.2.2. Cell Staining
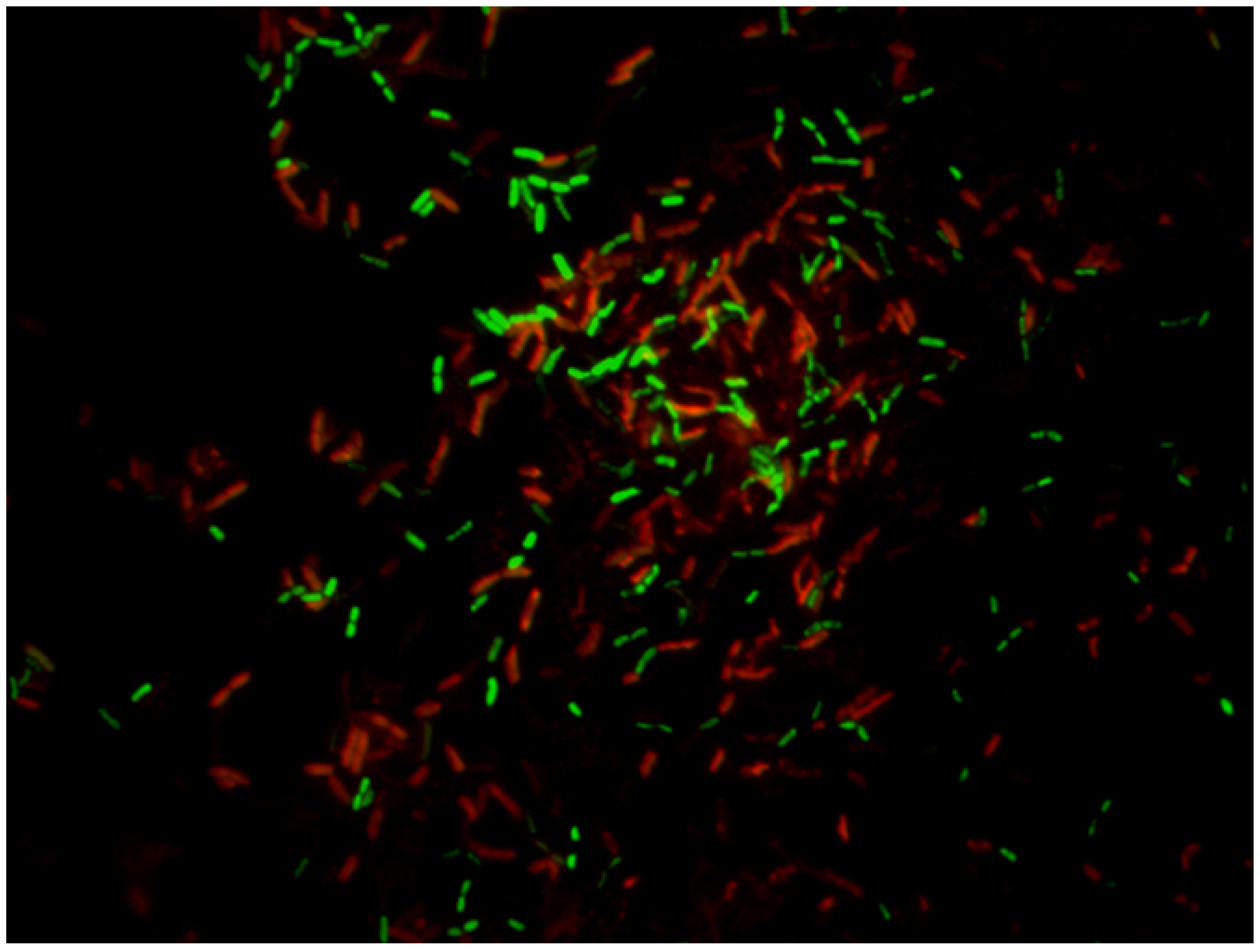
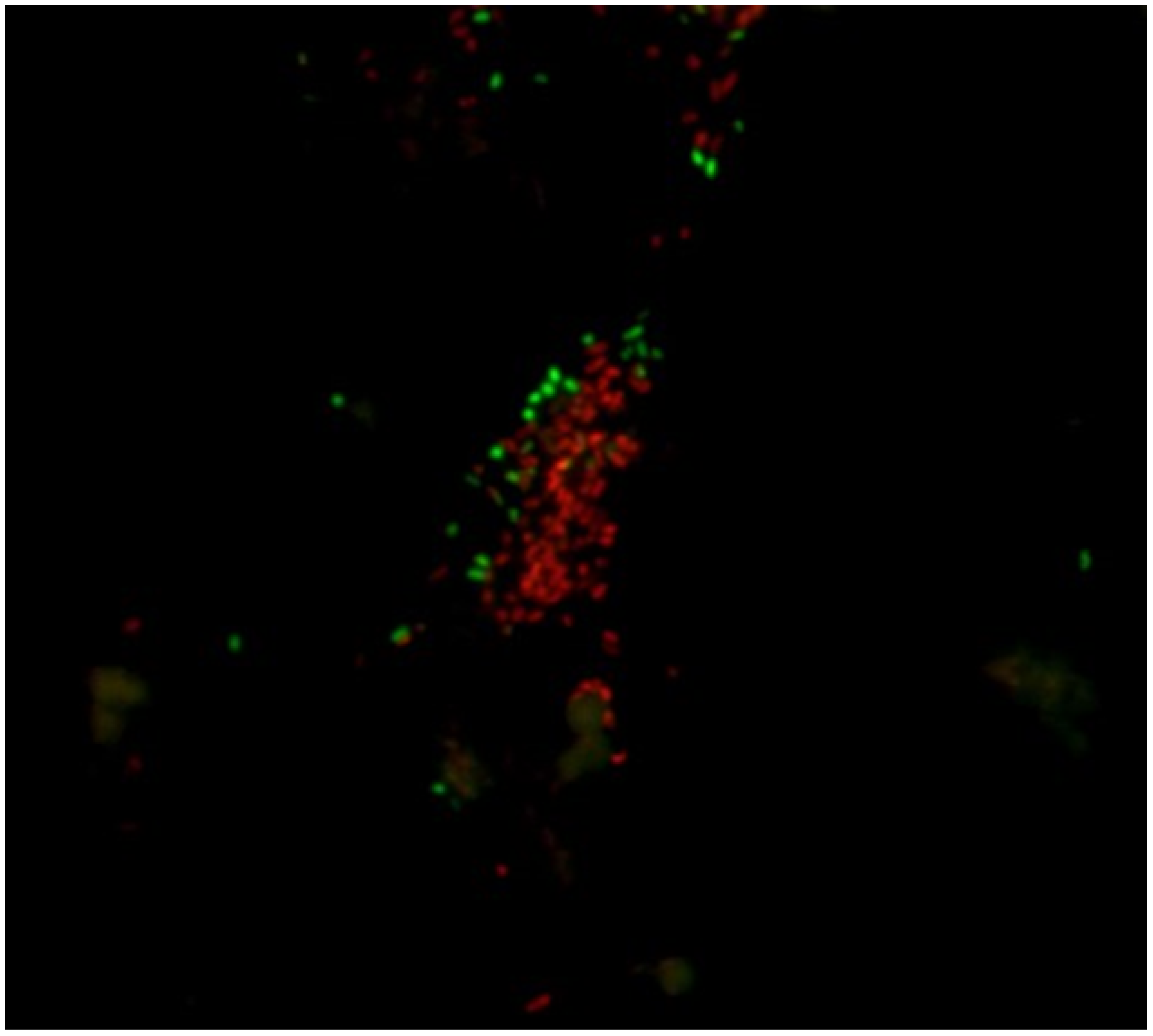
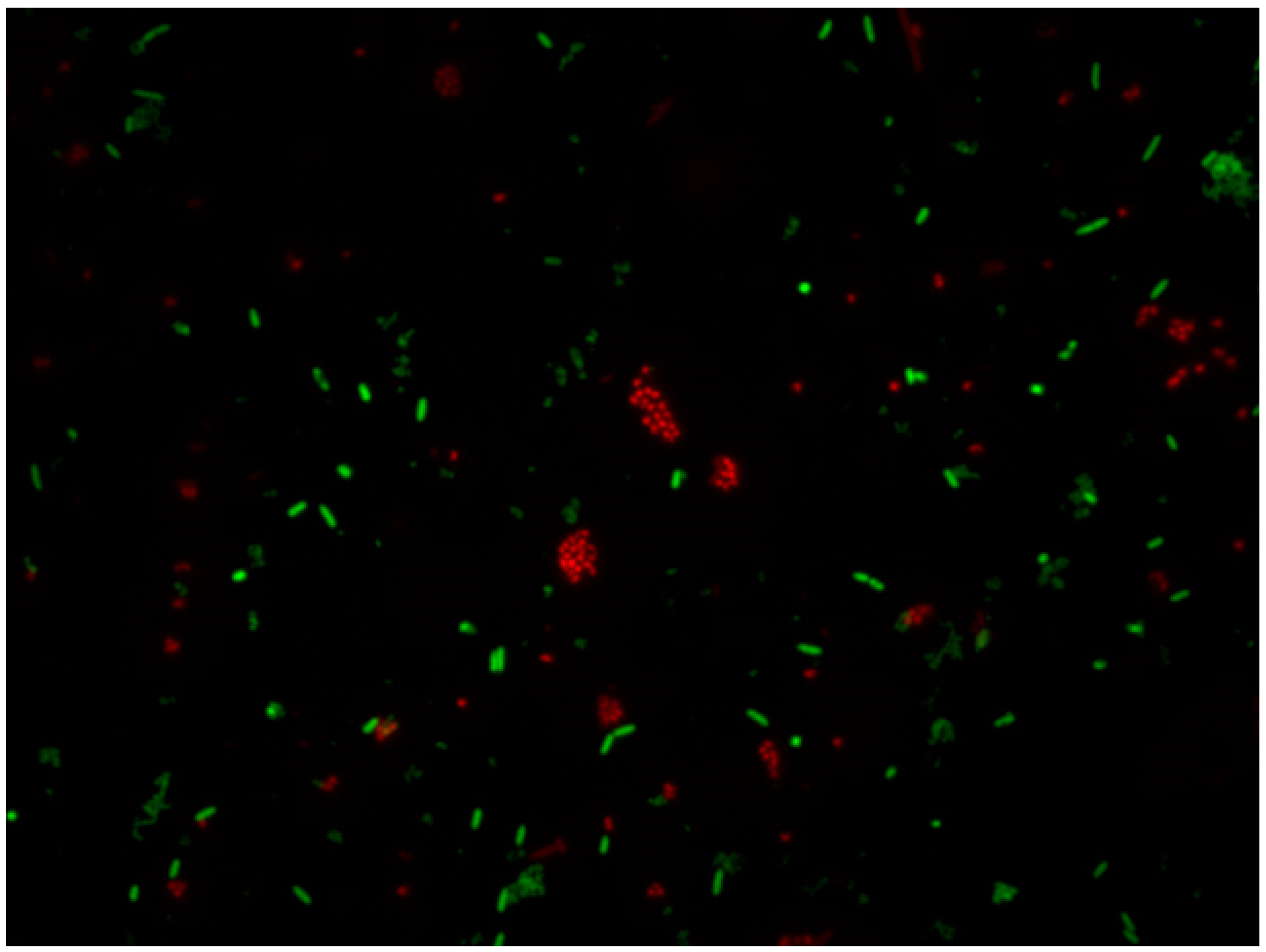
5.2.3. LIVE/DEAD
5.2.4. Different Fluorescents Stainings
5.2.5. Confocal Laser Scanning Microscopy (CLSM)
5.3. Raman Microscopy (RM)
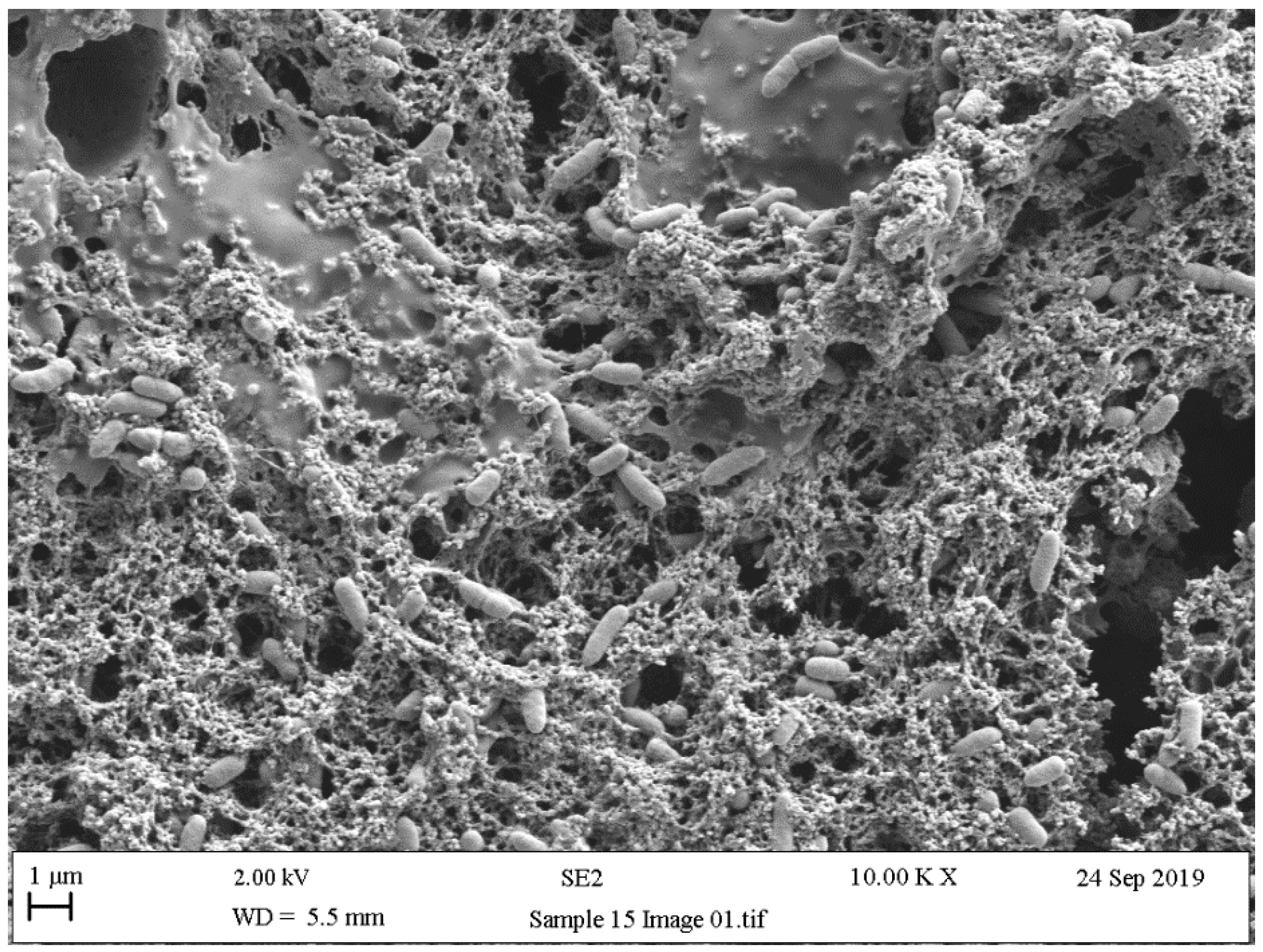
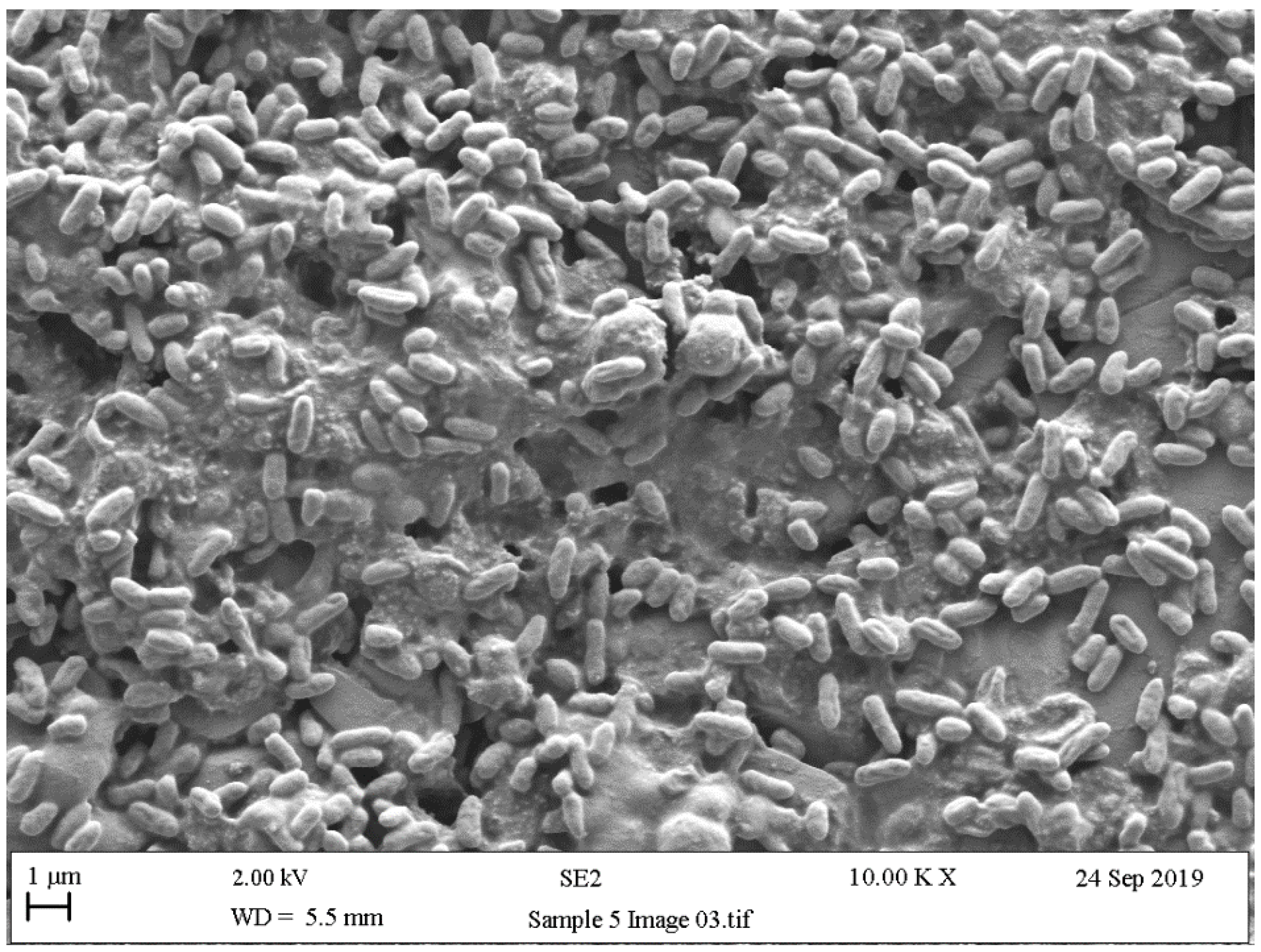
5.4. Scanning Electron Microscopy (SEM)
5.5. Microbiological Methods
6. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Satpathy, S.; Sen, S.K.; Pattanaik, S.; Raut, S. Review on bacterial biofilm: An universal cause of contamination. Biocatal. Agric. Biotechnol. 2016, 7, 56–66. [Google Scholar] [CrossRef]
- Flemming, H.C.; Wingender, J. The biofilm matrix. Nat. Rev. Microbiol. 2010, 8, 623–633. [Google Scholar] [CrossRef] [PubMed]
- Hall-Stoodley, L.J.; Costerton, W.; Stoodley, P. Bacterial biofilms: From the natural environment to infectious diseases. Nat. Rev. Microbiol. 2004, 2, 95–108. [Google Scholar] [CrossRef]
- Acker, H.V.; Dijck, P.V.; Coenye, P.V. Molecular mechanisms of antimicrobial tolerance and resistance in bacterial and fungal biofilms. Trends Microbiol. 2014, 22, 326–333. [Google Scholar] [CrossRef]
- Alvarez-Ordóñez, A.L.; Coughlan, M.; Briandet, R.; Cotter, P.D. Biofilms in food processing environments: Challenges and opportunities. Annu. Rev. Food Sci. Technol. 2019, 10, 173–195. [Google Scholar] [CrossRef]
- Brooks, J.D.; Flint, S.H. Biofilms in the food industry: Problems and potential solutions. Int. J. Food Sci. Technol. 2008, 43, 2163–2176. [Google Scholar] [CrossRef]
- Møretrø, M.; Langsrud, S. Residential bacteria on surfaces in the food industry and their implications for food safety and quality. Compr. Rev. Food Sci. Food Saf. 2017, 16, 1022–1041. [Google Scholar] [CrossRef]
- Yuan, L.; Sadiq, F.A.; Burmølle, M.; Liu, T.; He, G. Insights into bacterial milk spoilage with particular emphasis on the roles of heat-stable enzymes, biofilms, and quorum sensing. J. Food Prot. 2018, 81, 1651–1660. [Google Scholar] [CrossRef]
- Srey, S.; Jahid, I.K.; Ha, S.D. Biofilm formation in food industries: A food safety concern. Food Control 2013, 31, 572–585. [Google Scholar] [CrossRef]
- Anwar, H.; Strap, J.L.; Costerton, J.W. Establishment of aging biofilms: Possible mechanism of bacterial resistance to antimicrobial therapy. Antimicrob. Agents Chemother. 1992, 36, 1347. [Google Scholar] [CrossRef]
- Costerton, J.W.; Ellis, B.; Lam, K.; Johnson, F.; Khoury, A.E. Mechanism of electrical enhancement of efficacy of antibiotics in killing biofilm bacteria. Antimicrob. Agents Chemother. 1994, 38, 2803–2809. [Google Scholar] [CrossRef]
- Simoes, M.; Simoes, L.; Vieira, M. A review of current and emergent biofilm control strategies. LWT Food Sci. Technol. 2010, 43, 573–583. [Google Scholar] [CrossRef]
- Fung, D.Y.C. Rapid methods and automation in microbiology: A review. Ir. J. Agric. Food Res. 2000, 39, 301–307. [Google Scholar] [CrossRef]
- Bouix, M.; Leveau, J.Y. Les applications de la cytometrie en flux en microbiologie. L’Eurobiologiste 2002, 36, 31–43. [Google Scholar]
- Holah, J.T.; Taylor, J.H.; Dawson, D.J.; Hall, K.E. Biocide use in the food industry and the disinfectant resistance of persistent strains of Listeria monocytogenes and Escherichia Coli. J. Appl. Microbiol. 2002, 92, 111S–120S. [Google Scholar] [CrossRef] [PubMed]
- Kassa, H. An outbreak of Norwalk-like viral gastroenteritis in a frequently penalized food service operation: A case for mandatory training of food handlers in safety and hygiene. J. Environ. Health. 2001, 64. [Google Scholar]
- González-Rivas, F.; Ripolles-Avila, C.; Fontecha-Umaña, F.; Ríos-Castillo, A.G.; Rodríguez-Jerez, J.J. Biofilms in the Spotlight: Detection, Quantification, and Removal Methods. Compr. Rev. Food Sci. Food Saf. 2018, 17, 1261–1276. [Google Scholar] [CrossRef]
- Shi, X.; Zhu, X. Biofilm formation and food safety in food industries. Trends Food Sci. Technol. 2009, 20, 407–413. [Google Scholar] [CrossRef]
- Anand, S.; Singh, D.; Avadhanula, M.; Marka, S. Development and control of bacterial biofilms on dairy processing membranes. Compr. Rev. Food Sci. Food Saf. 2014, 13, 18–33. [Google Scholar] [CrossRef]
- Rabin, N.; Zheng, Y.; Opoku-Temeng, C.; Du, Y.; Bonsu, E.; Sintim, H.O. Biofilm formation mechanisms and targets for developing antibiofilm agents. Future Med. Chem. 2015, 7, 493–512. [Google Scholar] [CrossRef]
- Lindsay, D.; von Holy, A. What food safety professionals should know about bacterial biofilms. Br. Food J. 2006, 108, 27–37. [Google Scholar] [CrossRef]
- Tang, L.; Pillai, S.; Revsbech, N.P.; Schramm, A.; Bischoff, C.; Meyer, R.L. Biofilm retention on surfaces with variable roughness and hydrophobicity. Biofouling 2011, 27, 111–121. [Google Scholar] [CrossRef] [PubMed]
- Van Houdt, R.; Michiels, C.W. Biofilm formation and the food industry, a focus on the bacterial outer surface. J. Appl. Microbiol. 2010, 109, 1117–1131. [Google Scholar] [CrossRef]
- Govaert, M.; Smet, C.; Baka, M.; Janssens, T.; Van Impe, J. Influence of incubation conditions on the formation of model biofilms by Listeria monocytogenes and Salmonella Typhimurium on abiotic surfaces. J. Appl. Microbiol. 2018, 125, 1890–1900. [Google Scholar] [CrossRef] [PubMed]
- Makovcova, J.; Babak, V.; Kulich, P.; Masek, J.; Slany, M.; Cincarova, L. Dynamics of mono- and dual-species biofilm formation and interactions between Staphylococcus aureus and gramnegative bacteria. Microb. Biotechnol. 2017, 10, 819–832. [Google Scholar] [CrossRef]
- Araújo, E.A.; de Andrade, N.J.; da Silva, L.H.M.; de Carvalho, A.F.; da Silva, C.A.; Ramos, A.M. Control of microbial adhesion as a strategy for food and bioprocess technology. Food Bioprocess Technol. 2010, 3, 321–332. [Google Scholar] [CrossRef]
- Dhowlaghar, N.; Bansal, M.; Schilling, M.W.; Nannapaneni, R. Scanning electron microscopy of Salmonella biofilms on various food-contact surfaces in catfish mucus. Food Microbiol. 2018, 74, 143–150. [Google Scholar] [CrossRef] [PubMed]
- Jindal, S.; Anand, S.; Metzger, L.; Amamcharla, J. A comparison of biofilm development on stainless steel and modified-surface plate heat exchangers during a 17-h milk pasteurization run. J. Dairy Sci. 2018, 101, 2921–2926. [Google Scholar] [CrossRef] [PubMed]
- Gomes, L.C.; Silva, L.N.; Simoes, M.; Melo, L.F.; Mergulhao, F.J. Escherichia coli adhesion, biofilm development and antibiotic susceptibility on biomedical materials. J. Biomed. Mater. Res. 2015, 103, 1414–1423. [Google Scholar] [CrossRef] [PubMed]
- Veluz, G.A.; Pitchiah, S.; Alvarado, C.Z. Attachment of Salmonella serovars and Listeria monocytogenes to stainless steel and plastic conveyor belts. Poult. Sci. 2012, 91, 2004–2010. [Google Scholar] [CrossRef] [PubMed]
- Iñiguez-Moreno, M.M.; Gutiérrez-Lomelí, A.; Avila-Novoa, M.G. Kinetics of biofilm formation by pathogenic and spoilage microorganisms under conditions that mimic the poultry, meat, and egg processing industries. Int. J. Food Microbiol. 2019, 303, 32–41. [Google Scholar] [CrossRef] [PubMed]
- Dutra, T.V.; Fernandes, M.D.; Perdoncini, M.R.F.G.; dos Anjos, M.M.; Abreu Filho, B.A.D. Capacity of Escherichia coli and Staphylococcus aureus to produce biofilm on stainless steel surfaces in the presence of food residues. J. Food Process. Preserv. 2018, 42, e13574. [Google Scholar] [CrossRef]
- Duanis-Assaf, D.; Steinberg, D.; Chai, Y.; Shemesh, M. The LuxS based quorum sensing governs lactose induced biofilm formation by Bacillus subtilis. Front. Microbiol. 2016, 6, 1517. [Google Scholar] [CrossRef]
- Xue, T.; Chen, X.; Shang, F. Effects of lactose and milk on the expression of biofilm-associated genes in Staphylococcus aureus strains isolated from a dairy cow with mastitis. J. Dairy Sci. 2014, 97, 6129–6134. [Google Scholar] [CrossRef] [PubMed]
- Somerton, B.; Lindsay, D.; Palmer, J.; Brooks, J.; Flint, S. Changes in sodium, calcium, and magnesium ion concentrations that inhibit Geobacillus biofilms have no effect on Anoxybacillus flavithermus biofilms. Appl. Environ. Microbiol. 2015, 81, 5115–5122. [Google Scholar] [CrossRef]
- Wang, R.; Kalchayanand, N.; Bono, J.L. Sequence of colonization determines the composition of mixed biofilms by Escherichia coli O157:H7 and O111:H8 strains. J. Food Prot. 2015, 78, 1554–1559. [Google Scholar] [CrossRef]
- Abdallah, M.; Khelissa, O.; Ibrahim, A.; Benoliel, C.; Heliot, L.; Dhulster, P.; Chihib, N.E. Impact of growth temperature and surface type on the resistance of Pseudomonas aeruginosa and Staphylococcus aureus biofilms to disinfectants. Int. J. Food Microbiol. 2015, 214, 38–47. [Google Scholar] [CrossRef]
- Colagiorgi, A.; Bruini, I.; Di Ciccio, P.A.; Zanardi, E.; Ghidini, S.; Ianieri, A. Listeria Monocytogenes Biofilms in the Wonderland of Food Industry. Pathogens 2017, 6, 41. [Google Scholar] [CrossRef] [PubMed]
- Bottone, E.J. Bacillus Cereus, a Volatile Human Pathogen. Clin. Microbiol. Rev. 2010, 23, 382–398. [Google Scholar] [CrossRef]
- Grigore-Gurgu, L.; Bucur, F.I.; Borda, D.; Alexa, E.A.; Neagu, C.; Nicolau, A.I. Biofilms Formed by Pathogens in Food and Food Processing Environments. In Bacterial Biofilms; IntechOpen: London, UK, 2019. [Google Scholar]
- Klančnik, A.; Šimunović, K.; Sterniša, M.; Ramić, D.; Smole Možina, S.; Bucar, F. Anti-Adhesion Activity of Phytochemicals to Prevent Campylobacter Jejuni Biofilm Formation on Abiotic Surfaces. Phytochem. Rev. 2020. [Google Scholar] [CrossRef]
- EFSA; European Food Safety Authority and European Centre for Disease Prevention and Control (EFSA and ECDC). The European Union One Health 2018 Zoonoses Report. EFSA J. 2019, 17, e05926. [Google Scholar]
- Chlebicz, A.; Śliżewska, K. Campylobacteriosis, Salmonellosis, Yersiniosis, and Listeriosis as Zoonotic Foodborne Diseases: A Review. Int. J. Environ. Res. Public Health 2018, 15, 863. [Google Scholar] [CrossRef] [PubMed]
- Galié, S.; García-Gutiérrez, C.; Miguélez, E.M.; Villar, C.J.; Lombó, F. Biofilms in the Food Industry: Health Aspects and Control Methods. Front. Microbiol. 2018, 898. [Google Scholar] [CrossRef]
- CDC. Centre for Disease Control, USA: Listeria (Listeriosis). 2017. Available online: https://www.cdc.gov/listeria/outbreaks/index.html (accessed on 29 November 2020).
- Rothrock, M.J.; Davis, M.L.; Locatelli, A.; Bodie, A.; McIntosh, T.G.; Donaldson, J.R.; Ricke, S.C. Listeria Occurrence in Poultry Flocks: Detection and Potential Implications. Front. Vet. Sci. 2017, 4. [Google Scholar] [CrossRef]
- Nguyen, H.D.N.; Yang, Y.S.; Yuk, H.G. Biofilm Formation of Salmonella Typhimurium on Stainless Steel and Acrylic Surfaces as Affected by Temperature and PH Level. LWT Food Sci. Technol. 2014, 55, 383–388. [Google Scholar] [CrossRef]
- Duguid, J.P.; Anderson, E.S.; Campbell, I. Fimbriae and Adhesive Properties in Salmonellae. J. Pathol. Bacteriol. 1966, 92, 107–137. [Google Scholar] [CrossRef]
- Giaouris, E.; Heir, E.; Desvaux, M.; Hébraud, M.; Møretrø, T.; Langsrud, S.; Doulgeraki, A.; Nychas, G.-J.; Kačániová, M.; Czaczyk, K.; et al. Intra- and Inter-Species Interactions within Biofilms of Important Foodborne Bacterial Pathogens. Front. Microbiol. 2015, 6. [Google Scholar] [CrossRef]
- Kadariya, J.; Smith, T.C.; Thapaliya, D. Staphylococcus Aureus and Staphylococcal Food-Borne Disease: An Ongoing Challenge in Public Health. Biomed Res. Int. 2014. [Google Scholar] [CrossRef]
- Hammer, B.K.; Bassler, B.L. Quorum sensing controls biofilm formation in Vibrio cholerae. Mol. Microbiol. 2003, 50, 101–114. [Google Scholar] [CrossRef]
- Otto, M. Staphylococcal infections: Mechanisms of biofilm maturation and detachment as critical determinants of pathogenicity. Annu. Rev. Med. 2013, 64, 175–188. [Google Scholar] [CrossRef] [PubMed]
- Morgan, R.; Kohn, S.; Hwang, S.H.; Hassett, D.J.; Sauer, K. BdlA, a chemotaxis regulator essential for biofilm dispersion in Pseudomonas aeruginosa. J. Bacteriol. 2006, 188, 7335–7343. [Google Scholar] [CrossRef] [PubMed]
- Somerton, B.; Flint, S.; Palmer, J.; Brooks, J.; Lindsay, D. Preconditioning with cations increases the attachment of Anoxybacillus flavithermus and Geobacillus species to stainless steel. Appl. Environ. Microbiol. 2013, 79, 4186–4190. [Google Scholar] [CrossRef]
- Houry, A.; Briandet, R.; Aymerich, S.; Gohar, M. Involvement of Motility and Flagella in Bacillus Cereus Biofilm Formation. Microbiology 2010, 156, 1009–1018. [Google Scholar] [CrossRef]
- Téllez, S. Biofilms and their impact on food industry. In VISAVET Outreach Journal; Complutense University: Madrid, Spain, 2010. [Google Scholar]
- Tram, G.; Day, C.J.; Korolik, V. Bridging the Gap: A Role for Campylobacter Jejuni Biofilms. Microorganisms 2020, 8, 452. [Google Scholar] [CrossRef]
- Gould, L.H.; Mody, R.K.; Ong, K.L.; Clogher, P.; Cronquist, A.B.; Garman, K.N.; Lathrop, S.; Medus, C.; Spina, N.L.; Webb, T.H.; et al. Increased recognition of non-O157 Shiga toxin-producing Escherichia coli infections in the United States during 2000–2010: Epidemiologic features and comparison with E. coli O157 infections. Foodborne Pathog. Dis. 2013, 10, 453–460. [Google Scholar] [CrossRef] [PubMed]
- Lim, E.S.; Koo, O.K.; Kim, M.J.; Kim, J.S. Bio-enzymes for inhibition and elimination of Escherichia coli O157: H7 biofilm and their synergistic effect with sodium hypochlorite. Sci. Rep. 2019, 9, 9920. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.; Bansal, T.; Jayaraman, A.; Bentley, W.E.; Wood, T.K. Enterohemorrhagic Escherichia Coli Biofilms Are Inhibited by 7-Hydroxyindole and Stimulated by Isatin. Appl. Environ. Microbiol. 2007, 73, 4100–4109. [Google Scholar] [CrossRef]
- Milillo, S.R.; Friedly, E.C.; Saldivar, J.C.; Muthaiyan, A.; O’bryan, C.; Crandall, P.G.; Johnson, M.G.; Ricke, S.C. A Review of the Ecology, Genomics, and Stress Response of Listeria Innocua and Listeria Monocytogenes. Crit. Rev. Food Sci. Nutr. 2012, 52, 712–725. [Google Scholar] [CrossRef]
- Chmielewski, R.A.N.; Frank, J.F. Biofilm Formation and Control in Food Processing Facilities. Compr. Rev. Food Sci. Food Saf. 2003, 2, 22–32. [Google Scholar] [CrossRef]
- Raheem, D. Outbreaks of Listeriosis Associated with Deli Meats and Cheese: An Overview. Aims Microbiol. 2016, 2, 230–250. [Google Scholar] [CrossRef]
- Lemon, K.P.; Higgins, D.E.; Kolter, R. Flagellar Motility Is Critical for Listeria Monocytogenes Biofilm Formation. J. Bacteriol. 2007, 189, 4418–4424. [Google Scholar] [CrossRef] [PubMed]
- Giannella, R.A.; Baron, S.; Albrecht, T.; Castro, G.; Couch, R.B.; Davis, C.P.; Dianzani, F.; Mcginnis, M.R.; Niesel, D.W.; Woods, G.W. Salmonella. In Baron’s Medical Microbiology, 4th ed.; University of Texas Medical Branch: Galveston, TX, USA, 1996. [Google Scholar]
- Lamas, A.; Miranda, J.M.; Regal, P.; Vázquez, B.; Franco, C.M.; Cepeda, A. A Comprehensive Review of Non-Enterica Subspecies of Salmonella Enterica. Microbiol. Res. 2018, 60–73. [Google Scholar] [CrossRef]
- Ćwiek, K.; Bugla-Płoskońska, G.; Wieliczko, A. Salmonella Biofilm Development: Structure and Significance. Postepy Hig. I Med. Dosw. 2019, 937–943. [Google Scholar] [CrossRef]
- Wang, H.; Ding, S.; Wang, G.; Xu, X.; Zhou, G. In situ characterization and analysis of Salmonella biofilm formation under meat processing environments using a combined microscopic and spectroscopic approach. Int. J. Food Microbiol. 2013, 167, 293–302. [Google Scholar] [CrossRef]
- Schelin, J.; Susilo, Y.; Johler, S. Expression of Staphylococcal Enterotoxins under Stress Encountered during Food Production and Preservation. Toxins 2017, 9, 401. [Google Scholar] [CrossRef]
- Amina, M.; Bensoltane, A. Review of Pseudomonas Attachment and Biofilm Formation in Food Industry Pseudomonas Biofilm; Chemotaxis and Motility View Project. Poult. Fish Wildl. Sci. 2015, 3, 1. [Google Scholar] [CrossRef]
- Golovlev, E.L. The Mechanism of Formation of Pseudomonas Aeruginosa Biofilm, a Type of Structured Population. Microbiology 2002, 249–254. [Google Scholar] [CrossRef]
- Carrascosa, C.; Millán, R.; Jaber, J.R.; Lupiola, P.; del Rosario-Quintana, C.; Mauricio, C.; Sanjuán, E. Blue Pigment in Fresh Cheese Produced by Pseudomonas fluorescens. Food Control 2015, 54, 95–102. [Google Scholar] [CrossRef]
- Wu, P.; Guo, Y.; Golly, M.K.; Ma, H.; He, R.; Luo, S.; Zhang, C.; Zhang, L.; Zhu, J. Feasibility Study on Direct Fermentation of Soybean Meal by Bacillus Stearothermophilus under Non-sterile Conditions. J. Sci. Food Agric. 2019, 99, 3291–3298. [Google Scholar] [CrossRef]
- Flint, S.; Palmer, J.; Bloemen, K.; Brooks, J.; Crawford, R. The Growth of Bacillus Stearothermophilus on Stainless Steel. J. Appl. Microbiol. 2001, 90, 151–157. [Google Scholar] [CrossRef]
- Palmer, J.S.; Flint, S.H.; Schmid, J.; Brooks, J.D. The Role of Surface Charge and Hydrophobicity in the Attachment of Anoxybacillus Flavithermus Isolated from Milk Powder. J. Ind. Microbiol. Biotechnol. 2010, 37, 1111–1119. [Google Scholar] [CrossRef]
- Strejc, J.; Kyselova, L.; Cadkova, A.; Matoulkova, D.; Potocar, T.; Branyik, T. Experimental Adhesion of Geobacillus Stearothermophilus and Anoxybacillus Flavithermus to Stainless Steel Compared with Predictions from Interaction Models. Chem. Pap. 2020, 74, 297–304. [Google Scholar] [CrossRef]
- Murphy, P.M.; Lynch, D.; Kelly, P.M. Growth of Thermophilic Spore Forming Bacilli in Milk during the Manufacture of Low Heat Powders. Int. J. Dairy Technol. 1999, 52, 45–50. [Google Scholar] [CrossRef]
- Sadiq, F.A.; Flint, S.; Yuan, L.; Li, Y.; Liu, T.J.; He, G.Q. Propensity for Biofilm Formation by Aerobic Mesophilic and Thermophilic Spore Forming Bacteria Isolated from Chinese Milk Powders. Int. J. Food Microbiol. 2017, 262, 89–98. [Google Scholar] [CrossRef]
- Burgess, S.A.; Brooks, J.D.; Rakonjac, J.; Walker, K.M.; Flint, S.H. The Formation of Spores in Biofilms of Anoxybacillus Flavithermus. J. Appl. Microbiol. 2009, 107, 1012–1018. [Google Scholar] [CrossRef] [PubMed]
- Paradh, A.D.; Mitchell, W.J.; Hill, A.E. Occurrence of Pectinatus and Megasphaera in the Major UK Breweries. J. Inst. Brew. 2011, 117, 498–506. [Google Scholar] [CrossRef]
- Lee, S.Y.; Mabee, M.S.; Jangaard, N.O. Pectinatus, a New Genus of the Family Bacteroidaceae. Int. J. Syst. Bacteriol. 1978, 28, 582–594. [Google Scholar] [CrossRef]
- Sterniša, M.; Klančnik, A.; Smole Možina, S. Spoilage Pseudomonas biofilm with Escherichia coli protection in fish meat at 5 °C. J. Sci. Food Agric. 2019, 99, 4635–4641. [Google Scholar] [CrossRef]
- Mizan, M.F.R.; Jahid, I.K.; Ha, S.D. Microbial Biofilms in Seafood: A Food-Hygiene Challenge. Food Microbiol. 2015, 41–55. [Google Scholar] [CrossRef]
- Toushik, S.H.; Mizan, M.F.R.; Hossain, M.I.; Ha, S.D. Fighting with Old Foes: The Pledge of Microbe-Derived Biological Agents to Defeat Mono- and Mixed-Bacterial Biofilms Concerning Food Industries. Trends Food Sci. Technol. 2020, 413–425. [Google Scholar] [CrossRef]
- González, J.E.; Keshavan, N.D. Messing with Bacterial Quorum Sensing. Microbiol. Mol. Biol. Rev. 2006, 70, 859–875. [Google Scholar] [CrossRef] [PubMed]
- Zupančič, J.; Raghupathi, P.K.; Houf, K.; Burmølle, M.; Sørensen, S.J.; Gunde-Cimerman, N. Synergistic interactions in microbial biofilms facilitate the establishment of opportunistic pathogenic Fungi in household dishwashers. Front. Microbiol. 2018, 9, 21. [Google Scholar] [CrossRef] [PubMed]
- Pammi, M.; Liang, R.; Hicks, J.; Mistretta, T.A.; Versalovic, J. Biofilm extracellular DNA enhances mixed species biofilms of Staphylococcus epidermidis and Candida Albicans. BMC Microbiol. 2013, 13, 257. [Google Scholar] [CrossRef]
- Donlan, R.M.; Piede, J.A.; Heyes, C.D.; Sanii, L.; Murga, R.; Edmonds, P.; El-Sayed, I.; El-Sayed, M.A. Model System for Growing and Quantifying Streptococcus Pneumoniae Biofilms in Situ and in Real Time. Appl. Environ. Microbiol. 2004, 70, 4980–4988. [Google Scholar] [CrossRef] [PubMed]
- Heukelekian, H.; Heller, A. Relation between food concentration and surface forbacterial growth. J. Bacteriol. 1940, 40, 547–558. [Google Scholar] [CrossRef] [PubMed]
- Zobell, C.E. The effect of solid surfaces on bacterial activity. J. Bacteriol. 1943, 46, 39–56. [Google Scholar] [CrossRef] [PubMed]
- Characklis, W.G. Attached microbial growths-II. Frictional resistance due to microbial slimes. Water Res. 1973, 7, 1249–1258. [Google Scholar] [CrossRef]
- Moreno-García, J.; García-Martinez, T.; Moreno, J.; Mauricio, J.C.; Ogawa, M.; Luong, P.; Bisson, L.F. Impact of yeast flocculation and biofilm formation on yeast-fungus coadhesion in a novel immobilization system. Am. J. Enol. Vitic. 2018, 69, 278–288. [Google Scholar] [CrossRef]
- Ogawa, M.; Bisson, L.F.; García-Martínez, T.; Mauricio, J.C.; Moreno-García, J. New Insights on Yeast and Filamentous Fungus Adhesion in a Natural Co-Immobilization System: Proposed Advances and Applications in Wine Industry. Appl. Microbiol. Biotechnol. 2019, 103, 4723–4731. [Google Scholar] [CrossRef] [PubMed]
- Mowat, E.; Lang, S.; Williams, C.; McCulloch, E.; Jones, B.; Ramage, G. Phase-dependent antifungal activity against Aspergillus fumigatus developing multicelluar filamentous biofilms. J. Antimicrob. Chemother. 2008, 62, 1281–1284. [Google Scholar] [CrossRef]
- Mowat, E.; Butcher, J.; Lang, S.; Williams, C.; Ramage, G. Development of a simple model for studying the effects of antifungal agents on multicellular communities of Aspergillus fumigatus. J. Med. Microbiol. 2007, 56, 1205–1212. [Google Scholar] [CrossRef]
- European Hygienic Engineering and Design Group (EHEDG). Hygienic Design Principles, 3rd ed.; EHEDG Guidelines; European Hygienic Engineering & Design Group: Naarden, The Netherlands, 2018. [Google Scholar]
- Fortin, N.D. Regulatory Requirements in the United States on Hygiene Control in the Design, Construction, and Renovation of Food Processing Factories. Chapter 4 in Hygiene Control in the Design, Construction and Renovation of Food Processing Factories; Lelieveld, H.L.M., Holah, J., Eds.; Woodhead Publishing Ltd., 2011. Available online: https://ssrn.com/abstract=2497425 (accessed on 25 November 2020).
- Plieth, W. Electrochemistry for Materials Science; Elsevier: Amsterdam, The Netherlands, 2008; Chapter 12. [Google Scholar]
- Rosado-Castro, M.; da Silva Fernandes, M.; Kabuki, D.Y.; Kuaye, A.Y. Biofilm Formation of Enterococcus Faecium on Stainless Steel Surfaces: Modeling and Control by Disinfection Agents. J. Food Process Eng. 2018, 41, 1–9. [Google Scholar] [CrossRef]
- Rosenberg, M.; Kjelleberg, S. Hydrophobic interactions in bacterial adhesion. Adv. Microb. Ecol. 1986, 9, 353–393. [Google Scholar]
- Corpe, W.A. Microbial surface components involved in adsorption of microorganisms onto surfaces. In Adsorption of Microorganisms to Surfaces; Bitton, G., Marshall, K.C., Eds.; John Wiley & Sons: New York, NY, USA, 1980; pp. 105–144. [Google Scholar]
- Peña, W.E.L.; Andrade, N.J.; Soares, N.F.F.; Alvarenga, V.O.; Rodrigues Junior, S.; Granato, D.; Zuniga, A.D.G.; de Souza Sant’Ana, A. Modeling Bacillus cereus adhesion on stainless steel surface as affected by temperature, pH and time. Int. Dairy J. 2014, 34, 153–158. [Google Scholar] [CrossRef]
- Furukawa, S.; Akiyoshi, Y.; Komoriya, M.; Ogihara, H.; Morinaga, Y. Removing Staphylococcus aureus and Escherichia coli biofilms on stainless steel by cleaning-in-place (CIP) cleaning agents. Food Control 2010, 21, 669–672. [Google Scholar] [CrossRef]
- Bore, E.; Langsrud, S. Characterization of Micro-Organisms Isolated from Dairy Industry after Cleaning and Fogging Disinfection with Alkyl Amine and Peracetic Acid. J. Appl. Microbiol. 2005, 98, 96–105. [Google Scholar] [CrossRef]
- Hawkins, C.L.; Davies, M.J. Hypochlorite-induced oxidation of proteins in plasma: Formation of chloramines and nitrogen-centred radicals and their role in protein fragmentation. Biochem. J. 1999, 340, 539–548. [Google Scholar] [CrossRef] [PubMed]
- Nightingale, Z.D.; Lancha, A.H., Jr.; Handelman, S.K.; Dolnikowski, G.G.; Busse, S.C.; Dratz, E.A.; Blumberg, J.B.; Handelman, G.J. Relative reactivity of lysine and other peptide-bound amino acids to oxidation by hypochlorite. Free Radic. Biol. Med. 2000, 29, 425–433. [Google Scholar]
- Heinecke, J.W.; Li, W.; Daehnke, H.; Goldstein, J.A. Dityrosine, a specific marker of oxidation, is synthesized by the myeloperoxidase-hydrogen peroxide system of human neutrophils and macrophages. J. Biol. Chem. 1993, 268, 4069–4077. [Google Scholar] [CrossRef]
- Spickett, C.M.; Jerlich, A.; Panasenko, O.M.; Arnhold, J.; Pitt, A.R.; Stelmaszynska, T.; Schaur, R.J. The reactions of hypochlorous acid, the reactive oxygen species produced by myeloperoxidase, with lipids. Acta Biochim. Pol. 2000, 47, 889–899. [Google Scholar] [CrossRef]
- Prutz, W.A. Interactions of hypochlorous acid with pyrimidine nucleotides, and secondary reactions of chlorinated pyrimidines with GSH, NADH, and other substrates. Arch. Biochem. Biophys. 1998, 349, 183–191. [Google Scholar] [CrossRef] [PubMed]
- Fukuzaki, S. Mechanisms of actions of sodium hypochlorite in cleaning and disinfection processes. Biocontrol Sci. 2006, 11, 147–157. [Google Scholar] [CrossRef] [PubMed]
- Fernandes, M.S.; Fujimoto, G.; Souza, L.P.; Kabuki, D.Y.; Silva, M.J.; Kuaye, A.Y. Dissemination of Enterococcus faecalis and Enterococcus faecium in a ricotta processing plant and evaluation of pathogenic and antibiotic resistance profiles. J. Food Sci. 2015, 80, M765–M775. [Google Scholar] [CrossRef] [PubMed]
- Skowron, K.; Hulisz, K.; Gryń, G.; Olszewska, H.; Wiktorczyk, N.; Paluszak, Z. Comparison of selected disinfectants efficiency against Listeria monocytogenes biofilm formed on various surfaces. Int. Microbiol. 2018, 21, 23–33. [Google Scholar] [CrossRef] [PubMed]
- Ibusquisa, P.A.; Herrera, J.J.R.; Cabo, M.L. Resistance to benzalkonium chloride, peracetic acid and nisin during formation of mature biofilms by List. Monocytogenes. Food Microbiol. 2011, 28, 418–428. [Google Scholar] [CrossRef]
- Eterpi, M.; McDonnell, G.; Thomas, V. Disinfection efficacy against parvoviruses compared with reference viruses. J. Hosp. Infect. 2009, 73, 64–70. [Google Scholar] [CrossRef]
- Langsrud, S.; Sidhu, M.S.; Heir, E.; Holck, A.L. Bacterial Disinfectant Resistance—A Challenge for the Food Industry. Int. Biodeterior. Biodegrad. 2003, 51, 283–290. [Google Scholar] [CrossRef]
- Rasimus, S.; Kolari, M.; Rita, H.; Hoornstra, D.; Salkinoja-Salonen, M. Biofilm-Forming Bacteria with Varying Tolerance to Peracetic Acid from a Paper Machine. J. Ind. Microbiol. Biotechnol. 2011, 38, 1379–1390. [Google Scholar] [CrossRef] [PubMed]
- Russell, A.D. Similarities and differences in the responses of microorganisms to biocides. J. Antimicrob. Chemother. 2003, 52, 750–763. [Google Scholar] [CrossRef]
- Block, S.S. Peroxygen compounds. In Disinfection, Sterilization, and Preservation, 5th ed.; Lippincott Williams & Wilkins: Philadelphia, PA, USA, 2001; pp. 185–204. [Google Scholar]
- Marquis, R.E.; Rutherford, G.C.; Faraci, M.M.; Shin, S.Y. Sporicidal action of peracetic acid and protective eVects of transition metal ions. J. Ind. Microbiol. 1995, 15, 486–492. [Google Scholar] [CrossRef]
- Dunkin, N.; Weng, S.; Schwab, K.J.; McQuarrie, J.; Bell, K.; Jacangelo, J.G. Comparative inactivation of murine norovirus and MS2 bacteriophage by peracetic acid and monochloramine in municipal secondary wastewater effluent. Environ. Sci. Technol. 2017, 51, 2972–2981136. [Google Scholar] [CrossRef]
- Zhang, C.; Brown, P.J.B.; Miles, R.J.; White, T.A.; Grant, D.A.G.; Stalla, D.; Hu, Z. Inhibition of Regrowth of Planktonic and Biofilm Bacteria after Peracetic Acid Disinfection. Water Res. 2019, 149, 640–649. [Google Scholar] [CrossRef] [PubMed]
- Bridier, A.; Briandet, R.; Thomas, V.; Dubois-Brissonnet, F. Comparative biocidal activity of peracetic acid, benzalkonium chloride and ortho-phthalaldehyde on 77 bacterial strains. J. Hosp. Infect. 2011, 78, 208–213. [Google Scholar] [CrossRef] [PubMed]
- Vázquez-Sánchez, D.; Cabo, M.L.; Ibusquiza, P.S.; Rodríguez-Herrera, J.J. Biofilm-forming ability and resistance to industrial disinfectants of Staphylococcus aureus isolated from fishery products. Food Control. 2014, 39, 8–16. [Google Scholar] [CrossRef]
- Mah, T.F.C.; O’Toole, G.A. Mechanisms of biofilm resistance to antimicrobial agents. Trends Microbiol. 2001, 9, 34–39. [Google Scholar] [CrossRef]
- Marić, S.; Vraneš, J. Characteristics and Significance of Microbial Biofilm Formation. Period. Biol. 2007, 109, 115–121. [Google Scholar]
- Beveridge, T.J.; Makin, S.A.; Kadurugamuwa, J.L.; Li, Z. Interactions between biofilms and the environment. FEMS Microbiol. Rev. 1997, 20, 291–303. [Google Scholar] [CrossRef]
- Flemming, H.C. Biofilms and environmental protection. Water Sci. Technol. 1993, 27, 1–10. [Google Scholar] [CrossRef]
- Da Costa, J.B.; Rodgher, S.; Daniel, L.A.; Espíndola, E.L.G. Toxicity on aquatic organisms exposed to secondary effluent disinfected with chlorine, peracetic acid, ozone and UV radiation. Ecotoxicology 2014, 23, 1803–1813. [Google Scholar] [CrossRef]
- Ferriol-González, C.; Domingo-Calap, P. Phages for Biofilm Removal. Antibiotics 2020, 9, 268. [Google Scholar] [CrossRef]
- Hiom, S.J.; Lowe, C.; Oldcorne, M. Assessment of surface bioburden during hospital aseptic processing. Int. J. Pharm. Pract. 2003, 11, R62. [Google Scholar]
- Wang, Y.; Tan, X.; Xi, C.; Phillips, K.S. Removal of Staphylococcus aureus from skin using a combination antibiofilm approach. NPJ Biofilms Microbiomes. 2018, 4, 1–9. [Google Scholar] [CrossRef]
- Vaidya, M.Y.; McBain, A.J.; Butler, J.A.; Banks, C.E.; Whitehead, K.A. Antimicrobial efficacy and synergy of metal ions against Enterococcus faecium, Klebsiella pneumoniae and Acinetobacter baumannii in planktonic and biofilm phenotypes. Sci. Rep. 2017, 7, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Miao, J.; Liang, Y.; Chen, L.; Wang, W.; Wang, J.; Li, B.; Li, L.; Chen, D.; Xu, Z. Formation and development of Staphylococcus biofilm: With focus on food safety. J. Food Saf. 2017, 37, e12358. [Google Scholar] [CrossRef]
- Cacciatore, F.A.; Brandelli, A.; Malheiros, P.D.S. Combining natural antimicrobials and nanotechnology for disinfecting food surfaces and control microbial biofilm formation. Crit. Rev. Food Sci. Nutr. 2020, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Iacumin, L.; Manzano, M.; Comi, G. Phage Inactivation of Listeria Monocytogenes on San Daniele Dry-Cured Ham and Elimination of Biofilms from Equipment and Working Environments. Microorganisms 2016, 4, 4. [Google Scholar] [CrossRef] [PubMed]
- Greer, G.G. Bacteriophage control of foodborne bacteria. J. Food Prot. 2011, 68, 1102–1111. [Google Scholar] [CrossRef]
- Soni, K.A.; Nannapaneni, R. Bacteriophage significantly reduces L. monocytogenes on raw salmon fillet tissue. J. Food Prot. 2010, 73, 32–38. [Google Scholar] [CrossRef]
- Gironés, R.R.G.; Simmons, M.M. Evaluation of the safety and efficacy of ListexTM P100 for reduction of pathogens on different ready-to-eat (RTE) food products. EFSA J. 2016, 14, 4565. [Google Scholar]
- Marsden, J.L. The Effectiveness of Listex P100 in Reducing Listeria monocytogenes in RTE Food; Food Science Institute, Kansas State University; Available online: https://www.foodsafetynews.com/files/2013/05/Listex-P100-Final-Report-1-2.pdf (accessed on 1 December 2020).
- Choi, N.Y.; Baek, S.Y.; Yoon, J.H.; Choi, M.R.; Kang, D.H.; Lee, S.Y. Efficacy of aerosolized hydrogen peroxide-based sanitizer on the reduction of pathogenic bacteria on a stainless steel surface. Food Control. 2012, 27, 57–63. [Google Scholar] [CrossRef]
- Park, S.H.; Cheon, H.L.; Park, K.H.; Chung, M.S.; Choi, S.H.; Ryu, S.; Kang, D.H. Inactivation of biofilm cells of foodborne pathogen by aerosolized sanitizers. Int. J. Food Microbiol. 2012, 154, 130–134. [Google Scholar] [CrossRef]
- Del Pozo, J.L.; Patel, R. The challenge of treating biofilm-associated bacterial infections. Clin. Pharmacol. Ther. 2007, 82, 204–209. [Google Scholar] [CrossRef] [PubMed]
- Anghel, I.; Holban, A.M.; Grumezescu, A.M.; Andronescu, E.; Ficai, A.; Anghel, A.G.; Maganu, M.; Lazǎr, V.; Chifiriuc, M.C. Modified wound dressing with phyto-nanostructured coating to prevent staphylococcal and pseudomonal biofilm development. Nanoscale Res. Lett. 2012, 7, 1–8. [Google Scholar] [CrossRef] [PubMed]
- De Kievit, T.R.; Iglewski, B.H. Bacterial quorum sensing in pathogenic relationships. Infect. Immun. 2000, 68, 4839–4849. [Google Scholar] [CrossRef]
- Sajjan, U.; Moreira, J.; Liu, M.; Humar, A.; Chaparro, C.; Forstner, J.; Keshavjee, S. A novel model to study bacterial adherence to the transplanted airway: Inhibition of Burkholderia cepacia adherence to human airway by dextran and xylitol. J. Heart Lung Transplant. 2012, 23, 1382–1391. [Google Scholar] [CrossRef]
- Singh, P.K.; Schaefer, A.L.; Parsek, M.R.; Moninger, T.O.; Welsh, M.J.; Greenberg, E.P. Quorum-sensing signals indicate that cystic fibrosis lungs are infected with bacterial biofilms. Nature 2000, 407, 762–764. [Google Scholar] [CrossRef] [PubMed]
- Xiong, F.; Zhao, X.; Wen, D.; Li, Q. Effects of N-acyl-homoserine lactones-based quorum sensing on biofilm formation, sludge characteristics, and bacterial community during the start-up of bioaugmented reactors. Sci. Total Environ. 2020, 735, 139449. [Google Scholar] [CrossRef]
- Schuster, M.; Joseph Sexton, D.; Diggle, S.P.; Peter Greenberg, E. Acyl-homoserine lactone quorum sensing: From evolution to application. Annu. Rev. Microbiol. 2013, 67, 43–63. [Google Scholar] [CrossRef]
- Wang, J.; Liu, Q.; Ma, S.; Hu, H.; Wu, B.; Zhang, X.X.; Ren, H. Distribution characteristics of N-acyl homoserine lactones during the moving bed biofilm reactor biofilm development process: Effect of carbon/nitrogen ratio and exogenous quorum sensing signals. Bioresour. Technol. 2019, 289, 121591. [Google Scholar] [CrossRef]
- Maddela, N.R.; Sheng, B.; Yuan, S.; Zhou, Z.; Villamar-Torres, R.; Meng, F. Roles of quorum sensing in biological wastewater treatment: A critical review. Chemosphere 2019, 221, 616–629. [Google Scholar] [CrossRef]
- Pu, H.; Xu, Y.; Sun, D.-W.; Wei, Q.; Li, X. Optical nanosensors for biofilm detection in the food industry: Principles, applications and challenges. Crit. Rev. Food Sci. Nutr. 2020, 1–18. [Google Scholar] [CrossRef]
- Jayan, H.; Pu, H.; Sun, D.-W. Recent development in rapid detection techniques for microorganism activities in food matrices using bio-recognition: A review. Trends Food Sci. Technol. 2020, 95, 233–246. [Google Scholar] [CrossRef]
- Lv, M.; Liu, Y.; Geng, J.; Kou, X.; Xin, Z.; Yang, D. Engineering nanomaterials-based biosensors for food safety detection. Biosens. Bioelectron. 2018, 106, 122–128. [Google Scholar] [CrossRef] [PubMed]
- Zhang, P.; Chen, Y.P.; Qiu, J.H.; Dai, Y.Z.; Feng, B. Imaging the microprocesses in biofilm matrices. Trends Biotechnol. 2019, 37, 214–226. [Google Scholar] [CrossRef]
- Ivleva, N.P.; Kubryk, P.; Niessner, R. Raman microspectroscopy, surface-enhanced Raman scattering microspectroscopy, and stable-isotope Raman microspectroscopy for biofilm characterization. Anal. Bioanal. Chem. 2017, 409, 4353–4375. [Google Scholar] [CrossRef]
- Vanegas, D.C.; Gomes, C.L.; Cavallaro, N.D.; Giraldo-Escobar, D.; McLamore, E.S. Emerging biorecognition and transduction schemes for rapid detection of pathogenic bacteria in food. Compr. Rev. Food Sci. Food Saf. 2017, 16, 1188–1205. [Google Scholar] [CrossRef] [PubMed]
- Pantanella, F.; Valenti, P.; Natalizi, T.; Passeri, D.; Berlutti, F. Analytical Techniques to Study Microbial Biofilm on Abiotic Surfaces: Pros and Cons of the Main Techniques Currently in Use. Ann. Ig. Med. Prev. E Comunità 2013, 25, 31–42. [Google Scholar] [CrossRef]
- Azeredo, J.; Azevedo, N.F.; Briandet, R.; Cerca, N.; Coenye, T.; Costa, A.R.; Desvaux, M.; Di Bonaventura, G.; Hébraud, M.; Jaglic, Z.; et al. Critical Review on Biofilm Methods. Crit. Rev. Microbiol. 2017, 43, 313–351. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.; Chakraborty, S.; Liang, H. Methods to Probe the Formation of Biofilms: Applications in Foods and Related Surfaces. Anal. Methods 2020, 12, 416–432. [Google Scholar] [CrossRef]
- Nijland, R.; Hall, M.J.; Burgess, J.G. Dispersal of biofilms by secreted, matrix degrading, bacterial DNase. PLoS ONE 2010, 5, e15668. [Google Scholar] [CrossRef] [PubMed]
- Jamal, M.; Ahmad, W.; Andleeb, S.; Jalil, F.; Imran, M.; Nawaz, M.A.; Hussain, T.; Ali, M.; Rafiq, M.; Kamil, M.A. Bacterial Biofilm and Associated Infections. J. Chin. Med. Assoc. 2018, 81, 7–11. [Google Scholar] [CrossRef]
- Ansari, M.A.; Khan, H.M.; Khan, A.A.; Cameotra, S.S.; Alzohairy, M.A. Anti-biofilm efficacy of silver nanoparticles against MRSA and MRSE isolated from wounds in a tertiary care hospital. Indian J. Med. Microbiol. 2015, 33, 101. [Google Scholar] [CrossRef]
- Lawrence, J.R.; Neu, T.R. Confocal laser scanning microscopy for analysis of microbial biofilms. Methods Enzymol. 1999, 310, 131–144. [Google Scholar] [PubMed]
- Kania, R.E.; Lamers, G.E.; Vonk, M.J.; Huy, P.T.B.; Hiemstra, P.S.; Bloemberg, G.V.; Grote, J.J. Demonstration of bacterial cells and glycocalyx in biofilms on human tonsils. Arch. Otolaryngol. Head Neck Surg. 2007, 133, 115–121. [Google Scholar] [CrossRef]
- Ommen, P.; Zobek, N.; Meyer, R.L. Quantification of Biofilm Biomass by Staining: Non-Toxic Safranin Can Replace the Popular Crystal Violet. J. Microbiol. Methods 2017, 141, 87–89. [Google Scholar] [CrossRef]
- Stepanović, S.; Vuković, D.; Dakić, I.; Savić, B.; Švabić-Vlahović, M. A Modified Microtiter-Plate Test for Quantification of Staphylococcal Biofilm Formation. J. Microbiol. Methods 2000, 40, 175–179. [Google Scholar] [CrossRef]
- Doll, K.; Jongsthaphongpun, K.L.; Stumpp, N.S.; Winkel, A.; Stiesch, M. Quantifying Implant-Associated Biofilms: Comparison of Microscopic, Microbiologic and Biochemical Methods. J. Microbiol. Methods 2016, 130, 61–68. [Google Scholar] [CrossRef]
- Extremina, C.I.; Costa, L.; Aguiar, A.I.; Peixe, L.; Fonseca, A.P. Optimization of processing conditions for the quantification of enterococci biofilms using microtitreplates. J. Microbiol. Methods. 2010. [Google Scholar] [CrossRef]
- Stiefel, P.; Rosenberg, U.; Schneider, J.; Mauerhofer, S. Is biofilm removal properly assessed? Comparison of different quantification methods in a 96-well plate system. Appl. Microbiol. Biotechnol. 2016, 100, 4135–4145. [Google Scholar] [CrossRef]
- Romanova, N.A.; Gawande, P.V.; Brovko, L.Y.; Griffiths, M.W. Rapid Methods to Assess Sanitizing Efficacy of Benzalkonium Chloride to Listeria Monocytogenes Biofilms. J. Microbiol. Methods 2007, 71, 231–237. [Google Scholar] [CrossRef] [PubMed]
- Cell Staining. 2019. Available online: https://www.dojindo.com/ (accessed on 2 December 2020).
- Burton, E.; Yakandawala, N.; LoVetri, K.; Madhyastha, M.S. A Microplate Spectrofluorometric Assay for Bacterial Biofilms. J. Ind. Microbiol. Biotechnol. 2007, 34, 1–4. [Google Scholar] [CrossRef]
- Deng, Y.; Wang, L.; Chen, Y.; Long, Y. Optimization of Staining with SYTO 9/Propidium Iodide: Interplay, Kinetics and Impact on Brevibacillus Brevis. BioTechniques 2020, 69, 89–99. [Google Scholar] [CrossRef]
- Shi, L.; Gunther, S.; Hubschmann, T.; Wick, L.Y.; Harms, H.; Muller, S. Limits of propidium iodide as a cell viability indicator for environmental bacteria. Cytom. A 2007, 71, 592–598. [Google Scholar] [CrossRef] [PubMed]
- Munukka, E.; Leppäranta, O.; Korkeamäki, M.; Vaahtio, M.; Peltola, T.; Zhang, D.; Hupa, L.; Ylänen, H.; Salonen, J.I.; Viljanen, M.K.; et al. Bactericidal effects of bioactive glasses on clinically important aerobic bacteria. J. Mater. Sci. Mater. Med. 2008, 19, 27–32. [Google Scholar] [CrossRef] [PubMed]
- Larrosa, M.; Truchado, P.; Espin, J.C.; Tomás-Barberán, F.A.; Allende, A.; García-Conesa, M.T. Evaluation of Pseudomonas aeruginosa (PAO1) adhesion to human alveolar epithelial cells A549 using SYTO 9 dye. Mol. Cell. Probes. 2012, 26, 121–126. [Google Scholar] [CrossRef] [PubMed]
- Maukonen, J.; Mattila-Sandholm, T.; Wirtanen, G. Metabolic Indicators for Assessing Bacterial Viability in Hygiene Sampling Using Cells in Suspension and Swabbed Biofilm. LWT Food Sci. Technol. 2000, 33, 225–233. [Google Scholar] [CrossRef]
- Tawakoli, P.N.; Al-Ahmad, A.; Hoth-Hannig, W.; Hannig, M.; Hannig, C. Comparison of different live/dead stainings for detection and quantification of adherent microorganisms in the initial oral biofilm. Clin. Oral Investig. 2013, 17, 841–850. [Google Scholar] [CrossRef]
- Verran, J. Biofouling in food processing: Biofilm or biotransfer potential? Food Bioprod. Process. 2002, 80, 292–298. [Google Scholar] [CrossRef]
- Verran, J.; Whitehead, K.A. Assessment of organic materials and microbial components on hygienic surfaces. Food Bioprod. Process. 2006, 84, 260–264. [Google Scholar] [CrossRef][Green Version]
- Declerck, P.; Verelst, L.; Duvivier, L.; Van Damme, A.; Ollevier, F. A detection method for Legionella spp. in (cooling) water: Fluorescent in situ hybridisation (FISH) on whole bacteria. Water Sci. Technol. 2003, 47, 143–146. [Google Scholar] [CrossRef] [PubMed]
- Whitehead, K.A.; Benson, P.; Verran, J. Differential fluorescent staining of Listeria monocytogenes and a whey food soil for quantitative analysis of surface hygiene. Int. J. Food Microbiol. 2009, 135, 75–80. [Google Scholar] [CrossRef] [PubMed]
- Almeida, C.; Azevedo, N.F.; Santos, S.; Keevil, C.W.; Vieira, M.J. Discriminating Multi-Species Populations in Biofilms with Peptide Nucleic Acid Fluorescence in Situ Hybridization (PNA FISH). PLoS ONE 2011, 6. [Google Scholar] [CrossRef] [PubMed]
- Hansen, S.K.; Rainey, P.B.; Haagensen, J.A.; Molin, S. Evolution of species interactions in a biofilm community. Nature 2007, 445, 533–536. [Google Scholar] [CrossRef] [PubMed]
- Gu, F.; Lux, R.; Du-Thumm, L.; Stokes, I.; Kreth, J.; Anderson, M.H.; Wong, D.T.; Wolinsky, L.; Sullivan, R.; Shi, W. In situ and noninvasive detection of specific bacterial species in oral biofilms using fluorescently labeled monoclonal antibodies. J. Microbiol. Methods 2005, 62, 145–160. [Google Scholar] [CrossRef]
- Schlafer, S.; Meyer, R.L. Confocal Microscopy Imaging of the Biofilm Matrix. J. Microbiol. Methods. 2017, 138, 50–59. [Google Scholar] [CrossRef]
- Allesen-Holm, M.; Barken, K.B.; Yang, L.; Klausen, M.; Webb, J.S.; Kjelleberg, S.; Molin, S.; Givskov, M.; Tolker-Nielsen, T. A characterization of DNA release in Pseudomonas aeruginosa cultures and biofilms. Mol. Microbiol. 2006, 59, 1114–1128. [Google Scholar] [CrossRef]
- Conover, M.S.; Mishra, M.; Deora, R. Extracellular DNA is essential for maintaining Bordetella biofilm integrity on abiotic surfaces and in the upper respiratory tract of mice. PLoS ONE 2011, 6, e16861. [Google Scholar] [CrossRef]
- Okshevsky, M.; Meyer, R.L. Evaluation of fluorescent stains for visualizing extracellular DNA in biofilms. J. Microbiol. Methods 2014, 105, 102–104. [Google Scholar] [CrossRef]
- Schaeffer, C.R.; Woods, K.M.; Longo, G.M.; Kiedrowski, M.R.; Paharik, A.E.; Büttner, H.; Christner, M.; Boissy, R.J.; Horswill, A.R.; Rohde, H.; et al. Accumulation-associated protein enhances Staphylococcus epidermidis biofilm formation under dynamic conditions and is required for infection in a rat catheter model. Infect. Immun. 2015, 83, 214–226. [Google Scholar] [CrossRef]
- Büttner, H.; Mack, D.; Rohde, H. Structural basis of Staphylococcus epidermidis biofilm formation: Mechanisms and molecular interactions. Front. Cell. Infect. Microbiol. 2015, 5, 14. [Google Scholar] [CrossRef]
- Frank, K.L.; Patel, R. Poly-N-acetylglucosamine is not a major component of the extracellular matrix in biofilms formed by icaADBC-positive Staphylococcus lugdunensis isolates. Infect. Immun. 2007, 75, 4728–4742. [Google Scholar] [CrossRef]
- Toyofuku, M.; Roschitzki, B.; Riedel, K.; Eberl, L. Identification of proteins associated with the Pseudomonas aeruginosa biofilm extracellular matrix. J. Proteome Res. 2012, 11, 4906–4915. [Google Scholar] [CrossRef]
- Tseng, B.S.; Reichhardt, C.; Merrihew, G.E.; Araujo-Hernandez, S.A.; Harrison, J.J.; MacCoss, M.J.; Parsek, M.R. A biofilm matrix-associated protease inhibitor protects Pseudomonas aeruginosa from proteolytic attack. MBio 2018, 9. [Google Scholar] [CrossRef]
- Dige, I.; Nyengaard, J.R.; Kilian, M.; Nyvad, B. Application of stereological principles for quantification of bacteria in intact dental biofilms. Oral Microbiol. Immunol. 2009, 24, 69–75. [Google Scholar] [CrossRef] [PubMed]
- Malic, S.; Hill, K.E.; Hayes, A.; Percival, S.L.; Thomas, D.W.; Williams, D.W. Detection and identification of specific bacteria in wound biofilms using peptide nucleic acid fluorescent in situ hybridization (PNA FISH). Microbiology 2009, 155, 2603–2611. [Google Scholar] [CrossRef]
- Ivleva, N.P.; Wagner, M.; Horn, H.; Niessner, R.; Haisch, C. Towards a Nondestructive Chemical Characterization of Biofilm Matrix by Raman Microscopy. Anal. Bioanal. Chem. 2009, 393, 197–206. [Google Scholar] [CrossRef] [PubMed]
- Jung, G.B.; Nam, S.W.; Choi, S.; Lee, G.-J.; Park, H.-K. Evaluation of Antibiotic Effects on Pseudomonas aeruginosa Biofilm Using Raman Spectroscopy and Multivariate Analysis. Biomed. Opt. Express 2014, 5, 3238. [Google Scholar] [CrossRef]
- Fallatah, H.; Elhaneid, M.; Ali-Boucetta, H.; Overton, T.W.; El Kadri, H.; Gkatzionis, K. Antibacterial effect of graphene oxide (GO) nano-particles against Pseudomonas putida biofilm of variable age. Environ. Sci. Pollut. Res. 2019, 26, 25057–25070. [Google Scholar] [CrossRef]
- Vuotto, C.; Donelli, G. Field Emission Scanning Electron Microscopy of Biofilm-Growing Bacteria Involved in Nosocomial Infections. In Microbial Biofilms. Methods in Molecular Biology (Methods and Protocols); Donelli, G., Ed.; Humana Press: New York, NY, USA, 2014. [Google Scholar] [CrossRef]
- Kouider, N.; Hamadi, F.; Mallouki, B.; Bengoram, J.; Mabrouki, M.; Zekraoui, M.; Ellouali, M.; Latrache, H. Effect of stainless steel surface roughness on Staphylococcus aureus adhesion. Int. J. Pure Appl. Sci. 2010, 4, 17. [Google Scholar]
- Mallouki, B.; Latrache, H.; Mabrouki, M.; Outzourhit, A.; Hamadi, F.; Muller, D.; Ellouali, M. The inhibitory effect of fucans on adhesion and production of slime of Staphylococcus Aureus. Microbiol. Hygiène Aliment. 2007, 19, 6471. [Google Scholar]
- El Abed, S.; Ibnsouda, S.K.; Latrache, H.; Hamadi, F. Scanning electron microscopy (SEM) and environmental SEM: Suitable tools for study of adhesion stage and biofilm formation. In Scanning Electron Microscopy; Intechopen: London, UK, 2012. [Google Scholar]
- Agarwal, A.; Jern Ng, W.; Liu, Y. Removal of biofilms by intermittent low-intensity ultrasonication triggered bursting of microbubbles. Biofouling 2014, 30, 359–365. [Google Scholar] [CrossRef] [PubMed]
- Fricke, K.; Koban, I.; Tresp, H.; Jablonowski, L.; Schröder, K.; Kramer, A.; Weltmann, K.D.; von Woedtke, T.; Kocher, T. Atmospheric Pressure Plasma: A High-Performance Tool for the Efficient Removal of Biofilms. PLoS ONE 2012, 7, e42539. [Google Scholar] [CrossRef] [PubMed]
- Chan, T.F.; Esedoglu, S.; Nikolova, M. Algorithms for finding global minimizers of image segmentation and denoising models. Siam. J. Appl. Math. 2002, 66, 1632–1648. [Google Scholar] [CrossRef]
- Vyas, N.; Sammons, R.L.; Addison, O.; Dehghani, H.; Walmsley, A.D. A quantitative method to measure biofilm removal efficiency from complex biomaterial surfaces using SEM and image analysis. Sci. Rep. 2016, 6, 32694. [Google Scholar] [CrossRef]
- Hu, H.; Johani, K.; Gosbell, I.B.; Jacombs, A.S.W.; Almatroudi, A.; Whiteley, G.S.; Deva, A.K.; Jensen, S.; Vickery, K. Intensive care unit environmental surfaces are contaminated by multidrug-resistant bacteria in biofilms: Combined results of conventional culture, pyrosequencing, scanning electron microscopy, and confocal laser microscopy. J. Hosp. Infect. 2015, 91, 35–44. [Google Scholar] [CrossRef]
- Mohmmed, S.A.; Vianna, M.E.; Penny, M.R.; Hilton, S.T.; Mordan, N.; Knowles, J.C. Confocal laser scanning, scanning electron, and transmission electron microscopy investigation of Enterococcus faecalis biofilm degradation using passive and active sodium hypochlorite irrigation within a simulated root canal model. Microbiologyopen 2017, 6, e00455. [Google Scholar] [CrossRef] [PubMed]
- Bodelón, G.; Montes-García, V.; Costas, C.; Pérez-Juste, I.; Pérez-Juste, J.; Pastoriza-Santos, I.; Liz-Marzán, L.M. Imaging bacterial interspecies chemical interactions by surface-enhanced Raman scattering. ACS Nano 2017, 11, 4631–4640. [Google Scholar] [CrossRef]
- Whitehead, K.A.; Saubade, F.; Akhidime, I.D.; Liauw, C.M.; Benson, P.S.; Verran, J. The detection and quantification of food components on stainless steel surfaces following use in an operational bakery. Food Bioprod. Process. 2019, 116, 258–267. [Google Scholar] [CrossRef]
- Xu, H.S.; Roberts, N.; Singleton, F.L.; Attwell, R.W.; Grimes, D.J.; Colwell, R.R. Survival and viability of nonculturable Escherichia coli andVibrio cholerae in the estuarine and marine environment. Microb. Ecol. 1982, 8, 313–323. [Google Scholar] [CrossRef]
- Li, L.; Mendis, N.; Trigui, H.; Oliver, J.D.; Faucher, S.P. The importance of the viable but non-culturable state in human bacterial pathogens. Front. Microbiol. 2014, 5, 258. [Google Scholar] [CrossRef]
- Cerca, F.; Andrade, F.; França, Â.; Andrade, E.B.; Ribeiro, A.; Almeida, A.A.; Cerca, N.; Pier, G.; Azeredo, J.; Vilanova, M. Staphylococcus epidermidis biofilms with higher proportions of dormant bacteria induce a lower activation of murine macrophages. J. Med. Microbiol. 2011, 60, 1717–1724. [Google Scholar] [CrossRef]
- Oliveira, F.; Lima, C.A.; Brás, S.; França, Â.; Cerca, N. Evidence for interand intraspecies biofilm formation variability among a small group of coagulase-negative staphylococci. FEMS Microbiol. Lett. 2015, 362. [Google Scholar] [CrossRef]
- Bayoumi, M.A.; Kamal, R.M.; Abd El Aal, S.F.; Awad, E.I. Assessment of a regulatory sanitization process in Egyptian dairy plants in regard to the adherence of some food-borne pathogens and their biofilms. Int. J. Food Microbiol. 2012, 158, 225–231. [Google Scholar] [CrossRef] [PubMed]
- Khaddouj, A.; Fatima, H.; Rachida, M.; Hassan, L.; Khadija, A.; Youssef, N.; Mustapha, M. Evaluation of sodium hypochlorite efficiency on the elimination of Pseudomonas aeruginosa biofilm using two methods. Russ. Open Med. J. 2019, 8, e0302. [Google Scholar]
- Bremer, P.J.; Fillery, S.; McQuillan, A.J. Laboratory scale Clean-In-Place (CIP) studies on the effectiveness of different caustic and acid wash steps on the removal of dairy biofilms. Int. J. Food Microbiol. 2006, 106, 254–262. [Google Scholar] [CrossRef] [PubMed]
- Lebeaux, D.; Ghigo, J.M.; Beloin, C. Biofilm-related infections: Bridging the gap between clinical management and fundamental aspects of recalcitrance toward antibiotics. Microbiol. Mol. Biol. Rev. 2014, 78, 510–543. [Google Scholar] [CrossRef] [PubMed]
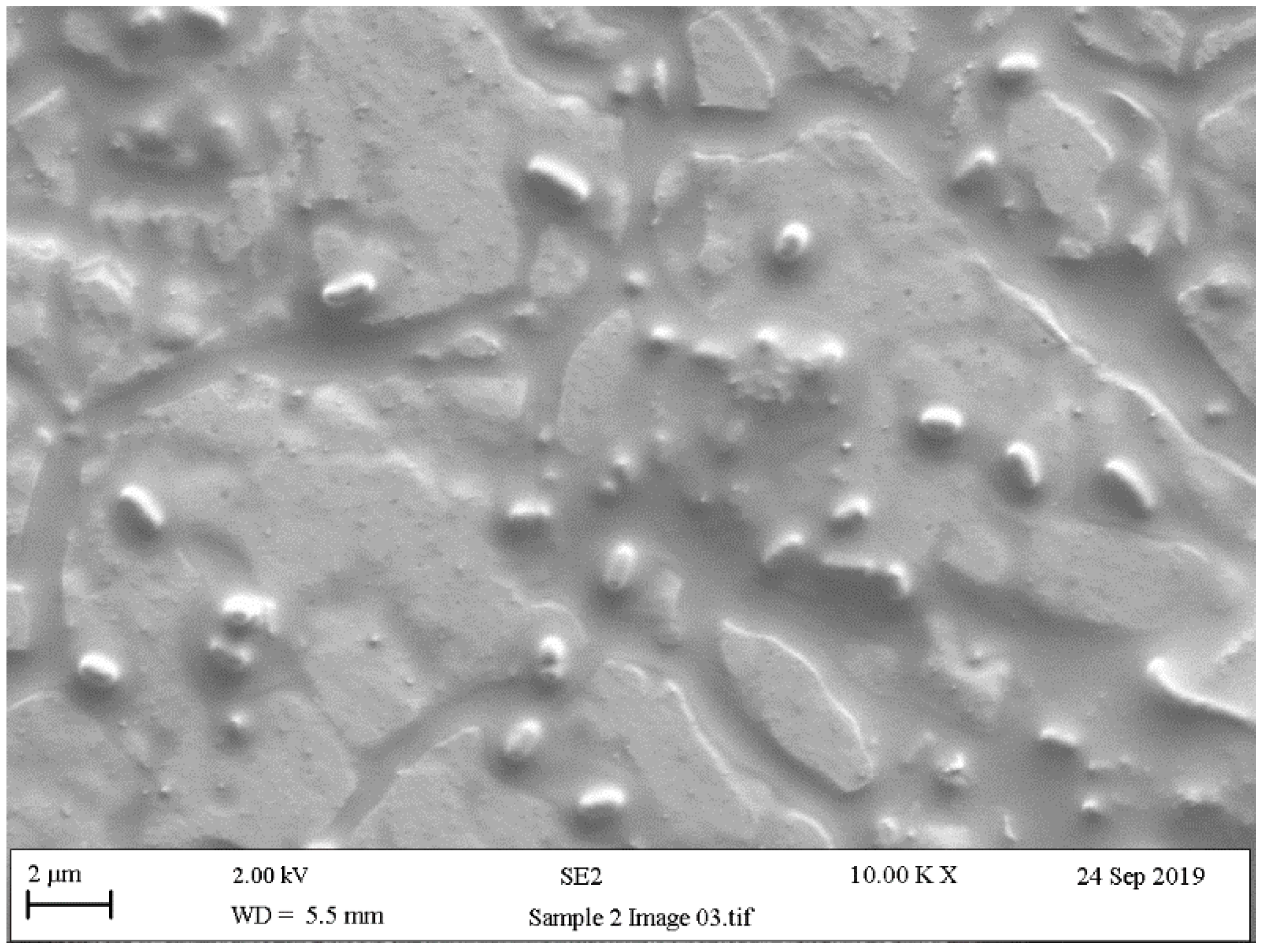

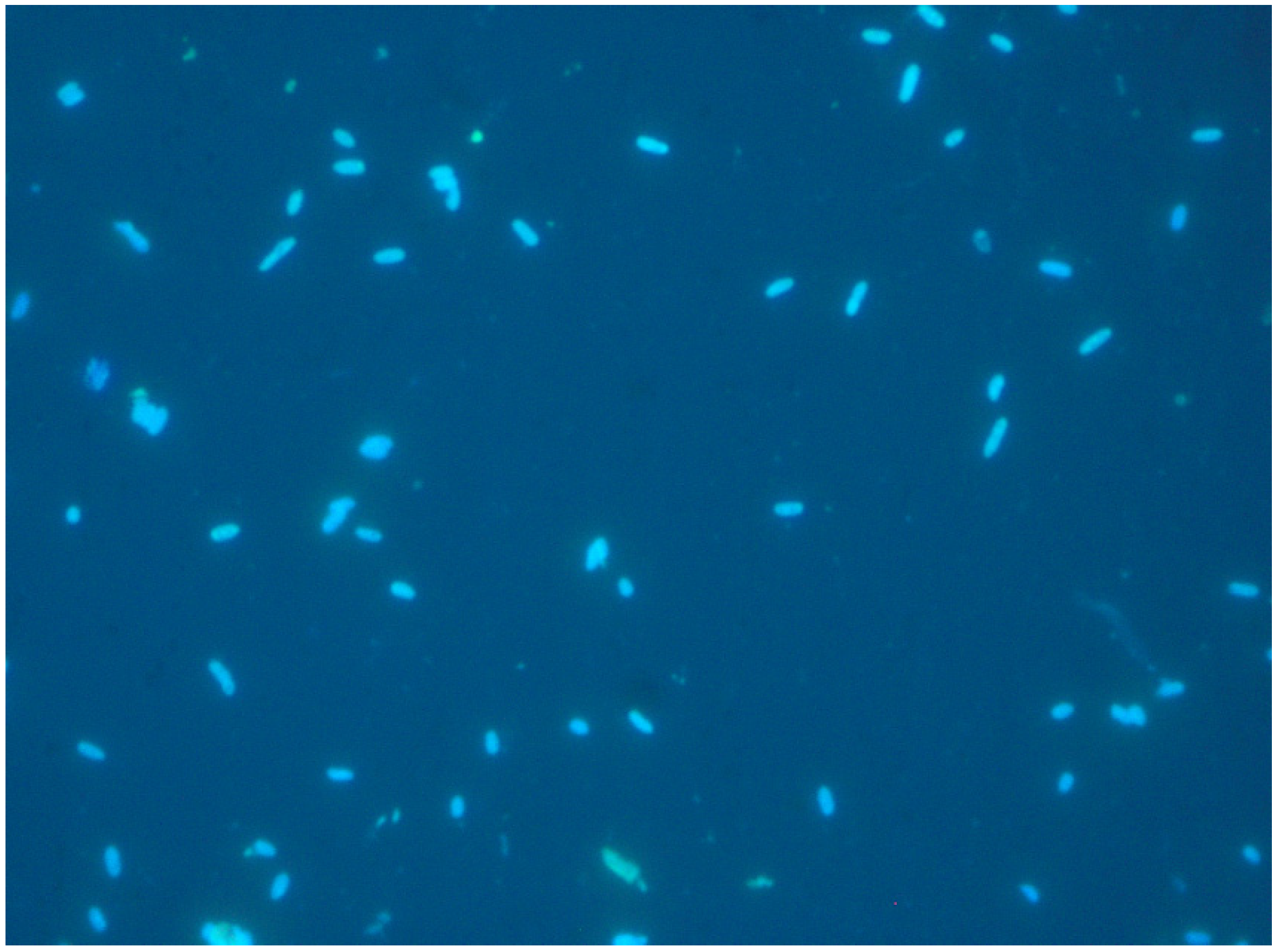
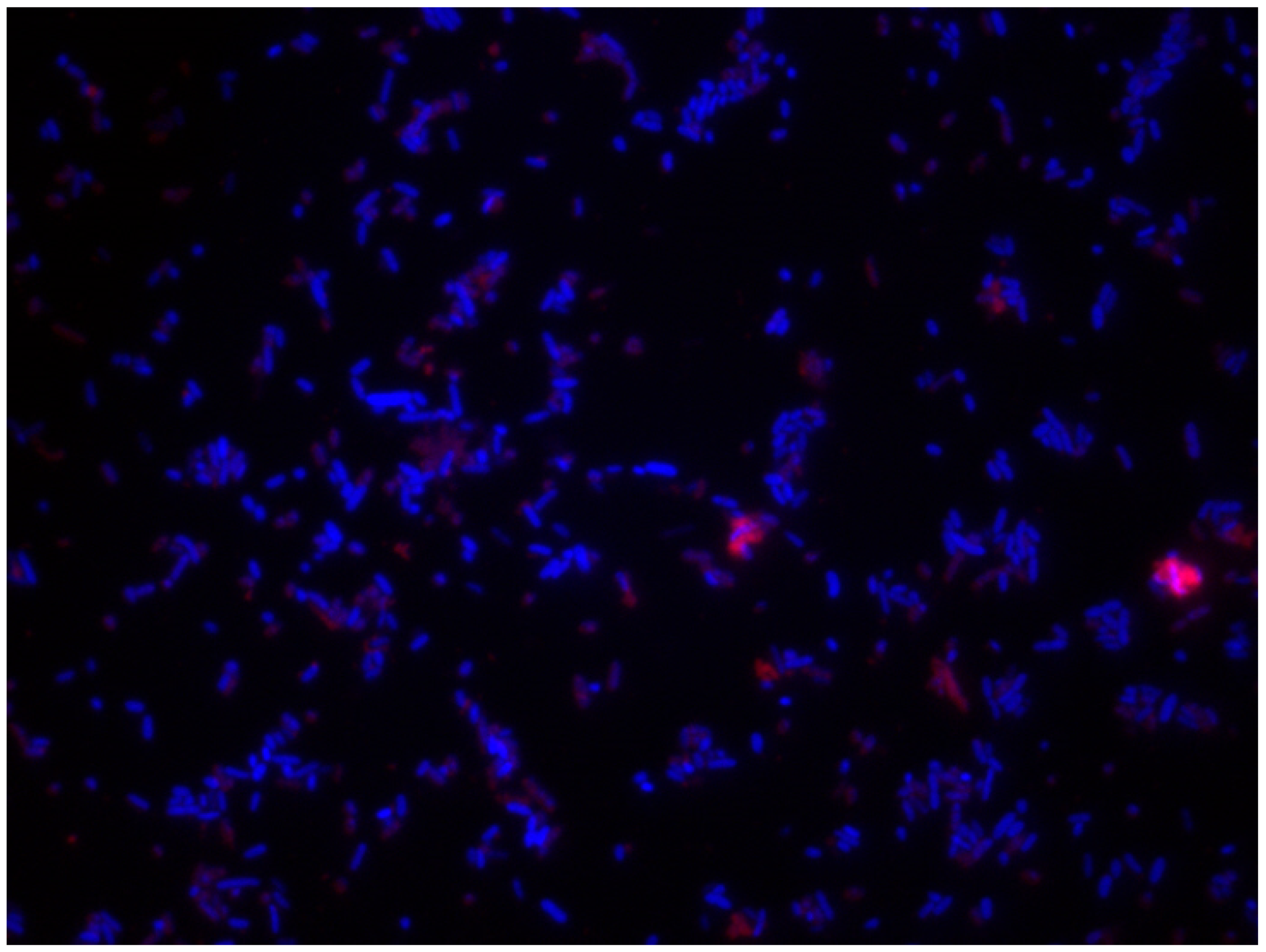
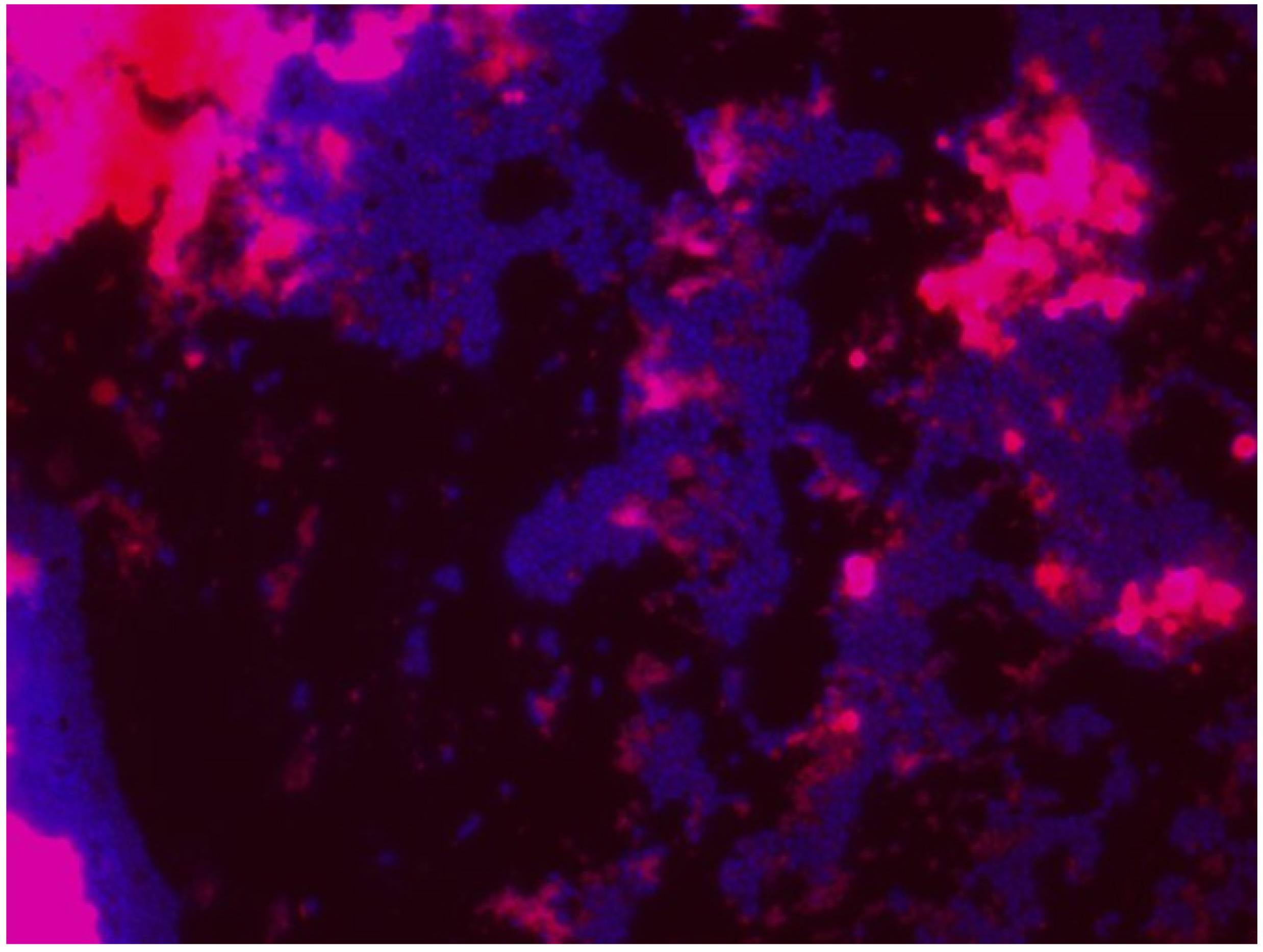



| Pathogen | Characteristics | Contaminated Food | Examples of Harmful Spoilage Effects | References |
|---|---|---|---|---|
| Bacillus cereus | Gram-positive, spore-forming, anaerobic, facultative anaerobic | dairy products, rice, vegetables, meat | diarrhoea and vomiting symptoms | [39,40] |
| Campylobacter jejuni | Gram-negative, aerobic and anaerobic | animals, poultry, unpasteurised milk | bloody diarrhoea, fever, stomach cramp, nausea and vomiting | [41,42,43] |
| Escherichia coli | Gram-negative, rod-shaped | raw milk, fresh meat, fruits and vegetables | diarrhoea outbreaks and haemolytic uremic syndrome | [44] |
| Listeria monocytogenes | Gram-positive, rod-shaped, facultative anaerobic | dairy products, meat, ready-to-eat products, fruit, soft cheeses, ice cream, unpasteurised milk, candied apples, frozen vegetables, poultry | listeriosis in the elderly, pregnant women and immune-compromised patients | [45,46] |
| Salmonella Enterica | Gram-negative, rod-shaped, flagellate, facultative aerobic | Poultry meat, bovine, ovine, porcine, fish | can cause gastroenteritis or septicaemia | [47,48] |
| Staphylococcus aureus | Gram-positive, non-spore forming, non-motile, facultative anaerobic | meat products, poultry, egg products, dairy products, salads, bakery products, especially cream-filled pastries and cakes, and sandwich fillings | methicillin resistance, can cause vomiting and diarrhoea | [49,50] |
| Pseudomonas spp. | psychrotrophic, motile, Gram-negative rod-shaped | fruits, vegetables, meat surfaces and low-acid dairy products | produces blue discolouration on fresh cheese. | [17] |
| Geobacillus stearothermophilus | thermophilic, Gram-positive, spore-forming, aerobic or facultative anaerobic | dried dairy products | production of acids or enzymes leading to off-flavours | [51,52] |
| Anoxybacillus flavithermus | thermophilic organism, Gram-positive, spore-forming, facultatively anaerobic, non-pathogenic | dried milk powder | an indicator of poor hygiene | [53,54] |
| Pectinatus spp. | Gram-negative, non-spore-forming, anaerobic | beer and brewery environment | rapid cell growth makes beer turbid and smells like rotten eggs due to production of sulphur compounds | [23] |
| Bacterial Species | Disinfectant | ppm or % | Surfaces | Biofilm. Log Reduction CFU | Reference |
|---|---|---|---|---|---|
| S. aureus | Sodium hypochlorite | 250 | Stainless steel/PP | 4.5/4.4 | [212] |
| Cronobacter Sakazakii | Sodium hypochlorite | 250 | Stainless steel/PP | 3.7/3.9 | [216] |
| S. Typhimurium | Sodium hypochlorite | 250 | Stainless steel/PP | 5.82/6.1 | [216] |
| S. aeruginosa | Sodium hypochlorite | 250, 500 | Stainless steel 316 | 2/100% | [217] |
| S. aeruginosa | Sodium hypochlorite | 750, 1000 | Stainless steel 316 | 100%/100% | [217] |
| B. cereus | NaOH and HNO3, 65 °C | 1% | CIP dairy | 2 | [218] |
| Enterococcus faecium | Sodium hypochlorite | 100 | Stainless steel | 3 | [98] |
| E. faecium | Peracetic acid | 300 | Stainless steel | 4 | [98] |
| Rhodococcus erythropolis | Alkyl amine | 1–1.3% | Stainless steel | >5 | [103] |
| R. erythropolis | Peracetic acid | 0.2% | Stainless steel | 0.48 | [103] |
| R. erythropolis | Sodium hypochlorite | 0.5–1% | Stainless steel | 4.51 | [103] |
| R. erythropolis | QAC | 200 | Stainless steel | >5 | [103] |
| Sphingomonas sp. | Alkyl amine | 1–1.3% | Stainless steel | >5 | [103] |
| Sphingomonas sp. | Peracetic acid | 0.2% | Stainless steel | >5 | [103] |
| Sphingomonas sp. | Sodium hypochlorite | 0.5–1% | Stainless steel | >5 | [103] |
| Sphingomonas sp. | QAC | 200 | Stainless steel | >5 | [103] |
| Methylobacterium rhodesianum | Alkyl amine | 1–1.3% | Stainless steel | 4.48 | [103] |
| M. rhodesianum | Peracetic acid | 0.2% | Stainless steel | >5 | [103] |
| M. rhodesianum | Sodium hypochlorite | 0.5–1% | Stainless steel | 0.01 | [103] |
| M. rhodesianum | QAC | 200 | Stainless steel | 0.64 | [103] |
| L. monocytogenes | Sodium hydroxide | 0.5% | Rubber | 0.66 | [112] |
| L. monocytogenes | QAC | 0.5% | Rubber | 1.72 | [112] |
| L. monocytogenes | Sodium hypochlorite | 0.5% | Rubber | 1.79 | [112] |
| L. monocytogenes | Peracetic acid | 0.5% | Rubber | 5.10 | [112] |
| L. monocytogenes | Sodium hydroxide | 0.5% | Polypropylene | 1.20 | [112] |
| L. monocytogenes | QAC | 0.5% | Polypropylene | 2.57 | [112] |
| L. monocytogenes | Sodium hypochlorite | 0.5% | Polypropylene | 2.74 | [112] |
| L. monocytogenes | Peracetic acid | 0.5% | Polypropylene | 6.62 | [112] |
| L. monocytogenes | Sodium hydroxide | 0.5% | Stainless steel | 1 | [112] |
| L. monocytogenes | QAC | 0.5% | Stainless steel | 4.06 | [112] |
| L. monocytogenes | Sodium hypochlorite | 0.5% | Stainless steel | 1.97 | [112] |
| L. monocytogenes | Peracetic acid | 0.5% | Stainless steel | 6.63 | [112] |
| L. monocytogenes | Sodium hydroxide | 0.5% | Aluminium foil | 0.52 | [112] |
| L. monocytogenes | QAC | 0.5% | Aluminium foil | 5.1 | [112] |
| L. monocytogenes | Sodium hypochlorite | 0.5% | Aluminium foil | 3.84 | [112] |
| L. monocytogenes | Peracetic acid | 0.5% | Aluminium foil | 6.54 | [112] |
| L. monocytogenes | Benzalkonium chloride | 100–10,000 | Polystyrene | 1–7 | [112] |
| L. monocytogenes | Benzalkonium chloride | 10 | Polystyrene | 100% | [170] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Carrascosa, C.; Raheem, D.; Ramos, F.; Saraiva, A.; Raposo, A. Microbial Biofilms in the Food Industry—A Comprehensive Review. Int. J. Environ. Res. Public Health 2021, 18, 2014. https://doi.org/10.3390/ijerph18042014
Carrascosa C, Raheem D, Ramos F, Saraiva A, Raposo A. Microbial Biofilms in the Food Industry—A Comprehensive Review. International Journal of Environmental Research and Public Health. 2021; 18(4):2014. https://doi.org/10.3390/ijerph18042014
Chicago/Turabian StyleCarrascosa, Conrado, Dele Raheem, Fernando Ramos, Ariana Saraiva, and António Raposo. 2021. "Microbial Biofilms in the Food Industry—A Comprehensive Review" International Journal of Environmental Research and Public Health 18, no. 4: 2014. https://doi.org/10.3390/ijerph18042014
APA StyleCarrascosa, C., Raheem, D., Ramos, F., Saraiva, A., & Raposo, A. (2021). Microbial Biofilms in the Food Industry—A Comprehensive Review. International Journal of Environmental Research and Public Health, 18(4), 2014. https://doi.org/10.3390/ijerph18042014










