Environmental Distribution of AR Class 1 Integrons in Upper Adige River Catchment (Northern Italy)
Abstract
:1. Introduction
2. Materials and Methods
2.1. Experimental Set-Up
2.2. Sampling Procedure
2.3. DNA Extraction and Quantification
2.4. PCR Amplifications
3. Results and Discussion
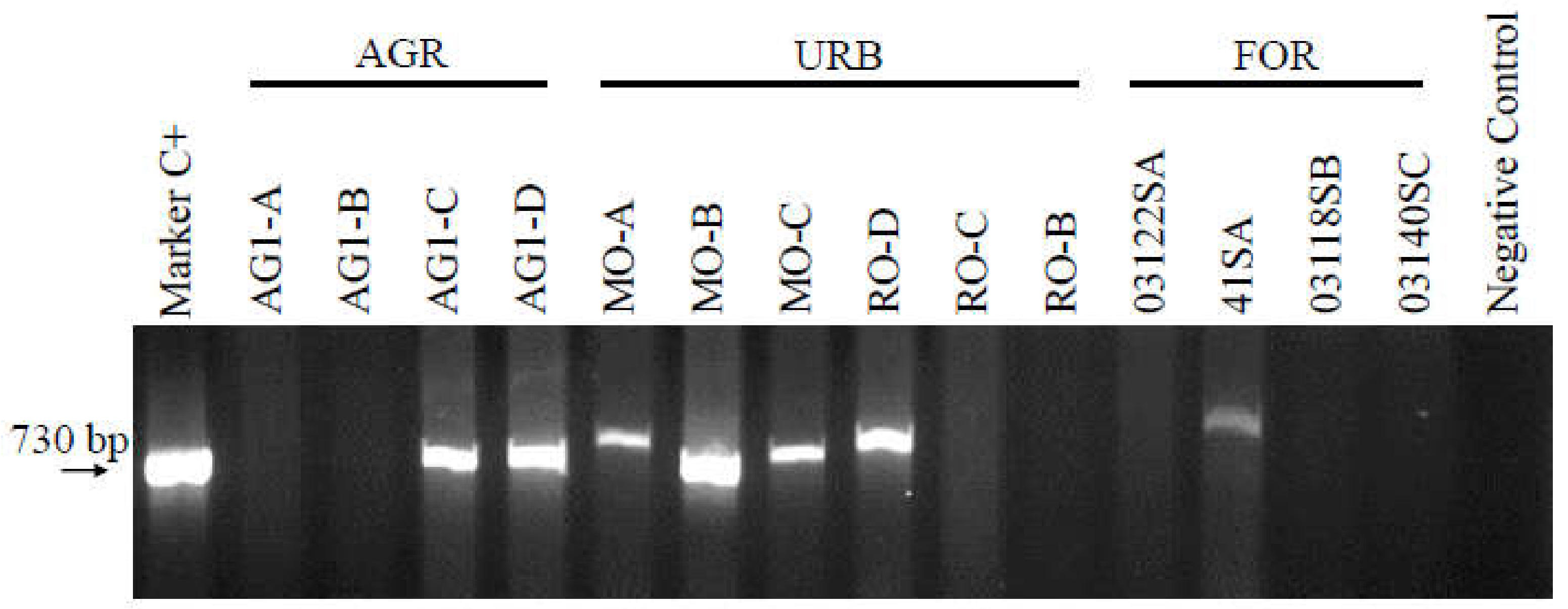
3.1. Intl1 Detection in Agricultural Channels and Apple Orchards
3.2. Intl1 Detection in Urban Channels
3.3. Intl1 Detection in Forest Soil, Leaf-Litter Samples and Post-Glacial Sediments
3.4. Gene Cassettes Detection
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Gillings, M.; Boucher, Y.; Labbate, M.; Holmes, A.; Krishnan, S.; Holley, M.; Stokes, H.W. The evolution of class 1 integrons and the rise of antibiotic resistance. J. Bacteriol. 2008, 190, 5095–5100. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Landers, T.F.; Cohen, B.; Wittum, T.E.; Larson, E.L. A review of antibiotic use in food animals: Perspective, policy, and potential. Public Health Rep. 2012, 127, 4–22. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- The Medicines Utilisation Monitoring Centre. National Report on Antibiotics Use in Italy. Year 2017; Italian Medicines Agency: Rome, Italy, 2019. [Google Scholar]
- EFSA. Scientific report of EFSA and ECDC. Antimicrobial resistance in zoonotic and indicator bacteria from humans, animals and food in the EU. EFSA J. 2015, 13, 1–178. [Google Scholar]
- Gullberg, E.; Albrecht, L.M.; Karlsson, C.; Sandegren, L.; Andersson, D.I. Selection of a multidrug resistance plasmid by sublethal levels of antibiotics and heavy metals. mBio 2014, 5, 19–23. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Martinez, J.L. The role of natural environments in the evolution of resistance traits in pathogenic bacteria. P Roy. Soc. B Biol. Sci. 2009, 276, 2521–2530. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ding, J.; Chen, Z.; Li, Y.; Zhang, Q.; Li, X. Detection of integrons in Escherichia coli producing plasmid-mediated AmpC β-lactamases. Int. J. Clin. Exp. Med. 2019, 12, 1690–1696. [Google Scholar]
- Boucher, Y.; Labbate, M.; Koenig, J.E.; Stokes, H.W. Integrons: Mobilizable platforms that promote genetic diversity in bacteria. Trends Microbiol. 2007, 15, 301–309. [Google Scholar] [CrossRef] [PubMed]
- Gillings, M.R. Integrons: Past, present, and future. Microbiol. Mol. Biol. Rev. 2014, 78, 257–277. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Borruso, L.; Harms, K.; Johnsen, P.J.; Nielsen, K.M.; Brusetti, L. Distribution of class 1 integrons in a highly impacted catchment. Sci. Total Environ. 2016, 566, 1588–1594. [Google Scholar] [CrossRef] [PubMed]
- Allen, H.K.; Donato, J.; Wang, H.H.; Cloud-Hansen, K.A.; Davies, J.; Handelsman, J. Call of the wild: Antibiotic resistance genes in natural environments. Nat. Rev. Microbiol. 2010, 8, 251–259. [Google Scholar] [CrossRef] [PubMed]
- Gillings, M.R. Class 1 integrons as invasive species. Curr. Opin. Microbiol. 2017, 38, 10–15. [Google Scholar] [CrossRef] [PubMed]
- Esposito, A.; Ciccazzo, S.; Borruso, L.; Zerbe, S.; Daffonchio, D.; Brusetti, L. A three-scale analysis of bacterial communities involved in rocks colonization and soil formation in high mountain environments. Curr. Microbiol. 2013, 67, 472–479. [Google Scholar] [CrossRef] [PubMed]
- Gillings, M.R.; Gaze, W.H.; Pruden, A.; Smalla, K.; Tiedje, J.M.; Zhu, Y.G. Using the class 1 integron-integrase gene as a proxy for anthropogenic pollution. ISME J. 2015, 9, 1269–1279. [Google Scholar] [CrossRef] [PubMed]
- Cheng, W.; Chen, H.; Su, C.; Yan, S. Abundance and persistence of antibiotic resistance genes in livestock farms: A comprehensive investigation in Eastern China. Environ. Int. 2013, 61, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Dealtry, S.; Holmsgaard, P.N.; Dunon, V.; Jechalke, S.; Ding, G.C.; Krögerrecklenfort, E.; Smalla, K. Shifts in abundance and diversity of mobile genetic elements after the introduction of diverse pesticides into an on-farm biopurification system over the course of a year. Appl. Env. Microb. 2014, 80, 4012–4020. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cordero, O.X.; Wildschutte, H.; Kirkup, B.; Proehl, S.; Ngo, L.; Hussain, F.; Polz, M.F. Ecological populations of bacteria act as socially cohesive units of antibiotic production and resistance. Science 2012, 337, 1228–1231. [Google Scholar] [CrossRef] [PubMed]
- Esposito, A.; Borruso, L.; Rattray, J.E.; Brusetti, L.; Ahmed, E. Taxonomic and functional insights into rock varnish microbiome using shotgun metagenomics. FEMS Microbiol. Ecol. 2019, 95, fiz180. [Google Scholar] [CrossRef] [PubMed]
- Ogawa, V.A.; Shah, C.M.; Hughes, J.M.; King, L.J. Prioritizing a one health approach in the immediate fight against antimicrobial resistance. EcoHealth 2019, 16, 410–413. [Google Scholar] [CrossRef] [PubMed]
- Chen, Z.; Zhang, W.; Yang, L.; Stedtfeld, R.D.; Peng, A.; Gu, C.; Boyd, S.A.; Li, H. Antibiotic resistance genes and bacterial communities in cornfield and pasture soils receiving swine and dairy manures. Environ. Poll. 2019, 248, 947–957. [Google Scholar] [CrossRef] [PubMed]
| Name | Site | Location | GPS | Land Use Matrix | N. of Samples |
|---|---|---|---|---|---|
| AGR1 | Mori | Rio Cameras Channel | 45°51′8.53″N 10°57′53.56″E | Agricultural sediments | 4 |
| MO | Mori | Rio Cameras Channel | 45°51′17.43″N 10°59′12.57″E | Urban sediments | 4 |
| RO | Rovereto | Leno Stream | 45°52′51.38″N 11° 1′20.54″E | Urban sediments | 4 |
| TRE | Trento | Fersina Stream | 46°2′38.80″N 11° 7′6.39″E | Urban sediments | 3 |
| AGR2 | Mezzolombardo | Noce River | 46°13′8.19″N 11° 6′6.42″E | Agricultural sediments | 3 |
| AGR3 | Ora-Auer | Lusina Channel | 46°21′24.91″N 11°17′55.99″E | Agricultural sediments | 3 |
| BO | Bolzano | Talvera Stream | 46°29′40.73″N 11°20′52.20″E | Urban sediments | 3 |
| CA | Caldaro-Kaltern | Apple orchard | 46°21′18.44″N 11°16′36.81″E | Agricultural soil and rhizosphere | 3 |
| ME1 | Sporminore | Apple orchard | 46°14′13.38″N 11° 2′23.03″E | Agricultural soil | 3 |
| ME2 | Sporminore | Apple orchard | 46°14′12.63″N 11° 2′18.88″E | Agricultural soil | 3 |
| ME3 | Ora-Auer | Apple orchard | 46°21′29.65″N 11°18′3.79″E | Agricultural soil | 3 |
| ME4 | Ora-Auer | Apple orchard | 46°21′32.14″N 11°18′4.97″E | Agricultural soil | 3 |
| MON1 | Monticolo-Montiggl | Oak forest | 46°25′28.78″N 11°17′7.35″E | Oak rhizosphere | 26 |
| MON2 | Monticolo-Montiggl | Oak forest | 46°25′28.78″N 11°17′7.35″E | Fresh leaf litter | 20 |
| VAL | Val di Mazia-Matschertal | Glacier moraine | 46°46′30.00″N 10°41′46.00″E | High mountain rhizosphere | 36 |
| Land Use | Site | Matrix | N. of Samples 1 | Intl1 Region n. of Bands | N. of Samples 2 | Gene Cassette n. of Bands | ||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 1 | 2 | 3 | 0 | 1 | 2 | 3 | |||||
| Vineyard | Mori | Sediments | 4 | 2 | 2(++) | - | - | 2 | - | 2(+) | - | - |
| Vineyard | Mezzolombardo | Sediments | 3 | 2 | 1(++) | - | - | 1 | - | 1(+) | - | - |
| Apple orchard | Ora-Auer | Sediments | 3 | 2 | 1(+) | - | - | 1 | 1 | - | - | - |
| Apple orchard | Caldaro-Kaltern | Rhizosphere and soil | 35 | 35 | - | - | - | None | n.a. | n.a. | n.a. | n.a. |
| Apple orchard | Val di Non | Soil | 6 | 3 | 2(+) | - | 1(+) | None | n.a. | n.a. | n.a. | n.a. |
| Apple orchard | Ora-Auer | Soil | 6 | 3 | 2(+) | 2(+) | - | 1 | - | 1(+) | - | - |
| Urban | Mori | Sediments | 4 | - | 3(++); 1(+++) | - | - | 4 | - | - | 4(++) | n.a. |
| Urban | Rovereto | Sediments | 4 | 3 | 1(+) | - | - | 1 | 1 | - | - | - |
| Urban | Trento | Sediments | 3 | 2 | 1(+) | - | - | 1 | 1 | - | - | - |
| Urban | Bolzano | Sediments | 3 | 3 | - | - | - | None | n.a. | n.a. | n.a. | n.a. |
| Forest | Monticolo-Montiggl | Rhizosphere | 26 | 14 | 10(+) | 2(+) | - | 6 | - | 1(+); 1(++) | 4(++) | - |
| Pristine | Val di Mazia-Matschertal | Mineral soil | 36 | 32 | 4(+) | - | - | 4 | - | 4(+) | - | - |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Piergiacomo, F.; Borruso, L.; Ciccazzo, S.; Rizzi, S.; Zerbe, S.; Brusetti, L. Environmental Distribution of AR Class 1 Integrons in Upper Adige River Catchment (Northern Italy). Int. J. Environ. Res. Public Health 2020, 17, 2336. https://doi.org/10.3390/ijerph17072336
Piergiacomo F, Borruso L, Ciccazzo S, Rizzi S, Zerbe S, Brusetti L. Environmental Distribution of AR Class 1 Integrons in Upper Adige River Catchment (Northern Italy). International Journal of Environmental Research and Public Health. 2020; 17(7):2336. https://doi.org/10.3390/ijerph17072336
Chicago/Turabian StylePiergiacomo, Federica, Luigimaria Borruso, Sonia Ciccazzo, Stefano Rizzi, Stefan Zerbe, and Lorenzo Brusetti. 2020. "Environmental Distribution of AR Class 1 Integrons in Upper Adige River Catchment (Northern Italy)" International Journal of Environmental Research and Public Health 17, no. 7: 2336. https://doi.org/10.3390/ijerph17072336
APA StylePiergiacomo, F., Borruso, L., Ciccazzo, S., Rizzi, S., Zerbe, S., & Brusetti, L. (2020). Environmental Distribution of AR Class 1 Integrons in Upper Adige River Catchment (Northern Italy). International Journal of Environmental Research and Public Health, 17(7), 2336. https://doi.org/10.3390/ijerph17072336







