Soil Bacterial Community Structure in Turfy Swamp and Its Response to Highway Disturbance
Abstract
1. Introduction
2. Materials and Methods
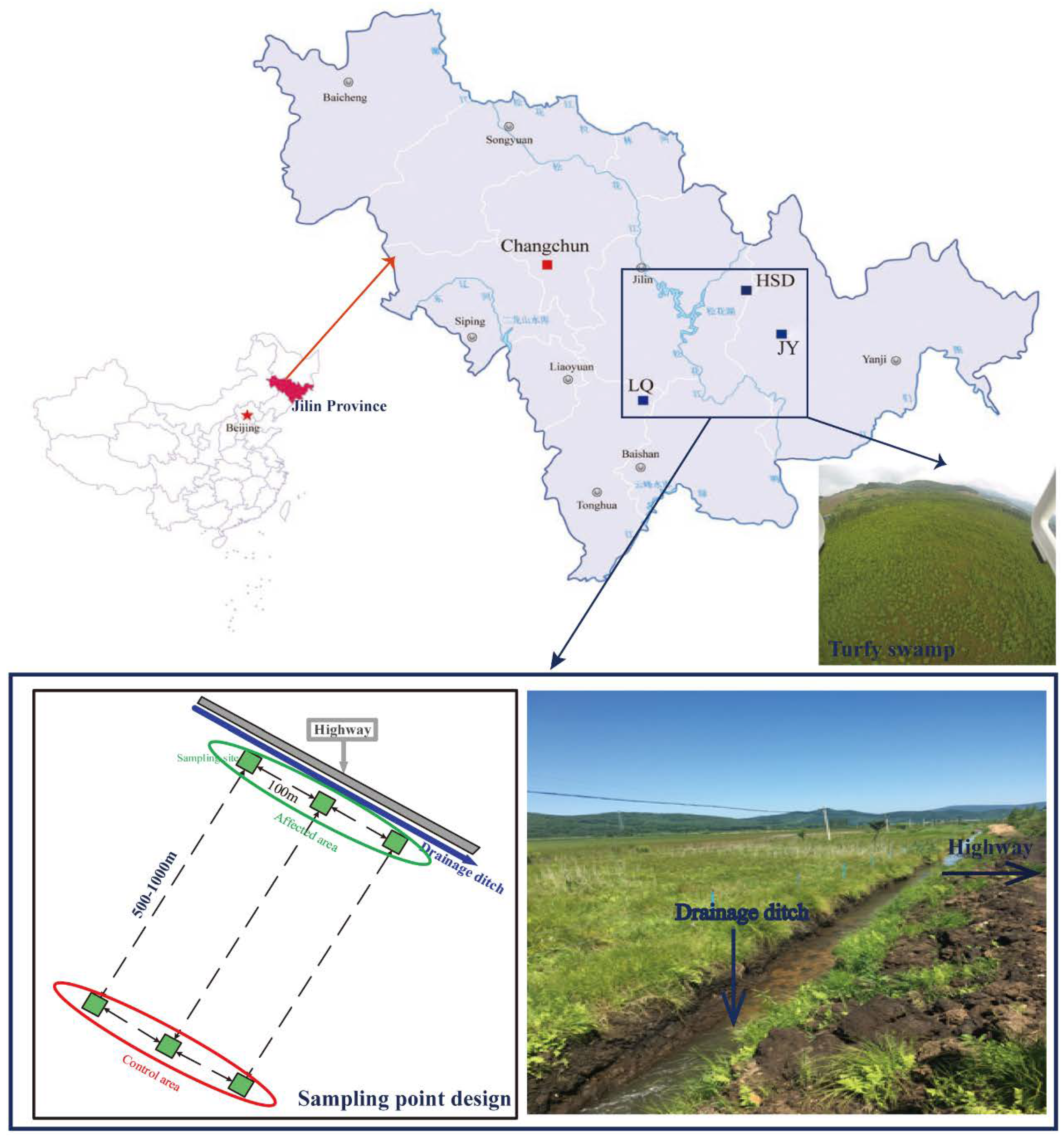
2.1. Site Description and Sample Collection
2.2. Physicochemical Properties Analyses
2.3. DNA Extraction, PCR Amplification and Pyrosequencing
2.4. Statistical and Bioinformatics Analysis
3. Results
3.1. Environmental Factors
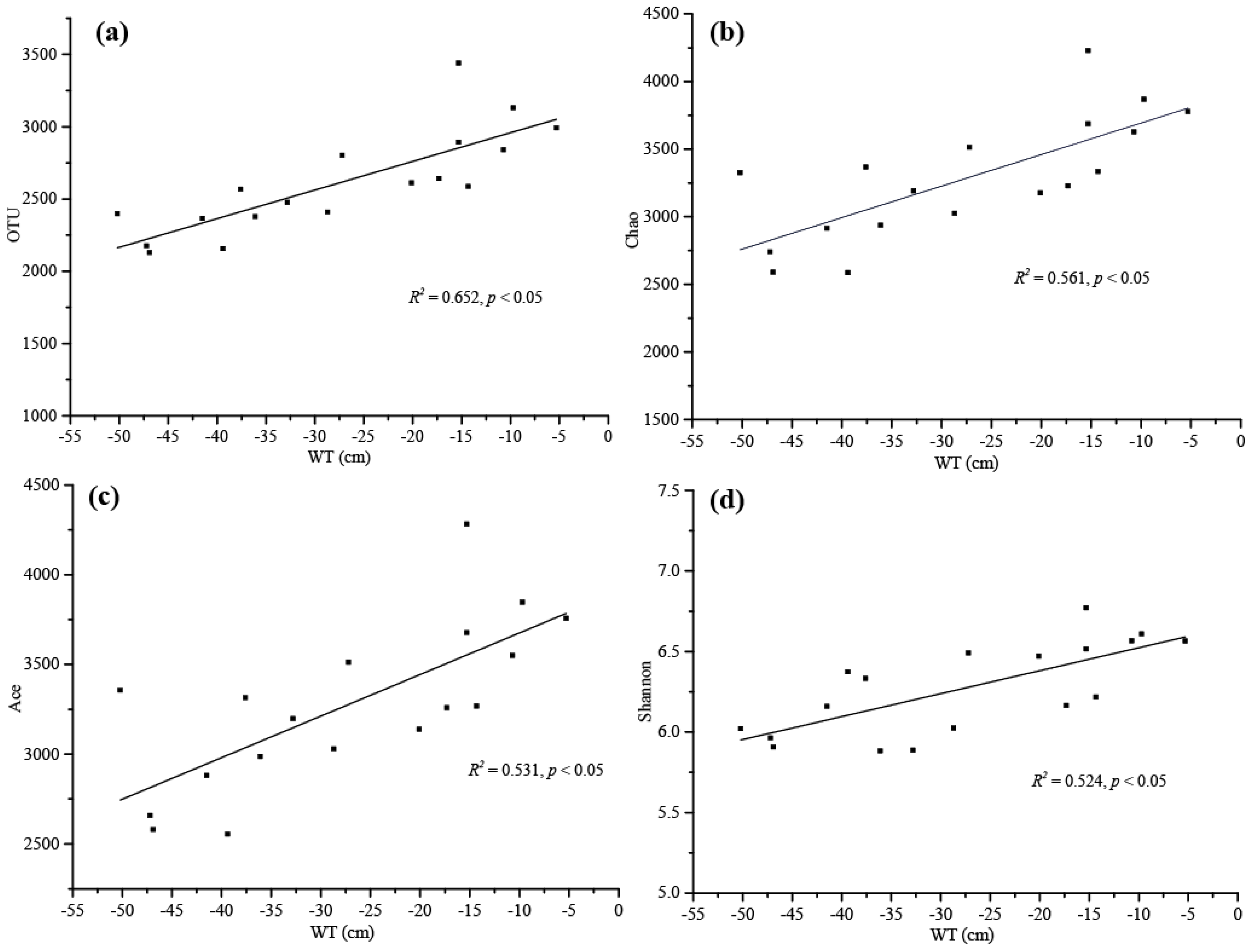
3.2. Bacterial Diversity
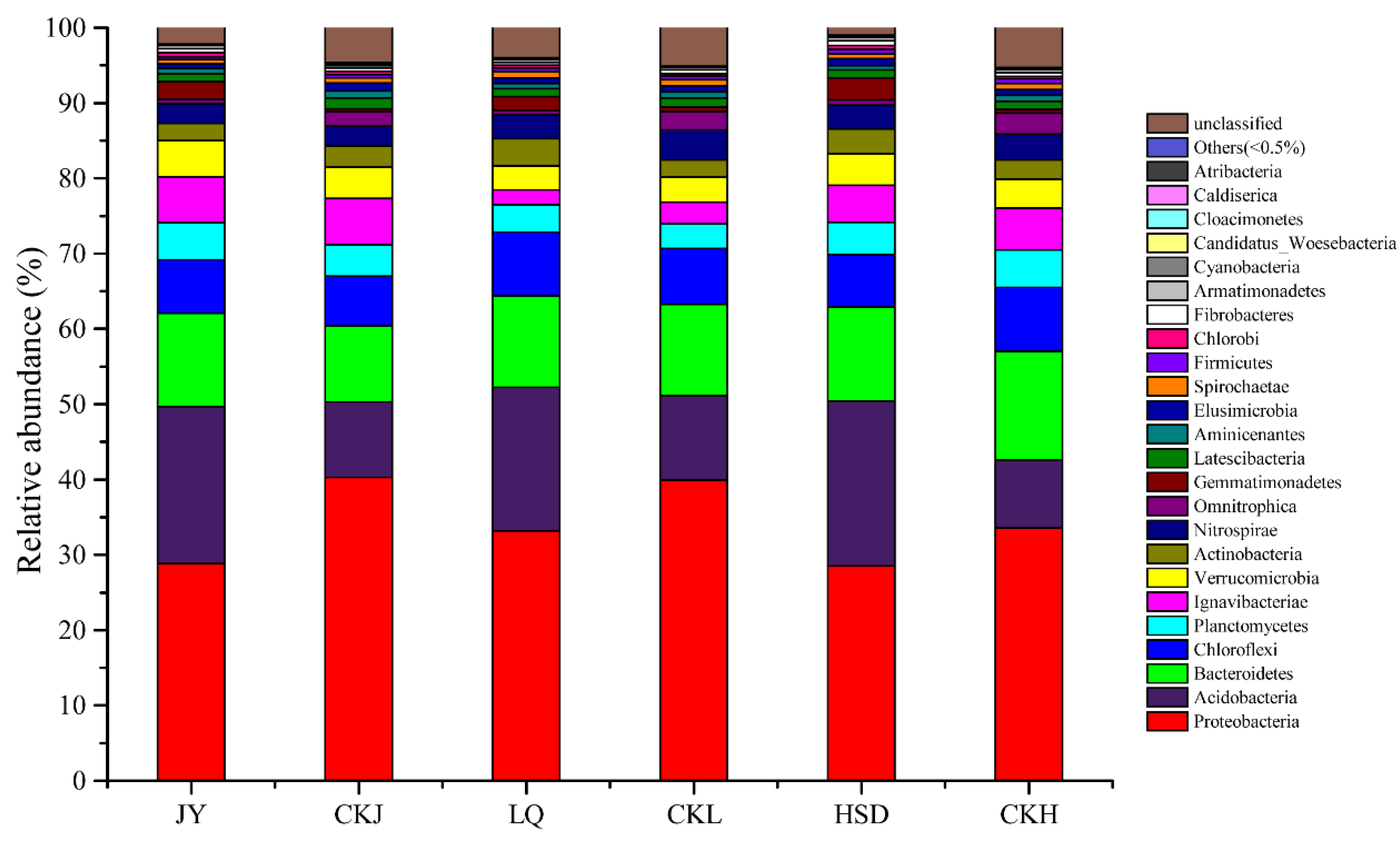
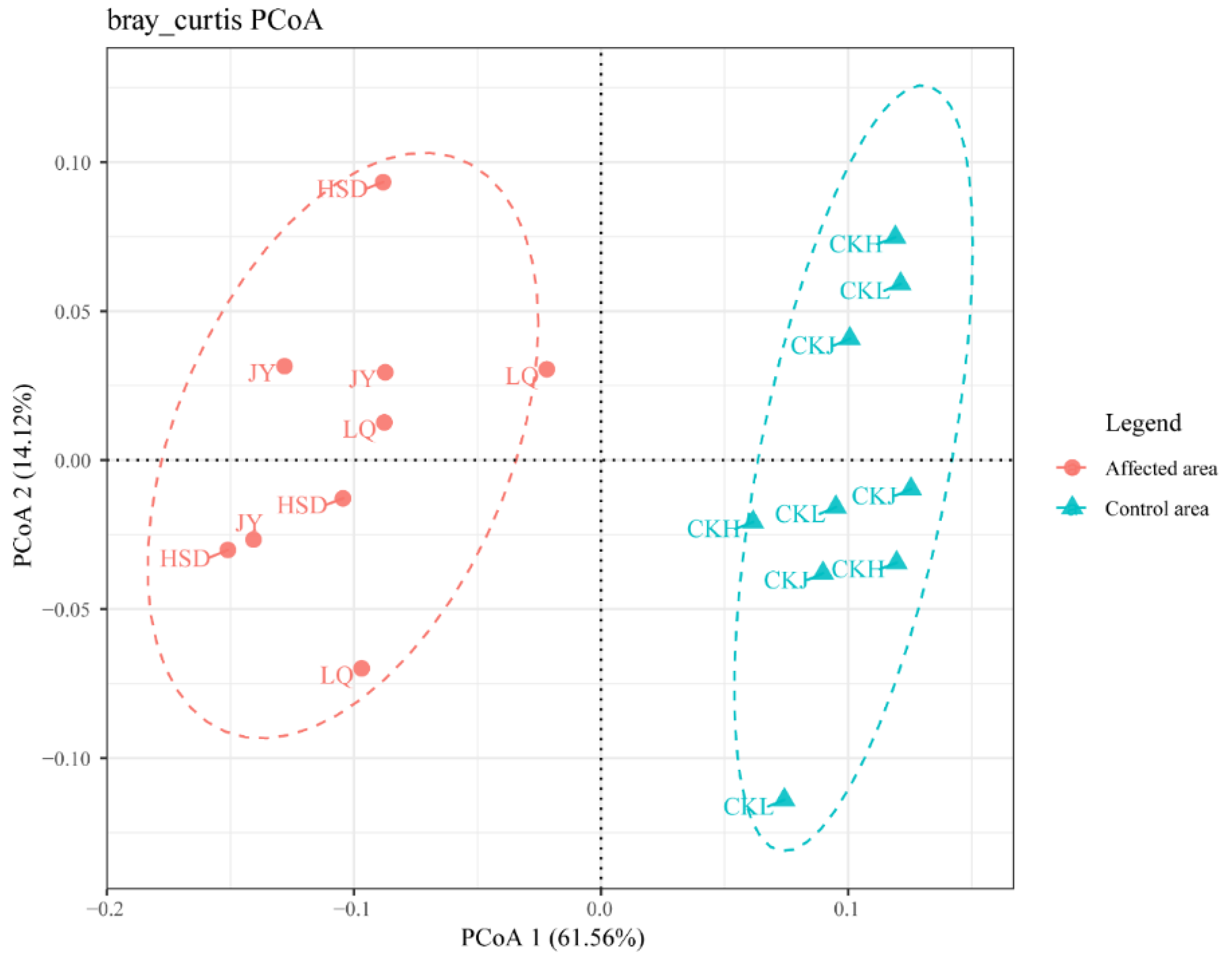
3.3. Bacterial Community Composition
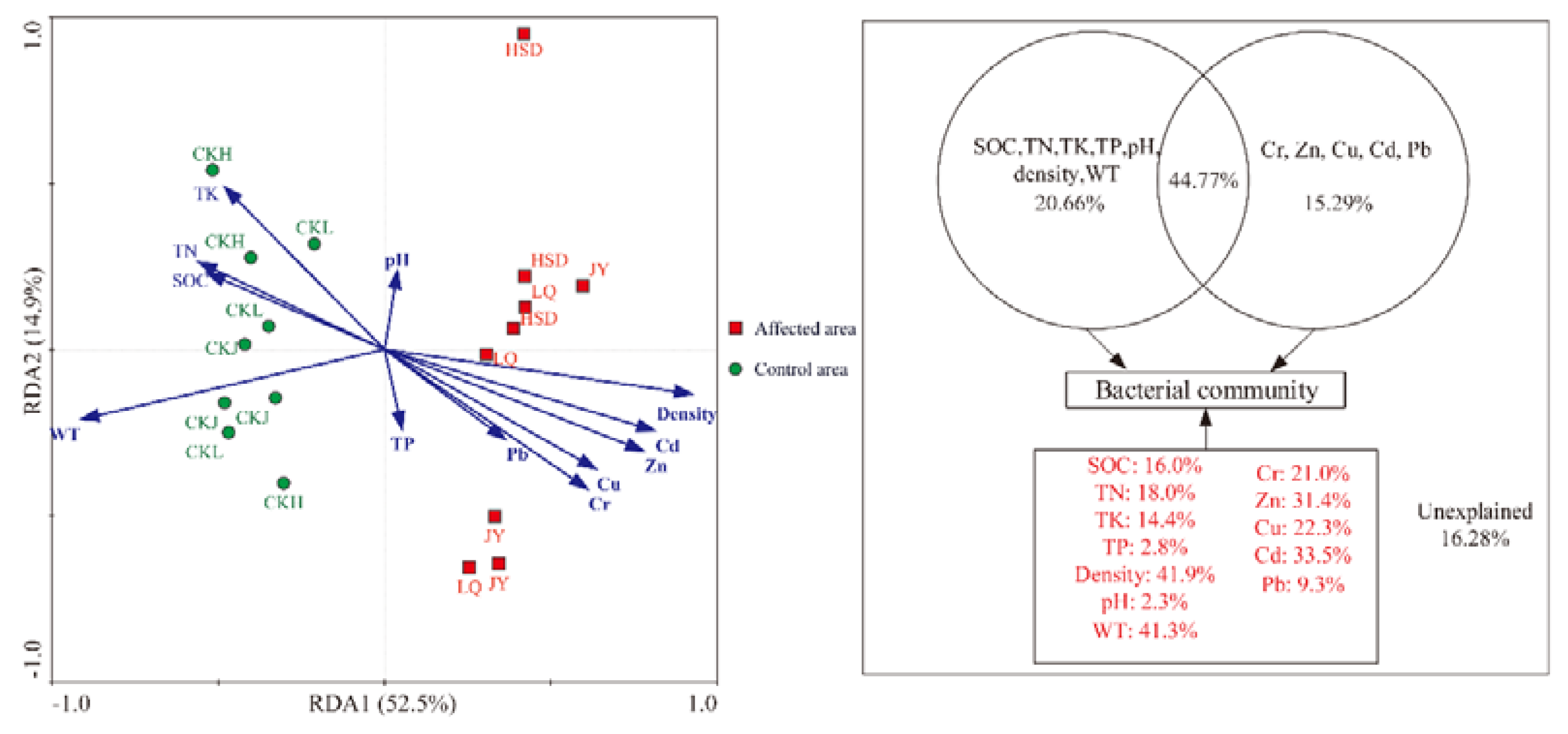
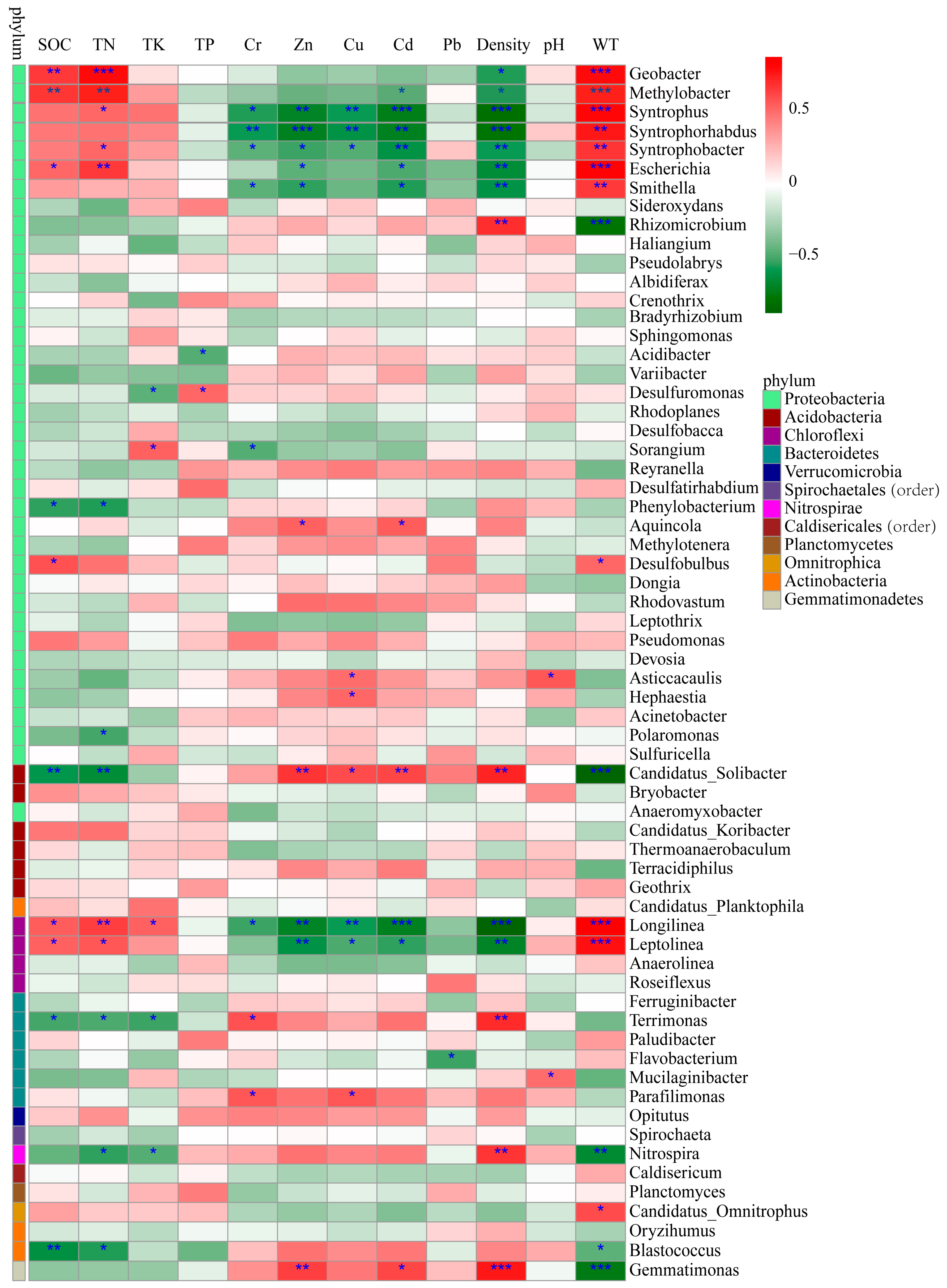
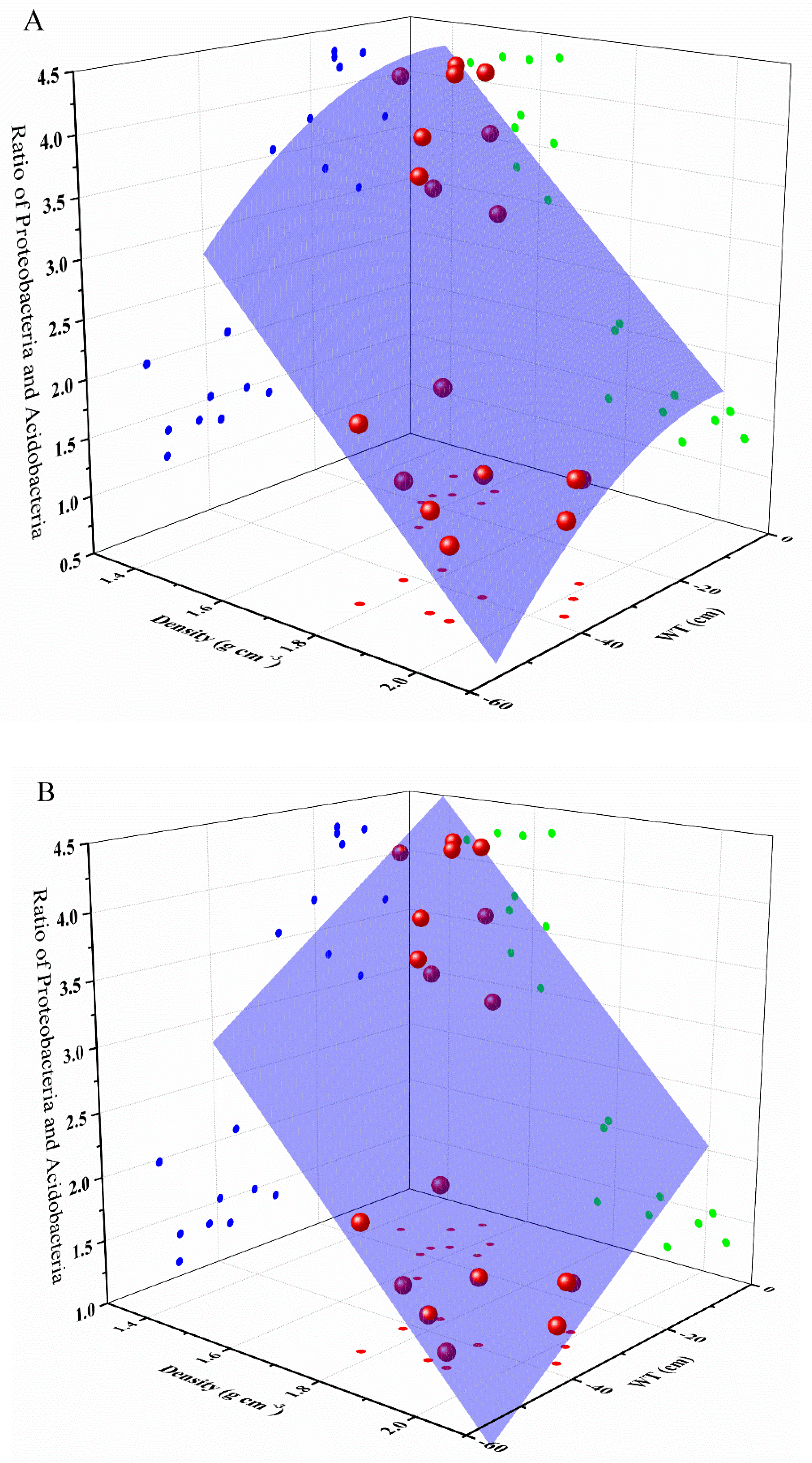
3.4. Relative Influences of Environmental Factors on Bacterial Community
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- McGlone, M.S. The late quaternary peat, vegetation and climate history of the southern oceanic islands of New Zealand. Quat. Sci. Rev. 2002, 21, 683–707. [Google Scholar] [CrossRef]
- Parfenova, L.N.; Selyanina, S.B.; Trufanova, M.V.; Bogolitsyn, K.G.; Orlov, A.S.; Volkova, N.N.; Ponomareva, T.I.; Sokolova, T.V. Influence of climatic and hydrological factors on structure and composition of peat from northern wetland territories with low anthropogenic impact. Sci. Total Environ. 2016, 551, 108–115. [Google Scholar] [CrossRef] [PubMed]
- Heikkinen, K.; Karppinen, A.; Karjalainen, S.M.; Postila, H.; Hadzic, M.; Tolkkinen, M.; Marttila, H.; Ihme, R.; Klove, B. Long-term purification efficiency and factors affecting performance in peatland-based treatment wetlands: An analysis of 28 peat extraction sites in Finland. Ecol. Eng. 2018, 117, 153–164. [Google Scholar] [CrossRef]
- Dai, J.; Wu, H.; Zhang, C.; Zeng, G.; Liang, J.; Guo, S.; Li, X.; Huang, L.; Lu, L.; Yuan, Y. Responses of soil microbial biomass and bacterial community structure to closed-off management (an ecological natural restoration measures): A case study of Dongting Lake wetland, middle China. J. Biosci. Bioeng. 2016, 122, 345–350. [Google Scholar] [CrossRef] [PubMed]
- Bocking, E.; Cooper, D.J.; Price, J. Using tree ring analysis to determine impacts of a road on a boreal peatland. For. Ecol. Manag. 2017, 404, 24–30. [Google Scholar] [CrossRef]
- Wang, H.; Nie, L.; Xu, Y.; Lv, Y. The Effect of Highway on Heavy Metal Accumulation in Soil in Turfy Swamps, Northeastern China. Water Air Soil Pollut. 2017, 228. [Google Scholar] [CrossRef]
- Bohemen, H.D.V.; Laak, W.H.J.V.D. The Influence of Road Infrastructure and Traffic on Soil, Water, and Air Quality. Environ. Manag. 2003, 31, 50–68. [Google Scholar] [CrossRef] [PubMed]
- Jaatinen, K.; Fritze, H.; Laine, J.; Laiho, R. Effects of short- and long-term water-level drawdown on the populations and activity of aerobic decomposers in a boreal peatland. Glob. Chang. Biol. 2007, 13, 491–510. [Google Scholar] [CrossRef]
- Fisk, M.C.; Ruether, K.F.; Yavitt, J.B. Microbial activity and functional composition among northern peatland ecosystems. Soil Biol. Biochem. 2003, 35, 591–602. [Google Scholar] [CrossRef]
- Thormann, M.N.; Currah, R.S.; Bayley, S.E. Patterns of distribution of microfungi in decomposing bog and fen plants. Can. J. Bot. Rev. Can. De Bot. 2004, 82, 710–720. [Google Scholar] [CrossRef]
- Smit, E.; Leeflang, P.; Gommans, S.; Broek, J.V.D.; Wernars, K. Diversity and Seasonal Fluctuations of the Dominant Members of the Bacterial Soil Community in a Wheat Field as Determined by Cultivation and Molecular Methods. Appl. Environ. Microbiol. 2001, 67, 2284–2291. [Google Scholar] [CrossRef] [PubMed]
- Sun, H.; Terhonen, E.; Koskinen, K.; Paulin, L.; Kasanen, R.; Asiegbu, F.O. Bacterial diversity and community structure along different peat soils in boreal forest. Appl. Soil Ecol. 2014, 74, 37–45. [Google Scholar] [CrossRef]
- Urbanová, Z.; Bárta, J.í. Effects of long-term drainage on microbial community composition vary between peatland types. Soil Biol. Biochem. 2016, 92, 16–26. [Google Scholar] [CrossRef]
- De Silva, S.; Ball, A.S.; Huynh, T.; Reichman, S.M. Metal accumulation in roadside soil in Melbourne, Australia: Effect of road age, traffic density and vehicular speed. Environ. Pollut. 2016, 208, 102–109. [Google Scholar] [CrossRef]
- Halonen, J.I.; Blangiardo, M.; Toledano, M.B.; Fecht, D.; Gulliver, J.; Ghosh, R.; Anderson, H.R.; Beevers, S.D.; Dajnak, D.; Kelly, F.J. Is long-term exposure to traffic pollution associated with mortality? A small-area study in London. Environ. Pollut. 2016, 208, 25–32. [Google Scholar] [CrossRef]
- Wang, H.; Zeng, Y.; Guo, C.; Bao, Y.; Lu, G.; Reinfelder, J.R.; Dang, Z. Bacterial, archaeal, and fungal community responses to acid mine drainage-laden pollution in a rice paddy soil ecosystem. Sci. Total Environ. 2018, 616, 107–116. [Google Scholar] [CrossRef] [PubMed]
- Yan, X.; Zhang, F.; Zeng, C.; Zhang, M.; Devkota, L.P.; Yao, T. Relationship between Heavy Metal Concentrations in Soils and Grasses of Roadside Farmland in Nepal. Int. J. Environ. Res. Public Health 2012, 9, 3209–3226. [Google Scholar] [CrossRef]
- Werkenthin, M.; Kluge, B.; Wessolek, G. Metals in European roadside soils and soil solution—A review. Environ. Pollut. 2014, 189, 98–110. [Google Scholar] [CrossRef]
- Shi, G.; Chen, Z.; Xu, S.; Zhang, J.; Wang, L.; Bi, C.; Teng, J. Potentially toxic metal contamination of urban soils and roadside dust in Shanghai, China. Environ. Pollut. 2008, 156, 251–260. [Google Scholar] [CrossRef] [PubMed]
- Zhao, X.; Wang, J.F.; Wang, Y.; Lu, X.; Liu, S.F.; Zhang, Y.B.; Guo, Z.H.; Xie, Z.K.; Wang, R.Y. Influence of proximity to the Qinghai-Tibet highway and railway on variations of soil heavy metal concentrations and bacterial community diversity on the Tibetan Plateau. Sci. Cold Arid Reg. 2019, 11, 407–418. [Google Scholar] [CrossRef]
- Zhang, C.; Nie, S.; Liang, J.; Zeng, G.; Wu, H.; Hua, S.; Liu, J.; Yuan, Y.; Xiao, H.; Deng, L.; et al. Effects of heavy metals and soil physicochemical properties on wetland soil microbial biomass and bacterial community structure. Sci. Total Environ. 2016, 557, 785–790. [Google Scholar] [CrossRef] [PubMed]
- Sun, Z.; Sun, W.; Tong, C.; Zeng, C.; Yu, X.; Mou, X. China’s coastal wetlands: Conservation history, implementation efforts, existing issues and strategies for future improvement. Environ. Int. 2015, 79, 25–41. [Google Scholar] [CrossRef] [PubMed]
- Forman, R.T.T.; Alexander, L.E. Roads and their major ecological effects. Annu. Rev. Ecol. Syst. 1998, 29, 207–231. [Google Scholar] [CrossRef]
- Assaeed, A.M.; Al-Rowaily, S.L.; El-Bana, M.I.; Abood, A.A.A.; Dar, B.A.M.; Hegazy, A.K. Impact of off-road vehicles on soil and vegetation in a desert rangeland in Saudi Arabia. Saudi J. Biol. Sci. 2019, 26, 1187–1193. [Google Scholar] [CrossRef]
- Saraswati, S.; Parsons, C.T.; Strack, M. Access roads impact enzyme activities in boreal forested peatlands. Sci. Total Environ. 2019, 651, 1405–1415. [Google Scholar] [CrossRef]
- Yang, J.Y.; Yang, X.E.; He, Z.L.; Li, T.Q.; Shentu, J.L.; Stoffella, P.J. Effects of pH, organic acids, and inorganic ions on lead desorption from soils. Environ. Pollut. 2006, 143, 9–15. [Google Scholar] [CrossRef]
- Ye, Q.; Wu, Y.; Zhu, Z.; Wang, X.; Li, Z.; Zhang, J. Bacterial diversity in the surface sediments of the hypoxic zone near the Changjiang Estuary and in the East China Sea. Microbiologyopen 2016, 5, 323–339. [Google Scholar] [CrossRef]
- Brown, R.; Zhang, Z.; Comeau, L.-P.; Bedard-Haughn, A. Effects of drainage duration on mineral wetland soils in a Prairie Pothole agroecosystem. Soil Tillage Res. 2017, 168, 187–197. [Google Scholar] [CrossRef]
- Yousefi, S.; Moradi, H.; Boll, J.; Schoenbrodt-Stitt, S. Effects of road construction on soil degradation and nutrient transport in Caspian Hyrcanian mixed forests. Geoderma 2016, 284, 103–112. [Google Scholar] [CrossRef]
- Shi, P.; Zhang, Y.; Hu, Z.; Ma, K.; Wang, H.; Chai, T. The response of soil bacterial communities to mining subsidence in the west China aeolian sand area. Appl. Soil Ecol. 2017, 121, 1–10. [Google Scholar] [CrossRef]
- Guo, H.; Nasir, M.; Lv, J.; Dai, Y.; Gao, J. Understanding the variation of microbial community in heavy metals contaminated soil using high throughput sequencing. Ecotoxicol. Environ. Saf. 2017, 144, 300–306. [Google Scholar] [CrossRef] [PubMed]
- Zhang, S.; Chang, J.; Liu, W.; Pan, Y.; Cui, K.; Chen, X.; Liang, P.; Zhang, X.; Wu, Q.; Qiu, Y.; et al. A novel bioaugmentation strategy to accelerate methanogenesis via adding Geobacter sulfurreducens PCA in anaerobic digestion system. Sci. Total Environ. 2018, 642, 322–326. [Google Scholar] [CrossRef]
- Mei, R.; Nobu, M.K.; Narihiro, T.; Yu, J.; Sathyagal, A.; Willman, E.; Liu, W.-T. Novel Geobacter species and diverse methanogens contribute to enhanced methane production in media-added methanogenic reactors. Water Res. 2018, 147, 403–412. [Google Scholar] [CrossRef] [PubMed]
- Wei, M.; Qiu, Q.; Qian, Y.; Cheng, L.; Guo, A. Methane oxidation and response of Methylobacter/Methylosarcina methanotrophs in flooded rice soil amended with urea. Appl. Soil Ecol. 2016, 101, 174–184. [Google Scholar] [CrossRef]
- Martinez-Cruz, K.; Leewis, M.-C.; Herriott, I.C.; Sepulveda-Jauregui, A.; Anthony, K.W.; Thalasso, F.; Leigh, M.B. Anaerobic oxidation of methane by aerobic methanotrophs in sub-Arctic lake sediments. Sci. Total Environ. 2017, 607, 23–31. [Google Scholar] [CrossRef] [PubMed]
- Franchi, O.; Bovio, P.; Ortega-Martinez, E.; Rosenkranz, F.; Chamy, R. Active and total microbial community dynamics and the role of functional genes bamA and mcrA during anaerobic digestion of phenol and p-cresol. Bioresour. Technol. 2018, 264, 290–297. [Google Scholar] [CrossRef]
- Narihiro, T.; Nobu, M.K.; Kim, N.-K.; Kamagata, Y.; Liu, W.-T. The nexus of syntrophy-associated microbiota in anaerobic digestion revealed by long-term enrichment and community survey. Environ. Microbiol. 2015, 17, 1707–1720. [Google Scholar] [CrossRef]
- Pham, H.N.; Pham, P.A.; Nguyen, T.T.H.; Meiffren, G.; Brothier, E.; Lamy, I.; Michalet, S.; Dijoux-Franca, M.-G.; Nazaret, S. Influence of metal contamination in soil on metabolic profiles of Miscanthus x giganteus belowground parts and associated bacterial communities. Appl. Soil Ecol. 2018, 125, 240–249. [Google Scholar] [CrossRef]
- Zeng, Q.; Dong, Y.; An, S. Bacterial Community Responses to Soils along a Latitudinal and Vegetation Gradient on the Loess Plateau, China. PLoS ONE 2016, 11, e0152894. [Google Scholar] [CrossRef]
- Kim, S.-Y.; Lee, S.-H.; Freeman, C.; Fenner, N.; Kang, H. Comparative analysis of soil microbial communities and their responses to the short-term drought in bog, fen, and riparian wetlands. Soil Biol. Biochem. 2008, 40, 2874–2880. [Google Scholar] [CrossRef]
- Peltoniemi, K.; Strakova, P.; Fritze, H.; Alvira Iraizoz, P.; Pennanen, T.; Laiho, R. How water-level drawdown modifies litter-decomposing fungal and actinobacterial communities in boreal peatlands. Soil Biol. Biochem. 2012, 51, 20–34. [Google Scholar] [CrossRef]
- Dang, P.; Gao, Y.; Liu, J.; Yu, S.; Zhao, Z. Effects of thinning intensity on understory vegetation and soil microbial communities of a mature Chinese pine plantation in the Loess Plateau. Sci. Total Environ. 2018, 630, 171–180. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Dang, P.; Gao, Y.; Zhu, H.; Zhu, H.; Zhao, F.; Zhao, Z. Effects of tree species and soil properties on the composition and diversity of the soil bacterial community following afforestation. For. Ecol. Manag. 2018, 427, 342–349. [Google Scholar] [CrossRef]
- Cui, H.; Liu, L.-L.; Dai, J.-R.; Yu, X.-N.; Guo, X.; Yi, S.J.; Zhou, D.Y.; Guo, W.H.; Du, N. Bacterial community shaped by heavy metals and contributing to health risks in cornfields. Ecotoxicol. Environ. Saf. 2018, 166, 259–269. [Google Scholar] [CrossRef]
- Weaver, M.A.; Zablotowicz, R.M.; Krutz, L.J.; Bryson, C.T.; Locke, M.A. Microbial and vegetative changes associated with development of a constructed wetland. Ecol. Indic. 2012, 13, 37–45. [Google Scholar] [CrossRef]
- Xie, B.; Cui, Y.; Yuan, Q.; Li, J. Pollutants Removal and Distribution of Microorganisms in a Reed Wetland of Shanghai Mengqing Park. Environ. Prog. Sustain. Energy 2009, 28, 240–248. [Google Scholar] [CrossRef]
- Li, G.; Chen, Y.; Zhang, M.; Gong, X.; Iop. Study on water environmental protection technology subsidence area ecological wetland. In Proceedings of the 2018 4th International Conference on Environmental Science and Material Application, Xi’an, China, 15–16 December 2018; Volume 252. [Google Scholar]

| Parameters | JY | LQ | HSD | |||
|---|---|---|---|---|---|---|
| Affected Area | Control Area | Affected Area | Control Area | Affected Area | Control Area | |
| SOC (g kg−1) | 130 ± 15.39 a | 147 ± 24.26 a | 72.93 ± 8.94 b | 135.67 ± 27.43 a | 101.8 ± 52.99ab | 132.33 ± 26.31 a |
| TN (g kg−1) | 14.73 ± 2.70 b | 20.17 ± 1.44 a | 8.03 ± 0.56 c | 13.33 ± 0.91 b | 12.19 ± 5.06 bc | 16.70 ± 2.77 ab |
| TK (g kg−1) | 5.61 ± 0.97 b | 6.44 ± 0.72 ab | 7.31 ± 0.69ab | 8.77 ± 0.12 a | 6.23 ± 1.15ab | 8.81 ± 3.09 a |
| TP (g kg−1) | 1.32 ± 0.09 a | 1.28 ± 0.14 a | 1.37 ± 0.46 a | 1.15 ± 0.07 a | 1.22 ± 0.19 a | 1.32 ± 0.04 a |
| Cr (mg kg−1) | 121.44 ± 9.36 a | 64.48 ± 10.78 b | 69.05 ± 2.32 b | 46.77 ± 1.09 c | 63.40 ± 5.94 b | 48.01 ± 5.05 c |
| Zn (mg kg−1) | 170.24 ± 27.89 a | 84.14 ± 3.44 bc | 158.43 ± 17.89 a | 69.01 ± 6.28 bc | 93.74 ± 4.95 b | 64.63 ± 4.01 c |
| Cu (mg kg−1) | 51.86 ± 9.91 a | 19.57 ± 1.09 b | 50.34 ± 9.91 a | 18.06 ± 2.76 b | 17.02 ± 1.88 b | 15.70 ± 0.37 b |
| Cd (mg kg−1) | 0.837 ± 0.11 a | 0.218 ± 0.06 cd | 0.667 ± 0.11 b | 0.152 ± 0.05 d | 0.349 ± 0.08 c | 0.138 ± 0.05 d |
| Pb (mg kg−1) | 22.45 ± 1.87 a | 19.81 ± 4.43 a | 21.55 ± 2.66 a | 20.25 ± 2.37 a | 19.43 ± 2.71 a | 17.56 ± 3.37 a |
| WT (cm) | −33.0 ± 4.5 b | −10.1 ± 5.0 a | −42.6 ± 13.1 bc | −17.7 ± 8.5 a | −44.5 ± 14.4 c | −17.2 ± 2.9 a |
| Density (g cm−3) | 2.01 ± 0.03 a | 1.54 ± 0.13 e | 1.77 ± 0.01 c | 1.63 ± 0.02 d | 1.89 ± 0.22 b | 1.52 ± 0.06 e |
| pH | 5.81 ± 0.34 a | 5.83 ± 0.07 a | 5.88 ± 0.39 a | 5.54 ± 0.52 a | 4.82 ± 0.78 a | 5.58 ± 0.48 a |
| Sample Code | Reads | 0.97 | ||||||
|---|---|---|---|---|---|---|---|---|
| Raw Sequences | Trimmed Sequences | Coverage | OTU | Ace | Chao | Shannon | Simpson | |
| JY-A | 50,260 | 40,118 | 0.98 | 2483 | 3180 ± 143 c | 3194 ± 171 c | 6.28 ± 0.23 a | 0.0072 ± 0.0113 a |
| JY-C | 49,895 | 38,528 | 0.97 | 3187 | 3961 ± 280 a | 3958 ± 238 a | 6.64 ± 0.31 a | 0.0080 ± 0.0011 a |
| LQ-A | 46,772 | 36,862 | 0.98 | 2380 | 3075 ± 249 c | 3059 ± 230 c | 6.12 ± 0.24 a | 0.0093 ± 0.0017 a |
| LQ-C | 48,862 | 37,321 | 0.98 | 2844 | 3579 ± 86 b | 3608 ± 87 b | 6.52 ± 0.04 a | 0.0079 ± 0.0013 a |
| HSD-A | 52,611 | 41,076 | 0.98 | 2153 | 2597 ± 54 d | 2637 ± 87 d | 6.28 ± 0.16 a | 0.0083 ± 0.0029 a |
| HSD-C | 50,237 | 38,937 | 0.98 | 2607 | 3221 ± 72 c | 3246 ± 81 c | 6.08 ± 0.35 a | 0.0069 ± 0.0014 a |
| JY Site | LQ Site | HSD Site | ||||
|---|---|---|---|---|---|---|
| Affected Area (JY) | Control Area (CKJ) | Affected Area (LQ) | Control Area (CKL) | Affected Area (HSD) | Control Area (CKH) | |
| Proteobacteria | 28.81 ± 1.08 c | 40.28 ± 1.22 a | 33.17 ± 3.45 b | 39.94 ± 3.12 a | 28.53 ± 1.65 c | 33.57 ± 2.90 b |
| Acidobacteria | 20.85 ± 2.27 a | 10.02 ± 1.11 b | 19.05 ± 3.83a | 11.15 ± 1.37 b | 21.90 ± 4.30 a | 9.02 ± 1.99 b |
| Bacteroidetes | 12.41 ± 1.97 a | 10.08 ± 1.76 a | 12.14 ± 3.59 a | 12.17 ± 4.91 a | 12.48 ± 4.20 a | 14.44 ± 2.81 a |
| Chloroflexi | 7.08 ± 2.94 a | 6.65 ± 1.24 a | 8.46 ± 0.72 a | 7.43 ± 1.27 a | 6.95 ± 2.17 a | 8.49 ± 2.05 a |
| Planctomycetes | 4.97 ± 0.20 a | 4.12 ± 0.91 a | 3.66 ± 1.68 a | 3.29 ± 0.50 a | 4.30 ± 1.05 a | 4.96 ± 1.24 a |
| Ignavibacteriae | 6.05 ± 0.50 a | 6.18 ± 1.44 a | 1.96 ± 0.67 b | 2.83 ± 1.03 b | 4.91 ± 1.07 a | 5.52 ± 1.33 a |
| Verrucomicrobia | 4.80 ± 1.20 a | 4.14 ± 1.07 ab | 3.16 ± 0.55 b | 3.30 ± 0.29 ab | 4.18 ± 0.27 ab | 3.89 ± 1.13 ab |
| Actinobacteria | 2.31 ± 1.94 a | 2.85 ± 1.45 a | 3.64 ± 1.63 a | 2.30 ± 0.48 a | 3.27 ± 0.54 a | 2.51 ± 0.99 a |
| Nitrospirae | 2.62 ± 1.05 a | 2.77 ± 2.23 a | 3.22 ± 1.12 a | 4.03 ± 2.38 a | 3.21 ± 0.41 a | 3.44 ± 0.95 a |
| Omnitrophica | 0.57 ± 0.21 c | 1.97 ± 0.13 b | 0.54 ± 0.22 c | 2.43 ± 0.29 ab | 0.66 ± 0.17 c | 2.82 ± 0.78 a |
| Gemmatimonadetes | 2.38 ± 0.61 ab | 0.34 ± 0.10 c | 1.83 ± 0.36 b | 0.57 ± 0.19 c | 2.91 ± 0.96 a | 0.51 ± 0.16 c |
| Latescibacteria | 1.04 ± 0.37 a | 1.46 ± 0.91 a | 1.05 ± 0.26 a | 1.22 ± 0.11 a | 1.06 ± 0.34 a | 1.07 ± 0.21 a |
| Aminicenantes | 0.71 ± 0.20 a | 0.96 ± 0.29 a | 0.66 ± 0.32 a | 0.78 ± 0.48 a | 0.59 ± 0.20 a | 0.79 ± 0.27 a |
| Elusimicrobia | 0.61 ± 0.33 a | 1.02 ± 0.43 a | 0.79 ± 0.29 a | 0.82 ± 0.18 a | 0.90 ± 0.22 a | 0.72 ± 0.06 a |
| Spirochaetae | 0.58 ± 0.14 a | 0.69 ± 0.04 a | 0.79 ± 0.13 a | 0.83 ± 0.21 a | 0.66 ± 0.27 a | 0.83 ± 0.33 a |
| Firmicutes | 0.38 ± 0.12 a | 0.45 ± 0.21 a | 0.49 ± 0.27 a | 0.48 ± 0.26 a | 0.62 ± 0.16 a | 0.64 ± 0.13 a |
| Chlorobi | 0.53 ± 0.08 a | 0.43 ± 0.27 a | 0.39 ± 0.14 a | 0.30 ± 0.07 a | 0.49 ± 0.32 a | 0.32 ± 0.16 a |
| Fibrobacteres | 0.47 ± 0.31 a | 0.44 ± 0.07 a | 0.38 ± 0.12 a | 0.48 ± 0.02 a | 0.59 ± 0.07 a | 0.49 ± 0.13 a |
| Armatimonadetes | 0.47 ± 0.12 ab | 0.31 ± 0.05 b | 0.36 ± 0.13 ab | 0.31 ± 0.05 b | 0.54 ± 0.14 a | 0.41 ± 0.13 ab |
| Cyanobacteria | 0.13 ± 0.01 a | 0.11 ± 0.04 a | 0.14 ± 0.09 a | 0.13 ± 0.05 a | 0.13 ± 0.05 a | 0.11 ± 0.04 a |
| Candidatus_Woesebacteria | 0.02 ± 0.02 a | 0.03 ± 0.03 a | 0.04 ± 0.04 a | 0.04 ± 0.02 a | 0.01 ± 0.00 a | 0.04 ± 0.01 a |
| Cloacimonetes | 0.03 ± 0.01 a | 0.02 ± 0.01 a | 0.04 ± 0.03 a | 0.04 ± 0.02 a | 0.03 ± 0.00 a | 0.06 ± 0.02 a |
| Caldiserica | 0.02 ± 0.01 a | 0.07 ± 0.01 a | 0.01 ± 0.00 a | 0.01 ± 0.00 a | 0.02 ± 0.01 a | 0.02 ± 0.02 a |
| Atribacteria | 0.00 ± 0.00 a | 0.21 ± 0.05 a | 0.00 ± 0.00 a | 0.01 ± 0.00 a | 0.01 ± 0.00 a | 0.00 ± 0.00 a |
| Others (<0.5%) | 0.01 ± 0.00 | 0.01 ± 0.01 | 0.01 ± 0.00 | 0.01 ± 0.00 | 0.01 ± 0.00 | 0.01 ± 0.00 |
| unclassified | 2.16 ± 1.51 | 4.58 ± 1.65 | 3.99 ± 1.87 | 5.08 ± 2.48 | 1.01 ± 0.63 | 5.32 ± 2.39 |
| JY Site | LQ Site | HSD Site | ||||
|---|---|---|---|---|---|---|
| Affected Area (JY) | Control Area (CKJ) | Affected Area (LQ) | Control Area (CKL) | Affected Area (HSD) | Control Area (CKH) | |
| Geobacter | 2.49 ± 1.59 bc | 7.23 ± 2.36 a | 1.16 ± 0.13 c | 3.97 ± 0.20 b | 1.22 ± 0.45 c | 3.10 ± 0.55 bc |
| Syntrophus | 0.06 ± 0.05 c | 1.67 ± 0.74 b | 0.09 ± 0.06 c | 1.84 ± 0.05 b | 0.06 ± 0.01 c | 2.84 ± 0.70 a |
| Escherichia | 0.44 ± 0.21 c | 3.84 ± 1.92 a | 0.39 ± 0.15 c | 2.07 ± 0.81 b | 0.25 ± 0.11 c | 1.48 ± 0.13 b |
| Sideroxydans | 0.15 ± 0.02 b | 0.15 ± 0.01 b | 0.45 ± 0.31 a | 0.42 ± 0.11 a | 0.14 ± 0.01 b | 0.15 ± 0.02 b |
| Rhizomicrobium | 0.59 ± 0.33 b | 0.09 ± 0.02 c | 0.59 ± 0.35 b | 0.15 ± 0.07 bc | 1.58 ± 0.35 a | 0.15 ± 0.08 bc |
| Haliangium | 0.80 ± 0.34 a | 0.80 ± 0.18 a | 0.48 ± 0.04 a | 0.41 ± 0.23 a | 0.99 ± 0.86 a | 0.83 ± 0.42 a |
| Pseudol abrys | 0.39 ± 0.28 a | 0.29 ± 0.12 a | 0.42 ± 0.09 a | 0.42 ± 0.33 a | 1.11 ± 0.97 a | 0.80 ± 0.62 a |
| Albidiferax | 0.63 ± 0.48 a | 0.61 ± 0.53 a | 0.88 ± 0.22 a | 0.91 ± 0.25 a | 0.53 ± 0.26 a | 0.55 ± 0.27 a |
| Crenothrix | 0.65 ± 0.37 a | 0.76 ± 0.47 a | 0.47 ± 0.31 a | 0.43 ± 0.29 a | 0.45 ± 0.30 a | 0.45 ± 0.12 a |
| Syntrophorh abdus | 0.02 ± 0.01 b | 0.73 ± 0.33 a | 0.02 ± 0.01 b | 0.81 ± 0.35 a | 0.09 ± 0.03 b | 1.03 ± 0.41 a |
| Smithella | 0.00 ± 0.00 b | 0.38 ± 0.14 ab | 0.04 ± 0.03 b | 0.79 ± 0.12 a | 0.00 ± 0.00 b | 1.08 ± 0.64 a |
| Syntrophobacter | 0.02 ± 0.01 b | 0.71 ± 0.36 a | 0.05 ± 0.03 b | 0.52 ± 0.27 a | 0.04 ± 0.03 b | 0.33 ± 0.13 ab |
| Bradyrhizobium | 0.19 ± 0.03 b | 0.22 ± 0.05 b | 0.32 ± 0.27 ab | 0.25 ± 0.18 b | 0.55 ± 0.16 a | 0.48 ± 0.16 ab |
| Sphingomonas | 0.10 ± 0.07 a | 0.11 ± 0.05 a | 0.33 ± 0.23 a | 0.47 ± 0.25 a | 0.14 ± 0.07 a | 0.14 ± 0.01 a |
| Acidibacter | 0.31 ± 0.16 a | 0.27 ± 0.19 a | 0.43 ± 0.42 a | 0.36 ± 0.27 a | 0.29 ± 0.13 a | 0.25 ± 0.12 a |
| Variibacter | 0.40 ± 0.18 a | 0.28 ± 0.18 a | 0.40 ± 0.35 a | 0.22 ± 0.13 a | 0.55 ± 0.38 a | 0.36 ± 0.30 a |
| Methylobacter | 0.19 ± 0.15 b | 0.58 ± 0.13 a | 0.10 ± 0.02 b | 0.38 ± 0.13 ab | 0.14 ± 0.01 b | 0.30 ± 0.14 b |
| Desulfuromonas | 0.03 ± 0.01 a | 0.02 ± 0.02 a | 0.02 ± 0.02 a | 0.01 ± 0.01 a | 0.01 ± 0.01 a | 0.03 ± 0.02 a |
| Rhodoplanes | 0.20 ± 0.09 a | 0.16 ± 0.07 a | 0.11 ± 0.06 a | 0.14 ± 0.12 a | 0.44 ± 0.38 a | 0.29 ± 0.26 a |
| Desulfobacca | 0.01 ± 0.00 a | 0.05 ± 0.04 ab | 0.03 ± 0.02 a | 0.13 ± 0.07 a | 0.33 ± 0.05 a | 0.27 ± 0.07 a |
| Sorangium | 0.04 ± 0.04 a | 0.05 ± 0.05 a | 0.11 ± 0.04 a | 0.11 ± 0.01 a | 0.14 ± 0.06 a | 0.16 ± 0.10 a |
| Reyranella | 0.10 ± 0.03 a | 0.06 ± 0.04 a | 0.11 ± 0.03 a | 0.06 ± 0.04 a | 0.10 ± 0.04 a | 0.06 ± 0.03 a |
| Desulfatirh abdium | 0.03 ± 0.02 a | 0.05 ± 0.01 a | 0.14 ± 0.04 a | 0.16 ± 0.15 a | 0.01 ± 0.01 a | 0.05 ± 0.04 a |
| Phenylobacterium | 0.06 ± 0.02 a | 0.03 ± 0.00 a | 0.06 ± 0.02 a | 0.06 ± 0.01 a | 0.09 ± 0.08 a | 0.05 ± 0.01 a |
| Aquincola | 0.29 ± 0.09 a | 0.12 ± 0.06 a | 0.14 ± 0.10 a | 0.07 ± 0.06 a | 0.14 ± 0.07 a | 0.08 ± 0.03 a |
| Methylotenera | 0.05 ± 0.04 ab | 0.04 ± 0.04 b | 0.12 ± 0.07 a | 0.02 ± 0.02 b | 0.02 ± 0.01 b | 0.02 ± 0.02 b |
| Desulfobulbus | 0.05 ± 0.01 a | 0.05 ± 0.05 a | 0.01 ± 0.01 a | 0.04 ± 0.04 a | 0.01 ± 0.01 a | 0.04 ± 0.03 a |
| Dongia | 0.03 ± 0.01 a | 0.02 ± 0.00 a | 0.03 ± 0.02 a | 0.01 ± 0.01 a | 0.05 ± 0.05 a | 0.03 ± 0.02 a |
| Rhodovastum | 0.03 ± 0.00 ab | 0.03 ± 0.03 ab | 0.07 ± 0.02 a | 0.04 ± 0.02 ab | 0.03 ± 0.01 ab | 0.02 ± 0.01 b |
| Leptothrix | 0.01 ± 0.00 a | 0.02 ± 0.01 a | 0.04 ± 0.05 a | 0.06 ± 0.06 a | 0.06 ± 0.05 a | 0.06 ± 0.04 a |
| Pseudomonas | 0.06 ± 0.03 a | 0.05 ± 0.03 a | 0.04 ± 0.02 a | 0.04 ± 0.02 a | 0.02 ± 0.02 a | 0.03 ± 0.01 a |
| Devosia | 0.03 ± 0.02 a | 0.02 ± 0.01 a | 0.03 ± 0.02 a | 0.03 ± 0.03 a | 0.06 ± 0.04 a | 0.04 ± 0.01 a |
| Asticcacaulis | 0.02 ± 0.00 a | 0.00 ± 0.00 a | 0.02 ± 0.02 a | 0.01 ± 0.01 a | 0.02 ± 0.02 a | 0.01 ± 0.01 a |
| Hephaestia | 0.01 ± 0.01 a | 0.02 ± 0.02 a | 0.05 ± 0.05 a | 0.00 ± 0.00 a | 0.01 ± 0.01 a | 0.00 ± 0.00 a |
| Acinetobacter | 0.02 ± 0.01 a | 0.02 ± 0.01 a | 0.02 ± 0.01 a | 0.02 ± 0.02 a | 0.01 ± 0.01 a | 0.01 ± 0.01 a |
| Polaromonas | 0.02 ± 0.01 a | 0.01 ± 0.01 a | 0.04 ± 0.02 a | 0.03 ± 0.01 a | 0.02 ± 0.02 a | 0.02 ± 0.03 a |
| Sulfuricella | 0.01 ± 0.01 ab | 0.02 ± 0.01 ab | 0.03 ± 0.04ab | 0.04 ± 0.02 a | 0.00 ± 0.00 b | 0.00 ± 0.00 b |
| Candidatus_Solibacter | 2.23 ± 0.24 a | 1.17 ± 0.26 b | 2.93 ± 0.52 a | 1.33 ± 0.46 b | 2.66 ± 0.58 a | 1.39 ± 0.47 b |
| Bryobacter | 0.76 ± 0.41 a | 0.71 ± 0.39 a | 0.70 ± 0.38 a | 0.63 ± 0.13 a | 1.53 ± 0.59 a | 1.20 ± 0.82 a |
| Anaeromyxobacter | 0.18 ± 0.02 a | 0.23 ± 0.14 a | 0.35 ± 0.22 a | 0.37 ± 0.07 a | 0.33 ± 0.16 a | 0.27 ± 0.10 a |
| Candidatus_Koribacter | 0.14 ± 0.05 ab | 0.12 ± 0.05 ab | 0.01 ± 0.01 b | 0.04 ± 0.02 b | 0.38 ± 0.29 a | 0.20 ± 0.18 ab |
| Thermoanaerobaculum | 0.02 ± 0.02 a | 0.02 ± 0.02 a | 0.08 ± 0.07 a | 0.10 ± 0.11 a | 0.07 ± 0.01 a | 0.11 ± 0.09 a |
| Terracidiphilus | 0.03 ± 0.02 a | 0.01 ± 0.00 a | 0.05 ± 0.04 a | 0.02 ± 0.02 a | 0.04 ± 0.03 a | 0.03 ± 0.02 a |
| Geothrix | 0.03 ± 0.01 a | 0.06 ± 0.04 a | 0.05 ± 0.05 a | 0.05 ± 0.03 a | 0.01 ± 0.01 a | 0.02 ± 0.02 a |
| Candidatus_Planktophila | 0.00 ± 0.00 a | 0.00 ± 0.00 a | 0.00 ± 0.00 a | 0.01 ± 0.01 a | 0.00 ± 0.00 a | 0.01 ± 0.01 a |
| Longilinea | 0.02 ± 0.02 b | 0.99 ± 0.44 a | 0.08 ± 0.05 b | 0.85 ± 0.16 a | 0.02 ± 0.01 b | 1.17 ± 0.15 a |
| Leptolinea | 0.07 ± 0.05 b | 0.32 ± 0.19 a | 0.03 ± 0.01 b | 0.26 ± 0.13 ab | 0.08 ± 0.03 b | 0.32 ± 0.20 a |
| Anaerolinea | 0.12 ± 0.08 a | 0.15 ± 0.11 a | 0.11 ± 0.05 a | 0.17 ± 0.13 a | 0.20 ± 0.06 a | 0.30 ± 0.26 a |
| Roseiflexus | 0.02 ± 0.01 a | 0.02 ± 0.02 a | 0.03 ± 0.00 a | 0.02 ± 0.02 a | 0.02 ± 0.01 a | 0.02 ± 0.02 a |
| Ferruginibacter | 0.43 ± 0.04 a | 0.45 ± 0.22 a | 0.41 ± 0.29 a | 0.34 ± 0.19 a | 0.49 ± 0.32 a | 0.33 ± 0.29 a |
| Terrimonas | 0.75 ± 0.26 a | 0.12 ± 0.14 b | 0.33 ± 0.10 ab | 0.12 ± 0.08 b | 0.75 ± 0.17 a | 0.08 ± 0.01 b |
| Paludibacter | 0.10 ± 0.06 a | 0.12 ± 0.10 a | 0.13 ± 0.20 a | 0.14 ± 0.11 a | 0.03 ± 0.03 a | 0.07 ± 0.04 a |
| Flavobacterium | 0.13 ± 0.07 a | 0.18 ± 0.08 a | 0.09 ± 0.07 a | 0.06 ± 0.05 a | 0.16 ± 0.07 a | 0.22 ± 0.07 a |
| Mucilaginibacter | 0.04 ± 0.03 a | 0.03 ± 0.01 a | 0.08 ± 0.05 a | 0.07 ± 0.05 a | 0.11 ± 0.06 a | 0.07 ± 0.05 a |
| Parafilimonas | 0.03 ± 0.02 a | 0.00 ± 0.00 b | 0.01 ± 0.01 ab | 0.00 ± 0.00 b | 0.01 ± 0.01 ab | 0.01 ± 0.01 ab |
| Opitutus | 3.53 ± 0.94 a | 2.94 ± 0.71 a | 2.58 ± 0.45 a | 2.30 ± 0.65 a | 2.74 ± 0.39 a | 2.82 ± 0.85 a |
| Spirochaeta | 0.57 ± 0.23 a | 0.57 ± 0.02 a | 0.58 ± 0.21 a | 0.54 ± 0.12 a | 0.53 ± 0.30 a | 0.66 ± 0.56 a |
| Nitrospira | 0.46 ± 0.13 ab | 0.04 ± 0.02 b | 0.54 ± 0.12 ab | 0.11 ± 0.09 b | 0.74 ± 0.12 a | 0.19 ± 0.13 b |
| Caldisericum | 0.00 ± 0.00 a | 0.01 ± 0.00 a | 0.00 ± 0.00 a | 0.02 ± 0.00 a | 0.00 ± 0.00 a | 0.09 ± 0.07 a |
| Planctomyces | 0.07 ± 0.00 a | 0.08 ± 0.07 a | 0.11 ± 0.06 a | 0.12 ± 0.06 a | 0.09 ± 0.04 a | 0.09 ± 0.02 a |
| Candidatus_Omnitrophus | 0.17 ± 0.05 a | 0.21 ± 0.07 a | 0.12 ± 0.07 a | 0.39 ± 0.14 a | 0.08 ± 0.06 a | 0.33 ± 0.06 a |
| Oryzihumus | 0.12 ± 0.04 a | 0.08 ± 0.07 a | 0.06 ± 0.03 a | 0.10 ± 0.06 a | 0.46 ± 0.14 a | 0.17 ± 0.07 a |
| Blastococcus | 0.00 ± 0.00 a | 0.00 ± 0.00 a | 0.001 ± 0.01 a | 0.00 ± 0.00 a | 0.01 ± 0.01 a | 0.00 ± 0.00 a |
| Gemmatimonas | 0.79 ± 0.58 a | 0.03 ± 0.03 b | 0.71 ± 0.40 ab | 0.05 ± 0.02 b | 1.06 ± 0.61 a | 0.05 ± 0.02 b |
| Model | Equation | R2 | Pr |
|---|---|---|---|
| Plane model | Ratio = 10.48 − 3.98 × density + 0.03 × WT | 0.873 | p < 0.05 |
| Poly2D model | Ratio = 11.12 − 5.04 × density − 0.01 × WT + 0.23 × density2 − 6.84 × WT2 + 0.003 × density × WT | 0.851 | p < 0.05 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
He, Y.; Xu, Y.; Lv, Y.; Nie, L.; Wang, H. Soil Bacterial Community Structure in Turfy Swamp and Its Response to Highway Disturbance. Int. J. Environ. Res. Public Health 2020, 17, 7822. https://doi.org/10.3390/ijerph17217822
He Y, Xu Y, Lv Y, Nie L, Wang H. Soil Bacterial Community Structure in Turfy Swamp and Its Response to Highway Disturbance. International Journal of Environmental Research and Public Health. 2020; 17(21):7822. https://doi.org/10.3390/ijerph17217822
Chicago/Turabian StyleHe, Yuanyuan, Yan Xu, Yan Lv, Lei Nie, and Hong Wang. 2020. "Soil Bacterial Community Structure in Turfy Swamp and Its Response to Highway Disturbance" International Journal of Environmental Research and Public Health 17, no. 21: 7822. https://doi.org/10.3390/ijerph17217822
APA StyleHe, Y., Xu, Y., Lv, Y., Nie, L., & Wang, H. (2020). Soil Bacterial Community Structure in Turfy Swamp and Its Response to Highway Disturbance. International Journal of Environmental Research and Public Health, 17(21), 7822. https://doi.org/10.3390/ijerph17217822





