Community-Acquired Antimicrobial Resistant Enterobacteriaceae in Central America: A One Health Systematic Review
Abstract
1. Introduction
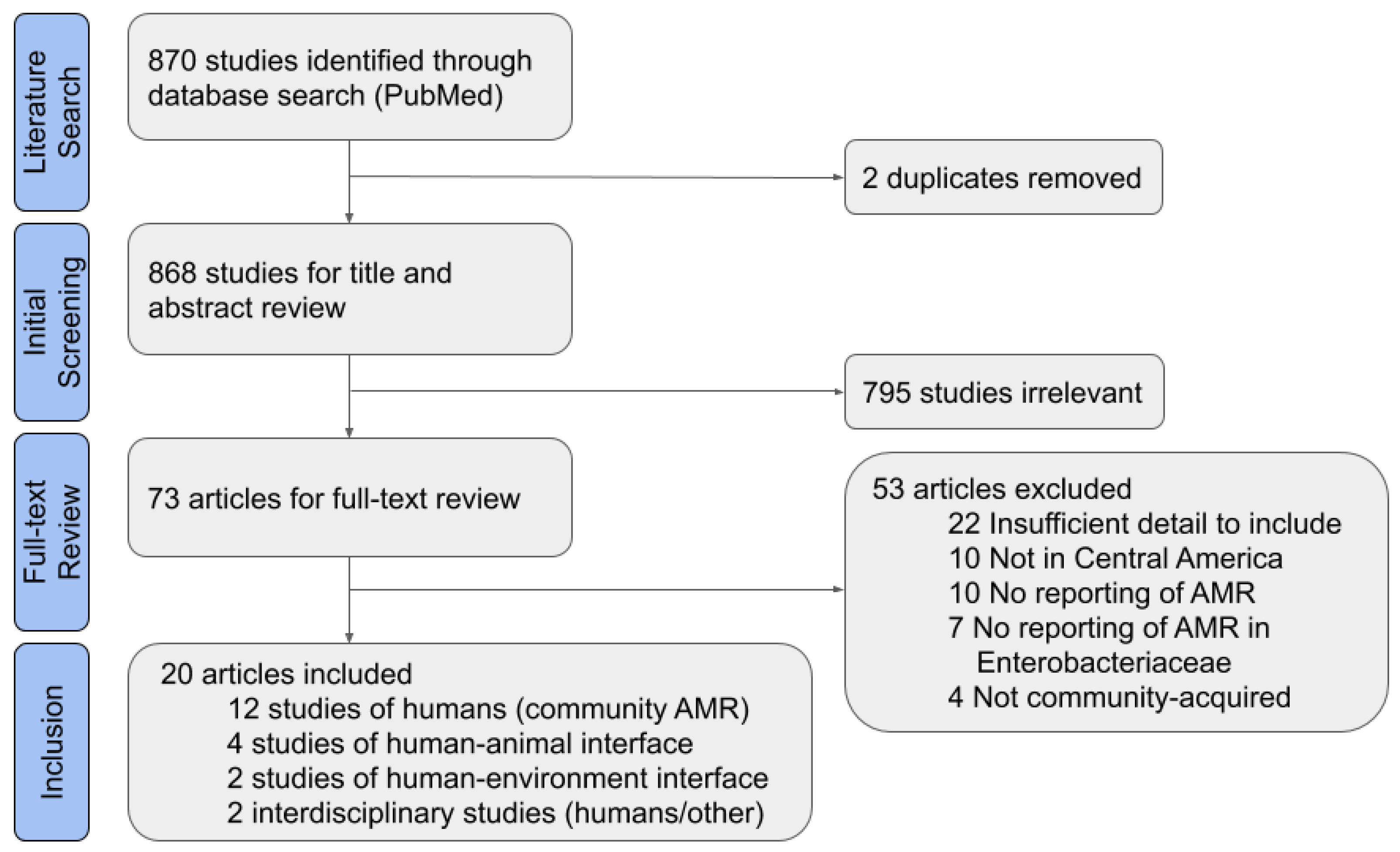
2. Materials and Methods
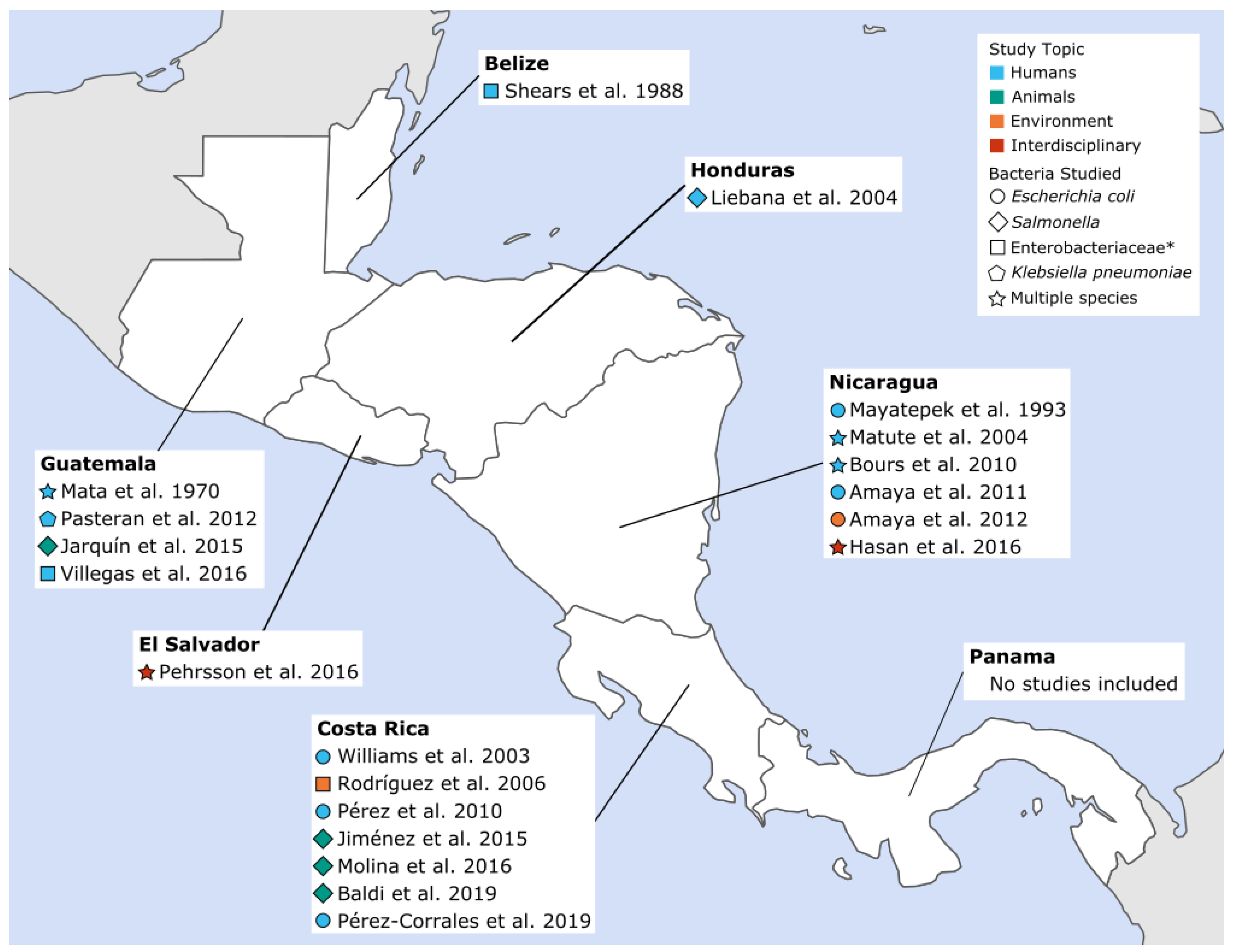
3. Results
3.1. CA-ARE in Humans
3.2. CA-ARE at the Human–Animal Interface
3.3. CA-ARE at the Human–Environment Interface
3.4. CA-ARE in Interdisciplinary Studies
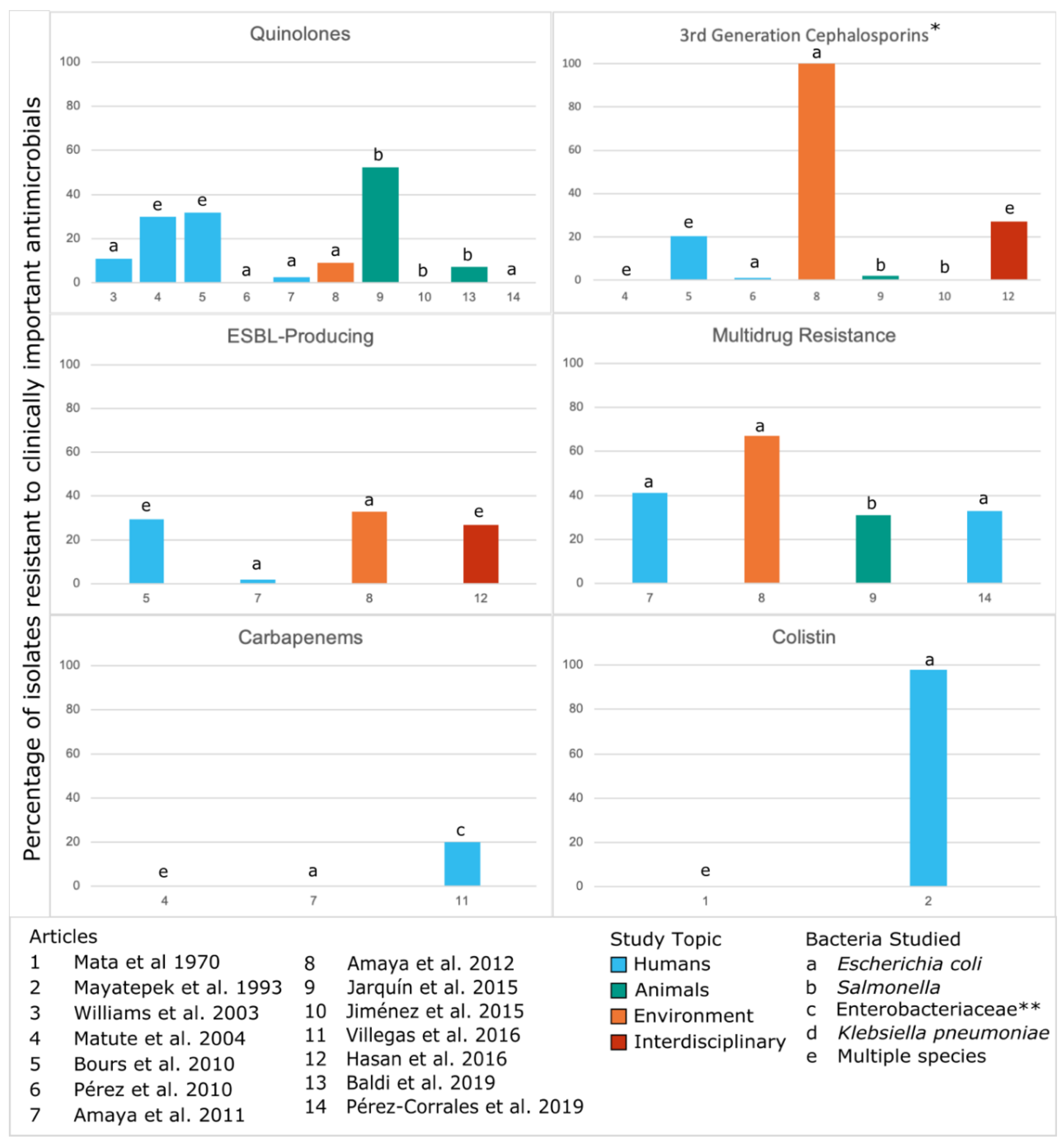
3.5. CA-ARE Across Studies
4. Discussion
5. Future Directions
6. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- World Health Organization. Global Priority List of Antibiotic-Resistant Bacteria to Guide Research, Discovery, and Development of New Antibiotics; World Health Organization: Switzerland, Geneva, 2017; Available online: https://www.who.int/medicines/publications/WHO-PPL-Short_Summary_25Feb-ET_NM_WHO.pdf (accessed on 16 October 2020).
- Van Boeckel, T.P.; Pires, J.; Silvester, R.; Zhao, C.; Song, J.; Criscuolo, N.G.; Gilbert, M.; Bonhoeffer, S.; Laxminarayan, R. Global trends in antimicrobial resistance in animals in low- and middle-income countries. Science 2019, 365. [Google Scholar] [CrossRef] [PubMed]
- World Health Organization. Worldwide Country Situation Analysis: Response to Antimicrobial Resistance; World Health Organization: Switzerland, Geneva, 2015. [Google Scholar]
- Holmes, A.H.; Moore, L.S.P.; Sundsfjord, A.; Steinbakk, M.; Regmi, S.; Karkey, A.; Guerin, P.J.; Piddock, L.J. Understanding the mechanisms and drivers of antimicrobial resistance. Lancet 2016, 387, 176–187. [Google Scholar] [CrossRef]
- Silbergeld, E.K.; Graham, J.; Price, L.B. Industrial food animal production, antimicrobial resistance, and human health. Annu. Rev. Public Health 2008, 29, 151–169. [Google Scholar] [CrossRef] [PubMed]
- Graham, J.P.; Eisenberg, J.N.S.; Trueba, G.; Zhang, L.; Johnson, T.J. Small-Scale Food Animal Production and Antimicrobial Resistance: Mountain, Molehill, or Something in-between? Environ. Health Perspect. 2017, 125, 104501. [Google Scholar] [CrossRef]
- Frost, I.; Van Boeckel, T.P.; Pires, J.; Craig, J.; Laxminarayan, R. Global geographic trends in antimicrobial resistance: The role of international travel. J. Travel. Med. 2019, 26. [Google Scholar] [CrossRef] [PubMed]
- Finley, R.L.; Collignon, P.; Larsson, D.G.J.; McEwen, S.A.; Li, X.-Z.; Gaze, W.H.; Reid-Smith, R.; Timinouni, M.; Graham, D.W.; Topp, E. The scourge of antibiotic resistance: The important role of the environment. Clin. Infect. Dis. 2013, 57, 704–710. [Google Scholar] [CrossRef] [PubMed]
- von Wintersdorff, C.J.H.; Penders, J.; van Niekerk, J.M.; Mills, N.D.; Majumder, S.; van Alphen, L.B.; Savelkoul, P.H.; Wolffs, P.F. Dissemination of Antimicrobial Resistance in Microbial Ecosystems through Horizontal Gene Transfer. Front. Microbiol. 2016, 7, 173. [Google Scholar] [CrossRef]
- Van Puyvelde, S.; Deborggraeve, S.; Jacobs, J. Why the antibiotic resistance crisis requires a One Health approach. Lancet Infect. Dis. 2018, 18, 132–134. [Google Scholar] [CrossRef]
- Lammie, S.L.; Hughes, J.M. Antimicrobial Resistance, Food Safety, and One Health: The Need for Convergence. Annu. Rev. Food Sci. Technol. 2016, 7, 287–312. [Google Scholar] [CrossRef]
- Collignon, P.J.; McEwen, S.A. One Health—Its Importance in Helping to Better Control Antimicrobial Resistance. Trop. Med. Infect. Dis. 2019, 4, 22. [Google Scholar] [CrossRef]
- Moreno, P.; Cerón, A.; Sosa, K.; Morales, M.; Grajeda, L.M.; Lopez, M.R.; McCraken, J.P.; Cordón-Rosales, C.; Palmer, G.H.; Call, D.R.; et al. Availability of over-the-counter antibiotics in Guatemalan corner stores. PLoS ONE 2020, 15, e0239873. [Google Scholar] [CrossRef] [PubMed]
- Ramay, B.M.; Lambour, P.; Cerón, A. Comparing antibiotic self-medication in two socio-economic groups in Guatemala City: A descriptive cross-sectional study. BMC Pharmacol. Toxicol. 2015, 16, 11. [Google Scholar] [CrossRef] [PubMed]
- Alvarado Sosa, J.L.; Villatoro, C.R. Resistencia bacteriana en infecciones del tracto urinario de origen comunitario. Rev. Med. Interma Guatem 2016, 20, 24–30. [Google Scholar]
- Ramay, B.M.; Caudell, M.A.; Cordón-Rosales, C.; Archila, L.D.; Palmer, G.H.; Jarquin, C.; Moreno, P.; McCracken, J.P.; Rosenkrantz, L.; Amram, O.; et al. Antibiotic use and hygiene interact to influence the distribution of antimicrobial-resistant bacteria in low-income communities in Guatemala. Sci. Rep. 2020, 10, 13767. [Google Scholar] [CrossRef] [PubMed]
- PAHO/WHO ReLAVRA. Available online: https://www.paho.org/hq/index.php?option=com_content&view=article&id=6221:2017-relavra-mas-informacion&Itemid=42432&lang=en (accessed on 10 August 2020).
- Annual Report of the Network for Monitoring/Surveillance of Antibiotic Resistance and Health Care Associated Infections—2014. Revista de Patologia Tropical. 2014. Available online: https://www.paho.org/hq/dmdocuments/2017/2014-cha-informe-anual-relavra.pdf (accessed on 16 October 2020).
- World Health Organization. Global Antimicrobial Resistance Surveillance System (GLASS) Report: Early Implementation 2017–2018; World Health Organization: Switzerland, Geneva, 2018; Available online: https://apps.who.int/iris/bitstream/handle/10665/279656/9789241515061-eng.pdf?ua=1 (accessed on 16 October 2020).
- World Health Organization. Global Antimicrobial Resistance and Use Surveillance System (GLASS) Report: Early Implementation 2020; World Health Organization: Switzerland, Geneva, 2020; Available online: https://apps.who.int/iris/bitstream/handle/10665/332081/9789240005587-eng.pdf?ua=1 (accessed on 16 October 2020).
- Plan de Accion Nacional de Lucha Contra la Resistencia a los Antimicrobianos Costa Rica 2018–2025. Available online: https://www.ministeriodesalud.go.cr/index.php/vigilancia-de-la-salud/normas-protocolos-y-guias/resistencia-microbiana/3811-plan-de-accion-nacional-de-lucha-contra-la-resistencia-a-los-antimicrobianos-costa-rica-2018-2025/file (accessed on 16 October 2020).
- Moher, D.; Liberati, A.; Tetzlaff, J.; Altman, D.G.; PRISMA Group. Preferred reporting items for systematic reviews and meta-analyses: The PRISMA statement. PLoS Med. 2009, 6, e1000097. [Google Scholar] [CrossRef] [PubMed]
- Covidence. Available online: http://www.covidence.org (accessed on 5 October 2020).
- Paterson, D.L. Resistance in gram-negative bacteria: Enterobacteriaceae. Am. J. Med. 2006, 119, S20–S28; discussion S62–S70. [Google Scholar] [CrossRef] [PubMed]
- Carattoli, A. Resistance plasmid families in Enterobacteriaceae. Antimicrob. Agents Chemother. 2009, 53, 2227–2238. [Google Scholar] [CrossRef]
- Bush, K. Alarming β-lactamase-mediated resistance in multidrug-resistant Enterobacteriaceae. Curr. Opin. Microbiol. 2010, 13, 558–564. [Google Scholar] [CrossRef]
- Amaya, E.; Reyes, D.; Vilchez, S.; Paniagua, M.; Möllby, R.; Nord, C.E.; Weintraub, A. Antibiotic resistance patterns of intestinal Escherichia coli isolates from Nicaraguan children. J. Med. Microbiol. 2011, 60, 216–222. [Google Scholar] [CrossRef]
- Mata, L.J.; Gangarosa, E.J.; Cáceres, A.; Perera, D.R.; Mejicanos, M.L. Epidemic Shiga bacillus dysentery in Central America. I. Etiologic investigations in Guatemala, 1969. J. Infect. Dis. 1970, 122, 170–180. [Google Scholar] [CrossRef]
- Mayatepek, E.; Seebass, E.; Hingst, V.; Kroeger, A.; Sonntag, H.G. Prevalence of enteropathogenic and enterotoxigenic Escherichia coli in children with and without diarrhoea in Esteli, Nicaragua. J. Diarrhoeal. Dis. Res. 1993, 11, 169–171. [Google Scholar] [PubMed]
- Pérez, C.; Gómez-Duarte, O.G.; Arias, M.L. Diarrheagenic Escherichia coli in children from Costa Rica. Am. J. Trop. Med. Hyg. 2010, 83, 292–297. [Google Scholar] [CrossRef]
- Pérez-Corrales, C.; Leandro-Sandí, K. Diarrheagenic Escherichia coli in Costa Rican children: A 9-year retrospective study. BMC Res. Notes 2019, 12, 297. [Google Scholar] [CrossRef]
- Shears, P.; Hart, C.A.; Suliman, G. A preliminary investigation of antibiotic resistance in Enterobacteriaceae isolated from children with diarrhoea from four developing countries. Ann. Trop. Med. Parasitol. 1988, 82, 185–188. [Google Scholar] [CrossRef] [PubMed]
- Bours, P.H.A.; Polak, R.; Hoepelman, A.I.M.; Delgado, E.; Jarquin, A.; Matute, A.J. Increasing resistance in community-acquired urinary tract infections in Latin America, five years after the implementation of national therapeutic guidelines. Int. J. Infect. Dis. 2010, 14, e770–e774. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Matute, A.J.; Hak, E.; Schurink, C.A.M.; McArthur, A.; Alonso, E.; Paniagua, M.; Van Asbeck, E.; Roskott, A.M.; Froeling, F.; Rozenberg-Arska, M.; et al. Resistance of uropathogens in symptomatic urinary tract infections in León, Nicaragua. Int. J. Antimicrob. Agents 2004, 23, 506–509. [Google Scholar] [CrossRef] [PubMed]
- Williams, D.N.; Sannes, M.R.; Eckhoff, A.A.; Peterson, P.K.; Johnson, J.R.; Sannes, M.R. Antimicrobial resistance in Escherichia coli causing urinary tract infections in Costa Rica: A clinical dilemma. Int. J. Antimicrob. Agents 2003, 21, 79–81. [Google Scholar] [CrossRef]
- Villegas, M.V.; Pallares, C.J.; Escandón-Vargas, K.; Hernández-Gómez, C.; Correa, A.; Álvarez, C.; Rosso, F.; Matta, L.; Luna, C.; Zurita, J.; et al. Characterization and Clinical Impact of Bloodstream Infection Caused by Carbapenemase-Producing Enterobacteriaceae in Seven Latin American Countries. PLoS ONE 2016, 11, e0154092. [Google Scholar] [CrossRef]
- Liebana, E.; Batchelor, M.; Torres, C.; Briñas, L.; Lagos, L.A.; Abdalhamid, B.; Hanson, N.D.; Martinez-Urtaza, J. Pediatric infection due to multiresistant Salmonella enterica serotype Infantis in Honduras. J. Clin. Microbiol. 2004, 42, 4885–4888. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Pasteran, F.; Albornoz, E.; Faccone, D.; Gomez, S.; Valenzuela, C.; Morales, M.; Estrada, P.; Valenzuela, L.; Matheu, J.; Guerriero, L.; et al. Emergence of NDM-1-producing Klebsiella pneumoniae in Guatemala. J. Antimicrob. Chemother. 2012, 67, 1795–1797. [Google Scholar] [CrossRef]
- Jiménez, R.R.; Barquero-Calvo, E.; Abarca, J.G.; Porras, L.P. Salmonella Isolates in the Introduced Asian House Gecko (Hemidactylus frenatus) with Emphasis on Salmonella Weltevreden, in Two Regions in Costa Rica. Vector. Borne Zoonotic. Dis. 2015, 15, 550–555. [Google Scholar] [CrossRef] [PubMed]
- Baldi, M.; Barquero Calvo, E.; Hutter, S.E.; Walzer, C. Salmonellosis detection and evidence of antibiotic resistance in an urban raccoon population in a highly populated area, Costa Rica. Zoonoses Public Health 2019, 66, 852–860. [Google Scholar] [CrossRef] [PubMed]
- Molina, A.; Granados-Chinchilla, F.; Jiménez, M.; Acuña-Calvo, M.T.; Alfaro, M.; Chavarría, G. Vigilance for Salmonella in Feedstuffs Available in Costa Rica: Prevalence, Serotyping and Tetracycline Resistance of Isolates Obtained from 2009 to 2014. Foodborne Pathog. Dis. 2016, 13, 119–127. [Google Scholar] [CrossRef]
- Jarquin, C.; Alvarez, D.; Morales, O.; Morales, A.J.; López, B.; Donado, P.; Valencia, M.F.; Arevalo, A.; Munoz, F.; Walls, I.; et al. Salmonella on Raw Poultry in Retail Markets in Guatemala: Levels, Antibiotic Susceptibility, and Serovar Distribution. J. Food Prot. 2015, 78, 1642–1650. [Google Scholar] [CrossRef]
- Rodríguez, C.; Lang, L.; Wang, A.; Altendorf, K.; García, F.; Lipski, A. Lettuce for Human Consumption Collected in Costa Rica Contains Complex Communities of Culturable Oxytetracycline- and Gentamicin-Resistant Bacteria. Appl. Environ. Microbiol. 2006, 72, 5870–5876. [Google Scholar] [CrossRef] [PubMed]
- Amaya, E.; Reyes, D.; Paniagua, M.; Calderón, S.; Rashid, M.-U.; Colque, P.; Kühn, I.; Möllby, R.; Weintraub, A.; Nord, C.E. Antibiotic resistance patterns of Escherichia coli isolates from different aquatic environmental sources in León, Nicaragua. Clin. Microbiol. Infect. 2012, 18, E347–E354. [Google Scholar] [CrossRef]
- Hasan, B.; Laurell, K.; Rakib, M.M.; Ahlstedt, E.; Hernandez, J.; Caceres, M.; Järhult, J.D. Fecal Carriage of Extended-Spectrum β-Lactamases in Healthy Humans, Poultry, and Wild Birds in León, Nicaragua-A Shared Pool of blaCTX-M Genes and Possible Interspecies Clonal Spread of Extended-Spectrum β-Lactamases-Producing Escherichia coli. Microb. Drug Resist. 2016, 22, 682–687. [Google Scholar] [CrossRef]
- Pehrsson, E.C.; Tsukayama, P.; Patel, S.; Mejía-Bautista, M.; Sosa-Soto, G.; Navarrete, K.M.; Calderon, M.; Cabrera, L.; Hoyos-Arango, W.; Bertoli, M.T.; et al. Interconnected microbiomes and resistomes in low-income human habitats. Nature 2016, 533, 212–216. [Google Scholar] [CrossRef]
- Committee for Medicinal Products for Veterinary Use; Committee for Medicinal Products for Human Use. Categorisation of antibiotics in the European Union; European Medicines Agency: London, UK, 2019; Available online: https://www.ema.europa.eu/en/documents/report/categorisation-antibiotics-european-union-answer-request-european-commission-updating-scientific_en.pdf (accessed on 16 October 2020).
- Mitchell, M.E.V.; Alders, R.; Unger, F.; Nguyen-Viet, H.; Le, T.T.H.; Toribio, J.-A. The challenges of investigating antimicrobial resistance in Vietnam—What benefits does a One Health approach offer the animal and human health sectors? BMC Public Health 2020, 20, 213. [Google Scholar] [CrossRef]
- Rüegg, S.R.; Nielsen, L.R.; Buttigieg, S.C.; Santa, M.; Aragrande, M.; Canali, M.; Ehlinger, T.; Chantziaras, I.; Boriani, E.; Radeski, M.; et al. A Systems Approach to Evaluate One Health Initiatives. Front. Vet. Sci. 2018, 5, 23. [Google Scholar] [CrossRef]
- Rozwandowicz, M.; Brouwer, M.S.M.; Fischer, J.; Wagenaar, J.A.; Gonzalez-Zorn, B.; Guerra, B.; Mevius, D.J.; Hordijk, J. Plasmids carrying antimicrobial resistance genes in Enterobacteriaceae. J. Antimicrob. Chemother. 2018, 73, 1121–1137. [Google Scholar] [CrossRef] [PubMed]
- Lynch, J.P., III; Clark, N.M.; Zhanel, G.G. Evolution of antimicrobial resistance among Enterobacteriaceae (focus on extended spectrum β-lactamases and carbapenemases). Expert Opin. Pharmacother. 2013, 14, 199–210. [Google Scholar] [CrossRef] [PubMed]
- Bonelli, R.R.; Moreira, B.M.; Picão, R.C. Antimicrobial resistance among Enterobacteriaceae in South America: History, current dissemination status and associated socioeconomic factors. Drug Resist. Update 2014, 17, 24–36. [Google Scholar] [CrossRef] [PubMed]
| References | Measure (No. of Isolates) | Aminoglycosides | Penicillin ꞵ-lactams | Cephalosporins | Carbapenems | Chloramphenicols | Polymyxins | Nitrofurans | Sulfonamides | Tetracyclines | Quinolones |
|---|---|---|---|---|---|---|---|---|---|---|---|
| Belize | |||||||||||
| Shears et al. 1988 [32] | % resistance in Enterobacteriaceae isolates from diarrhea (ND) | 0 | 0 | ND | ND | 0 | ND | ND | 11 | 22 | ND |
| Guatemala | |||||||||||
| Mata et al. 1970 [28] | % resistance in Shigella dysenteriae strains from diarrhea (n = 53) | 98 | 23 | 0 | ND | 42 | 0 | 0 | 100 | 98 | ND |
| Jarquín et al. 2015 [42] | % resistance in Salmonella spp. isolates from chicken carcasses (n = 103) | 36 | 11 | 2 | ND | 4 | ND | ND | 38 | 41 | 52 |
| Villegas et al. 2016 [36] | % prevalence of carbapenemase-producing Enterobacteriaceae from blood samples (n = 20) | ND | ND | ND | 20 c | ND | ND | ND | ND | ND | ND |
| Nicaragua | |||||||||||
| Mayatepek et al. 1993 [29] | % resistance in E. coli isolates from diarrhea (ND) | 94 | 16 | ND | ND | 56 | 98 | ND | 18 | 4 | ND |
| Matute et al. 2004 [34] | % resistance in E. coli isolates from urine samples (UTI) (n = 35) | 11 | 74 | 58 | 0 | ND | ND | 0 | 63 | ND | 30 |
| Bours et al. 2010 [33] | % resistance in E. coli from urine samples (UTI) (n = 44) | 25 | 61 | 46 | ND | ND | ND | 7 | 39 | ND | 32 |
| Amaya et al. 2011 [27] | % resistance in E. coli isolates from stool samples (n = 727) | 3 b | 60 | ND | 0 | 11 | ND | ND | 64 | ND | 3 b |
| Amaya et al. 2012 [44] | % resistance in E. coli isolates from a hospital sewage sample (n = 32) | 69 | 100 | 100 | ND | 97 | ND | ND | 100 | ND | 9 |
| Hasan et al. 2016 [45] | % prevalence of ESBL-producing organisms from human stool (n = 28) | ND | ND | 27 | ND | ND | ND | ND | ND | ND | ND |
| Costa Rica | |||||||||||
| Williams et al. 2003 [35] | % resistance in E. coli isolates from outpatient urine samples (n = 171) | ND | ND | ND | ND | ND | ND | 8 | 40 | ND | 11 |
| Rodríguez et al. 2006 [43] | % resistance in Enterobacteriaceae isolates from lettuce (ND) | 100 | ND | ND | ND | ND | ND | ND | ND | 100 | ND |
| Pérez et al. 2010 [30] | % resistance in pathenogenic E. coli strains from diarrhea (n = 52) | ND a | 40 | 11 | ND | ND | ND | 0 | 13 | 11 | 0 |
| Jiménez et al. 2015 [39] | % resistance in Salmonella spp. isolates from gecko gut content (ND) | 50 | 0 | 0 | ND | 0 | ND | 0 | 50 | 0 | 0 |
| Molina et al. 2016 [41] | % resistance in Salmonella spp. isolates from animal feed (n = 110) | ND | ND | ND | ND | ND | ND | ND | ND | 7 | ND |
| Baldi et al. 2019 [40] | % resistance in Salmonella spp. isolates from raccoon fecal samples (n = 42) | ND | ND | ND | ND | ND | ND | ND | ND | ND | 7 |
| Pérez-Corrales et al. 2019 [31] | % resistance in enteroaggregative E. coli isolates from diarrhea (n = 189) | ND | 54 | ND | ND | ND | ND | ND | 34 | ND | 0 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
O’Neal, L.; Alvarez, D.; Mendizábal-Cabrera, R.; Ramay, B.M.; Graham, J. Community-Acquired Antimicrobial Resistant Enterobacteriaceae in Central America: A One Health Systematic Review. Int. J. Environ. Res. Public Health 2020, 17, 7622. https://doi.org/10.3390/ijerph17207622
O’Neal L, Alvarez D, Mendizábal-Cabrera R, Ramay BM, Graham J. Community-Acquired Antimicrobial Resistant Enterobacteriaceae in Central America: A One Health Systematic Review. International Journal of Environmental Research and Public Health. 2020; 17(20):7622. https://doi.org/10.3390/ijerph17207622
Chicago/Turabian StyleO’Neal, Lauren, Danilo Alvarez, Renata Mendizábal-Cabrera, Brooke M. Ramay, and Jay Graham. 2020. "Community-Acquired Antimicrobial Resistant Enterobacteriaceae in Central America: A One Health Systematic Review" International Journal of Environmental Research and Public Health 17, no. 20: 7622. https://doi.org/10.3390/ijerph17207622
APA StyleO’Neal, L., Alvarez, D., Mendizábal-Cabrera, R., Ramay, B. M., & Graham, J. (2020). Community-Acquired Antimicrobial Resistant Enterobacteriaceae in Central America: A One Health Systematic Review. International Journal of Environmental Research and Public Health, 17(20), 7622. https://doi.org/10.3390/ijerph17207622






