Distribution of Antibiotic Resistance Genes in Three Different Natural Water Bodies-A Lake, River and Sea
Abstract
1. Introduction
2. Materials and Methods
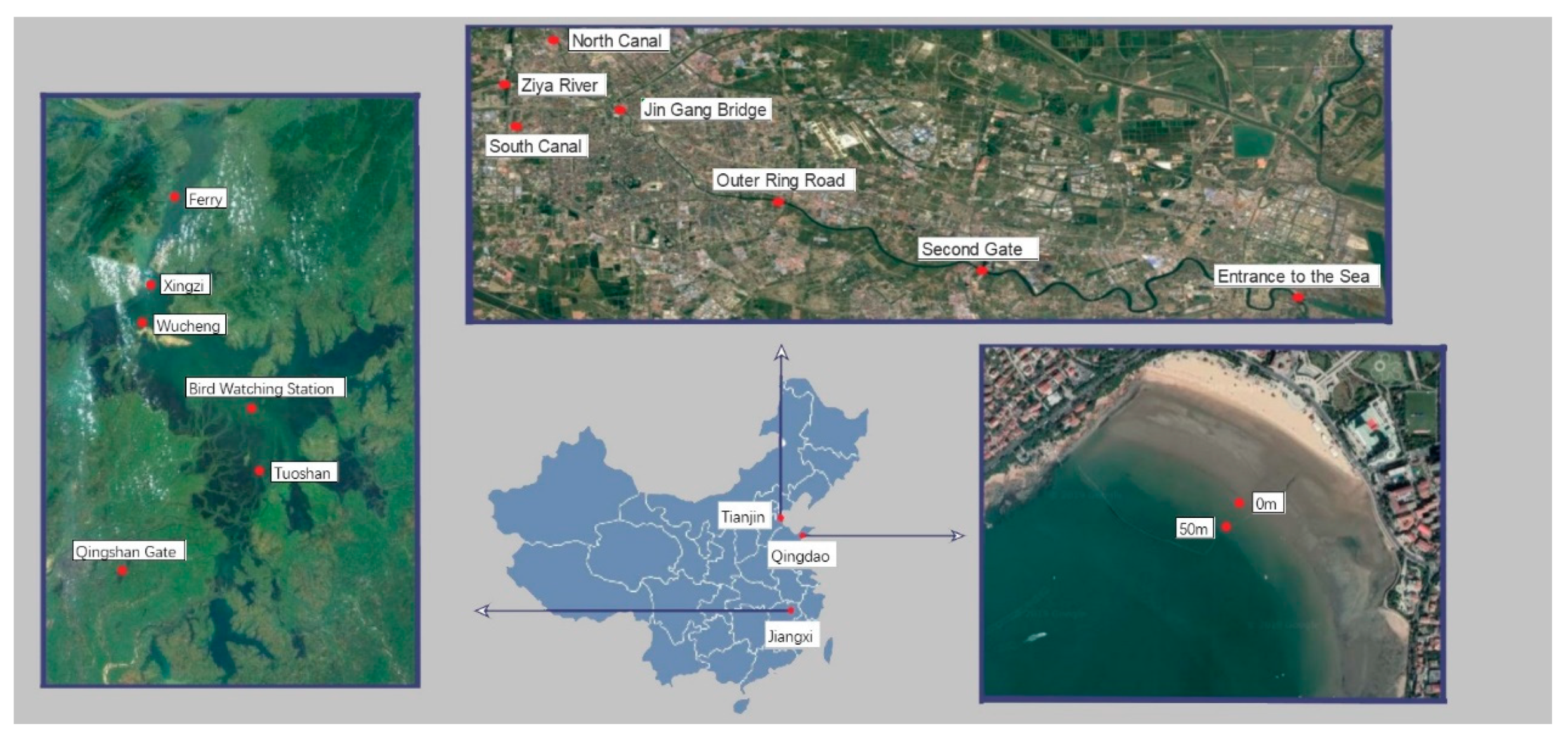
2.1. Sampling Sites and Time
2.2. DNA Extraction, Conventional PCR and qPCR
2.2.1. DNA Extraction
2.2.2. Conventional PCR
2.2.3. qPCR
2.3. Statistical Analysis
3. Results
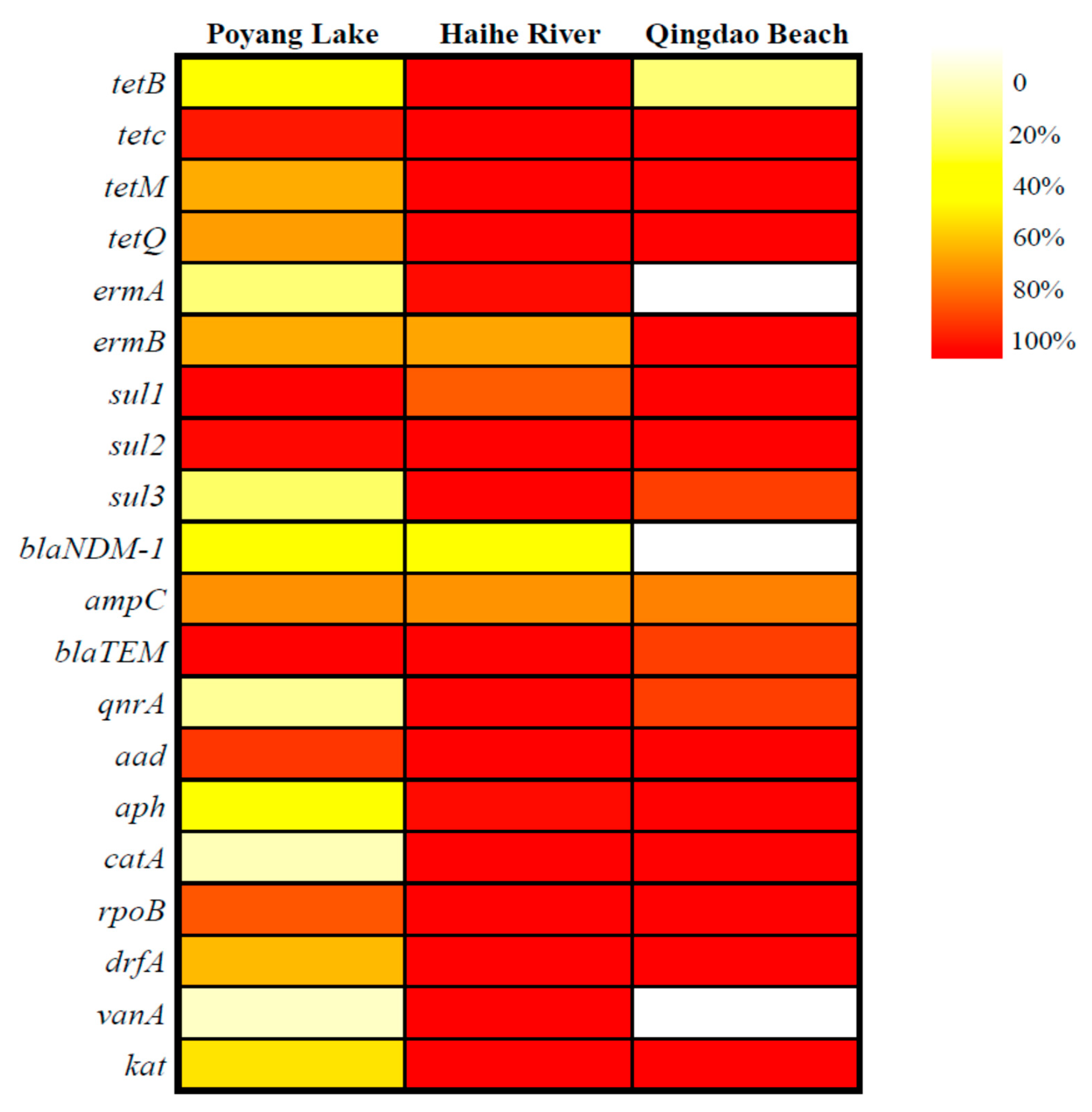
3.1. ARGs Occurrence in Water Bodies
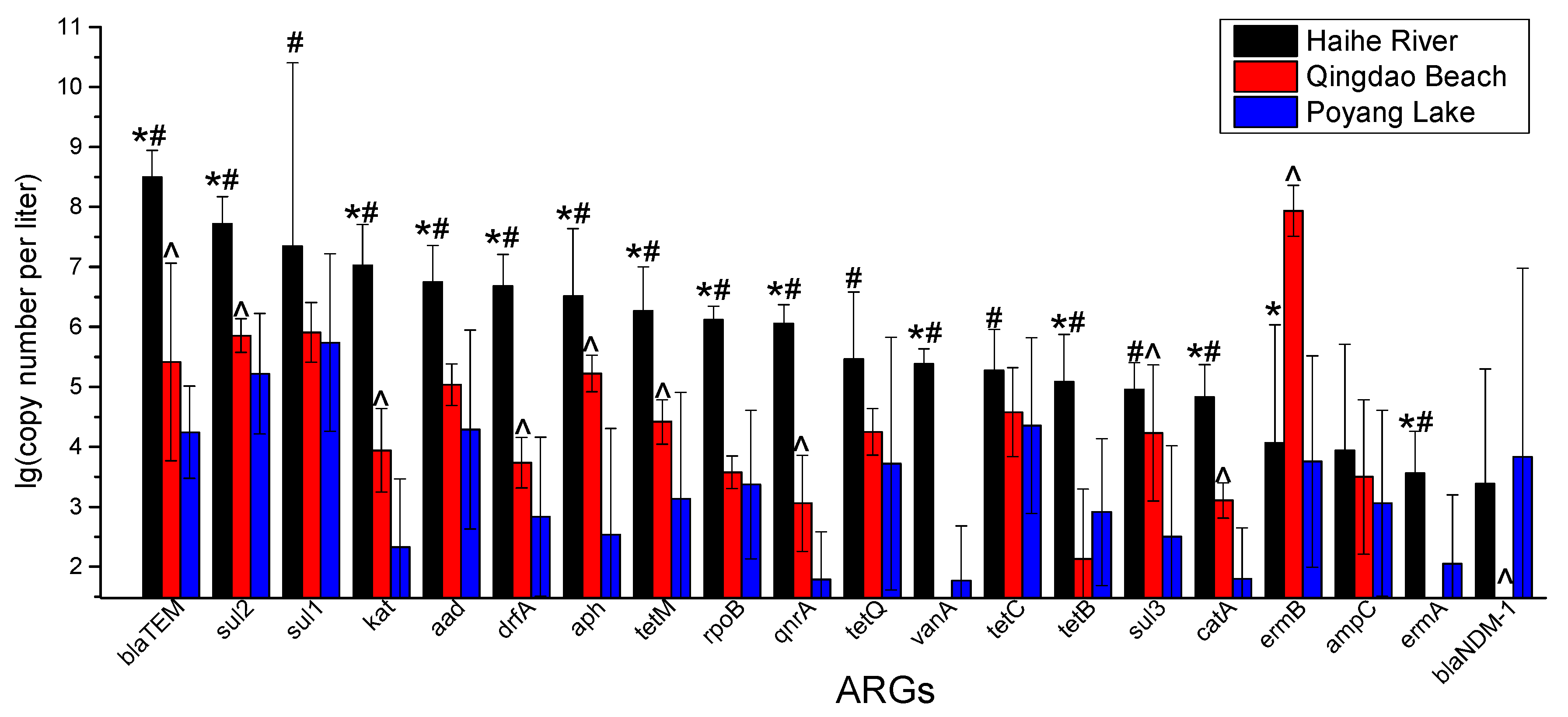
3.2. RT-qPCR Analysis of ARGs
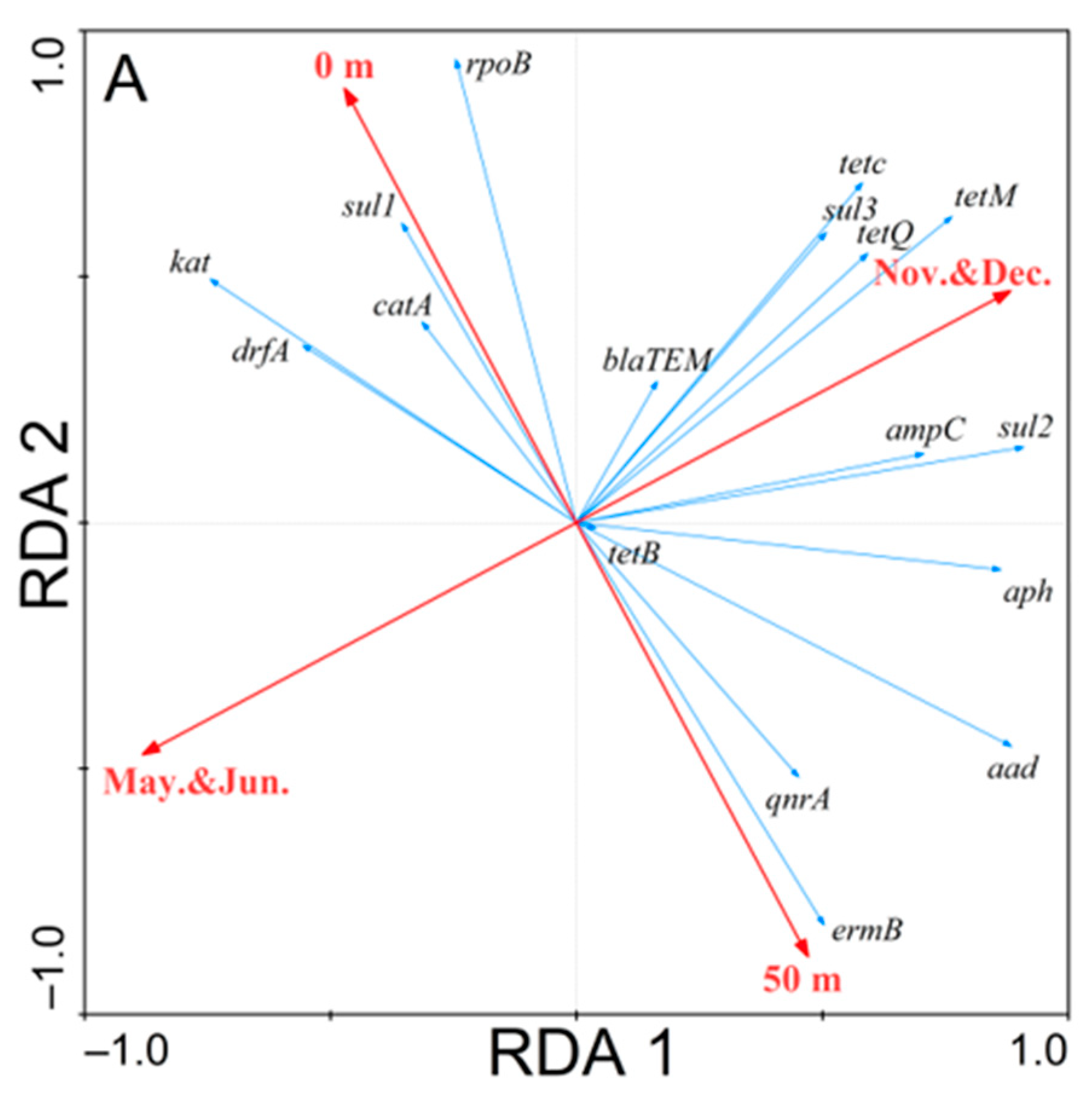
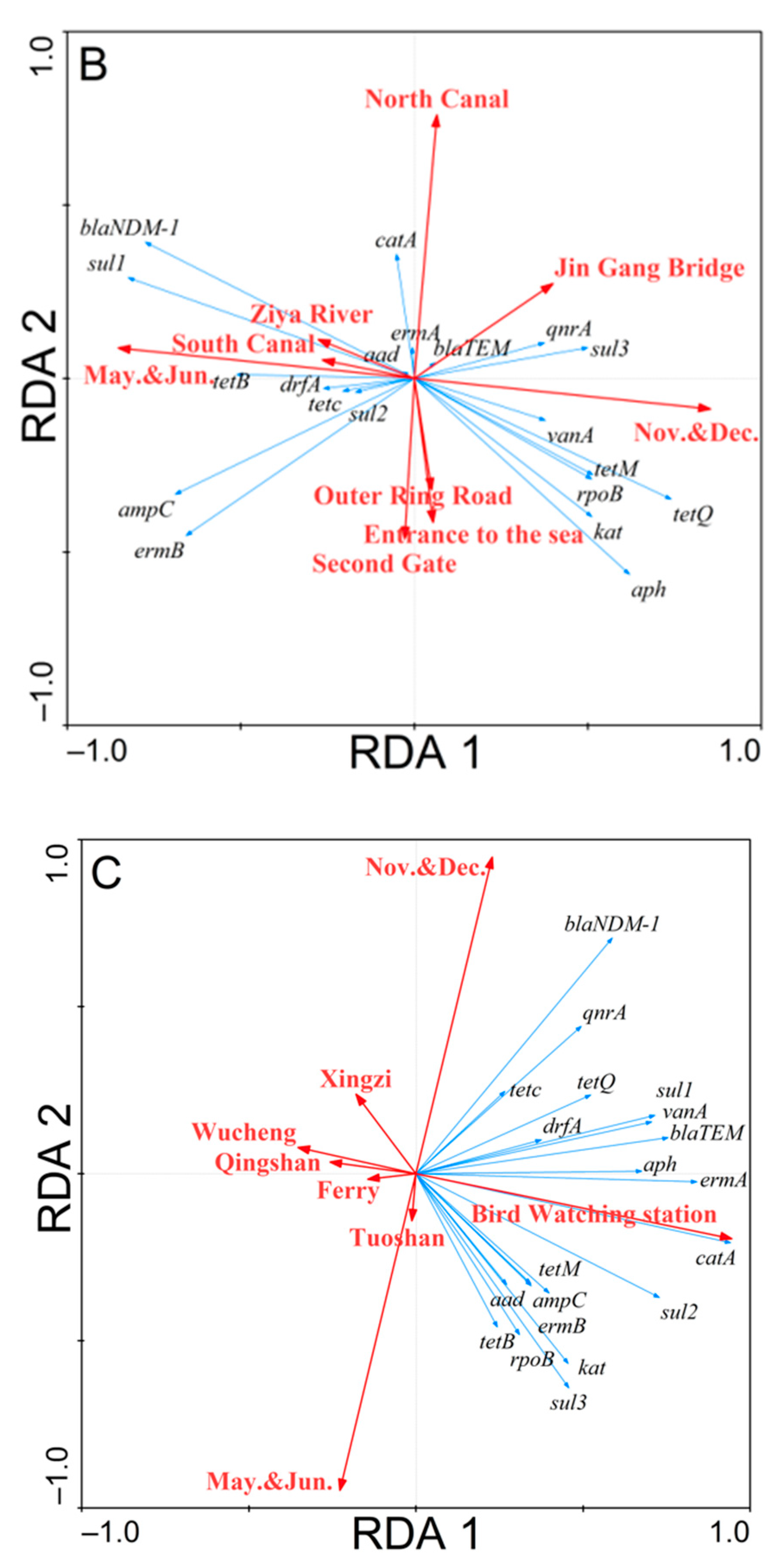
3.3. Redundancy Analysis of the Correlations
4. Discussion
4.1. Detection of the ARGs
4.2. Absolute Abundance of ARGs
4.3. Time and Spatial Distribution of ARGs
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Zhang, X.X.; Zhang, T.; Fang, H.H.P. Antibiotic resistance genes in water environment. Appl. Microbiol. Biotechnol. 2009, 82, 397–414. [Google Scholar] [CrossRef]
- Kolar, M.; Urbanek, K.; Latal, T. Antibiotic selective pressure and development of bacterial resistance. Int. J. Antimicrob. Agents 2001, 17, 357–363. [Google Scholar] [CrossRef]
- Karkman, A.; Parnanen, K.; Larsson, D. Fecal pollution can explain antibiotic resistance gene abundances in anthropogenically impacted environments. Nat. Commun. 2019, 10, 80. [Google Scholar] [CrossRef] [PubMed]
- Spellberg, B.; Gilbert, D.N. The future of antibiotics and resistance: A tribute to a career of leadership by John Bartlett. Clin. Infect. Dis. 2014, 59 (Suppl. 2), S71–S75. [Google Scholar] [CrossRef] [PubMed]
- Jasovský, D.; Littmann, J.; Zorzet, A.; Cars, O. Antimicrobial resistance—A threat to the world’s sustainable development. Upsala J. Med. Sci. 2016, 121, 159–164. [Google Scholar] [CrossRef]
- Golkar, Z.; Bagasra, O.; Pace, D.G. Bacteriophage therapy: A potential solution for the antibiotic resistance crisis. J. Infect. Dev. Ctries. 2014, 8, 129–136. [Google Scholar] [CrossRef]
- Matlou, D.P.; Bissong, M.E.A.; Tchatchouang, C.K.; Adem, M.R.; Foka, F.E.T.; Kumar, A.; Ateba, C.N. Virulence profiles of vancomycin-resistant enterococci isolated from surface and ground water utilized by humans in the North West Province, South Africa: A public health perspective. Environ. Sci. Pollut. Res. Int. 2019, 26, 15105–15114. [Google Scholar] [CrossRef]
- Rossolini, G.M.; Arena, F.; Pecile, P.; Pollini, S. Update on the antibiotic resistance crisis. Curr. Opin. Pharmacol. 2014, 18, 56–60. [Google Scholar] [CrossRef] [PubMed]
- Watkins, R.R.; Van Duin, D. Current trends in the treatment of pneumonia due to multidrug-resistant Gram-negative bacteria. F1000Research 2019, 8. [Google Scholar] [CrossRef]
- Singer, A.C.; Shaw, H.; Rhodes, V.; Hart, A. Review of Antimicrobial Resistance in the Environment and Its Relevance to Environmental Regulators. Front. Microbiol. 2016, 7, 1728. [Google Scholar] [CrossRef] [PubMed]
- Aminov, R.I. A brief history of the antibiotic era: Lessons learned and challenges for the future. Front. Microbiol. 2010, 1, 134. [Google Scholar] [CrossRef] [PubMed]
- Tyrrell, C.; Burgess, C.M.; Brennan, F.P.; Walsh, F. Antibiotic Resistance in Grass and Soil. Biochem. Soc. Trans. 2019, 47, 477–486. [Google Scholar] [CrossRef] [PubMed]
- Di Cesare, A.; Eckert, E.M.; Rogora, M.; Corno, G. Rainfall increases the abundance of antibiotic resistance genes within a riverine microbial community. Environ. Pollut. 2017, 226, 473–478. [Google Scholar] [CrossRef] [PubMed]
- Zervaa, I.; Alexandropouloub, I.; Panopouloub, M.; Melidisa, P.; Ntougiasa, S. Antibiotic resistance gene profiles at various treatment stages of a full-scale municipal sewage plant. Desalin. Water Treat. 2019, 167, 412–421. [Google Scholar] [CrossRef]
- Yang, Y.; Xu, C.; Cao, X.; Lin, H.; Wang, J. Antibiotic resistance genes in surface water of eutrophic urban lakes are related to heavy metals, antibiotics, lake morphology and anthropic impact. Ecotoxicology 2017, 26, 831–840. [Google Scholar] [CrossRef] [PubMed]
- Ma, L.; Li, B.; Jiang, X.T.; Wang, Y.L.; Xia, Y.; Li, A.D.; Zhang, T. Catalogue of antibiotic resistome and host-tracking in drinking water deciphered by a large scale survey. Microbiome 2017, 5, 154. [Google Scholar] [CrossRef]
- Du, J.; Zhao, H.; Wang, Y.; Xie, H.; Zhu, M.; Chen, J. Presence and environmental risk assessment of selected antibiotics in coastal water adjacent to mariculture areas in the Bohai Sea. Ecotoxicol. Environ. Saf. 2019, 177, 117–123. [Google Scholar] [CrossRef]
- Su, H.C.; Liu, Y.S.; Pan, C.G.; Chen, J.; He, L.Y.; Ying, G.G. Persistence of antibiotic resistance genes and bacterial community changes in drinking water treatment system: From drinking water source to tap water. Sci. Total Environ. 2018, 616, 453–461. [Google Scholar] [CrossRef]
- Caucci, S.; Karkman, A.; Cacace, D.; Rybicki, M.; Timpel, P.; Voolaid, V.; Gurke, R.; Virta, M.; Berendonk, T.U. Seasonality of antibiotic prescriptions for outpatients and resistance genes in sewers and wastewater treatment plant outflow. FEMS Microbiol. Ecol. 2016, 92, w60. [Google Scholar] [CrossRef]
- Son, D.I.; Aleta, P.; Park, M.; Yoon, H.; Cho, K.H.; Kim, Y.M.; Kim, S. Seasonal Changes in Antibiotic Resistance Genes in Rivers and Reservoirs in South Korea. J. Environ. Qual. 2018, 47, 1079–1085. [Google Scholar] [CrossRef]
- Guo, X.-P.; Liu, X.; Niu, Z.-S.; Lu, D.-P.; Zhao, S.; Sun, X.-L.; Wu, J.-Y.; Chen, Y.-R.; Tou, F.-Y.; Hou, L.; et al. Seasonal and spatial distribution of antibiotic resistance genes in the sediments along the Yangtze Estuary, China. Environ. Pollut. 2018, 242 Pt A, 576–584. [Google Scholar] [CrossRef] [PubMed]
- Seiler, C.; Berendonk, T.U. Heavy metal driven co-selection of antibiotic resistance in soil and water bodies impacted by agriculture and aquaculture. Front. Microbiol. 2012, 3, 399. [Google Scholar] [CrossRef] [PubMed]
- Wang, F.H.; Qiao, M.; Lv, Z.E.; Guo, G.X.; Jia, Y.; Su, Y.H.; Zhu, Y.G. Impact of reclaimed water irrigation on antibiotic resistance in public parks, Beijing, China. Environ. Pollut. 2014, 184, 247–253. [Google Scholar] [CrossRef] [PubMed]
- Walker, T.M.; Kohl, T.A.; Omar, S.V.; Hedge, J.; Del Ojo Elias, C.; Bradley, P.; Iqbal, Z.; Feuerriegel, S.; Niehaus, K.E.; Wilson, D.J.; et al. Whole-genome sequencing for prediction of Mycobacterium tuberculosis drug susceptibility and resistance: A retrospective cohort study. Lancet Infect. Dis. 2015, 15, 1193–1202. [Google Scholar] [CrossRef]
- Liu, Z.; Klumper, U.; Liu, Y.; Yang, Y.; Wei, Q.; Lin, J.G.; Gu, J.D.; Li, M. Metagenomic and metatranscriptomic analyses reveal activity and hosts of antibiotic resistance genes in activated sludge. Environ. Int. 2019, 129, 208–220. [Google Scholar] [CrossRef]
- Hsu, J.T.; Chen, C.Y.; Young, C.W.; Chao, W.L.; Li, M.H.; Lin, C.M.; Ying, C. Prevalence of sulfonamide-resistant bacteria, resistance genes and integron-associated horizontal gene transfer in natural water bodies and soils adjacent to a swine feedlot in northern taiwan. J. Hazard. Mater. 2014, 277, 34–43. [Google Scholar] [CrossRef]
- Liu, M.; Zhang, Y.; Yang, M.; Tian, Z.; Ren, L.; Zhang, S. Abundance and distribution of tetracycline resistance genes and mobile elements in an oxytetracycline production wastewater treatment system. Environ. Sci. Technol. 2012, 46, 7551–7557. [Google Scholar] [CrossRef]
- Zhang, T.; Zhang, M.; Zhang, X.; Fang, H.H. Tetracycline resistance genes and tetracycline resistant lactose-fermenting Enterobacteriaceae in activated sludge of sewage treatment plants. Environ. Sci. Technol. 2009, 43, 3455–3460. [Google Scholar] [CrossRef]
- Liu, Y.F.; Fu, H.M.; Wu, H.M.; Janapatle, R.P.; Wang, C.H.; Wu, J.J. P990 Presence of plasmid pA15 correlates with prevalence of constitutive MLSB resistance in group A Streptococci isolates at a university hospital in southern Taiwan. Int. J. Antimicrob. Agents 2007, 29, S263. [Google Scholar] [CrossRef]
- Hayes, J.R.; Wagner, D.D.; English, L.L.; Carr, L.E.; Joseph, S.W. Distribution of streptogramin resistance determinants among Enterococcus faecium from a poultry production environment of the USA. J. Antimicrob. Chemother. 2005, 55, 123–126. [Google Scholar] [CrossRef]
- Laport, M.S.; Pontes, P.V.; Dos Santos, D.S.; Santos-Gandelman Jde, F.; Muricy, G.; Bauwens, M.; Giambiagi-deMarval, M.; George, I. Antibiotic resistance genes detected in the marine sponge Petromica citrina from Brazilian coast. Braz. J. Microbiol. 2016, 47, 617–620. [Google Scholar] [CrossRef] [PubMed]
- Shen, Y.; Xiao, W.Q.; Gong, J.M.; Pan, J.; Xu, Q.X. Detection of New Delhi Metallo-Beta-Lactamase (Encoded by blaNDM-1) in Enterobacter aerogenes in China. J. Clin. Lab. Anal. 2017, 31. [Google Scholar] [CrossRef] [PubMed]
- Bailey, J.K.; Pinyon, J.L.; Anantham, S.; Hall, R.M. Distribution of the blaTEM gene and blaTEM-containing transposons in commensal Escherichia coli. J. Antimicrob. Chemother. 2011, 66, 745–751. [Google Scholar] [CrossRef] [PubMed]
- Fukutsu, N.; Kawasaki, T.; Saito, K.; Nakazawa, H. Application of high-performance liquid chromatography hyphenated techniques for identification of degradation products of cefpodoxime proxetil. J. Chromatogr. A 2006, 1129, 153–159. [Google Scholar] [CrossRef]
- Arabi, H.; Pakzad, I.; Nasrollahi, A.; Hosainzadegan, H.; Azizi Jalilian, F.; Taherikalani, M.; Samadi, N.; Monadi Sefidan, A. Sulfonamide Resistance Genes (sul) M in Extended Spectrum Beta Lactamase (ESBL) and Non-ESBL Producing Escherichia coli Isolated from Iranian Hospitals. Jundishapur J. Microbiol. 2015, 8, e19961. [Google Scholar] [CrossRef]
- Luo, D.; Chen, Q.; Xiong, G.; Peng, Y.; Liu, T.; Chen, X.; Zeng, L.; Chen, K. Prevalence and molecular characterization of multidrug-resistant M. tuberculosis in Jiangxi province, China. Sci. Rep. 2019, 9, 7315. [Google Scholar] [CrossRef]
- Kendall, E.A.; Azman, A.S.; Cobelens, F.G.; Dowdy, D.W. MDR-TB treatment as prevention: The projected population-level impact of expanded treatment for multidrug-resistant tuberculosis. PLoS ONE 2017, 12, e0172748. [Google Scholar] [CrossRef]
- Garner, E.; Benitez, R.; von Wagoner, E.; Sawyer, R.; Schaberg, E.; Hession, W.C.; Krometis, L.H.; Badgley, B.D.; Pruden, A. Stormwater loadings of antibiotic resistance genes in an urban stream. Water Res. 2017, 123, 144–152. [Google Scholar] [CrossRef]
- Cole, D.; Drum, D.J.; Stalknecht, D.E.; White, D.G.; Lee, M.D.; Ayers, S.; Sobsey, M.; Maurer, J.J. Free-living Canada geese and antimicrobial resistance. Emerg. Infect. Dis. 2005, 11, 935–938. [Google Scholar] [CrossRef]
- Dolejska, M.; Cizek, A.; Literak, I. High prevalence of antimicrobial-resistant genes and integrons in Escherichia coli isolates from Black-headed Gulls in the Czech Republic. J. Appl. Microbiol. 2007, 103, 11–19. [Google Scholar] [CrossRef]
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Su, S.; Li, C.; Yang, J.; Xu, Q.; Qiu, Z.; Xue, B.; Wang, S.; Zhao, C.; Xiao, Z.; Wang, J.; et al. Distribution of Antibiotic Resistance Genes in Three Different Natural Water Bodies-A Lake, River and Sea. Int. J. Environ. Res. Public Health 2020, 17, 552. https://doi.org/10.3390/ijerph17020552
Su S, Li C, Yang J, Xu Q, Qiu Z, Xue B, Wang S, Zhao C, Xiao Z, Wang J, et al. Distribution of Antibiotic Resistance Genes in Three Different Natural Water Bodies-A Lake, River and Sea. International Journal of Environmental Research and Public Health. 2020; 17(2):552. https://doi.org/10.3390/ijerph17020552
Chicago/Turabian StyleSu, Sicong, Chenyu Li, Jiping Yang, Qunying Xu, Zhigang Qiu, Bin Xue, Shang Wang, Chen Zhao, Zhonghai Xiao, Jingfeng Wang, and et al. 2020. "Distribution of Antibiotic Resistance Genes in Three Different Natural Water Bodies-A Lake, River and Sea" International Journal of Environmental Research and Public Health 17, no. 2: 552. https://doi.org/10.3390/ijerph17020552
APA StyleSu, S., Li, C., Yang, J., Xu, Q., Qiu, Z., Xue, B., Wang, S., Zhao, C., Xiao, Z., Wang, J., & Shen, Z. (2020). Distribution of Antibiotic Resistance Genes in Three Different Natural Water Bodies-A Lake, River and Sea. International Journal of Environmental Research and Public Health, 17(2), 552. https://doi.org/10.3390/ijerph17020552



