Oral Microbiome of Children Living in an Isolated Area in Myanmar
Abstract
1. Introduction
2. Materials and Methods
2.1. Subject
2.2. Oral Examination
2.3. Sample Collection
2.4. Microbial DNA Extraction
2.5. Microbial Community Analysis
2.6. Bioinformatics Analysis
2.7. Ethical Approval
3. Results
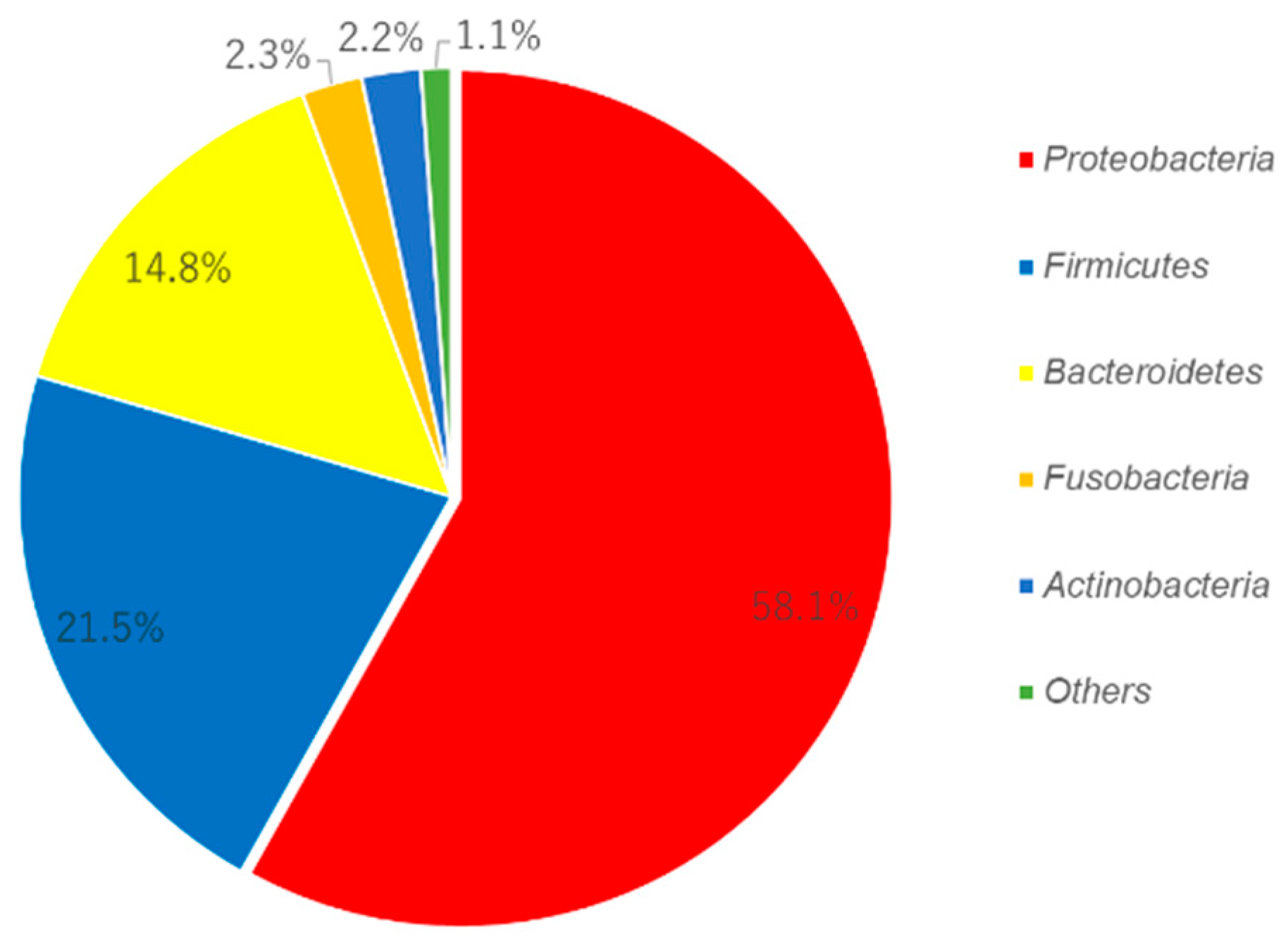
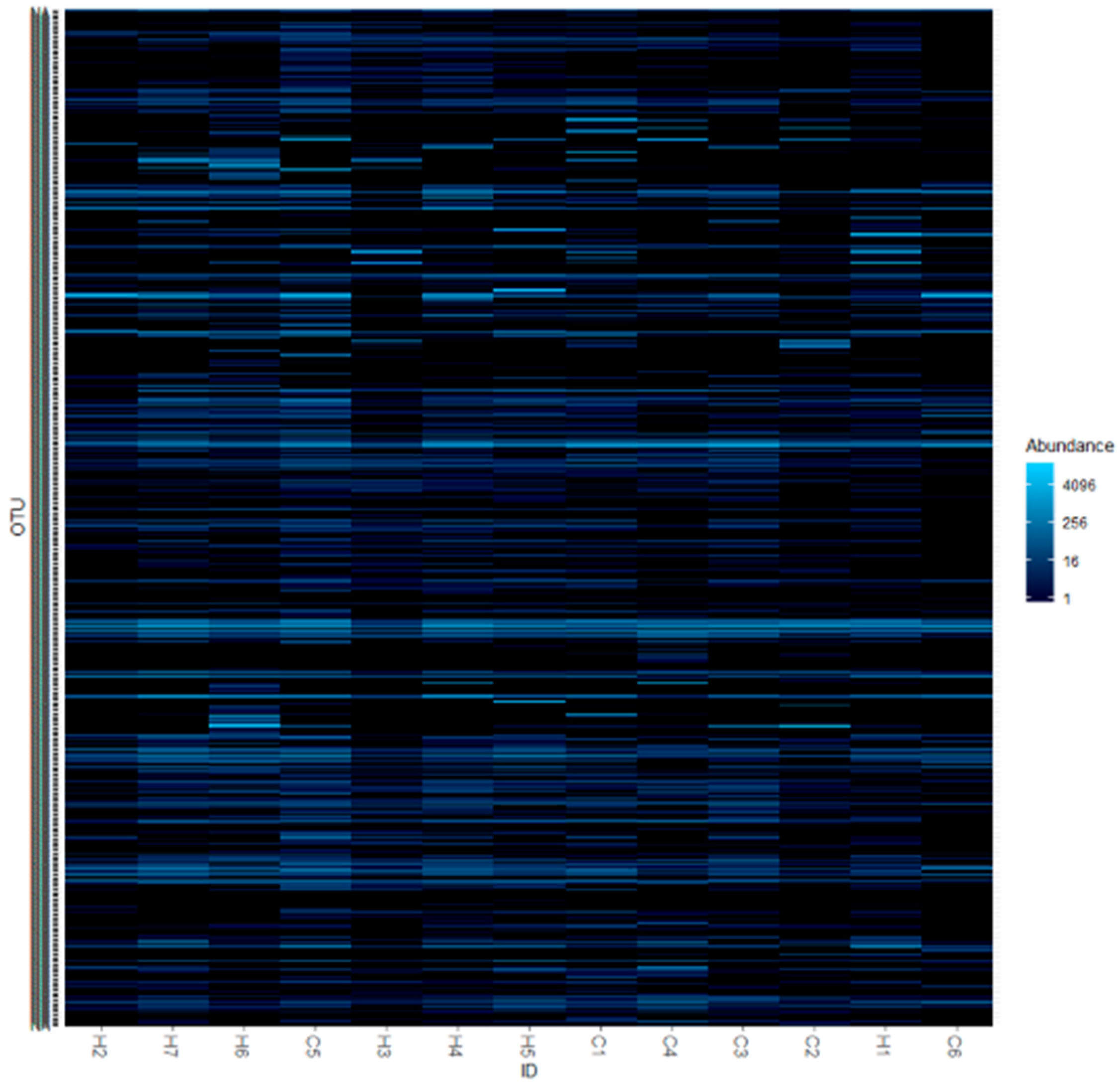
3.1. Candidate for the Core Microbiome
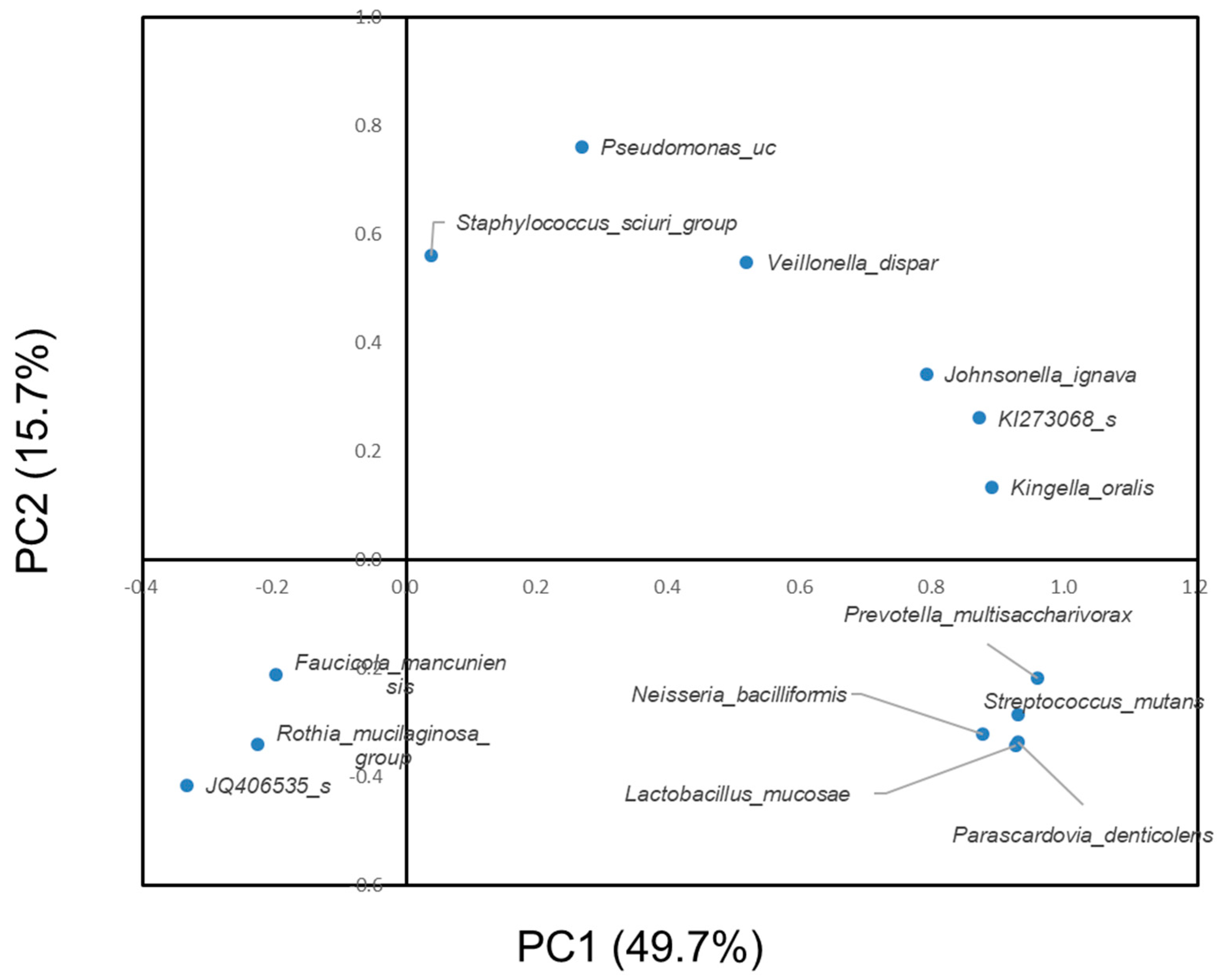
3.2. Difference in the Oral Microbiome between Subjects with or without Dental Caries
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Aas, J.A.; Paster, B.J.; Stokes, L.N.; Olsen, I.; Dewhirst, F.E. Defining the normal bacterial flora of the oral cavity. J. Clin. Microbiol. 2005, 43, 5721–5732. [Google Scholar] [CrossRef]
- Keijser, B.J.; Zaura, E.; Huse, S.M.; van der Vossen, J.M.; Schuren, F.H.; Montijn, R.C.; ten Cate, J.M.; Crielaard, W. Pyrosequencing analysis of the oral microflora of healthy adults. J. Dent. Res. 2008, 87, 1016–1020. [Google Scholar] [CrossRef]
- Wade, W.G. The oral microbiome in health and disease. Pharmacol. Res. 2013, 69, 137–143. [Google Scholar] [CrossRef]
- Costalonga, M.; Herzberg, M.C. The oral microbiome and the immunobiology of periodontal disease and caries. Immunol. Lett. 2014, 162, 22–38. [Google Scholar] [CrossRef] [PubMed]
- Greenwood, D.; Afacan, B.; Emingil, G.; Bostanci, N.; Belibasakis, G.N. Salivary Microbiome Shifts in Response to Periodontal Treatment Outcome. Proteom. Clin. Appl. 2020, 29, e2000011. [Google Scholar] [CrossRef] [PubMed]
- Sampaio-Maia, B.; Caldas, I.M.; Pereira, M.L.; Pérez-Mongiovi, D.; Araujo, R. The Oral Microbiome in Health and Its Implication in Oral and Systemic Diseases. Adv. Appl. Microbiol. 2016, 97, 171–210. [Google Scholar] [CrossRef] [PubMed]
- Patini, R. Oral Microbiota: Discovering and Facing the New Associations with Systemic Diseases. Pathogens 2020, 9, 313. [Google Scholar] [CrossRef] [PubMed]
- Grusell, E.N.; Dahlén, G.; Ruth, M.; Bergquist, H.; Bove, M. The Cultivable Bacterial Flora of the Esophagus in Subjects with Esophagitis. Scand. J. Gastroenterol. 2018, 53, 650–656. [Google Scholar] [CrossRef]
- Olsen, I.; Hicks, S.D. Oral Microbiota and Autism Spectrum Disorder (ASD). J. Oral Microbiol. 2019, 12, 1702806. [Google Scholar] [CrossRef]
- Bars, P.L.; Matamoros, S.; Montassier, E.; Vacon, F.L.; Potel, G.; Soueidan, A.; Jordana, F.; Cochetière, M.F.L. The Oral Cavity Microbiota: Between Health, Oral Disease, and Cancers of the Aerodigestive Tract. Can. J. Microbiol. 2017, 63, 475–492. [Google Scholar] [CrossRef]
- Gao, L.; Xu, T.; Huang, G.; Jiang, S.; Gu, Y.; Chen, F. Oral Microbiomes: More and More Importance in Oral Cavity and Whole Body. Protein Cell. 2018, 9, 488–500. [Google Scholar] [CrossRef]
- Krishnan, K.; Chen, T.; Paster, B.J. A Practical Guide to the Oral Microbiome and Its Relation to Health and Disease. Oral Dis. 2017, 3, 276–286. [Google Scholar] [CrossRef]
- Segata, N.; Haake, S.K.; Mannon, P.; Lemon, K.P.; Waldron, L.; Gevers, D.; Huttenhower, C.; Izard, J. Composition of the adult digestive tract bacterial microbiome based on seven mouth surfaces, tonsils, throat and stool samples. Genome Biol. 2012, 13, R42. [Google Scholar] [CrossRef] [PubMed]
- Griffen, A.L.; Beall, C.J.; Firestone, N.D.; Gross, E.L.; Difranco, J.M.; Hardman, J.H.; Vriesendorp, B.; Faust, R.A.; Janies, D.A.; Leys, E.J. CORE: A phylogenetically-curated 16S rDNA database of the core oral microbiome. PLoS ONE 2011, 6, e19051. [Google Scholar] [CrossRef] [PubMed]
- Zaura, E.; Keijser, B.J.; Huse, S.M.; Crielaard, W. Defining the healthy “core microbiome” of oral microbial communities. BMC Microbiol. 2009, 9, 259. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Quinque, D.; Horz, H.P.; Li, M.; Rzhetskaya, M.; Raff, J.A.; Hayes, M.G.; Stoneking, M. Comparative analysis of the human saliva microbiome from different climate zones: Alaska Germany and Africa. BMC Microbiol. 2014, 14, 316. [Google Scholar] [CrossRef] [PubMed]
- Takeshita, T.; Matsuo, K.; Furuta, M.; Shibata, Y.; Fukami, K.; Shimazaki, Y.; Akifusa, S.; Han, D.H.; Kim, H.D.; Yokoyama, T.; et al. Distinct composition of the oral indigenous microbiota in South Korean and Japanese adults. Sci. Rep. 2014, 4, 6990. [Google Scholar] [CrossRef] [PubMed]
- Myint, Z.C.K.; Zaitsu, T.; Oshiro, A.; Ueno, M.; Soe, K.K.; Kawaguchi, Y. Risk Indicators of Dental Caries and Gingivitis Among 10–11-year-old Students in Yangon, Myanmar. Int. Dent. J. 2019. [Google Scholar] [CrossRef]
- Phyo, A.Z.Z.; Chansatitporn, N.; Narksawat, K. Oral Health Status and Oral Hygiene Habits Among Children Aged 12–13 Years in Yangon, Myanmar. Southeast. Asian J. Trop. Med. Public Health 2013, 44, 1108–1114. [Google Scholar]
- Chu, C.H.; Chau, A.M.; Wong, Z.S.; Hui, B.S.; Lo, E.C. Oral Health Status and Behaviours of Children in Myanmar—A Pilot Study in Four Villages in Rural Areas. Oral Health Prev. Dent. 2012, 10, 365–371. [Google Scholar]
- Nomura, Y.; Maung, K.; Khine, E.M.K.; Sint, K.M.; Lin, M.P.; Myint, M.K.W.; Aung, T.; Sogabe, K.; Otsuka, R.; Okada, A.; et al. Prevalence of Dental Caries in 5- and 6-Year-Old Myanmar Children. Int. J. Dent. 2019, 2019, 5948379. [Google Scholar] [CrossRef] [PubMed]
- Gross, E.L.; Beall, C.J.; Kutsch, S.R.; Firestone, N.D.; Leys, E.J.; Griffen, A.L. Beyond Streptococcus mutans: Dental caries onset linked to multiple species by 16S rRNA community analysis. PLoS ONE 2012, 7, e47722. [Google Scholar] [CrossRef] [PubMed]
- Hughes, T.; Bockmann, M.; Townsend, G.; Salehi, H.; Adler, C.J. Preliminary study of the oral mycobiome of children with and without dental caries. J. Oral Microbiol. 2018, 11, 1536182. [Google Scholar] [CrossRef]
- Hurley, E.; Barrett, M.P.J.; Kinirons, M.; Whelton, H.; Ryan, C.A.; Stanton, C.; Harris, H.M.B.; O’Toole, P.W. Comparison of the salivary and dentinal microbiome of children with severe-early childhood caries to the salivary microbiome of caries-free children. BMC Oral Health 2019, 19, 13. [Google Scholar] [CrossRef] [PubMed]
- World Health Organization. Oral Health Surveys: Basic Methods, 5th ed.; WHO: Geneva, Switzerland, 2013. [Google Scholar]
- Okada, M.; Hayashi, F.; Nagasaka, N. PCR detection of 5 Putative periodontal pathogens in dental plaque samples from children 2 to 12 years of age. J. Clin. Periodontol. 2001, 28, 576–582. [Google Scholar] [CrossRef] [PubMed]
- Okada, A.; Sogabe, K.; Takeuchi, H.; Okamoto, M.; Nomura, Y.; Hanada, N. Characterization of specimens obtained by different sampling methods for evaluation of periodontal bacteria. J. Oral Sci. 2017, 59, 491–498. [Google Scholar] [CrossRef] [PubMed]
- Kim, O.S.; Cho, Y.J.; Lee, K.; Yoon, S.H.; Kim, M.; Na, H.; Park, S.C.; Jeon, Y.S.; Lee, J.H.; Yi, H.; et al. Introducing EzTaxon-e: A prokaryotic 16S rRNA gene sequence database with phylotypes that represent uncultured species. Int. J. Syst. Evol. Microbiol. 2012, 62, 716–721. [Google Scholar] [CrossRef] [PubMed]
- Yoon, S.H.; Ha, S.M.; Kwon, S.; Lim, J.; Kim, Y.; Seo, H.; Chun, J. Introducing EzBioCloud: A taxonomically united database of 16S rRNA and whole genome assemblies. Int. J. Syst. Evol. Microbiol. 2017, 67, 1613–1617. [Google Scholar] [CrossRef] [PubMed]
- Verma, D.; Garg, P.K.; Dubey, A.K. Insights into the human oral microbiome. Arch. Microbiol. 2018, 200, 525–540. [Google Scholar] [CrossRef]
- Bik, E.M.; Long, C.D.; Armitage, G.C.; Loomer, P.; Emerson, J.; Mongodin, E.F.; Nelson, K.E.; Gill, S.R.; Fraser-Liggett, C.M.; Relman, D.A. Bacterial diversity in the oral cavity of 10 healthy individuals. ISME J. 2010, 4, 962–974. [Google Scholar] [CrossRef]
- Palmer, R.J. Composition and development of oral bacterial com-munities. Periodontol. 2000 2014, 64, 20–39. [Google Scholar] [CrossRef] [PubMed]
- Dewhirst, F.E.; Chen, T.; Izard, J.; Paster, B.J.; Tanner, A.C.; Yu, W.H.; Lakshmanan, A.; Wade, W.G. The human oral microbiome. J. Bacteriol. 2010, 192, 5002–5017. [Google Scholar] [CrossRef] [PubMed]
- Mager, D.L.; Haffajee, A.D.; Devlin, P.M.; Norris, C.M.; Posner, M.R.; Goodson, J.M. The salivary microbiota as a diagnostic indicator of oral cancer: A descriptive non-randomized study of cancer-free and oral squamous cell carcinoma subjects. J. Transl. Med. 2005, 3, 27. [Google Scholar] [CrossRef] [PubMed]
- Shi, W.; Tian, J.; Xu, H.; Zhou, Q.; Qin, M. Distinctions and associations between the microbiota of saliva and supragingival plaque of permanent and deciduous teeth. PLoS ONE 2018, 13, e0200337. [Google Scholar] [CrossRef]
- Kersters, K.; Lisdiyanti, P.; Kmagata, K.; Swings, J. The Family Acetobacteracea: The Genera Acetobacter; Acidomonas; Asaia; Gluconacetobacter; Gluconobacter; and Kozakia. In The Prokaryotes; Dworkin, M., Falkow, S., Rosenberg, E., Eds.; Springer: New York, NY, USA, 2006; Volume 5, pp. 163–200. [Google Scholar]
- Sampaio-Maia, B.; Monteiro-Silva, F. Acquisition and maturation of oral microbiome throughout childhood: An update. Dent. Res. J. (Isfahan) 2014, 11, 291–301. [Google Scholar]
- Cephas, K.D.; Kim, J.; Mathai, R.A.; Barry, K.A.; Dowd, S.E.; Meline, B.S.; Swanson, K.S. Comparative analysis of salivary bacterial microbiome diversity in edentulous infants and their mothers or primary care givers using pyrosequencing. PLoS ONE 2011, 6, e23503. [Google Scholar] [CrossRef]
- Shi, W.; Qin, M.; Chen, F.; Xia, B. Supragingival Microbial Profiles of Permanent and Deciduous Teeth in Children with Mixed Dentition. PLoS ONE 2016, 11, e0146938. [Google Scholar] [CrossRef] [PubMed]
- Lassalle, F.; Spagnoletti, M.; Fumagalli, M.; Shaw, L.; Dyble, M.; Walker, C.; Thomas, M.G.; Migliano, A.B.; Balloux, F. Oral microbiomes from hunter-gatherers and traditional farmers reveal shifts in commensal balance and pathogen load linked to diet. Mol. Ecol. 2018, 27, 182–195. [Google Scholar] [CrossRef] [PubMed]
- Xu, X.; He, J.; Xue, J.; Wang, Y.; Li, K.; Zhang, K.; Guo, Q.; Liu, X.; Zhou, Y.; Cheng, L.; et al. Oral cavity contains distinct niches with dynamic microbial communities. Environ. Microbiol. 2015, 17, 699–710. [Google Scholar] [CrossRef] [PubMed]
- Crielaard, W.; Zaura, E.; Schuller, A.A.; Huse, S.M.; Montijn, R.C.; Keijser, B.J. Exploring the oral microbiota of children at various developmental stages of their dentition in the relation to their oral health. BMC Med. Genom. 2011, 4, 22. [Google Scholar] [CrossRef]
- Štšepetova, J.; Truu, J.; Runnel, R.; Nõmmela, R.; Saag, M.; Olak, J.; Nõlvak, H.; Preem, J.K.; Oopkaup, K.; Krjutškov, K.; et al. Impact of polyols on Oral microbiome of Estonian schoolchildren. BMC Oral Health. 2019, 19, 60. [Google Scholar] [CrossRef]
- Xu, Y.; Jia, Y.H.; Chen, L.; Huang, W.M.; Yang, D.Q. Metagenomic analysis of oral microbiome in young children aged 6–8 years living in a rural isolated Chinese province. Oral Dis. 2018, 24, 1115–1125. [Google Scholar] [CrossRef]
- Gao, X.; Jiang, S.; Koh, D.; Hsu, C.Y. Salivary biomarkers for dental caries. Periodontol. 2000 2016, 70, 128–141. [Google Scholar] [CrossRef] [PubMed]
- Pagano, S.; Rabbit, M.; Valenti, C.; Negri, P.; Lombardo, G.; Costanzi, E.; Cianetti, S.; Montaseri, A.; Marinucci, L. Biological effects of resin monomers on oral cell populations: Descriptive analysis of literature. Eur. J. Paediatr. Dent. 2019, 20, 224–232. [Google Scholar] [CrossRef] [PubMed]
- Bromo, F.; Guida, A.; Santoro, G.; Peciarolo, M.R.; Eramo, S. Pit and Fissure Sealants: Review of Literature and Application Technique. Minerva Stomatol. 2011, 60, 529–541. [Google Scholar]
- Bunick, F.J.; Kashket, S. Enolases from fluoride-sensitive and fluoride-resistant streptococci. Infect. Immun. 1981, 34, 856–863. [Google Scholar] [CrossRef] [PubMed]
- Yamada, T.; Takahashi-Abbe, S.; Abbe, K. Effects of oxygen on pyruvate formate-lyase in situ and sugar metabolism of Streptococcus mutans and Streptococcus sanguis. Infect. Immun. 1985, 47, 129–134. [Google Scholar] [CrossRef]
- Nyvad, B.; Kilian, M. Microbiology of the early colonization of human enamel and root surfaces in vivo. Eur. J. Oral Sci. 1987, 95, 369–380. [Google Scholar] [CrossRef]
- Sharma, N.; Bhatia, S.; Sodhi, A.S.; Batra, N. Oral microbiome and health. AIMS Microbiol. 2018, 4, 42–66. [Google Scholar] [CrossRef]
- Zachary, D.; Moye, L.Z.; Robert, A.B. Fueling the caries process: Carbohydrate metabolism and gene regulation by Streptococcus mutans. J. Oral Microbiol. 2014, 6, 24878. [Google Scholar] [CrossRef]

| Taxon Name | Abundance (%) |
|---|---|
| Veillonella parvula group | 5.22% (0.48–21.92%) |
| Neisseria sicca group | 4.72% (0.09–31.89%) |
| Streptococcus pneumoniae group | 4.40% (0.37–8.80%) |
| Haemophilus parainfluenzae group | 3.60% (0.21–8.34%) |
| Lautropia mirabilis | 2.99% (0.27–9.79%) |
| Streptococcus sanguinis group | 1.89% (0.18–4.00%) |
| Veillonella dispar | 1.78% (0.03–7.47%) |
| Streptococcus parasanguinis group | 1.24% (0.15–2.89%) |
| Granulicatella adiacens group | 0.99% (0.05–3.57%) |
| Aggregatibacter aphrophilus | 0.92% (0.02–5.49%) |
| Fusobacterium nucleatum group | 0.74% (0.07–1.55%) |
| Veillonella rogosae | 0.72% (0.02–3.17%) |
| Porphyromonas pasteri | 0.72% (0.01–2.35%) |
| Streptococcus peroris group | 0.61% (0.03–1.30%) |
| Gemella morbillorum | 0.57% (0.02–2.05%) |
| Leptotrichia buccalis group | 0.54% (0.02–1.97%) |
| Aggregatibacter segnis | 0.54% (0.01–1.63%) |
| Capnocytophaga granulosa | 0.47% (0.04–1.49%) |
| Streptococcus gordonii group | 0.46% (0.04–2.15%) |
| Prevotella loescheii | 0.46% (0.03–2.12%) |
| Abiotrophia defectiva | 0.41% (0.01–1.40%) |
| Capnocytophaga sputigena | 0.39% (0.02–1.27%) |
| KI259256_s | 0.36% (0.02–1.90%) |
| Streptococcus_uc | 0.36% (0.01–0.90%) |
| Streptococcus sinensis group | 0.34% (0.03–1.07%) |
| ADCM_s | 0.31% (0.02–0.54%) |
| JQ463704_s | 0.28% (0.02–1.14%) |
| Gemella haemolysans group | 0.25% (0.02–0.90%) |
| Veillonella_uc | 0.24% (0.01–0.90%) |
| CP017038_s | 0.22% (0.03–0.53%) |
| Campylobacter gracilis | 0.21% (0.02–0.65%) |
| Cardiobacterium hominis | 0.19% (0.02–0.40%) |
| JF239777_s | 0.18% (0–0.98%) |
| Unclassified in a higher taxonomic rank | 0.16% (0.03–0.45%) |
| Streptococcus anginosus group | 0.16% (0.02–0.65%) |
| Prevotella oris | 0.12% (0.01–0.50%) |
| Actinomyces odontolyticus | 0.11% (0.01–0.47%) |
| Corynebacterium matruchotii | 0.09% (0.01–0.51%) |
| Dialister invisus | 0.09% (0.01–0.27%) |
| Actinomyces oris | 0.08% (<0.01–0.29%) |
| Granulicatella elegans | 0.07% (<0.01–0.23%) |
| Campylobacter concisus group | 0.07% (0.02–0.16%) |
| Actinomyces naeslundii | 0.06% (<0.01–0.21%) |
| Prevotella maculosa | 0.06% (<0.01–0.13%) |
| Dental Caries | p-Value | ||||
|---|---|---|---|---|---|
| − | + | ||||
| OTUs | Mean ± SD | Median (25th–75th%) | Mean ± SD | Median (25th–75th%) | |
| Faucicola mancuniensis | 0.4136 ± 1.0826 | 0.0043 (0–0.0181) | - | 0.036 | |
| Johnsonella ignava | 0.0019 ± 0.0051 | 0 (0–0) | 0.0082 ± 0.00686 | 0.0029 (0.0029–0.0156) | 0.030 |
| JQ406535 s | 0.0321 ± 0.0435 | 0.0136 (0–0.048) | - | 0.015 | |
| KI273068 s | 0.0043 ± 0.00539 | 0.0019 (0–0.0104) | 0.042 | ||
| Kingella oralis | 0.0037 ± 0.0063 | 0 (0–0.0045) | 0.0360 ± 0.0276 | 0.0319 (0.0095–0.0627) | 0.006 |
| Lactobacillus mucosae | - | 0.0563 ± 0.12536 | 0.0049 (0–0.0900) | 0.042 | |
| Neisseria bacilliformis | - | 0.0083 ± 0.01179 | 0.0036 (0–0.0168) | 0.042 | |
| Parascardovia denticolens | - | 0.0217 ± 0.04818 | 0.0025 (0–0.0340) | 0.042 | |
| Prevotella multisaccharivorax | - | 0.0535 ± 0.09038 | 0.0208 (0–0.0927) | 0.015 | |
| Pseudomonas uc | - | 0.4038 ± 0.87311 | 0.0255 (0–0.6889) | 0.015 | |
| Rothia mucilaginosa group | 0.0469 ± 0.0660 | 0.0175 (0.0086–0.0815) | 0.0038 ± 0.00585 | 0 (0–0.011) | 0.030 |
| Staphylococcus sciuri group | - | 0.0039 ± 0.00459 | 0.0025 (0–0.0084) | 0.042 | |
| Streptococcus mutans | 0.0062 ± 0.0050 | 0.0086 (0–0.0099) | 0.0960 ± 0.1302 | 0.0670 (0.0093–0.1455) | 0.031 |
| Veillonella dispar | 0.4027 ± 0.6048 | 0.1981 (0.0481–0.5183) | 3.3924 ± 2.6177 | 2.9546 (0.9421–5.8599) | 0.007 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Nomura, Y.; Otsuka, R.; Hasegawa, R.; Hanada, N. Oral Microbiome of Children Living in an Isolated Area in Myanmar. Int. J. Environ. Res. Public Health 2020, 17, 4033. https://doi.org/10.3390/ijerph17114033
Nomura Y, Otsuka R, Hasegawa R, Hanada N. Oral Microbiome of Children Living in an Isolated Area in Myanmar. International Journal of Environmental Research and Public Health. 2020; 17(11):4033. https://doi.org/10.3390/ijerph17114033
Chicago/Turabian StyleNomura, Yoshiaki, Ryoko Otsuka, Ryo Hasegawa, and Nobuhiro Hanada. 2020. "Oral Microbiome of Children Living in an Isolated Area in Myanmar" International Journal of Environmental Research and Public Health 17, no. 11: 4033. https://doi.org/10.3390/ijerph17114033
APA StyleNomura, Y., Otsuka, R., Hasegawa, R., & Hanada, N. (2020). Oral Microbiome of Children Living in an Isolated Area in Myanmar. International Journal of Environmental Research and Public Health, 17(11), 4033. https://doi.org/10.3390/ijerph17114033






