Characteristics of Bacterial Community and Function in Paddy Soil Profile around Antimony Mine and Its Response to Antimony and Arsenic Contamination
Abstract
1. Introduction
2. Material and Methods
2.1. Site Description
2.2. Geochemical Properties Analysis
2.3. Antimony and Arsenic Contamination Fractions Analysis
2.4. DNA Extraction and PCR Amplification
2.5. Data Analysis
3. Results
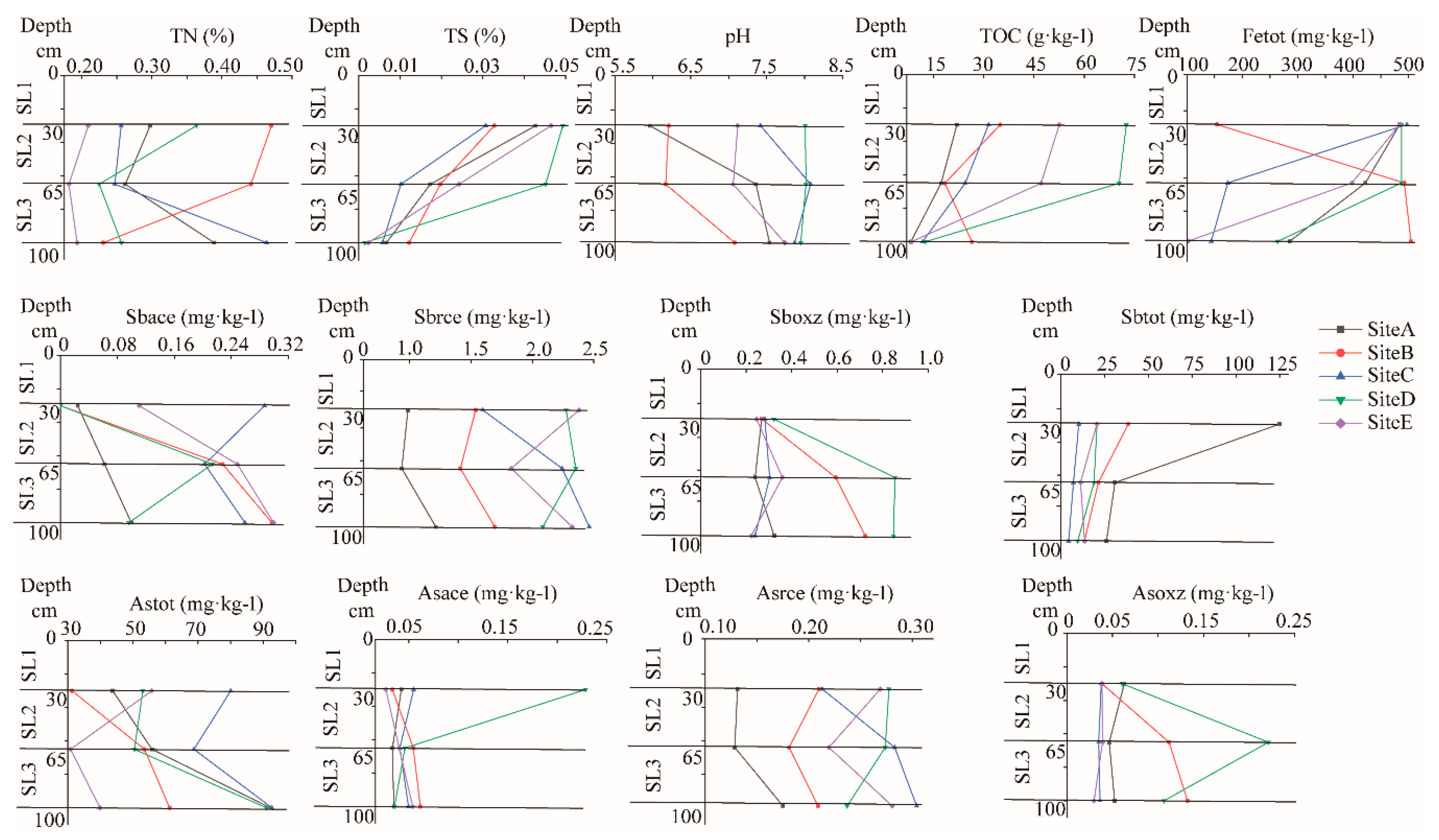
3.1. Geochemical Properties
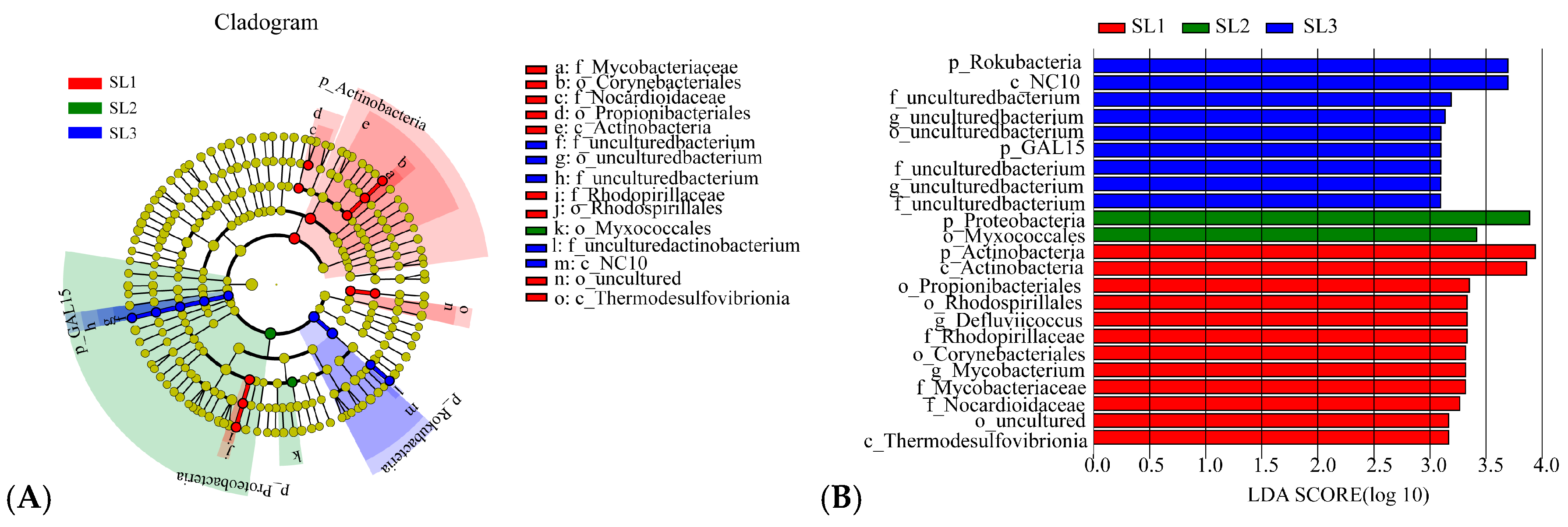
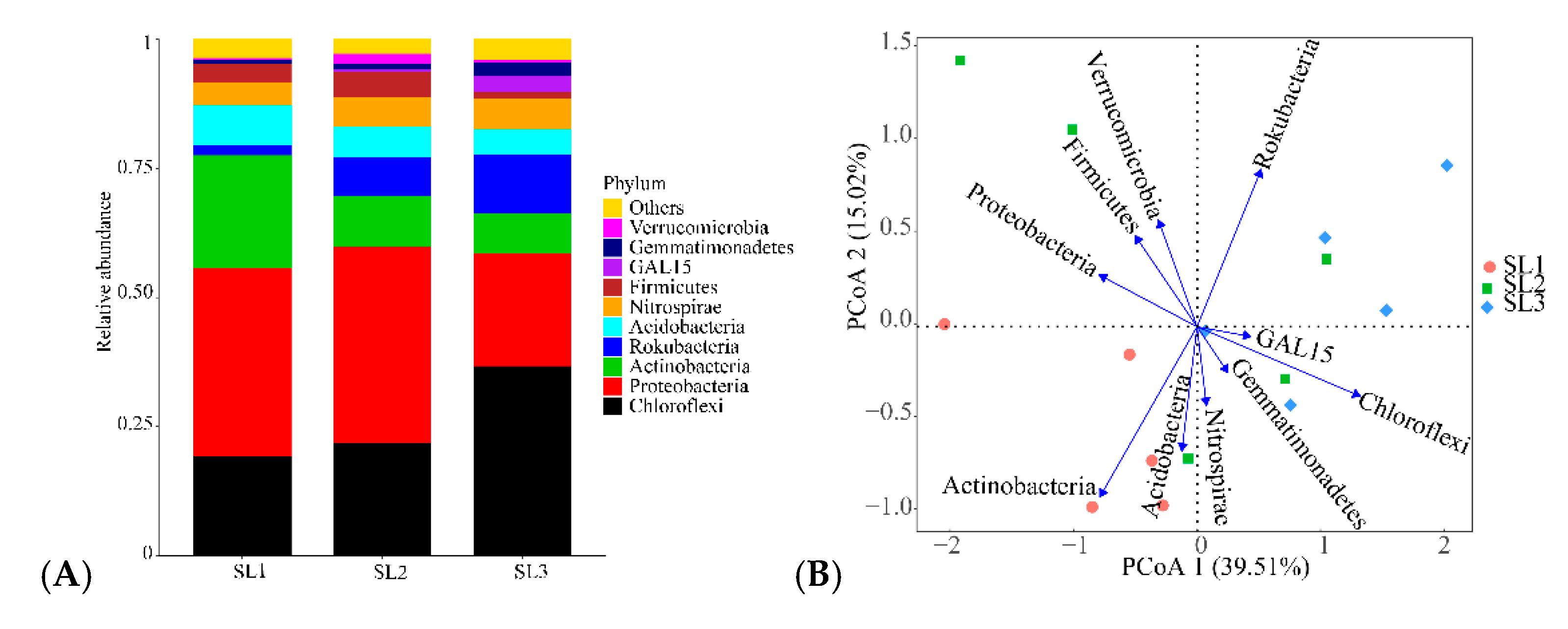
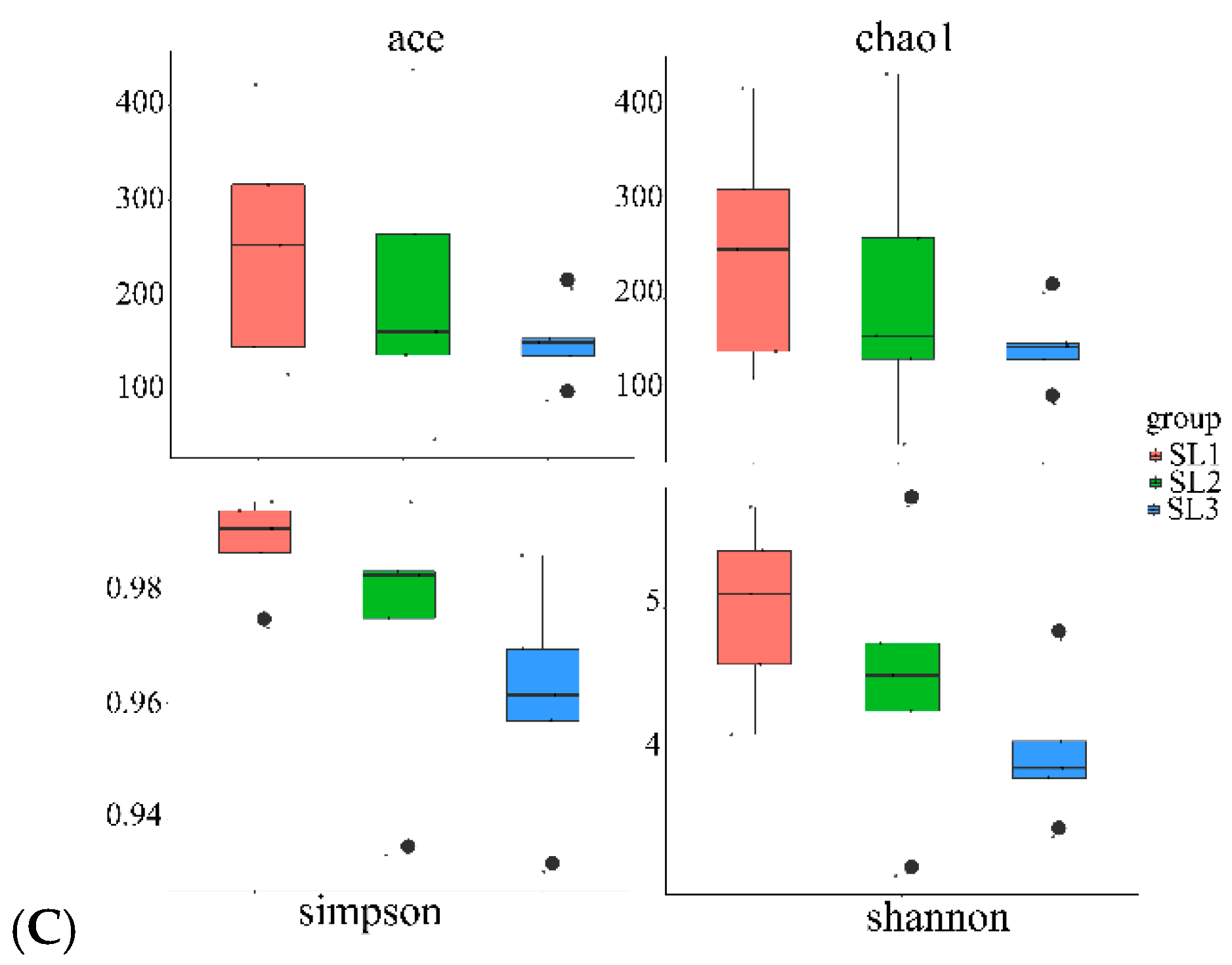
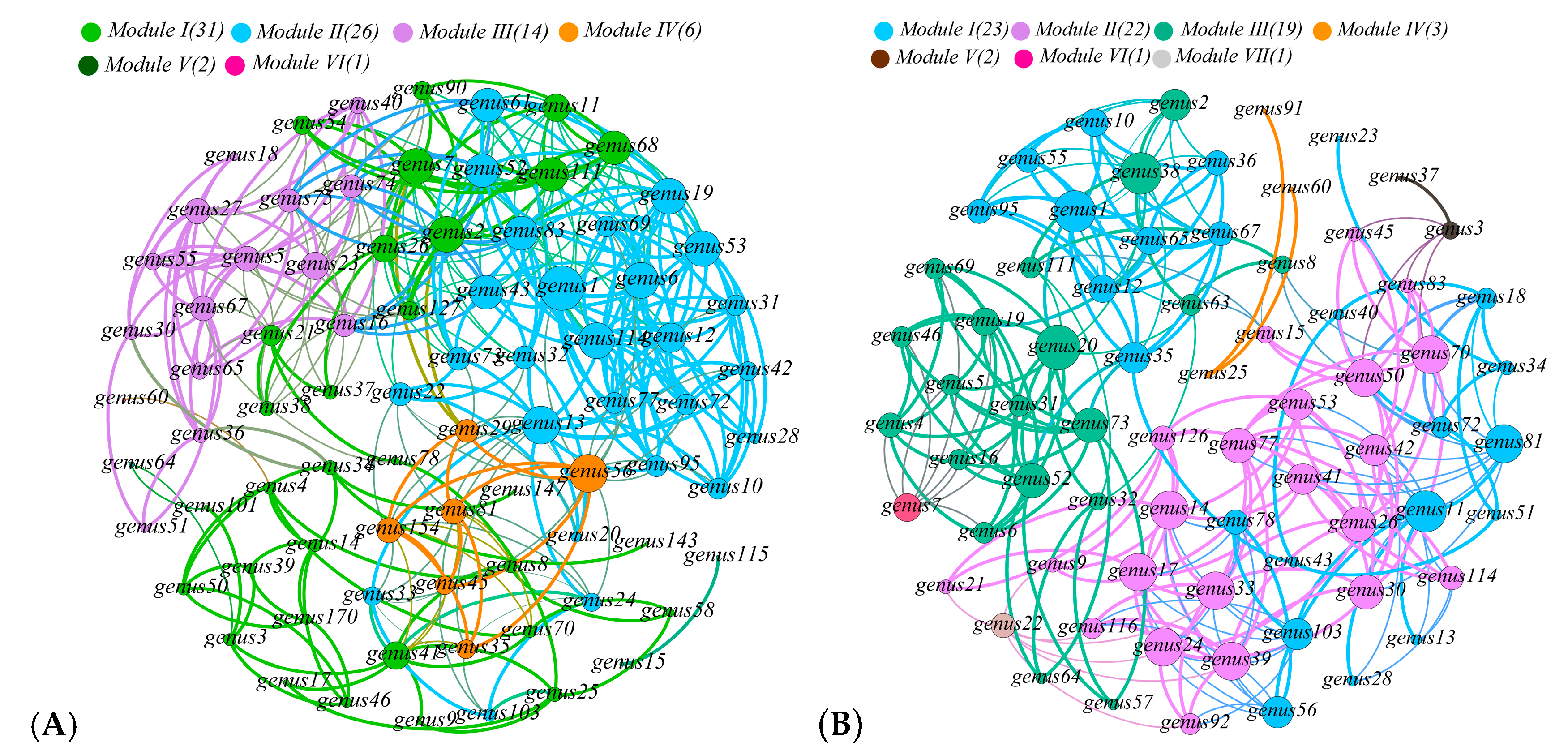
3.2. Bacterial Community Diversity and Structure Analysis
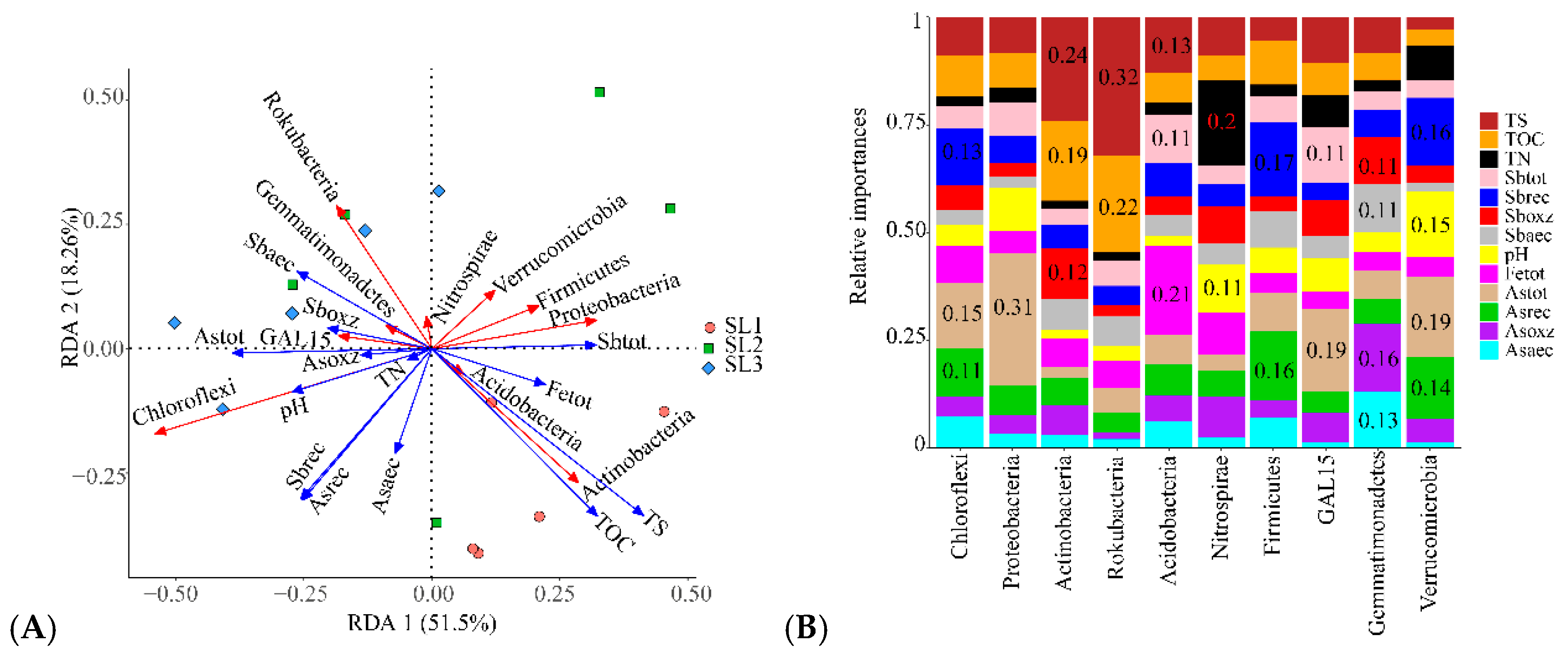
3.3. The Relationship between Geochemical Properties and Bacterial Communities
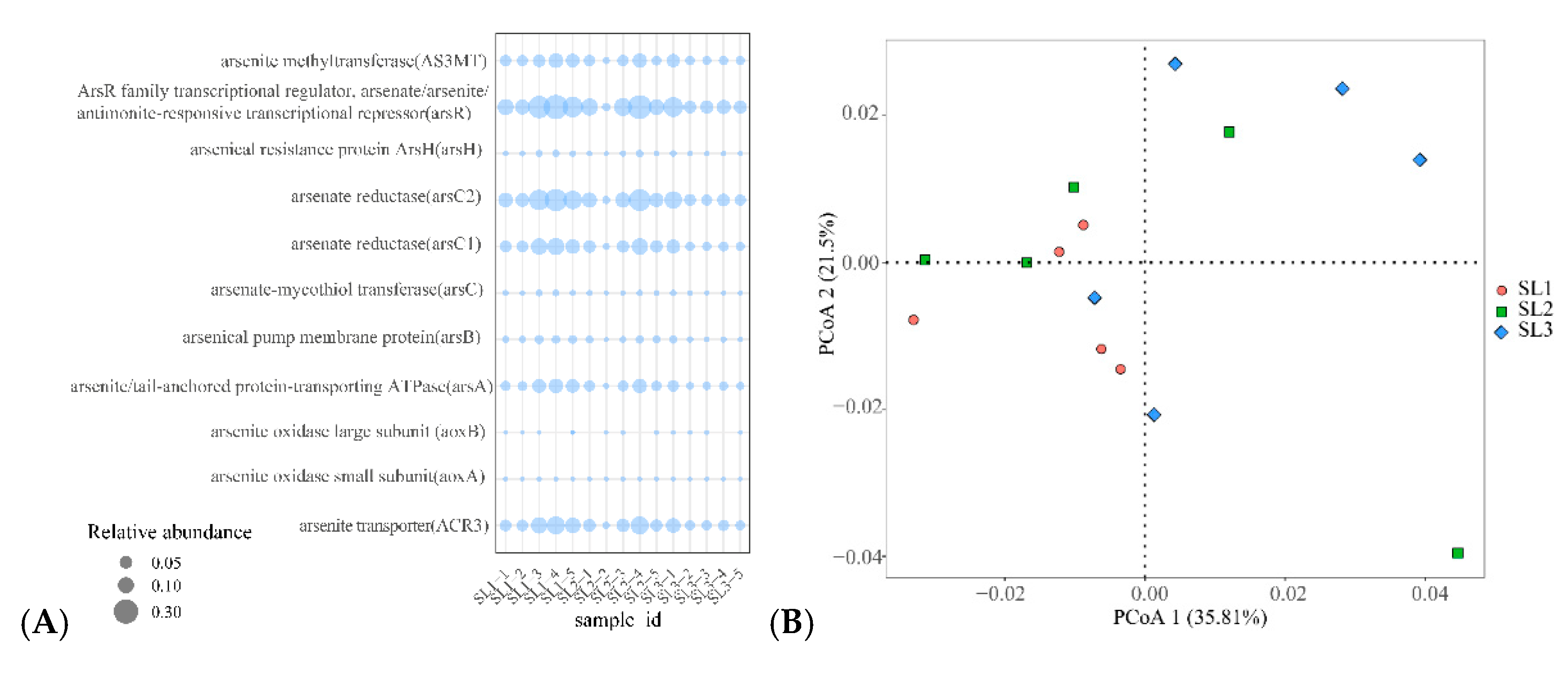
3.4. The Relationship between Geochemical Properties and Bacterial Function
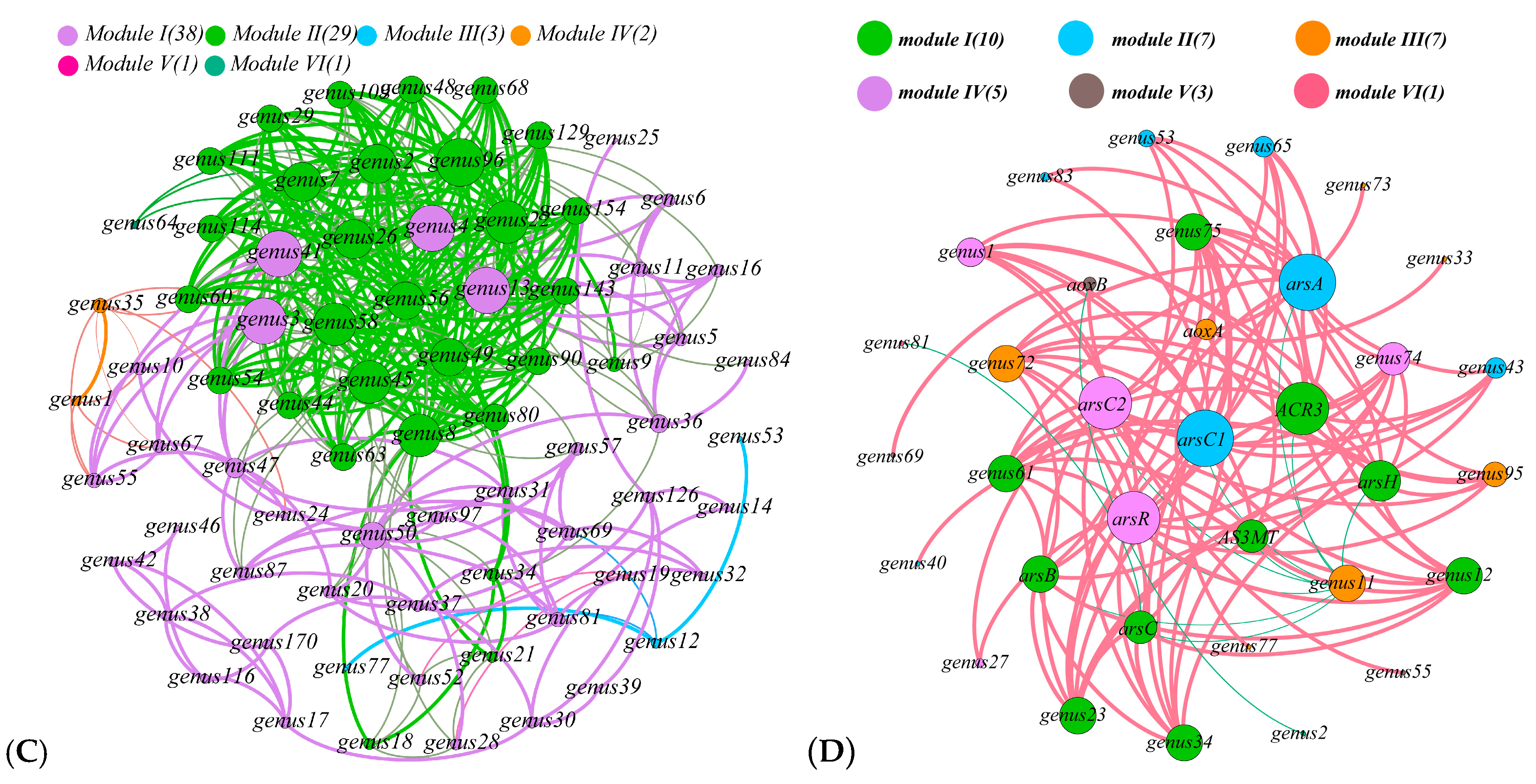
3.5. Relationship with Community Structure and Bacterial Function
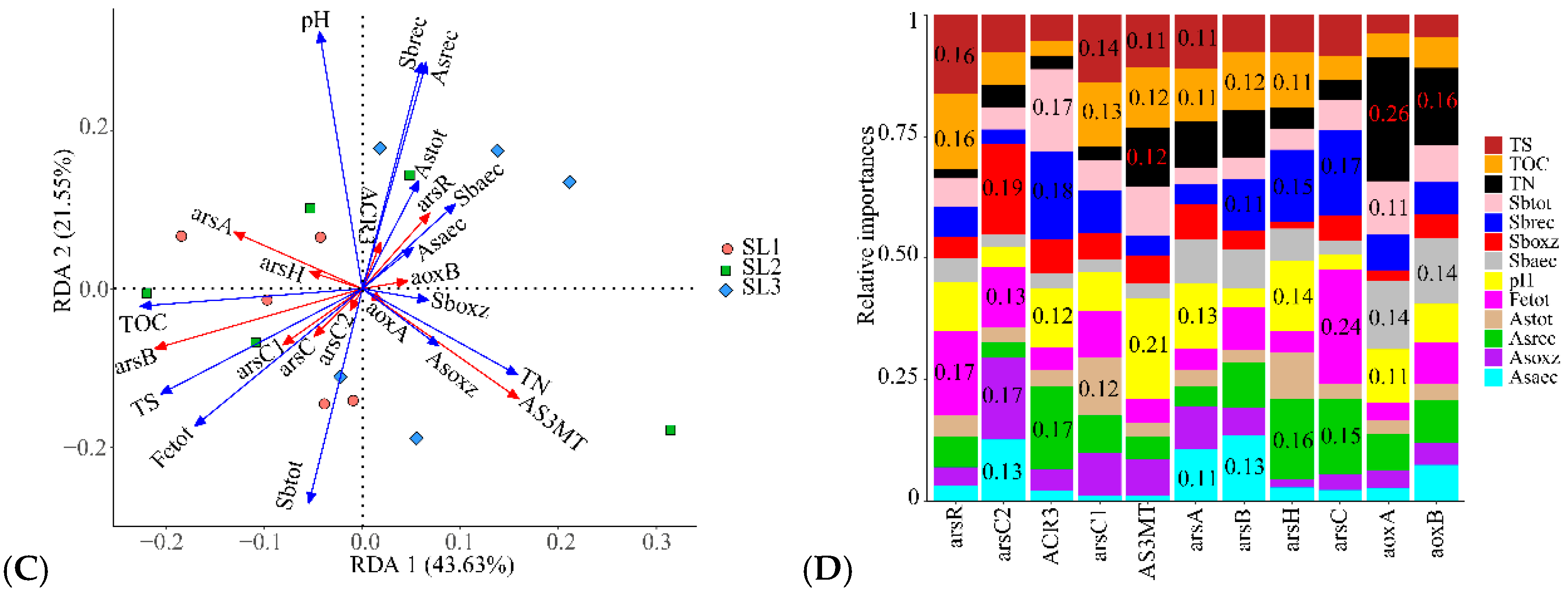
3.6. Determining the Relationship between Geochemical Properties, Bacterial Community, and Function
4. Discussion
4.1. Distribution Characteristics of Geochemical Properties in Paddy Soil Profiles
4.2. Effects of Sb and As Contamination Fractions on Bacterial Community
4.3. Effects of Bacterial Community on Sb and As Related Function
4.4. Effects of Geochemical Properties on Sb and As related Bacterial Function
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Fu, Z.; Wu, F.; Mo, C.; Deng, Q.; Meng, W.; Giesy, J.P. Comparison of arsenic and antimony biogeochemical behavior in water, soil and tailings from Xikuangshan, China. Sci. Total Environ. 2016, 539, 97–104. [Google Scholar] [CrossRef] [PubMed]
- Wang, S. Present situation and sustainable development of antimony resources in China. China Met. Bull. 2002, 47, 6–10. [Google Scholar]
- Sun, W.; Xiao, E.; Dong, Y.; Tang, S.; Krumins, V.; Ning, Z.; Sun, M.; Zhao, Y.; Wu, S.; Xiao, T. Profiling microbial community in a watershed heavily contaminated by an active antimony (Sb) mine in Southwest China. Sci. Total Environ. 2016, 550, 297–308. [Google Scholar] [CrossRef] [PubMed]
- Ning, Z.; Xiao, T.; Xiao, E. Antimony in the Soil-Plant System in an Sb Mining/Smelting Area of Southwest China. Int. J. Phytoremediat. 2015, 17, 1081–1089. [Google Scholar] [CrossRef] [PubMed]
- Xiao, E.; Krumins, V.; Dong, Y.; Xiao, T.; Ning, Z.; Xiao, Q.; Sun, W. Microbial diversity and community structure in an antimony-rich tailings dump. Appl. Microbiol. Biotechnol. 2016, 100, 7751–7763. [Google Scholar] [CrossRef] [PubMed]
- Li, L.; Tu, H.; Zhang, S.; Wu, L.; Wu, M.; Tang, Y.; Wu, P. Geochemical behaviors of antimony in mining-affected water environment (Southwest China). Env. Geochem. Health 2019, 41, 2397–2411. [Google Scholar] [CrossRef] [PubMed]
- Mitsunobu, S.; Harada, T.; Takahashi, Y. Comparison of antimony behavior with that of arsenic under various soil redox conditions. Environ. Sci. Technol. 2006, 40, 7270–7276. [Google Scholar] [CrossRef]
- Majzlan, J.; Lalinská, B.; Chovan, M.; Bläß, U.; Brecht, B.; Göttlicher, J.; Steininger, R.; Hug, K.; Ziegler, S.; Gescher, J. A mineralogical, geochemical, and microbiogical assessment of the antimony-and arsenic-rich neutral mine drainage tailings near Pezinok, Slovakia. Am. Mineral. 2011, 96, 1–13. [Google Scholar] [CrossRef]
- Zhang, S.Y.; Zhao, F.J.; Sun, G.X.; Su, J.Q.; Yang, X.R.; Li, H.; Zhu, Y.G. Diversity and abundance of arsenic biotransformation genes in paddy soils from southern China. Environ. Sci. Technol. 2015, 49, 4138–4146. [Google Scholar] [CrossRef]
- Cai, L.; Yu, K.; Yang, Y.; Chen, B.W.; Li, X.D.; Zhang, T. Metagenomic exploration reveals high levels of microbial arsenic metabolism genes in activated sludge and coastal sediments. Appl. Microbiol. Biotechnol. 2013, 97, 9579–9588. [Google Scholar] [CrossRef]
- Ma, L.; Wang, L.; Jia, Y.; Yang, Z. Arsenic speciation in locally grown rice grains from Hunan Province, China: Spatial distribution and potential health risk. Sci. Total Environ. 2016, 557–558, 438–444. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Zeng, X.C.; Wang, J.; Deng, Y.; Ma, T.; Guoji, E.; Mu, Y.; Yang, Y.; Li, H.; Wang, Y. Microbial communities involved in arsenic mobilization and release from the deep sediments into groundwater in Jianghan plain, Central China. Sci. Total Environ. 2017, 579, 989–999. [Google Scholar] [CrossRef] [PubMed]
- Wang, N.; Zhang, S.; He, M. Bacterial community profile of contaminated soils in a typical antimony mining site. Environ. Sci. Pollut. Res. Int. 2018, 25, 141–152. [Google Scholar] [CrossRef] [PubMed]
- Xiao, E.; Krumins, V.; Tang, S.; Xiao, T.; Ning, Z.; Lan, X.; Sun, W. Correlating microbial community profiles with geochemical conditions in a watershed heavily contaminated by an antimony tailing pond. Environ. Pollut. 2016, 215, 141–153. [Google Scholar] [CrossRef] [PubMed]
- Sun, W.; Xiao, E.; Xiao, T.; Krumins, V.; Wang, Q.; Haggblom, M.; Dong, Y.; Tang, S.; Hu, M.; Li, B.; et al. Response of Soil Microbial Communities to Elevated Antimony and Arsenic Contamination Indicates the Relationship between the Innate Microbiota and Contaminant Fractions. Environ. Sci. Technol. 2017, 51, 9165–9175. [Google Scholar] [CrossRef]
- Hamamura, N.; Fukushima, K.; Itai, T. Identification of antimony- and arsenic-oxidizing bacteria associated with antimony mine tailing. Microbes Environ. 2013, 28, 257–263. [Google Scholar] [CrossRef]
- Dovick, M.A.; Kulp, T.R.; Arkle, R.S.; Pilliod, D.S. Bioaccumulation trends of arsenic and antimony in a freshwater ecosystem affected by mine drainage. Environ. Chem. 2016, 13, 149–159. [Google Scholar] [CrossRef]
- Majzlan, J.; Števko, M.; Lánczos, T. Soluble secondary minerals of antimony in Pezinok and Kremnica (Slovakia) and the question of mobility or immobility of antimony in mine waters. Environ. Chem. 2016, 13, 927–935. [Google Scholar] [CrossRef]
- Zhang, S.; Rensing, C.; Zhu, Y.G. Cyanobacteria-mediated arsenic redox dynamics is regulated by phosphate in aquatic environments. Environ. Sci. Technol. 2014, 48, 994–1000. [Google Scholar] [CrossRef]
- Ye, J.; Rensing, C.; Rosen, B.P.; Zhu, Y.G. Arsenic biomethylation by photosynthetic organisms. Trends Plant Sci. 2012, 17, 155–162. [Google Scholar] [CrossRef]
- Shi, X.Z.; Yu, D.S.; Warner, E.D.; Sun, W.X.; Petersen, G.W.; Gong, Z.T.; Lin, H. Cross-Reference System for Translating Between Genetic Soil Classification of China and Soil Taxonomy. Soil Sci. Soc. Am. J. 2006, 70, 78–83. [Google Scholar] [CrossRef]
- Liu, C.; Yu, H.-Y.; Liu, C.; Li, F.; Xu, X.; Wang, Q. Arsenic availability in rice from a mining area: Is amorphous iron oxide-bound arsenic a source or sink? Environ. Pollut. 2015, 199, 95–101. [Google Scholar] [CrossRef]
- Tamura, H.; Goto, K.; Yotsuyanagi, T.; Nagayama, M. Spectrophotometric determination of iron(II) with 1,10-phenanthroline in the presence of large amounts of iron(III). Talanta 1974, 21, 314–318. [Google Scholar] [CrossRef]
- Pascaud, G.; Leveque, T.; Soubrand, M.; Boussen, S.; Joussein, E.; Dumat, C. Environmental and health risk assessment of Pb, Zn, As and Sb in soccer field soils and sediments from mine tailings: Solid speciation and bioaccessibility. Environ. Sci. Pollut. Res. Int. 2014, 21, 4254–4264. [Google Scholar] [CrossRef] [PubMed]
- Guo, X.; Wang, K.; He, M.; Liu, Z.; Yang, H.; Li, S. Antimony smelting process generating solid wastes and dust: Characterization and leaching behaviors. J. Environ. Sci. 2014, 26, 1549–1556. [Google Scholar] [CrossRef] [PubMed]
- Bolyen, E.; Rideout, J.R.; Dillon, M.R.; Bokulich, N.A.; Abnet, C.C.; Al-Ghalith, G.A.; Alexander, H.; Alm, E.J.; Arumugam, M.; Asnicar, F.; et al. Author Correction: Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat. Biotechnol. 2019, 37, 1091. [Google Scholar] [CrossRef]
- Callahan, B.J.; McMurdie, P.J.; Rosen, M.J.; Han, A.W.; Johnson, A.J.; Holmes, S.P. DADA2: High-resolution sample inference from Illumina amplicon data. Nat. Methods 2016, 13, 581–583. [Google Scholar] [CrossRef]
- Paradis, E.; Claude, J.; Strimmer, K. APE: Analyses of Phylogenetics and Evolution in R language. Bioinformatics 2004, 20, 289–290. [Google Scholar] [CrossRef]
- Oksanen, J.; Kindt, R.; Legendre, P.; O’Hara, B.; Stevens, M.H.H.; Oksanen, M.J.; Suggests, M. The vegan package. Community Ecol. Package 2007, 10, 631–637. [Google Scholar]
- Revelle, W.R. psych: Procedures for Personality and Psychological Research; Northwestern University: Evanston, IL, USA, 2017. [Google Scholar]
- Wickham, H. reshape2: Flexibly reshape data: A reboot of the reshape package. R Package Version 2012, 1, 2. [Google Scholar]
- Grömping, U. Relative importance for linear regression in R: The package relaimpo. J. Stat. Softw. 2006, 17, 1–27. [Google Scholar] [CrossRef]
- Csardi, G.; Nepusz, T. The igraph software package for complex network research. Int. J. Complex Syst. 2006, 1695, 1–9. [Google Scholar]
- Sanchez, G. PLS Path Modeling with R; Trowchez Editions: Berkeley, CA, USA, 2013; Volume 383. [Google Scholar]
- Wickham, H. ggplot2: Elegant Graphics for Data Analysis; Springer: Berlin/Heidelberg, Germany, 2016. [Google Scholar]
- Wemheuer, F.; Taylor, J.A.; Daniel, R.; Johnston, E.; Meinicke, P.; Thomas, T.; Wemheuer, B. Tax4Fun2: A R-based tool for the rapid prediction of habitat-specific functional profiles and functional redundancy based on 16S rRNA gene marker gene sequences. bioRxiv 2018, 490037. [Google Scholar] [CrossRef]
- Jurburg, S.D.; Salles, J.F. Functional redundancy and ecosystem function—The soil microbiota as a case study. In Biodiversity in Ecosystems: Linking Structure and Function; Intech: Rijeka, Croatia, 2015; pp. 29–49. [Google Scholar]
- Hammel, W.; Debus, R.; Steubing, L. Mobility of antimony in soil and its availability to plants. Chemosphere 2000, 41, 1791–1798. [Google Scholar] [CrossRef]
- He, M.; Wang, X.; Wu, F.; Fu, Z. Antimony pollution in China. Sci. Total Environ. 2012, 421–422, 41–50. [Google Scholar] [CrossRef] [PubMed]
- Hiller, E.; Lalinská, B.; Chovan, M.; Jurkovič, Ľ.; Klimko, T.; Jankulár, M.; Hovorič, R.; Šottník, P.; Fľaková, R.; Ženišová, Z. Arsenic and antimony contamination of waters, stream sediments and soils in the vicinity of abandoned antimony mines in the Western Carpathians, Slovakia. Appl. Geochem. 2012, 27, 598–614. [Google Scholar] [CrossRef]
- Buanuam, J.; Wennrich, R. Dynamic flow-through sequential extraction for assessment of fractional transformation and inter-element associations of arsenic in stabilized soil and sludge. J. Hazard. Mater. 2010, 184, 849–854. [Google Scholar] [CrossRef]
- Wang, X.; He, M.; Xi, J.; Lu, X. Antimony distribution and mobility in rivers around the world’s largest antimony mine of Xikuangshan, Hunan Province, China. Microchem. J. 2011, 97, 4–11. [Google Scholar] [CrossRef]
- Kim, E.J.; Yoo, J.C.; Baek, K. Arsenic speciation and bioaccessibility in arsenic-contaminated soils: Sequential extraction and mineralogical investigation. Environ. Pollut. 2014, 186, 29–35. [Google Scholar] [CrossRef]
- Wenzel, W.W.; Kirchbaumer, N.; Prohaska, T.; Stingeder, G.; Lombi, E.; Adriano, D.C. Arsenic fractionation in soils using an improved sequential extraction procedure. Anal. Chim. Acta 2001, 436, 309–323. [Google Scholar] [CrossRef]
- Yi, Y.; Wen, J.; Zeng, G.; Zhang, T.; Huang, F.; Qin, H.; Tian, S. A comparative study for the stabilisation of heavy metal contaminated sediment by limestone, MnO2 and natural zeolite. Environ. Sci. Pollut. Res. Int. 2017, 24, 795–804. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.; Lu, Y.; Hong, J.; Ye, M.; Wang, Y.; Lu, H. Metal and metalloid contaminant availability in Yundang Lagoon sediments, Xiamen Bay, China, after 20 years continuous rehabilitation. J. Hazard. Mater. 2010, 175, 1048–1055. [Google Scholar] [CrossRef] [PubMed]
- Zhu, X.; Yao, J.; Wang, F.; Yuan, Z.; Liu, J.; Jordan, G.; Knudsen, T.S.; Avdalovic, J. Combined effects of antimony and sodium diethyldithiocarbamate on soil microbial activity and speciation change of heavy metals. Implications for contaminated lands hazardous material pollution in nonferrous metal mining areas. J. Hazard. Mater. 2018, 349, 160–167. [Google Scholar] [CrossRef] [PubMed]
- Filella, M. Antimony interactions with heterogeneous complexants in waters, sediments and soils: A review of data obtained in bulk samples. Earth Sci. Rev. 2011, 107, 325–341. [Google Scholar] [CrossRef]
- Xiao, E.; Krumins, V.; Xiao, T.; Dong, Y.; Tang, S.; Ning, Z.; Huang, Z.; Sun, W. Depth-resolved microbial community analyses in two contrasting soil cores contaminated by antimony and arsenic. Environ. Pollut. 2017, 221, 244–255. [Google Scholar] [CrossRef] [PubMed]
- Mendez-Garcia, C.; Pelaez, A.I.; Mesa, V.; Sanchez, J.; Golyshina, O.V.; Ferrer, M. Microbial diversity and metabolic networks in acid mine drainage habitats. Front. Microbiol. 2015, 6, 475. [Google Scholar]
- Sheik, C.S.; Mitchell, T.W.; Rizvi, F.Z.; Rehman, Y.; Faisal, M.; Hasnain, S.; McInerney, M.J.; Krumholz, L.R. Exposure of soil microbial communities to chromium and arsenic alters their diversity and structure. PLoS ONE 2012, 7, e40059. [Google Scholar] [CrossRef]
- Wang, Q.; He, M.; Wang, Y. Influence of combined pollution of antimony and arsenic on culturable soil microbial populations and enzyme activities. Ecotoxicology 2011, 20, 9–19. [Google Scholar] [CrossRef]
- He, M.; Wang, N.; Long, X.; Zhang, C.; Ma, C.; Zhong, Q.; Wang, A.; Wang, Y.; Pervaiz, A.; Shan, J. Antimony speciation in the environment: Recent advances in understanding the biogeochemical processes and ecological effects. J. Environ. Sci 2019, 75, 14–39. [Google Scholar] [CrossRef]
- Stebbing, A.R. Hormesis—The stimulation of growth by low levels of inhibitors. Sci. Total Environ. 1982, 22, 213–234. [Google Scholar] [CrossRef]
- Cai, L.; Liu, G.; Rensing, C.; Wang, G. Genes involved in arsenic transformation and resistance associated with different levels of arsenic-contaminated soils. BMC Microbiol. 2009, 9, 4. [Google Scholar] [CrossRef] [PubMed]
- Xiao, K.Q.; Li, L.G.; Ma, L.P.; Zhang, S.Y.; Bao, P.; Zhang, T.; Zhu, Y.G. Metagenomic analysis revealed highly diverse microbial arsenic metabolism genes in paddy soils with low-arsenic contents. Environ. Pollut. 2016, 211, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Mewis, K.; Armstrong, Z.; Song, Y.C.; Baldwin, S.A.; Withers, S.G.; Hallam, S.J. Biomining active cellulases from a mining bioremediation system. J. Biotechnol. 2013, 167, 462–471. [Google Scholar] [CrossRef] [PubMed]
- Wilson, R.M.; Cherrier, J.; Sarkodee-Adoo, J.; Bosman, S.; Mickle, A.; Chanton, J.P. Tracing the intrusion of fossil carbon into coastal Louisiana macrofauna using natural 14C and 13C abundances. Deep Sea Res. Part II Top. Stud. Oceanogr. 2016, 129, 89–95. [Google Scholar] [CrossRef]
- Gu, Y.; Van Nostrand, J.D.; Wu, L.; He, Z.; Qin, Y.; Zhao, F.J.; Zhou, J. Bacterial community and arsenic functional genes diversity in arsenic contaminated soils from different geographic locations. PLoS ONE 2017, 12, e0176696. [Google Scholar]
- Hartmann, M.; Frey, B.; Mayer, J.; Mäder, P.; Widmer, F. Distinct soil microbial diversity under long-term organic and conventional farming. ISME J. 2015, 9, 1177. [Google Scholar] [CrossRef] [PubMed]
- Kudo, K.; Yamaguchi, N.; Makino, T.; Ohtsuka, T.; Kimura, K.; Dong, D.T.; Amachi, S. Release of arsenic from soil by a novel dissimilatory arsenate-reducing bacterium, Anaeromyxobacter sp. strain PSR-1. Appl. Environ. Microbiol. 2013, 79, 4635–4642. [Google Scholar] [CrossRef]
- Giloteaux, L.; Holmes, D.E.; Williams, K.H.; Wrighton, K.C.; Wilkins, M.J.; Montgomery, A.P.; Smith, J.A.; Orellana, R.; Thompson, C.A.; Roper, T.J.; et al. Characterization and transcription of arsenic respiration and resistance genes during in situ uranium bioremediation. ISME J. 2013, 7, 370–383. [Google Scholar] [CrossRef]
- Kinegam, S.; Yingprasertchai, T.; Tanasupawat, S.; Leepipatpiboon, N.; Akaracharanya, A.; Kim, K.-W. Isolation and characterization of arsenite-oxidizing bacteria from arsenic-contaminated soils in Thailand. World J. Microbiol. Biotechnol. 2008, 24, 3091–3096. [Google Scholar] [CrossRef]
- Rosenstein, R.; Peschel, A.; Wieland, B.; Gotz, F. Expression and regulation of the antimonite, arsenite, and arsenate resistance operon of Staphylococcus xylosus plasmid pSX267. J. Bacteriol. 1992, 174, 3676–3683. [Google Scholar] [CrossRef]
- Terry, L.R.; Kulp, T.R.; Wiatrowski, H.; Miller, L.G.; Oremland, R.S. Microbiological oxidation of antimony(III) with oxygen or nitrate by bacteria isolated from contaminated mine sediments. Appl. Environ. Microbiol. 2015, 81, 8478–8488. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Wang, Q.; Zhang, S.; Qin, D.; Wang, G. Phylogenetic and genome analyses of antimony-oxidizing bacteria isolated from antimony mined soil. Int. Biodeterior. Biodegrad. 2013, 76, 76–80. [Google Scholar] [CrossRef]
- Meng, Y.L.; Liu, Z.; Rosen, B.P. As(III) and Sb(III) uptake by GlpF and efflux by ArsB in Escherichia coli. J. Biol. Chem. 2004, 279, 18334–18341. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Li, P.; Jiang, Z.; Liu, H.; Wei, D.; Wang, H.; Wang, Y. Diversity and abundance of arsenic methylating microorganisms in high arsenic groundwater from Hetao Plain of Inner Mongolia, China. Ecotoxicology 2018, 27, 1047–1057. [Google Scholar] [CrossRef]
- Zhou, Y.; Qin, Y.; Liu, X.; Feng, Z.; Zhu, H.; Yao, Q. Soil Bacterial Function Associated With Stylo (Legume) and Bahiagrass (Grass) Is Affected More Strongly by Soil Chemical Property Than by Bacterial Community Composition. Front. Microbiol. 2019, 10, 798. [Google Scholar] [CrossRef]

© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Huang, B.; Long, J.; Liao, H.; Liu, L.; Li, J.; Zhang, J.; Li, Y.; Wang, X.; Yang, R. Characteristics of Bacterial Community and Function in Paddy Soil Profile around Antimony Mine and Its Response to Antimony and Arsenic Contamination. Int. J. Environ. Res. Public Health 2019, 16, 4883. https://doi.org/10.3390/ijerph16244883
Huang B, Long J, Liao H, Liu L, Li J, Zhang J, Li Y, Wang X, Yang R. Characteristics of Bacterial Community and Function in Paddy Soil Profile around Antimony Mine and Its Response to Antimony and Arsenic Contamination. International Journal of Environmental Research and Public Health. 2019; 16(24):4883. https://doi.org/10.3390/ijerph16244883
Chicago/Turabian StyleHuang, Bocong, Jian Long, Hongkai Liao, Lingfei Liu, Juan Li, Jumei Zhang, Yirong Li, Xian Wang, and Rui Yang. 2019. "Characteristics of Bacterial Community and Function in Paddy Soil Profile around Antimony Mine and Its Response to Antimony and Arsenic Contamination" International Journal of Environmental Research and Public Health 16, no. 24: 4883. https://doi.org/10.3390/ijerph16244883
APA StyleHuang, B., Long, J., Liao, H., Liu, L., Li, J., Zhang, J., Li, Y., Wang, X., & Yang, R. (2019). Characteristics of Bacterial Community and Function in Paddy Soil Profile around Antimony Mine and Its Response to Antimony and Arsenic Contamination. International Journal of Environmental Research and Public Health, 16(24), 4883. https://doi.org/10.3390/ijerph16244883



