Single-Cell Analysis Reveals that Chronic Silver Nanoparticle Exposure Induces Cell Division Defects in Human Epithelial Cells
Abstract
1. Introduction
2. Materials and Methods
2.1. Silver Nanoparticle Characterization
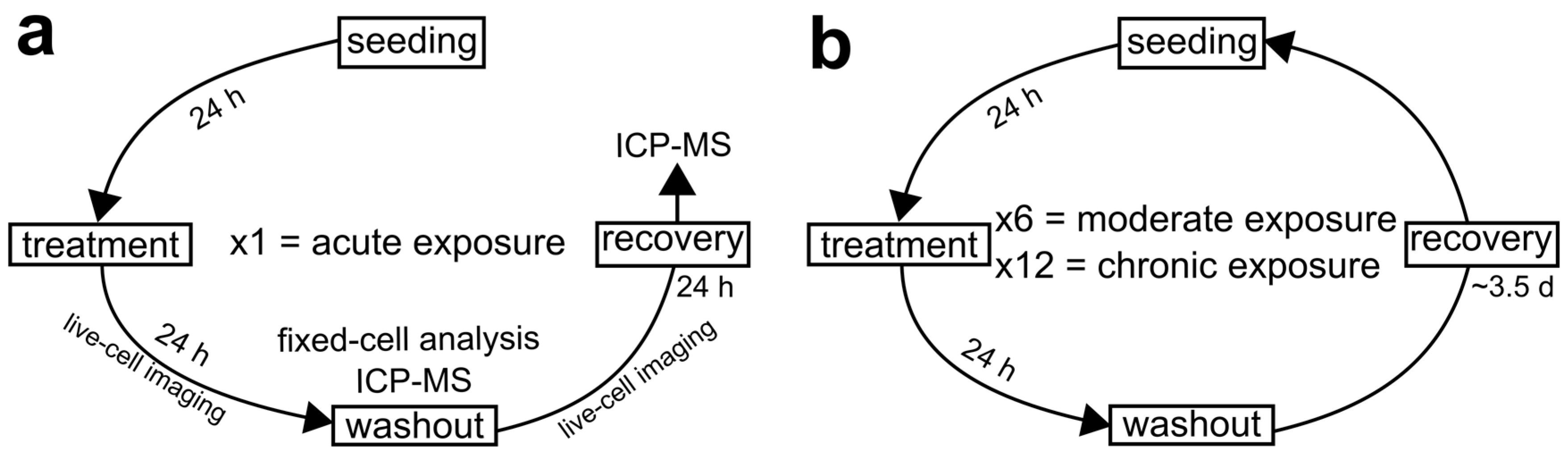
2.2. Cell Culture and Experimental Design
2.3. ICP-MS
2.4. Transmission Electron Microscopy
2.5. Growth Curves
2.6. Live Cell Imaging and Analysis
2.7. Trypan Blue Assay for Cell Viability
2.8. Immunostaining, Fluorescence Microscopy, and Fixed-Cell Analysis
3. Results
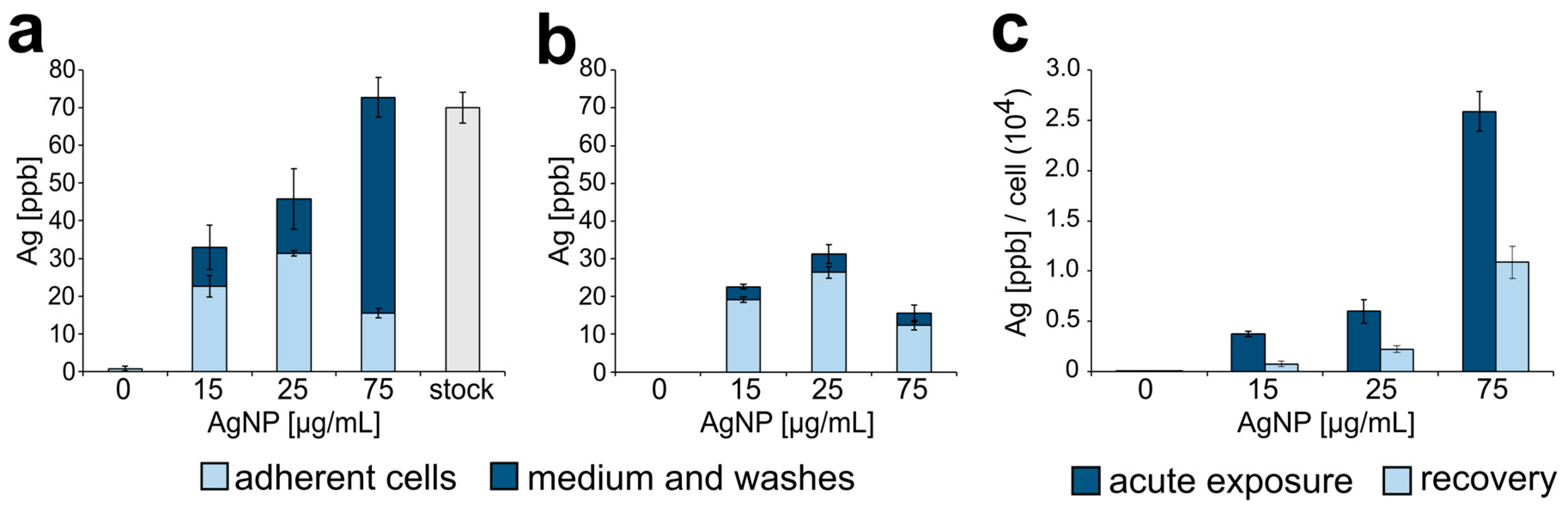
3.1. Silver Nanoparticle Internalization and Accumulation
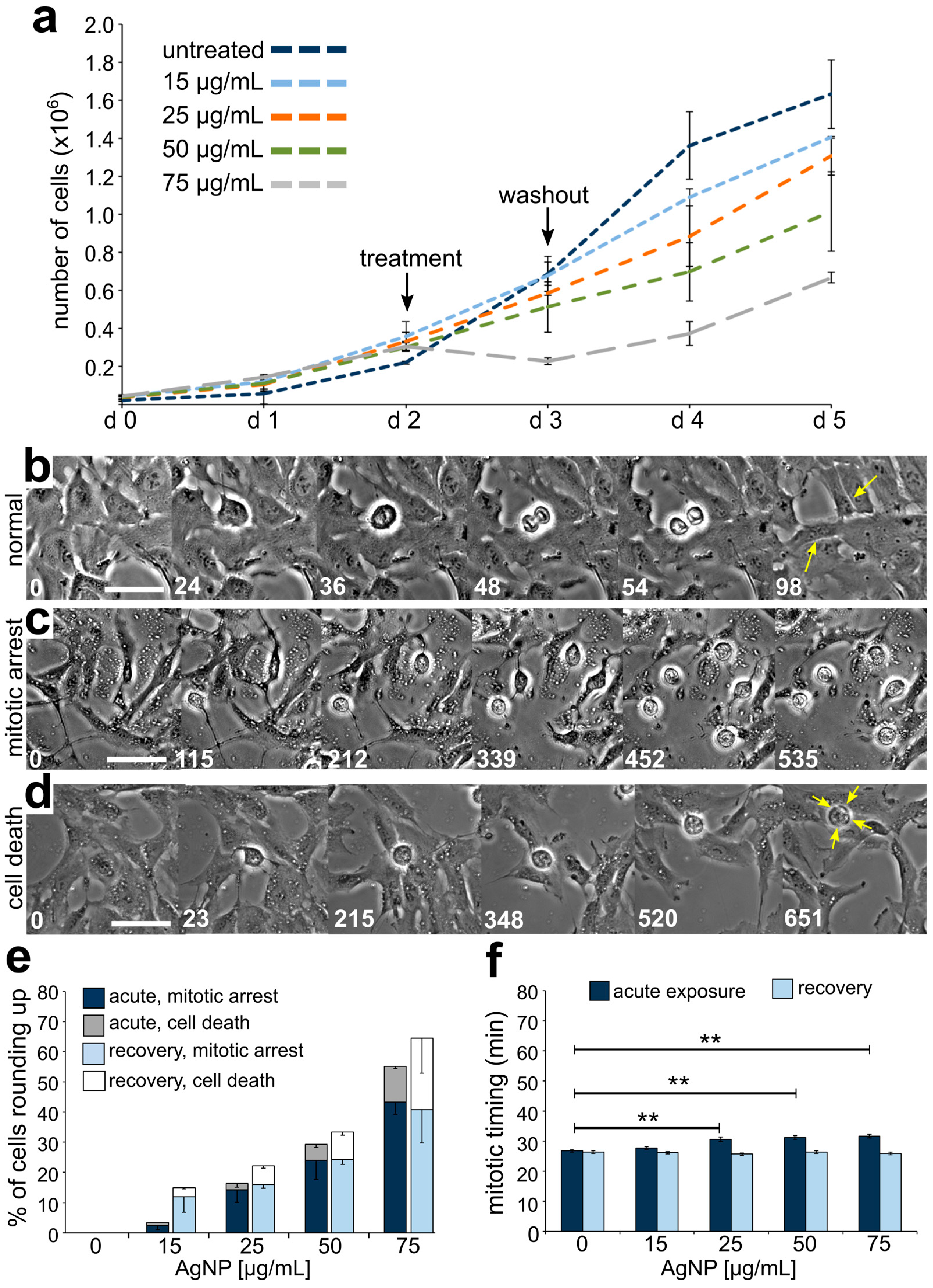
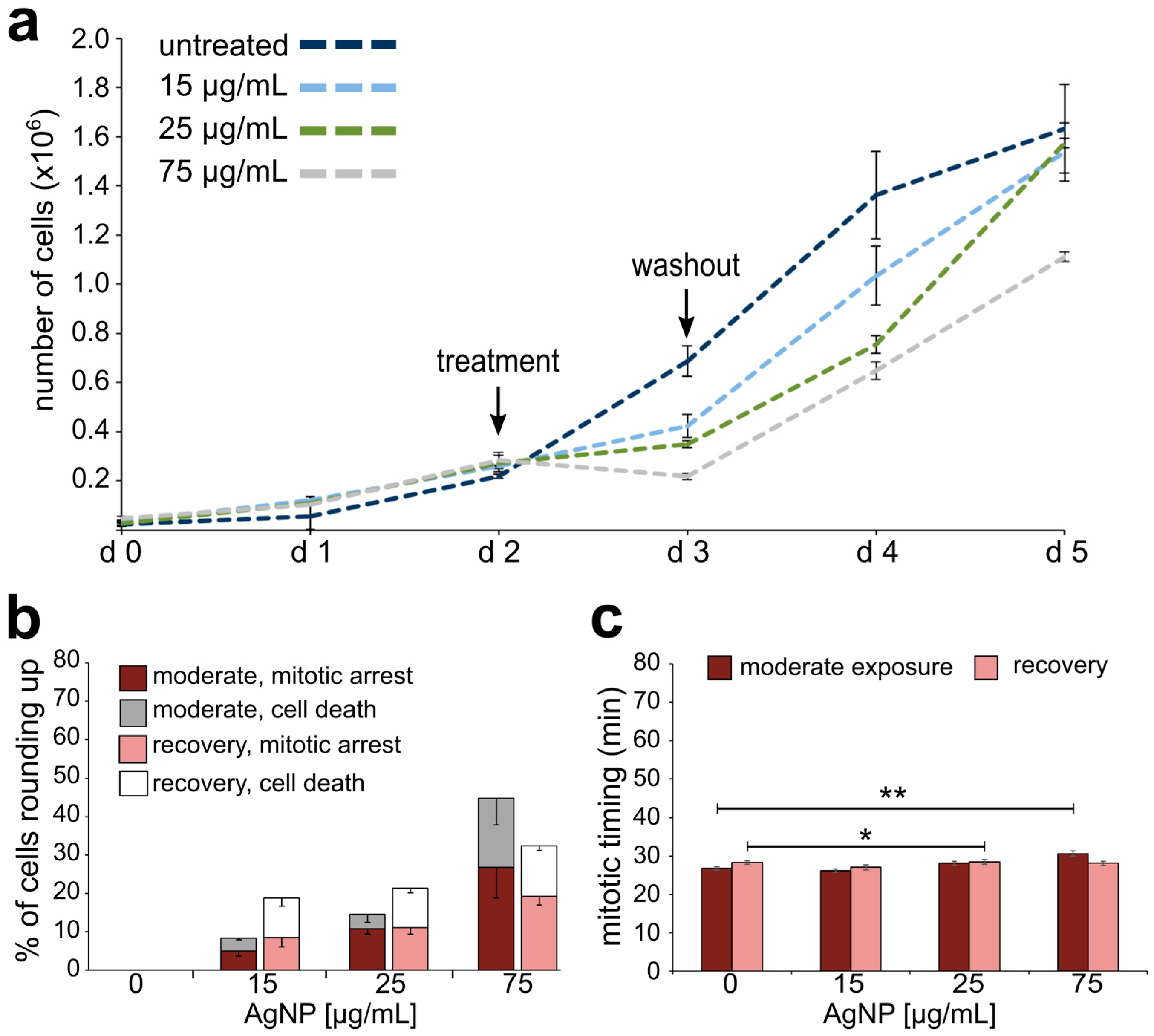
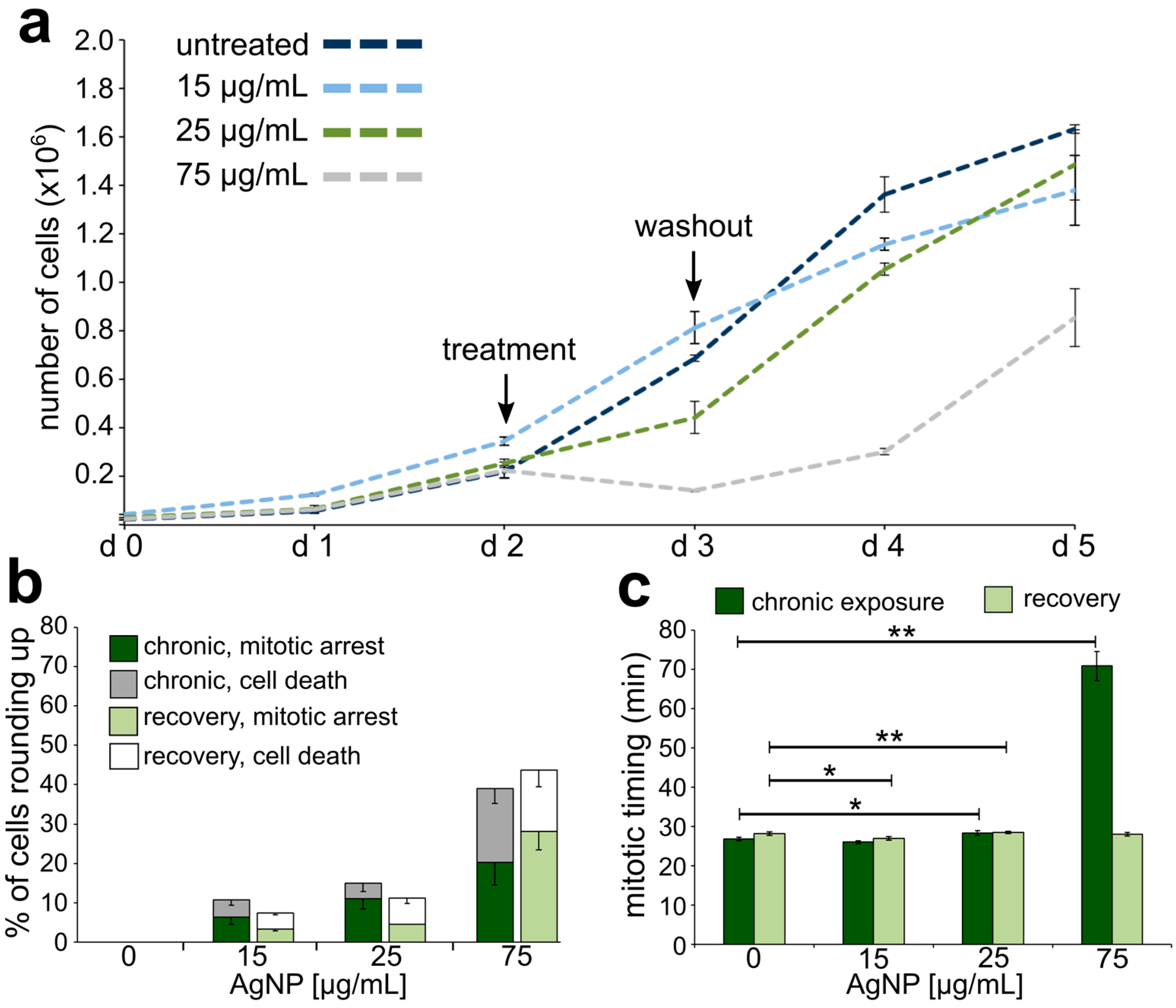
3.2. Reduced Cell Viability and Mitotic Delay from AgNP Exposure
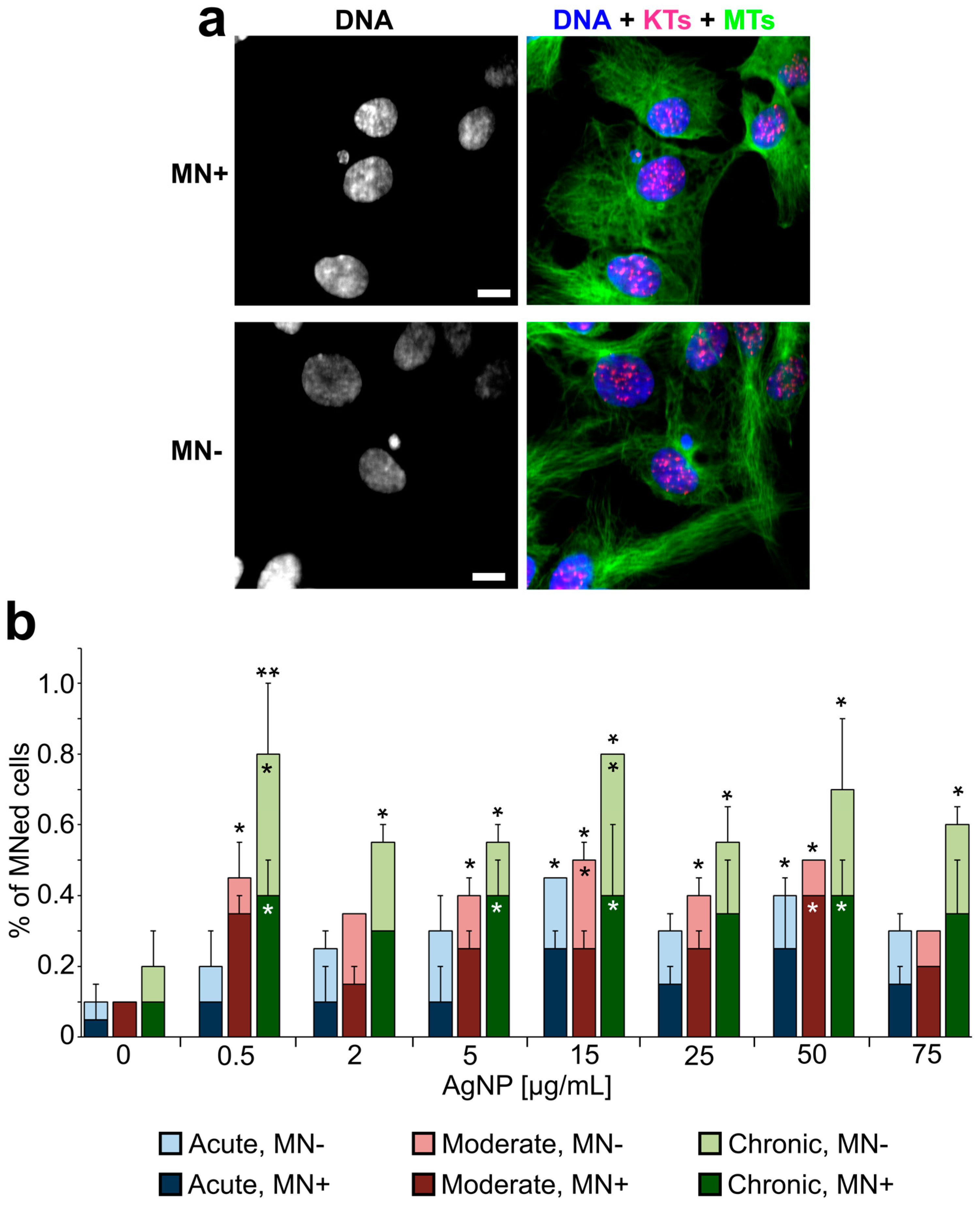
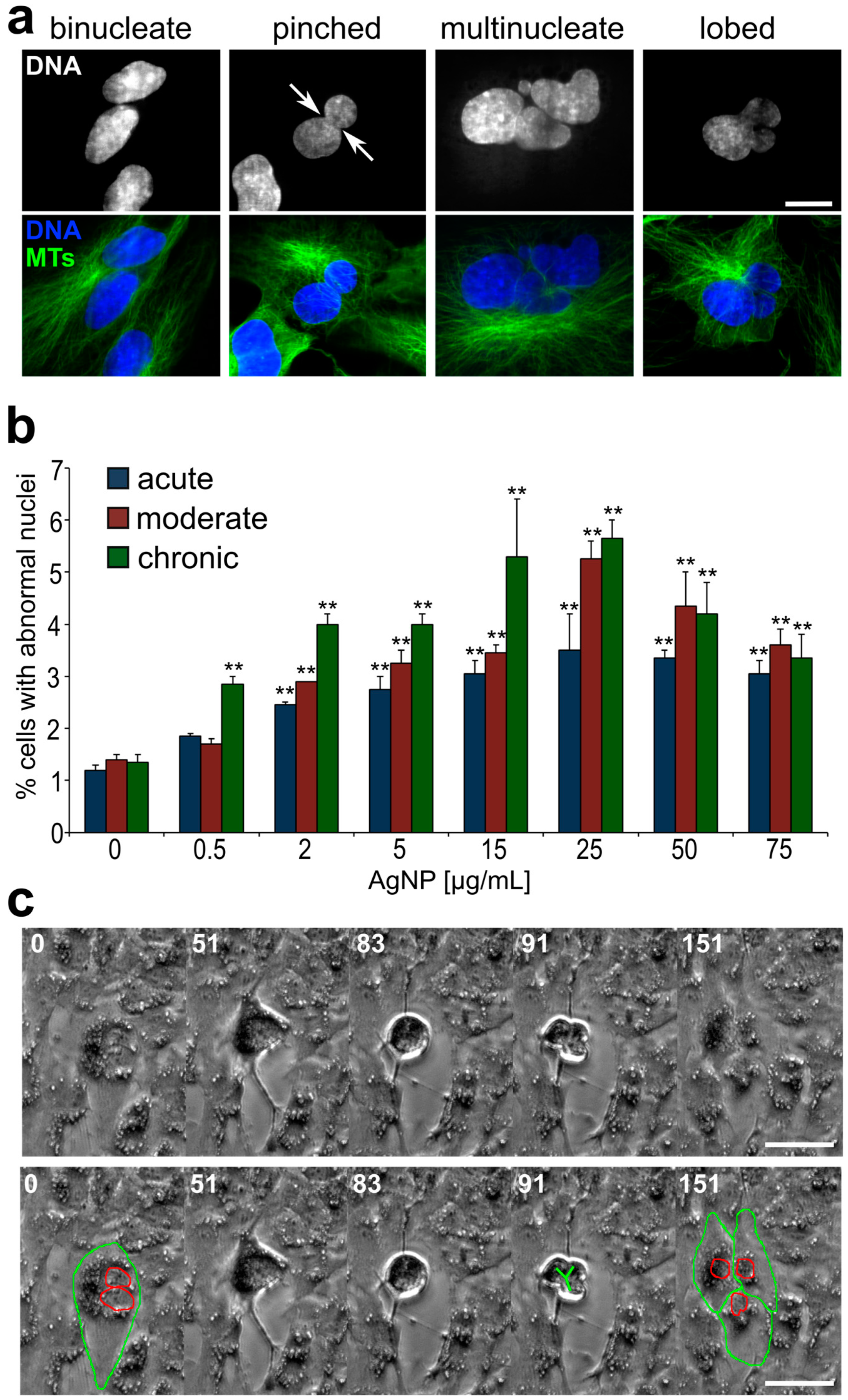
3.3. Emergence of Abnormal Nuclear Phenotypes in AgNP-Treated Cell Populations
4. Discussion
4.1. Long-Term in vitro Exposure to AgNPs, Even at Very Low Doses, Leads to the Accumulation of Cells with Abnormal DNA Content
4.2. Cells with Abnormal DNA Content Emerge as a Result of AgNP-Induced Cell Division Defects
4.3. The Health Risk Associated with Cell Division-Dependent Defects
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- United States Enviornmental Protection Agency. Chemical Data Report 2016; EPA: Washington, DC, USA, 2016.
- Hudson-Shore, M. Statistics of Scientific Procedures on Living Animals in Great Britain 2015- highlighting an ongoing upward trend in animal use and missed opportunities. Altern. Lab. Anim. 2016, 44, 569–580. [Google Scholar] [CrossRef] [PubMed]
- Krewski, D.; Acosta, D., Jr.; Andersen, M.; Anderson, H.; Bailar, J., III; Boekelheide, K.; Kelsey, K. Toxicity testing in the 21st century: A vision and a strategy. J. Toxicol. Environ. Health B Crit. Rev. 2010, 13, 51–138. [Google Scholar] [CrossRef] [PubMed]
- Taylor, K.; Gordon, N.; Langley, G.; Higgins, W. Estimates for worldwide laboratory animal use in 2005. Altern. Lab. Anim. 2008, 36, 327–342. [Google Scholar] [CrossRef] [PubMed]
- Zainzinger, V. Animal tests surge under new US chemical law. Science 2018, 360, 585–586. [Google Scholar] [CrossRef] [PubMed]
- Van Vliet, E. Current standing and future prospects for the technologies proposed to transform toxicity testing in the 21st century. Altern. Anim. Exp. 2011, 28, 17–44. [Google Scholar]
- Burden, N.; Sewell, F.; Chapman, K. Testing chemical safety: What is needed to ensure the widespread application of non-animal approaches? PLoS Biol. 2015, 13, e1002156. [Google Scholar] [CrossRef] [PubMed]
- Prior, H.; Casey, W.; Kimber, I.; Whelan, M.; Sewell, F. Reflections on the progress towards non-animal methods for acute toxicity testing of chemicals. Regul. Toxicol. Pharmacol. 2018, 102, 30–33. [Google Scholar] [CrossRef]
- Burden, N.; Aschberger, K.; Chaudhry, Q.; Clift, M.; Doak, S.; Fowler, P.; Johnston, H.; Landsiedel, R.; Rowland, J.; Stone, V. The 3Rs as a framework to support a 21st century approach for nanosafety assessment. Nano Today 2017, 12, 10–13. [Google Scholar] [CrossRef]
- Alves, V.; Muratov, E.; Zakharov, A.; Muratov, N.; Andrade, C.; Tropsha, A. Chemical toxicity prediction for major classes of industrial chemicals: Is it possible to develop universal models covering cosmetics, drugs, and pesticides? Food Chem. Toxicol. 2018, 112, 526–534. [Google Scholar] [CrossRef]
- Jain, A.; Singh, D.; Dubey, K.; Maurya, R.; Mittal, S.; Pandey, A. Models and Methods for In Vitro Toxicity. In In Vitro Toxicology; Academic Press: Cambridge, MA, USA, 2018; pp. 45–65. [Google Scholar]
- Eskes, C.; Boström, A.; Bowe, G.; Coecke, S.; Hartung, T.; Hendriks, G.; Pamies, D.; Piton, A.; Rovida, C. Good cell culture practices & in vitro toxicology. Toxicol. In Vitro 2017, 45, 272–277. [Google Scholar]
- Coleman, J.; Liu, R.; Wang, K.; Kumar, A. Detecting Apoptosis, Autophagy, and Necrosis. In Apoptosis Methods in Toxicology; Humana Press: New York, NY, USA, 2016. [Google Scholar]
- Smart, R.; Hodgson, E. Molecular and Biochemical Toxicology; John Wiley & Sons: Hoboken, NJ, USA, 2018. [Google Scholar]
- Dhawan, A.; Anderson, D. The Comet Assay in Toxicology; Royal Society of Chemistry: London, UK, 2016. [Google Scholar]
- Corton, J.; Williams, A.; Yauk, C. Using a gene expression biomarker to identify DNA damage-inducing agents in microarray profiles. Environ. Mol. Mutagen. 2018, 59, 772–784. [Google Scholar] [CrossRef] [PubMed]
- Pozarowski, P.; Darzynkiewicz, Z. Analysis of cell cycle by flow cytometry. Checkp. Control. Cancer 2010, 281, 301–311. [Google Scholar]
- Fenech, M. The in vitro micronucleus technique. Mutat. Res. 2000, 455, 81–95. [Google Scholar] [CrossRef]
- Smith-Roe, S.; Swartz, C.; Shepard, K.; Bryce, S.; Dertinger, S.; Waidyanatha, S.; Kissling, G.; Auerbach, S.; Witt, K.L. Black cohosh extracts and powders induce micronuclei, a biomarker of genetic damage, in human cells. Environ. Mol. Mutagen. 2018, 59, 416–426. [Google Scholar] [CrossRef]
- Ly, P.; Cleveland, D. Rebuilding Chromosomes After Catastrophe: Emerging Mechanisms of Chromothripsis. Trends Cell Biol. 2017, 27, 917–930. [Google Scholar] [CrossRef] [PubMed]
- Russo, A.; Pacchierotti, F.; Cimini, D.; Ganem, N.; Genescà, A.; Natarajan, A.; Degrassi, F. Genomic instability: Crossing pathways at the origin of structural and numerical chromosome changes. Environ. Mol. Mutagen. 2015, 56, 563–580. [Google Scholar] [CrossRef] [PubMed]
- Benameur, L.; Orsière, T.; Rose, J.; Botta, A. Detection of environmental clastogens and aneugens in human fibroblasts by cytokinesis-blocked micronucleus assay associated with immunofluorescent staining of CENP-A in micronuclei. Chemosphere 2011, 84, 676–680. [Google Scholar] [CrossRef]
- Hanahan, D.; Weinberg, R. Hallmarks of cancer: The next generation. Cell 2011, 144, 646–674. [Google Scholar] [CrossRef]
- Durán, N.; Durán, M.; de Jesus, M.; Seabra, A.; Fávaro, W.; Nakazato, G. Silver nanoparticles: A new view on mechanistic aspects on antimicrobial activity. Biol. Med. 2016, 12, 789–799. [Google Scholar] [CrossRef]
- Souza, L.; da Silva, V.; Franchi, L.; de Souza, T. Toxic and Beneficial Potential of Silver Nanoparticles: The Two Sides of the Same Coin. In Cellular and Molecular Toxicology of Nanoparticles; Springer: Cham, Switzerland, 2018; pp. 251–262. [Google Scholar]
- Chernousova, S.; Epple, M. Silver as Antibacterial Agent: Ion, Nanoparticle, and Metal. Angew. Chem. Int. Ed. Engl. 2013, 52, 1636–1653. [Google Scholar] [CrossRef]
- Vance, M.; Kuiken, T.; Vejerano, E.; Mcginnis, S.; Hochella, M.; Rejeski, D.; Hull, M. Nanotechnology in the real world: Redeveloping the nanomaterial consumer products inventory. Beilstein J. Nanotechnol. 2015, 6, 1769–1780. [Google Scholar] [CrossRef] [PubMed]
- Majeed, A.; Ullah, W.; Anwar, A.; Shuaib, A.; Ilyas, U.; Khalid, P.; Mustafa, G.; Junaid, M.; Faheem, B.; Ali, S. Cost-effective biosynthesis of silver nanoparticles using different organs of plants and their antimicrobial applications: A review. Mater. Technol. 2018, 33, 313–320. [Google Scholar] [CrossRef]
- Franci, G.; Falanga, A.; Galdiero, S.; Palomba, L.; Rai, M.; Morelli, G.; Galdiero, M. Silver nanoparticles as potential antibacterial agents. Molecules 2015, 20, 8856–8874. [Google Scholar] [CrossRef] [PubMed]
- Shrivastava, S.; Bera, T.; Roy, A.; Singh, G.; Ramachandrarao, P.; Dash, D. Characterization of enhanced antibacterial effects of novel silver nanoparticles. Nanotechnology 2007, 18, 225103. [Google Scholar] [CrossRef]
- Karthiga, P.; Rajeshkumar, S.; Annadurai, G. Mechanism of Larvicidal Activity of Antimicrobial Silver Nanoparticles Synthesized Using Garcinia mangostana Bark Extract. J. Clust. Sci. 2018, 29, 1233–1241. [Google Scholar] [CrossRef]
- Foldbjerg, R.; Autrup, H. Mechanisms of Silver Nanoparticle Toxicity. Arch. Basic Appl. Med. 2013, 1, 5–15. [Google Scholar]
- Hadrup, N.; Sharma, A.; Loeschner, K. Toxicity of silver ions, metallic silver, and silver nanoparticle materials after in vivo dermal and mucosal surface exposure: A review. Regul. Toxicol. Pharmacol. 2018, 98, 257–267. [Google Scholar] [CrossRef] [PubMed]
- Naasz, S.; Altenburger, R.; Kühnel, D. Environmental mixtures of nanomaterials and chemicals: The Trojan-horse phenomenon and its relevance for ecotoxicity. Sci. Total Environ. 2018, 635, 1170–1181. [Google Scholar] [CrossRef] [PubMed]
- Hsiao, I.; Hsieh, Y.; Wang, C.; Chen, I.; Huang, Y. Trojan-horse mechanism in the cellular uptake of silver nanoparticles verified by direct intra-and extracellular silver speciation analysis. Environ. Sci. Technol. 2015, 49, 3813–3821. [Google Scholar] [CrossRef]
- You, F.; Tang, W.; Yung, L.Y. Real-time monitoring of the Trojan-horse effect of silver nanoparticles by using a genetically encoded flourescent cell sensor. Nanoscale 2018, 10, 7726–7735. [Google Scholar] [CrossRef]
- Ahamed, M.; Alsalhi, M.; Siddiqui, M. Silver nanoparticle applications and human health. Clin. Chim. Acta 2010, 411, 1841–1848. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.; Ryu, D. Silver nanoparticle-induced oxidative stress, genotoxicity and apoptosis in cultured cells and animal tissues. J. Appl. Toxicol. 2013, 33, 78–89. [Google Scholar] [CrossRef] [PubMed]
- Asharani, P.; Low, G.; Mun, K.; Hande, M.; Valiyaveettil, S. Cytotoxicity and Genotoxicity of Silver. ACS Nano 2009, 3, 279–290. [Google Scholar] [CrossRef] [PubMed]
- Schneider, T.; Westermann, M.; Glei, M. In vitro uptake and toxicity studies of metal nanoparticles and metal oxide nanoparticles in human HT29 cells. Arch. Toxicol. 2017, 91, 3517–3527. [Google Scholar] [CrossRef] [PubMed]
- Harvanova, M.; Jiravova, J.; Malohlava, J.; Tomankova, K.; Jirova, D.; Kolarova, H. Ramen imaging of cellular uptake and studies of silver nanoparticles effect in BJ human fibroblasts cell lines. Int. J. Pharm. 2017, 528, 280–286. [Google Scholar] [CrossRef] [PubMed]
- Xie, H.; Wang, P.; Wu, J. Effect of exposure of osteoblast-like cells to low-dose silver nanoparticles: Uptake, retention and osteogenic activity. Artif. Cells Nanomed. Biotechnol. 2018, 47, 260–267. [Google Scholar] [CrossRef] [PubMed]
- Castro-Gamboa, S.; Garcia-Garcia, M.R.; PiñonZarate, G.; Rojas-Lemus, M.; Jarquin-Yañez, K.; Herrera-Enriquez, M.; Fortoul, T.; Toledano-Magaña, Y.; Garcia-Iglesias, T.; Pestryakov, A.; et al. Toxicity of silver nanoparticles in mouse bone marrow-derived dendritic cells: Implications for phenotype. J. Immunotoxicol. 2019. [Google Scholar] [CrossRef]
- Tang, J.; Lu, X.; Chen, B.; Cai, E.; Liu, W.; Jiang, J.; Chen, F.; Shan, X.; Zhang, H. Mechanisms of silver nanoparticles-induced cytotoxicity and apoptosis in rat tracheal epithelial cells. J. Toxicol. Sci. 2019, 44, 155–165. [Google Scholar] [CrossRef]
- Wang, X.; Li, T.; Su, X.; Li, J.; Li, W.; Gan, J.; Wu, T.; Kong, L.; Zhang, T.; Tang, M.; et al. Genotoxic effects of silver nanoparticles with/without coating in human liver HepG2 cells and in mice. J. Appl. Toxicol. 2019, 39, 908–918. [Google Scholar] [CrossRef]
- Zhang, J.; Chen, Y.; Gao, M.; Wang, Z.; Liu, R.; Xia, T.; Liu, S. Silver Nanoparticles Compromise Female Embryonic Stem Cell Differentiation through Disturbing X Chromosome Inactivation. ACS Nano 2019, 13, 2050–2061. [Google Scholar] [CrossRef]
- Piao, M.; Kang, K.; Lee, I.; Kim, H.; Kim, S.; Choi, J.; Hyun, J. Silver nanoparticles induce oxidative cell damage in human liver cells through inhibition of reduced glutathione and induction of mitochondria-involved apoptosis. Toxicol. Lett. 2011, 201, 92–100. [Google Scholar] [CrossRef] [PubMed]
- Park, E.; Yi, J.; Kim, Y.; Choi, K.; Park, K. Silver nanoparticles induce cytotoxicity by a Trojan-horse type mechanism. Toxicol. In Vitro 2010, 24, 872–878. [Google Scholar] [CrossRef] [PubMed]
- Liu, W.; Wu, Y.; Wang, C.; Li, H.; Wang, T.; Liao, C.; Jiang, G. Impact of silver nanoparticles on human cells: Effect of particle size. Nanotoxicology 2010, 4, 319–330. [Google Scholar] [CrossRef] [PubMed]
- Chairuangkitti, P.; Lawanprasert, S.; Roytrakul, S.; Aueviriyavit, S.; Phummiratch, D.; Kulthong, K.; Chanvorachote, P.; Maniratanachote, R. Silver nanoparticles induce toxicity in A549 cells via ROS-dependent and ROS-independent pathways. Toxicol. In Vitro 2013, 27, 330–338. [Google Scholar] [CrossRef] [PubMed]
- Eom, H.; Choi, J. p38 MAPK Activation, DNA Damage, Cell Cycle Arrest and Apoptosis As Mechanisms of Toxicity of Silver Nanoparticles in Jurkat T Cells. Environ. Sci. Technol. 2010, 44, 8337–8342. [Google Scholar] [CrossRef] [PubMed]
- Wei, L.; Tang, J.; Zhang, Z.; Chen, Y.; Zhou, G.; Xi, T. Investigation of the cytotoxicity mechanism of silver nanoparticles in vitro. Biomed. Mater. 2010, 5, 044103. [Google Scholar] [CrossRef] [PubMed]
- Bastos, V.; Duarte, I.; Santos, C.; Oliveira, H. Genotoxicity of citrate-coated silver nanoparticles to human keratinocytes assessed by the comet assay and cytokinesis blocked micronucleus assay. Environ. Sci. Pollut. Res. 2017, 24, 5039–5048. [Google Scholar] [CrossRef]
- Luther, E.; Koehler, Y.; Diendorf, J.; Epple, M.; Dringen, R. Accumulation of silver nanoparticles by cultured primary brain astrocytes. Nanotechnology 2011, 22, 375101. [Google Scholar] [CrossRef]
- Arora, S.; Tyagi, N.; Bhardwaj, A.; Rusu, L.; Palanki, R.; Vig, K.; Singh, S. Silver nanoparticles protect human keratinocytes against UVB radiation-induced DNA damage and apoptosis: Potential for prevention of skin carcinogenesis. Nanomedicine 2015, 11, 1265–1275. [Google Scholar] [CrossRef]
- Baudoin, N.; Cimini, D. A guide to classifying mitotic stages and mitotic defects in fixed cells. Chromosoma 2018, 127, 215–227. [Google Scholar] [CrossRef]
- Krystek, P.; Kettler, K.; van der Wagt, B.; de Jong, W. Exploring influences on the cellular uptake of medium-sized silver nanoparticles into THP-1 cells. Microchem. J. 2015, 120, 45–50. [Google Scholar] [CrossRef]
- Maiorano, G.; Sabella, S.; Sorce, B.; Brunetti, V.; Malvindi, M.; Cingolani, R.; Pompa, P. Effects of cell culture media on the dynamic formation of protein−nanoparticle complexes and influence on the cellular response. ACS Nano 2010, 4, 7481–7491. [Google Scholar] [CrossRef] [PubMed]
- Müller, L.; Steiner, S.; Rodriguez-Lorenzo, L.; Petri-Fink, A.; Rothen-Rutishauser, B.; Latzin, P. Exposure to silver nanoparticles affects viability and function of natural killer cells, mostly via the release of ions. Cell Biol. Toxicol. 2018, 34, 167–176. [Google Scholar] [CrossRef] [PubMed]
- Degrassi, F.; Tanzarella, C. Immunofluorescent staining of kinetochores in micronuclei: A new assay for the detection of aneuploidy. Mutat. Res. 1988, 203, 339–345. [Google Scholar] [CrossRef]
- Sung, J.; Lee, E.; Lee, J.; Yu, I. Lung Function Changes in Sprague-Dawley Rats After Prolonged Inhalation Exposure to Silver Nanoparticles. Inhal. Toxicol. 2008, 20, 567–574. [Google Scholar] [CrossRef] [PubMed]
- Santoro, C.; Duchsherer, N.; Grainger, D. Minimal In Vitro Antimicrobial Efficacy and Ocular Cell Toxicity from Silver Nanoparticles. Nanobiotechnology 2007, 3, 55–65. [Google Scholar] [CrossRef]
- Kim, H.; Kim, M.; Lee, S.; Oh, S.; Chung, K. Genotoxic effects of silver nanoparticles stimulated by oxidative stress in human normal bronchial epithelial (BEAS-2B) cells. Mutat. Res. 2011, 726, 129–135. [Google Scholar] [CrossRef]
- Bakhoum, S.; Kabeche, L.; Wood, M.; Laucius, C.; Qu, D.; Laughney, A.; Zaki, B. Numerical chromosomal instability mediates susceptibility to radiation treatment. Nat. Commun. 2015, 6, 5990. [Google Scholar] [CrossRef]
- Hodgson, J.; Darnton, A. The quantitative risks of mesothelioma and lung cancer in relation to asbestos exposure. Ann. Occup. Hyg. 2000, 44, 565–601. [Google Scholar] [CrossRef]
- Selikoff, I.; Churg, J.; Hammond, E. Relation between exposure to asbestos and mesothelioma. N. Engl. J. Med. 1965, 272, 560–565. [Google Scholar] [CrossRef]
- Oshimura, M.; Hesterberg, T.W.; Tsutsui, T.; Barrett, J.C. Correlation of asbestos-induced cytogenetic effects with cell transformation of Syrian hamster embryo cells in culture. Cancer Res. 1984, 1, 5017–5022. [Google Scholar]
- Walker, C.; Everitt, J.; Barrett, J. Possible cellular and molecular mechanisms for asbestos carcinogenicity. Am. J. Ind. Med. 1992, 21, 253–273. [Google Scholar] [CrossRef] [PubMed]
- Ault, J.; Cole, R.; Jensen, C.; Jensen, L.; Bachert, L.; Rieder, C. Behavior of crocidolite asbestos during mitosis in living vertebrate lung epithelial cells. Cancer Res. 1995, 55, 792–798. [Google Scholar] [PubMed]
- Jensen, C.; Jensen, L.; Rieder, C.; Cole, R.; Ault, J. Long crocidolite asbestos fibers cause polyploidy by sterically blocking cytokinesis. Carcinogenesis 1996, 17, 2013–2021. [Google Scholar] [CrossRef] [PubMed]
- Wang, F.; Chen, Y.; Wang, Y.; Yin, Y.; Qu, G.; Song, M.; Wang, H. Ultra-long silver nanowires induced mitotic abnormalities and cytokinetic failure in A549 cells. Nanotoxicology 2019, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Johnston, H.; Hutchison, G.; Christensen, F.; Peters, S.; Hankin, S.; Aschberger, K.; Stone, V. A critical review of the biological mechanisms underlying the in vivo and in vitro toxicity of carbon nanotubes: The contribution of physico-chemical characteristics. Nanotoxicology 2010, 4, 207–246. [Google Scholar] [CrossRef] [PubMed]
- Sargent, L.; Hubbs, A.; Young, S.; Kashon, M.; Dinu, C.; Salisbury, J.; Benkovic, S.; Lowry, D.; Murray, A.; Kisin, E.; et al. Single-walled carbon nanotube-induced mitotic disruption. Mutat. Res. 2012, 745, 28–37. [Google Scholar] [CrossRef] [PubMed]
- Bhatia, A.; Kumar, Y. Cancer cell micronucleus: An update on clinical and diagnostic applications. APMIS 2013, 121, 569–581. [Google Scholar] [CrossRef]
- Bonassi, S.; Znaor, A.; Ceppi, M.; Lando, C.; Chang, W.; Holland, N.; Kirsch-Volders, M.; Zeiger, E.; Ban, S.; Barale, R.; et al. An increased micronucleus frequency in peripheral blood lymphocytes predicts the risk of cancer in humans. Carcinogenesis 2007, 28, 625–631. [Google Scholar] [CrossRef] [PubMed]
- Bakhoum, S.; Silkworth, W.; Nardi, I.; Nicholson, J.; Compton, D.; Cimini, D. The mitotic origin of chromosomal instability. Curr. Biol. 2014, 24, 148–149. [Google Scholar] [CrossRef]
- Thompson, S.; Compton, D. Examining the link between chromosomal instability and aneuploidy in human cells. J. Cell Biol. 2008, 180, 665–672. [Google Scholar] [CrossRef] [PubMed]
- Cimini, D.; Fioravanti, D.; Salmon, E.; Degrassi, F. Merotelic kinetochore orientation versus chromosome mono-orientation in the origin of lagging chromosomes in human primary cells. J. Cell Sci. 2002, 115, 507–515. [Google Scholar] [PubMed]
- Hoffelder, D.; Luo, L.; Burke, N.; Watkins, S.; Gollin, S.; Saunders, W. Resolution of anaphase bridges in cancer cells. Chromosoma 2004, 112, 389–397. [Google Scholar] [CrossRef] [PubMed]
- Terradas, M.; Martin, M.; Hernandez, L.; Tusell, L.; Genesca, A. Nuclear envelope defects impede a proper response to micronuclear DNA lesions. Mutat. Res. 2012, 729, 35–40. [Google Scholar] [CrossRef] [PubMed]
- Crasta, K.; Ganem, N.; Dagher, R.; Lantermann, A.; Ivanova, E.; Pan, Y.; Nezi, L.; Protopopov, A.; Chowdhury, D.; Pellman, D. DNA breaks and chromosome pulverization from errors in mitosis. Nature 2012, 482, 53–58. [Google Scholar] [CrossRef] [PubMed]
- Leibowitz, M.; Zhang, C.; Pellman, D. Chromothripsis: A New Mechanism for Rapid Karyotype Evolution. Annu. Rev. Genet. 2015, 49, 183–1047. [Google Scholar] [CrossRef] [PubMed]
- Fujiwara, T.; Bandi, M.; Nitta, M.; Ivanova, E.; Bronson, R.; Pellman, D. Cytokinesis failure generating tetraploids promotes tumorigenesis in p53-null cells. Nature 2005, 437, 1043–1047. [Google Scholar] [CrossRef] [PubMed]
- Zack, T.; Schumacher, S.; Carter, S.; Cherniack, A.; Saksena, G.; Tabak, B.; Lawrence, M.; Zhsng, C.; Wala, J.; Mermel, C.; et al. Pan-cancer patterns of somatic copy number alteration. Nat. Genet. 2013, 45, 1134–1140. [Google Scholar] [CrossRef]
- Wangsa, D.; Quintanilla, I.; Torabi, K.; Vila- Casadesus, M.; Ercilla, A.; Lus, G.; Yuce, Z.; Galofre, C.; Cuatrecasas, M.; Lozano, J.; et al. Near-tetraploid cancer cells show chromosome instability triggered by replication stress and exhibit enhanced invasiveness. FASEB J. 2018, 32, 3502–3517. [Google Scholar] [CrossRef]

© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Garcia, E.B.; Alms, C.; Hinman, A.W.; Kelly, C.; Smith, A.; Vance, M.; Loncarek, J.; Marr, L.C.; Cimini, D. Single-Cell Analysis Reveals that Chronic Silver Nanoparticle Exposure Induces Cell Division Defects in Human Epithelial Cells. Int. J. Environ. Res. Public Health 2019, 16, 2061. https://doi.org/10.3390/ijerph16112061
Garcia EB, Alms C, Hinman AW, Kelly C, Smith A, Vance M, Loncarek J, Marr LC, Cimini D. Single-Cell Analysis Reveals that Chronic Silver Nanoparticle Exposure Induces Cell Division Defects in Human Epithelial Cells. International Journal of Environmental Research and Public Health. 2019; 16(11):2061. https://doi.org/10.3390/ijerph16112061
Chicago/Turabian StyleGarcia, Ellen B., Cynthia Alms, Albert W. Hinman, Conor Kelly, Adam Smith, Marina Vance, Jadranka Loncarek, Linsey C. Marr, and Daniela Cimini. 2019. "Single-Cell Analysis Reveals that Chronic Silver Nanoparticle Exposure Induces Cell Division Defects in Human Epithelial Cells" International Journal of Environmental Research and Public Health 16, no. 11: 2061. https://doi.org/10.3390/ijerph16112061
APA StyleGarcia, E. B., Alms, C., Hinman, A. W., Kelly, C., Smith, A., Vance, M., Loncarek, J., Marr, L. C., & Cimini, D. (2019). Single-Cell Analysis Reveals that Chronic Silver Nanoparticle Exposure Induces Cell Division Defects in Human Epithelial Cells. International Journal of Environmental Research and Public Health, 16(11), 2061. https://doi.org/10.3390/ijerph16112061




