Data-Driven Analysis of Antimicrobial Resistance in Foodborne Pathogens from Six States within the US
Abstract
1. Introduction
2. Materials and Methods
2.1. Data from the NCBI Pathogen Detection Isolates Browser (NPDIB)
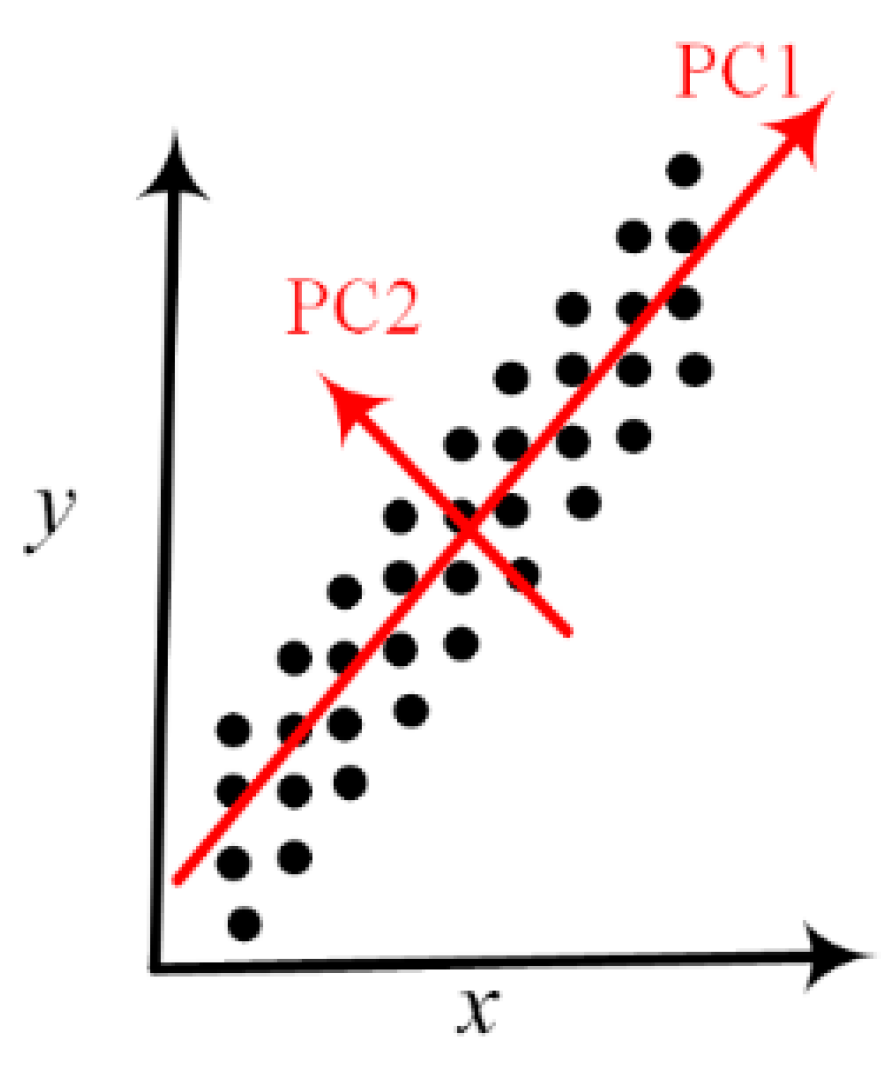
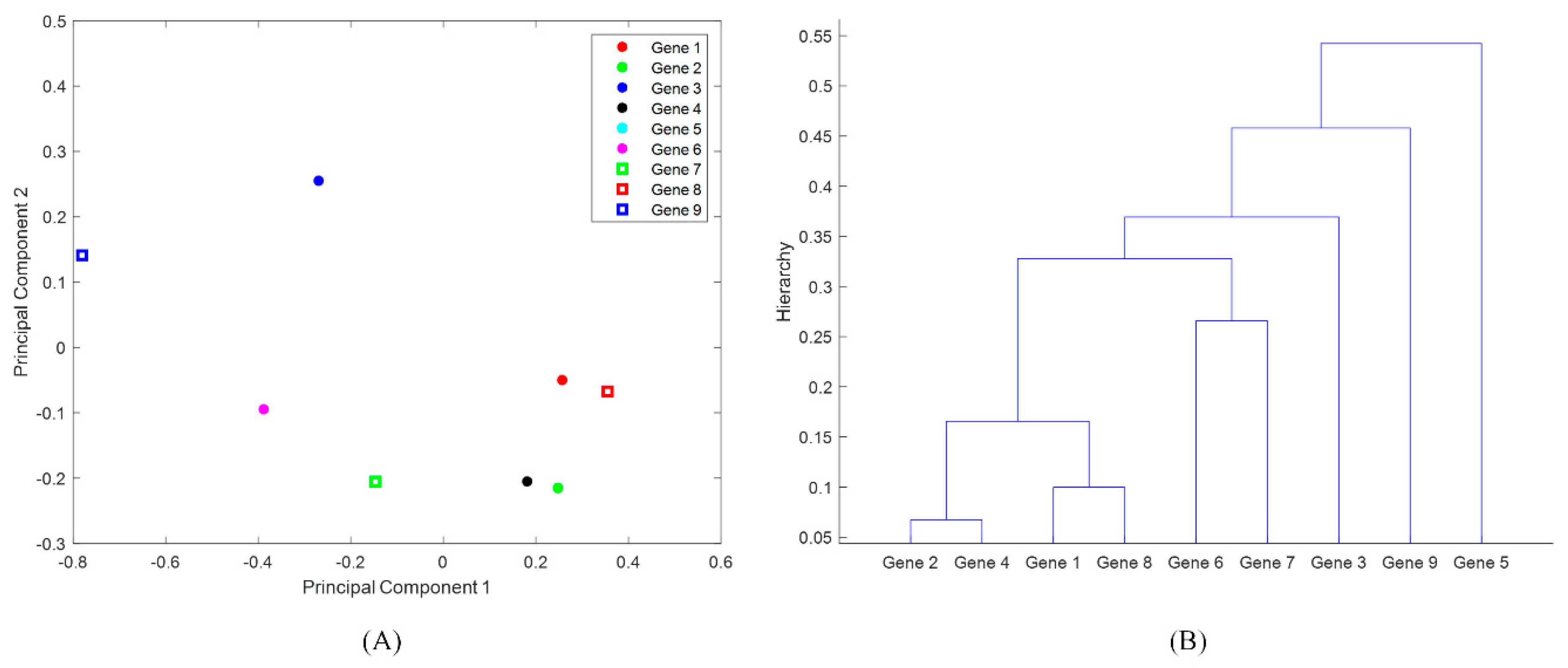
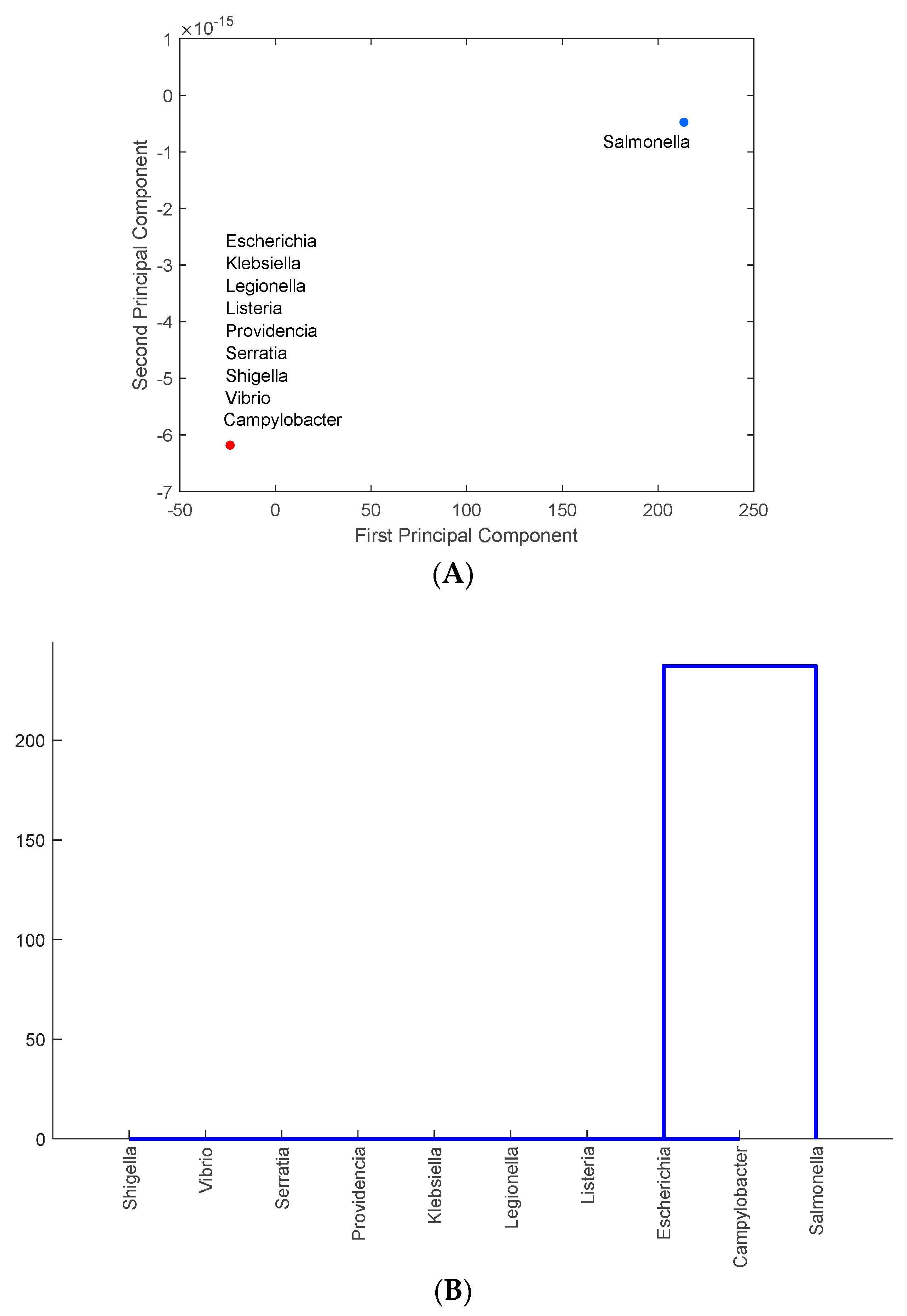
2.2. Multivariate Statistical Analysis: Principal Component Analysis and Hierarchical Clustering
3. Results
3.1. Identification of Antimicrobials to Which Pathogens Show Most Resistance in States PA, MD, NY, NM, MN, and CA
3.2. Identification of Antimicrobial-Resistance Genes Most Common in States PA, MD, NY, NM, MN, and CA
3.3. Identification of Foodborne Pathogens Mostly Carrying Antimicrobial-Resistance Genes in PA, MD, NY, NM, MN, and CA
3.4. Identification of Meats Mostly Carrying Antimicrobial-Resistance Genes in States PA, MD, NY, NM, MN, and CA
4. Discussion
4.1. Overuse of Antimicrobials and Antimicrobial-Resistance Genes
4.2. The Impact of Geographic Location on the Distribution of Antimicrobial-Resistance Genes
4.3. Limitations and Future Work
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Hoffmann, S.; Maculloch, B.; Batz, M. Economic burden of major foodborne illnesses acquired in the United States. Curr. Politics Econ. U. S. Can. Mex. 2015, 17, 543–616. [Google Scholar]
- Scallan, E.; Hoekstra, R.M.; Angulo, F.J.; Tauxe, R.V.; Widdowson, M.A.; Roy, S.L.; Jones, J.L.; Griffin, P.M. Foodborne Illness Acquired in the United States-Major Pathogens. Emerg. Infect. Dis. 2011, 17, 7–15. [Google Scholar] [CrossRef]
- Mezal, E.H.; Sabol, A.; Khan, M.A.; Ali, N.; Stefanova, R.; Khan, A.A. Isolation and molecular characterization of Salmonella enterica serovar Enteritidis from poultry house and clinical samples during 2010. Food Microbiol. 2014, 38, 67–74. [Google Scholar] [CrossRef] [PubMed]
- Tenover, F.C. Mechanisms of antimicrobial resistance in bacteria. Am. J. Med. 2006, 119, S3–S10. [Google Scholar] [CrossRef]
- Barriere, S.L. Clinical, economic and societal impact of antibiotic resistance. Expert Opin. Pharm. 2015, 16, 151–153. [Google Scholar] [CrossRef] [PubMed]
- Jain, R.; Rivera, M.C.; Lake, J.A. Horizontal gene transfer among genomes: The complexity hypothesis. Proc. Natl. Acad. Sci. USA 1999, 96, 3801–3806. [Google Scholar] [CrossRef]
- White, D.G.; Zhao, S.H.; Simjee, S.; Wagner, D.D.; McDermott, P.F. Antimicrobial resistance of foodborne pathogens. Microbes Infect. 2002, 4, 405–412. [Google Scholar] [CrossRef]
- Sivapalasingam, S.; Nelson, J.M.; Joyce, K.; Hoekstra, M.; Angulo, F.J.; Mintz, E.D. High prevalence of antimicrobial resistance among Shigella isolates in the United States tested by the National Antimicrobial Resistance Monitoring System from 1999 to 2002. Antimicrob. Agents Chem. 2006, 50, 49–54. [Google Scholar] [CrossRef] [PubMed]
- Zhao, S.; Blickenstaff, K.; Glenn, A.; Ayers, S.L.; Friedman, S.L.; Abbott, J.W.; McDermott, P.F. beta-Lactam Resistance in Salmonella Strains Isolated from Retail Meats in the United States by the National Antimicrobial Resistance Monitoring System between 2002 and 2006. Appl. Environ. Microbiol. 2009, 75, 7624–7630. [Google Scholar] [CrossRef]
- Qin, S.J. Statistical process monitoring: Basics and beyond. J. Chem. 2003, 17, 480–502. [Google Scholar] [CrossRef]
- Kourti, T. Application of latent variable methods to process control and multivariate statistical process control in industry. Int. J. Adapt. Control 2005, 19, 213–246. [Google Scholar] [CrossRef]
- Arnau, V.; Mars, S.; Marin, I. Iterative cluster analysis of protein interaction data. Bioinformatics 2005, 21, 364–378. [Google Scholar] [CrossRef]
- Wang, X.Z.; Smith, K.; Hyndman, R. Characteristic-based clustering for time series data. Data Min. Knowl. Discov. 2006, 13, 335–364. [Google Scholar] [CrossRef]
- Bar-Joseph, Z.; Demaine, E.D.; Gifford, D.K.; Srebro, N.; Hamel, A.M.; Jaakkola, T.S. K-ary clustering with optimal leaf ordering for gene expression data. Bioinformatics 2003, 19, 1070–1078. [Google Scholar] [CrossRef]
- Marshall, B.M.; Levy, S.B. Food Animals and Antimicrobials: Impacts on Human Health. Clin. Microbiol. Rev. 2011, 24, 718. [Google Scholar] [CrossRef]
- Van Boeckel, T.P.; Brower, C.; Gilbert, M.; Grenfell, B.T.; Levin, S.A.; Robinson, T.P.; Teillant, A.; Laxminarayan, R. Global trends in antimicrobial use in food animals. Proc. Natl. Acad. Sci. USA 2015, 112, 5649–5654. [Google Scholar] [CrossRef]
- USDA Poultry Production Data. Available online: https://www.usda.gov/sites/default/files/documents/nass-poultry-stats-factsheet.pdf (accessed on 1 June 2018).
- Martens, E.; Demain, A.L. The antibiotic resistance crisis, with a focus on the United States. J. Antibiot. 2017, 70, 520–526. [Google Scholar] [CrossRef]
- Jolliffe, I.T.; Cadima, J. Principal component analysis: A review and recent developments. Philos. Trans R. Soc. A 2016, 374. [Google Scholar] [CrossRef]
- Ueno, Y.; Forghani, B.; Forghani, R.; Dohan, A.; Zeng, X.Z.; Chamming’s, F.; Arseneau, J.; Fu, L.L.; Gilbert, L.; Gallix, B.; et al. Dimensional Reduction and Feature Selection: Principal Component Analysis for Data Mining Response. Radiology 2017, 285, 1055–1056. [Google Scholar]
- Kashanian, H.; Dabaghi, E. Feature Dimension Reduction of Multisensor Data Fusion using Principal Component Fuzzy Analysis. Int. J. Eng.-Iran 2017, 30, 493–499. [Google Scholar] [CrossRef]
- Chambers, J.M. Software for Data Analysis: Programming with R. Stat. Comput. Ser. 2008. [Google Scholar] [CrossRef]
- Wise, R. Antimicrobial resistance: Priorities for action. J. Antimicrob. Chem. 2002, 49, 585–586. [Google Scholar] [CrossRef]
- Ventola, C.L. The Antibiotic Resistance Crisis- Part 1: Causes and Threats. Pharm. Ther. 2015, 40, 277–283. [Google Scholar]
- Donabedian, S.M.; Thal, L.A.; Hershberger, E.; Perri, M.B.; Chow, J.W.; Bartlett, P.; Jones, R.; Joyce, K.; Rossiter, S.; Gay, K.; et al. Molecular characterization of gentamicin-resistant enterococci in the United States: Evidence of spread from animals to humans through food. J. Clin. Microbiol. 2003, 41, 1109–1113. [Google Scholar] [CrossRef]
- Trends in U.S. Antibiotic Use: New Data Needed to Improve Prescribing, Combat Threat of Antibiotic Resistance. Available online: http://www.pewtrusts.org/en/research-and-analysis/issue-briefs/2017/03/trends-in-us-antibiotic-use (accessed on 1 June 2018).
- Halloran, J. The Overuse of Antibiotics in Food Animals Threatens Public Health. Available online: https://consumersunion.org/research/the-overuse-of-antibiotics-in-food-animals-threatens-public-health/ (accessed on 1 June 2018).
- CDC. Investigation Update: Multistate Outbreak of Human Salmonella Heidelberg Infections Linked to Ground Turkey; CDC: Atlanta, GA, USA, 2011.
- The Union of Concerned Scientists. Consumer Reports: How Safe Is That Chicken. Available online: https://www.pbs.org/wgbh/pages/frontline/shows/meat/safe/overview.html (accessed on 1 June 2018).



| States | PA | NY | MD | NM | MN | CA | |
|---|---|---|---|---|---|---|---|
| Antimicrobials | |||||||
| Ampicillin | |||||||
| Streptomycin | |||||||
| Gentamicin | |||||||
| Kanamycin | |||||||
| Cefoxitin | |||||||
| Sulfisoxazole | |||||||
| Tetracycline | |||||||
| Ciprofloxacin | |||||||
| States | aadA | aph(3’’) | aph(3’’)-Ib | aph(6)-I | aph(6)-Id | bla | blaCMY | tet | tet(A) | sul2 |
|---|---|---|---|---|---|---|---|---|---|---|
| PA | ||||||||||
| NY | ||||||||||
| MD | ||||||||||
| NM | ||||||||||
| MN | ||||||||||
| CA |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhang, N.; Liu, E.; Tang, A.; Ye, M.C.; Wang, K.; Jia, Q.; Huang, Z. Data-Driven Analysis of Antimicrobial Resistance in Foodborne Pathogens from Six States within the US. Int. J. Environ. Res. Public Health 2019, 16, 1811. https://doi.org/10.3390/ijerph16101811
Zhang N, Liu E, Tang A, Ye MC, Wang K, Jia Q, Huang Z. Data-Driven Analysis of Antimicrobial Resistance in Foodborne Pathogens from Six States within the US. International Journal of Environmental Research and Public Health. 2019; 16(10):1811. https://doi.org/10.3390/ijerph16101811
Chicago/Turabian StyleZhang, Nina, Emily Liu, Alexander Tang, Martin Cheng Ye, Kevin Wang, Qian Jia, and Zuyi Huang. 2019. "Data-Driven Analysis of Antimicrobial Resistance in Foodborne Pathogens from Six States within the US" International Journal of Environmental Research and Public Health 16, no. 10: 1811. https://doi.org/10.3390/ijerph16101811
APA StyleZhang, N., Liu, E., Tang, A., Ye, M. C., Wang, K., Jia, Q., & Huang, Z. (2019). Data-Driven Analysis of Antimicrobial Resistance in Foodborne Pathogens from Six States within the US. International Journal of Environmental Research and Public Health, 16(10), 1811. https://doi.org/10.3390/ijerph16101811





