Antibiotic Sensitivity Profiling and Virulence Potential of Campylobacter jejuni Isolates from Estuarine Water in the Eastern Cape Province, South Africa
Abstract
1. Introduction
1.1. Virulence Determinants
1.1.1. Cytolethal Distending Toxin (CDT)
1.1.2. Campylobacter Invasion Antigens (ciaB)
1.1.3. Fibronectin-Binding Protein (cadF)
1.1.4. Sialyltransferases (cstII)
1.1.5. Post Transcriptional Regulator (csrA)
1.1.6. ATP-Dependent Endopeptidase Protease (clpP)
1.1.7. Periplasmic Chaperon (htrB)
1.2. Treatment and Drug Resistance
2. Materials and Methods
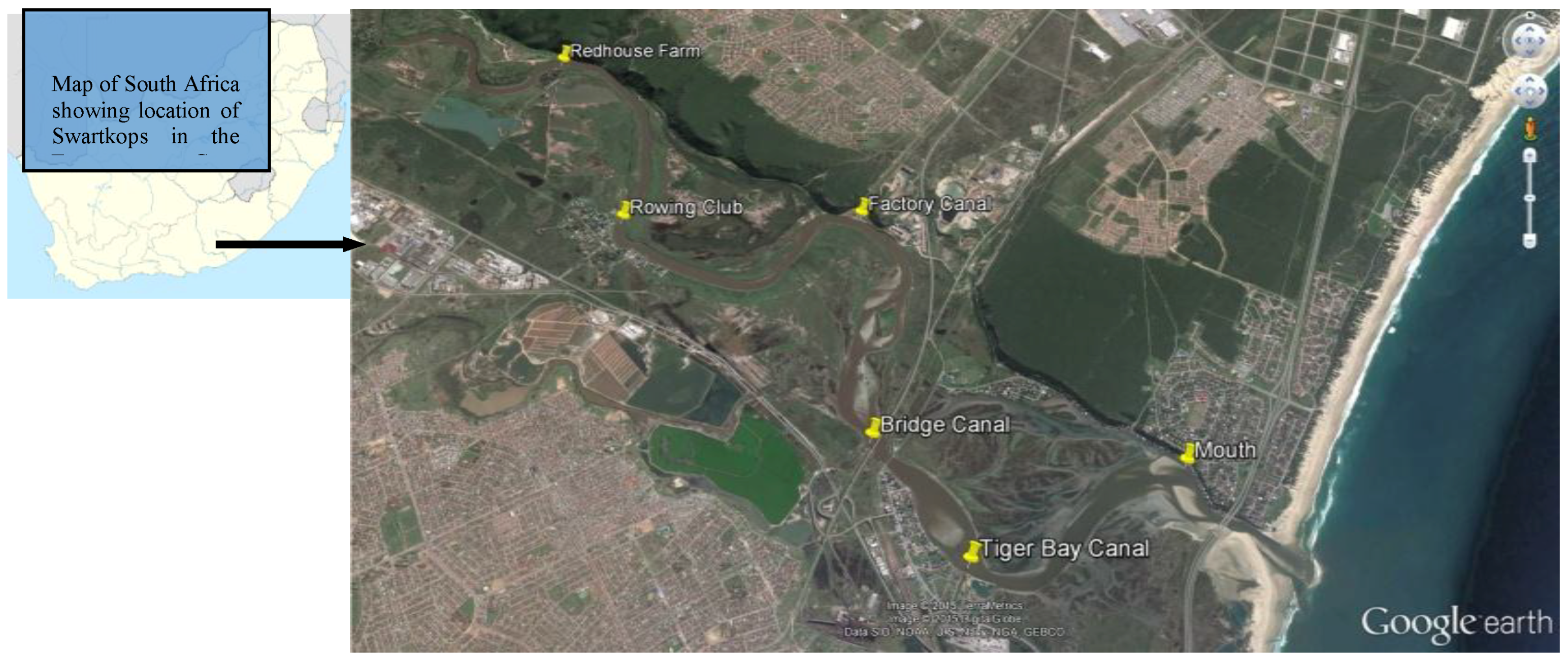
2.1. Study Area
2.2. Sampling and Isolation of CJ
Sampling and Isolation
2.3. PCR Confirmation of CJ and Detection of Virulence Genes
2.4. Antimicrobial Sensitivity Testing
Detection of Multidrug Resistance Genes (cmeA, cmeB and cmeC)
3. Results
3.1. Physicochemical Analyses
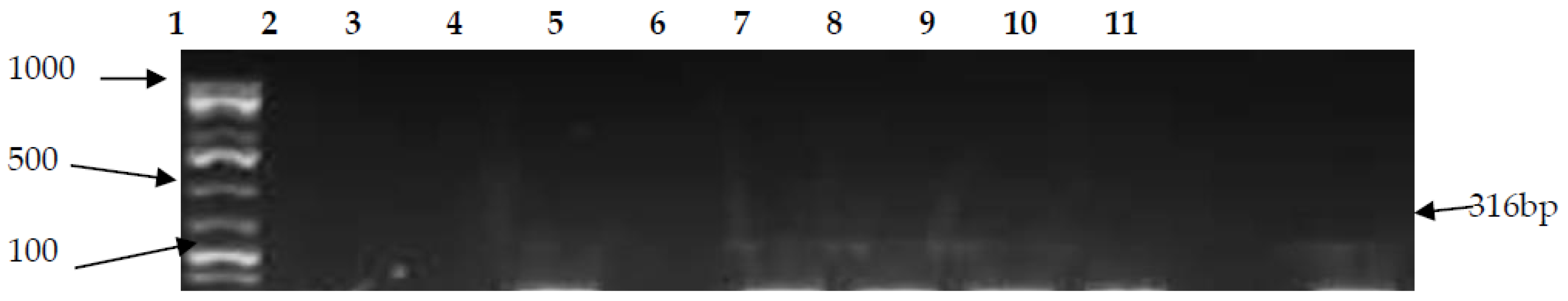
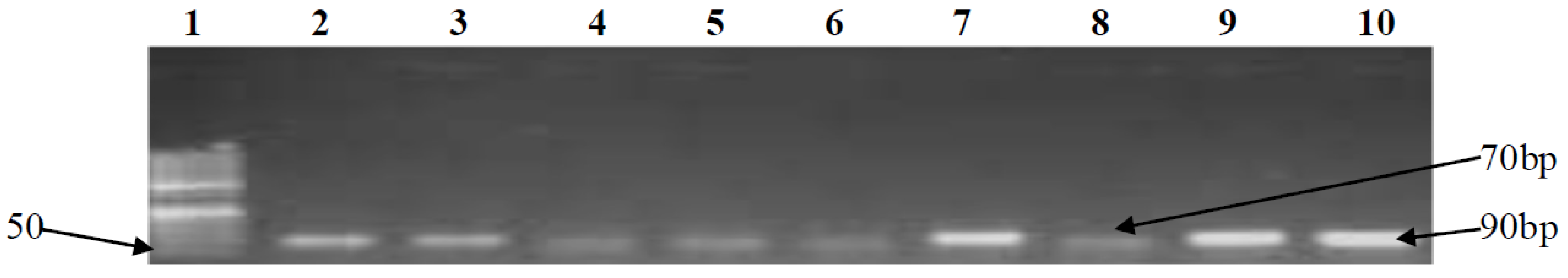
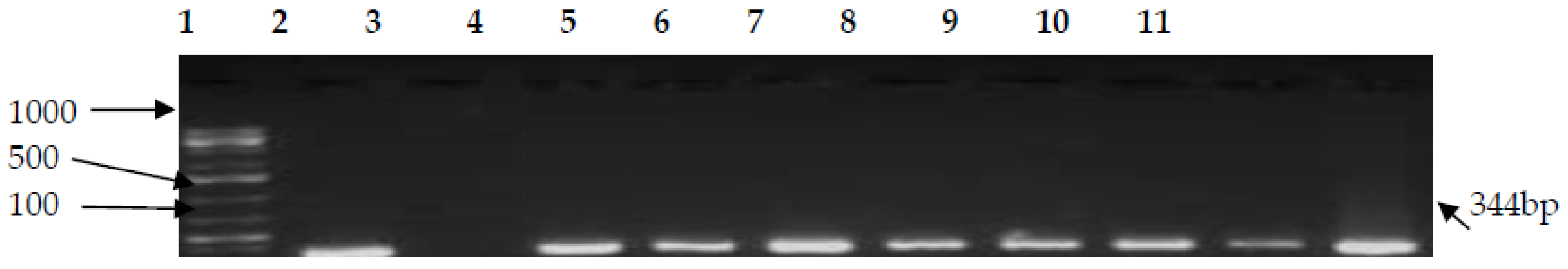
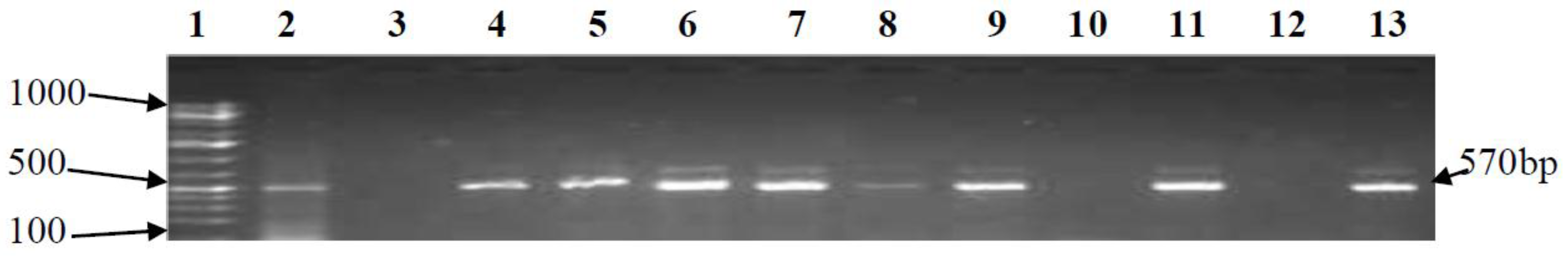
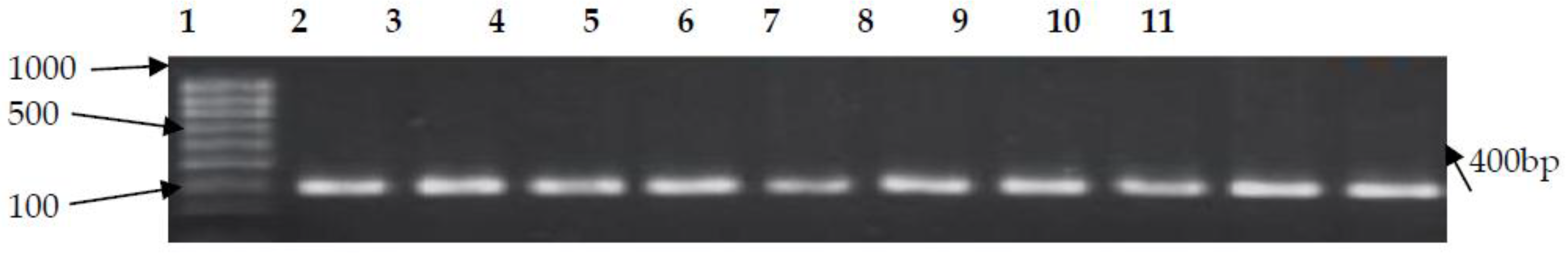
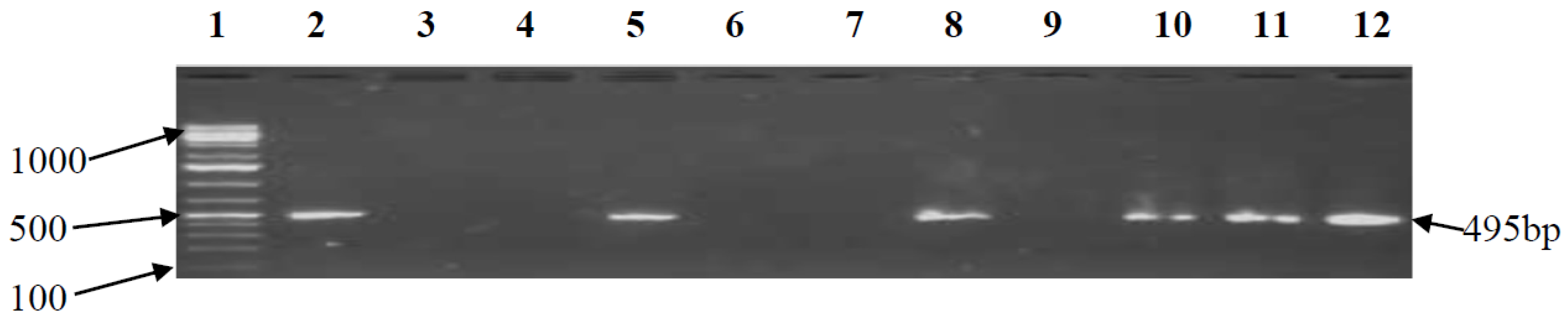
3.2. PCR Confirmation of CJ and Detection of Virulence Genes
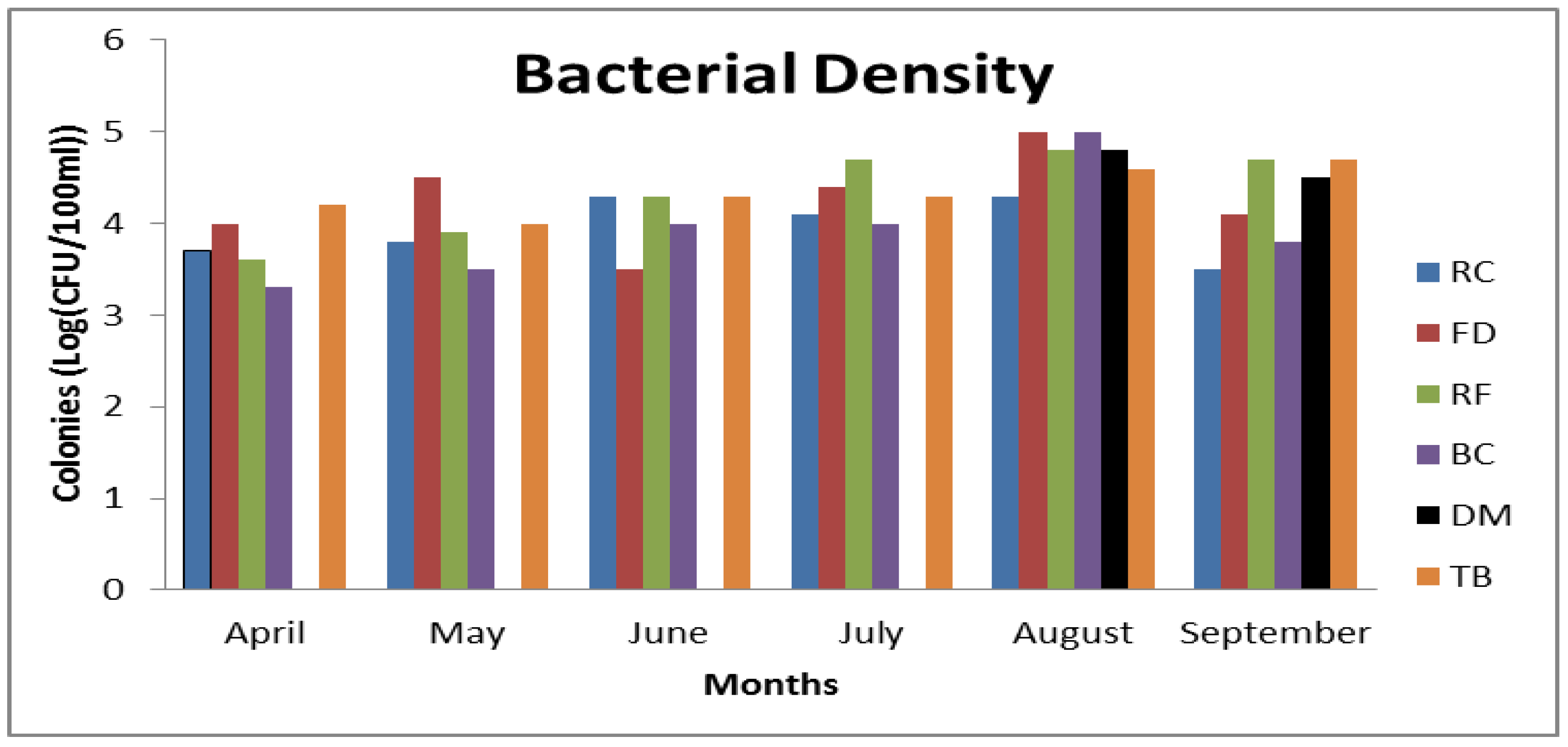
3.3. Frequency of CJ Isolation
3.4. Physicochemical Parameters and Occurrence of C. jejuni
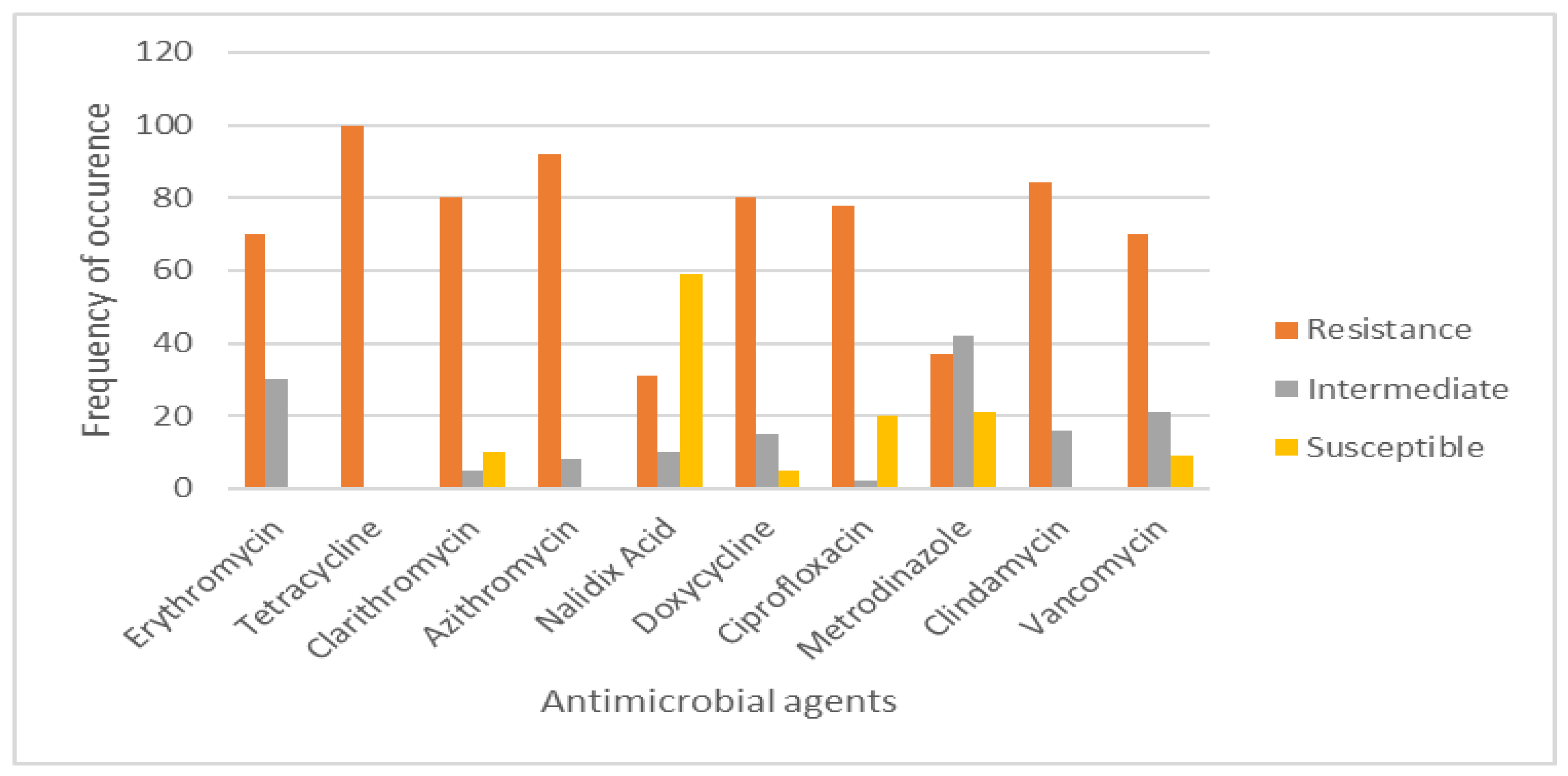
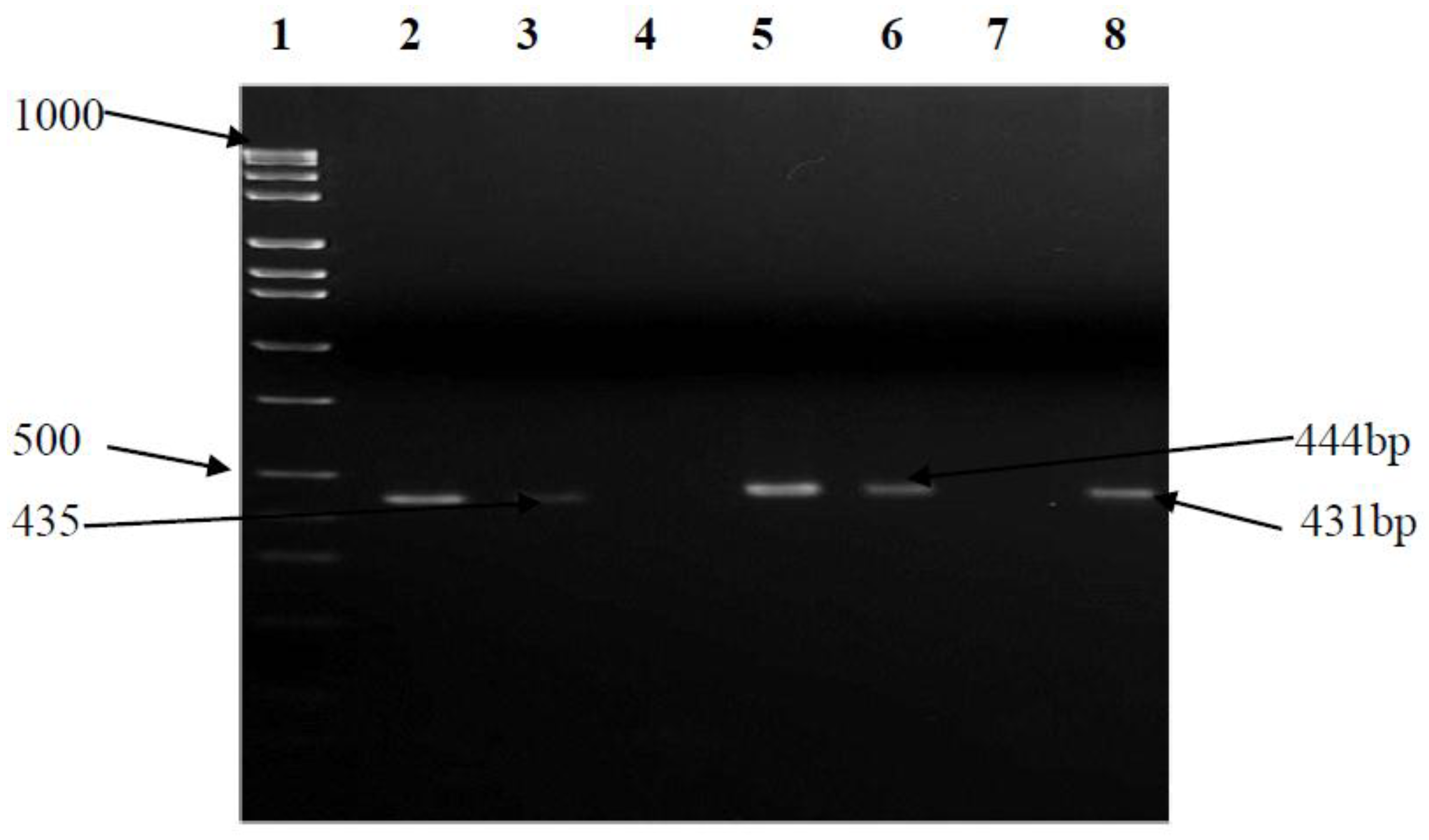
3.5. Antimicrobial Sensitivity Testing and Prevalence of Multidrug Resistance (MDR) Efflux Pump Genes
4. Discussion
4.1. Survival of Organism
4.2. Drug Resistance
5. Conclusions
6. Future Direction of Study
Author Contributions
Acknowledgments
Conflicts of Interest
References
- Bester, L.; Essack, S. Prevalence of antibiotic resistance in Campylobacter isolates from commercial poultry suppliers in KwaZulu-Natal, South Africa. J. Antimicrob. Chemother. 2008, 62, 1298–1300. [Google Scholar] [CrossRef] [PubMed]
- Nichols, G.; Richardson, J.; Sheppard, S.; Lane, K.; Sarran, C. Campylobacter epidemiology: A descriptive study reviewing 1 million cases in England and Wales between 1989 and 2011. BMJ 2012, 2, e001179. [Google Scholar] [CrossRef] [PubMed]
- Croinin, O.; Backert, S. Host epithelial cell invasion by Campylobacter jejuni: Trigger of zipper mechanism? Research Advances in the study of Campylobacter, Helicobacter and related organisms. Front. Cell. Infect. Microbiol. 2012, 2, 25. [Google Scholar] [CrossRef] [PubMed]
- World Health Organization. The Increasing Incidence of Human Campylobacteriosis Report and Proceedings of a WHO Consultation of Experts, Copenhagen, Denmark 21–25 November 2000; World Health Organization: Copenhagen, Denmark; pp. 21–25.
- EFSA (European Food Safety Authority); ECDC (European Centre for Disease Prevention and Control). The European Union summary report on trends and sources of zoonoses, zoonotic agents and food-borne outbreaks in 2012. EFSA J. 2014, 12, 3547. [Google Scholar] [CrossRef]
- Skirrow, M.; Blaser, M. Clinical aspects of Campylobacter infection. In Campylobacter, 2nd ed.; Nachamkin, I., Blaser, M.J., Eds.; ASM Press: Washington, DC, USA, 2000; pp. 69–88. [Google Scholar]
- Engberg, J.; Aarestrup, F.; Taylor, D.; Gerner-Smidt, P.; Nachamkin, I. Quinolone and macrolide resistance in Campylobacter jejuni and C. coli: Resistance mechanisms and trends in human isolates. Emerg. Infect. Dis. 2001, 7, 24–34. [Google Scholar] [CrossRef] [PubMed]
- Sopwith, W.; Birtles, A.; Matthews, M.; Fox, A.; Gee, S.; Painter, M. Campylobacter jejuni multilocus sequence types in humans, northwest England, 2003–2004. Emerg. Infect. Dis. 2006, 12, 1500–1507. [Google Scholar] [CrossRef] [PubMed]
- DefFraites, F.; Sanchez, L.; Brandt, A.; Kadlec, P.; Haberberger, L.; Lin, J.; Taylor, N. An outbreak of Campylobacter enteritis associated with a community water supply on a US military installation. MSMR 2014, 21, 10–15. [Google Scholar]
- Perera, V.; Nachamkin, I.; Ung, H.; Patterson, J.; McConville, M. Molecular mimicry in Campylobacter jejuni: Role of the lipo-oligosaccharide core oligosaccharide in inducing anti-ganglioside antibodies. FEMS Immunol. Med. Microbiol. 2007, 50, 27–36. [Google Scholar] [CrossRef] [PubMed]
- Samosornsuk, W.; Asakura, M.; Yoshida, E.; Taguchi, T.; Eampokalap, B.; Chaicumpa, W.; Yamasaki, S. Isolation and characterization of Campylobacter strains from diarrheal patients in Bangkok and its suburb in Thailand. Jpn. J. Infect. Dis. 2015, 68, 209–215. [Google Scholar] [CrossRef] [PubMed]
- Saouaf, J.; Li, B.; Zhang, G.; Shen, Y.; Furuuchi, N.; Hancock, W.; Greene, I. Deacetylase inhibition increases regulatory T cell function and decreases incidence and severity of collagen-induced arthritis. Exp. Mol. Pathol. 2009, 87, 99–104. [Google Scholar] [CrossRef] [PubMed]
- Sails, A.; Bolton, F.; Fox, A.; Wareing, D.; Greenway, D. Detection of Campylobacter jejuni and Campylobacter coli in environmental waters by PCR enzyme- linked immunosorbent assay. Appl. Environ. Microbiol. 2002, 68, 1319–1324. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Patrone, V.; Campana, R.; Vallorani, L.; Dominici, S.; Federici, S.; Casadei, L.; Gioacchini, A.M.; Stocchi, V.; Baffone, W. CadF expression in Campylobacter jejuni strains incubated under low-temperature water microcosm conditions which induce the viable but non-culturable (VBNC) state. Antonie Van Leeuwenhoek 2013, 103, 979–988. [Google Scholar] [CrossRef] [PubMed]
- Pérez-Boto, D.; López-Portolés, J.; Simón, C.; Valdezate, S.; Echeita, M. Study of the molecular mechanisms involved in high-level macrolide resistance of Spanish Campylobacter jejuni and Campylobacter coli strains. J. Antimicrob. Chemother. 2010, 65, 2083–2088. [Google Scholar]
- Bach, J. Infections and autoimmune diseases. J. Autoimmun. 2005, 25, 74–80. [Google Scholar] [CrossRef] [PubMed]
- Fields, J.; Thompson, S. Campylobacter jejuni CsrA mediates oxidative stress responses, biofilm formation, and host cell invasion. J. Bacteriol. 2008, 190, 3411–3416. [Google Scholar] [CrossRef] [PubMed]
- González-Hein, G.; Huaracán, B.; García, P.; Figueroa, G. Prevalence of virulence genes in strains of Campylobacter jejuni isolated from human, bovine and broiler. Braz. J. Microbiol. 2013, 44, 1223–1229. [Google Scholar]
- Thakur, S.; Zhao, S.; McDermott, P.; Harbottle, H.; Abbott, J.; Gebreyes, W.; White, D. Antimicrobial resistance, virulence, and genotypic profile comparison of Campylobacter jejuni and Campylobacter coli isolated from humans and retail meats. Foodborne Pathog. Dis. 2010, 7, 835–844. [Google Scholar] [CrossRef] [PubMed]
- Hlaváček, O.; Vachova, L. ATP- dependent proteinases in bacteria. Folia Microbiol. 2002, 47, 203–212. [Google Scholar] [CrossRef]
- Frees, D.; Brøndsted, L.; Ingmer, H. Bacterial proteases and virulence. In Regulated Proteolysis in Microorg; Springer Science: Dordrecht, The Netherlands, 2013; pp. 161–192. [Google Scholar]
- Hughes, R.; Cornblath, D. Guillain-Barre´ syndrome. Lancet 2005, 366, 1653–1666. [Google Scholar] [CrossRef]
- Parkhill, J.; Wren, B.W.; Mungall, K.; Ketley, J.M.; Churcher, C.; Basham, D.; Chillingworth, T.; Davies, R.M.; Feltwell, T.; Holroyd, S.; et al. The genome sequence of the food-borne pathogen Campylobacter jejuni reveals hypervariable sequences. Nature 2000, 403, 665–668. [Google Scholar] [CrossRef] [PubMed]
- Hausdorf, L.; Neumann, M.; Bergmann, I.; Sobiella, K.; Mundt, K.; Fröhling, A.; Schlüter, O.; Klocke, M. Occurrence and genetic diversity of Arcobacter spp. in a spinach processing plant and evaluation of two Arcobacter-specific quantitative PCR assays. Syst. Appl. Microbiol. 2013, 36, 235–243. [Google Scholar] [CrossRef] [PubMed]
- Mihaljevic, R.; Sikic, M.; Klancnik, A.; Brumini, G.; Mozina, S.; Abram, M. Environmental stress factors affecting survival and virulence of Campylobacter jejuni. Microb. Pathog. 2007, 43, 120–125. [Google Scholar] [CrossRef] [PubMed]
- Sheppard, S.; Dallas, J.; Strachan, N.; MacRae, M.; McCarthy, N.; Wilson, D.; Gormley, F.J.; Falush, D.; Ogden, I.; Maiden, M.; et al. Campylobacter genotyping to determine the source of human infection. Clin. Infect. Dis. 2009, 48, 1072–1078. [Google Scholar] [CrossRef] [PubMed]
- Koolman, L.; Whyte, P.; Burgess, C.; Bolton, D. Distribution of virulence- associated genes in a selection of Campylobacter isolates. Foodborne Pathol. Dis. 2016, 12, 424–432. [Google Scholar] [CrossRef] [PubMed]
- Igimi, S.; Okada, Y.; Ishiwa, A.; Yamasaki, M.; Morisaki, N.; Kubo, Y.; Asakura, H.; Yamamoto, S. Antimicrobial resistance of Campylobacter: Prevalence and trends in Japan. Food Addit. Contam. 2008, 25, 1080–1083. [Google Scholar] [CrossRef]
- Rożynek, E.; Dzierżanowska-Fangrat, K.; Szczepańska, B.; Wardak, S.; Szych, J.; Konieczny, P.; Albrecht, P.; Dzierżanowska, D. Trends in antimicrobial susceptibility of Campylobacter isolates in Poland (2000–2007). Polskie Tow. Mikrobiol. Pol. Soc. Microbiol. 2009, 58, 111–115. [Google Scholar]
- Jonker, A.; Picard, J. Antimicrobial susceptibility in thermophilic Campylobacter spp. isolated from pigs and chickens in South Africa. J. S. Afr. Vet. Assoc. 2010, 81, 228–236. [Google Scholar] [CrossRef] [PubMed]
- Coker, O.; Adefeso, O. The changing patterns of Campylobacter jejuni/coli in Lagos, Nigeria after ten years. East African Med. J. 1994, 71, 437–440. [Google Scholar]
- Ghunaim, H.; Behnke, J.; Aigha, I.; Sharma, A.; Doiphode, S.; Deshmukh, A.; Abu-Madi, M. Analysis of Resistance to Antimicrobials and Presence of Virulence/Stress Response Genes in Campylobacter Isolates from Patients with Severe Diarrhoea. PLoS ONE 2015, 10, e0119268. [Google Scholar] [CrossRef] [PubMed]
- Nachamkin, I.; Mishu-Allos, B.; Ho, T. Campylobacter species and Guillain-Barre´ Syndrome. J. Clin. Microbiol. 1998, 11, 555–567. [Google Scholar]
- Vacher, S.; Menard, A.; Bernard, E.; Santos, A.; Megraud, F. Detection of mutation associated with macrolide resistance in thermophilic Campylobacter spp. By real-time PCR. Microb. Drug Resist. 2005, 11, 40–47. [Google Scholar] [CrossRef] [PubMed]
- Qin, S.; Wang, Y.; Zhang, Q.; Chen, X.; Shen, Z.; Deng, F. Identification of a novel genomic island conferring resistance to multiple amino glycoside antibiotics in Campylobacter coli. Antimicrob. Agents Chemother. 2012, 56, 5332–5339. [Google Scholar] [CrossRef] [PubMed]
- Iovine, N. Resistance mechanisms in Campylobacter jejuni. Virulence 2013, 4, 230–240. [Google Scholar] [CrossRef] [PubMed]
- Gibreel, A.; Taylor, D.E. Macrolide resistance in Campylobacter jejuni and Campylobacter coli. J. Antimicrob. Chemother. 2006, 58, 243–255. [Google Scholar] [CrossRef] [PubMed]
- Department of Water Affairs. Revision of general authorizations in terms of Section 39 of the National Water Act, 1998 (At No. 36 of 1998); Published under Government Notice 665 in Government Gazette 36820 No. 665; Department of Water Affairs: Cape Town, South Africa, 2013.
- Scharler, M.; Baird, D. A comparison of selected ecosystem attributes of three South African estuaries with different freshwater inflow regimes using network analysis. J. Mar. Syst. 2005, 56, 283–308. [Google Scholar] [CrossRef]
- Baird, D.; Marais, J.; Martin, P. The Swartkops Estuary: Proceedings of a Symposium at the University of Port Elizabeth; Foundation for Research Development: Pretoria, South Africa, 2007; p. 8. [Google Scholar]
- Heip, C.; Herman, P. Major biological processes in European tidal estuaries: A synthesis of the JEEP-92 Project. Hydrobiologia 1995, 311, 1–7. [Google Scholar] [CrossRef]
- EPA. Guidelines for Water Reuse; EPA/625/R-04/108; Environmental Protection Agency, Municipal Support Division Office of Wastewater Management Office of Water: Washington, DC, USA; Agency for International Development: Washington, DC, USA, 2012.
- Vacher, S.; Menard, A.; Bernard, E.; Mégraud, F. PCR-Restriction Fragment Polymorphism analysis for detection of point mutations associated with macrolide resistance in Campylobacter spp. Antimicrob. Agents Chemother. 2003, 47, 1125–1128. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Persson, S.; Olsen, K.E. Multiplex PCR for identification of Campylobacter coli and Campylobacter jejuni from pure cultures and directly on stool samples. J. Med. Microbiol. 2005, 54, 1043–1047. [Google Scholar] [CrossRef] [PubMed]
- Phongsisay, V. Campylobacter jejuni and the Guillain-Barré Syndrome. Ph.D. Thesis, RMIT University, Melbourne, Australia, 2006. [Google Scholar]
- Cohn, M.; Ingmer, H.; Mulholland, F.; Jørgensen, K.; Wells, J.; Brøndsted, L. Contribution of conserved ATP-dependent proteases of Campylobacter jejuni to stress tolerance and virulence. Appl. Environ. Microbiol. 2007, 73, 7803–7813. [Google Scholar] [CrossRef] [PubMed]
- Lin, J.; Overbye Michel, L.; Zhang, Q. CmeABC functions as a multidrug efflux system in Campylobacter jejuni. Antimicrob. Agents Chemother. 2002, 46, 2124–2131. [Google Scholar] [CrossRef] [PubMed]
- CLSI. Methods for Antimicrobial Dilution and Disk Susceptibility Testing of infrequently Isolated or Fastidious Bacteria, Approved Guidelines-2nd ed.; CLSI: Wayne, PA, USA, 2012. [Google Scholar]
- De Vries, S.P.; Gupta, S.; Baig, A.; L’Heureux, J.; Pont, E.; Wolanska, D.P.; Maskell, D.J.; Grant, A.J. Motility defects in Campylobacter jejuni defined gene deletion mutants caused by second-site mutations. Microbiology 2015, 161, 2316–2327. [Google Scholar] [CrossRef] [PubMed]


| Name of Gene | Sequence (5′-3′) | Product (bp) | Annealing Temperature (°C) | References |
|---|---|---|---|---|
| Campy 23S | F-AATTGATGGGGTTAGCATTAGC R-CAACAATGGCTCATATACAACTGG | 316 | 55 | [43] |
| hipO | F-AGAGTTTGATCCTGGCTCAG R-ACGGCTACCTTGTTACGACTT | 344 | 58 | [44] |
| cdtB | F-CAC GGT TAA AAT CCC CTG CT R-GCA CTT GGA ATT TGC AAG GC | 495 | 52 | [18] |
| htrB | F-CGC ACC CAA TTT GAC ATA GAA R-TTT TTA GAG CGC TTA GCA TTT GTC T | 70 | 52 | [45] |
| clpP | F-TCG GAG CAT TTT TGC TTA GTT G R-CTC CAC CTA AAG GTT GAT GAA TCA T | 90 | 52 | [46] |
| csrA | F-CAC AGT CAG TGA AGG TGC TT R-ACT CGC ACA ATC GCT ACT TC | 878 | 52 | [47] |
| cstII | F-CAG CTT TCT ATT GCC CTT GC R-ACA CAT ATA GAC CCC TGA GG | 570 | 52 | [18] |
| cadF | F-TTGAAGGTAATTTAGATATG R-CTAATACCTAAAGTTGAAAC | 400 | 42 | [10] |
| Target Genes | Primer Sequences 5’-3’ | Annealing Temp (°C) | Amplicon Size (bp) |
|---|---|---|---|
| CmeA | F-TAGCGGCGTAATAGTAAATAAAC R-ATAAAGAAATCTGCGTAAATAGGA | 50 | 435 |
| CmeB | F-AGGCGGTTTTGAAATGTATGTT R-TGTGCCGCTGGGAAAAG | 50 | 444 |
| CmeC | F-CAAGTTGGCGCTGTAGGTGAA R-CCCCAATGAAAAATAGGCAGAGTA | 52 | 431 |
| Sample Stations | Parameters | Temp (°C) | pH | Salinity (psu) | DO (mg/L) | Turbidity (NTU) | Conductivity (ms/cm) |
|---|---|---|---|---|---|---|---|
| RC | M | 14.7 | 8.32 | 13.9 | 53.8 | 37.2 | 20.9 |
| Cv | 0.16 | 0.02 | 0.52 | 0.68 | 0.85 | 0.49 | |
| FD | M | 14.8 | 8.33 | 19.2 | 60.5 | 66.9 | 25.7 |
| Cv | 0.16 | 0.01 | 0.4 | 0.67 | 0.74 | 0.43 | |
| RF | M | 14.9 | 8.27 | 14.9 | 55.3 | 48.4 | 22 |
| Cv | 0.17 | 0.01 | 0.15 | 0.7 | 0.78 | 0.13 | |
| BC | M | 14.9 | 8.31 | 21.4 | 56.9 | 31.9 | 26.3 |
| Cv | 0.15 | 0.02 | 0.5 | 0.69 | 0.4 | 0.55 | |
| DM | M | 15.6 | 8.29 | 32.8 | 58.9 | 4.2 | 26.3 |
| Cv | 0.13 | 0.01 | 0.19 | 0.69 | 0.7 | 0.23 | |
| TB | M | 14.9 | 8.36 | 25.4 | 62.9 | 24.2 | 30 |
| Cv | 0.15 | 0.01 | 0.38 | 0.68 | 1.5 | 0.44 |
| Sample Source | No. of Samples | No. of Isolates Confirmed as Campylobacter Genus for Samples (%) | No. of Isolates Confirmed as CJ (%) | Genes Detected in CJ Isolates (% Positive) | |||||
|---|---|---|---|---|---|---|---|---|---|
| Estuarine water | 244 | 23S rRNA | HipO | cdtB | cadF | cstII | csrA | htrB | ClpP |
| 120 (49.2) | 95 (79.2) | 80 (84.2) | 49 (51.6) | 60 (63.2) | 0 (0) | 30 (31.6) | 45 (47.4) | ||
| Temp | PH | Salinity | DO | Turbidity | Conductivity | CJ | |
|---|---|---|---|---|---|---|---|
| Temp | 1 | ||||||
| pH | −0.30 | 1 | |||||
| Salinity | −0.60 | 0.46 | 1 | ||||
| DO | 0.19 | 0.07 | −0.31 | 1 | |||
| Turbidity | −0.08 | −0.03 | −0.12 | −0.43 | 1 | ||
| Conductivity | −0.53 | 0.48 | 0.87 | −0.32 | 0.08 | 1 | |
| CJ | 0.17 | 0.02 | −0.39 | 0.21 | 0.23 | −0.28 | 1 |
| Genes | No. of Isolates (n = 95) | |
|---|---|---|
| a (%) | b (%) | |
| cmeA | 11/20 (55) | 9/20 (45) |
| cmeB | 18/20 (90) | 2/20 (20) |
| cmeC | 14/20 (70) | 7/20 (30) |
| cmeABC | 69/95 (72.6) | 6/95 (6.3) |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Otigbu, A.C.; Clarke, A.M.; Fri, J.; Akanbi, E.O.; Njom, H.A. Antibiotic Sensitivity Profiling and Virulence Potential of Campylobacter jejuni Isolates from Estuarine Water in the Eastern Cape Province, South Africa. Int. J. Environ. Res. Public Health 2018, 15, 925. https://doi.org/10.3390/ijerph15050925
Otigbu AC, Clarke AM, Fri J, Akanbi EO, Njom HA. Antibiotic Sensitivity Profiling and Virulence Potential of Campylobacter jejuni Isolates from Estuarine Water in the Eastern Cape Province, South Africa. International Journal of Environmental Research and Public Health. 2018; 15(5):925. https://doi.org/10.3390/ijerph15050925
Chicago/Turabian StyleOtigbu, Anthony C., Anna M. Clarke, Justine Fri, Emmanuel O. Akanbi, and Henry A. Njom. 2018. "Antibiotic Sensitivity Profiling and Virulence Potential of Campylobacter jejuni Isolates from Estuarine Water in the Eastern Cape Province, South Africa" International Journal of Environmental Research and Public Health 15, no. 5: 925. https://doi.org/10.3390/ijerph15050925
APA StyleOtigbu, A. C., Clarke, A. M., Fri, J., Akanbi, E. O., & Njom, H. A. (2018). Antibiotic Sensitivity Profiling and Virulence Potential of Campylobacter jejuni Isolates from Estuarine Water in the Eastern Cape Province, South Africa. International Journal of Environmental Research and Public Health, 15(5), 925. https://doi.org/10.3390/ijerph15050925





