Aerobic Mesophilic, Coliform, Escherichia coli, and Staphylococcus aureus Counts of Raw Meat from the Formal and Informal Meat Sectors in South Africa
Abstract
1. Introduction
2. Materials and Methods
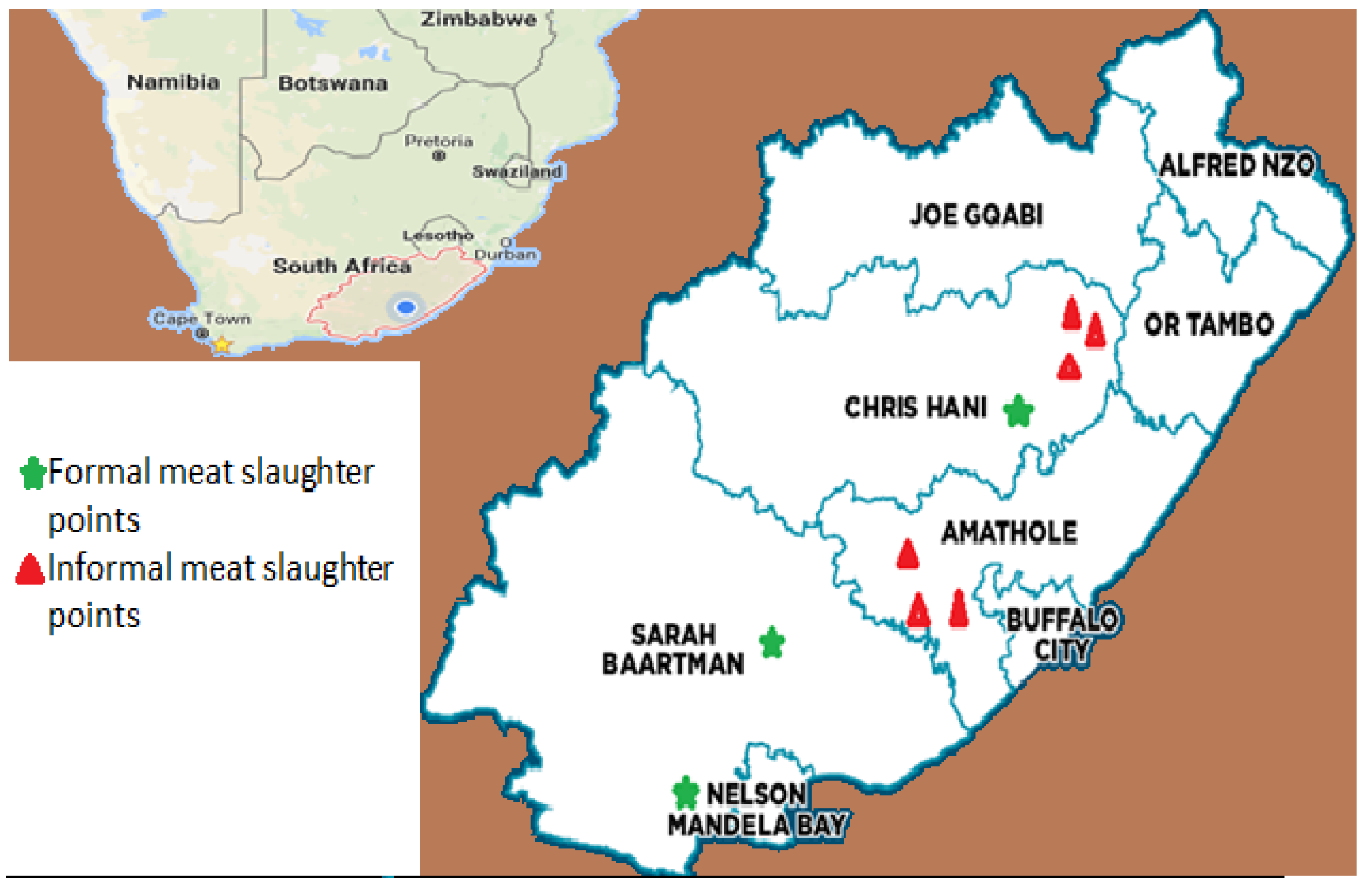
2.1. Sample Collection
2.2. Isolation and Identification of Organisms
2.3. Bacterial DNA Extraction
2.4. Molecular Identification of S. aureus and E. coli Isolates
2.5. Statistical Analysis
2.6. Ethical Approval
3. Results
3.1. Aerobic Colony Counts and Total Coliform Count
3.2. Escherichia coli and Staphylococcus aureus Count
4. Discussion
5. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Angkititrakul, S.; Polpakdee, A.; Chuanchuen, R. Prevalence of Salmonella enterica, Escherichia coli and Staphylococcus aureus in Raw Meat in Thai Self-Service Style Restaurants in Khon Kaen Municipality. Thai J. Vet. Med. 2013, 43, 265–268. [Google Scholar]
- Fayemi, P.O.; Muchenje, V. Meat in African context: From history to science. Afr. J. Biotechnol. 2012, 11, 1298–1306. [Google Scholar] [CrossRef]
- Hagen-Zanker, J.; Morgan, J.; Meth, C. South Africa’s Social Security System: Expanding Coverage of Grants and Limiting Increases in Inequality; Over-Seas Development Institute: London, UK, 2011. [Google Scholar]
- Cawthorn, D.-M.; Steinman, H.A.; Hoffman, L.C. A high incidence of species substitution and mislabelling detected in meat products sold in South Africa. Food Control 2013, 32, 440–449. [Google Scholar] [CrossRef]
- FAO. The State of Food and Agriculture: Livestock in the Balance (Food and Agriculture Organization of the United Nations); FAO: Rome, Italy, 2009; Available online: www.fao.org/docrep/012/i0680e/i0680e.pdf (accessed on 23 September 2016).
- Ahmed, A.M.; Shimamoto, T. Isolation and molecular characterization of Salmonella enterica, Escherichia coli O157:H7 and Shigella spp. from meat and dairy products in Egypt. Int. J. Food Microbiol. 2014, 168–169, 57–62. [Google Scholar] [CrossRef] [PubMed]
- Ali, N.H.; Farooqui, A.; Khan, A.; Khan, A.Y.; Kazmi, S.U. Microbial contamination of raw meat and its environment in retail shops in Karachi, Pakistan. J. Infect. Dev. Ctries. 2010, 4, 382–388. [Google Scholar]
- Kh, H.; Boukhors, K.T.; Dahmani, A.; Zenia, S.; Aissi, M. Survey of hygiene in ovine slaughterhouses of Algiers region by bacteriological analysis of carcasses. Afr. J. Microbiol. Res. 2012, 6, 4722–4726. [Google Scholar]
- Caine, L.-A.; Nwodo, U.U.; Okoh, A.I.; Ndip, R.N.; Green, E. Occurrence of virulence genes associated with diarrheagenic Escherichia coli isolated from raw cow’s milk from two commercial dairy farms in the Eastern Cape Province, South Africa. Int. J. Environ. Res. Public Health 2014, 11, 11950–11963. [Google Scholar] [CrossRef] [PubMed]
- Mhone, T.A.; Matope, G.; Saidi, P.T. Aerobic bacterial, coliform, Escherichia coli and Staphylococcus aureus counts of raw and processed milk from selected smallholder dairy farms of Zimbabwe. Int. J. Food Microbiol. 2011, 151, 223–228. [Google Scholar] [CrossRef] [PubMed]
- Jindal, A.K.; Pandya, K.; Khan, I.D. Antimicrobial resistance: A public health challenge. Med. J. Armed Forces India 2015, 71, 178–181. [Google Scholar] [CrossRef] [PubMed]
- Havelaar, A.; Martyn, D.K.; Paul, R.; Torgerson Herman, J.G.; Tine, H.; Lake, R.J.; Praet, N.; Bellinger, D.C.; de Silva, N.R.; Gargouri, N.; et al. World Health Organization Global Estimates and Regional Comparisons of the Burden of Foodborne Disease in 2010. PLoS Med. 2015, 12, 1–23. [Google Scholar] [CrossRef] [PubMed]
- Casagrande Proietti, P.; Coppola, G.; Bietta, A.; Luisa Marenzoni, M.; Hyatt, D.R.; Coletti, M.; Passamonti, F. Characterization of genes encoding virulence determinants and toxins in Staphylococcus aureus from bovine milk in Central Italy. J. Vet. Med. Sci. 2010, 72, 1443–1448. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Shuiep, E.S.; Kanbar, T.; Eissa, N.; Alber, J.; Lämmler, C.; Zschöck, M.; El Zubeir, I.E.M.; Weiss, R. Phenotypic and genotypic characterization of Staphylococcus aureus isolated from raw camel milk samples. Res. Vet. Sci. 2009, 86, 211–215. [Google Scholar] [CrossRef] [PubMed]
- Jiamboonsri, P.; Pithayanukul, P.; Bavovada, R.; Chomnawang, M.T. The inhibitory potential of thai mango seed kernel extract against methicillin-resistant Staphylococcus aureus. Molecules 2011, 16, 6255–6270. [Google Scholar] [CrossRef] [PubMed]
- Ahmed, A.M.; Shimamoto, T.; Shimamoto, T. Characterization of integrons and resistance genes in multidrug-resistant Salmonella enterica isolated from meat and dairy products in Egypt. Int. J. Food Microbiol. 2014, 189, 39–44. [Google Scholar] [CrossRef] [PubMed]
- Nyamakwere, F.; Muchenje, V.; Mushonga, B.; Makepe, M.; Mutero, G. Assessment of Salmonella, Escherichia coli, Enterobacteriaceae and aerobic colony counts contamination levels during the beef slaughter. J. Food Saf. 2016, 36, 548–556. [Google Scholar] [CrossRef]
- Bolton, D.J.; Pearce, R.A.; Sheridan, J.J.; Blair, I.S.; McDowell, D.A.; Harrington, D. Washing and chilling as critical control points in pork slaughter hazard analysis and critical control point (HACCP) systems. J. Appl. Microbiol. 2002, 92, 893–902. [Google Scholar] [CrossRef] [PubMed]
- Zweifel, C.; Capek, M.; Stephan, R. Microbiological contamination of cattle carcasses at different stages of slaughter in two abattoirs. Meat Sci. 2014, 98, 198–202. [Google Scholar] [CrossRef] [PubMed]
- Wheatley, P.; Giotis, E.S.; McKevitt, A.I. Effects of slaughtering operations on carcass contamination in an Irish pork production plant. Ir. Vet. J. 2014, 67, 1. [Google Scholar] [CrossRef] [PubMed]
- Bello, M.; Lawan, M.K.; Kwaga, J.K.P.; Raji, M.A. Assessment of carcass contamination with E. coli O157 before and after washing with water at abattoirs in Nigeria. Int. J. Food Microbiol. 2011, 150, 184–186. [Google Scholar] [CrossRef] [PubMed]
- Ahmad, M.U.D.; Sarwar, A.; Najeeb, M.I.; Nawaz, M.; Anjum, A.A.; Ali, M.A.; Mansur, N. Assessment of microbial load of raw meat at abattoirs and retail outlets. J. Anim. Plant Sci. 2013, 23, 745–748. [Google Scholar]
- Pathare, N.A.; Asogan, H.; Tejani, S.; Al Mahruqi, G.; Al Fakhri, S.; Zafarulla, R.; Pathare, A.V. Prevalence of methicillin resistant Staphylococcus aureus [MRSA] colonization or carriage among health-care workers. J. Infect. Public Health 2016, 9, 571–576. [Google Scholar] [CrossRef] [PubMed]
- NDVQPH. Standard for the Microbiologival Monitoring of Meat, Process Hygiene and Cleaning; National Directorate for Veterinary Quarantine and Public Health: Pretoria, South Africa, 2010. [Google Scholar]
- Iwu, C.J.; Iweriebor, B.C.; Obi, L.C.; Okoh, A.I. Occurrence of non-O157 Shiga toxin-producing Escherichia coli in two commercial swine farms in the Eastern Cape Province, South Africa. Comp. Immunol. Microbiol. Infect. Dis. 2016, 44, 48–53. [Google Scholar] [CrossRef] [PubMed]
- Graber, H.U.; Pfister, S.; Burgener, P.; Boss, R.; Meylan, M.; Hummerjohann, J. Bovine Staphylococcus aureus: Diagnostic properties of specific media. Res. Vet. Sci. 2013, 95, 38–44. [Google Scholar] [CrossRef] [PubMed]
- Rinttilä, T.; Lyra, A.; Krogius-Kurikka, L.; Palva, A. Real-time PCR analysis of enteric pathogens from fecal samples of irritable bowel syndrome subjects. Gut Pathog. 2011, 3, 6. [Google Scholar] [CrossRef] [PubMed]
- Ali, R.; Al-Achkar, K.; Al-Mariri, A.; Safi, M. Role of Polymerase Chain Reaction (PCR) in the detection of antibiotic-resistant Staphylococcus aureus. Egypt. J. Med. Hum. Genet. 2014, 15, 293–298. [Google Scholar] [CrossRef]
- Saxena, T.; Kaushik, P.; Krishna Mohan, M. Prevalence of E. coli O157:H7 in water sources: An overview on associated diseases, outbreaks and detection methods. Diagn. Microbiol. Infect. Dis. 2015, 82, 249–264. [Google Scholar] [CrossRef] [PubMed]
- Oguttu, J.W.; McCrindle, C.M.E.; Makita, K.; Grace, D. Investigation of the food value chain of ready-to-eat chicken and the associated risk for staphylococcal food poisoning in Tshwane Metropole, South Africa. Food Control 2014, 45, 87–94. [Google Scholar] [CrossRef]
- Grace, D. Food safety in low and middle income countries. Int. J. Environ. Res. Public Health 2015, 12, 10490–10507. [Google Scholar] [CrossRef] [PubMed]
- Doulgeraki, A.I.; Ercolini, D.; Villani, F.; Nychas, G.J.E. Spoilage microbiota associated to the storage of raw meat in different conditions. Int. J. Food Microbiol. 2012, 157, 130–141. [Google Scholar] [CrossRef] [PubMed]
- Hammad, A.M.; Watanabe, W.; Fujii, T.; Shimamoto, T. Occurrence and characteristics of methicillin-resistant and -susceptible Staphylococcus aureus and methicillin-resistant coagulase-negative staphylococci from Japanese retail ready-to-eat raw fish. Int. J. Food Microbiol. 2012, 156, 286–289. [Google Scholar] [CrossRef] [PubMed]
- Hessain, A.M.; Al-Arfaj, A.A.; Zakri, A.M.; El-Jakee, J.K.; Al-Zogibi, O.G.; Hemeg, H.A.; Ibrahim, I.M. Molecular characterization of Escherichia coli O157:H7 recovered from meat and meat products relevant to human health in Riyadh, Saudi Arabia. Saudi J. Biol. Sci. 2015, 22, 725–729. [Google Scholar] [CrossRef] [PubMed]
- Iwu, C.J.; Iweriebor, B.C.; Obi, L.C.; Basson, A.K.; Okoh, A.I. Multidrug-Resistant Salmonella isolates from Swine in the Eastern Cape Province, South Africa. J. Food Prot. 2016, 79, 1234–1239. [Google Scholar] [CrossRef] [PubMed]
- Buncic, S.; Nychas, G.J.; Lee, M.R.F.; Koutsoumanis, K.; Hébraud, M.; Desvaux, M.; Chorianopoulos, N.; Bolton, D.; Blagojevic, B.; Antic, D. Microbial pathogen control in the beef chain: Recent research advances. Meat Sci. 2014, 97, 288–297. [Google Scholar] [CrossRef] [PubMed]
- Jaja, I.F.; Mushonga, B.; Green, E.; Muchenje, V. A Quantitative Assessment of Causes of Bovine Liver Condemnation and Its Implication for Food Security in the Eastern Cape Province South Africa. Sustainability 2017, 9, 736. [Google Scholar] [CrossRef]
- FAO. Expert Consultation on Community-Based Veterinary Public Health Systems. Available online: http://www.fao.org/docrep/016/y5405e/y5405e00.htm (accessed on 13 May 2016).
- Nastasijevic, I.; Mitrovic, R.; Buncic, S. The occurrence of Escherichia coli O157 in/on faeces, carcasses and fresh meats from cattle. Meat Sci. 2009, 82, 101–105. [Google Scholar] [CrossRef] [PubMed]
- Magwedere, K.; Shilangale, R.; Mbulu, R.S.; Hemberger, Y.; Hoffman, L.C.; Dziva, F. Microbiological quality and potential public health risks of export meat from springbok (Antidorcas marsupialis) in Namibia. Meat Sci. 2013, 93, 73–78. [Google Scholar] [CrossRef] [PubMed]
- Thomas, K.M.; McCann, M.S.; Collery, M.M.; Logan, A.; Whyte, P.; McDowell, D.A.; Duffy, G. Tracking verocytotoxigenic Escherichia coli O157, O26, O111, O103 and O145 in Irish cattle. Int. J. Food Microbiol. 2012, 153, 288–296. [Google Scholar] [CrossRef] [PubMed]
- Byrne, B.; Dunne, G.; Lyng, J.; Bolton, D.J. Microbiological carcass sampling methods to achieve compliance with 2001/471/EC and new hygiene regulations. Res. Microbiol. 2005, 156, 104–106. [Google Scholar] [CrossRef] [PubMed]
- Zweifel, C.; Fischer, R.; Stephan, R. Microbiological contamination of pig and cattle carcasses in different small-scale Swiss abattoirs. Meat Sci. 2008, 78, 225–231. [Google Scholar] [CrossRef] [PubMed]
- FAO. National Food Safety Systems in Africa—A Situation Analysis. FAO/WHO Reg. Conf. Food Saf. Africa 2005. Available online: www.fao.org/tempref/docrep/fao/meeting/010/j6122e.pdf (accessed on 25 Febuary 2016).
- Uyttendaele, M.; Franz, E.; Schlüter, O. Food Safety, a Global Challenge. Int. J. Environ. Res. Public Health 2015, 13, 67. [Google Scholar] [CrossRef]
- Fahrion, A.S.; Jamir, L.; Richa, K.; Begum, S.; Rutsa, V.; Ao, S.; Padmakumar, V.P.; Deka, R.P.; Grace, D. Food-safety hazards in the pork chain in Nagaland, north east India: Implications for human health. Int. J. Environ. Res. Public Health 2013, 11, 403–417. [Google Scholar] [CrossRef] [PubMed]
- Thomas, M.K.; Murray, R.; Flockhart, L.; Pintar, K.; Pollari, F.; Fazil, A.; Nesbitt, A.; Marshall, B. Estimates of the Burden of Foodborne Illness in Canada for 30 Specified Pathogens and Unspecified Agents, Circa 2006. Foodborne Pathog. Dis. 2013, 10, 639–648. [Google Scholar] [CrossRef] [PubMed]
- Qekwana, D.; McCrindle, C.; Oguttu, J.; Grace, D. Assessment of the Occupational Health and Food Safety Risks Associated with the Traditional Slaughter and Consumption of Goats in Gauteng, South Africa. Int. J. Environ. Res. Public Health 2017, 14, 420. [Google Scholar] [CrossRef] [PubMed]
- Gohar, A.; Abdeltawab, N.F.; Fahmy, A.; Amin, M.A. Development of safe, effective and immunogenic vaccine candidate for diarrheagenic Escherichia coli main pathotypes in a mouse model. BMC Res. Notes 2016, 9, 80. [Google Scholar] [CrossRef] [PubMed]
- Songe, M.M.; Hang’ombe, B.M.; Knight-Jones, T.J.D.; Grace, D. Antimicrobial resistant enteropathogenic Escherichia coli and Salmonella spp. in houseflies infesting fish in food markets in Zambia. Int. J. Environ. Res. Public Health 2017, 14, 21. [Google Scholar] [CrossRef] [PubMed]
- Fischer Walker, C.L.; Sack, D.; Black, R.E. Etiology of diarrhea in older children, adolescents and adults: A systematic review. PLoS Negl. Trop. Dis. 2010, 4. [Google Scholar] [CrossRef] [PubMed]
- NICD. Foodborne Disease Outbreaks Reported to the National Institute of Communicable Diseases. January–December 2017. Available online: www.nicd.ac.za (accessed on 5 Janurary 2018).
- Kitai, S.; Shimizu, A.; Kawano, J.; Sato, E.; Nakano, C.; Kitagawa, H.; Fujio, K.; Matsumura, K.; Yasuda, R.; Inamoto, T. Prevalence and characterization of Staphylococcus aureus and enterotoxigenic Staphylococcus aureus in retail raw chicken meat throughout Japan. J. Vet. Med. Sci. 2005, 67, 269–274. [Google Scholar] [CrossRef] [PubMed]
- Nel, S.; Lues, J.F.R.; Buys, E.M.; Venter, P. Bacterial populations associated with meat from the deboning room of a high throughput red meat abattoir. Meat Sci. 2004, 66, 667–674. [Google Scholar] [CrossRef]
- Abdalrahman, L.S.; Wells, H.; Fakhr, M.K. Staphylococcus aureus is More Prevalent in Retail Beef Livers than in Pork and other Beef Cuts. Pathogens 2015, 4, 182–198. [Google Scholar] [CrossRef] [PubMed]
- De Melo, C.B.; de Sa, M.E.P.; Sabino, V.M.; de Fatima Boechat-Fernandes, M.; Santiago, M.T.; Schwingel, F.F.; Freitas, C.; Magioli, C.A.; Cabral-Pinto, S.; McManus, C.; et al. Microbiological detection of bacteria in animal products seized in baggage of international air passengers to Brazil. Prev. Vet. Med. 2015, 118, 22–27. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Jans, C.; Merz, A.; Johler, S.; Younan, M.; Tanner, S.A.; Kaindi, D.W.M.; Wangoh, J.; Bonfoh, B.; Meile, L.; Tasara, T. East and West African milk products are reservoirs for human and livestock-associated Staphylococcus aureus. Food Microbiol. 2017, 65, 64–73. [Google Scholar] [CrossRef] [PubMed]
- Kadariya, J.; Smith, T.C.; Thapaliya, D. Staphylococcus aureus and staphylococcal food-borne disease: An ongoing challenge in public health. BioMed Res. Int. 2014, 2014, 827965. [Google Scholar] [CrossRef] [PubMed]
- Stewart, G.C. Staphylococcal Food Poisoning, 3rd ed.; Elsevier Inc.: Columbia, MO, USA, 2017. [Google Scholar]
- Hauge, S.J.; Nafstad, O.; Rotterud, O.J.; Nesbakken, T. The hygienic impact of categorisation of cattle by hide cleanliness in the abattoir. Food Control 2012, 27, 100–107. [Google Scholar] [CrossRef]
- Yilmaz, A.; Gun, H.; Ugur, M.; Turan, N.; Yilmaz, H. Detection and frequency of VT1, VT2 and eaeA genes in Escherichia coli O157 and O157:H7 strains isolated from cattle, cattle carcasses and abattoir environment in Istanbul. Int. J. Food Microbiol. 2006, 106, 213–217. [Google Scholar] [CrossRef] [PubMed]
- Hugas, M.; Tsigarida, E. Pros and cons of carcass decontamination: The role of the European Food Safety Authority. Meat Sci. 2008, 78, 43–52. [Google Scholar] [CrossRef] [PubMed]
- Reid, C.-A.; Avery, S.W.; Hutchison, M.L.; Buncic, S. Evaluation of sampling methods to assess the microbiological status of cattle hides. Food Control 2002, 13, 405–410. [Google Scholar] [CrossRef]
| Gene | Sequence | Product Size (bp) | PCR Conditions | References |
|---|---|---|---|---|
| Nuc gene | F 5′-GCGATTGATGGTGATACGGTT-3′ | 270 | Initial denaturation at 95 °C for 5 min was followed by 37 cycles of amplification (denaturation at 95 °C for 30 s, annealing at 55 °C for 30 s, and extension at 72 °C for 60 s) and ending with a final extension at 72 °C for 10 min. | [27,28] |
| R 5′-AGCCAAGCCTTGAACGAACTAAAGC-3′ | ||||
| UidA gene | F 5′AAAACGGCAAGAAAAAGCAG-3′ | 147 | Initial denaturation at 94 °C for 2 min followed by 25 cycles of denaturation at 94 °C for 1 min, annealing at 58 °C for 1 min, and extension at 72 °C for 1 min, and ended with a final extension at 72 °C for 2 min. Holding was at 4 °C. | [29] |
| R 5′ACGCGTGGTTAACAGTCTTGCG-3′ |
| Meat Sector | Specie | Sampling Point | Number of Carcasses | Log10 Mean Total Bacterial Counts ± SD (log10 cfu/cm2) | |||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| ACC | TCC | E. coli | S. aureus | ||||||||
| n | BW | AW | BW | AW | BW | AW | BW | AW | |||
| HT1 | cattle | Rump | 40 | 4.2 ± 2.2 | 2.7 ± 1.6 | 5.0 ± 2.1 | 4.6 ± 2.5 | 4.1 ± 2.1 | 3.2 ± 2.0 | 3.9 ± 2.4 | 3.2 ± 2.4 |
| Flank | 40 | 2.5 ± 1.5 | 2.1 ± 1.3 | 4.1 ± 2.4 | 3.8 ± 2.5 | 3.6 ± 2.8 | 2.9 ± 1.9 | 4.2 ± 2.6 | 3.8 ± 2.4 | ||
| Brisket | 40 | 2.7 ± 1.5 | 2.6 ± 1.3 | 4.3 ± 2.7 | 4.3 ± 2.7 | 3.7 ± 2.2 | 3.3 ± 1.9 | 3.7 ± 2.2 | 2.8 ± 1.8 | ||
| Neck | 40 | 5.8 ± 2.6 | 4.3 ± 2.5 | 6.3 ± 2.4 | 5.4 ± 3.1 | 5.0 ± 2.7 | 4.2 ± 2.6 | 3.9 ± 2.3 | 3.6 ± 2.3 | ||
| HT2 | Sheep | Perineal | 40 | 3.2 ± 2.4 | 2.9 ± 2.4 | 4.6 ± 2.5 | 5.7 ± 6.9 | 2.8 ± 2.6 | 2.8 ± 2.5 | 4.2 ± 2.6 | 3.4 ± 2.2 |
| Flank | 40 | 2.4 ± 1.6 | 2.3 ± 1.7 | 3.5 ± 2.4 | 3.5 ± 2.3 | 2.5 ± 2.0 | 2.6 ± 1.9 | 4.9 ± 2.7 | 4.0 ± 2.5 | ||
| Brisket | 40 | 2.2 ± 1.5 | 2.3 ± 1.8 | 3.9 ± 2.3 | 3.6 ± 2.2 | 2.1 ± 1.6 | 2.7 ± 2.2 | 4.1 ± 2.4 | 2.9 ± 1.7 | ||
| Neck | 40 | 4.7 ± 2.7 | 4.3 ± 2.5 | 4.5 ± 2.6 | 4.2 ± 2.8 | 3.5 ± 2.3 | 3.2 ± 2.0 | 3.9 ± 1.9 | 3.7 ± 2.3 | ||
| Pig | Ham | 20 | 3.6 ± 1.8 | 3.5 ± 2.7 | 4.8 ± 2.4 | 3.2 ± 1.9 | 2.5 ± 1.5 | 3.0 ± 1.7 | 3.5 ± 1.5 | 2.7 ± 1.5 | |
| Back | 20 | 2.7 ± 1.0 | 2.7 ± 0.9 | 3.1 ± 0.9 | 1.1 ± 1.0 | 3.7 ± 2.0 | 2.9 ± 1.1 | 4.3 ± 2.3 | 3.2 ± 1.7 | ||
| Belly | 20 | 2.9 ± 1.1 | 2.7 ± 1.5 | 3.9 ± 2.2 | 3.0 ± 1.5 | 2.6 ± 1.4 | 3.6 ± 1.6 | 2.9 ± 1.2 | 3.3 ± 1.5 | ||
| Jowl | 20 | 3.7 ± 1.3 | 4.5 ± 1.9 | 4.2 ± 2.0 | 2.8 ± 2.1 | 2.6 ± 1.7 | 2.7 ± 1.1 | 5.3 ± 2.0 | 3.2 ± 1.6 | ||
| Meat Sector | Specie | Sampling Point | Number of Carcasses | Log10 Mean Total Bacterial Counts ± SD (log10 cfu/cm2) | |||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| ACC | TCC | E. coli | S. aureus | ||||||||
| BW | AW | BW | AW | BW | AW | BW | AW | ||||
| INMS | Cattle | Rump | 15 | 6.4 ± 3.6 | 5.2 ± 2.5 | 5.4 ± 3.5 | 4.9 ± 2.6 | 6.5 ± 3.0 | 5.4 ± 2.7 | 5.1 ± 2.2 | 4.3 ± 1.3 |
| Flank | 15 | 6.2 ± 3.1 | 4.8 ± 2.5 | 4.5 ± 2.0 | 3.7 ± 1.9 | 4.7 ± 2.3 | 4.0 ± 3.0 | 5.6 ± 2.4 | 4.8 ± 2.0 | ||
| Brisket | 15 | 6.6 ± 3.1 | 4.8 ± 3.0 | 6.9 ± 3.2 | 4.0 ± 2.2 | 5.3 ± 2.6 | 3.8 ± 2.3 | 5.2 ± 2.5 | 5.1 ± 2.4 | ||
| Neck | 15 | 6.7 ± 3.5 | 5.1 ± 3.3 | 5.3 ± 3.1 | 5.5 ± 2.4 | 5.0 ± 2.6 | 4.3 ± 2.1 | 5.3 ± 2.5 | 4.9 ± 1.4 | ||
| INMS | Sheep | Perineal | 13 | 4.4 ± 3.4 | 4.1 ± 2.8 | 4.7 ± 3.0 | 3.8 ± 2.3 | 6.3 ± 3.5 | 5.7 ± 2.4 | 4.6 ± 1.9 | 4.2 ± 1.3 |
| Flank | 13 | 3.2 ± 1.5 | 2.8 ± 1.2 | 4.7 ± 2.2 | 4.3 ± 1.8 | 4.0 ± 2.3 | 3.0 ± 1.6 | 5.7 ± 1.7 | 4.2 ± 1.5 | ||
| Brisket | 13 | 4.7 ± 1.3 | 4.0 ± 0.9 | 5.6 ± 2.8 | 3.6 ± 1.7 | 4.5 ± 2.7 | 5.0 ± 2.7 | 4.8 ± 2.7 | 4.7 ± 2.1 | ||
| Neck | 13 | 4.4 ± 1.1 | 3.7 ± 0.8 | 4.8 ± 1.5 | 4.8 ± 1.5 | 4.8 ± 2.2 | 5.8 ± 2.9 | 4.5 ± 2.2 | 4.5 ± 0.9 | ||
| Abattoirs | Sampling Points | Sample Size | Presumptive Isolates (%) | Confirmed with PCR (%) |
|---|---|---|---|---|
| HT1 | Cattle | 168 | 109 (56.2) | 32 (16.5) |
| slaughtermen | 12 | 12 (6.2) | 7 (3.6) | |
| Door handle | 4 | 3 (1.5) | 0 (0) | |
| Knife | 8 | 7 (3.6) | 5 (2.6) | |
| Saw | 2 | 1 (0.5) | 0 (0) | |
| Total | 194 | 132 (68) | 44 (22.7) | |
| HT2 | Sheep | 36 | 27 (30.3) | 14 (15.7) |
| Pig | 36 | 21 (23.6) | 8 (9) | |
| slaughtermen | 8 | 8 (9) | 7 (7.9) | |
| Door handle | 2 | 1 (1.1) | 0 (0) | |
| Knife | 5 | 3 (3.4) | 1 (1.1) | |
| Saw | 2 | 0 (0) | 0 (0) | |
| Total | 89 | 60 (67.4) | 30 (33.7) | |
| INF | Cattle | 20 | 16 (16) | 16 (16) |
| Sheep | 52 | 44 (44) | 32 (32) | |
| Slaughter men | 16 | 15 (15) | 15 (15) | |
| Knife | 12 | 8 (8) | 5 (5) | |
| Total | 100 | 83 (83) | 68 (68) |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Jaja, I.F.; Green, E.; Muchenje, V. Aerobic Mesophilic, Coliform, Escherichia coli, and Staphylococcus aureus Counts of Raw Meat from the Formal and Informal Meat Sectors in South Africa. Int. J. Environ. Res. Public Health 2018, 15, 819. https://doi.org/10.3390/ijerph15040819
Jaja IF, Green E, Muchenje V. Aerobic Mesophilic, Coliform, Escherichia coli, and Staphylococcus aureus Counts of Raw Meat from the Formal and Informal Meat Sectors in South Africa. International Journal of Environmental Research and Public Health. 2018; 15(4):819. https://doi.org/10.3390/ijerph15040819
Chicago/Turabian StyleJaja, Ishmael Festus, Ezekiel Green, and Voster Muchenje. 2018. "Aerobic Mesophilic, Coliform, Escherichia coli, and Staphylococcus aureus Counts of Raw Meat from the Formal and Informal Meat Sectors in South Africa" International Journal of Environmental Research and Public Health 15, no. 4: 819. https://doi.org/10.3390/ijerph15040819
APA StyleJaja, I. F., Green, E., & Muchenje, V. (2018). Aerobic Mesophilic, Coliform, Escherichia coli, and Staphylococcus aureus Counts of Raw Meat from the Formal and Informal Meat Sectors in South Africa. International Journal of Environmental Research and Public Health, 15(4), 819. https://doi.org/10.3390/ijerph15040819






