Association between Promoter Methylation of Gene ERCC3 and Benzene Hematotoxicity
Abstract
:1. Introduction
2. Materials and Methods
2.1. Study Population and Biological Sample Collection
2.2. DNA Methylation Analysis
2.3. Target Prediction of Transcription Factors
2.4. Statistical Analysis
3. Results
3.1. Demographic Characteristics and Benzene Exposure of the Study Population
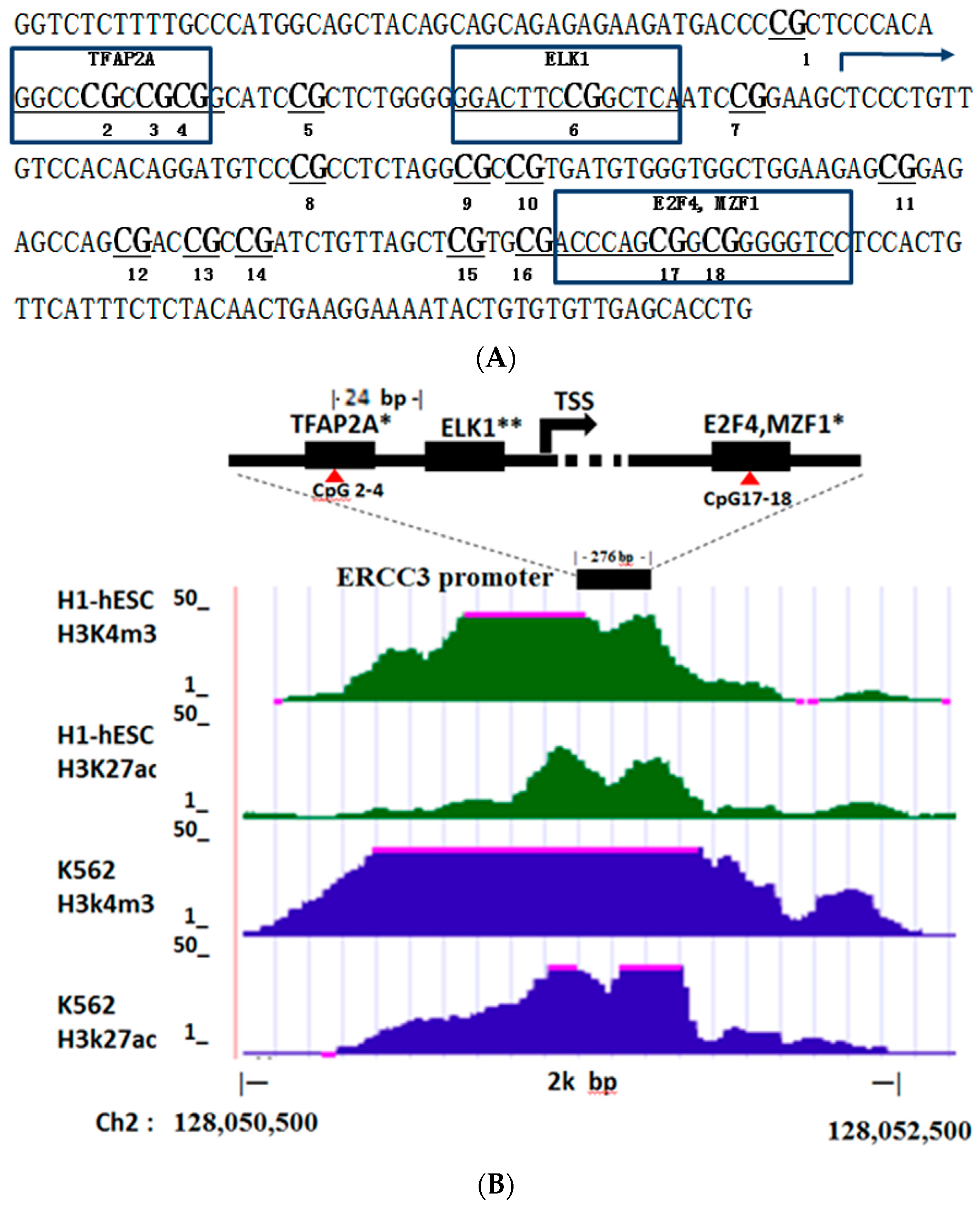
3.2. Methylation of Gene ERCC3
3.3. Correlation between Hematotoxicity and Aberrant CpG Methylation Induced by Benzene
4. Discussion
5. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Yin, S.N.; Hayes, R.B.; Linet, M.S.; Li, G.L.; Dosemeci, M.; Travis, L.B.; Li, C.Y.; Zhang, Z.N.; Li, D.G.; Chow, W.H.; et al. A cohort study of cancer among benzene-exposed workers in China: Overall results. Am. J. Ind. Med. 1996, 29, 227–235. [Google Scholar] [CrossRef]
- Yin, S.N.; Hayes, R.B.; Linet, M.S.; Li, G.L.; Dosemeci, M.; Travis, L.B.; Zhang, Z.N.; Li, D.G.; Chow, W.H.; Wacholder, S.; et al. An expanded cohort study of cancer among benzene-exposed workers in China. Environ. Health Perspect. 1996, 104, 1339–1341. [Google Scholar] [CrossRef] [PubMed]
- Hayes, R.B.; Yin, S.N.; Dosemeci, M.; Li, G.L.; Wacholder, S.; Travis, L.B.; Li, C.Y.; Rothman, N.; Hoover, R.N.; Linet, M.S. Benzene and the dose-related incidence of hematologic neoplasms in China. Chinese Academy of Preventive Medicine—National Cancer Institute Benzene Study Group. J. Natl. Cancer Inst. 1997, 89, 1065–1071. [Google Scholar] [CrossRef] [PubMed]
- Bollati, V.; Baccarelli, A.; Hou, L.; Bonzini, M.; Fustinoni, S.; Cavallo, D.; Byun, H.M.; Jiang, J.; Marinelli, B.; Pesatori, A.C.; et al. Changes in DNA methylation patterns in subjects exposed to low-dose benzene. Cancer Res. 2007, 67, 876–880. [Google Scholar] [CrossRef] [PubMed]
- Xing, C.H.; Wang, Q.F.; Li, B.; Tian, H.Y.; Ni, Y.; Yin, S.N.; Li, G.L. Methylation and expression analysis of tumor suppressor genes p15 and p16 in benzene poisoning. Chem. Biol. Interact. 2010, 184, 306–309. [Google Scholar] [CrossRef] [PubMed]
- Gao, A.; Zuo, X.; Liu, Q.; Lu, X.; Guo, W.; Tian, L. Methylation of PARP-1 promoter involved in the regulation of benzene-induced decrease of PARP-1 mRNA expression. Toxicol. Lett. 2010, 195, 114–118. [Google Scholar] [CrossRef] [PubMed]
- Ji, Z.; Zhang, L.; Peng, V.; Ren, X.; McHale, C.M.; Smith, M.T. A comparison of the cytogenetic alterations and global DNA hypomethylation induced by the benzene metabolite, hydroquinone, with those induced by melphalan and etoposide. Leukemia 2010, 24, 986–991. [Google Scholar] [CrossRef] [PubMed]
- Jaitovich-Groisman, I.; Benlimame, N.; Slagle, B.L.; Perez, M.H.; Alpert, L.; Song, D.J.; Fotouhi-Ardakani, N.; Galipeau, J.; Alaoui-Jamali, M.A. Transcriptional regulation of the TFIIH transcription repair components XPB and XPD by the hepatitis B virus x protein in liver cells and transgenic liver tissue. J. Biol. Chem. 2001, 276, 14124–14132. [Google Scholar] [CrossRef] [PubMed]
- Hosgood, H.D., 3rd; Zhang, L.; Shen, M.; Berndt, S.I.; Vermeulen, R.; Li, G.L.; Yin, S.N.; Yeager, M.; Yuenger, J.; Rothman, N.; et al. Association between genetic variants in VEGF, ERCC3 and occupational benzene haematotoxicity. Occup. Environ. Med. 2009, 66, 848–853. [Google Scholar] [CrossRef] [PubMed]
- Bélanger, A.S.; Tojcic, J.; Harvey, M.; Guillemette, C. Regulation of UGT1A1 and HNF1 transcription factor gene expression by DNA methylation in colon cancer cells. BMC Mol. Biol. 2010, 11, 9. [Google Scholar] [CrossRef] [PubMed]
- Mancini, D.N.; Singh, S.M.; Archer, T.K.; Rodenhiser, D.I. Site-specific DNA methylation in the neurofibromatosis (NF1) promoter interferes with binding of CREB and SP1 transcription factors. Oncogene 1999, 18, 4108–4119. [Google Scholar] [CrossRef] [PubMed]
- Yang, P.; Ma, J.X.; Zhang, B.; Duan, H.W.; He, Z.N.; Zeng, J.L.; Zeng, X.W.; Li, D.C.; Wang, Q.; Xiao, Y.M.; et al. CpG site-specific hypermethylation of p16INK4αin peripheral blood lymphocytes of PAH-exposed workers. Cancer Epidemiol. Biomark. Prev. 2012, 21, 182–190. [Google Scholar] [CrossRef] [PubMed]
- Carter, J.A.; Gorecki, D.C.; Mein, C.A.; Ljungberg, B.; Hafizi, S. CpG dinucleotide-specific hypermethylation of the TNS3 gene promoter in human renal cell carcinoma. Epigenetics 2013, 8, 739–747. [Google Scholar] [CrossRef] [PubMed]
- Xing, C.H.; Chen, Q.; Li, G.L.; Zhang, L.Y.; Zheng, M.; Zou, Z.; Hou, L.; Wang, Q.F.; Liu, X.; Guo, X. Microsomal epoxide hydrolase (EPHX1) polymorphisms are associated with aberrant promoter methylation of ERCC3 and hematotoxicity in benzene-exposed workers. Environ. Mol. Mutagen. 2013, 54, 397–405. [Google Scholar] [CrossRef] [PubMed]
- Rosenbloom, K.R.; Sloan, C.A.; Malladi, V.S.; Dreszer, T.R.; Learned, K.; Kirkup, V.M.; Wong, M.C.; Maddren, M.; Fang, R.; Heitner, S.G.; et al. ENCODE data in the UCSC Genome Browser: Year 5 update. Nuclei. Acids Res. 2013, 41, D56–D63. [Google Scholar] [CrossRef] [PubMed]
- Mathelier, A.; Fornes, O.; Arenillas, D.J.; Chen, C.Y.; Denay, G.; Lee, J.; Shi, W.; Shyr, C.; Tan, G.; Worsley-Hunt, R.; et al. JASPAR 2016: A major expansion and update of the open-access database of transcription factor binding profiles. Nucleic Acids Res. 2016, 44, D110–D115. [Google Scholar] [CrossRef] [PubMed]
- Benjamini, Y.; Hochberg, Y. Controlling the false discovery rate: A practical and powerful approach to multiple testing. J. R. Stat. Soc. Ser. B Methodol. 1995, 57, 289–300. [Google Scholar]
- Ram, O.; Goren, A.; Amit, I.; Shoresh, N.; Yosef, N.; Ernst, J.; Kellis, M.; Gymrek, M.; Issner, R.; Coyne, M.; et al. Combinatorial patterning of chromatin regulators uncovered by genome-wide location analysis in human cells. Cell 2011, 147, 1628–1639. [Google Scholar] [CrossRef] [PubMed]
- Xing, C.H.; Zhang, L.Y.; Zheng, M.; Tian, H.Y.; Zhou, G.; Li, G.L.; Wang, Q.F. Correlation of the down-regulation of tumor suppressor gene p15 and hematotoxicity from benzene exposure. J. Hyg. Res. 2012, 41, 489–492. [Google Scholar]
- Okitsu, C.Y.; Hsieh, C.L. DNA methylation dictates histone H3K4 methylation. Mol. Cell. Biol. 2007, 27, 2746–2757. [Google Scholar] [CrossRef] [PubMed]
- Kuroda, A.; Rauch, T.A.; Todorov, I.; Ku, H.T.; Al-Abdullah, I.H.; Kandeel, F.; Mullen, Y.; Pfeifer, G.P.; Ferreri, K. Insulin gene expression is regulated by DNA methylation. PLoS ONE 2009, 4, e6953. [Google Scholar] [CrossRef]
- Butta, N.; Larrucea, S.; Alonso, S.; Rodriguez, R.B.; Arias-Salgado, E.G.; Ayuso, M.S.; Gonzalez-Manchon, C.; Parrilla, R. Role of transcription factor Sp1 and CpG methylation on the regulation of the human podocalyxin gene promoter. BMC Mol. Biol. 2006, 7, 17. [Google Scholar] [CrossRef] [PubMed]
- Hallberg, A.R.; Vorrink, S.U.; Hudachek, D.R.; Cramer-Morales, K.; Milhem, M.M.; Cornell, R.A.; Domann, F.E. Aberrant CpG methylation of the TFAP2A gene constitutes a mechanism for loss of TFAP2A expression in human metastatic melanoma. Epigenetics 2014, 9, 1641–1647. [Google Scholar] [CrossRef] [PubMed]
- Gaubatz, S.; Lindeman, G.J.; Ishida, S.; Jakoi, L.; Nevins, J.R.; Livingston, D.M.; Rempel, R.E. E2F4 and E2F5 play an essential role in pocket protein-mediated G1 control. Mol. Cell 2000, 6, 729–735. [Google Scholar] [CrossRef]
- Piszczatowski, R.T.; Rafferty, B.J.; Rozado, A.; Parziale, J.V.; Lents, N.H. Myeloid Zinc Finger 1 (MZF-1) Regulates Expression of the CCN2/CTGF and CCN3/NOV Genes in the Hematopoietic Compartment. J. Cell. Physiol. 2015, 230, 2634–2639. [Google Scholar] [CrossRef] [PubMed]
- Domcke, S.; Bardet, A.F.; Adrian Ginno, P.; Hartl, D.; Burger, L.; Schubeler, D. Competition between DNA methylation and transcription factors determines binding of NRF1. Nature 2015, 528, 575–579. [Google Scholar] [CrossRef] [PubMed]
- Akhtar, M.M.; Scala, G.; Cocozza, S.; Miele, G.; Monticelli, A. CpG islands under selective pressure are enriched with H3K4me3, H3K27ac and H3K36me3 histone modifications. BMC Evol. Biol. 2013, 13, 145. [Google Scholar] [CrossRef] [PubMed]
- Compe, E.; Egly, J.M. TFIIH: When transcription met DNA repair, Nature reviews. Nat. Rev. Mol. Cell Biol. 2012, 13, 343–354. [Google Scholar] [CrossRef] [PubMed]
- Vijai, J.; Topka, S.; Villano, D.; Ravichandran, V.; Maxwell, K.N.; Maria, A.; Thomas, T.; Gaddam, P.; Lincoln, A.; Kazzaz, S.; et al. A Recurrent ERCC3 Truncating Mutation Confers Moderate Risk for Breast Cancer. Cancer Discov. 2016, 6, 1267–1275. [Google Scholar] [CrossRef] [PubMed]
- Ministry of Health of the People’s Republic of China. Diagnostic Criteria of Occupational Benzene Poisoning GBZ 68-2013; Ministry of Health of the People’s Republic of China Standards Press of China: Beijing, China, 2013. [Google Scholar]
- Liu, Y.; Aryee, M.J.; Padyukov, L.; Fallin, M.D.; Hesselberg, E.; Runarsson, A.; Reinius, L.; Acevedo, N.; Taub, M.; Ronninger, M.; et al. Epigenome-wide association data implicate DNA methylation as an intermediary of genetic risk in rheumatoid arthritis. Nat. Biotechnol. 2013, 31, 142–147. [Google Scholar] [CrossRef] [PubMed]
- Ly, A.; Hoyt, L.; Crowell, J.; Kim, Y.I. Folate and DNA methylation. Antioxid. Redox Signal. 2012, 17, 302–326. [Google Scholar] [CrossRef] [PubMed]
- Xue, G.; Lu, C.J.; Pan, S.J.; Zhang, Y.L.; Miao, H.; Shan, S.; Zhu, X.T.; Zhang, Y. DNA hypomethylation of CBS promoter induced by folate deficiency is a potential noninvasive circulating biomarker for colorectal adenocarcinomas. Oncotarget 2017, 8, 51387–51401. [Google Scholar] [CrossRef] [PubMed]
- Jacob, R.A. Folate, DNA methylation, and gene expression: Factors of nature and nurture. Am. J. Clin. Nutr. 2000, 72, 903–904. [Google Scholar] [PubMed]
- Stern, L.L.; Mason, J.B.; Selhub, J.; Choi, S.W. Genomic DNA hypomethylation, a characteristic of most cancers, is present in peripheral leukocytes of individuals who are homozygous for the C677T polymorphism in the methylenetetrahydrofolate reductase gene. Cancer Epidemiol. Biomark. Prev. 2000, 9, 849–853. [Google Scholar]
- Friso, S.; Chow, S.W.; Girelli, D.; Mason, J.B.; Dolnikowski, G.G.; Baqley, P.J.; Olivieri, O.; Jacques, P.F.; Rosenberg, I.H.; Corrocher, R.; et al. A common mutation in the 5,10-methylenetetrahydrofolate reductase gene affects genomic DNA methylation through an interaction with folate status. Proc. Natl. Acad. Sci. USA 2002, 99, 5606–5611. [Google Scholar] [CrossRef] [PubMed]
- Van den Donk, M.; Van Enqeland, M.; Pellis, L.; Witteman, B.J.; Kok, F.J.; Keijer, J.; Kampman, E. Dietary folate intake in combination with MTHFR C677T genotype and promoter methylation of tumor suppressor and DNA repair genes in sporadic colorectal adenomas. Cancer Epidemiol. Biomark. Prev. 2007, 16, 327–333. [Google Scholar] [CrossRef] [PubMed]
| CpG Unit a | In Relation to TSS | Control (n = 24) Mean ± SD (Cm%) | Exposure (n = 76) Mean ± SD (Cm%) | Beta | SE | p Value | FDR Value |
|---|---|---|---|---|---|---|---|
| 1 | −62 bp | 3.04 ± 2.69 | 3.87 ± 3.94 | 0.07 | 0.19 | 0.725 | 0.725 |
| 2–4 | −43 bp | 4.92 ± 1.53 | 5.86 ± 2.77 | 0.19 | 0.09 | 0.032 * | 0.176 |
| 5 | −36 bp | 1.38 ± 1.66 | 2.51 ± 3.44 | 0.32 | 0.18 | 0.082 | 0.226 |
| 6 | −19 bp | 1.21 ± 2.02 | 2.74 ± 3.90 | 0.28 | 0.20 | 0.153 | 0.281 |
| 7 | −9 bp | 4.54 ± 3.11 | 4.69 ± 4.03 | 0.09 | 0.18 | 0.626 | 0.689 |
| 9–10 | +30 bp | 6.96 ± 3.13 | 7.74 ± 3.49 | 0.15 | 0.09 | 0.107 | 0.235 |
| 11 | +56 bp | 1.15 ± 1.68 | 2.81 ± 4.36 | 0.21 | 0.38 | 0.576 | 0.689 |
| 13–14 | +71 bp | 6.46 ± 4.96 | 7.47 ± 5.50 | 0.19 | 0.19 | 0.317 | 0.436 |
| 15–16 | +87 bp | 1.54 ± 2.06 | 3.04 ± 3.85 | 0.39 | 0.20 | 0.059 | 0.216 |
| 17–18 | +99 bp | 6.79 ± 2.50 | 8.45 ± 4.09 | 0.24 | 0.11 | 0.024 * | 0.176 |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zheng, M.; Lin, F.; Hou, F.; Li, G.; Zhu, C.; Xu, P.; Xing, C.; Wang, Q. Association between Promoter Methylation of Gene ERCC3 and Benzene Hematotoxicity. Int. J. Environ. Res. Public Health 2017, 14, 921. https://doi.org/10.3390/ijerph14080921
Zheng M, Lin F, Hou F, Li G, Zhu C, Xu P, Xing C, Wang Q. Association between Promoter Methylation of Gene ERCC3 and Benzene Hematotoxicity. International Journal of Environmental Research and Public Health. 2017; 14(8):921. https://doi.org/10.3390/ijerph14080921
Chicago/Turabian StyleZheng, Min, Feiliang Lin, Fenxia Hou, Guilan Li, Caiying Zhu, Peiyu Xu, Caihong Xing, and Qianfei Wang. 2017. "Association between Promoter Methylation of Gene ERCC3 and Benzene Hematotoxicity" International Journal of Environmental Research and Public Health 14, no. 8: 921. https://doi.org/10.3390/ijerph14080921
APA StyleZheng, M., Lin, F., Hou, F., Li, G., Zhu, C., Xu, P., Xing, C., & Wang, Q. (2017). Association between Promoter Methylation of Gene ERCC3 and Benzene Hematotoxicity. International Journal of Environmental Research and Public Health, 14(8), 921. https://doi.org/10.3390/ijerph14080921





