Antimicrobial Resistant Enteropathogenic Escherichia coli and Salmonella spp. in Houseflies Infesting Fish in Food Markets in Zambia
Abstract
:1. Introduction
2. Materials and Methods
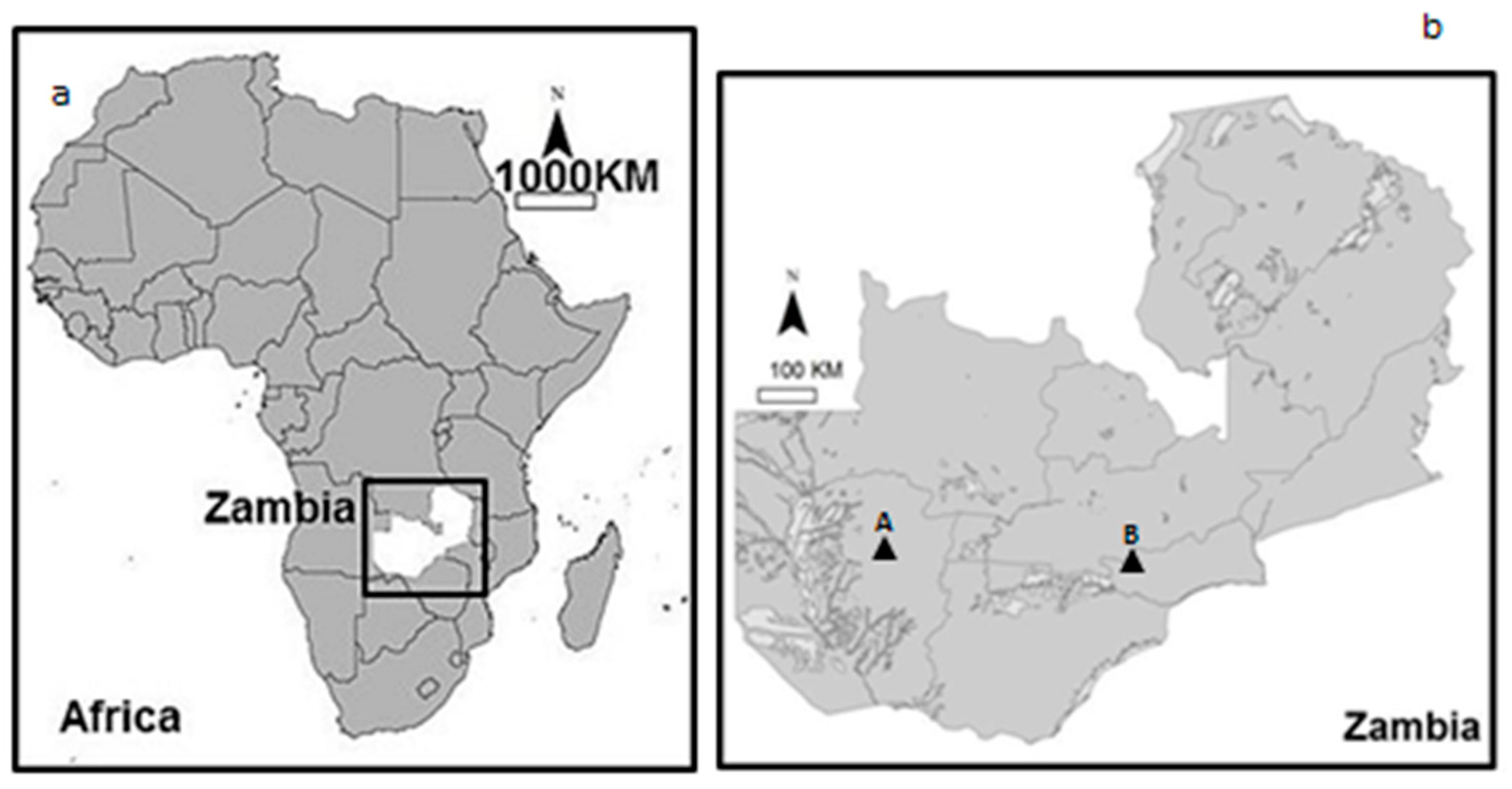
2.1. Site Selection and Semi-Structured Interviews
- (1)
- How the vendor/consumer feels about the presence of high numbers of flies at fish stalls.
- (a)
- Whether or not the vendor has tried some control measures against the flies
- (b)
- The consumer’s view of a fish stall that appears to have some intervention against flies in place.
- (2)
- What interventions those vendors that have had no means of reducing or getting rid of flies would like to try, given a chance.
- (3)
- What the vendors think about covering the fish with netting material to keep flies away from the fish.
2.2. Fly Collection and Sampling
2.3. Culture, Isolation and PCR Detection of Salmonella spp. and Enteropathogenic E. coli
2.4. Determination of Extended Spectrum Beta-Lactamase (ESBL) Producing E. coli and Antibiotic Susceptibility Patterns
3. Results
3.1. Semi-Structured Interviews
3.2. Salmonella spp. and E. coli in Flies
3.3. ESBL and Antimicrobial Susceptibility Patterns of Salmonella spp. and E. coli
4. Discussion
5. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Walker, C.L.; Rudan, I.; Liu, L.; Nair, H.; Theodoratou, E.; Bhutta, Z.A.; O’Brien, K.L.; Campbell, H.; Black, R.E. Global burden of childhood pneumonia and diarrhoea. Lancet 2013, 381, 1405–1416. [Google Scholar] [CrossRef]
- Gohar, A.; Abdeltawab, N.F.; Fahmy, A.; Amin, M.A. Development of safe, effective and immunogenic vaccine candidate for diarrheagenic Escherichia coli main pathotypes in a mouse model. BMC Res. Notes 2016, 9, 80. [Google Scholar] [CrossRef] [PubMed]
- Havelaar, A.H.; Kirk, M.D.; Torgerson, P.R.; Gibb, H.J.; Hald, T.; Lake, R.J.; Praet, N.; Bellinger, D.C.; de Silva, N.R.; Gargouri, N.; et al. World Health Organization global estimates and regional comparisons of the burden of foodborne disease in 2010. PLoS Med. 2015, 12, e1001923. [Google Scholar] [CrossRef] [PubMed]
- Roesel, K.; Grace, D. Food Safety and Informal Markets: Animal Products in Sub-Saharan Africa; Routledge: London, UK, 2014. [Google Scholar]
- Greenberg, B. Flies and Disease. In Biology and Disease Transmission; Princeton University Press: Princeton, NJ, USA, 1973. [Google Scholar]
- Fischer, O.; Mátlová, L.; Dvorská, L.; Švástová, P.; Bartl, J.; Melichárek, I.; Weston, R.T.; Pavlík, I. Diptera as vector of mycobacterial infections in cattle and pigs. Med. Vet. Entomol. 2001, 15, 208–211. [Google Scholar] [CrossRef] [PubMed]
- Olsen, A.R.; Hammack, T.S. Isolation of Salmonella spp. from the housefly, Musca domestica L., and the dump fly, Hydrotaea aenescens (Wiedemann) (Diptera: Muscidae), at caged-layer houses. J. Food Prot. 2000, 63, 958–960. [Google Scholar] [CrossRef] [PubMed]
- Banjo, A.D.; Lawal, O.A.; Adeduji, O.O. Bacteria and fungi isolated from housefly (Musca domestica L.) larvae. Afr. J. Biotechnol. 2005, 4, 780–784. [Google Scholar]
- Damriyasa, I.M.; Failing, K.; Volmer, R.; Zahner, H.; Bauer, C. Prevalence, risk factors and economic importance of infestations with Sarcoptes scabiei and Haematopinus suis in sows of pig breeding farms in Hesse, Germany. Med. Vet. Entomol. 2004, 18, 361–367. [Google Scholar] [CrossRef] [PubMed]
- Forster, M.; Klimpel, S.; Mehlhorn, H.; Sievert, K.; Messler, S.; Pfeffer, K. Pilot study on synanthropic flies (e.g., Musca, Sarcophaga, Calliphora, Fannia, Lucilia, Stomoxys) as vectors of pathogenic microorganisms. Parasitol. Res. 2007, 101, 243–246. [Google Scholar] [CrossRef] [PubMed]
- Forster, M.; Klimpel, S.; Sievert, K. The housefly (Musca domestica) as a potential vector of metazoan parasites caught in a pig-pen in Germany. Vet. Parasitol. 2009, 160, 163–167. [Google Scholar] [CrossRef] [PubMed]
- Gestmann, F.; Förster, M.; Mehlhorn, H.; Sievert, K.; Messler, S.; Neuhausen, N.; Petersdorf, S.; Pfeffer, K. Flies as vectors of microorganisms potentially inducing severe diseases in humans and animals. In Arthropods as Vectors of Emerging Diseases; Mehlhorn, H., Ed.; Springer: Heidelberg, Germany, 2012; pp. 195–226. [Google Scholar]
- Alexander, K.A.; Carzolio, M.; Goodin, D.G.; Vance, E. Climate change is likely to worsen the public health threat of diarrheal disease in Botswana. Int. J. Environ. Res. Public Health 2013, 10, 1202–1230. [Google Scholar] [CrossRef] [PubMed]
- Butaye, P.; Michael, G.; Schwartz, S.; Barett, T.; Brisabois, A.; White, D. The clonal spread of multidrug-resistant non-typhi Salmonella serotypes. Microbes Infect. 2006, 8, 1891–1897. [Google Scholar] [CrossRef] [PubMed]
- Ryan, K.J.; Ray, C.G. Sherris Medical Microbiology: An Introduction to Infectious Diseases, 4th ed.; McGraw Hill: New York, NY, USA, 2004; pp. 362–368. [Google Scholar]
- Nakhjavani, F.A.; Emaneini, M.; Hosseini, H.; Iman-Eini, H.; Aligholi, M.; Jabalameli, F.; Haghi-Ashtiani, M.T.; Taherikalani, M.; Mirsalehian, A. Molecular analysis of typical and atypical enteropathogenic Escherichia coli (EPEC) isolated from children with diarrhea. J. Med. Microbiol. 2013, 62, 191–195. [Google Scholar] [CrossRef] [PubMed]
- Erb, A.; Stürmer, T.; Marre, R.; Brenner, H. Prevalence of antibiotic resistance in Escherichia coli: Overview of geographical, temporal, and methodological variations. Eur. J. Clin. Microbiol. Infect. Dis. 2007, 26, 83–90. [Google Scholar] [CrossRef] [PubMed]
- US Food and Drug Administration. National Antimicrobial Resistance Monitoring System Enteric Bacteria (NARMS): 2008 Executive Report. Rockville (MD), 2010. Available online: http://www.fda.gov/AnimalVeterinary/SafetyHealth/AntimicrobialResistance/NationalAntimicrobialResistanceMonitoringSystem/default.htm (accessed on 13 February 2012). [Google Scholar]
- Holt, P.S.; Geden, C.J.; Moore, R.W.; Gast, R.K. Isolation of Salmonella enterica Serovar Enteritidis from Houseflies (Musca domestica) found in Rooms Containing Salmonella serovar Enteritidis-Challenged Hens. Appl. Environ. Microbiol. 2007, 73, 6030–6035. [Google Scholar] [CrossRef] [PubMed]
- Herigstad, B.; Hamilton, M.; Heersink, J. How to optimize the drop plate method for enumerating bacteria. J. Microbiol. Methods 2001, 44, 121–129. [Google Scholar] [CrossRef]
- Rayamajhi, N.; Kang, G.S.; Lee, Y.D.; Kang, L.M.; Lee, I.S.; Park, Y.K.; Lee, S.H.; Yoo, S.H. Characterization of TEM, SHV and AmpC-type beta-lactamases from cephalosporin-resistant Enterobacteriacea isolated from swine. Int. J. Food Microbiol. 2008, 124, 183–187. [Google Scholar] [CrossRef] [PubMed]
- Batchelor, M.; Hopkins, K.; Threlfal, J.E.; Cliford-Hadley, A.F.; Stallwood, R.; Davies, H.; Liebana, E. blaCTX-M genes in clinical Salmonella isolates recovered from humans in England and Wales from 1992 to 2003. Antimicrob. Agents Chemother. 2005, 49, 1319–1322. [Google Scholar] [CrossRef] [PubMed]
- Kuroda, K.; Suzuki, R.; Ihara, K.; Miyagi, H.; Watanabe, H.; Sato, K.; Hang’ombe, B.M.; Mubita, C.; Isogai, N.; Mulenga, E.; et al. Detection of virulence genes of Escherichia coli and Salmonella spp. from fecal samples of Kafue lechwe (Kobus leche kafuensis) and pastoral cattle in the interface areas of Zambia. Afr. J. Microbiol. Res. 2013, 7, 504–508. [Google Scholar]
- Changkaew, K.; Intarapuk, A.; Utrarachkij, F.; Nakajima, C.; Suthienkul, O.; Suzuki, Y. Antimicrobial resistance, extended spectrum b-lactamase productivity, and class 1 integrons in Escherichia coli from healthy swine. J. Food Prot. 2015, 78, 1443–1450. [Google Scholar] [CrossRef] [PubMed]
- Isogai, E.; Makungu, C.; Yabe, J.; Sinkala, P.; Nambota, A.; Isogai, H.; Fukushi, H.; Silungwe, M.; Mubita, C.; Syakalima, M.; et al. Detection of Salmonella invA by isothermal and chimeric primer-initiated amplification of nucleic acids (ICAN) in Zambia. Comp. Immunol. Microbiol. Infect. Dis. 2005, 28, 363–370. [Google Scholar] [CrossRef] [PubMed]
- Reich, F.; Atanassova, V.; Klein, G. Extended spectrum beta-lactamase and AmpC-producing Enterobacteria in healthy broiler chickens, Germany. Emerg. Infect. Dis. 2013, 19, 1253–1259. [Google Scholar] [CrossRef] [PubMed]
- Chishimba, K.; Hang’ombe, B.M.; Muzandu, K.; Mshana, S.E.; Matee, M.I.; Nakajima, C.; Suzuki, Y. Detection of extended-spectrum beta-lactamase-producing Escherichia coli in market-ready chickens in Zambia. Int. J. Microbiol. 2016, 2016, 5275724. [Google Scholar] [CrossRef] [PubMed]
- Clinical and Laboratory Standards Institute. Performance Standards for Antimicrobial Susceptibility Testing. 19th Informational Supplement M100-S19; Clinical and Laboratory Standards Institute: Wayne, PA, USA, 2009. [Google Scholar]
- Bailey, J.S.; Stern, N.J.; Fedorka-Cray, P.; Craven, S.E.; Cox, N.A.; Cosby, D.E.; Ladely, S.; Musgrove, M.T. Sources and movement of Salmonella through integrated poultry operations: A multistate epidemiological investigation. J. Food Prot. 2001, 64, 1690–1697. [Google Scholar] [CrossRef] [PubMed]
- Hald, B.; Skovgard, H.D.D.; Bang, D.D.; Pedersen, K.; Dybdahl, J.; Jespersen, J.B.; Madsen, M. Flies and Campylobacter infection of broiler flocks. Emerg. Infect. Dis. 2004, 10, 1490–1492. [Google Scholar] [CrossRef] [PubMed]
- Mian, L.S.; Maag, H.; Tacal, V. Isolation of Salmonella from muscoid flies at commercial animal establishments in San Bernardino County, California. J. Vector Ecol. 2002, 27, 82–85. [Google Scholar] [PubMed]
- Rosef, O.; Kapperud, G. Houseflies (Musca domestica) as possible vectors of Campylobacter fetus subsp. jejuni. Appl. Environ. Microbiol. 1983, 45, 381–383. [Google Scholar] [PubMed]
- Wales, A.; Breslin, M.; Davies, R. Assessment of cleaning and disinfection in Salmonella-contaminated poultry layer houses using qualitative and semi-quantitative culture techniques. Vet. Microbiol. 2006, 116, 283–293. [Google Scholar] [CrossRef] [PubMed]
- Sun, Y.; Zeng, Z.; Chen, S.; Ma, J.; He, L.; Liu, Y.; Deng, Y.; Lei, T.; Zhao, J.; Liu, J.H. High prevalence of blaCTX-M extended-spectrum b-lactamase genes in Escherichia coli isolates from pets and emergence of CTX-M-64 in China. Clin. Microbiol. Infect. 2010, 16, 1475–1481. [Google Scholar] [CrossRef] [PubMed]
- Perez, F.; Endimiani, A.; Hujer, K.M.; Bonomo, R.A. The continuing challenge of ESBLs. Curr. Opin. Pharmacol. 2007, 7, 459–469. [Google Scholar] [CrossRef] [PubMed]
- Holmberg, S.D.; Solomon, S.L.; Blake, P.A. Health and Economic Impacts of Antimicrobial Resistance. Rev. Infect. Dis. 1987, 9, 1065–1078. [Google Scholar] [CrossRef] [PubMed]
- Rubin, R.J.; Harrington, C.A.; Poon, A.; Dietrich, K.; Greene, J.A.; Moiduddin, A. The Economic Impact of Staphylococcus aureus in New York City Hospitals. Emerg. Infect. Dis. 1999, 5, 9–17. [Google Scholar] [CrossRef] [PubMed]
- Laxminarayan, R.; Bhutta, Z.; Duse, A.; Jenkins, P.; O’Brien, T.; Okeke, I.N.; Pablo-Mendez, A.; Klugman, K.P. Drug Resistance. In Disease Control Priorities in Developing Countries; Jamison, D.T., Breman, J.G., Measham, A.R., Eds.; World Bank: Washington, DC, USA, 2006. [Google Scholar]
| Total Number of Flies Positive for E. coli | EAST + ve | stx-1 + ve (%) | stx-2 + ve (%) |
|---|---|---|---|
| Lusaka (n = 221) | 72/221 = 32.6% | 12/221 = 5.4% | 23/221 = 10.4% |
| Mongu (n = 86) | 26/86 = 30.2% | 8/86 = 9.3% | 6/86 = 7.0% |
| Gene | Number of Isolates | Percentage (%) |
|---|---|---|
| blaTEM | 18 | 42.9 |
| blaSHV | 15 | 35.7 |
| blaTEM and blaSHV | 9 | 21.4 |
| Total | 42 | 100 |
| Antibiotic | No. of Isolates Showing Resistance | Percent (%) |
|---|---|---|
| Ampicillin (AMP) | 42 | 100 |
| Sulfamethoxazole-trimethoprim (SXT) | 31 | 74.0 |
| Streptomycin (STR) | 37 | 88.1 |
| Tetracycline (TET) | 38 | 91.0 |
| Gentamycin (GEN) | 38 | 91.0 |
| Nalidixic acid (NAL) | 39 | 93.0 |
| Ceftazidime (CAZ) | 40 | 95.2 |
| Chloramphenicol (CHL) | 34 | 81.0 |
| Norfloxacin (NOR) | 38 | 91.0 |
| Ciprofloxacin (CIP) | 40 | 95.2 |
| Cefotaxime (CTX) | 42 | 100 |
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Songe, M.M.; Hang’ombe, B.M.; Knight-Jones, T.J.D.; Grace, D. Antimicrobial Resistant Enteropathogenic Escherichia coli and Salmonella spp. in Houseflies Infesting Fish in Food Markets in Zambia. Int. J. Environ. Res. Public Health 2017, 14, 21. https://doi.org/10.3390/ijerph14010021
Songe MM, Hang’ombe BM, Knight-Jones TJD, Grace D. Antimicrobial Resistant Enteropathogenic Escherichia coli and Salmonella spp. in Houseflies Infesting Fish in Food Markets in Zambia. International Journal of Environmental Research and Public Health. 2017; 14(1):21. https://doi.org/10.3390/ijerph14010021
Chicago/Turabian StyleSonge, Mwansa M., Bernard M. Hang’ombe, Theodore J. D. Knight-Jones, and Delia Grace. 2017. "Antimicrobial Resistant Enteropathogenic Escherichia coli and Salmonella spp. in Houseflies Infesting Fish in Food Markets in Zambia" International Journal of Environmental Research and Public Health 14, no. 1: 21. https://doi.org/10.3390/ijerph14010021
APA StyleSonge, M. M., Hang’ombe, B. M., Knight-Jones, T. J. D., & Grace, D. (2017). Antimicrobial Resistant Enteropathogenic Escherichia coli and Salmonella spp. in Houseflies Infesting Fish in Food Markets in Zambia. International Journal of Environmental Research and Public Health, 14(1), 21. https://doi.org/10.3390/ijerph14010021







