Biodegradation of Methyl tert-Butyl Ether by Co-Metabolism with a Pseudomonas sp. Strain
Abstract
:1. Introduction
2. Experimental Section
2.1. Strains and Media
2.2. Chemicals
2.3. Identification of the Isolated Strain
2.4. The Co-Metabolic Culture Conditions of the Microorganisms
2.5. Kinetic Analysis
2.6. Analytical Method
3. Results and Discussion
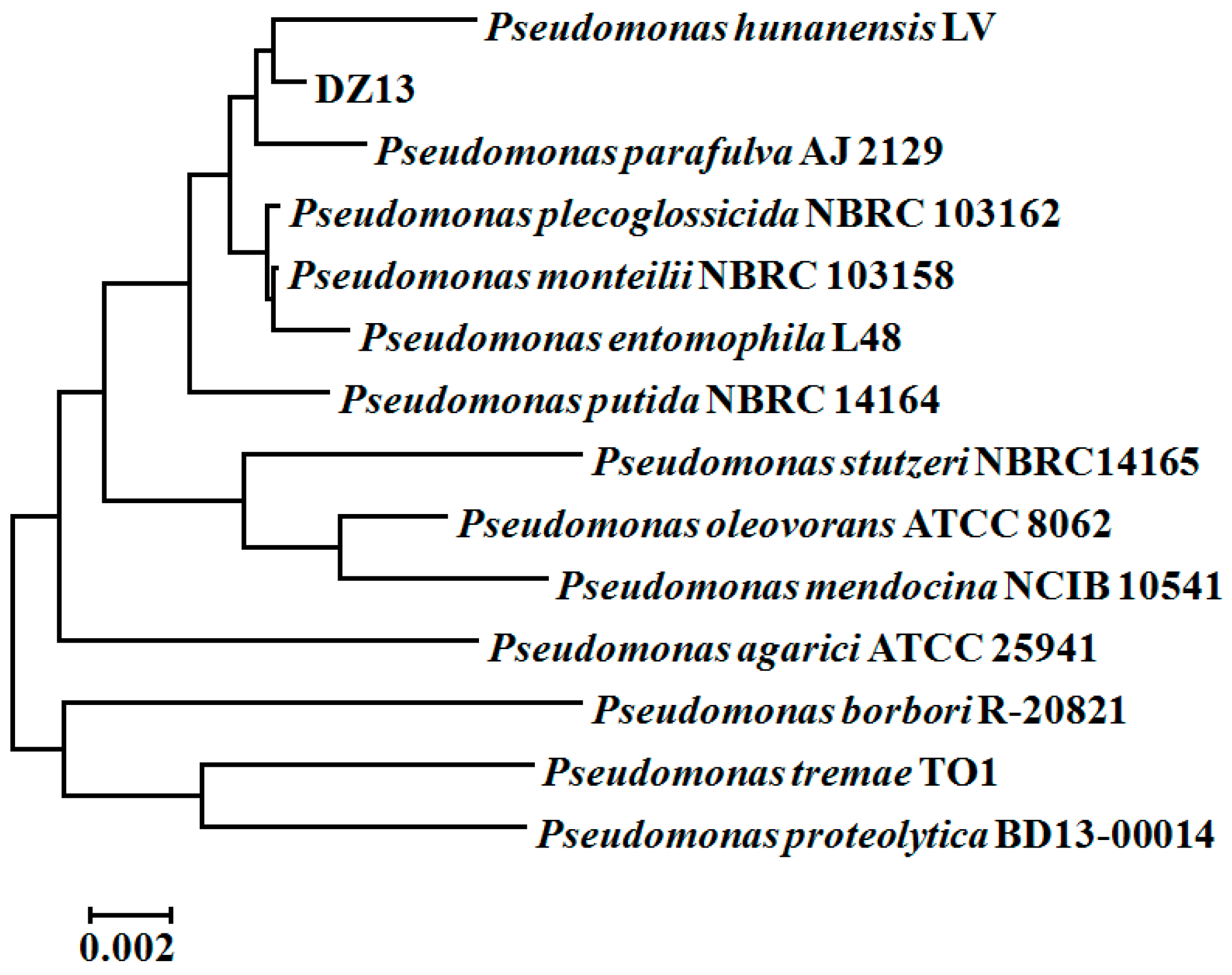
3.1. The Isolation and Identification of MTBE-Cometabolic-Degrading Strain
3.2. Co-Metabolic Degradation of MTBE with Different n-Alkanes as Growth Substrate
3.3. The Kinetic Characteristics of the Co-Metabolism of MTBE by Pseudomonas sp. DZ13 on n-Alkane
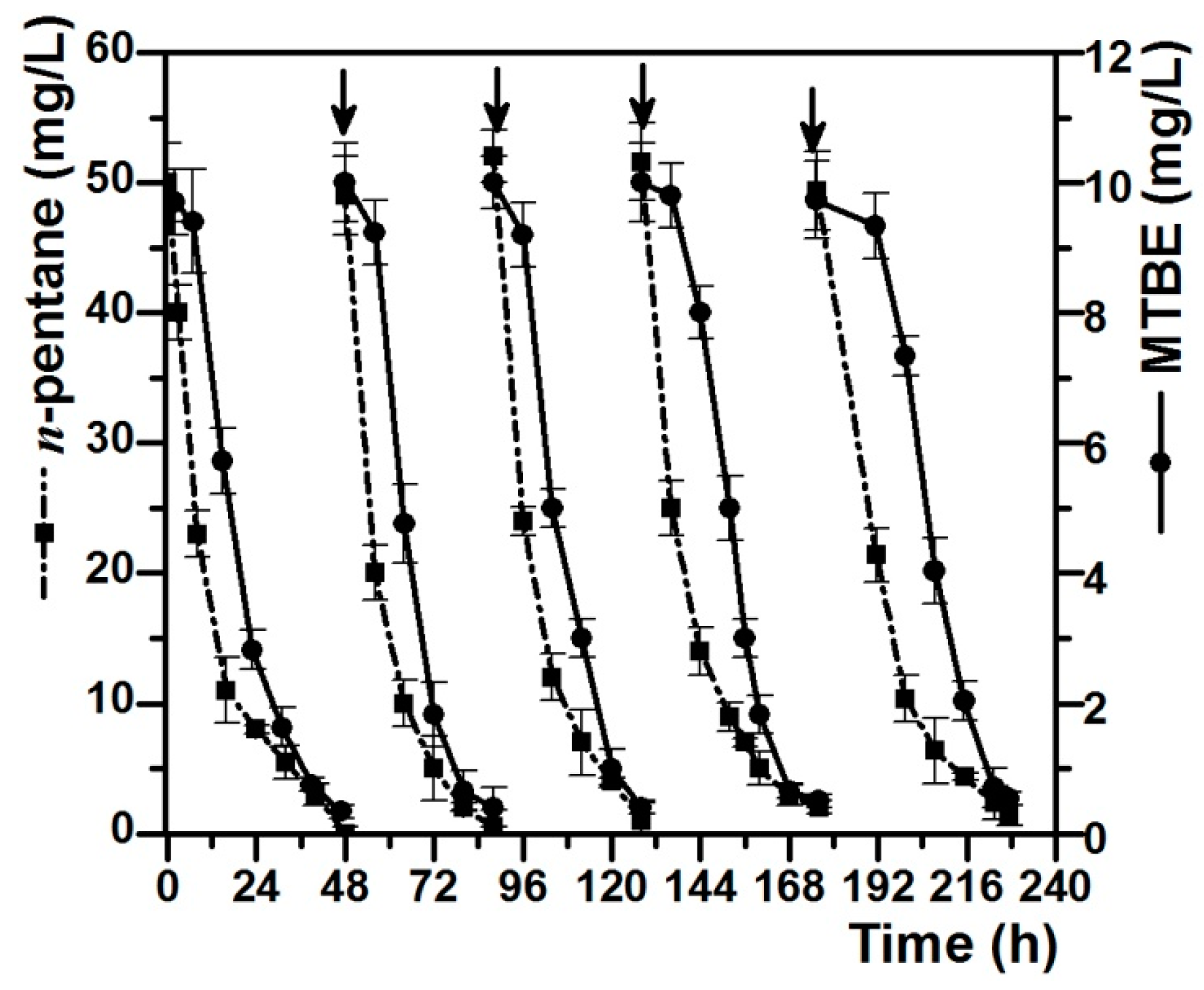
3.4. Degradation of MTBE by Pseudomonas sp. DZ13 Grown on n-Pentane in Continuous Culture
4. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Tran, N.H.; Urase, T.; Ngo, H.H.; Hu, J.Y.; Ong, S.L. Insight into metabolic and cometabolic activities of autotrophic and heterotrophic microorganisms in the biodegradation of emerging trace organic contaminants. Bioresour. Technol. 2013, 146, 721–731. [Google Scholar] [CrossRef] [PubMed]
- Tran, N.H.; Urase, T.; Kusakabe, O. The characteristics of enriched nitrifier culture in the degradation of selected pharmaceutically active compounds. J. Hazard. Mater. 2009, 171, 1051–1057. [Google Scholar] [CrossRef] [PubMed]
- Johnson, R.; Pankow, J.; Bender, D.; Price, C.; Zogorski, J. Peer reviewed: Mtbe-to what extent will past releases contaminate community water supply wells? Environ. Sci. Technol. 2000, 34, 210A–217A. [Google Scholar] [CrossRef] [PubMed]
- Morales, M.; Nava, V.; Velasquez, E.; Razo-Flores, E.; Revah, S. Mineralization of methyl tert-butyl ether and other gasoline oxygenates by pseudomonads using short n-alkanes as growth source. Biodegradation 2009, 20, 271–280. [Google Scholar] [CrossRef] [PubMed]
- Kane, S.R.; Chakicherla, A.Y.; Chain, P.S.; Schmidt, R.; Shin, M.W.; Legler, T.C.; Scow, K.M.; Larimer, F.W.; Lucas, S.M.; Richardson, P.M.; et al. Whole-genome analysis of the methyl tert-butyl ether-degrading beta-proteobacterium methylibium petroleiphilum PM1. J. Bacteriol. 2007, 189, 1931–1945. [Google Scholar] [CrossRef] [PubMed]
- Rosell, M.; Barcelo, D.; Rohwerder, T.; Breuer, U.; Gehre, M.; Richnow, H.H. Variations in 13c/12c and d/h enrichment factors of aerobic bacterial fuel oxygenate degradation. Environ. Sci. Technol. 2007, 41, 2036–2043. [Google Scholar] [CrossRef] [PubMed]
- Streger, S.H.; Vainberg, S.; Dong, H.; Hatzinger, P.B. Enhancing transport of hydrogenophaga flava env735 for bioaugmentation of aquifers contaminated with methyl tert-butyl ether. Appl. Environ. Microbiol. 2002, 68, 5571–5579. [Google Scholar] [CrossRef] [PubMed]
- Salazar, M.; Morales, M.; Revah, S. Biodegradation of methyl tert-butyl ether by cometabolism with hexane in biofilters inoculated with Pseudomonas aeruginosa. J. Environ. Sci. Health A Toxic Hazard. Subst. Environ. Eng. 2012, 47, 1017–1026. [Google Scholar] [CrossRef] [PubMed]
- Steffan, R.J.; Vainberg, S.; Condee, C.W.; McClay, K.; Hatzinger, P.B. Biotreatment of MTBE with a New Bacterial Isolate; Battelle press: Columbus, OH, USA, 2000. [Google Scholar]
- Smith, C.A.; Hyman, M.R. Oxidation of methyl tert-butyl ether by alkane hydroxylase in dicyclopropylketone-induced and n-octane-grown Pseudomonas putida GPo1. Appl. Environ. Microbiol. 2004, 70, 4544–4550. [Google Scholar] [CrossRef] [PubMed]
- Malandain, C.; Fayolle-Guichard, F.; Vogel, T.M. Cytochromes p450-mediated degradation of fuel oxygenates by environmental isolates. FEMS Microbiol. Ecol. 2010, 72, 289–296. [Google Scholar] [CrossRef] [PubMed]
- Chauvaux, S.; Chevalier, F.; Le Dantec, C.; Fayolle, F.; Miras, I.; Kunst, F.; Beguin, P. Cloning of a genetically unstable cytochrome p-450 gene cluster involved in degradation of the pollutant ethyl tert-butyl ether by rhodococcus ruber. J. Bacteriol. 2001, 183, 6551–6557. [Google Scholar] [CrossRef] [PubMed]
- Johnson, E.L.; Hyman, M.R. Propane and n-butane oxidation by Pseudomonas putida gpo1. Appl. Environ. Microbiol. 2006, 72, 950–952. [Google Scholar] [CrossRef] [PubMed]
- Li, S.; Li, D.; Yan, W. Cometabolism of methyl tert-butyl ether by a new microbial consortium ers. Environ. Sci. Pollut. Res. Int. 2015, 22, 10196–10205. [Google Scholar] [CrossRef] [PubMed]
- Pevsner, J. Basic Local Alignment Search Tool (BLAST); John Wiley & Sons, Inc.: Malden, MA, USA, 2005; pp. 87–125. [Google Scholar]
- Kumar, S.; Nei, M.; Dudley, J.; Tamura, K. Mega: A biologist-centric software for evolutionary analysis of DNA and protein sequences. Brief. Bioinform. 2008, 9, 299–306. [Google Scholar] [CrossRef] [PubMed]
- Larkin, M.A.; Blackshields, G.; Brown, N.P.; Chenna, R.; McGettigan, P.A.; McWilliam, H.; Valentin, F.; Wallace, I.M.; Wilm, A.; Lopez, R.; et al. Clustal W and Clustal X version 2.0. Bioinformatics 2007, 23, 2947–2948. [Google Scholar] [CrossRef] [PubMed]
- Peterson, G.L. Review of the folin phenol protein quantitation method of lowry, rosebrough, farr and randall. Anal. Biochem. 1979, 100, 201–220. [Google Scholar] [CrossRef]
- Li, S.; Zhao, H.; Li, Y.; Niu, S.; Cai, B. Complete genome sequence of the naphthalene-degrading Pseudomonas putida strain ND6. J. Bacteriol. 2012, 194, 5154–5155. [Google Scholar] [CrossRef] [PubMed]
- Ferreira, L.; Rosales, E.; Sanroman, M.A.; Pazos, M. Preliminary testing and design of permeable bioreactive barrier for phenanthrene degradation by Pseudomonas stutzeri CECT 930 immobilized in hydrogel matrices. J. Chem. Technol. Biotechnol. 2015, 90, 500–506. [Google Scholar] [CrossRef]
- Hasan, S.A.; Jabeen, S. Degradation kinetics and pathway of phenol by Pseudomonas and bacillus species. Biotechnol. Biotechnol. Equip. 2015, 29, 45–53. [Google Scholar] [CrossRef] [PubMed]
- Vallon, T.; Simon, O.; Rendgen-Heugle, B.; Frana, S.; Muckschel, B.; Broicher, A.; Siemann-Herzberg, M.; Pfannenstiel, J.; Hauer, B.; Huber, A.; et al. Applying systems biology tools to study n-butanol degradation in Pseudomonas putida KT2440. Eng. Life Sci. 2015, 15, 760–771. [Google Scholar] [CrossRef]
- Wang, X.Q.; Wu, C.; Liu, N.; Li, S.J.; Li, W.; Chen, J.M.; Chen, D.Z. Degradation of ethyl mercaptan and its major intermediate diethyl disulfide by Pseudomonas sp. strain WL2. Appl. Microbiol. Biotechnol. 2015, 99, 3211–3220. [Google Scholar] [CrossRef] [PubMed]
- Elango, V.; Kurtz, H.D.; Freedman, D.L. Aerobic cometabolism of trichloroethene and cis-dichloroethene with benzene and chlorinated benzenes as growth substrates. Chemosphere 2011, 84, 247–253. [Google Scholar] [CrossRef] [PubMed]
- Arp, D.J.; Yeager, C.M.; Hyman, M.R. Molecular and cellular fundamentals of aerobic cometabolism of trichloroethylene. Biodegradation 2001, 12, 81–103. [Google Scholar] [CrossRef] [PubMed]
- Jones, M.D.; Rodgers-Vieira, E.A.; Hu, J.; Aitken, M.D. Association of growth substrates and bacterial genera with benzo(a)pyrene mineralization in contaminated soil. Environ. Eng. Sci. 2014, 31, 689–697. [Google Scholar] [CrossRef] [PubMed]
- Hadibarata, T.; Kristanti, R.A.; Fulazzaky, M.A.; Nugroho, A.E. Characterization of pyrene biodegradation by white-rot fungus polyporus sp. S133. Biotechnol. Appl. Biochem. 2012, 59, 465–470. [Google Scholar] [CrossRef] [PubMed]
- Fayolle, F.; Vandecasteele, J.P.; Monot, F. Microbial degradation and fate in the environment of methyl tert-butyl ether and related fuel oxygenates. Appl. Microbiol. Biotechnol. 2001, 56, 339–349. [Google Scholar] [CrossRef] [PubMed]
- Volpe, A.; Del Moro, G.; Rossetti, S.; Tandoi, V.; Lopez, A. Enhanced bioremediation of methyl tert-butyl ether (MTBE) by microbial consortia obtained from contaminated aquifer material. Chemosphere 2009, 75, 149–155. [Google Scholar] [CrossRef] [PubMed]
- Garnier, P.M.; Auria, R. Cometabolic biodegradation of methyl tert-butyl ether by a soil consortium: Effect of components present in gasoline. J. Gen. Appl. Microbiol. 2000, 46, 79–84. [Google Scholar] [CrossRef] [PubMed]
- Smith, C.A.; O’Reilly, K.T.; Hyman, M.R. Cometabolism of methyl tertiary butyl ether and gaseous n-alkanes by Pseudomonas mendocina KR-1 grown on C5 to C8 n-alkanes. Appl. Environ. Microbiol. 2003, 69, 7385–7394. [Google Scholar] [CrossRef] [PubMed]
- Tran, N.H.; Nguyen, V.T.; Urase, T.; Ngo, H.H. Role of nitrification in the biodegradation of selected artificial sweetening agents in biological wastewater treatment process. Bioresour. Technol. 2014, 161, 40–46. [Google Scholar] [CrossRef] [PubMed]
- Sander, R. Compilation of henry’s law constants (version 4.0) for water as solvent. Atmos. Chem. Phys. 2015, 15, 4399–4981. [Google Scholar] [CrossRef]
- Liu, C.Y.; Speitel, G.E.; Georgiou, G. Kinetics of methyl t-butyl ether cometabolism at low concentrations by pure cultures of butane-degrading bacteria. Appl. Environ. Microbiol. 2001, 67, 2197–2201. [Google Scholar] [CrossRef] [PubMed]
- Nava, V.; Morales, M.; Revah, S. Cometabolism of methyl tert-butyl ether (MTBE) with alkanes. Rev. Environ. Sci. Biotechnol. 2007, 6, 339–352. [Google Scholar] [CrossRef]
- Fortin, N.Y.; Morales, M.; Nakagawa, Y.; Focht, D.D.; Deshusses, M.A. Methyl tert-butyl ether (MTBE) degradation by a microbial consortium. Environ. Microbiol. 2001, 3, 407–416. [Google Scholar] [CrossRef] [PubMed]

| Potential Growth Substrates a | OD600 for 5 Days Cultivation b | |||
|---|---|---|---|---|
| DZ2 | DZ6 | DZ9 | DZ13 | |
| n-Alkanes | ||||
| Methane | <0.01 | <0.01 | <0.01 | <0.01 |
| Ethane | <0.01 | <0.01 | <0.01 | <0.01 |
| Propane | <0.01 | <0.01 | <0.01 | <0.01 |
| n-Butane | <0.01 | <0.01 | <0.01 | <0.01 |
| n-Pentane | 0.086 ± 0.003 | 0.132 ± 0.019 | 0.192 ± 0.018 | 0.285 ± 0.013 |
| n-Hexane | 0.124 ± 0.007 | 0.196 ± 0.033 | 0.227 ± 0.024 | 0.238 ± 0.010 |
| n-Heptane | 0.136 ± 0.018 | 0.123 ± 0.006 | 0.175 ± 0.016 | 0.211 ± 0.013 |
| n-Octane | 0.179 ± 0.021 | 0.145 ± 0.013 | 0.184 ± 0.026 | 0.175 ± 0.026 |
| Aromatics | ||||
| Benzene | 0.086 ± 0.011 | 0.045 ± 0.004 | 0.062 ± 0.006 | 0.022 ± 0.008 |
| Toluene | 0.075 ± 0.017 | 0.031 ± 0.008 | 0.095 ± 0.004 | 0.074 ± 0.013 |
| o-Xylene | 0.096 ± 0.006 | 0.051 ± 0.006 | 0.033 ± 0.007 | 0.094 ± 0.006 |
| p-Xylene | 0.114 ± 0.021 | 0.062 ± 0.016 | 0.038 ± 0.009 | 0.088 ± 0.018 |
| m-Xylene | 0.089 ± 0.014 | 0.057 ± 0.012 | 0.045 ± 0.003 | 0.082 ± 0.003 |
| MTBE and its metabolites | ||||
| MTBE | <0.01 | 0.012 ± 0.005 | <0.01 | <0.01 |
| TBA | <0.01 | <0.01 | <0.01 | <0.01 |
| Carbon Source a | Biomass (mgprotein/d) | Degradation Rate of Substrates b (mgsubstrate/gprotein/h) | Degradation Rate of MTBE b (mgMTBE/gprotein/h) | CC (mgMTBE/mgsubstrate) | Residual TBA c (mg/L) |
|---|---|---|---|---|---|
| Aromatic substrate | |||||
| Benzene | 0.3 ± 0.1 | 3.6 ± 0.2 | 0 | - | - |
| Toluene | 8.3 ± 0.7 | 247.3 ± 7.9 | 4.6 ± 0.1 | 0.03 ± 0.01 | 0.12 ± 0.06 |
| Xylene | 9.5 ± 0.5 | 435.8 ± 10.5 | 12.3 ± 0.9 | 0.11 ± 0.06 | 1.34 ± 0.1 |
| n-Alkanes | |||||
| n-Pentane | 15.2 ± 1.1 | 1393.2 ± 78.1 | 178.4 ± 8.4 | 0.72 ± 0.2 | 0.14 ± 0.1 |
| n-Hexane | 14.6 ± 1.4 | 1096.7 ± 56.4 | 130.2 ± 9.2 | 0.56 ± 0.1 | 0.56 ± 0.1 |
| n-Heptane | 12.2 ± 0.4 | 841.7 ± 53.9 | 72.3 ± 7.1 | 0.44 ± 0.1 | 0.67 ± 0.1 |
| n-Octane | 10.9 ± 0.8 | 681.6 ± 45.8 | 59.3 ± 6.9 | 0.26 ± 0.1 | 0.26 ± 0.1 |
| Others | |||||
| MTBE | - | - | - | - | - |
| TBA | - | - | - | - | 9.8 ± 0.3 |
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Li, S.; Wang, S.; Yan, W. Biodegradation of Methyl tert-Butyl Ether by Co-Metabolism with a Pseudomonas sp. Strain. Int. J. Environ. Res. Public Health 2016, 13, 883. https://doi.org/10.3390/ijerph13090883
Li S, Wang S, Yan W. Biodegradation of Methyl tert-Butyl Ether by Co-Metabolism with a Pseudomonas sp. Strain. International Journal of Environmental Research and Public Health. 2016; 13(9):883. https://doi.org/10.3390/ijerph13090883
Chicago/Turabian StyleLi, Shanshan, Shan Wang, and Wei Yan. 2016. "Biodegradation of Methyl tert-Butyl Ether by Co-Metabolism with a Pseudomonas sp. Strain" International Journal of Environmental Research and Public Health 13, no. 9: 883. https://doi.org/10.3390/ijerph13090883
APA StyleLi, S., Wang, S., & Yan, W. (2016). Biodegradation of Methyl tert-Butyl Ether by Co-Metabolism with a Pseudomonas sp. Strain. International Journal of Environmental Research and Public Health, 13(9), 883. https://doi.org/10.3390/ijerph13090883






