Nonylphenol Toxicity Evaluation and Discovery of Biomarkers in Rat Urine by a Metabolomics Strategy through HPLC-QTOF-MS
Abstract
:1. Introduction
2. Materials and Methods
2.1. Animal Treatment
2.2. Monitoring the NP Exposure Levels
2.2.1. Sample Preparation
2.2.2. NP Analysis by HPLC-MS/MS
2.3. Metabolomics Analysis in Urine with HPLC-QTOF-MS
2.3.1. Sample Preparation
2.3.2. HPLC-QTOF-MS Data Acquisition
2.4. Processing and Statistical Analysis of MS Spectra Data
2.4.1. Data Extraction and Normalization
2.4.2. Multivariate Data Analysis
2.5. Identification of Metabolites
2.6. HPLC-MS/MS-Based Validation Test
2.7. Quantitation of the Oxidative Stress Biomarker 8-oxo-deoxyguanosine (8-oxodG) in Urine
3. Results
3.1. Determination of the Level of Exposure to Pesticides
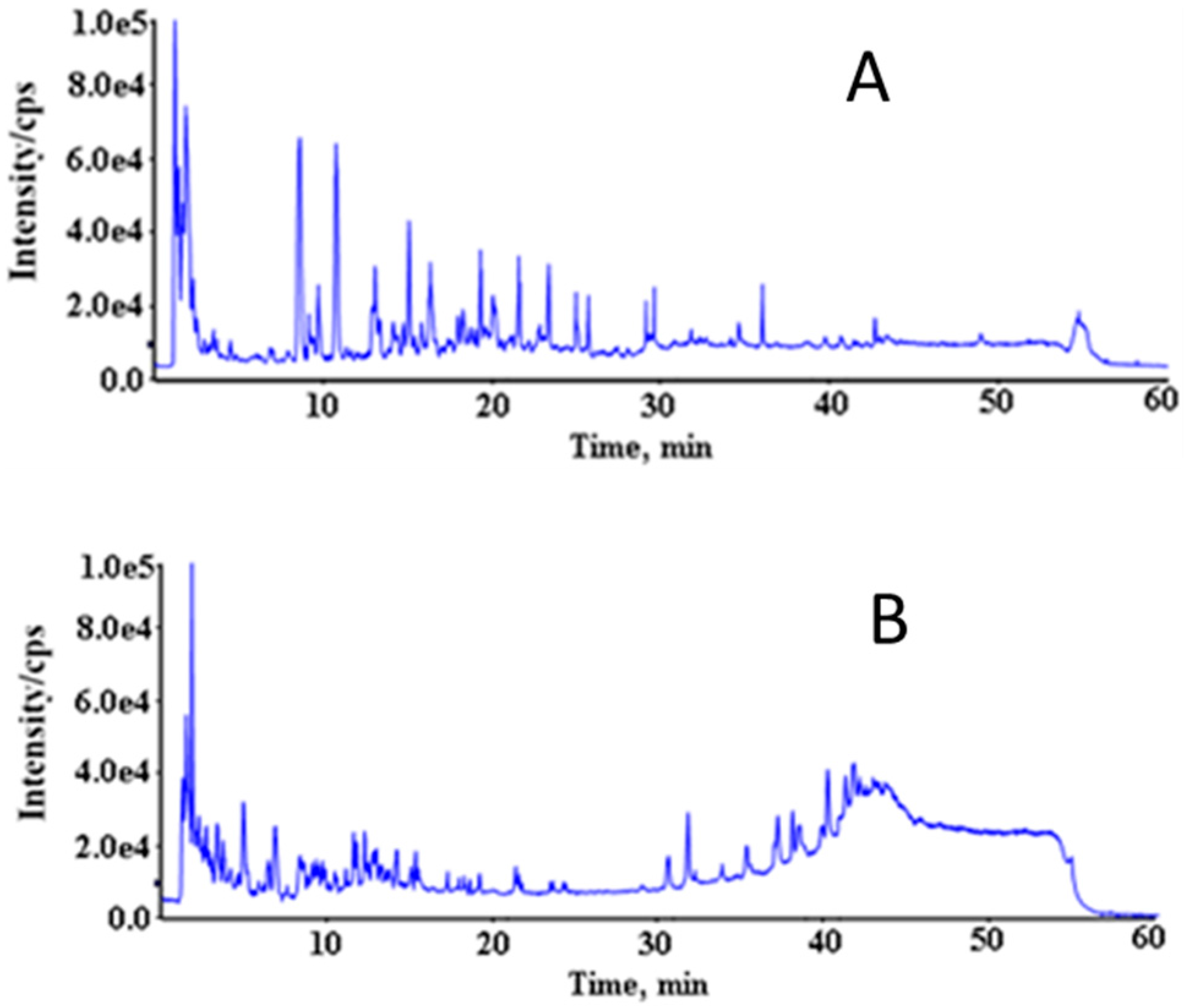
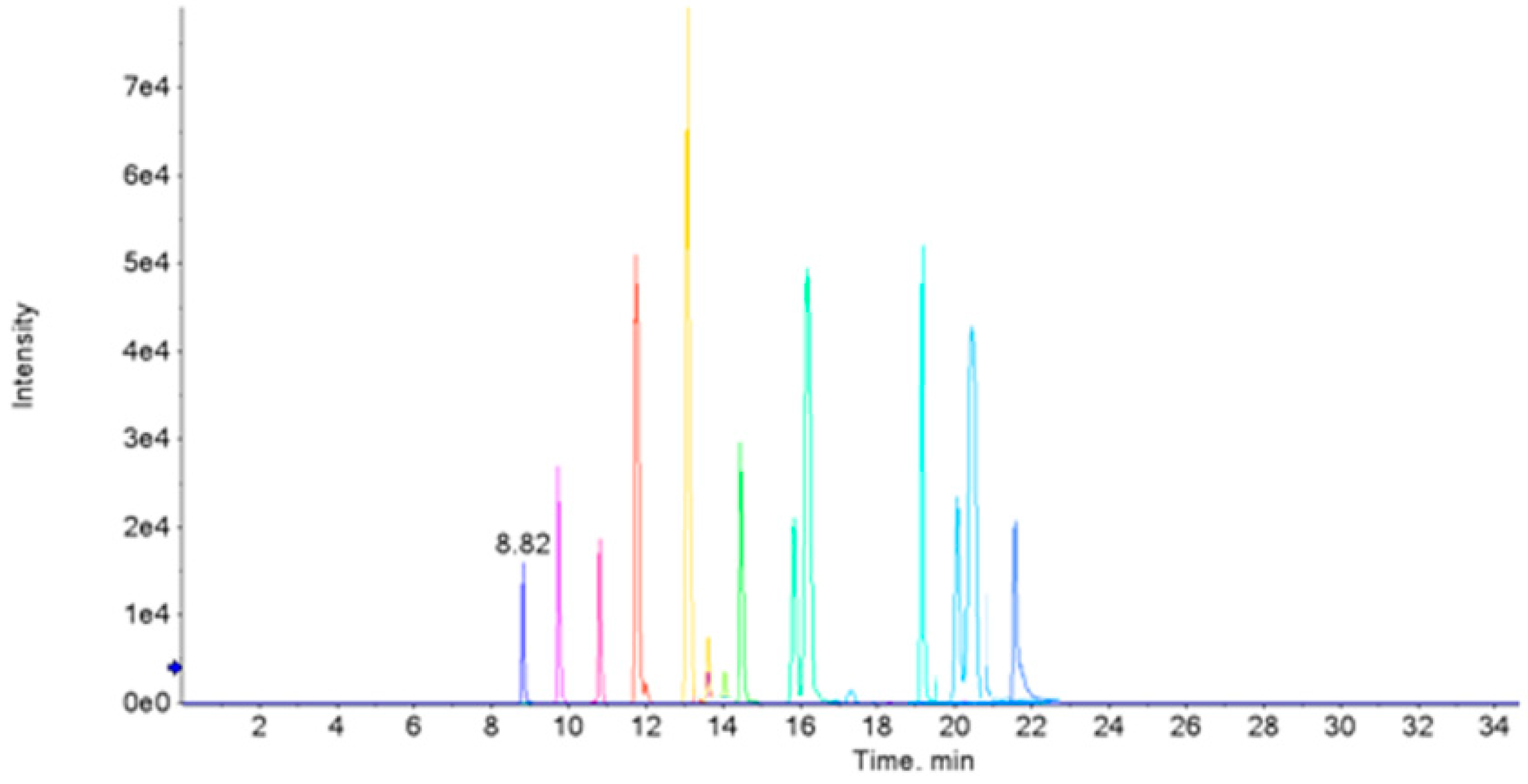
3.2. Acquisition of Mass Spectrum Data
3.3. Dose Selection
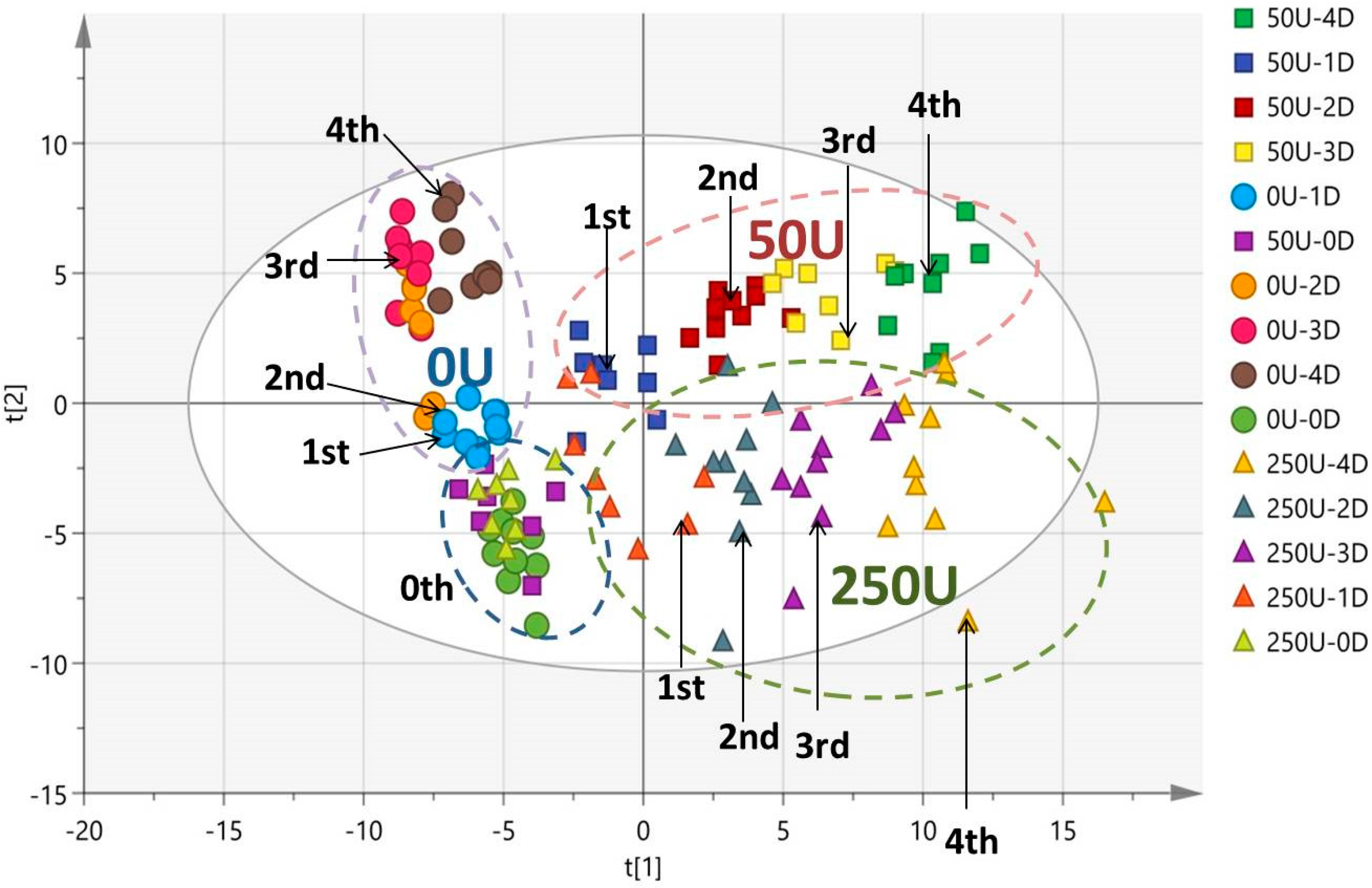
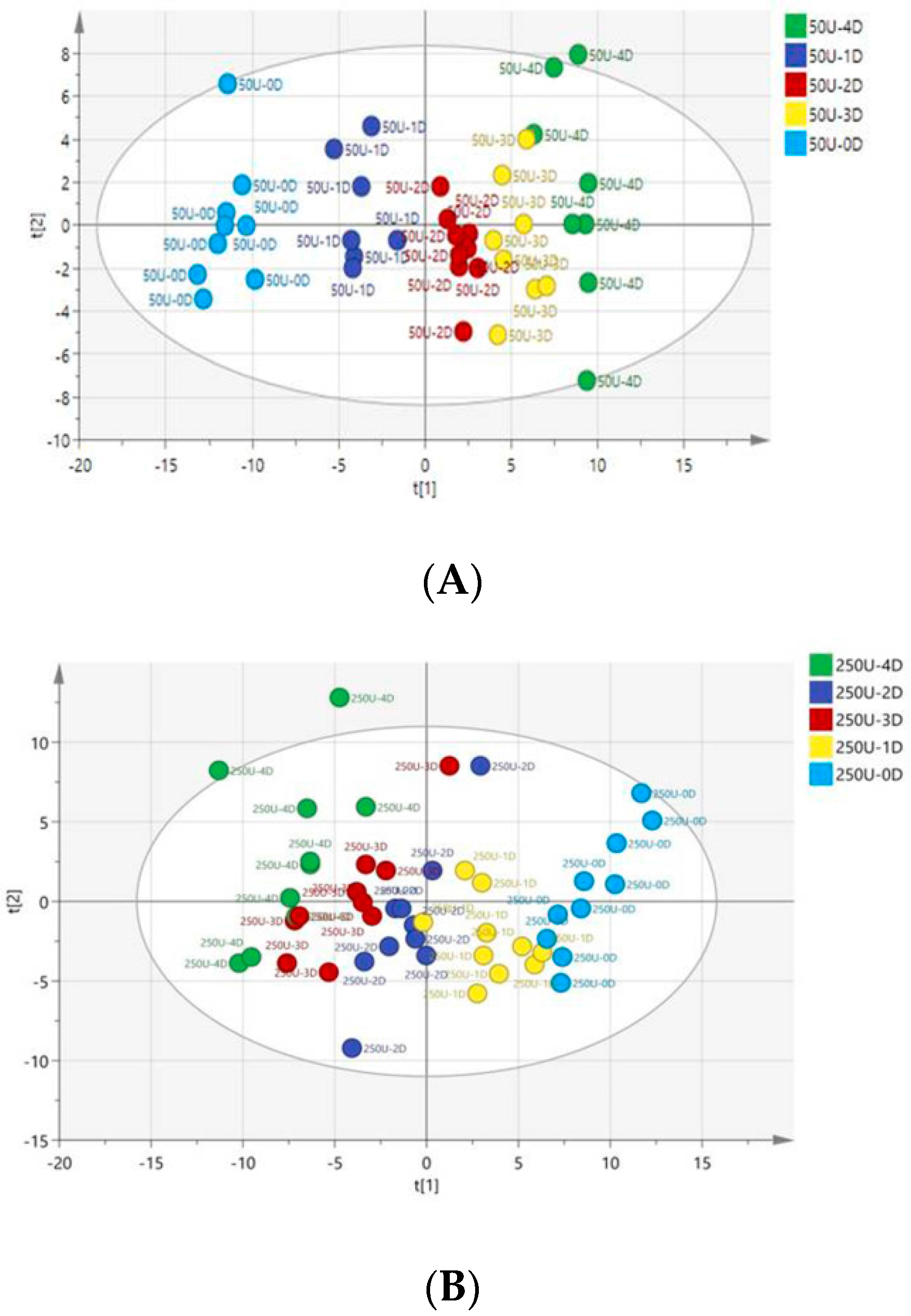
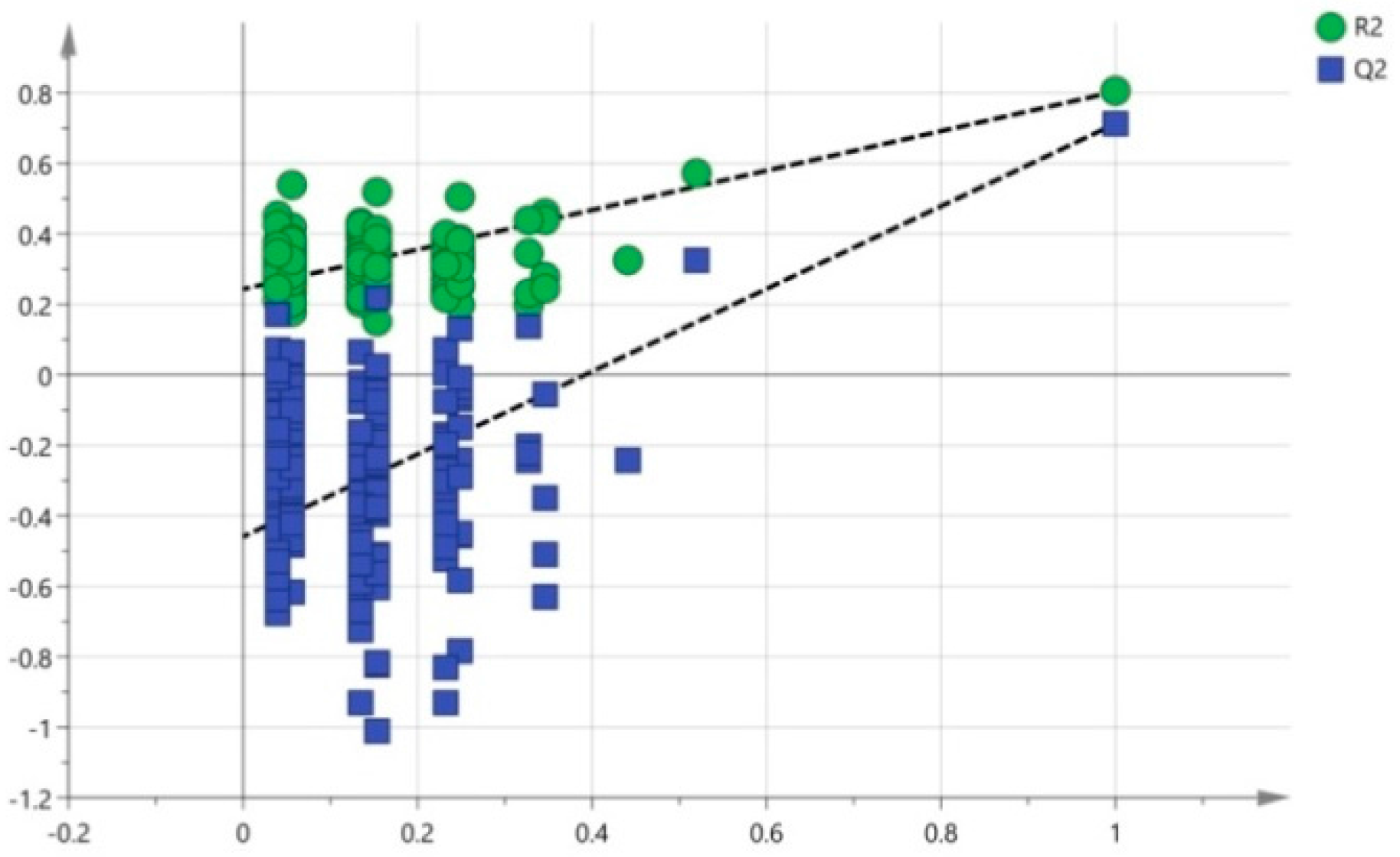
3.4. Multivariate Data Analysis of HPLC-TOF-MS Spectra
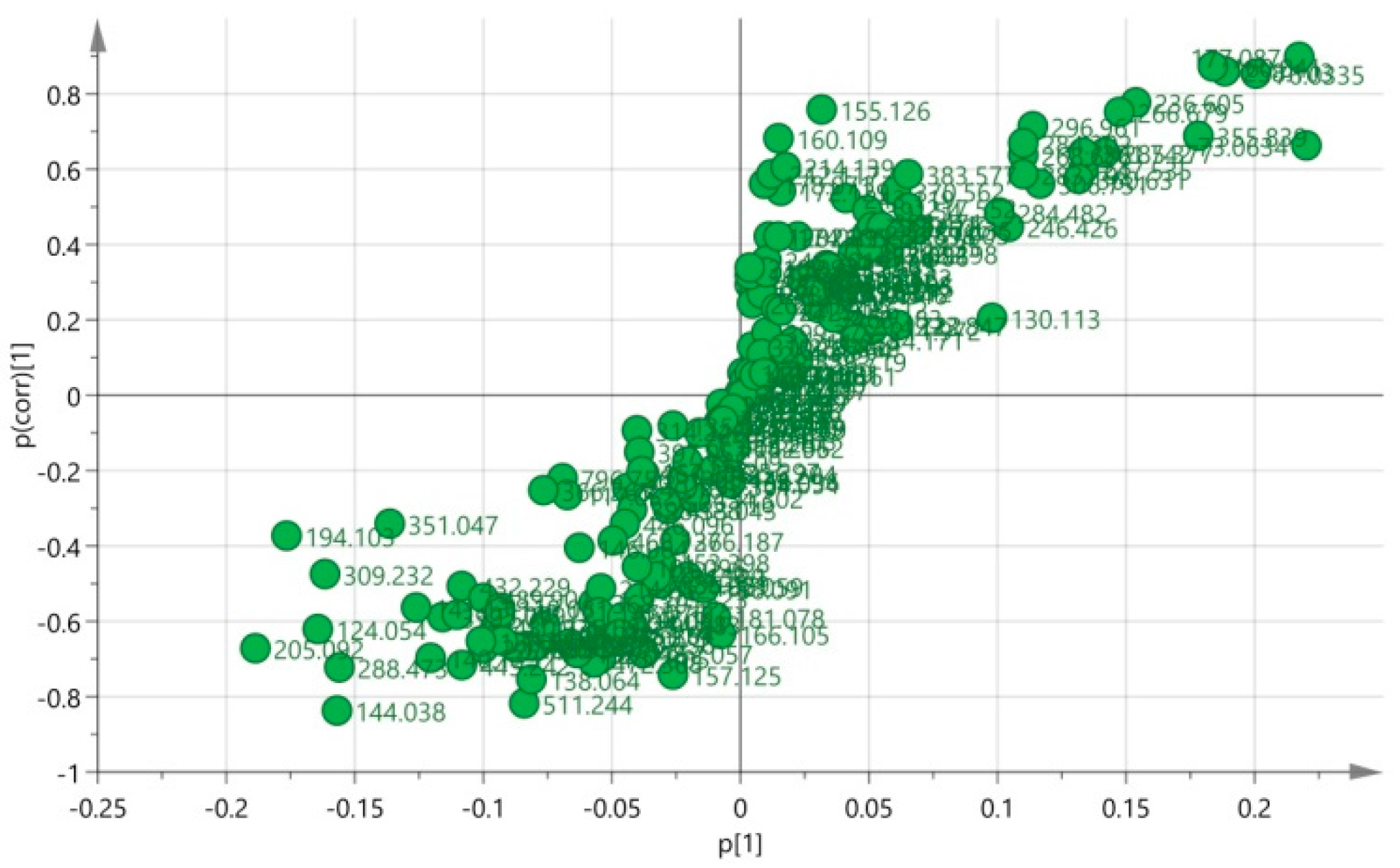
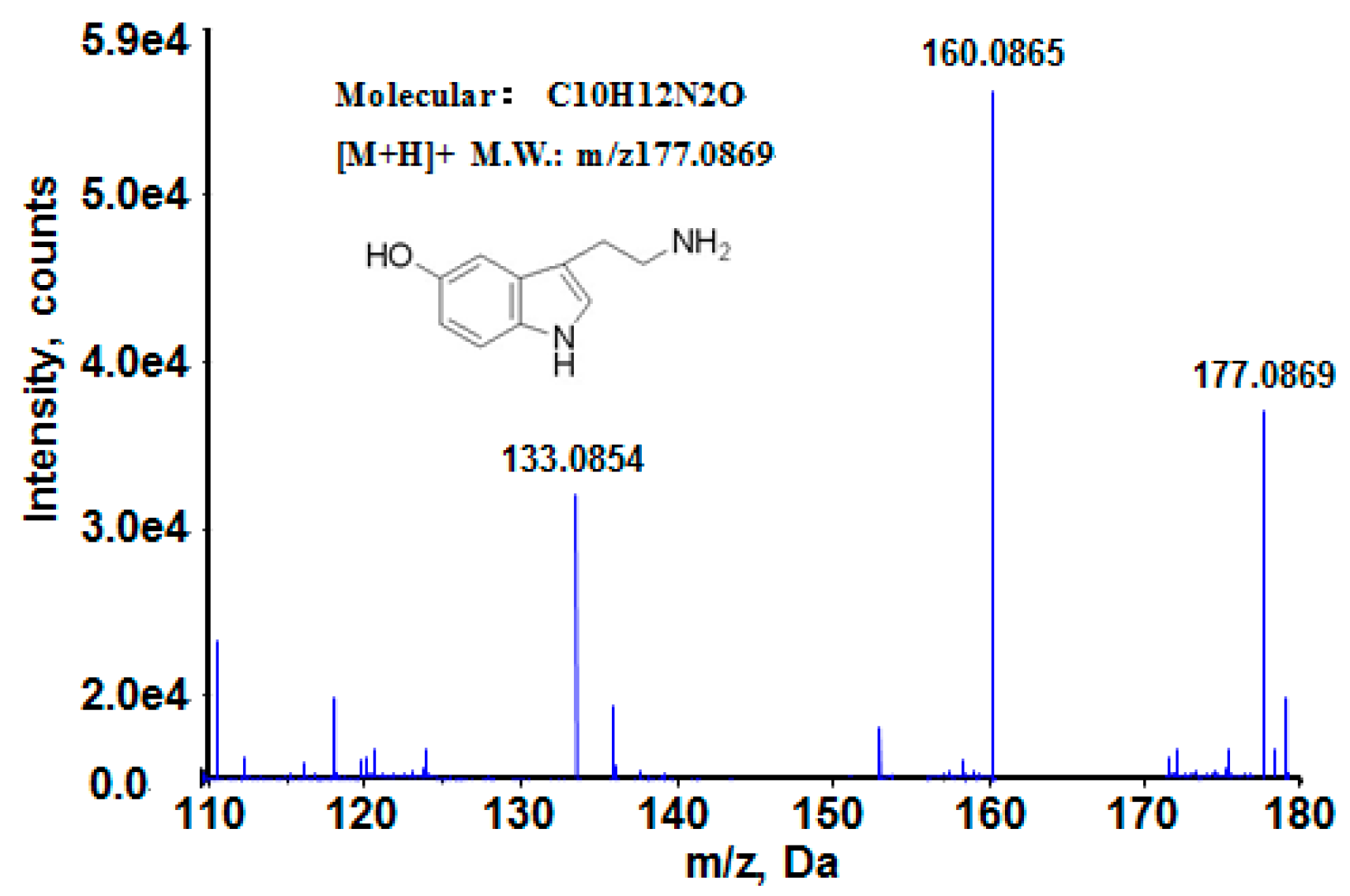
3.5. Biomarkers Identification
3.6. Potential Biomarkers Validation by Targeted Metabolomics Based on HPLC-MS/MS
3.7. Association of NP Exposure and Oxidative Stress
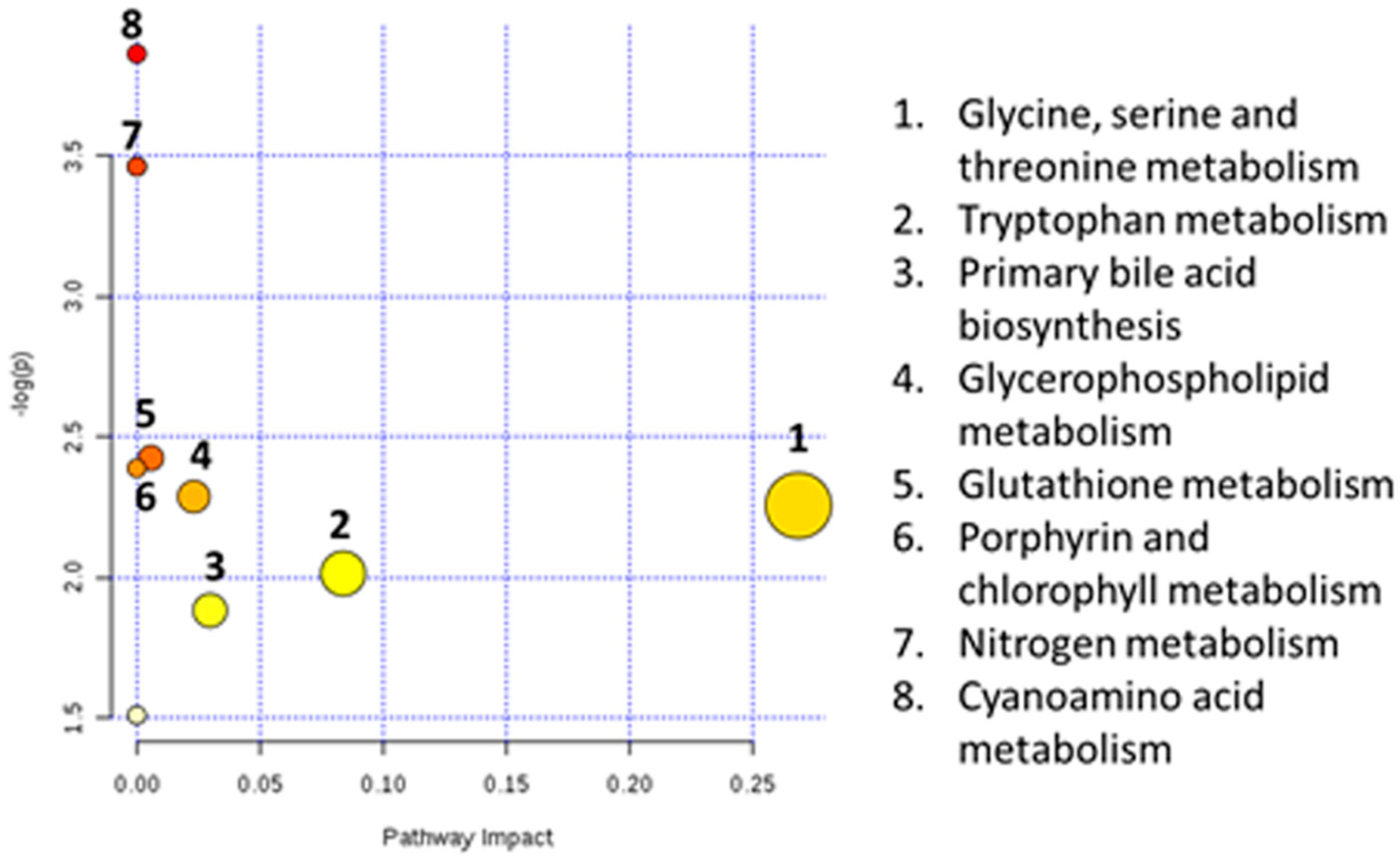
4. Discussion
5. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
Abbreviations
| NP | Nonylphenol |
| ESI | Electrospray ionization |
| MS | Mass spectrometry |
| Q-TOF MS | Quadrupole time-of-flight mass spectrometer |
| HPLC | Ultra high performance liquid chromatography |
| IDA | information-dependent acquisition |
| MVDA | multivariate data analysis |
| PCA | principal component analysis |
| OPLS-DA | orthogonal partial least-squares discrimination analysis |
| 8-oxodG | 8-oxo-deoxyguanosine |
References
- Vazquez-Duhalt, R.; Marquez-Rocha, F.; Ponce, E.; Licea, A.F.; Viana, M.T. Nonylphenol, an integrated vision of a pollutant: Scientific review. Appl. Ecol. Environ. Res. 2005, 4, 1–25. [Google Scholar] [CrossRef]
- David, A.; Fenet, H.; Gomez, E. Alkylphenols in marine environments: Distribution monitoring strategies and detection considerations. Mar. Pollut. Bull. 2009, 58, 953–960. [Google Scholar] [CrossRef] [PubMed]
- Hawker, D.W.; Cumming, J.L.; Neale, P.A.; Bartkow, M.E.; Escher, B.I. A screening level fate model of organic contaminants from advanced water treatment in a potable water supply reservoir. Water Res. 2011, 45, 768–780. [Google Scholar] [CrossRef] [PubMed]
- Gatidou, G.; Vassalou, E.; Thomaidis, N.S. Bioconcentration of selected endocrine disrupting compounds in the mediterranean mussel, mytilus galloprovincialis. Mar. Pollut. Bull. 2010, 60, 2111–2116. [Google Scholar] [CrossRef] [PubMed]
- Chen, G.W.; Ding, W.H.; Ku, H.Y.; Chao, H.R.; Chen, H.Y.; Huang, M.C.; Wang, S.L. Alkylphenols in human milk and their relations to dietary habits in central Taiwan. Food Chem. Toxicol. 2010, 48, 1939–1944. [Google Scholar] [CrossRef] [PubMed]
- Tao, M.S.; Chang, C.H.; Lin, C.L.; Hou, J.W. The association between nonylphenols and sexual hormones levels among pregnant women: A cohort study in Taiwan. PLoS ONE 2014, 9, E104245. [Google Scholar]
- Lu, Z.; Gan, J. Analysis, toxicity, occurrence and biodegradation of nonylphenol isomers: A review. Environ. Int. 2014, 73, 334–345. [Google Scholar] [CrossRef] [PubMed]
- Litwa, E.; Rzemieniec, J.; Wnuk, A.; Lason, W.; Krzeptowski, W.; Kajta, M. Apoptotic and neurotoxic actions of 4-para-nonylphenol are accompanied by activation of retinoid x receptor and impairment of classical estrogen receptor signaling. J. Steroid. Biochem. Mol. Biol. 2014, 144 Pt B, 334–347. [Google Scholar] [CrossRef] [PubMed]
- Mao, Z.; Zheng, Y.L.; Zhang, Y.Q. Behavioral impairment and oxidative damage induced by chronic application of nonylphenol. Int. J. Mol. Sci. 2010, 12, 114–127. [Google Scholar] [CrossRef] [PubMed]
- Kumaran, S.S.; Kavitha, C.; Ramesha, M.; Grummtb, T. Toxicity studies of nonylphenol and octylphenol: Hormonal, hematological and biochemical effects in clarias gariepinus. J. Appl. Toxicol. 2011, 31, 752–761. [Google Scholar] [CrossRef] [PubMed]
- Woo, G.H.; Shibutani, M.; Ichiki, T.; Hamamura, M.; Lee, K.Y.; Inoue, K.; Hirose, M. A repeated 28-day oral dose toxicity study of nonylphenol in rats, based on the “enhanced oecd test guideline 407” for screening of endocrine-disrupting chemicals. Arch. Toxicol. 2007, 81, 77–88. [Google Scholar] [CrossRef] [PubMed]
- Sakazaki, H.; Ueno, H.; Nakamuro, K. Estrogen receptor in mouse splenic lymphocytes: Possible involvement in immunity. Toxicol. Lett. 2002, 133, 221–229. [Google Scholar] [CrossRef]
- Laurenzanaa, E.M.; Weisa, C.C.; Bryanta, C.W.; Newboldb, R.; Delclosa, K.B. Effect of dietary administration of genistein, nonylphenol or ethinyl estradiol on hepatic testosterone metabolism, cytochrome P-450 enzymes, and estrogen receptor alpha expression. Food Chem. Toxicol. 2002, 40, 53–63. [Google Scholar] [CrossRef]
- Bino, R.J.; Hall, R.D.; Fiehn, O.; Kopka, J.; Saito, K.; Draper, J.; Nikolau, B.J.; Mendes, P.; Roessner-Tunali, U.; Beale, M.H.; et al. Potential of metabolomics as a functional genomics tool. Trends Plant. Sci. 2004, 9, 418–425. [Google Scholar] [CrossRef] [PubMed]
- Vlaanderen, J.; Moore, L.E.; Smith, M.T.; Lan, Q.; Zhang, L.; Skibola, C.F.; Rothman, N.; Vermeulen, R. Application of omics technologies in occupational and environmental health research: Current status and projections. Occup. Environ. Med. 2010, 67, 136–143. [Google Scholar] [CrossRef] [PubMed]
- Robertson, D.G.; Watkins, P.B.; Reily, M.D. Metabolomics in toxicology: Preclinical and clinical applications. Toxicol. Sci. 2011, 1, 146–170. [Google Scholar] [CrossRef] [PubMed]
- Dettmer, K.; Hammock, B.D. Metabolomics—A new exciting field within the omics sciences. Environ. Health Persp. 2004, 112, 396–397. [Google Scholar]
- Jeanneret, F.; Tonoli, D.; Hochstrasser, D.; Saurat, J.H.; Sorg, O.; Boccard, J.; Rudaz, S. Evaluation and identification of dioxin exposure biomarkers in human urine by high-resolution metabolomics, multivariate analysis and in vitro synthesis. Toxicol. Lett. 2016, 240, 22–31. [Google Scholar] [CrossRef] [PubMed]
- Tufi, S.; Stel, J.M.; de Boer, J.; Lamoree, M.H.; Leonards, P.E. Metabolomics to explore imidacloprid-induced toxicity in the central nervous system of the freshwater snail lymnaea stagnalis. Environ. Sci. Technol. 2015, 49, 14529–14536. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.H.; Woo, H.M.; Jung, B.H.; Lee, J.; Kwon, O.S.; Pyo, H.S.; Choi, M.H.; Chung, B.C. Metabolomic approach to evaluate the toxicological effects of nonylphenol with rat urine. Anal. Chem. 2007, 79, 6102–6110. [Google Scholar] [CrossRef] [PubMed]
- Dettmer, K.; Aronov, P.A.; Hammock, B.D. Mass spectrometry-based metabolomics. Mass Spectrom. Rev. 2007, 26, 51–78. [Google Scholar] [CrossRef] [PubMed]
- Chen, M.; Zhu, P.; Xu, B.; Zhao, R.; Qiao, S.; Chen, X.; Tang, R.; Wu, D.; Song, L.; Wang, S.; et al. Determination of nine environmental phenols in urine by ultra-high-performance liquid chromatography-tandem mass spectrometry. J. Anal. Toxicol. 2012, 36, 608–615. [Google Scholar] [CrossRef] [PubMed]
- Ye, X.Y.; Kuklenyik, Z.; Needham, L.L.; Calafat, A.M. Automated on-line column-switching HPLC-MS/MS method with peak focusing for the determination of nine environmental phenols in urine. Anal. Chem. 2005, 77, 5407–5413. [Google Scholar] [CrossRef] [PubMed]
- Scripps Center for Metabolomics. Metlin Metabolite Database. Available online: http://metlin.scripps.edu/ (accessed on 22 March 2016).
- MassBank. Available online: http://massbank.eu/MassBank/ (accessed on 22 March 2016).
- The Human Metabolome Database (HMDB). Available online: http://hmdb.ca/ (accessed on 22 March 2016).
- Asimakopoulos, A.G.; Thomaidis, N.S.; Koupparis, M.A. Recent trends in biomonitoring of bisphenol a, 4-t-octylphenol, and 4-nonylphenol. Toxicol. Lett. 2012, 210, 141–154. [Google Scholar] [CrossRef] [PubMed]
- Gika, H.G.; Wingate, J.E.; Wilson, I.D. Within-Day reproducibility of an HPLC-MS-Based method for metabonomic analysis: Application to human urine. J. Proteome. Res. 2007, 6, 3291–3303. [Google Scholar] [CrossRef] [PubMed]
- Kumar, B.; Prakash, A.; Ruhela, R.K.; Medhi, B. Potential of metabolomics in preclinical and clinical drug development. Pharmacol. Rep. 2014, 66, 956–963. [Google Scholar] [CrossRef] [PubMed]
- Robotti, E.; Marengo, E. Chemometric multivariate tools for candidate biomarker identification: LDA, PLS-DA, SIMCA, Ranking-PCA. Methods Mol. Biol. 2016, 1384, 237–267. [Google Scholar] [PubMed]
- Loft, S.; Danielsen, P.H.; Mikkelsen, L.; Risom, L.; Forchhammer, L.; Moller, P. Biomarkers of oxidative damage to DNA and repair. Biochem. Soc. Trans. 2008, 36, 1071–1076. [Google Scholar] [CrossRef] [PubMed]
- Mbongue, J.C.; Zhang, K.L.; Kim, N.S. Induction of indoleamine 2,3-dioxygenase in human dendritic cells by a cholera toxin b subunit—Proinsulin vaccine. PLOS ONE 2015, 2, 23. [Google Scholar]
- Maddison, D.C.; Giorgini, F. The kynurenine pathway and neurodegenerative disease. Semin. Cell. Dev. Biol. 2015, 40, 134–141. [Google Scholar] [CrossRef] [PubMed]
- Quak, J.; Doornbos, B.; Roest, A.M.; Duivis, H.E.; Vogelzangs, N.; Nolen, W.A.; Penninx, B.W.; Kema, I.P.; de Jonge, P. Does tryptophan degradation along the kynurenine pathway mediate the association between pro-inflammatory immune activity and depressive symptoms? Psychoneuroendocrinology 2014, 45, 202–210. [Google Scholar] [CrossRef] [PubMed]
- Hashimoto, T.; Perlot, T.; Rehman, A.; Trichereau, J.; Ishiguro, H.; Paolino, M.; Sigl, V.; Hanada, T.; Hanada, R.; Lipinski, S.; et al. ACE2 links amino acid malnutrition to microbial ecology and intestinal inflammation. Nature 2012, 487, 477–481. [Google Scholar] [CrossRef] [PubMed]
- Dairam, A.; Antunes, E.M.; Saravanan, K.S.; Daya, S. Non-steroidal anti-inflammatory agents, tolmetin and sulindac, inhibit liver tryptophan 2,3-dioxygenase activity and alter brain neurotransmitter levels. Life Sci. 2006, 79, 2269–2274. [Google Scholar] [CrossRef] [PubMed]
- Wang, P.W.; Chen, M.L.; Huang, L.W.; Yang, W.; Wu, K.Y.; Huang, Y.F. Prenatal nonylphenol exposure, oxidative and nitrative stress, and birth outcomes: A cohort study in Taiwan. Environ. Pollut. 2015, 207, 145–151. [Google Scholar] [CrossRef] [PubMed]
- Kowalczyk-Pachel, D.; Iciek, M.; Wydra, K.; Nowak, E.; Gorny, M.; Filip, M.; Wlodek, L.; Lorenc-Koci, E. Cysteine metabolism and oxidative processes in the rat liver and kidney after acute and repeated cocaine treatment. PLOS ONE 2016, 11, E0147238. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.E.; Park, J.H.; Shin, I.C.; Koh, H.C. Reactive oxygen species regulated mitochondria-mediated apoptosis in PC12 cells exposed to chlorpyrifos. Toxicol. Appl. Pharmacol. 2012, 263, 148–162. [Google Scholar] [CrossRef] [PubMed]
- Klein, J. Membrane breakdown in acute and chronic neurodegeneration: Focus on choline-containing phospholipids. J. Neural. Transm. 2000, 107, 1027–1063. [Google Scholar] [CrossRef] [PubMed]
- Steuer, A.E.; Boxler, M.I.; Stock, L.; Kraemer, T. Inhibition potential of 3,4-methylenedioxymethamphetamine (MDMA) and its metabolites on the in vitro monoamine oxidase (MAO)-catalyzed deamination of the neurotransmitters serotonin and dopamine. Toxicol. Lett. 2016, 243, 48–55. [Google Scholar] [CrossRef] [PubMed]
- Yen, C.H.; Lin, K.C.; Leu, S.; Sun, C.K.; Chang, L.T.; Chai, H.T.; Chung, S.Y.; Chang, H.W.; Ko, S.F.; Chen, Y.T.; et al. Chronic exposure to environmental contaminant nonylphenol exacerbates adenine-induced chronic renal insufficiency: Role of signaling pathways and therapeutic impact of rosuvastatin. Eur. J. Pharm. Sci. 2012, 46, 455–467. [Google Scholar] [CrossRef] [PubMed]
- Yen, C.H.; Leu, S.; Wallace, C.G.; Lin, Y.C.; Chang, L.T. Continuing exposure to low-dose nonylphenol aggravates adenine-induced chronic renal dysfunction and role of rosuvastatin therapy. J. Transl. Med. 2012, 10, 147. [Google Scholar] [CrossRef] [PubMed]
- Kim, Y.S.; Hwang, K.A.; Hyun, S.H.; Nam, K.H.; Lee, C.K.; Choi, K.C. Bisphenol A and nonylphenol have the potential to stimulate the migration of ovarian cancer cells by inducing epithelial–mesenchymal transition via an estrogen receptor dependent pathway. Chem. Res. Toxicol. 2015, 28, 662–671. [Google Scholar] [CrossRef] [PubMed]
- Naderi, M.; Safahieh, A.; Madiseh, S.D.; Zolgharnein, H.; Ghatrami, E.R. Induction of vitellogenin synthesis in immature male yellowfin seabream (acanthopagrus latus) exposed to 4-nonylphenol and 17beta-estradiol. Toxicol. Ind. Health. 2015, 31, 209–220. [Google Scholar] [CrossRef] [PubMed]
- Sayed, A.E.D.; Mahmoud, U.M.; Mekkawy, I.A. Reproductive biomarkers to identify endocrine disruption in Clarias gariepinus exposed to 4-nonylphenol. Ecotoxicol. Environ. Saf. 2012, 78, 310–319. [Google Scholar] [CrossRef] [PubMed]
- Verderame, M.; Limatola, E. Interferences of an environmental pollutant with estrogen-like action in the male reproductive system of the terrestrial vertebrate podarcis sicula. Gen. Comp. Endocrinol. 2015, 213, 9–15. [Google Scholar] [CrossRef] [PubMed]
- Asimakopoulos, A.G.; Xue, J.; De Carvalho, B.P.; Iyer, A.; Abualnaja, K.O.; Yaghmoor, S.S.; Kumosani, T.A.; Kannan, K. Urinary biomarkers of exposure to 57 xenobiotics and its association with oxidative stress in a population in Jeddah, Saudi Arabia. Environ. Res. 2015. [Google Scholar] [CrossRef] [PubMed]



| Metabolite | Molecular Formula | VIP Score | p-Value | Monitored Structure | Monitored M.W. | Exact Mass | Trend |
|---|---|---|---|---|---|---|---|
| 5-hydroxytryptamine | C10H12N2O | 2.59281 | 0.036 | M + H | 177.0869 | 176.0950 | upward |
| tryptophan | C11H12N2O2 | 2.39113 | 0.027 | M + H | 205.0918 | 204.0899 | downward |
| glycine | C2H5NO2 | 2.40796 | 0.041 | M + H | 76.0335 | 75.0320 | upward |
| glycerophosphocholine | C8H20NO6P | 2.2673 | 0.032 | M + H | 258.1031 | 257.1028 | upward |
| malonaldehyde | C3H4O2 | 2.89664 | 0.023 | M + H | 73.0634 | 72.0211 | upward |
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhang, Y.-X.; Yang, X.; Zou, P.; Du, P.-F.; Wang, J.; Jin, F.; Jin, M.-J.; She, Y.-X. Nonylphenol Toxicity Evaluation and Discovery of Biomarkers in Rat Urine by a Metabolomics Strategy through HPLC-QTOF-MS. Int. J. Environ. Res. Public Health 2016, 13, 501. https://doi.org/10.3390/ijerph13050501
Zhang Y-X, Yang X, Zou P, Du P-F, Wang J, Jin F, Jin M-J, She Y-X. Nonylphenol Toxicity Evaluation and Discovery of Biomarkers in Rat Urine by a Metabolomics Strategy through HPLC-QTOF-MS. International Journal of Environmental Research and Public Health. 2016; 13(5):501. https://doi.org/10.3390/ijerph13050501
Chicago/Turabian StyleZhang, Yan-Xin, Xin Yang, Pan Zou, Peng-Fei Du, Jing Wang, Fen Jin, Mao-Jun Jin, and Yong-Xin She. 2016. "Nonylphenol Toxicity Evaluation and Discovery of Biomarkers in Rat Urine by a Metabolomics Strategy through HPLC-QTOF-MS" International Journal of Environmental Research and Public Health 13, no. 5: 501. https://doi.org/10.3390/ijerph13050501
APA StyleZhang, Y.-X., Yang, X., Zou, P., Du, P.-F., Wang, J., Jin, F., Jin, M.-J., & She, Y.-X. (2016). Nonylphenol Toxicity Evaluation and Discovery of Biomarkers in Rat Urine by a Metabolomics Strategy through HPLC-QTOF-MS. International Journal of Environmental Research and Public Health, 13(5), 501. https://doi.org/10.3390/ijerph13050501








