On the Chemistry, Toxicology and Genetics of the Cyanobacterial Toxins, Microcystin, Nodularin, Saxitoxin and Cylindrospermopsin
Abstract
:1. Microcystin
1.1. Introduction
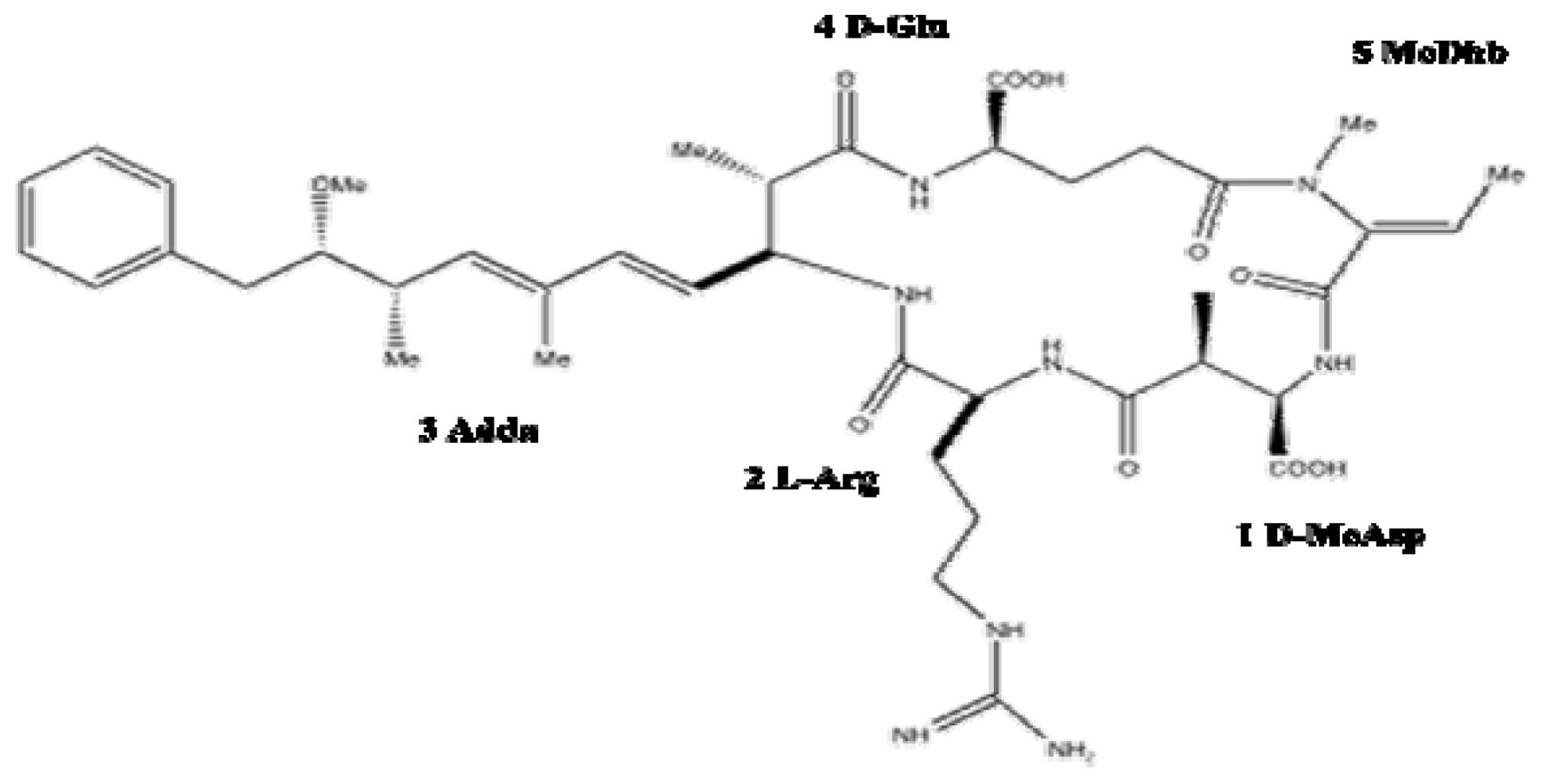
1.2. Chemistry
1.3. Toxicology
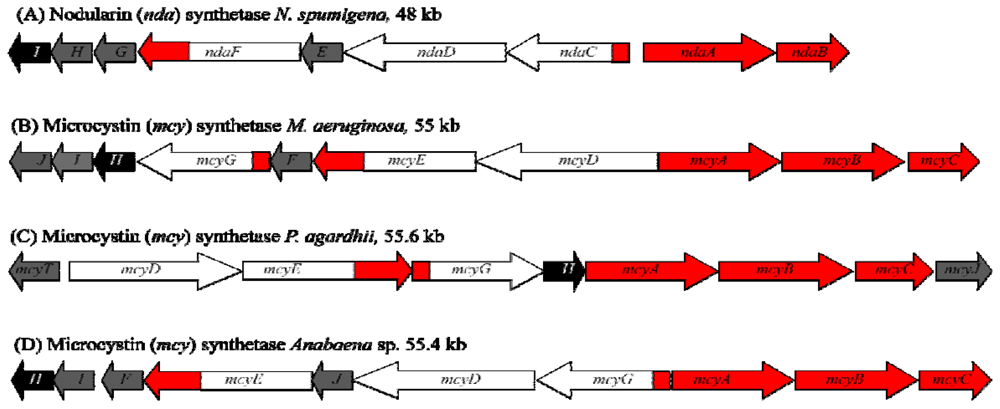
1.4. Biosynthesis and Genetics
2. Nodularin
2.1. Introduction
2.2. Chemistry
2.3. Toxicology
2.4. Biosynthesis and Genetics
3. Saxitoxin
3.1. Introduction
3.2. Chemistry
3.3. Toxicology
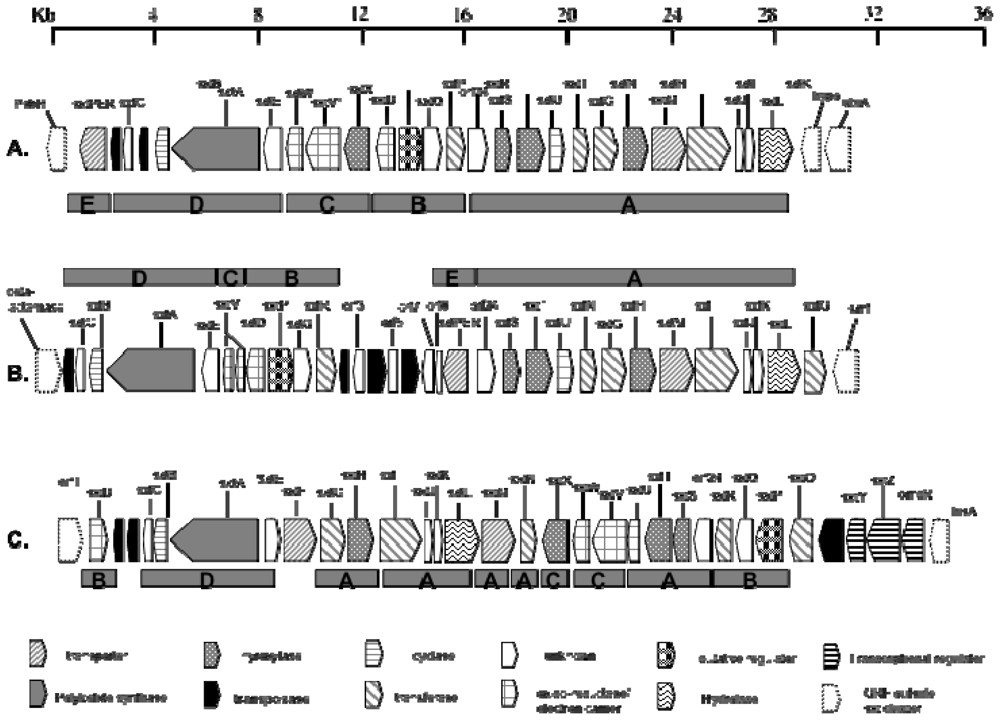
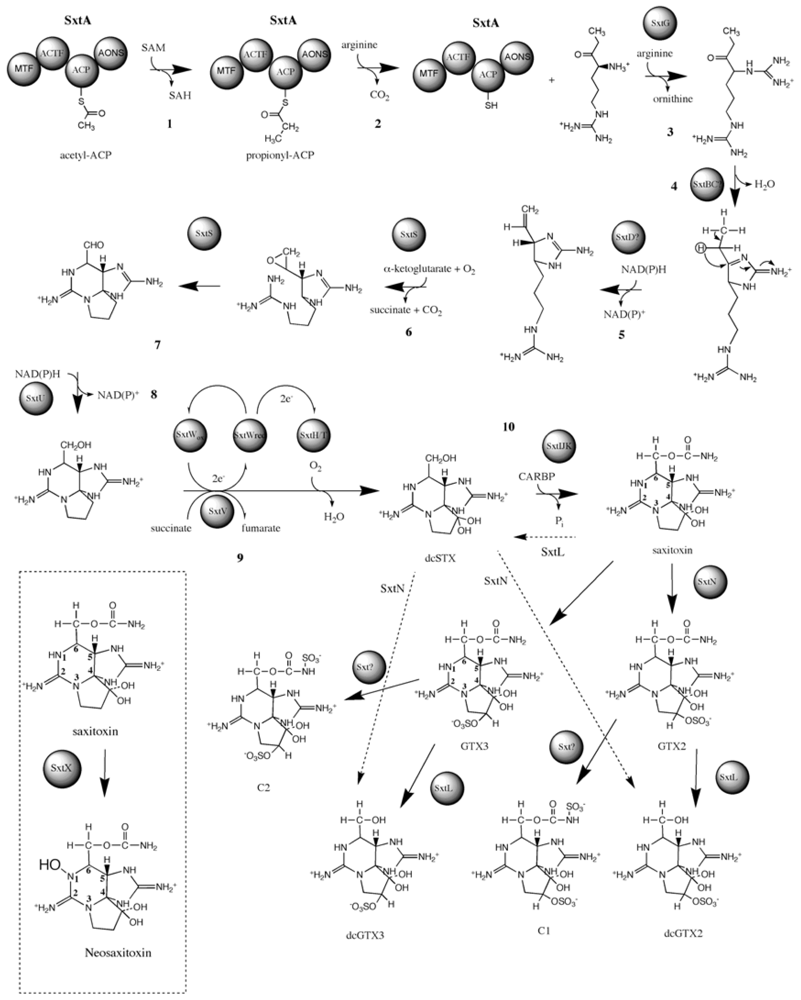
3.4. Biosynthesis and Genetics
4. Cylindrospermopsin
4.1. Introduction
4.2. Chemistry
4.3. Toxicology
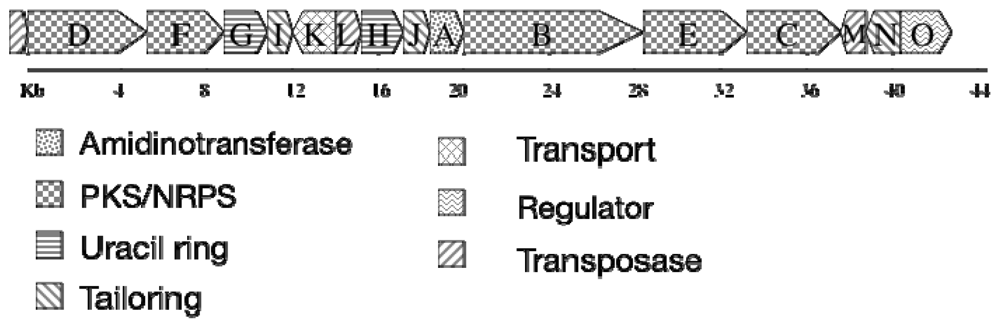
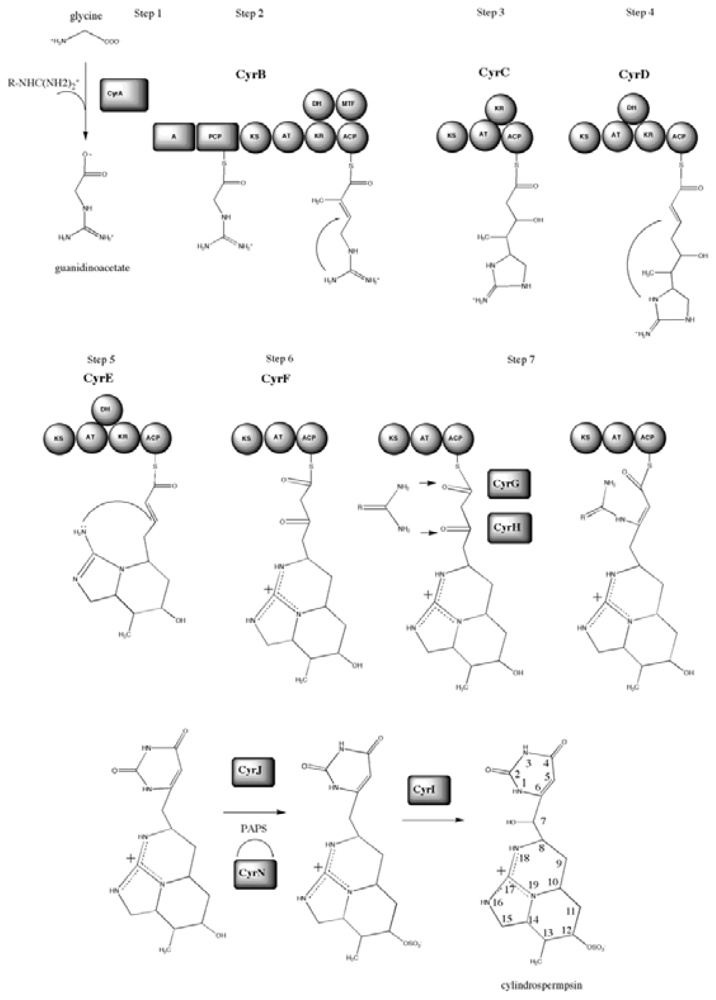
4.4. Biosynthesis and Genetics
References and Notes
- Sivonen, K; Jones, G. Toxic Cyanobacteria in Water: A Guide to Their Public Health consequences, Monitoring and Management; E and FN Spon: New York, NY, USA, 1999; Volume 1, pp. 40–111. [Google Scholar]
- Welker, M; von Dohren, H. Cyanobacterial peptides -nature’s own combinatorial biosynthesis. FEMS Microbiol Rev 2006, 30, 530–563. [Google Scholar]
- Botes, D; Wessels, P; Kruger, H; Runnegar, M; Santikarn, S; Smith, R; Barna, J; Williams, D. Structural studies on cyanoginosins-LR, -YR, -YA, and -YM, peptide toxins from Microcystis aeruginosa. J Chem Soc 1985, 1, 2747–2748. [Google Scholar]
- Namikoshi, M; Yuan, M; Sivonen, K; Carmichael, WW; Rinehart, KL; Rouhiainen, L; Sun, F; Brittain, S; Otsuki, A. Seven new microcystins possessing two l-glutamic acid units, isolated from Anabaena sp. strain 186. Chem Res Toxicol 1998, 11, 143–149. [Google Scholar]
- Rinehart, K; Namikoshi, N; Choi, B. Structure and biosynthesis of toxins from blue-green algae (cyanobacteria). J App Phycol 1994, 6, 159–176. [Google Scholar]
- Sivonen, K. Cyanobacterial toxins and toxin production. Phycologia 1996, 35, 12–24. [Google Scholar]
- Chorus, I; Bartram, J. Toxic Cyanobacteria in Water. In A Guide to their Public Health Consequences, Monitoring and Management; E & FN Spon: London, UK, 1999. [Google Scholar]
- Jochimsen, EM; Carmichael, WW; An, JS; Cardo, DM; Cookson, ST; Holmes, CE; Antunes, MB; de Melo Filho, DA; Lyra, TM; Barreto, VS; Azevedo, SM; Jarvis, WR. Liver failure and death after exposure to microcystins at a hemodialysis center in Brazil. N Engl J Med 1998, 338, 873–878. [Google Scholar]
- Runnegar, MT; Gerdes, RG; Falconer, IR. The uptake of the cyanobacterial hepatotoxin microcystin by isolated rat hepatocytes. Toxicon 1991, 29, 43–51. [Google Scholar]
- Runnegar, M; Berndt, N; Kaplowitz, N. Microcystin uptake and inhibition of protein phosphatases: effects of chemoprotectants and self-inhibition in relation to known hepatic transporters. Toxicol Appl Pharmacol 1995, 134, 264–272. [Google Scholar]
- Dawson, RM. The toxicology of microcystins. Toxicon 1998, 36, 953–962. [Google Scholar]
- Eriksson, JE; Toivola, D; Meriluoto, JA; Karaki, H; Han, YG; Hartshorne, D. Hepatocyte deformation induced by cyanobacterial toxins reflects inhibition of protein phosphatases. Biochem Biophys Res Commun 1990, 173, 1347–1353. [Google Scholar]
- Sahin, A; Tencalla, FG; Dietrich, DR; Mez, K; Naegeli, H. Enzymatic analysis of liver samples from rainbow trout for diagnosis of blue-green algae-induced toxicosis. Am J Vet Res 1995, 56, 1110–1115. [Google Scholar]
- Krishnamurthy, T; Carmichael, WW; Sarver, EW. Toxic peptides from freshwater cyanobacteria (blue-green algae). I. Isolation, purification and characterization of peptides from Microcystis aeruginosa and Anabaena flosaquae. Toxicon 1986, 24, 865–873. [Google Scholar]
- Watanabe, MF; Oishi, S; Harda, K; Matsuura, K; Kawai, H; Suzuki, M. Toxins contained in Microcystis species of cyanobacteria (blue-green algae). Toxicon 1988, 26, 1017–1025. [Google Scholar]
- Yoshizawa, S; Matsushima, R; Watanabe, MF; Harada, K; Ichihara, A; Carmichael, WW; Fujiki, H. Inhibition of protein phosphatases by microcystins and nodularin associated with hepatotoxicity. J Cancer Res Clin Oncol 1990, 116, 609–614. [Google Scholar]
- Nishiwaki, S; Fujiki, H; Suganuma, M; Nishiwaki-Matsushima, R; Sugimura, T. Rapid purification of protein phosphatase 2A from mouse brain by microcystin-affinity chromatography. FEBS Lett 1991, 279, 115–118. [Google Scholar]
- Falconer, IR. Tumor promotion and liver injury caused by oral consumption of cyanobacteria. Environ Toxicol Water Qual 1991, 6, 177–184. [Google Scholar]
- Nishiwaki-Matsushima, R; Ohta, T; Nishiwaki, S; Suganuma, M; Kohyama, K; Ishikawa, T; Carmichael, WW; Fujiki, H. Liver tumor promotion by the cyanobacterial cyclic peptide toxin microcystin-LR. J Cancer Res Clin Oncol 1992, 118, 420–424. [Google Scholar]
- Yu, S. Primary prevention of hepatocellular carcinoma. J Gastroenterol Hepatol 1995, 10, 674–682. [Google Scholar]
- Yu, S. Drinking Water and Primary Liver Cancer; China academic publishers: New York, NY, USA, 1989. [Google Scholar]
- Tillett, D; Dittmann, E; Erhard, M; von Dohren, H; Borner, T; Neilan, BA. Structural organization of microcystin biosynthesis in Microcystis aeruginosa PCC7806: an integrated peptide-polyketide synthetase system. Chem Biol 2000, 7, 753–764. [Google Scholar]
- Christiansen, G; Fastner, J; Erhard, M; Borner, T; Dittmann, E. Microcystin biosynthesis in planktothrix: genes, evolution, and manipulation. J Bacteriol 2003, 185, 564–572. [Google Scholar]
- Rouhiainen, L; Vakkilainen, T; Siemer, BL; Buikema, W; Haselkorn, R; Sivonen, K. Genes coding for hepatotoxic heptapeptides (microcystins) in the cyanobacterium Anabaena strain 90. Appl Environ Microbiol 2004, 70, 686–692. [Google Scholar]
- Hicks, LM; Moffitt, MC; Beer, LL; Moore, B; Kelleher, NL. Structural characterisation of in vitro and in vivo intermediates on the loading module of microcystin synthetase. ACS Chem Biol 2006, 1, 93–102. [Google Scholar]
- Nishizawa, T; Asayama, M; Shirai, M. Cyclic heptapeptide microcystin biosynthesis requires the glutamate racemase gene. Microbiology 2001, 147, 1235–1241. [Google Scholar]
- Sielaff, H; Dittmann, E; Tandeau De Marsac, N; Bouchier, C; Von Dohren, H; Borner, T; Schwecke, T. The mcyF gene of the microcystin biosynthetic gene cluster from Microcystis aeruginosa encodes an aspartate racemase. Biochem J 2003, 373, 909–916. [Google Scholar]
- Pearson, LA; Barrow, KD; Neilan, BA. Characterization of the 2-hydroxy-acid dehydrogenase McyI, encoded within the microcystin biosynthesis gene cluster of Microcystis aeruginosa PCC7806. J Biol Chem 2007, 282, 4681–4692. [Google Scholar]
- Pearson, LA; Hisbergues, M; Borner, T; Dittmann, E; Neilan, BA. Inactivation of an ABC transporter gene, mcyH, results in loss of microcystin production in the cyanobacterium Microcystis aeruginosa PCC 7806. Appl Environ Microbiol 2004, 70, 6370–6378. [Google Scholar]
- Shi, L; Carmichael, WW; Miller, I. Immuno-gold localization of hepatotoxins in cyanobacterial cells. Arch Microbiol 1995, 163, 7–15. [Google Scholar]
- Young, FM; Thomson, C; Metcalf, JS; Lucocq, JM; Codd, GA. Immunogold localisation of microcystins in cryosectioned cells of Microcystis. J Struct Biol 2005, 151, 208–214. [Google Scholar]
- Kaebernick, M; Neilan, BA; Borner, T; Dittmann, E. Light and the transcriptional response of the microcystin biosynthesis gene cluster. Appl Environ Microbiol 2000, 66, 3387–3392. [Google Scholar]
- Mikalsen, B; Boison, G; Skulberg, OM; Fastner, J; Davies, W; Gabrielsen, TM; Rudi, K; Jakobsen, KS. Natural variation in the microcystin synthetase operon mcyABC and impact on microcystin production in Microcystis strains. J Bacteriol 2003, 185, 2774–2785. [Google Scholar]
- Tooming-Klunderud, A; Mikalsen, B; Kristensen, T; Jakobsen, KS. The mosaic structure of the mcyABC operon in Microcystis. Microbiology 2008, 154, 1886–1899. [Google Scholar]
- Sivonen, K. Effects of light, temperature, nitrate, orthophosphate, and bacteria on growth of and hepatotoxin production by Oscillatoria agardhii strains. Appl Environ Microbiol 1990, 56, 2658–2666. [Google Scholar]
- Lukac, M; Aegerter, R. Influence of trace metals on growth and toxin production of Microcystis aeruginosa. Toxicon 1993, 31, 293–305. [Google Scholar]
- van der Westhuizen, AJ; Eloff, JN. Effect of temperature and light on the toxicity and growth of the blue-green alga Microcystis aeruginosa (UV-006). Planta 1985, 163, 55–59. [Google Scholar]
- Song, L; Sano, T; Li, R; Watanabe, M; Liu, Y; Kaya, K. Microcystin production of Microcystis viridis (cyanobacteria) under different culture conditions. Phycol Res 1998, 42, 19. [Google Scholar]
- Davis, TW; Berry, DL; Boyer, GL; Gobler, CJ. The effects of temperature and nutrients on the growth and dynamics of toxic and non-toxic strains of Microcystis during cyanobacteria blooms. Harmful Algae 2009, 8, 715–725. [Google Scholar]
- Tonk, L; Visser, PM; Christiansen, G; Dittmann, E; Snelder, EO; Wiedner, C; Mur, LR; Huisman, J. The microcystin composition of the cyanobacterium Planktothrix agardhii changes toward a more toxic variant with increasing light intensity. Appl Environ Microbiol 2005, 71, 5177–5181. [Google Scholar]
- Sevilla, E; Martin-Luna, B; Vela, L; Bes, MT; Fillat, MF; Peleato, ML. Iron availability affects mcyD expression and microcystin-LR synthesis in Microcystis aeruginosa PCC7806. Environ Microbiol 2008, 10, 2476–2483. [Google Scholar]
- Kaebernick, M; Dittmann, E; Borner, T; Neilan, BA. Multiple alternate transcripts direct the biosynthesis of microcystin, a cyanobacterial nonribosomal peptide. Appl Environ Microbiol 2002, 68, 449–455. [Google Scholar]
- Sivonen, K; Kononen, K; Carmichael, WW; Dahlem, AM; Rinehart, KL; Kiviranta, J; Niemela, SI. Occurrence of the hepatotoxic cyanobacterium Nodularia spumigena in the Baltic Sea and structure of the toxin. Appl Environ Microbiol 1989, 55, 1990–1995. [Google Scholar]
- Jones, GJ; Blackburn, SI; Parker, NS. A toxic bloom of Nodularia spumigena mertens in Orielton Lagoon, Tasmania. Aust J Mar Freshwater Res 1994, 45, 787–800. [Google Scholar]
- Heresztyn, T; Nicholson, BC. Nodularin concentrations in Lakes Alexandrina and Albert, South Australia, during a bloom of the cyanobacterium (blue-green alga) Nodularia spumigena and degradation of the toxin. Environ Toxicol Water Qual 1997, 12, 273–282. [Google Scholar]
- Nehring, S. Mortality of dogs associated with a mass development of Nodularia spumigena (Cyanophyceae) in a brackish lake at the German North Sea coast. J Plankton Res 1993, 15, 867–872. [Google Scholar]
- Carmichael, WW; Eschedor, JT; Patterson, GM; Moore, RE. Toxicity and partial structure of a hepatotoxic peptide produced by the cyanobacterium Nodularia spumigena Mertens emend. L575 from New Zealand. Appl Environ Microbiol 1988, 54, 2257–2263. [Google Scholar]
- Galat, DL; Verdin, JP; Sims, LL. Large-scale patterns of Nodularia spumigena blooms in Pyramid Lake, Nevada, determined from Landsat imagery: 1972–1986. Hydrobiologia 1990, 197, 147–164. [Google Scholar]
- Baker, PD; Humpage, AR. Toxicity associated with commonly occurring cyanobacteria in surface waters of the Murray-Darling Basin, Australia. Aust J Mar Freshwater Res 1994, 45, 773–786. [Google Scholar]
- Karjalainen, M; Engstrom-Ost, J; Korpinen, S; Peltonen, H; Paakkonen, JP; Ronkkonen, S; Suikkanen, S; Viitasalo, M. Ecosystem consequences of cyanobacteria in the northern Baltic Sea. Ambio 2007, 36, 195–202. [Google Scholar]
- Carmichael, WW. The Toxins of Cyanobacteria. Sci Am 1994, 64–72. [Google Scholar]
- Francis, G. Poisonous Australian lake. Nature 1878, 18, 11–12. [Google Scholar]
- Ohta, T; Sueoka, E; Iida, N; Komori, A; Suganuma, M; Nishiwaki, R; Tatematsu, M; Kim, SJ; Carmichael, WW; Fujiki, H. Nodularin, a potent inhibitor of protein phosphatases 1 and 2A, is a new environmental carcinogen in male F344 rat liver. Cancer Res 1994, 54, 6402–6406. [Google Scholar]
- Van Buynder, PG; Oughtred, T; Kirkby, B; Phillips, S; Eaglesham, G; Thomas, K; Burch, M. Nodularin uptake by seafood during a cyanobacterial bloom. Environ Toxicol 2001, 16, 468–471. [Google Scholar]
- Falconer, IR; Choice, A; Hosja, W. Toxicity of edible mussels Mytilus edulis growing naturally in an estuary during a water bloom of the blue-green alga Nodularia spumigena. Environ Toxicol Water Qual 1992, 7, 119–123. [Google Scholar]
- Mazur-Marzec, H; Tyminska, A; Szafranek, J; Plinski, M. Accumulation of nodularin in sediments, mussels, and fish from the Gulf of Gdansk, southern Baltic Sea. Environ Toxicol 2007, 22, 101–111. [Google Scholar]
- Rinehart, KL; Harada, K; Namikoshi, M; Chen, C; Harvis, CA; Munro, MHG; Blunt, JW; Mulligan, PE; Beasley, VR; Dahlem, AM; Carmicheal, WW. Nodularin, microcystin, and the configuration of Adda. J Am Chem Soc 1988, 110, 8557–8558. [Google Scholar]
- Saito, K; Konno, A; Ishii, H; Saito, H; Nishida, F; Abe, T; Chen, C. Nodularin-Har: a new nodularin from Nodularia. J Nat Prod 2001, 64, 139–141. [Google Scholar]
- Beattie, KA; Kaya, K; Codd, GA. The cyanobacterium Nodularia PCC7804, of freshwater origin, produces [l-Har2]nodularin. Phytochemistry 2000, 54, 57–61. [Google Scholar]
- deSilva, ED; Williams, DE; Anderson, RJ; Klix, H; Holmes, CFB; Allen, TM. Motuporin, a potent protein phosphatase inhibitor isolated from the Papua New Guinea sponge Theonella swinhoei Gray. Tetrahedron Lett 1992, 33, 1561–1564. [Google Scholar]
- Eriksson, JE; Meriluoto, JA; Kujari, HP; Osterlund, K; Fagerlund, K; Hallbom, L. Preliminary characterization of a toxin isolated from the cyanobacterium Nodularia spumigena. Toxicon 1988, 26, 161–166. [Google Scholar]
- Honkanen, RE; Dukelow, M; Zwiller, J; Moore, RE; Khatra, BS; Boynton, AL. Cyanobacterial nodularin is a potent inhibitor of type 1 and type 2A protein phosphatases. Mol Pharmacol 1991, 40, 577–583. [Google Scholar]
- An, J; Carmichael, WW. Use of a colorimetric protein phosphatase inhibition assay and enzyme linked immunosorbent assay for the study of microcystins and nodularins. Toxicon 1994, 32, 1495–1507. [Google Scholar]
- MacKintosh, RW; Dalby, KN; Campbell, DG; Cohen, PT; Cohen, P; MacKintosh, C. The cyanobacterial toxin microcystin binds covalently to cysteine-273 on protein phosphatase 1. FEBS Lett 1995, 371, 236–240. [Google Scholar]
- Kelker, MS; Page, R; Peti, W. Crystal structures of protein phosphatase-1 bound to nodularin-R and tautomycin: a novel scaffold for structure-based drug design of serine/threonine phosphatase inhibitors. J Mol Biol 2009, 385, 11–21. [Google Scholar]
- Runnegar, M; Berndt, N; Kong, SM; Lee, EY; Zhang, L. In vivo and in vitro binding of microcystin to protein phosphatases 1 and 2A. Biochem Biophys Res Commun 1995, 216, 162–169. [Google Scholar]
- Bagu, JR; Sykes, BD; Craig, MM; Holmes, CF. A molecular basis for different interactions of marine toxins with protein phosphatase-1. Molecular models for bound motuporin, microcystins, okadaic acid, and calyculin A. J Biol Chem 1997, 272, 5087–5097. [Google Scholar]
- Karjalainen, M; Pääkkönen, J; Peltonen, H; Sipiä, V; Valtonen, T; Viitasalo, M. Nodularin concentrations in Baltic Sea zooplankton and fish during a cyanobacterial bloom. Mar Biol 2008, 155, 483–491. [Google Scholar]
- Persson, K; Legrand, C; Olsson, T. Detection of nodularin in European flounder (Platichthys flesus) in the west coast of Sweden: Evidence of nodularin mediated oxidative stress. Harmful Algae 2009, 8, 832–838. [Google Scholar]
- Vuorinen, PJ; Sipia, VO; Karlsson, K; Keinanen, M; Furey, A; Allis, O; James, K; Perttila, U; Rimaila-Parnanen, E; Meriluoto, JA. Accumulation and effects of nodularin from a single and repeated oral doses of cyanobacterium Nodularia spumigena on flounder (Platichthys flesus L.). Arch Environ Contam Toxicol 2009, 57, 164–173. [Google Scholar]
- Moffitt, MC; Neilan, BA. Characterization of the nodularin synthetase gene cluster and proposed theory of the evolution of cyanobacterial hepatotoxins. Appl Environ Microbiol 2004, 70, 6353–6362. [Google Scholar]
- Kleinkauf, H; Von Dohren, H. A nonribosomal system of peptide biosynthesis. Eur J Biochem 1996, 236, 335–351. [Google Scholar]
- Copp, JN; Roberts, AA; Marahiel, MA; Neilan, BA. Characterization of PPTNs, a cyanobacterial phosphopantetheinyl transferase from Nodularia spumigena NSOR10. J Bacteriol 2007, 189, 3133–3139. [Google Scholar]
- Jonasson, S; Vintila, S; Sivonen, K; El-Shehawy, R. Expression of the nodularin synthetase genes in the Baltic Sea bloom-former cyanobacterium Nodularia spumigena strain AV1. FEMS Microbiol Ecol 2008, 65, 31–39. [Google Scholar]
- Negri, AP; Jones, GJ; Hindmarsh, M. Sheep mortality associated with paralytic shellfish poisons from the cyanobacterium Anabaena circinalis. Toxicon 1995, 33, 1321–1329. [Google Scholar]
- Reyero, M; Cacho, E; Martínez, A; Vázquez, J; Marina, A; Fraga, S; Franco, J. Evidence of saxitoxin derivatives as causative agents in the 1997 mass mortality of monk seals in the Cape Blanc Peninsula. Nat Toxins 1999, 7, 311–315. [Google Scholar]
- Sawyer, PJ; Gentile, JH; Sasner, JJJ. Demonstration of a toxin from Aphanisomenon flosaquae (L.) Ralfs. Can J Microbiol 1968, 14, 1199–1204. [Google Scholar]
- Shumway, SE. Phycotoxin-related shellfish poisoning: Bivalve molluscs are not the only vectors. Rev Fish Sci 1995, 3, 1–31. [Google Scholar]
- Shumway, SE; Cucci, TL; Yentsch, CM; Newell, RC; Gainey, L. The effects of the toxic dinoflagellate, Protogonyaulax tamarensis, on the physiology and behavior of marine molluscs. J Shellfish Res 1988, 7, 132–133. [Google Scholar]
- Hallegraeff, GM. Hallegraeff, GM, Anderson, DM, Cembella, AD, Eds.; Harmful algal blooms: A global overview. In Manual on Harmful Marine Microalgae; UNESCO: Paris, France, 1995; pp. 1–22. [Google Scholar]
- Daly, JW. Marine toxins and nonmarine toxins: convergence or symbiotic organisms? J Nat Prod 2004, 67, 1211–1215. [Google Scholar]
- Baker, TR; Doucette, GJ; Powell, CL; Boyer, GL; Plumley, FG. GTX(4) imposters: characterization of fluorescent compounds synthesized by Pseudomonas stutzeri SF/PS and Pseudomonas/Alteromonas PTB-1, symbionts of saxitoxin-producing Alexandrium spp. Toxicon 2003, 41, 339–347. [Google Scholar]
- Kodoma, M; Ogata, T; Sakamoto, S; Sato, S; Honda, T; Miwatani, T. Production of paralytic shellfish toxins by a bacterium Moraxella sp. isolated from Protogonyaulax tamarensis. Toxicon 1990, 28, 707–714. [Google Scholar]
- Martins, CA; Alvito, P; Tavares, MJ; Pereira, P; Doucette, G; Franca, S. Reevaluation of production of paralytic shellfish toxin by bacteria associated with dinoflagellates of the Portuguese coast. Appl Environ Microbiol 2003, 69, 5693–5698. [Google Scholar]
- Castro, D; Vera, D; Lagos, N; Garcia, C; Vasquez, M. The effect of temperature on growth and production of paralytic shellfish poisoning toxins by the cyanobacterium Cylindrospermopsis raciborskii C10. Toxicon 2004, 44, 483–489. [Google Scholar]
- Negri, A; Stirling, D; Quilliam, M; Blackburn, S; Bolch, C; Burton, I; Eaglesham, G; Thomas, K; Walter, J; Willis, R. Three novel hydroxybenzoate saxitoxin analogues isolated from the dinoflagellate Gymnodinium catenatum. Chem Res Toxicol 2003, 16, 1029–1033. [Google Scholar]
- Sako, Y; Kim, CH; Ishida, Y. Mendelian inheritance of paralytic shellfish poisoning toxin in the marine dinoflagellateAlexandrium. Biosci Biotechnol Biochem 1992, 56692–56694. [Google Scholar]
- Sako, Y; Naya, N; Yoshida, Y; Kim, CH; Ushida, A; Ishida, Y. Lassus, P, Arzul, G, Erard, P, Gentien, C, Marcaillou-Le Baut, C, Eds.; Studies on the stability and heredity of PSP toxin composition in the toxic dinoflagellateAlexandrium. In Harmful Marine Algal Blooms; London-Paris-New York: Nantes, France, 1995; pp. 345–350. [Google Scholar]
- Velzeboer, RMA; Baker, PD; Rositano, J; Heresztyn, T; Codd, GA; Raggett, SL. Geographical patterns of occurrence and composition of saxitoxins in the cyanobacterial genus Anabaena (Nostocales, Cyanophyta) in Australia. Phycologia 2000, 39, 395–407. [Google Scholar]
- Mahmood, NA; Carmichael, WW. Paralytic shellfish poisons produced by the freshwater cyanobacterium Aphanizomenon flosaquae NH-5. Toxicon 1986, 24, 175–186. [Google Scholar]
- Harada, T; Oshima, Y; Yasumoto, T. Structure of two paralytic shellfish toxins, gonyautoxins V and VI, isolated from a tropical dinoflagellate Pyrodinium bahamense var. compressa. Agric Biol Chem 1982, 46, 1861–1864. [Google Scholar]
- Oshima, Y; Hasegawa, M; Yasumoto, T; Hallegaeff, G; Blackburn, S. Dinoflagellate Gimnodium catenatum as the source of paralytic shellfishtoxins in Tasmanian shellfish. Toxicon 1987, 25, 1105–1111. [Google Scholar]
- Shimizu, Y. Faulkner, D, Fenical, W, Eds.; Chemistry and Distribution of Deleterious Dinoflagellate Toxins; Plenum: New York, NY, USA, 1977; pp. 261–269. [Google Scholar]
- Carmichael, WW; Evans, WR; Yin, QQ; Bell, P; Moczydlowski, E. Evidence for paralytic shellfish poisons in the freshwater cyanobacterium Lyngbya wollei (Farlow ex Gomont) comb. nov. Appl Environ Microbiol 1997, 63, 3104–3110. [Google Scholar]
- Humpage, AR; Rositano, JAB; Brown, R; Baker, P; Nicholson, BC; Steffensen, DA. Paralytic shellfish poisons from australian cyanobacterial blooms. Aust J Mar Freshwater Res 1994, 45, 761–771. [Google Scholar]
- Lagos, N; Onodera, H; Zagatto, PA; Andrinolo, D; Azevedo, SM; Oshima, Y. The first evidence of paralytic shellfish toxins in the fresh water cyanobacterium Cylindrospermopsis raciborskii, isolated from Brazil. Toxicon 1999, 37, 1359–1373. [Google Scholar]
- Ballot, A; Fastner, J; Wiedner, C. Paralytic shellfish poisoning toxin-producing cyanobacterium Aphanizomenon gracile in northeast Germany. Appl Environ Microbiol 2010, 76, 1173–1180. [Google Scholar]
- Llewellyn, LE. Saxitoxin, a toxic marine natural product that targets a multitude of receptors. Nat Prod Rep 2006, 23, 200–222. [Google Scholar]
- Llewellyn, LE; Negri, AP; Doyle, J; Baker, PD; Beltran, EC; Neilan, BA. Radioreceptor assays for sensitive detection and quantitation of saxitoxin and its analogues from strains of the freshwater cyanobacterium, Anabaena circinalis. Environ Sci Technol 2001, 35, 1445–1451. [Google Scholar]
- Oshima, Y. Postcolumn derivatization liquid chromatographic method for paralytic shellfish toxins. J AOAC Int 1995, 78, 528–532. [Google Scholar]
- Oshima, Y; Blackburn, S; Hallegraeff, GM. Comparative study on paralytic shellfish toxin profiles of the dinoflagellate Gymnodinium catenatum from three different countries. Mar Biol 1993, 116, 471–476. [Google Scholar]
- Onodera, H; Satake, M; Oshima, Y; Yasumoto, T; Carmichael, WW. New saxitoxin analogues from the freshwater filamentous cyanobacterium Lyngbya wollei. Nat Toxins 1997, 5, 146–151. [Google Scholar]
- Arakawa, O; Nishio, S; Noguchi, T; Shida, Y; Onoue, Y. A new saxitoxin analogue from a xanthid crab Atergatis floridus. Toxicon 1995, 33, 1577–1584. [Google Scholar]
- Arakawa, O; Noguchi, T; Shida, Y; Onoue, Y. Occurrence of carbamoyl-N-hydroxy derivatives of saxitoxin and neosaxitoxin in a xanthid crab Zosimus aeneus. Toxicon 1994, 32, 175–183. [Google Scholar]
- Zaman, L; Arakawa, O; Shimosu, A; Shida, Y; Onoue, Y. Occurrence of a methyl derivative of saxitoxin in Bangladeshi freshwater puffers. Toxicon 1998, 36, 627–630. [Google Scholar]
- Yotsu-Yamashita, M; Kim, YH; Dudley, SC, Jr; Choudhary, G; Pfahnl, A; Oshima, Y; Daly, JW. The structure of zetekitoxin AB, a saxitoxin analog from the golden frog Atelopus zeteki: a potent sodium channel blocker. Proc Natl Acad Sci USA 2004, 101, 4346–4351. [Google Scholar]
- Kao, CY. Falconer, IR, Ed.; Paralytic shellfish poisoning. In Algal Toxins in Seafood and Drinking Water; Academic Press: London, UK, 1993; pp. 75–86. [Google Scholar]
- Kao, CY; Levinson, SR. Tetrodotoxin, Saxitoxin, and the Molecular Biology of the Sodium Channel; The New York Academy of Science: New York, NY, USA, 1986; Volume 479. [Google Scholar]
- Su, Z; Sheets, M; Ishida, H; Li, F; Barry, WH. Saxitoxin blocks l-type ICa. J Pharmacol Exp Ther 2004, 308, 324–329. [Google Scholar]
- Wang, J; Salata, JJ; Bennett, PB. Saxitoxin is a gating modifier of HERG K+ channels. J Gen Physiol 2003, 121, 583–598. [Google Scholar]
- Halstead, BW; Schantz, EJ. Paralytic shellfish poisoning. WHO Offset Publ 1984, 1–59. [Google Scholar]
- Evans, MH. Mechanism of saxitoxin and tetrodotoxin poisoning. Br Med Bull 1969, 25, 263–267. [Google Scholar]
- Satin, J; Kyle, JW; Chen, M; Bell, P; Cribbs, LL; Fozzard, HA; Rogart, RB. A mutant of TTX-resistant cardiac sodium channels with TTX-sensitive properties. Science 1992, 256, 1202–1205. [Google Scholar]
- Strichartz, G. Structural determinabts of the affinity of saxitoxin for neuronal sodium channels. J Gen Physiol 1984, 84, 281–305. [Google Scholar]
- Tikhonov, DB; Zhorov, BS. Modeling P-loops domain of sodium channel: homology with potassium channels and interaction with ligands. Biophys J 2005, 88, 184–197. [Google Scholar]
- Jiang, Y; Lee, A; Chen, J; Cadene, M; Chait, BT; MacKinnon, R. The open pore conformation of potassium channels. Nature 2002, 417, 523–526. [Google Scholar]
- Hall, S; Strichartz, G; Moczydlowski, E; Ravindran, A; Reichardt, PB. Hall, S, Reichardt, PB, Eds.; The saxitoxins: sources, chemistry and pharmacology. In Marine Toxins Origin, Structure and Pharmacology; American Chemical Society: Washington DC, USA, 1990; pp. 29–69. [Google Scholar]
- Cembella, AD; Shumway, SE; Larocque, R. Sequestering and putative biotransformation of paralytic shellfish toxins by the sea scallop Placopecten magellanicus -seasonal and spatial scales in natural populations. J Exp Mar Biol Ecol 1994, 180, 1–22. [Google Scholar]
- Cembella, AD; Shumway, SE; Lewis, NI. Anatomical distribution and spatiotemporal variation in paralytic shellfish toxin composition in two bivalve species from the Gulf of Maine. J Shellfish Res 1993, 12, 389–403. [Google Scholar]
- Kellmann, R; Mihali, TK; Jeon, YJ; Pickford, R; Pomati, F; Neilan, BA. Biosynthetic intermediate analysis and functional homology reveal a saxitoxin gene cluster in cyanobacteria. Appl Environ Microbiol 2008, 74, 4044–4053. [Google Scholar]
- Neuwald, AF; Landsman, D. GCN5-related histone N-acetyltransferases belong to a diverse superfamily that includes the yeast SPT10 protein. Trends Biochem Sci 1997, 22, 154–155. [Google Scholar]
- Hederstedt, L; Rutberg, L. Succinate dehydrogenase--a comparative review. Microbiol Rev 1981, 45, 542–555. [Google Scholar]
- Mahmood, NA; Carmichael, WW. The pharmacology of anatoxin-a(s), a neurotoxin produced by the freshwater cyanobacterium Anabaena flos-aquae NRC 525–17. Toxicon 1986, 24, 425–434. [Google Scholar]
- Sako, Y; Yoshida, T; Uchida, A; Arakawa, O; Noguchi, T; Ishida, Y. Purification and characterization of a sulfotransferase specific to N-21 of saxitoxin and gonyautoxin 2+3 from the toxic dinoflagellate Gymnodinium catenatum (Dinophyceae). J Phycol 2001, 37, 1044–1051. [Google Scholar]
- Yoshida, T; Sako, Y; Uchida, A; Kakutani, T; Arakawa, O; Noguchi, T; Ishida, Y. Purification and characterization of sulfotransferase specific to O-22 of 11-hydroxy saxitoxin from the toxic dinoflagellate Gymnodinium catenatum (dinophyceae). Fish Sci 2002, 68, 634–642. [Google Scholar]
- Schwedock, JS; Long, SR. Rhizobium meliloti genes involved in sulfate activation: the two copies of nodPQ and a new locus, sua. Genetics 1992, 132, 899–909. [Google Scholar]
- Akoh, CC; Lee, GC; Liaw, YC; Huang, TH; Shaw, JF. GDSL family of serine esterases/lipases. Prog Lipid Res 2004, 43, 534–552. [Google Scholar]
- Pomati, F; Rossetti, C; Manarolla, G; Burns, BP; Neilan, BA. Interactions between intracellular Na+ levels and saxitoxin production in Cylindrospermopsis raciborskii T3. Microbiology 2004, 150, 455–461. [Google Scholar]
- Brown, MH; Paulsen, IT; Skurray, RA. The multidrug efflux protein NorM is a prototype of a new family of transporters. Mol Microbiol 1999, 31, 394–395. [Google Scholar]
- Anderson, DM; Kulis, DM; Sullivan, JJ; Hall, S; Lee, C. Dynamics and physiology of saxitoxin production by the dinoflagellates Alexandrium spp. Mar Biol 1990, 104, 511–524. [Google Scholar]
- Dias, E; Pereira, P; Franca, S. Production of paralytic shellfish toxins by Aphanizomenon sp. LMECYA 31 (cyanobacteria). J Phycol 2002, 38, 705–712. [Google Scholar]
- Gedaria, AI; Luckas, B; Reinhardt, K; Azanza, RV. Growth response and toxin concentration of cultured Pyrodinium bahamense var. compressum to varying salinity and temperature conditions. Toxicon 2007, 50, 518–529. [Google Scholar]
- Steed, PM; Wanner, BL. Use of the rep technique for allele replacement to construct mutants with deletions of the pstSCAB-phoU operon: evidence of a new role for the PhoU protein in the phosphate regulon. J Bacteriol 1993, 175, 6797–6809. [Google Scholar]
- Ehria, S; Ohmori, M. NrrA, a nitrogen-responsive response regulator facilitates heterocyst development in the cyanobacterium Anabaena sp. strain PCC 7120. Mol Microbiol 2006, 59, 1692–1703. [Google Scholar]
- Forst, S; Delgado, J; Inouye, M. Phosphorylation of OmpR by the Osmosensor EnvZ Modulates Expression of the ompF and ompC Genes in Escherichia coli. Proc Natl Acad Sci USA 1989, 86, 6052–6056. [Google Scholar]
- Mihali, TK; Kellmann, R; Neilan, BA. Characterisation of the paralytic shellfish toxin biosynthesis gene clusters in Anabaena circinalis AWQC131C and Aphanizomenon sp. NH-5. BMC Biochem 2009, 10, 8. [Google Scholar]
- Kellmann, R; Mihali, TK; Neilan, BA. Identification of a saxitoxin biosynthesis gene with a history of frequent horizontal gene transfers. J Mol Evol 2008, 67, 526–538. [Google Scholar]
- Bourke, ATC; Hawes, RB; Neilson, A; Stallman, ND. An outbreak of hepato-enteritis (the Palm Island mystery disease) possibly caused by algal intoxication. Toxicon 1983, 21, 45–48. [Google Scholar]
- Byth, S. Palm Island mystery disease. Med J Aust 1980, 2, 40–42. [Google Scholar]
- Saker, ML; Thomas, AD; Norton, JH. Cattle mortality attributed to the toxic cyanobacterium Cylindrospermopsis raciborskii in an outback region of North Queensland. Environ Toxicol Water Qual 1999, 14, 179–182. [Google Scholar]
- Banker, R; Carmeli, S; Hadas, O; Teltsch, B; Porat, R; Sukenik, A. Identification of cylindrospermopsin in Aphanizomenon ovalisporum (Cyanophyceae) isolated from Lake Kinneret, Israel. J Phycol 1997, 33, 613–616. [Google Scholar]
- Harada, K-I; Ohtani, I; Iwamoto, K; Suzuki, M; Watanabe, MF; Watanabe, M; Terao, K. Isolation of cylindrospermopsin from a cyanobacterium Umezakia natans and its screening method. Toxicon 1994, 32, 73–84. [Google Scholar]
- Li, R; Carmichael, WW; Brittain, S; Eaglesham, GK; Shaw, GR; Liu, Y; Watanabe, MM. First report of the cyanotoxins cylindrospermopsin and deoxycylindrospermopsin from Raphidiopsis curvata (Cyanobacteria). J Phycol 2001, 37, 1121–1126. [Google Scholar]
- Li, R; Carmichael, WW; Brittain, S; Eaglesham, GK; Shaw, GR; Mahakhant, A; Noparatnaraporn, N; Yongmanitchai, W; Kaya, K; Watanabe, MM. Isolation and identification of the cyanotoxin cylindrospermopsin and deoxy-cylindrospermopsin from a Thailand strain of Cylindrospermopsis raciborskii (Cyanobacteria). Toxicon 2001, 39, 973–980. [Google Scholar]
- Preussel, K; Stuken, A; Wiedner, C; Chorus, I; Fastner, J. First report on cylindrospermopsin producing Aphanizomenon flos-aquae (Cyanobacteria) isolated from two German lakes. Toxicon 2006, 47, 156–162. [Google Scholar]
- Schembri, MA; Neilan, BA; Saint, CP. Identification of genes implicated in toxin production in the cyanobacterium Cylindrospermopsis raciborskii. Environ Toxicol 2001, 16, 413–421. [Google Scholar]
- Spoof, L; Berg, KA; Rapala, J; Lahti, K; Lepisto, L; Metcalf, JS; Codd, GA; Meriluoto, J. First observation of cylindrospermopsin in Anabaena lapponica isolated from the boreal environment (Finland). Environ Toxicol 2006, 21, 552–560. [Google Scholar]
- Seifert, M; McGregor, G; Eaglesham, G; Wickramasinghe, W; Shaw, G. First evidence for the production of cylindrospermopsin and deoxy-cylindrospermopsin by the freshwater benthic cyanobacterium, Lyngbya wollei (Farlow ex Gomont) Speziale and Dyck. Harmful Algae 2007, 6, 73–80. [Google Scholar]
- Neilan, BA; Saker, ML; Fastner, J; Torokne, A; Burns, BP. Phylogeography of the invasive cyanobacterium Cylindrospermopsis raciborskii. Mol Ecol 2003, 12, 133–140. [Google Scholar]
- Ohtani, I; Moore, RE; Runnegar, MTC. Cylindrospermopsin: A potent hepatotoxin from the blue-green alga Cylindrospermopsis raciborskii. J Am Chem Soc 1992, 114, 7941–7942. [Google Scholar]
- Banker, R; Carmeli, S; Werman, M; Teltsch, B; Porat, R; Sukenik, A. Uracil Moiety is Required for Toxicity of the Cyanobacterial Hepatotoxin Cylindrospermopsin. J Toxicol Environ Health Part A 2001, 62, 281–288. [Google Scholar]
- Runnegar, MT; Kong, SM; Zhong, YZ; Lu, SC. Inhibition of reduced glutathione synthesis by cyanobacterial alkaloid cylindrospermopsin in cultured rat hepatocytes. Biochem Pharmacol 1995, 49, 219–225. [Google Scholar]
- Runnegar, MT; Kong, SM; Zhong, YZ; Ge, JL; Lu, SC. The role of glutathione in the toxicity of a novel cyanobacterial alkaloid cylindrospermopsin in cultured rat hepatocytes. Biochem Biophys Res Commun 1994, 201, 235–241. [Google Scholar]
- Runnegar, MT; Xie, C; Snider, BB; Wallace, GA; Weinreb, SM; Kuhlenkamp, J. In vitro Hepatotoxicity of the Cyanobacterial Alkaloid Cylindrospermopsin and Related Synthetic Analogues. Toxicol Sci 2002, 67, 81–87. [Google Scholar]
- Kiss, T; Vehovszky, A; Hiripi, L; Kovacs, A; Voros, L. Membrane effects of toxins isolated from a cyanobacterium, Cylindrospermopsis raciborskii, on identified molluscan neurones. Comp Biochem Physiol C: Toxicol Pharmacol 2002, 131, 167–176. [Google Scholar]
- Humpage, AR; Fenech, M; Thomas, P; Falconer, IR. Micronucleus induction and chromosome loss in transformed human white cells indicate clastogenic and aneugenic action of the cyanobacterial toxin, cylindrospermopsin. Mutat Res 2000, 472, 155–161. [Google Scholar]
- Froscio, SM; Humpage, AR; Burcham, PC; Falconer, IR. Cylindrospermopsin-induced protein synthesis inhibition and its dissociation from acute toxicity in mouse hepatocytes. Environ Toxicol Water Qual 2003, 18, 243–251. [Google Scholar]
- Norris, RL; Eaglesham, GK; Pierens, P; Shaw, GR; Smith, MJ; Chiswell, RK; Seawright, AA; Moore, MR. Deoxycylindrospermopsin, an analog of cylindrospermopsin from Cylindrospermopsis raciborskii. Environ Toxicol Water Qual 1999, 14, 163–165. [Google Scholar]
- Terao, K; Ohmori, S; Igarashi, K; Ohtani, I; Watanabe, MF; Harada, KI; Ito, E; Watanabe, M. Electron microscopic studies on experimental poisoning in mice induced by cylindrospermopsin isolated from blue-green alga Umezakia natans. Toxicon 1994, 32, 833–843. [Google Scholar]
- Wiegand, C; Pflugmacher, S. Ecotoxicological effects of selected cyanobacterial secondary metabolites: a short review. Toxicol Appl Pharmacol 2005, 203, 201–218. [Google Scholar]
- Mihali, TK; Kellmann, R; Muenchhoff, J; Barrow, KD; Neilan, BA. Characterization of the gene cluster responsible for cylindrospermopsin biosynthesis. Appl Environ Microbiol 2008, 74, 716–722. [Google Scholar]
- Burgoyne, DL; Hemscheidt, TK; Moore, RE; Runnegar, MT. Biosynthesis of cylindrospermopsin. J Org Chem 2000, 65, 152–156. [Google Scholar]
- Kellmann, R; Mills, T; Neilan, BA. Functional modeling and phylogenetic distribution of putative cylindrospermopsin biosynthesis enzymes. J Mol Evol 2006, 62, 267–280. [Google Scholar]
- Shalev-Alon, G; Sukenik, A; Livnah, O; Schwarz, R; Kaplan, A. A novel gene encoding amidinotransferase in the cylindrospermopsin producing cyanobacterium Aphanizomenon ovalisporum. FEMS Microbiol Lett 2002, 209, 87–91. [Google Scholar]
- Baldwin, JE; Thomas, RC; Kruse, LI; Silberman, L. Rules for ring closure: ring formation by conjugate addition of oxygen nucleophiles. J Org Chem 1977, 42, 3846–3852. [Google Scholar]
- Norris, RL; Seawright, AA; Shaw, GR; Senogles, P; Eaglesham, GK; Smith, MJ; Chiswell, RK; Moore, MR. Hepatic xenobiotic metabolism of cylindrospermopsin in vivo in the mouse. Toxicon 2002, 40, 471–476. [Google Scholar]
- Saker, ML; Neilan, BA; Griffiths, DJ. Two morphological forms of cylindrospermopsis raciborskii (cyanobacteria) isolated from Solomon dam, palm island, Queensland. J Phycol 1999, 35, 599–606. [Google Scholar]
- Hansel, A; Axelsson, R; Lindberg, P; Troshina, OY; Wunschiers, R; Lindblad, P. Cloning and characterisation of a hyp gene cluster in the filamentous cyanobacterium Nostoc sp. strain PCC 73102. FEMS Microbiol Lett 2001, 201, 59–64. [Google Scholar]
- Tamagnini, P; Axelsson, R; Lindberg, P; Oxelfelt, F; Wünschiers, R; Lindblad, P. Hydrogenases and Hydrogen Metabolism of Cyanobacteria. Microbiol Mol Biol Rev 2002, 66, 1–20. [Google Scholar]





© 2010 by the authors; licensee Molecular Diversity Preservation International, Basel, Switzerland This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Pearson, L.; Mihali, T.; Moffitt, M.; Kellmann, R.; Neilan, B. On the Chemistry, Toxicology and Genetics of the Cyanobacterial Toxins, Microcystin, Nodularin, Saxitoxin and Cylindrospermopsin. Mar. Drugs 2010, 8, 1650-1680. https://doi.org/10.3390/md8051650
Pearson L, Mihali T, Moffitt M, Kellmann R, Neilan B. On the Chemistry, Toxicology and Genetics of the Cyanobacterial Toxins, Microcystin, Nodularin, Saxitoxin and Cylindrospermopsin. Marine Drugs. 2010; 8(5):1650-1680. https://doi.org/10.3390/md8051650
Chicago/Turabian StylePearson, Leanne, Troco Mihali, Michelle Moffitt, Ralf Kellmann, and Brett Neilan. 2010. "On the Chemistry, Toxicology and Genetics of the Cyanobacterial Toxins, Microcystin, Nodularin, Saxitoxin and Cylindrospermopsin" Marine Drugs 8, no. 5: 1650-1680. https://doi.org/10.3390/md8051650
APA StylePearson, L., Mihali, T., Moffitt, M., Kellmann, R., & Neilan, B. (2010). On the Chemistry, Toxicology and Genetics of the Cyanobacterial Toxins, Microcystin, Nodularin, Saxitoxin and Cylindrospermopsin. Marine Drugs, 8(5), 1650-1680. https://doi.org/10.3390/md8051650



